Project
Inflammatory response time course, HUVEC (Wada, 2009)
Navigation
Downloads
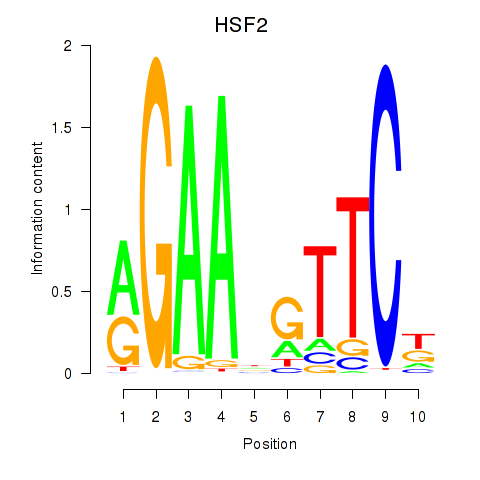
Results for HSF2
Z-value: 0.37
Transcription factors associated with HSF2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HSF2
|
ENSG00000025156.13 | HSF2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HSF2 | hg38_v1_chr6_+_122399536_122399568, hg38_v1_chr6_+_122399621_122399692 | 0.06 | 7.7e-01 | Click! |
Activity profile of HSF2 motif
Sorted Z-values of HSF2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HSF2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr8_-_80171106 | 0.58 |
ENST00000519303.6
|
TPD52
|
tumor protein D52 |
| chr4_-_185956652 | 0.47 |
ENST00000355634.9
|
SORBS2
|
sorbin and SH3 domain containing 2 |
| chr8_-_80171496 | 0.41 |
ENST00000379096.9
ENST00000518937.6 |
TPD52
|
tumor protein D52 |
| chr4_-_185956348 | 0.40 |
ENST00000431902.5
ENST00000284776.11 ENST00000415274.5 |
SORBS2
|
sorbin and SH3 domain containing 2 |
| chr6_-_142946312 | 0.35 |
ENST00000367604.6
|
HIVEP2
|
HIVEP zinc finger 2 |
| chrX_-_19965142 | 0.30 |
ENST00000340625.3
|
BCLAF3
|
BCLAF1 and THRAP3 family member 3 |
| chrX_+_136148440 | 0.28 |
ENST00000627383.2
ENST00000630084.2 |
FHL1
|
four and a half LIM domains 1 |
| chr11_+_46295126 | 0.27 |
ENST00000534787.1
|
CREB3L1
|
cAMP responsive element binding protein 3 like 1 |
| chrX_+_136197020 | 0.27 |
ENST00000370676.7
|
FHL1
|
four and a half LIM domains 1 |
| chr9_-_13279407 | 0.26 |
ENST00000546205.5
|
MPDZ
|
multiple PDZ domain crumbs cell polarity complex component |
| chr2_-_187554473 | 0.26 |
ENST00000453013.5
ENST00000417013.5 |
TFPI
|
tissue factor pathway inhibitor |
| chrX_+_136196750 | 0.22 |
ENST00000539015.5
|
FHL1
|
four and a half LIM domains 1 |
| chrX_+_136197039 | 0.21 |
ENST00000370683.6
|
FHL1
|
four and a half LIM domains 1 |
| chr11_-_72080472 | 0.21 |
ENST00000537217.5
ENST00000366394.7 ENST00000358965.10 ENST00000546131.1 ENST00000393695.8 ENST00000543937.5 ENST00000368959.9 ENST00000541641.5 |
NUMA1
|
nuclear mitotic apparatus protein 1 |
| chr11_-_72080680 | 0.20 |
ENST00000613205.4
|
NUMA1
|
nuclear mitotic apparatus protein 1 |
| chr7_+_130486324 | 0.20 |
ENST00000427521.6
ENST00000378576.9 |
MEST
|
mesoderm specific transcript |
| chr7_+_130486171 | 0.19 |
ENST00000341441.9
ENST00000416162.7 |
MEST
|
mesoderm specific transcript |
| chr2_+_218959635 | 0.19 |
ENST00000302625.6
|
CDK5R2
|
cyclin dependent kinase 5 regulatory subunit 2 |
| chr3_-_15798184 | 0.19 |
ENST00000624145.3
|
ANKRD28
|
ankyrin repeat domain 28 |
| chr7_+_76302665 | 0.18 |
ENST00000248553.7
ENST00000674638.1 ENST00000674547.1 ENST00000675226.1 ENST00000675538.1 ENST00000676231.1 ENST00000675134.1 ENST00000675906.1 ENST00000674650.1 |
HSPB1
|
heat shock protein family B (small) member 1 |
| chrX_+_106920393 | 0.18 |
ENST00000336803.2
|
CLDN2
|
claudin 2 |
| chr5_+_57174016 | 0.17 |
ENST00000514387.6
ENST00000506184.7 |
GPBP1
|
GC-rich promoter binding protein 1 |
| chr3_-_15797930 | 0.17 |
ENST00000683139.1
|
ANKRD28
|
ankyrin repeat domain 28 |
| chr9_+_125748175 | 0.16 |
ENST00000491787.7
ENST00000447726.6 |
PBX3
|
PBX homeobox 3 |
| chr2_-_187554351 | 0.16 |
ENST00000437725.5
ENST00000409676.5 ENST00000233156.9 ENST00000339091.8 ENST00000420747.1 |
TFPI
|
tissue factor pathway inhibitor |
| chr15_-_70096260 | 0.15 |
ENST00000558201.5
|
TLE3
|
TLE family member 3, transcriptional corepressor |
| chr6_-_53148822 | 0.15 |
ENST00000259803.8
|
GCM1
|
glial cells missing transcription factor 1 |
| chr6_+_32741382 | 0.15 |
ENST00000374940.4
|
HLA-DQA2
|
major histocompatibility complex, class II, DQ alpha 2 |
| chr16_+_68085861 | 0.15 |
ENST00000570212.5
ENST00000562926.5 |
NFATC3
|
nuclear factor of activated T cells 3 |
| chr2_+_74421721 | 0.14 |
ENST00000409791.5
ENST00000348227.4 |
WDR54
|
WD repeat domain 54 |
| chr5_+_57173948 | 0.14 |
ENST00000424459.7
|
GPBP1
|
GC-rich promoter binding protein 1 |
| chr5_+_80035341 | 0.13 |
ENST00000350881.6
|
THBS4
|
thrombospondin 4 |
| chr4_-_22515932 | 0.13 |
ENST00000502482.1
|
ADGRA3
|
adhesion G protein-coupled receptor A3 |
| chr18_-_812230 | 0.13 |
ENST00000314574.5
|
YES1
|
YES proto-oncogene 1, Src family tyrosine kinase |
| chr12_-_14567714 | 0.13 |
ENST00000240617.10
|
PLBD1
|
phospholipase B domain containing 1 |
| chr11_+_86302211 | 0.12 |
ENST00000533986.5
ENST00000278483.8 |
HIKESHI
|
heat shock protein nuclear import factor hikeshi |
| chr20_+_33168148 | 0.12 |
ENST00000354932.6
|
BPIFA2
|
BPI fold containing family A member 2 |
| chr2_+_28392802 | 0.12 |
ENST00000379619.5
ENST00000264716.9 |
FOSL2
|
FOS like 2, AP-1 transcription factor subunit |
| chr7_+_31529675 | 0.12 |
ENST00000451887.6
ENST00000454513.5 |
ITPRID1
|
ITPR interacting domain containing 1 |
| chr20_+_4686320 | 0.12 |
ENST00000430350.2
|
PRNP
|
prion protein |
| chr15_-_70096604 | 0.12 |
ENST00000559048.5
ENST00000560939.5 ENST00000440567.7 ENST00000557907.5 ENST00000558379.5 ENST00000559929.5 |
TLE3
|
TLE family member 3, transcriptional corepressor |
| chr12_+_69585434 | 0.12 |
ENST00000299300.11
ENST00000544368.6 |
CCT2
|
chaperonin containing TCP1 subunit 2 |
| chr4_-_163473732 | 0.12 |
ENST00000280605.5
|
TKTL2
|
transketolase like 2 |
| chr5_-_160312756 | 0.12 |
ENST00000644313.1
|
CCNJL
|
cyclin J like |
| chr7_+_157336961 | 0.11 |
ENST00000429029.6
|
DNAJB6
|
DnaJ heat shock protein family (Hsp40) member B6 |
| chr9_+_125747345 | 0.11 |
ENST00000342287.9
ENST00000373489.10 ENST00000373487.8 |
PBX3
|
PBX homeobox 3 |
| chr17_-_78577383 | 0.11 |
ENST00000389840.7
|
DNAH17
|
dynein axonemal heavy chain 17 |
| chr5_-_160312524 | 0.11 |
ENST00000520748.1
ENST00000257536.13 ENST00000393977.7 |
CCNJL
|
cyclin J like |
| chr4_-_151015263 | 0.11 |
ENST00000510413.5
ENST00000507224.5 ENST00000651943.2 |
LRBA
|
LPS responsive beige-like anchor protein |
| chr1_-_228425350 | 0.10 |
ENST00000366696.2
|
H3-4
|
H3.4 histone |
| chr9_+_89311187 | 0.10 |
ENST00000314355.7
|
CKS2
|
CDC28 protein kinase regulatory subunit 2 |
| chr3_-_184017863 | 0.10 |
ENST00000427120.6
ENST00000334444.11 ENST00000392579.6 ENST00000382494.6 ENST00000265586.10 ENST00000446941.2 |
ABCC5
|
ATP binding cassette subfamily C member 5 |
| chr4_+_182448995 | 0.10 |
ENST00000510504.1
|
TENM3
|
teneurin transmembrane protein 3 |
| chr12_+_106684696 | 0.10 |
ENST00000229387.6
|
RFX4
|
regulatory factor X4 |
| chr11_-_83286377 | 0.10 |
ENST00000455220.6
ENST00000529689.5 |
CCDC90B
|
coiled-coil domain containing 90B |
| chr2_-_96740034 | 0.10 |
ENST00000264963.9
ENST00000377079.8 |
LMAN2L
|
lectin, mannose binding 2 like |
| chr11_+_12674397 | 0.10 |
ENST00000527636.7
|
TEAD1
|
TEA domain transcription factor 1 |
| chr2_-_197499857 | 0.09 |
ENST00000428204.6
ENST00000678170.1 ENST00000676933.1 ENST00000678621.1 |
HSPD1
|
heat shock protein family D (Hsp60) member 1 |
| chr7_+_23598144 | 0.09 |
ENST00000410069.1
|
CCDC126
|
coiled-coil domain containing 126 |
| chr12_+_69585666 | 0.09 |
ENST00000543146.2
|
CCT2
|
chaperonin containing TCP1 subunit 2 |
| chr2_-_177263522 | 0.08 |
ENST00000448782.5
ENST00000446151.6 |
NFE2L2
|
nuclear factor, erythroid 2 like 2 |
| chr6_-_32154326 | 0.08 |
ENST00000475826.1
ENST00000485392.5 ENST00000494332.5 ENST00000498575.1 ENST00000428778.5 |
ENSG00000284954.1
ENSG00000285085.1
|
novel transcript novel protein |
| chr15_-_43493076 | 0.08 |
ENST00000413546.1
|
TP53BP1
|
tumor protein p53 binding protein 1 |
| chr2_-_85354500 | 0.08 |
ENST00000449375.1
ENST00000409984.2 ENST00000295802.9 |
RETSAT
|
retinol saturase |
| chr2_-_197499826 | 0.08 |
ENST00000439605.2
ENST00000388968.8 ENST00000418022.2 |
HSPD1
|
heat shock protein family D (Hsp60) member 1 |
| chr17_-_58417547 | 0.08 |
ENST00000577716.5
|
RNF43
|
ring finger protein 43 |
| chr13_+_42138042 | 0.08 |
ENST00000628433.2
ENST00000536612.3 |
DGKH
|
diacylglycerol kinase eta |
| chr6_+_31815532 | 0.08 |
ENST00000375651.7
ENST00000608703.1 |
HSPA1A
|
heat shock protein family A (Hsp70) member 1A |
| chr8_-_119592954 | 0.08 |
ENST00000522167.5
|
ENPP2
|
ectonucleotide pyrophosphatase/phosphodiesterase 2 |
| chr12_+_2794961 | 0.08 |
ENST00000001008.6
|
FKBP4
|
FKBP prolyl isomerase 4 |
| chr18_+_3451647 | 0.08 |
ENST00000345133.9
ENST00000330513.10 ENST00000549546.5 |
TGIF1
|
TGFB induced factor homeobox 1 |
| chr6_+_44247866 | 0.08 |
ENST00000371554.2
|
HSP90AB1
|
heat shock protein 90 alpha family class B member 1 |
| chr1_-_119648165 | 0.07 |
ENST00000421812.3
|
ZNF697
|
zinc finger protein 697 |
| chr11_-_414948 | 0.07 |
ENST00000530494.1
ENST00000528209.5 ENST00000528058.1 ENST00000431843.7 |
SIGIRR
|
single Ig and TIR domain containing |
| chr3_-_112641292 | 0.07 |
ENST00000439685.6
|
CCDC80
|
coiled-coil domain containing 80 |
| chr7_-_36724543 | 0.07 |
ENST00000612871.4
|
AOAH
|
acyloxyacyl hydrolase |
| chr6_-_123636979 | 0.07 |
ENST00000662930.1
|
TRDN
|
triadin |
| chr17_-_58417521 | 0.07 |
ENST00000584437.5
ENST00000407977.7 |
RNF43
|
ring finger protein 43 |
| chr9_-_4666495 | 0.07 |
ENST00000475086.5
|
SPATA6L
|
spermatogenesis associated 6 like |
| chr6_-_32371912 | 0.07 |
ENST00000612031.4
|
TSBP1
|
testis expressed basic protein 1 |
| chr21_+_34073569 | 0.07 |
ENST00000399312.3
ENST00000381151.5 ENST00000362077.4 |
MRPS6
SLC5A3
ENSG00000272657.1
|
mitochondrial ribosomal protein S6 solute carrier family 5 member 3 novel transcript |
| chr7_+_142670734 | 0.07 |
ENST00000390398.3
|
TRBV25-1
|
T cell receptor beta variable 25-1 |
| chr7_+_157336988 | 0.07 |
ENST00000262177.9
ENST00000417758.5 ENST00000443280.5 |
DNAJB6
|
DnaJ heat shock protein family (Hsp40) member B6 |
| chr1_-_168729187 | 0.07 |
ENST00000367817.4
|
DPT
|
dermatopontin |
| chr17_-_76141240 | 0.07 |
ENST00000322957.7
|
FOXJ1
|
forkhead box J1 |
| chr17_-_41505597 | 0.07 |
ENST00000336861.7
ENST00000246635.8 ENST00000587544.5 ENST00000587435.1 |
KRT13
|
keratin 13 |
| chr6_-_32371897 | 0.07 |
ENST00000442822.6
ENST00000375015.8 ENST00000533191.5 |
TSBP1
|
testis expressed basic protein 1 |
| chr6_-_32371872 | 0.07 |
ENST00000527965.5
ENST00000532023.5 ENST00000447241.6 ENST00000534588.1 |
TSBP1
|
testis expressed basic protein 1 |
| chr3_+_14947568 | 0.07 |
ENST00000413118.5
ENST00000425241.5 |
NR2C2
|
nuclear receptor subfamily 2 group C member 2 |
| chr3_+_14947680 | 0.06 |
ENST00000435454.5
ENST00000323373.10 |
NR2C2
|
nuclear receptor subfamily 2 group C member 2 |
| chr9_+_5629025 | 0.06 |
ENST00000251879.10
ENST00000414202.7 ENST00000418622.7 |
RIC1
|
RIC1 homolog, RAB6A GEF complex partner 1 |
| chr11_+_66509079 | 0.06 |
ENST00000419755.3
|
ENSG00000256349.1
|
novel protein |
| chr7_-_36724457 | 0.06 |
ENST00000617537.5
ENST00000435386.1 |
AOAH
|
acyloxyacyl hydrolase |
| chr10_-_99913971 | 0.06 |
ENST00000543621.6
|
DNMBP
|
dynamin binding protein |
| chr4_-_152679984 | 0.06 |
ENST00000304385.8
ENST00000504064.1 |
TMEM154
|
transmembrane protein 154 |
| chr19_+_14031746 | 0.06 |
ENST00000263379.4
|
IL27RA
|
interleukin 27 receptor subunit alpha |
| chr13_-_31162341 | 0.06 |
ENST00000445273.6
ENST00000630972.2 |
HSPH1
|
heat shock protein family H (Hsp110) member 1 |
| chr7_-_36724380 | 0.06 |
ENST00000617267.4
|
AOAH
|
acyloxyacyl hydrolase |
| chr2_+_197500398 | 0.06 |
ENST00000604458.1
|
HSPE1-MOB4
|
HSPE1-MOB4 readthrough |
| chr20_+_63235883 | 0.06 |
ENST00000342412.10
|
BIRC7
|
baculoviral IAP repeat containing 7 |
| chr21_+_43728851 | 0.06 |
ENST00000327574.4
|
PDXK
|
pyridoxal kinase |
| chr20_+_63235899 | 0.06 |
ENST00000217169.8
|
BIRC7
|
baculoviral IAP repeat containing 7 |
| chr19_-_48746797 | 0.06 |
ENST00000602105.1
ENST00000332955.7 |
IZUMO1
|
izumo sperm-egg fusion 1 |
| chr1_-_197775435 | 0.06 |
ENST00000620048.6
|
DENND1B
|
DENN domain containing 1B |
| chr18_+_3451585 | 0.06 |
ENST00000551541.5
|
TGIF1
|
TGFB induced factor homeobox 1 |
| chr15_+_96325935 | 0.05 |
ENST00000421109.6
|
NR2F2
|
nuclear receptor subfamily 2 group F member 2 |
| chr21_-_29073565 | 0.05 |
ENST00000431234.1
ENST00000286788.9 ENST00000540844.5 |
CCT8
|
chaperonin containing TCP1 subunit 8 |
| chr11_-_124310837 | 0.05 |
ENST00000357821.2
|
OR8D1
|
olfactory receptor family 8 subfamily D member 1 |
| chr4_+_122239965 | 0.05 |
ENST00000446180.5
|
KIAA1109
|
KIAA1109 |
| chr2_-_10837977 | 0.05 |
ENST00000404824.2
|
PDIA6
|
protein disulfide isomerase family A member 6 |
| chr19_-_14518383 | 0.05 |
ENST00000254322.3
ENST00000595139.2 |
DNAJB1
|
DnaJ heat shock protein family (Hsp40) member B1 |
| chr11_-_123062022 | 0.05 |
ENST00000532182.5
ENST00000524590.5 ENST00000528292.5 ENST00000533540.5 ENST00000534624.6 ENST00000525463.5 |
HSPA8
|
heat shock protein family A (Hsp70) member 8 |
| chr6_+_152697888 | 0.05 |
ENST00000367245.5
ENST00000529453.1 |
MYCT1
|
MYC target 1 |
| chr14_+_88594430 | 0.05 |
ENST00000406216.7
ENST00000557737.1 |
ZC3H14
|
zinc finger CCCH-type containing 14 |
| chr3_-_36739791 | 0.05 |
ENST00000416516.2
|
DCLK3
|
doublecortin like kinase 3 |
| chr6_+_42564060 | 0.05 |
ENST00000372903.6
|
UBR2
|
ubiquitin protein ligase E3 component n-recognin 2 |
| chr1_+_158831323 | 0.05 |
ENST00000368141.5
|
MNDA
|
myeloid cell nuclear differentiation antigen |
| chr9_-_134917872 | 0.05 |
ENST00000616356.4
ENST00000371806.4 |
FCN1
|
ficolin 1 |
| chr14_+_88594406 | 0.05 |
ENST00000555900.5
|
ZC3H14
|
zinc finger CCCH-type containing 14 |
| chr1_+_18631513 | 0.04 |
ENST00000400661.3
|
PAX7
|
paired box 7 |
| chr12_-_44875647 | 0.04 |
ENST00000395487.6
|
NELL2
|
neural EGFL like 2 |
| chr3_-_112641128 | 0.04 |
ENST00000206423.8
|
CCDC80
|
coiled-coil domain containing 80 |
| chr11_+_66002754 | 0.04 |
ENST00000527348.1
|
BANF1
|
BAF nuclear assembly factor 1 |
| chr13_-_31161890 | 0.04 |
ENST00000320027.10
|
HSPH1
|
heat shock protein family H (Hsp110) member 1 |
| chr14_+_88594395 | 0.04 |
ENST00000318308.10
|
ZC3H14
|
zinc finger CCCH-type containing 14 |
| chr7_-_26200734 | 0.04 |
ENST00000354667.8
ENST00000618183.5 |
HNRNPA2B1
|
heterogeneous nuclear ribonucleoprotein A2/B1 |
| chr11_-_83285965 | 0.04 |
ENST00000529073.5
ENST00000529611.5 |
CCDC90B
|
coiled-coil domain containing 90B |
| chr22_+_39901075 | 0.04 |
ENST00000344138.9
|
GRAP2
|
GRB2 related adaptor protein 2 |
| chr3_-_47513677 | 0.04 |
ENST00000296149.9
|
ELP6
|
elongator acetyltransferase complex subunit 6 |
| chr14_+_103334339 | 0.04 |
ENST00000558316.5
ENST00000558265.5 |
EIF5
|
eukaryotic translation initiation factor 5 |
| chr2_+_197500371 | 0.04 |
ENST00000409468.1
ENST00000233893.10 |
HSPE1
|
heat shock protein family E (Hsp10) member 1 |
| chr12_-_122526929 | 0.04 |
ENST00000331738.12
ENST00000528279.1 ENST00000344591.8 ENST00000526560.6 |
RSRC2
|
arginine and serine rich coiled-coil 2 |
| chr19_-_55180242 | 0.04 |
ENST00000592470.1
ENST00000354308.8 |
SYT5
|
synaptotagmin 5 |
| chr11_-_133956957 | 0.04 |
ENST00000533871.8
|
IGSF9B
|
immunoglobulin superfamily member 9B |
| chr7_-_42152444 | 0.04 |
ENST00000479210.1
|
GLI3
|
GLI family zinc finger 3 |
| chr18_-_21704763 | 0.03 |
ENST00000580981.5
ENST00000289119.7 |
ABHD3
|
abhydrolase domain containing 3, phospholipase |
| chr14_+_103334176 | 0.03 |
ENST00000560338.5
ENST00000560763.5 ENST00000216554.8 |
EIF5
|
eukaryotic translation initiation factor 5 |
| chr19_-_55180104 | 0.03 |
ENST00000537500.5
|
SYT5
|
synaptotagmin 5 |
| chr6_+_32153441 | 0.03 |
ENST00000414204.5
ENST00000361568.6 ENST00000395523.5 |
PPT2
|
palmitoyl-protein thioesterase 2 |
| chr6_+_106087580 | 0.03 |
ENST00000424894.1
ENST00000648754.1 |
PRDM1
|
PR/SET domain 1 |
| chr13_+_45702411 | 0.03 |
ENST00000610924.1
|
CBY2
|
chibby family member 2 |
| chr15_+_78264086 | 0.03 |
ENST00000394855.7
ENST00000489435.2 |
DNAJA4
|
DnaJ heat shock protein family (Hsp40) member A4 |
| chr7_-_42152396 | 0.03 |
ENST00000642432.1
ENST00000647255.1 ENST00000677288.1 |
GLI3
|
GLI family zinc finger 3 |
| chr11_-_111910830 | 0.03 |
ENST00000526167.5
ENST00000651650.1 |
CRYAB
|
crystallin alpha B |
| chr13_+_45702306 | 0.03 |
ENST00000533564.1
ENST00000310521.6 |
CBY2
|
chibby family member 2 |
| chr13_-_31161927 | 0.03 |
ENST00000380405.7
|
HSPH1
|
heat shock protein family H (Hsp110) member 1 |
| chr11_-_65557740 | 0.03 |
ENST00000526927.5
|
LTBP3
|
latent transforming growth factor beta binding protein 3 |
| chr8_+_86866415 | 0.03 |
ENST00000518476.6
|
CNBD1
|
cyclic nucleotide binding domain containing 1 |
| chr5_+_119356011 | 0.03 |
ENST00000504771.3
ENST00000415806.2 |
TNFAIP8
|
TNF alpha induced protein 8 |
| chr12_-_44876294 | 0.03 |
ENST00000429094.7
ENST00000551601.5 ENST00000549027.5 ENST00000452445.6 |
NELL2
|
neural EGFL like 2 |
| chr11_-_111910790 | 0.03 |
ENST00000533280.6
|
CRYAB
|
crystallin alpha B |
| chr9_-_86100123 | 0.03 |
ENST00000388711.7
ENST00000466178.1 |
GOLM1
|
golgi membrane protein 1 |
| chr3_+_39467598 | 0.03 |
ENST00000428261.5
ENST00000420739.5 ENST00000415443.5 ENST00000447324.5 ENST00000383754.7 |
MOBP
|
myelin associated oligodendrocyte basic protein |
| chr14_-_25010604 | 0.02 |
ENST00000550887.5
|
STXBP6
|
syntaxin binding protein 6 |
| chr7_-_33040893 | 0.02 |
ENST00000643244.1
ENST00000409467.6 ENST00000449201.5 |
NT5C3A
|
5'-nucleotidase, cytosolic IIIA |
| chr11_+_75562274 | 0.02 |
ENST00000532356.5
ENST00000524558.5 |
SERPINH1
|
serpin family H member 1 |
| chr11_-_111910888 | 0.02 |
ENST00000525823.1
ENST00000528961.6 |
CRYAB
|
crystallin alpha B |
| chr13_+_111115303 | 0.02 |
ENST00000646102.2
ENST00000449979.5 |
ARHGEF7
|
Rho guanine nucleotide exchange factor 7 |
| chr11_+_75562056 | 0.02 |
ENST00000533603.5
|
SERPINH1
|
serpin family H member 1 |
| chr5_-_35089620 | 0.02 |
ENST00000511486.5
ENST00000310101.9 ENST00000231423.7 ENST00000513753.5 ENST00000348262.7 |
PRLR
|
prolactin receptor |
| chr15_+_67542668 | 0.02 |
ENST00000178640.10
|
MAP2K5
|
mitogen-activated protein kinase kinase 5 |
| chr16_+_715092 | 0.02 |
ENST00000568223.7
|
METRN
|
meteorin, glial cell differentiation regulator |
| chr6_+_152697934 | 0.02 |
ENST00000532295.1
|
MYCT1
|
MYC target 1 |
| chr3_-_185938006 | 0.02 |
ENST00000342294.4
ENST00000453386.7 ENST00000382191.4 |
TRA2B
|
transformer 2 beta homolog |
| chr14_-_23155302 | 0.02 |
ENST00000529705.6
|
SLC7A8
|
solute carrier family 7 member 8 |
| chr6_-_155314444 | 0.02 |
ENST00000367166.5
|
TFB1M
|
transcription factor B1, mitochondrial |
| chr11_+_66002225 | 0.02 |
ENST00000445560.6
ENST00000530204.1 |
BANF1
|
BAF nuclear assembly factor 1 |
| chr11_-_83286328 | 0.02 |
ENST00000525503.5
|
CCDC90B
|
coiled-coil domain containing 90B |
| chr14_-_56810380 | 0.02 |
ENST00000672125.1
ENST00000673481.1 |
OTX2
|
orthodenticle homeobox 2 |
| chr13_+_23570370 | 0.02 |
ENST00000403372.6
ENST00000248484.9 |
TNFRSF19
|
TNF receptor superfamily member 19 |
| chr8_-_140921188 | 0.02 |
ENST00000517887.5
|
PTK2
|
protein tyrosine kinase 2 |
| chr6_+_146029131 | 0.02 |
ENST00000355289.8
|
GRM1
|
glutamate metabotropic receptor 1 |
| chr7_+_149239641 | 0.02 |
ENST00000335870.7
|
ZNF212
|
zinc finger protein 212 |
| chr2_-_174682854 | 0.02 |
ENST00000409415.7
ENST00000359761.7 ENST00000272746.9 |
WIPF1
|
WAS/WASL interacting protein family member 1 |
| chr18_-_46917432 | 0.02 |
ENST00000324794.11
ENST00000545673.5 ENST00000585916.6 |
PIAS2
|
protein inhibitor of activated STAT 2 |
| chr19_+_17305801 | 0.02 |
ENST00000602206.1
ENST00000252602.2 |
MRPL34
|
mitochondrial ribosomal protein L34 |
| chr11_-_123062335 | 0.02 |
ENST00000453788.6
ENST00000527387.5 |
HSPA8
|
heat shock protein family A (Hsp70) member 8 |
| chr7_-_76409682 | 0.02 |
ENST00000275560.4
|
SSC4D
|
scavenger receptor cysteine rich family member with 4 domains |
| chr4_-_48134200 | 0.02 |
ENST00000264316.9
|
TXK
|
TXK tyrosine kinase |
| chr6_+_146029059 | 0.02 |
ENST00000282753.6
|
GRM1
|
glutamate metabotropic receptor 1 |
| chr11_+_66002475 | 0.02 |
ENST00000312175.7
ENST00000533166.5 |
BANF1
|
BAF nuclear assembly factor 1 |
| chr11_+_59511368 | 0.02 |
ENST00000641278.1
|
OR4D9
|
olfactory receptor family 4 subfamily D member 9 |
| chr15_+_67543178 | 0.02 |
ENST00000395476.6
|
MAP2K5
|
mitogen-activated protein kinase kinase 5 |
| chr7_+_128830399 | 0.01 |
ENST00000325888.13
ENST00000346177.6 |
FLNC
|
filamin C |
| chr12_+_57745017 | 0.01 |
ENST00000547992.5
ENST00000552816.5 ENST00000257910.8 ENST00000547472.5 |
TSPAN31
|
tetraspanin 31 |
| chr17_-_1628341 | 0.01 |
ENST00000571650.5
|
SLC43A2
|
solute carrier family 43 member 2 |
| chr3_+_45026296 | 0.01 |
ENST00000296130.5
|
CLEC3B
|
C-type lectin domain family 3 member B |
| chr9_-_4666347 | 0.01 |
ENST00000381890.9
ENST00000682582.1 |
SPATA6L
|
spermatogenesis associated 6 like |
| chr6_+_106086316 | 0.01 |
ENST00000369091.6
ENST00000369096.9 |
PRDM1
|
PR/SET domain 1 |
| chr15_-_56245074 | 0.01 |
ENST00000674082.1
|
RFX7
|
regulatory factor X7 |
| chr8_+_133113483 | 0.01 |
ENST00000521107.1
|
TG
|
thyroglobulin |
| chr16_+_176659 | 0.01 |
ENST00000320868.9
ENST00000397797.1 |
HBA1
|
hemoglobin subunit alpha 1 |
| chr4_-_158723355 | 0.01 |
ENST00000307720.4
|
PPID
|
peptidylprolyl isomerase D |
| chr15_+_78264552 | 0.01 |
ENST00000394852.8
ENST00000343789.7 |
DNAJA4
|
DnaJ heat shock protein family (Hsp40) member A4 |
| chr6_+_32154010 | 0.01 |
ENST00000375137.6
|
PPT2
|
palmitoyl-protein thioesterase 2 |
| chr6_+_32154131 | 0.01 |
ENST00000375143.6
ENST00000324816.11 ENST00000424499.1 |
PPT2
|
palmitoyl-protein thioesterase 2 |
| chr1_+_192575765 | 0.01 |
ENST00000469578.2
ENST00000367459.8 |
RGS1
|
regulator of G protein signaling 1 |
| chr2_-_216695540 | 0.01 |
ENST00000233813.5
|
IGFBP5
|
insulin like growth factor binding protein 5 |
| chr14_-_56810448 | 0.01 |
ENST00000339475.10
ENST00000555006.5 ENST00000672264.2 ENST00000554559.5 ENST00000555804.1 |
OTX2
|
orthodenticle homeobox 2 |
| chrX_-_119606412 | 0.00 |
ENST00000304449.8
|
NKRF
|
NFKB repressing factor |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0071988 | regulation of spindle elongation(GO:0032887) regulation of mitotic spindle elongation(GO:0032888) anastral spindle assembly(GO:0055048) protein localization to spindle pole body(GO:0071988) regulation of protein localization to spindle pole body(GO:1902363) positive regulation of protein localization to spindle pole body(GO:1902365) positive regulation of mitotic spindle elongation(GO:1902846) |
| 0.1 | 0.2 | GO:0021718 | superior olivary nucleus development(GO:0021718) superior olivary nucleus maturation(GO:0021722) |
| 0.1 | 0.2 | GO:0002368 | B cell cytokine production(GO:0002368) protein import into mitochondrial intermembrane space(GO:0045041) |
| 0.0 | 0.4 | GO:0007598 | blood coagulation, extrinsic pathway(GO:0007598) |
| 0.0 | 0.1 | GO:0071603 | endothelial cell-cell adhesion(GO:0071603) |
| 0.0 | 0.3 | GO:0007386 | compartment pattern specification(GO:0007386) |
| 0.0 | 0.2 | GO:0060018 | astrocyte fate commitment(GO:0060018) |
| 0.0 | 0.2 | GO:0038018 | Wnt receptor catabolic process(GO:0038018) |
| 0.0 | 0.2 | GO:0099640 | axo-dendritic protein transport(GO:0099640) |
| 0.0 | 0.3 | GO:0070278 | extracellular matrix constituent secretion(GO:0070278) |
| 0.0 | 0.1 | GO:1990535 | neuron projection maintenance(GO:1990535) |
| 0.0 | 0.2 | GO:0090666 | scaRNA localization to Cajal body(GO:0090666) |
| 0.0 | 0.1 | GO:1901389 | regulation of transforming growth factor beta activation(GO:1901388) negative regulation of transforming growth factor beta activation(GO:1901389) |
| 0.0 | 0.2 | GO:0060715 | syncytiotrophoblast cell differentiation involved in labyrinthine layer development(GO:0060715) |
| 0.0 | 0.1 | GO:0060367 | subpallium cell proliferation in forebrain(GO:0022012) lateral ganglionic eminence cell proliferation(GO:0022018) lambdoid suture morphogenesis(GO:0060366) sagittal suture morphogenesis(GO:0060367) anterior semicircular canal development(GO:0060873) lateral semicircular canal development(GO:0060875) |
| 0.0 | 0.8 | GO:0061049 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.0 | 0.1 | GO:0046223 | mycotoxin catabolic process(GO:0043387) aflatoxin catabolic process(GO:0046223) organic heteropentacyclic compound catabolic process(GO:1901377) regulation of glutathione biosynthetic process(GO:1903786) positive regulation of glutathione biosynthetic process(GO:1903788) |
| 0.0 | 0.1 | GO:1903751 | regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:1903750) negative regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:1903751) |
| 0.0 | 0.0 | GO:0002752 | cell surface pattern recognition receptor signaling pathway(GO:0002752) |
| 0.0 | 0.1 | GO:0090158 | endoplasmic reticulum membrane organization(GO:0090158) |
| 0.0 | 0.0 | GO:0014813 | skeletal muscle satellite cell commitment(GO:0014813) |
| 0.0 | 0.0 | GO:0033082 | regulation of extrathymic T cell differentiation(GO:0033082) sebum secreting cell proliferation(GO:1990654) |
| 0.0 | 0.1 | GO:0009443 | pyridoxal 5'-phosphate salvage(GO:0009443) |
| 0.0 | 0.1 | GO:0009956 | radial pattern formation(GO:0009956) |
| 0.0 | 0.1 | GO:1904764 | negative regulation of fibril organization(GO:1902904) chaperone-mediated autophagy translocation complex disassembly(GO:1904764) |
| 0.0 | 0.1 | GO:0021914 | negative regulation of smoothened signaling pathway involved in ventral spinal cord patterning(GO:0021914) |
| 0.0 | 0.1 | GO:0031437 | regulation of mRNA cleavage(GO:0031437) regulation of mRNA endonucleolytic cleavage involved in unfolded protein response(GO:1904720) |
| 0.0 | 0.1 | GO:0045079 | negative regulation of chemokine biosynthetic process(GO:0045079) |
| 0.0 | 1.0 | GO:0043268 | positive regulation of potassium ion transport(GO:0043268) |
| 0.0 | 0.1 | GO:0015798 | myo-inositol transport(GO:0015798) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0055028 | cortical microtubule(GO:0055028) |
| 0.0 | 0.2 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 0.0 | 0.1 | GO:0071148 | TEAD-1-YAP complex(GO:0071148) |
| 0.0 | 0.1 | GO:1990913 | sperm head plasma membrane(GO:1990913) ooplasm(GO:1990917) |
| 0.0 | 0.2 | GO:0046696 | cyclin-dependent protein kinase activating kinase holoenzyme complex(GO:0019907) lipopolysaccharide receptor complex(GO:0046696) |
| 0.0 | 0.2 | GO:0002199 | zona pellucida receptor complex(GO:0002199) |
| 0.0 | 0.1 | GO:1990769 | proximal neuron projection(GO:1990769) |
| 0.0 | 0.3 | GO:0043220 | Schmidt-Lanterman incisure(GO:0043220) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0050528 | acyloxyacyl hydrolase activity(GO:0050528) |
| 0.1 | 0.2 | GO:0016534 | cyclin-dependent protein kinase 5 activator activity(GO:0016534) |
| 0.0 | 0.2 | GO:0008426 | protein kinase C inhibitor activity(GO:0008426) |
| 0.0 | 0.4 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.0 | 0.9 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 0.2 | GO:0034186 | apolipoprotein A-I binding(GO:0034186) |
| 0.0 | 0.1 | GO:0002135 | CTP binding(GO:0002135) |
| 0.0 | 0.1 | GO:0071074 | eukaryotic initiation factor eIF2 binding(GO:0071074) |
| 0.0 | 0.1 | GO:1903135 | cupric ion binding(GO:1903135) |
| 0.0 | 0.1 | GO:0004802 | transketolase activity(GO:0004802) |
| 0.0 | 0.1 | GO:0047391 | alkylglycerophosphoethanolamine phosphodiesterase activity(GO:0047391) |
| 0.0 | 0.1 | GO:0032767 | copper-dependent protein binding(GO:0032767) |
| 0.0 | 0.1 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 0.0 | 0.1 | GO:0008478 | pyridoxal kinase activity(GO:0008478) |
| 0.0 | 0.1 | GO:0030144 | alpha-1,6-mannosylglycoprotein 6-beta-N-acetylglucosaminyltransferase activity(GO:0030144) |
| 0.0 | 0.0 | GO:0052739 | phosphatidylserine 1-acylhydrolase activity(GO:0052739) |
| 0.0 | 0.3 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.0 | 0.1 | GO:0032395 | MHC class II receptor activity(GO:0032395) |
| 0.0 | 0.1 | GO:0061649 | ubiquitinated histone binding(GO:0061649) |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.0 | 1.1 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |