Project
Inflammatory response time course, HUVEC (Wada, 2009)
Navigation
Downloads
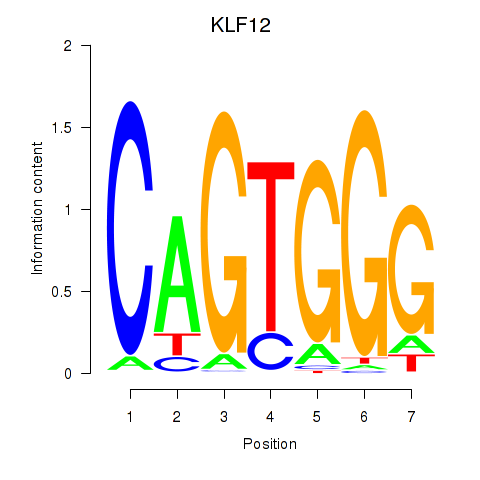
Results for KLF12
Z-value: 0.87
Transcription factors associated with KLF12
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
KLF12
|
ENSG00000118922.18 | KLF12 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| KLF12 | hg38_v1_chr13_-_74133892_74133941 | 0.36 | 7.8e-02 | Click! |
Activity profile of KLF12 motif
Sorted Z-values of KLF12 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of KLF12
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_+_138465472 | 3.87 |
ENST00000239938.5
|
EGR1
|
early growth response 1 |
| chr3_+_194136138 | 2.04 |
ENST00000232424.4
|
HES1
|
hes family bHLH transcription factor 1 |
| chr4_-_74099187 | 1.96 |
ENST00000508487.3
|
CXCL2
|
C-X-C motif chemokine ligand 2 |
| chr2_-_224402097 | 1.74 |
ENST00000409685.4
|
FAM124B
|
family with sequence similarity 124 member B |
| chr2_-_224401994 | 1.73 |
ENST00000389874.3
|
FAM124B
|
family with sequence similarity 124 member B |
| chr1_+_153778178 | 1.58 |
ENST00000532853.5
|
SLC27A3
|
solute carrier family 27 member 3 |
| chr15_-_52652031 | 1.57 |
ENST00000546305.6
|
FAM214A
|
family with sequence similarity 214 member A |
| chr4_-_74038681 | 1.41 |
ENST00000296026.4
|
CXCL3
|
C-X-C motif chemokine ligand 3 |
| chr4_+_73869385 | 1.33 |
ENST00000395761.4
|
CXCL1
|
C-X-C motif chemokine ligand 1 |
| chr2_-_219571241 | 1.33 |
ENST00000373876.5
ENST00000603926.5 ENST00000373873.8 ENST00000289656.3 |
OBSL1
|
obscurin like cytoskeletal adaptor 1 |
| chr6_+_116511626 | 1.31 |
ENST00000368599.4
|
CALHM5
|
calcium homeostasis modulator family member 5 |
| chr17_+_28744034 | 1.27 |
ENST00000444415.7
ENST00000262396.10 |
TRAF4
|
TNF receptor associated factor 4 |
| chr9_-_131276499 | 1.25 |
ENST00000372271.4
|
FAM78A
|
family with sequence similarity 78 member A |
| chr20_-_40689228 | 1.23 |
ENST00000373313.3
|
MAFB
|
MAF bZIP transcription factor B |
| chr1_+_203305510 | 1.21 |
ENST00000290551.5
|
BTG2
|
BTG anti-proliferation factor 2 |
| chr19_+_45469841 | 1.20 |
ENST00000592811.5
ENST00000586615.5 |
FOSB
|
FosB proto-oncogene, AP-1 transcription factor subunit |
| chr12_-_95790755 | 1.19 |
ENST00000343702.9
ENST00000344911.8 |
NTN4
|
netrin 4 |
| chr10_-_79445617 | 1.18 |
ENST00000372336.4
|
ZCCHC24
|
zinc finger CCHC-type containing 24 |
| chr1_+_60865259 | 1.08 |
ENST00000371191.5
|
NFIA
|
nuclear factor I A |
| chr17_-_49764123 | 1.06 |
ENST00000240364.7
ENST00000506156.1 |
FAM117A
|
family with sequence similarity 117 member A |
| chr14_-_91946989 | 1.05 |
ENST00000556154.5
|
FBLN5
|
fibulin 5 |
| chr2_+_233195433 | 1.01 |
ENST00000417661.1
|
INPP5D
|
inositol polyphosphate-5-phosphatase D |
| chr14_+_58298497 | 0.96 |
ENST00000348476.7
ENST00000355431.8 ENST00000395168.7 |
ARID4A
|
AT-rich interaction domain 4A |
| chr1_-_46132650 | 0.94 |
ENST00000372006.5
ENST00000425892.2 ENST00000420542.5 |
PIK3R3
|
phosphoinositide-3-kinase regulatory subunit 3 |
| chr3_-_149086488 | 0.92 |
ENST00000392912.6
ENST00000465259.5 ENST00000310053.10 ENST00000494055.5 |
HLTF
|
helicase like transcription factor |
| chr11_-_61361834 | 0.87 |
ENST00000544118.5
ENST00000294072.9 ENST00000545361.5 ENST00000539128.5 ENST00000546151.5 ENST00000447532.6 |
CYB561A3
|
cytochrome b561 family member A3 |
| chr11_+_7513966 | 0.86 |
ENST00000299492.9
|
PPFIBP2
|
PPFIA binding protein 2 |
| chr12_+_93572664 | 0.85 |
ENST00000551556.2
|
SOCS2
|
suppressor of cytokine signaling 2 |
| chr17_+_1762052 | 0.85 |
ENST00000254722.9
ENST00000576406.5 ENST00000571149.5 |
SERPINF1
|
serpin family F member 1 |
| chr20_+_36154630 | 0.84 |
ENST00000338074.7
ENST00000636016.2 ENST00000373945.5 |
EPB41L1
|
erythrocyte membrane protein band 4.1 like 1 |
| chr2_+_24049673 | 0.83 |
ENST00000380991.8
|
FKBP1B
|
FKBP prolyl isomerase 1B |
| chr17_-_5500997 | 0.81 |
ENST00000568641.2
|
ENSG00000286190.2
|
novel protein |
| chr14_+_63204436 | 0.80 |
ENST00000316754.8
|
RHOJ
|
ras homolog family member J |
| chr2_+_24049705 | 0.80 |
ENST00000380986.9
ENST00000452109.1 |
FKBP1B
|
FKBP prolyl isomerase 1B |
| chr16_+_29807775 | 0.78 |
ENST00000568411.5
ENST00000563012.1 ENST00000562557.5 |
MAZ
|
MYC associated zinc finger protein |
| chr19_-_4558417 | 0.75 |
ENST00000586965.1
|
SEMA6B
|
semaphorin 6B |
| chr11_-_62707581 | 0.72 |
ENST00000684475.1
ENST00000683296.1 ENST00000684067.1 ENST00000682223.1 |
BSCL2
|
BSCL2 lipid droplet biogenesis associated, seipin |
| chr8_-_6563409 | 0.71 |
ENST00000325203.9
|
ANGPT2
|
angiopoietin 2 |
| chr8_-_6563238 | 0.70 |
ENST00000629816.3
ENST00000523120.2 |
ANGPT2
|
angiopoietin 2 |
| chr16_+_29807536 | 0.70 |
ENST00000567444.5
|
MAZ
|
MYC associated zinc finger protein |
| chr8_-_6563044 | 0.69 |
ENST00000338312.10
|
ANGPT2
|
angiopoietin 2 |
| chr17_-_8630749 | 0.69 |
ENST00000379980.8
ENST00000269243.8 |
MYH10
|
myosin heavy chain 10 |
| chr16_+_29808051 | 0.69 |
ENST00000568544.5
ENST00000569978.1 |
MAZ
|
MYC associated zinc finger protein |
| chr17_-_8630713 | 0.68 |
ENST00000411957.1
ENST00000360416.8 |
MYH10
|
myosin heavy chain 10 |
| chr16_+_69105636 | 0.67 |
ENST00000569188.6
|
HAS3
|
hyaluronan synthase 3 |
| chr16_+_29808125 | 0.67 |
ENST00000568282.1
|
MAZ
|
MYC associated zinc finger protein |
| chr22_+_25111810 | 0.67 |
ENST00000637069.1
|
KIAA1671
|
KIAA1671 |
| chr14_+_73950489 | 0.66 |
ENST00000554320.1
|
COQ6
|
coenzyme Q6, monooxygenase |
| chr11_+_104036624 | 0.65 |
ENST00000302259.5
|
DDI1
|
DNA damage inducible 1 homolog 1 |
| chr17_-_78903193 | 0.65 |
ENST00000322630.3
ENST00000586713.5 |
CEP295NL
|
CEP295 N-terminal like |
| chr2_+_172860038 | 0.64 |
ENST00000538974.5
ENST00000540783.5 |
RAPGEF4
|
Rap guanine nucleotide exchange factor 4 |
| chr11_-_102530738 | 0.62 |
ENST00000260227.5
|
MMP7
|
matrix metallopeptidase 7 |
| chr17_-_40994159 | 0.62 |
ENST00000391586.3
|
KRTAP3-3
|
keratin associated protein 3-3 |
| chr6_+_11537738 | 0.61 |
ENST00000379426.2
|
TMEM170B
|
transmembrane protein 170B |
| chr11_-_65606959 | 0.60 |
ENST00000532507.5
|
MAP3K11
|
mitogen-activated protein kinase kinase kinase 11 |
| chr1_-_46132616 | 0.60 |
ENST00000423209.5
ENST00000262741.10 |
PIK3R3
|
phosphoinositide-3-kinase regulatory subunit 3 |
| chr11_-_62707413 | 0.60 |
ENST00000360796.10
ENST00000449636.6 |
BSCL2
|
BSCL2 lipid droplet biogenesis associated, seipin |
| chr2_+_148021404 | 0.59 |
ENST00000638043.2
|
MBD5
|
methyl-CpG binding domain protein 5 |
| chr1_+_92029971 | 0.59 |
ENST00000370383.5
|
EPHX4
|
epoxide hydrolase 4 |
| chr5_-_81751022 | 0.58 |
ENST00000509013.2
ENST00000505980.5 ENST00000509053.5 |
SSBP2
|
single stranded DNA binding protein 2 |
| chr16_+_2969270 | 0.57 |
ENST00000293978.12
|
PAQR4
|
progestin and adipoQ receptor family member 4 |
| chr2_+_202634960 | 0.57 |
ENST00000392238.3
|
FAM117B
|
family with sequence similarity 117 member B |
| chr16_+_2969307 | 0.57 |
ENST00000576565.1
ENST00000318782.9 |
PAQR4
|
progestin and adipoQ receptor family member 4 |
| chr2_+_188291661 | 0.56 |
ENST00000409843.5
|
GULP1
|
GULP PTB domain containing engulfment adaptor 1 |
| chr11_+_121590388 | 0.56 |
ENST00000527934.1
|
SORL1
|
sortilin related receptor 1 |
| chrX_-_45200895 | 0.55 |
ENST00000377934.4
|
DIPK2B
|
divergent protein kinase domain 2B |
| chr14_-_22819721 | 0.53 |
ENST00000554517.5
ENST00000285850.11 ENST00000397529.6 ENST00000555702.5 |
SLC7A7
|
solute carrier family 7 member 7 |
| chr5_-_81751085 | 0.53 |
ENST00000515395.5
|
SSBP2
|
single stranded DNA binding protein 2 |
| chr16_-_31508370 | 0.52 |
ENST00000430477.6
ENST00000567994.5 ENST00000570164.5 ENST00000327237.7 |
RUSF1
|
RUS family member 1 |
| chr10_+_61901678 | 0.52 |
ENST00000644638.1
ENST00000681100.1 ENST00000279873.12 |
ARID5B
|
AT-rich interaction domain 5B |
| chr12_+_12725897 | 0.50 |
ENST00000326765.10
|
APOLD1
|
apolipoprotein L domain containing 1 |
| chr8_+_96493803 | 0.50 |
ENST00000518385.5
ENST00000302190.9 |
SDC2
|
syndecan 2 |
| chr2_+_36696686 | 0.50 |
ENST00000379242.7
ENST00000389975.7 |
VIT
|
vitrin |
| chr19_+_43612083 | 0.49 |
ENST00000417606.3
|
SRRM5
|
serine/arginine repetitive matrix 5 |
| chr7_-_27185223 | 0.48 |
ENST00000517402.1
ENST00000006015.4 |
HOXA11
|
homeobox A11 |
| chr6_-_31897200 | 0.48 |
ENST00000395728.7
ENST00000375528.8 |
EHMT2
|
euchromatic histone lysine methyltransferase 2 |
| chr22_+_46620380 | 0.48 |
ENST00000406902.6
|
GRAMD4
|
GRAM domain containing 4 |
| chr11_-_119340816 | 0.47 |
ENST00000528368.3
|
C1QTNF5
|
C1q and TNF related 5 |
| chr14_+_63204859 | 0.46 |
ENST00000555125.1
|
RHOJ
|
ras homolog family member J |
| chr12_+_7061206 | 0.46 |
ENST00000423384.5
ENST00000413211.5 |
C1S
|
complement C1s |
| chrX_-_71254106 | 0.46 |
ENST00000373984.7
ENST00000314425.9 ENST00000373982.5 |
ZMYM3
|
zinc finger MYM-type containing 3 |
| chr14_+_75002903 | 0.46 |
ENST00000266126.10
|
EIF2B2
|
eukaryotic translation initiation factor 2B subunit beta |
| chr2_+_178194460 | 0.45 |
ENST00000392505.6
ENST00000359685.7 ENST00000357080.8 ENST00000190611.9 ENST00000409045.7 |
OSBPL6
|
oxysterol binding protein like 6 |
| chrX_+_54809060 | 0.45 |
ENST00000396224.1
|
MAGED2
|
MAGE family member D2 |
| chr17_+_2337480 | 0.44 |
ENST00000268989.8
ENST00000426855.6 |
SGSM2
|
small G protein signaling modulator 2 |
| chr7_-_122995700 | 0.44 |
ENST00000249284.3
|
TAS2R16
|
taste 2 receptor member 16 |
| chr5_+_75337192 | 0.43 |
ENST00000680160.1
|
HMGCR
|
3-hydroxy-3-methylglutaryl-CoA reductase |
| chr5_+_75337261 | 0.43 |
ENST00000680940.1
|
HMGCR
|
3-hydroxy-3-methylglutaryl-CoA reductase |
| chr5_+_75337211 | 0.43 |
ENST00000287936.9
ENST00000343975.9 |
HMGCR
|
3-hydroxy-3-methylglutaryl-CoA reductase |
| chr7_-_27662836 | 0.42 |
ENST00000265395.7
|
HIBADH
|
3-hydroxyisobutyrate dehydrogenase |
| chr11_-_119340544 | 0.42 |
ENST00000530681.2
|
C1QTNF5
|
C1q and TNF related 5 |
| chr5_+_75337348 | 0.42 |
ENST00000681271.1
|
HMGCR
|
3-hydroxy-3-methylglutaryl-CoA reductase |
| chr16_+_31459950 | 0.42 |
ENST00000564900.1
|
ARMC5
|
armadillo repeat containing 5 |
| chr18_-_55585773 | 0.41 |
ENST00000563824.5
ENST00000626425.2 ENST00000566514.5 ENST00000568673.5 ENST00000562847.5 ENST00000568147.5 |
TCF4
|
transcription factor 4 |
| chr2_+_218880844 | 0.41 |
ENST00000258411.8
|
WNT10A
|
Wnt family member 10A |
| chr9_+_34652167 | 0.41 |
ENST00000441545.7
ENST00000553620.5 |
IL11RA
|
interleukin 11 receptor subunit alpha |
| chr2_-_55296361 | 0.40 |
ENST00000647547.1
|
CCDC88A
|
coiled-coil domain containing 88A |
| chr2_-_127643212 | 0.40 |
ENST00000409286.5
|
LIMS2
|
LIM zinc finger domain containing 2 |
| chr21_-_43060546 | 0.40 |
ENST00000430013.1
|
CBS
|
cystathionine beta-synthase |
| chr19_+_36111151 | 0.40 |
ENST00000633214.1
ENST00000585332.3 |
OVOL3
|
ovo like zinc finger 3 |
| chr16_+_28863812 | 0.39 |
ENST00000684370.1
|
SH2B1
|
SH2B adaptor protein 1 |
| chr10_-_102120246 | 0.39 |
ENST00000425280.2
|
LDB1
|
LIM domain binding 1 |
| chr3_-_49685090 | 0.39 |
ENST00000448220.5
|
MST1
|
macrophage stimulating 1 |
| chr1_-_203175783 | 0.39 |
ENST00000621380.1
ENST00000255416.9 |
MYBPH
|
myosin binding protein H |
| chr10_-_89643870 | 0.38 |
ENST00000322191.10
ENST00000342512.3 |
PANK1
|
pantothenate kinase 1 |
| chr12_+_57089094 | 0.38 |
ENST00000342556.6
ENST00000300131.8 |
NAB2
|
NGFI-A binding protein 2 |
| chr17_+_997101 | 0.38 |
ENST00000327158.5
|
TIMM22
|
translocase of inner mitochondrial membrane 22 |
| chr20_+_35699442 | 0.38 |
ENST00000374072.5
ENST00000397416.1 ENST00000336695.4 |
ROMO1
|
reactive oxygen species modulator 1 |
| chr16_+_75222609 | 0.38 |
ENST00000495583.1
|
CTRB1
|
chymotrypsinogen B1 |
| chr11_-_1757452 | 0.37 |
ENST00000427721.3
|
ENSG00000250644.3
|
novel protein |
| chrX_-_10576901 | 0.37 |
ENST00000380779.5
|
MID1
|
midline 1 |
| chr1_+_150067668 | 0.36 |
ENST00000611412.4
ENST00000644510.2 ENST00000643611.1 |
VPS45
|
vacuolar protein sorting 45 homolog |
| chr19_+_40627033 | 0.36 |
ENST00000599225.1
ENST00000598166.2 |
LTBP4
|
latent transforming growth factor beta binding protein 4 |
| chr12_-_6663083 | 0.36 |
ENST00000467678.5
ENST00000493873.1 ENST00000412586.6 ENST00000423703.6 ENST00000444704.5 ENST00000341550.9 |
ING4
|
inhibitor of growth family member 4 |
| chr9_-_96302142 | 0.35 |
ENST00000648799.1
|
HSD17B3
|
hydroxysteroid 17-beta dehydrogenase 3 |
| chr1_-_202808406 | 0.35 |
ENST00000650569.1
ENST00000367265.9 ENST00000649770.1 |
KDM5B
|
lysine demethylase 5B |
| chr5_-_108381109 | 0.35 |
ENST00000619412.4
|
FBXL17
|
F-box and leucine rich repeat protein 17 |
| chr3_-_128493173 | 0.35 |
ENST00000498200.1
ENST00000341105.7 |
GATA2
|
GATA binding protein 2 |
| chr19_+_49877660 | 0.35 |
ENST00000535102.6
|
TBC1D17
|
TBC1 domain family member 17 |
| chr11_+_62707668 | 0.35 |
ENST00000294117.6
|
GNG3
|
G protein subunit gamma 3 |
| chr1_-_45206594 | 0.35 |
ENST00000359600.6
|
ZSWIM5
|
zinc finger SWIM-type containing 5 |
| chr15_+_40239420 | 0.35 |
ENST00000560346.5
|
PAK6
|
p21 (RAC1) activated kinase 6 |
| chr12_+_6951271 | 0.35 |
ENST00000456013.5
|
PTPN6
|
protein tyrosine phosphatase non-receptor type 6 |
| chr18_+_58864866 | 0.34 |
ENST00000588456.5
ENST00000591808.6 ENST00000589481.1 ENST00000591049.1 |
ZNF532
|
zinc finger protein 532 |
| chr1_-_153628180 | 0.34 |
ENST00000339556.8
ENST00000440685.7 |
S100A13
|
S100 calcium binding protein A13 |
| chr2_+_36696790 | 0.34 |
ENST00000497382.5
ENST00000404084.5 ENST00000379241.7 ENST00000401530.5 |
VIT
|
vitrin |
| chr16_-_740980 | 0.34 |
ENST00000251588.7
|
CIAO3
|
cytosolic iron-sulfur assembly component 3 |
| chr16_-_740934 | 0.34 |
ENST00000540986.5
|
CIAO3
|
cytosolic iron-sulfur assembly component 3 |
| chr17_-_81869934 | 0.33 |
ENST00000580685.5
|
ARHGDIA
|
Rho GDP dissociation inhibitor alpha |
| chr11_-_44950151 | 0.33 |
ENST00000533940.5
ENST00000533937.1 |
TP53I11
|
tumor protein p53 inducible protein 11 |
| chr12_+_6946468 | 0.33 |
ENST00000543115.5
ENST00000399448.5 |
PTPN6
|
protein tyrosine phosphatase non-receptor type 6 |
| chrX_+_154429092 | 0.33 |
ENST00000619046.5
|
ATP6AP1
|
ATPase H+ transporting accessory protein 1 |
| chr13_-_67230313 | 0.33 |
ENST00000377865.7
|
PCDH9
|
protocadherin 9 |
| chr19_+_49877694 | 0.33 |
ENST00000221543.10
|
TBC1D17
|
TBC1 domain family member 17 |
| chr1_+_15756659 | 0.33 |
ENST00000375771.5
|
FBLIM1
|
filamin binding LIM protein 1 |
| chr6_-_43053832 | 0.33 |
ENST00000265348.9
ENST00000674134.1 ENST00000674100.1 |
CUL7
|
cullin 7 |
| chr16_+_31459479 | 0.33 |
ENST00000268314.9
|
ARMC5
|
armadillo repeat containing 5 |
| chrX_-_130165825 | 0.33 |
ENST00000675240.1
ENST00000319908.8 ENST00000674546.1 ENST00000287295.8 |
AIFM1
|
apoptosis inducing factor mitochondria associated 1 |
| chr1_+_150067820 | 0.33 |
ENST00000419023.3
ENST00000644526.1 |
VPS45
|
vacuolar protein sorting 45 homolog |
| chr15_+_40239042 | 0.32 |
ENST00000558055.5
ENST00000455577.6 |
PAK6
|
p21 (RAC1) activated kinase 6 |
| chr15_-_72783611 | 0.32 |
ENST00000563907.5
|
ADPGK
|
ADP dependent glucokinase |
| chr1_-_202808464 | 0.32 |
ENST00000648469.1
ENST00000648338.1 ENST00000367264.7 ENST00000648473.1 ENST00000648056.1 ENST00000650368.1 |
KDM5B
|
lysine demethylase 5B |
| chr12_+_6951345 | 0.32 |
ENST00000318974.14
ENST00000541698.5 ENST00000542462.1 |
PTPN6
|
protein tyrosine phosphatase non-receptor type 6 |
| chr22_-_31346317 | 0.32 |
ENST00000266269.10
|
PATZ1
|
POZ/BTB and AT hook containing zinc finger 1 |
| chrX_-_130165873 | 0.32 |
ENST00000676229.1
|
AIFM1
|
apoptosis inducing factor mitochondria associated 1 |
| chrX_+_152831054 | 0.32 |
ENST00000370274.8
|
NSDHL
|
NAD(P) dependent steroid dehydrogenase-like |
| chr12_-_108731505 | 0.31 |
ENST00000261401.8
ENST00000552871.5 |
CORO1C
|
coronin 1C |
| chr14_+_57390544 | 0.31 |
ENST00000555166.5
ENST00000556492.6 ENST00000554703.1 |
NAA30
|
N-alpha-acetyltransferase 30, NatC catalytic subunit |
| chr15_-_72783685 | 0.31 |
ENST00000456471.3
ENST00000311669.12 |
ADPGK
|
ADP dependent glucokinase |
| chr16_+_2969548 | 0.31 |
ENST00000572687.1
|
PAQR4
|
progestin and adipoQ receptor family member 4 |
| chr9_+_113444725 | 0.31 |
ENST00000374140.6
|
RGS3
|
regulator of G protein signaling 3 |
| chr17_-_39401593 | 0.31 |
ENST00000394294.7
ENST00000264658.11 ENST00000583610.5 ENST00000647139.1 |
FBXL20
|
F-box and leucine rich repeat protein 20 |
| chr15_+_41493860 | 0.30 |
ENST00000260386.7
|
ITPKA
|
inositol-trisphosphate 3-kinase A |
| chrX_-_112840815 | 0.30 |
ENST00000304758.5
ENST00000371959.9 |
AMOT
|
angiomotin |
| chr16_-_27549887 | 0.30 |
ENST00000561623.5
ENST00000356183.9 |
GTF3C1
|
general transcription factor IIIC subunit 1 |
| chr13_-_67230377 | 0.30 |
ENST00000544246.5
ENST00000377861.4 |
PCDH9
|
protocadherin 9 |
| chr11_+_66593194 | 0.30 |
ENST00000310190.8
|
CCS
|
copper chaperone for superoxide dismutase |
| chr10_-_102120318 | 0.30 |
ENST00000673968.1
|
LDB1
|
LIM domain binding 1 |
| chr7_+_37920602 | 0.30 |
ENST00000199448.9
ENST00000423717.1 |
EPDR1
|
ependymin related 1 |
| chr14_+_104724221 | 0.30 |
ENST00000330877.7
|
ADSS1
|
adenylosuccinate synthase 1 |
| chr2_+_233636502 | 0.30 |
ENST00000373445.1
|
UGT1A10
|
UDP glucuronosyltransferase family 1 member A10 |
| chr16_-_19886133 | 0.30 |
ENST00000568214.1
ENST00000569479.5 |
GPRC5B
|
G protein-coupled receptor class C group 5 member B |
| chrX_-_130165664 | 0.30 |
ENST00000535724.6
ENST00000346424.6 ENST00000676436.1 |
AIFM1
|
apoptosis inducing factor mitochondria associated 1 |
| chr6_+_20401864 | 0.29 |
ENST00000346618.8
ENST00000613242.4 |
E2F3
|
E2F transcription factor 3 |
| chr17_+_28662183 | 0.29 |
ENST00000347486.8
ENST00000314616.11 |
SUPT6H
|
SPT6 homolog, histone chaperone and transcription elongation factor |
| chr16_+_57976435 | 0.29 |
ENST00000290871.10
ENST00000441824.4 |
TEPP
|
testis, prostate and placenta expressed |
| chr17_+_2337622 | 0.29 |
ENST00000574563.5
|
SGSM2
|
small G protein signaling modulator 2 |
| chr20_+_63235883 | 0.29 |
ENST00000342412.10
|
BIRC7
|
baculoviral IAP repeat containing 7 |
| chr14_+_51847145 | 0.29 |
ENST00000615906.4
|
GNG2
|
G protein subunit gamma 2 |
| chr9_-_125189721 | 0.28 |
ENST00000456642.1
ENST00000373547.9 ENST00000415905.5 ENST00000451402.5 |
PPP6C
|
protein phosphatase 6 catalytic subunit |
| chr2_-_113241779 | 0.28 |
ENST00000497038.6
|
PAX8
|
paired box 8 |
| chr1_+_155127866 | 0.28 |
ENST00000368406.2
ENST00000368407.8 |
EFNA1
|
ephrin A1 |
| chr9_-_96302170 | 0.28 |
ENST00000375263.8
|
HSD17B3
|
hydroxysteroid 17-beta dehydrogenase 3 |
| chr14_+_103715724 | 0.28 |
ENST00000216602.10
|
ZFYVE21
|
zinc finger FYVE-type containing 21 |
| chr14_+_103715767 | 0.27 |
ENST00000311141.7
|
ZFYVE21
|
zinc finger FYVE-type containing 21 |
| chr16_+_6483379 | 0.27 |
ENST00000552089.5
|
RBFOX1
|
RNA binding fox-1 homolog 1 |
| chr10_+_97713694 | 0.27 |
ENST00000285605.8
|
MARVELD1
|
MARVEL domain containing 1 |
| chrX_-_84188148 | 0.27 |
ENST00000262752.5
|
RPS6KA6
|
ribosomal protein S6 kinase A6 |
| chr17_+_82519694 | 0.27 |
ENST00000335255.10
|
FOXK2
|
forkhead box K2 |
| chr12_-_70754631 | 0.26 |
ENST00000440835.6
ENST00000549308.5 ENST00000550661.1 ENST00000378778.5 |
PTPRR
|
protein tyrosine phosphatase receptor type R |
| chr1_+_109249530 | 0.26 |
ENST00000271332.4
|
CELSR2
|
cadherin EGF LAG seven-pass G-type receptor 2 |
| chr19_-_55175031 | 0.26 |
ENST00000587067.1
|
SYT5
|
synaptotagmin 5 |
| chr20_-_54070520 | 0.26 |
ENST00000371435.6
ENST00000395961.7 |
BCAS1
|
brain enriched myelin associated protein 1 |
| chr17_+_44557476 | 0.26 |
ENST00000315323.5
|
FZD2
|
frizzled class receptor 2 |
| chr17_+_4715438 | 0.26 |
ENST00000571206.1
|
ARRB2
|
arrestin beta 2 |
| chr19_-_45768627 | 0.26 |
ENST00000560160.1
|
SIX5
|
SIX homeobox 5 |
| chr20_+_35699368 | 0.26 |
ENST00000374077.8
|
ROMO1
|
reactive oxygen species modulator 1 |
| chr16_-_30787169 | 0.26 |
ENST00000262525.6
|
ZNF629
|
zinc finger protein 629 |
| chr16_-_67944113 | 0.25 |
ENST00000264005.10
|
LCAT
|
lecithin-cholesterol acyltransferase |
| chr6_-_166167832 | 0.25 |
ENST00000366876.7
|
TBXT
|
T-box transcription factor T |
| chrX_-_130165699 | 0.25 |
ENST00000676328.1
ENST00000675857.1 ENST00000675427.1 ENST00000675092.1 |
AIFM1
|
apoptosis inducing factor mitochondria associated 1 |
| chr11_+_66593171 | 0.25 |
ENST00000533244.6
|
CCS
|
copper chaperone for superoxide dismutase |
| chr17_+_28744002 | 0.25 |
ENST00000618771.1
ENST00000262395.10 ENST00000422344.5 |
TRAF4
|
TNF receptor associated factor 4 |
| chr19_+_49877425 | 0.25 |
ENST00000622860.4
|
TBC1D17
|
TBC1 domain family member 17 |
| chrX_-_135052114 | 0.24 |
ENST00000370775.3
|
RTL8A
|
retrotransposon Gag like 8A |
| chr6_-_107115493 | 0.24 |
ENST00000369042.6
|
BEND3
|
BEN domain containing 3 |
| chr20_+_35699227 | 0.23 |
ENST00000374078.5
|
ROMO1
|
reactive oxygen species modulator 1 |
| chr16_-_9943182 | 0.23 |
ENST00000535259.6
|
GRIN2A
|
glutamate ionotropic receptor NMDA type subunit 2A |
| chr9_-_136050502 | 0.23 |
ENST00000371753.5
|
NACC2
|
NACC family member 2 |
| chr10_+_102644462 | 0.23 |
ENST00000643721.2
ENST00000302424.12 |
TRIM8
|
tripartite motif containing 8 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 3.9 | GO:0050720 | regulation of interleukin-1 biosynthetic process(GO:0045360) positive regulation of interleukin-1 biosynthetic process(GO:0045362) interleukin-1 beta biosynthetic process(GO:0050720) response to interleukin-8(GO:0098758) cellular response to interleukin-8(GO:0098759) regulation of progesterone biosynthetic process(GO:2000182) |
| 0.7 | 2.1 | GO:0050928 | negative regulation of positive chemotaxis(GO:0050928) |
| 0.7 | 2.0 | GO:0042668 | trochlear nerve development(GO:0021558) auditory receptor cell fate determination(GO:0042668) negative regulation of auditory receptor cell differentiation(GO:0045608) regulation of timing of neuron differentiation(GO:0060164) negative regulation of pro-B cell differentiation(GO:2000974) |
| 0.3 | 1.4 | GO:0021592 | fourth ventricle development(GO:0021592) |
| 0.3 | 1.2 | GO:0035284 | rhombomere 5 development(GO:0021571) central nervous system segmentation(GO:0035283) brain segmentation(GO:0035284) |
| 0.3 | 1.2 | GO:1904045 | cellular response to aldosterone(GO:1904045) |
| 0.3 | 1.0 | GO:0045659 | regulation of neutrophil differentiation(GO:0045658) negative regulation of neutrophil differentiation(GO:0045659) |
| 0.2 | 3.4 | GO:0061179 | negative regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0061179) |
| 0.2 | 0.7 | GO:0043973 | histone H3-K4 acetylation(GO:0043973) |
| 0.2 | 1.2 | GO:0035426 | extracellular matrix-cell signaling(GO:0035426) |
| 0.2 | 1.0 | GO:0097368 | establishment of Sertoli cell barrier(GO:0097368) |
| 0.2 | 0.6 | GO:1902948 | regulation of choline O-acetyltransferase activity(GO:1902769) positive regulation of choline O-acetyltransferase activity(GO:1902771) negative regulation of tau-protein kinase activity(GO:1902948) positive regulation of early endosome to recycling endosome transport(GO:1902955) negative regulation of aspartic-type endopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902960) negative regulation of neurofibrillary tangle assembly(GO:1902997) negative regulation of aspartic-type peptidase activity(GO:1905246) |
| 0.2 | 1.0 | GO:0033277 | abortive mitotic cell cycle(GO:0033277) |
| 0.1 | 0.7 | GO:0034499 | late endosome to Golgi transport(GO:0034499) |
| 0.1 | 0.9 | GO:0071279 | negative regulation of epithelial cell proliferation involved in prostate gland development(GO:0060770) cellular response to cobalt ion(GO:0071279) |
| 0.1 | 0.5 | GO:0046379 | extracellular polysaccharide biosynthetic process(GO:0045226) extracellular polysaccharide metabolic process(GO:0046379) |
| 0.1 | 4.0 | GO:0090023 | positive regulation of neutrophil chemotaxis(GO:0090023) |
| 0.1 | 1.3 | GO:0061299 | retina vasculature morphogenesis in camera-type eye(GO:0061299) |
| 0.1 | 0.6 | GO:0060399 | positive regulation of growth hormone receptor signaling pathway(GO:0060399) |
| 0.1 | 1.7 | GO:0071688 | striated muscle myosin thick filament assembly(GO:0071688) |
| 0.1 | 0.3 | GO:0014028 | notochord formation(GO:0014028) |
| 0.1 | 0.6 | GO:0015680 | intracellular copper ion transport(GO:0015680) |
| 0.1 | 0.4 | GO:0010725 | regulation of primitive erythrocyte differentiation(GO:0010725) eosinophil fate commitment(GO:0035854) |
| 0.1 | 0.5 | GO:0008218 | bioluminescence(GO:0008218) |
| 0.1 | 1.5 | GO:0090073 | positive regulation of protein homodimerization activity(GO:0090073) |
| 0.1 | 0.5 | GO:0051552 | flavone metabolic process(GO:0051552) |
| 0.1 | 0.4 | GO:0016480 | negative regulation of transcription from RNA polymerase III promoter(GO:0016480) |
| 0.1 | 0.5 | GO:0000821 | regulation of arginine metabolic process(GO:0000821) |
| 0.1 | 0.3 | GO:0061086 | negative regulation of histone H3-K27 methylation(GO:0061086) |
| 0.1 | 0.1 | GO:2000297 | negative regulation of synapse maturation(GO:2000297) |
| 0.1 | 0.3 | GO:0070460 | thyroid-stimulating hormone secretion(GO:0070460) regulation of thyroid-stimulating hormone secretion(GO:2000612) |
| 0.1 | 0.2 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 0.1 | 0.4 | GO:0006574 | valine catabolic process(GO:0006574) |
| 0.1 | 0.4 | GO:1903566 | positive regulation of protein localization to cilium(GO:1903566) |
| 0.1 | 0.5 | GO:0014859 | negative regulation of skeletal muscle cell proliferation(GO:0014859) negative regulation of skeletal muscle satellite cell proliferation(GO:1902723) |
| 0.1 | 0.4 | GO:0006535 | cysteine biosynthetic process from serine(GO:0006535) cysteine catabolic process(GO:0009093) L-cysteine catabolic process(GO:0019448) homocysteine catabolic process(GO:0043418) L-cysteine metabolic process(GO:0046439) |
| 0.1 | 0.3 | GO:0002032 | desensitization of G-protein coupled receptor protein signaling pathway by arrestin(GO:0002032) |
| 0.1 | 0.2 | GO:2000591 | positive regulation of metanephric mesenchymal cell migration by platelet-derived growth factor receptor-beta signaling pathway(GO:0035793) metanephric mesangial cell differentiation(GO:0072209) metanephric glomerular mesangial cell differentiation(GO:0072254) regulation of metanephric mesenchymal cell migration by platelet-derived growth factor receptor-beta signaling pathway(GO:1900238) positive regulation of metanephric mesenchymal cell migration(GO:2000591) |
| 0.1 | 0.3 | GO:0090107 | regulation of high-density lipoprotein particle assembly(GO:0090107) |
| 0.1 | 1.6 | GO:2000178 | negative regulation of neural precursor cell proliferation(GO:2000178) |
| 0.1 | 0.2 | GO:0010133 | proline catabolic process to glutamate(GO:0010133) |
| 0.1 | 0.2 | GO:0071140 | resolution of recombination intermediates(GO:0071139) resolution of mitotic recombination intermediates(GO:0071140) |
| 0.1 | 0.9 | GO:0051715 | cytolysis in other organism(GO:0051715) |
| 0.1 | 0.6 | GO:0007256 | activation of JNKK activity(GO:0007256) |
| 0.1 | 0.6 | GO:0070120 | ciliary neurotrophic factor-mediated signaling pathway(GO:0070120) |
| 0.1 | 0.3 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.0 | 0.3 | GO:0070345 | negative regulation of fat cell proliferation(GO:0070345) |
| 0.0 | 1.3 | GO:0034389 | lipid particle organization(GO:0034389) |
| 0.0 | 0.3 | GO:0098532 | histone H3-K27 trimethylation(GO:0098532) |
| 0.0 | 1.0 | GO:0040015 | negative regulation of multicellular organism growth(GO:0040015) |
| 0.0 | 0.1 | GO:1990927 | short-term synaptic potentiation(GO:1990926) calcium ion regulated lysosome exocytosis(GO:1990927) |
| 0.0 | 1.5 | GO:2001275 | positive regulation of glucose import in response to insulin stimulus(GO:2001275) |
| 0.0 | 0.3 | GO:1903385 | regulation of homophilic cell adhesion(GO:1903385) |
| 0.0 | 0.4 | GO:0061370 | testosterone biosynthetic process(GO:0061370) |
| 0.0 | 0.2 | GO:0033484 | nitric oxide homeostasis(GO:0033484) |
| 0.0 | 0.2 | GO:0033058 | directional locomotion(GO:0033058) |
| 0.0 | 0.4 | GO:0045039 | protein import into mitochondrial inner membrane(GO:0045039) |
| 0.0 | 0.1 | GO:0036233 | glycine import(GO:0036233) |
| 0.0 | 0.1 | GO:0031446 | regulation of fast-twitch skeletal muscle fiber contraction(GO:0031446) positive regulation of fast-twitch skeletal muscle fiber contraction(GO:0031448) maintenance of mitochondrion location(GO:0051659) |
| 0.0 | 1.2 | GO:0051412 | response to corticosterone(GO:0051412) |
| 0.0 | 0.3 | GO:0042791 | 5S class rRNA transcription from RNA polymerase III type 1 promoter(GO:0042791) tRNA transcription from RNA polymerase III promoter(GO:0042797) |
| 0.0 | 0.6 | GO:0070734 | histone H3-K27 methylation(GO:0070734) |
| 0.0 | 1.0 | GO:2000121 | regulation of removal of superoxide radicals(GO:2000121) |
| 0.0 | 0.3 | GO:0060907 | positive regulation of macrophage cytokine production(GO:0060907) |
| 0.0 | 0.3 | GO:2001206 | positive regulation of osteoclast development(GO:2001206) |
| 0.0 | 0.1 | GO:0010609 | mRNA localization resulting in posttranscriptional regulation of gene expression(GO:0010609) platelet alpha granule organization(GO:0070889) regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902162) positive regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902164) |
| 0.0 | 0.3 | GO:0017196 | N-terminal peptidyl-methionine acetylation(GO:0017196) |
| 0.0 | 0.2 | GO:0006420 | arginyl-tRNA aminoacylation(GO:0006420) |
| 0.0 | 1.1 | GO:0072189 | ureter development(GO:0072189) |
| 0.0 | 0.2 | GO:1902231 | positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage(GO:1902231) |
| 0.0 | 0.3 | GO:0021999 | neural plate anterior/posterior regionalization(GO:0021999) |
| 0.0 | 0.2 | GO:0051697 | protein delipidation(GO:0051697) |
| 0.0 | 0.4 | GO:0048733 | sebaceous gland development(GO:0048733) |
| 0.0 | 0.3 | GO:0050703 | interleukin-1 alpha secretion(GO:0050703) |
| 0.0 | 0.3 | GO:1903361 | protein localization to basolateral plasma membrane(GO:1903361) |
| 0.0 | 0.5 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.0 | 0.5 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.0 | 0.5 | GO:0060272 | embryonic skeletal joint morphogenesis(GO:0060272) |
| 0.0 | 0.6 | GO:0097118 | neuroligin clustering involved in postsynaptic membrane assembly(GO:0097118) |
| 0.0 | 0.3 | GO:2000381 | negative regulation of mesoderm development(GO:2000381) |
| 0.0 | 0.2 | GO:0051533 | positive regulation of NFAT protein import into nucleus(GO:0051533) |
| 0.0 | 2.9 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.0 | 0.1 | GO:0044027 | hypermethylation of CpG island(GO:0044027) |
| 0.0 | 0.4 | GO:0043983 | histone H4-K12 acetylation(GO:0043983) |
| 0.0 | 0.3 | GO:0042074 | cell migration involved in gastrulation(GO:0042074) |
| 0.0 | 0.1 | GO:0071931 | positive regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0071931) |
| 0.0 | 0.4 | GO:0035372 | protein localization to microtubule(GO:0035372) |
| 0.0 | 0.3 | GO:0044387 | negative regulation of protein kinase activity by regulation of protein phosphorylation(GO:0044387) |
| 0.0 | 0.1 | GO:1904640 | response to methionine(GO:1904640) |
| 0.0 | 0.6 | GO:0006743 | ubiquinone metabolic process(GO:0006743) ubiquinone biosynthetic process(GO:0006744) quinone biosynthetic process(GO:1901663) |
| 0.0 | 0.6 | GO:0046597 | negative regulation of viral entry into host cell(GO:0046597) |
| 0.0 | 0.5 | GO:0044849 | estrous cycle(GO:0044849) |
| 0.0 | 0.2 | GO:0002084 | protein depalmitoylation(GO:0002084) |
| 0.0 | 0.2 | GO:0045647 | negative regulation of erythrocyte differentiation(GO:0045647) |
| 0.0 | 0.3 | GO:0061734 | parkin-mediated mitophagy in response to mitochondrial depolarization(GO:0061734) |
| 0.0 | 0.3 | GO:0060088 | auditory receptor cell stereocilium organization(GO:0060088) |
| 0.0 | 0.2 | GO:0036309 | protein localization to M-band(GO:0036309) |
| 0.0 | 0.3 | GO:0031580 | membrane raft polarization(GO:0001766) membrane raft distribution(GO:0031580) |
| 0.0 | 0.1 | GO:0030047 | actin modification(GO:0030047) |
| 0.0 | 0.4 | GO:0045721 | negative regulation of gluconeogenesis(GO:0045721) |
| 0.0 | 0.2 | GO:0061343 | cell adhesion involved in heart morphogenesis(GO:0061343) |
| 0.0 | 0.1 | GO:0071105 | response to interleukin-11(GO:0071105) |
| 0.0 | 0.5 | GO:0042118 | endothelial cell activation(GO:0042118) |
| 0.0 | 0.1 | GO:0016240 | autophagosome docking(GO:0016240) |
| 0.0 | 0.2 | GO:0019227 | neuronal action potential propagation(GO:0019227) action potential propagation(GO:0098870) |
| 0.0 | 0.0 | GO:0060739 | mesenchymal-epithelial cell signaling involved in prostate gland development(GO:0060739) |
| 0.0 | 0.2 | GO:0031087 | nuclear-transcribed mRNA catabolic process, deadenylation-independent decay(GO:0031086) deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
| 0.0 | 0.1 | GO:1901315 | negative regulation of histone ubiquitination(GO:0033183) regulation of histone H2A K63-linked ubiquitination(GO:1901314) negative regulation of histone H2A K63-linked ubiquitination(GO:1901315) |
| 0.0 | 0.2 | GO:0034036 | purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 0.0 | 0.2 | GO:0034380 | high-density lipoprotein particle assembly(GO:0034380) |
| 0.0 | 0.8 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.0 | 0.0 | GO:0070634 | transepithelial ammonium transport(GO:0070634) |
| 0.0 | 0.2 | GO:0060346 | bone trabecula formation(GO:0060346) |
| 0.0 | 0.3 | GO:0060022 | hard palate development(GO:0060022) |
| 0.0 | 0.3 | GO:1990001 | inhibition of cysteine-type endopeptidase activity involved in apoptotic process(GO:1990001) |
| 0.0 | 0.4 | GO:0034033 | coenzyme A biosynthetic process(GO:0015937) nucleoside bisphosphate biosynthetic process(GO:0033866) ribonucleoside bisphosphate biosynthetic process(GO:0034030) purine nucleoside bisphosphate biosynthetic process(GO:0034033) |
| 0.0 | 0.1 | GO:0070294 | renal sodium ion transport(GO:0003096) renal sodium ion absorption(GO:0070294) |
| 0.0 | 0.5 | GO:0030252 | growth hormone secretion(GO:0030252) |
| 0.0 | 0.3 | GO:0060716 | labyrinthine layer blood vessel development(GO:0060716) |
| 0.0 | 0.4 | GO:0009235 | cobalamin metabolic process(GO:0009235) |
| 0.0 | 0.1 | GO:0008343 | adult feeding behavior(GO:0008343) |
| 0.0 | 0.0 | GO:0070682 | proteasome regulatory particle assembly(GO:0070682) |
| 0.0 | 0.2 | GO:0030210 | heparin metabolic process(GO:0030202) heparin biosynthetic process(GO:0030210) |
| 0.0 | 0.7 | GO:0060612 | adipose tissue development(GO:0060612) |
| 0.0 | 0.1 | GO:0033623 | regulation of integrin activation(GO:0033623) |
| 0.0 | 0.1 | GO:0070315 | G1 to G0 transition involved in cell differentiation(GO:0070315) |
| 0.0 | 0.1 | GO:0055059 | asymmetric neuroblast division(GO:0055059) |
| 0.0 | 0.2 | GO:0002862 | negative regulation of inflammatory response to antigenic stimulus(GO:0002862) |
| 0.0 | 0.7 | GO:0071377 | cellular response to glucagon stimulus(GO:0071377) |
| 0.0 | 0.2 | GO:0006388 | tRNA splicing, via endonucleolytic cleavage and ligation(GO:0006388) |
| 0.0 | 0.9 | GO:0070206 | protein trimerization(GO:0070206) |
| 0.0 | 0.2 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 0.0 | 0.7 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.0 | 0.2 | GO:0035414 | negative regulation of catenin import into nucleus(GO:0035414) |
| 0.0 | 0.2 | GO:0010225 | response to UV-C(GO:0010225) |
| 0.0 | 0.1 | GO:0043461 | proton-transporting ATP synthase complex assembly(GO:0043461) proton-transporting ATP synthase complex biogenesis(GO:0070272) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.7 | GO:1990393 | 3M complex(GO:1990393) |
| 0.3 | 1.4 | GO:0097513 | myosin II filament(GO:0097513) |
| 0.2 | 0.5 | GO:0036117 | hyaluranon cable(GO:0036117) |
| 0.2 | 0.6 | GO:0097059 | CNTFR-CLCF1 complex(GO:0097059) |
| 0.1 | 1.0 | GO:0071953 | elastic fiber(GO:0071953) |
| 0.1 | 1.0 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 0.1 | 0.9 | GO:0043203 | axon hillock(GO:0043203) |
| 0.1 | 0.2 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.1 | 1.8 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.1 | 0.4 | GO:0042721 | mitochondrial inner membrane protein insertion complex(GO:0042721) |
| 0.1 | 0.3 | GO:0031417 | NatC complex(GO:0031417) |
| 0.1 | 0.3 | GO:1990769 | proximal neuron projection(GO:1990769) |
| 0.0 | 0.4 | GO:0005851 | eukaryotic translation initiation factor 2B complex(GO:0005851) |
| 0.0 | 0.3 | GO:0000127 | transcription factor TFIIIC complex(GO:0000127) |
| 0.0 | 0.9 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.0 | 0.6 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.0 | 0.1 | GO:0036156 | inner dynein arm(GO:0036156) |
| 0.0 | 0.5 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.0 | 0.1 | GO:0014802 | terminal cisterna(GO:0014802) |
| 0.0 | 0.3 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.0 | 0.2 | GO:0033063 | Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 0.0 | 0.1 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 0.0 | 1.7 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.0 | 0.6 | GO:0010369 | chromocenter(GO:0010369) |
| 0.0 | 0.3 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.0 | 0.3 | GO:0016600 | flotillin complex(GO:0016600) |
| 0.0 | 1.8 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.0 | 0.4 | GO:0032982 | myosin filament(GO:0032982) |
| 0.0 | 0.0 | GO:0031251 | PAN complex(GO:0031251) |
| 0.0 | 0.3 | GO:0032426 | stereocilium tip(GO:0032426) |
| 0.0 | 0.3 | GO:0097025 | MPP7-DLG1-LIN7 complex(GO:0097025) |
| 0.0 | 1.2 | GO:1904724 | tertiary granule lumen(GO:1904724) |
| 0.0 | 0.2 | GO:0036057 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.0 | 0.1 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.0 | 0.1 | GO:0070775 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.0 | 0.3 | GO:0035327 | transcriptionally active chromatin(GO:0035327) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 3.7 | GO:0044729 | hemi-methylated DNA-binding(GO:0044729) |
| 0.4 | 1.7 | GO:0042282 | hydroxymethylglutaryl-CoA reductase (NADPH) activity(GO:0004420) hydroxymethylglutaryl-CoA reductase activity(GO:0042282) |
| 0.3 | 1.6 | GO:0005219 | ryanodine-sensitive calcium-release channel activity(GO:0005219) |
| 0.3 | 4.5 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.2 | 1.0 | GO:0016314 | phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase activity(GO:0016314) |
| 0.2 | 1.5 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.2 | 1.4 | GO:0043237 | laminin-1 binding(GO:0043237) |
| 0.1 | 0.6 | GO:0004706 | JUN kinase kinase kinase activity(GO:0004706) |
| 0.1 | 0.6 | GO:0016532 | superoxide dismutase copper chaperone activity(GO:0016532) |
| 0.1 | 0.9 | GO:0008269 | JAK pathway signal transduction adaptor activity(GO:0008269) |
| 0.1 | 0.3 | GO:0000252 | C-3 sterol dehydrogenase (C-4 sterol decarboxylase) activity(GO:0000252) sterol-4-alpha-carboxylate 3-dehydrogenase (decarboxylating) activity(GO:0047012) |
| 0.1 | 0.4 | GO:0004616 | phosphogluconate dehydrogenase (decarboxylating) activity(GO:0004616) |
| 0.1 | 0.7 | GO:0047045 | testosterone 17-beta-dehydrogenase (NADP+) activity(GO:0047045) |
| 0.1 | 0.3 | GO:0000992 | polymerase III regulatory region sequence-specific DNA binding(GO:0000992) RNA polymerase III type 1 promoter sequence-specific DNA binding(GO:0001002) RNA polymerase III type 2 promoter sequence-specific DNA binding(GO:0001003) |
| 0.1 | 0.4 | GO:0070025 | oxidoreductase activity, acting on other nitrogenous compounds as donors, cytochrome as acceptor(GO:0016662) nitrite reductase (NO-forming) activity(GO:0050421) carbon monoxide binding(GO:0070025) nitrite reductase activity(GO:0098809) |
| 0.1 | 1.2 | GO:0016174 | NAD(P)H oxidase activity(GO:0016174) |
| 0.1 | 0.5 | GO:0050501 | hyaluronan synthase activity(GO:0050501) |
| 0.1 | 1.5 | GO:0031996 | thioesterase binding(GO:0031996) |
| 0.1 | 0.3 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.1 | 1.6 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.1 | 0.6 | GO:0000285 | 1-phosphatidylinositol-3-phosphate 5-kinase activity(GO:0000285) |
| 0.1 | 1.8 | GO:0097493 | structural molecule activity conferring elasticity(GO:0097493) |
| 0.1 | 0.3 | GO:0031826 | type 2A serotonin receptor binding(GO:0031826) |
| 0.1 | 0.6 | GO:0004897 | ciliary neurotrophic factor receptor activity(GO:0004897) |
| 0.1 | 0.6 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.1 | 1.4 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.1 | 0.4 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 0.1 | 0.7 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.1 | 0.1 | GO:0004999 | vasoactive intestinal polypeptide receptor activity(GO:0004999) |
| 0.1 | 0.2 | GO:0008665 | 2'-phosphotransferase activity(GO:0008665) |
| 0.1 | 0.8 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.0 | 0.2 | GO:0033857 | diphosphoinositol-pentakisphosphate kinase activity(GO:0033857) |
| 0.0 | 0.3 | GO:0034186 | apolipoprotein A-I binding(GO:0034186) |
| 0.0 | 1.1 | GO:0071949 | FAD binding(GO:0071949) |
| 0.0 | 1.3 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.0 | 0.3 | GO:0004996 | thyroid-stimulating hormone receptor activity(GO:0004996) |
| 0.0 | 0.6 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.0 | 0.2 | GO:0004814 | arginine-tRNA ligase activity(GO:0004814) |
| 0.0 | 0.3 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.0 | 0.4 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.0 | 0.3 | GO:0005094 | Rho GDP-dissociation inhibitor activity(GO:0005094) |
| 0.0 | 0.5 | GO:0015174 | basic amino acid transmembrane transporter activity(GO:0015174) |
| 0.0 | 0.1 | GO:0015375 | glycine:sodium symporter activity(GO:0015375) |
| 0.0 | 0.3 | GO:0004972 | NMDA glutamate receptor activity(GO:0004972) |
| 0.0 | 0.3 | GO:0000990 | transcription factor activity, core RNA polymerase binding(GO:0000990) |
| 0.0 | 0.2 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 0.0 | 0.2 | GO:0003998 | acylphosphatase activity(GO:0003998) |
| 0.0 | 0.2 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.0 | 0.2 | GO:0070699 | type II activin receptor binding(GO:0070699) |
| 0.0 | 0.2 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.0 | 0.3 | GO:0005225 | volume-sensitive anion channel activity(GO:0005225) |
| 0.0 | 0.7 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.0 | 0.4 | GO:0001011 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.0 | 0.3 | GO:0031702 | type 1 angiotensin receptor binding(GO:0031702) |
| 0.0 | 0.4 | GO:0043422 | protein kinase B binding(GO:0043422) |
| 0.0 | 0.2 | GO:0000182 | rDNA binding(GO:0000182) |
| 0.0 | 0.3 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 0.0 | 0.2 | GO:0008821 | crossover junction endodeoxyribonuclease activity(GO:0008821) |
| 0.0 | 0.3 | GO:0097016 | L27 domain binding(GO:0097016) |
| 0.0 | 0.3 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 0.3 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.0 | 0.2 | GO:0098599 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 0.0 | 0.3 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 2.2 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.0 | 0.4 | GO:0033130 | acetylcholine receptor binding(GO:0033130) |
| 0.0 | 0.1 | GO:0004522 | ribonuclease A activity(GO:0004522) |
| 0.0 | 0.6 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.0 | 0.5 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.0 | 0.3 | GO:0031005 | filamin binding(GO:0031005) |
| 0.0 | 1.1 | GO:0004407 | histone deacetylase activity(GO:0004407) |
| 0.0 | 0.1 | GO:0061628 | H3K27me3 modified histone binding(GO:0061628) |
| 0.0 | 0.1 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.0 | 0.1 | GO:0035368 | selenocysteine insertion sequence binding(GO:0035368) |
| 0.0 | 0.3 | GO:0070742 | C2H2 zinc finger domain binding(GO:0070742) |
| 0.0 | 0.1 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.0 | 1.0 | GO:0048365 | Rac GTPase binding(GO:0048365) |
| 0.0 | 0.1 | GO:1990460 | leptin receptor binding(GO:1990460) |
| 0.0 | 0.5 | GO:0032452 | histone demethylase activity(GO:0032452) |
| 0.0 | 4.8 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.0 | 0.1 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.0 | 0.6 | GO:0030552 | cAMP binding(GO:0030552) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.7 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.0 | 2.1 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.0 | 1.8 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.0 | 1.1 | PID IL23 PATHWAY | IL23-mediated signaling events |
| 0.0 | 1.3 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 0.8 | PID PI3K PLC TRK PATHWAY | Trk receptor signaling mediated by PI3K and PLC-gamma |
| 0.0 | 1.3 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.0 | 1.0 | PID IL3 PATHWAY | IL3-mediated signaling events |
| 0.0 | 0.3 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
| 0.0 | 0.8 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.0 | 0.7 | PID SHP2 PATHWAY | SHP2 signaling |
| 0.0 | 0.3 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.0 | 1.2 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.0 | 0.9 | PID ERBB1 RECEPTOR PROXIMAL PATHWAY | EGF receptor (ErbB1) signaling pathway |
| 0.0 | 0.9 | PID IL2 1PATHWAY | IL2-mediated signaling events |
| 0.0 | 1.1 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 0.1 | PID ERBB2 ERBB3 PATHWAY | ErbB2/ErbB3 signaling events |
| 0.0 | 1.0 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 1.4 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.0 | 0.7 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 2.7 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.1 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.1 | 1.5 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.1 | 4.5 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.1 | 1.0 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.0 | 2.0 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.7 | REACTOME ANDROGEN BIOSYNTHESIS | Genes involved in Androgen biosynthesis |
| 0.0 | 3.9 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 1.2 | REACTOME GROWTH HORMONE RECEPTOR SIGNALING | Genes involved in Growth hormone receptor signaling |
| 0.0 | 0.5 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.0 | 0.7 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.0 | 0.4 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.0 | 0.9 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.0 | 1.0 | REACTOME THROMBIN SIGNALLING THROUGH PROTEINASE ACTIVATED RECEPTORS PARS | Genes involved in Thrombin signalling through proteinase activated receptors (PARs) |
| 0.0 | 1.9 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.0 | 1.2 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 0.5 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.0 | 0.5 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.0 | 0.2 | REACTOME SHC MEDIATED SIGNALLING | Genes involved in SHC-mediated signalling |
| 0.0 | 0.5 | REACTOME HYALURONAN METABOLISM | Genes involved in Hyaluronan metabolism |
| 0.0 | 0.6 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.0 | 0.2 | REACTOME UNBLOCKING OF NMDA RECEPTOR GLUTAMATE BINDING AND ACTIVATION | Genes involved in Unblocking of NMDA receptor, glutamate binding and activation |
| 0.0 | 0.4 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.0 | 0.9 | REACTOME TRAFFICKING OF AMPA RECEPTORS | Genes involved in Trafficking of AMPA receptors |
| 0.0 | 0.5 | REACTOME RNA POL I TRANSCRIPTION INITIATION | Genes involved in RNA Polymerase I Transcription Initiation |
| 0.0 | 0.4 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.0 | 0.5 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |