Project
Inflammatory response time course, HUVEC (Wada, 2009)
Navigation
Downloads
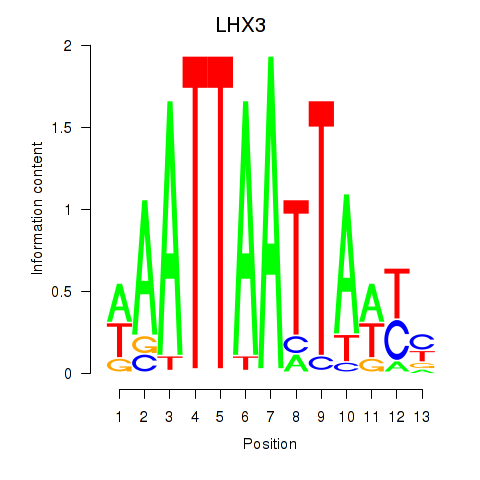
Results for LHX3
Z-value: 0.47
Transcription factors associated with LHX3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
LHX3
|
ENSG00000107187.17 | LHX3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| LHX3 | hg38_v1_chr9_-_136203183_136203194, hg38_v1_chr9_-_136205122_136205133 | -0.34 | 9.3e-02 | Click! |
Activity profile of LHX3 motif
Sorted Z-values of LHX3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of LHX3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr9_+_72577939 | 1.26 |
ENST00000645773.1
|
TMC1
|
transmembrane channel like 1 |
| chr3_-_142029108 | 1.00 |
ENST00000497579.5
|
TFDP2
|
transcription factor Dp-2 |
| chr4_+_41613476 | 0.81 |
ENST00000508466.1
|
LIMCH1
|
LIM and calponin homology domains 1 |
| chr19_+_14583076 | 0.72 |
ENST00000547437.5
ENST00000417570.6 |
CLEC17A
|
C-type lectin domain containing 17A |
| chr2_-_187448244 | 0.63 |
ENST00000392370.8
ENST00000410068.5 ENST00000447403.5 ENST00000410102.5 |
CALCRL
|
calcitonin receptor like receptor |
| chr1_-_150235972 | 0.61 |
ENST00000534220.1
|
ANP32E
|
acidic nuclear phosphoprotein 32 family member E |
| chr9_+_72577788 | 0.56 |
ENST00000645208.2
|
TMC1
|
transmembrane channel like 1 |
| chr9_+_72577369 | 0.54 |
ENST00000651183.1
|
TMC1
|
transmembrane channel like 1 |
| chr9_+_79573162 | 0.39 |
ENST00000425506.5
|
TLE4
|
TLE family member 4, transcriptional corepressor |
| chr8_-_30812867 | 0.37 |
ENST00000518243.5
|
PPP2CB
|
protein phosphatase 2 catalytic subunit beta |
| chr9_-_72953047 | 0.36 |
ENST00000297785.8
ENST00000376939.5 |
ALDH1A1
|
aldehyde dehydrogenase 1 family member A1 |
| chr1_-_160954801 | 0.35 |
ENST00000368029.4
|
ITLN2
|
intelectin 2 |
| chr7_-_86965872 | 0.33 |
ENST00000398276.6
ENST00000416314.5 ENST00000425689.1 |
ELAPOR2
|
endosome-lysosome associated apoptosis and autophagy regulator family member 2 |
| chr17_-_41467386 | 0.33 |
ENST00000225899.4
|
KRT32
|
keratin 32 |
| chr17_-_40937641 | 0.26 |
ENST00000209718.8
|
KRT23
|
keratin 23 |
| chr7_-_105691637 | 0.24 |
ENST00000472195.1
|
ATXN7L1
|
ataxin 7 like 1 |
| chr17_-_40937445 | 0.24 |
ENST00000436344.7
ENST00000485751.1 |
KRT23
|
keratin 23 |
| chr18_-_55635948 | 0.24 |
ENST00000565124.4
ENST00000398339.5 |
TCF4
|
transcription factor 4 |
| chr15_-_19988117 | 0.23 |
ENST00000558565.2
|
IGHV3OR15-7
|
immunoglobulin heavy variable 3/OR15-7 (pseudogene) |
| chr5_-_88823763 | 0.23 |
ENST00000635898.1
ENST00000626391.2 ENST00000628656.2 |
MEF2C
|
myocyte enhancer factor 2C |
| chr1_+_81306096 | 0.22 |
ENST00000370721.5
ENST00000370727.5 ENST00000370725.5 ENST00000370723.5 ENST00000370728.5 ENST00000370730.5 |
ADGRL2
|
adhesion G protein-coupled receptor L2 |
| chr20_-_31390580 | 0.22 |
ENST00000339144.3
ENST00000376321.4 |
DEFB119
|
defensin beta 119 |
| chr15_-_75455767 | 0.21 |
ENST00000360439.8
|
SIN3A
|
SIN3 transcription regulator family member A |
| chr9_+_122370523 | 0.21 |
ENST00000643810.1
ENST00000540753.6 |
PTGS1
|
prostaglandin-endoperoxide synthase 1 |
| chr3_-_20012250 | 0.20 |
ENST00000389050.5
|
PP2D1
|
protein phosphatase 2C like domain containing 1 |
| chr15_+_65550819 | 0.20 |
ENST00000569894.5
|
HACD3
|
3-hydroxyacyl-CoA dehydratase 3 |
| chr9_+_96928310 | 0.20 |
ENST00000354649.7
|
NUTM2G
|
NUT family member 2G |
| chr1_-_150235943 | 0.19 |
ENST00000533654.5
|
ANP32E
|
acidic nuclear phosphoprotein 32 family member E |
| chr2_-_88860605 | 0.18 |
ENST00000390238.2
|
IGKJ5
|
immunoglobulin kappa joining 5 |
| chr18_-_55423757 | 0.17 |
ENST00000675707.1
|
TCF4
|
transcription factor 4 |
| chr19_-_51417700 | 0.16 |
ENST00000529627.1
ENST00000439889.6 |
SIGLEC10
|
sialic acid binding Ig like lectin 10 |
| chr1_+_207325629 | 0.15 |
ENST00000618707.2
|
CD55
|
CD55 molecule (Cromer blood group) |
| chr7_-_36724543 | 0.15 |
ENST00000612871.4
|
AOAH
|
acyloxyacyl hydrolase |
| chr17_-_10469558 | 0.15 |
ENST00000255381.2
|
MYH4
|
myosin heavy chain 4 |
| chr1_+_15341744 | 0.15 |
ENST00000444385.5
|
FHAD1
|
forkhead associated phosphopeptide binding domain 1 |
| chr5_+_148312416 | 0.15 |
ENST00000274565.5
|
SPINK7
|
serine peptidase inhibitor Kazal type 7 |
| chr1_-_182671853 | 0.15 |
ENST00000367556.5
|
RGS8
|
regulator of G protein signaling 8 |
| chr1_-_182671902 | 0.15 |
ENST00000483095.6
|
RGS8
|
regulator of G protein signaling 8 |
| chr3_+_2892199 | 0.14 |
ENST00000397459.6
|
CNTN4
|
contactin 4 |
| chr11_-_63608542 | 0.14 |
ENST00000540943.1
|
PLAAT3
|
phospholipase A and acyltransferase 3 |
| chr7_-_4862015 | 0.14 |
ENST00000404991.2
|
PAPOLB
|
poly(A) polymerase beta |
| chr7_-_36724457 | 0.14 |
ENST00000617537.5
ENST00000435386.1 |
AOAH
|
acyloxyacyl hydrolase |
| chr19_-_51417791 | 0.14 |
ENST00000353836.9
|
SIGLEC10
|
sialic acid binding Ig like lectin 10 |
| chr20_-_31390483 | 0.13 |
ENST00000376315.2
|
DEFB119
|
defensin beta 119 |
| chr17_-_5035418 | 0.12 |
ENST00000254853.10
ENST00000424747.1 |
SLC52A1
|
solute carrier family 52 member 1 |
| chr8_-_18083278 | 0.12 |
ENST00000636691.1
|
ASAH1
|
N-acylsphingosine amidohydrolase 1 |
| chr9_-_113303271 | 0.12 |
ENST00000297894.5
ENST00000489339.2 |
RNF183
|
ring finger protein 183 |
| chr17_+_37491464 | 0.12 |
ENST00000613659.1
|
DUSP14
|
dual specificity phosphatase 14 |
| chr16_+_15395745 | 0.12 |
ENST00000287594.7
ENST00000396385.4 ENST00000568766.1 |
MPV17L
ENSG00000261130.5
|
MPV17 mitochondrial inner membrane protein like novel protein |
| chr10_+_87357720 | 0.12 |
ENST00000412718.3
ENST00000381697.7 |
NUTM2D
|
NUT family member 2D |
| chr17_+_70075317 | 0.12 |
ENST00000589377.1
|
KCNJ16
|
potassium inwardly rectifying channel subfamily J member 16 |
| chr1_-_158426237 | 0.12 |
ENST00000641042.1
|
OR10K2
|
olfactory receptor family 10 subfamily K member 2 |
| chr5_-_16916400 | 0.11 |
ENST00000513882.5
|
MYO10
|
myosin X |
| chr14_+_88385714 | 0.11 |
ENST00000045347.11
|
SPATA7
|
spermatogenesis associated 7 |
| chr3_+_141387801 | 0.11 |
ENST00000514251.5
|
ZBTB38
|
zinc finger and BTB domain containing 38 |
| chr13_-_60013178 | 0.10 |
ENST00000498416.2
ENST00000465066.5 |
DIAPH3
|
diaphanous related formin 3 |
| chr5_+_55853314 | 0.10 |
ENST00000354961.8
ENST00000297015.7 |
IL31RA
|
interleukin 31 receptor A |
| chr3_+_158110052 | 0.10 |
ENST00000295930.7
ENST00000471994.5 ENST00000482822.3 ENST00000476899.6 ENST00000683899.1 ENST00000684604.1 ENST00000682164.1 ENST00000464171.5 ENST00000611884.5 ENST00000312179.10 ENST00000475278.6 |
RSRC1
|
arginine and serine rich coiled-coil 1 |
| chr8_+_91249307 | 0.10 |
ENST00000309536.6
ENST00000276609.8 |
SLC26A7
|
solute carrier family 26 member 7 |
| chr17_+_1771688 | 0.09 |
ENST00000572048.1
ENST00000573763.1 |
SERPINF1
|
serpin family F member 1 |
| chr3_+_68006224 | 0.09 |
ENST00000496687.1
|
TAFA1
|
TAFA chemokine like family member 1 |
| chr5_-_1882902 | 0.09 |
ENST00000231357.7
|
IRX4
|
iroquois homeobox 4 |
| chr15_-_55408245 | 0.09 |
ENST00000563171.5
ENST00000425574.7 ENST00000442196.8 ENST00000564092.1 |
CCPG1
|
cell cycle progression 1 |
| chr2_-_98663464 | 0.09 |
ENST00000414521.6
|
MGAT4A
|
alpha-1,3-mannosyl-glycoprotein 4-beta-N-acetylglucosaminyltransferase A |
| chr6_-_49744434 | 0.09 |
ENST00000433368.6
ENST00000354620.4 |
CRISP3
|
cysteine rich secretory protein 3 |
| chr1_-_93681829 | 0.09 |
ENST00000260502.11
|
BCAR3
|
BCAR3 adaptor protein, NSP family member |
| chr6_-_49744378 | 0.09 |
ENST00000371159.8
ENST00000263045.9 |
CRISP3
|
cysteine rich secretory protein 3 |
| chr18_-_24311495 | 0.09 |
ENST00000357041.8
|
OSBPL1A
|
oxysterol binding protein like 1A |
| chr12_+_49346911 | 0.09 |
ENST00000395069.3
|
DNAJC22
|
DnaJ heat shock protein family (Hsp40) member C22 |
| chr1_-_92486916 | 0.08 |
ENST00000294702.6
|
GFI1
|
growth factor independent 1 transcriptional repressor |
| chr8_+_106726115 | 0.08 |
ENST00000521592.5
|
OXR1
|
oxidation resistance 1 |
| chr1_+_244051275 | 0.08 |
ENST00000358704.4
|
ZBTB18
|
zinc finger and BTB domain containing 18 |
| chr14_+_21997531 | 0.08 |
ENST00000390445.2
|
TRAV17
|
T cell receptor alpha variable 17 |
| chr19_+_49513154 | 0.08 |
ENST00000426395.7
ENST00000600273.5 ENST00000599988.5 |
FCGRT
|
Fc fragment of IgG receptor and transporter |
| chr3_+_46877705 | 0.08 |
ENST00000449590.6
|
PTH1R
|
parathyroid hormone 1 receptor |
| chr3_-_142000353 | 0.08 |
ENST00000499676.5
|
TFDP2
|
transcription factor Dp-2 |
| chr6_+_72216442 | 0.07 |
ENST00000425662.6
ENST00000453976.6 |
RIMS1
|
regulating synaptic membrane exocytosis 1 |
| chr6_+_152697888 | 0.07 |
ENST00000367245.5
ENST00000529453.1 |
MYCT1
|
MYC target 1 |
| chr5_-_39274515 | 0.07 |
ENST00000510188.1
|
FYB1
|
FYN binding protein 1 |
| chr11_-_790062 | 0.07 |
ENST00000330106.5
|
CEND1
|
cell cycle exit and neuronal differentiation 1 |
| chr6_+_122996227 | 0.07 |
ENST00000275162.10
|
CLVS2
|
clavesin 2 |
| chr4_+_53377630 | 0.07 |
ENST00000337488.11
ENST00000358575.9 ENST00000507922.5 ENST00000306932.10 |
FIP1L1
|
factor interacting with PAPOLA and CPSF1 |
| chr6_+_106360668 | 0.06 |
ENST00000633556.3
|
CRYBG1
|
crystallin beta-gamma domain containing 1 |
| chr6_-_31120437 | 0.06 |
ENST00000376288.3
|
CDSN
|
corneodesmosin |
| chr17_-_44066595 | 0.06 |
ENST00000585388.2
ENST00000293406.8 |
LSM12
|
LSM12 homolog |
| chr8_-_109974688 | 0.06 |
ENST00000297404.1
|
KCNV1
|
potassium voltage-gated channel modifier subfamily V member 1 |
| chr3_+_101724602 | 0.06 |
ENST00000341893.8
|
CEP97
|
centrosomal protein 97 |
| chr13_-_83882456 | 0.06 |
ENST00000674365.1
|
SLITRK1
|
SLIT and NTRK like family member 1 |
| chr19_+_49513353 | 0.06 |
ENST00000596975.5
|
FCGRT
|
Fc fragment of IgG receptor and transporter |
| chr13_+_53028806 | 0.05 |
ENST00000219022.3
|
OLFM4
|
olfactomedin 4 |
| chr6_+_130018565 | 0.05 |
ENST00000361794.7
ENST00000526087.5 ENST00000533560.5 |
L3MBTL3
|
L3MBTL histone methyl-lysine binding protein 3 |
| chr16_+_67570741 | 0.05 |
ENST00000644753.1
ENST00000642819.1 ENST00000645306.1 |
CTCF
|
CCCTC-binding factor |
| chr19_+_48695952 | 0.05 |
ENST00000522966.2
ENST00000425340.3 ENST00000391876.5 |
FUT2
|
fucosyltransferase 2 |
| chr3_-_64019334 | 0.05 |
ENST00000480205.5
|
PSMD6
|
proteasome 26S subunit, non-ATPase 6 |
| chr19_+_11346499 | 0.05 |
ENST00000458408.6
ENST00000586451.5 ENST00000588592.5 |
CCDC159
|
coiled-coil domain containing 159 |
| chr12_-_86256299 | 0.04 |
ENST00000552808.6
ENST00000547225.5 |
MGAT4C
|
MGAT4 family member C |
| chr8_-_18083184 | 0.04 |
ENST00000636269.1
|
ASAH1
|
N-acylsphingosine amidohydrolase 1 |
| chr6_+_26103922 | 0.04 |
ENST00000377803.4
|
H4C3
|
H4 clustered histone 3 |
| chr9_+_35042213 | 0.04 |
ENST00000378745.3
ENST00000312292.6 |
C9orf131
|
chromosome 9 open reading frame 131 |
| chr4_+_112647059 | 0.04 |
ENST00000511529.1
|
LARP7
|
La ribonucleoprotein 7, transcriptional regulator |
| chr5_+_141177790 | 0.04 |
ENST00000239444.4
ENST00000623995.1 |
PCDHB8
ENSG00000279472.1
|
protocadherin beta 8 novel transcript |
| chr2_+_233917371 | 0.04 |
ENST00000324695.9
ENST00000433712.6 |
TRPM8
|
transient receptor potential cation channel subfamily M member 8 |
| chr18_-_24272179 | 0.04 |
ENST00000399443.7
|
OSBPL1A
|
oxysterol binding protein like 1A |
| chr2_-_99255107 | 0.04 |
ENST00000333017.6
ENST00000626374.2 ENST00000409679.5 ENST00000423306.1 |
LYG2
|
lysozyme g2 |
| chr1_-_45522870 | 0.04 |
ENST00000424390.2
|
PRDX1
|
peroxiredoxin 1 |
| chr4_+_154563003 | 0.04 |
ENST00000302068.9
ENST00000509493.1 |
FGB
|
fibrinogen beta chain |
| chrX_+_85003863 | 0.04 |
ENST00000373173.7
|
APOOL
|
apolipoprotein O like |
| chr16_-_28623330 | 0.04 |
ENST00000677940.1
|
ENSG00000288656.1
|
novel protein |
| chr4_+_168631597 | 0.04 |
ENST00000504519.5
ENST00000512127.5 |
PALLD
|
palladin, cytoskeletal associated protein |
| chr10_-_27240505 | 0.04 |
ENST00000375888.5
ENST00000676732.1 |
ACBD5
|
acyl-CoA binding domain containing 5 |
| chr7_+_134986483 | 0.04 |
ENST00000436302.6
ENST00000435976.6 ENST00000455283.2 |
AGBL3
|
ATP/GTP binding protein like 3 |
| chr12_-_86256267 | 0.04 |
ENST00000620241.4
|
MGAT4C
|
MGAT4 family member C |
| chr13_+_30422487 | 0.04 |
ENST00000638137.1
ENST00000635918.2 |
UBE2L5
|
ubiquitin conjugating enzyme E2 L5 |
| chr2_-_178108339 | 0.04 |
ENST00000358450.8
|
PDE11A
|
phosphodiesterase 11A |
| chr12_-_10130143 | 0.03 |
ENST00000298523.9
ENST00000396484.6 ENST00000310002.4 ENST00000304084.13 |
CLEC7A
|
C-type lectin domain containing 7A |
| chr19_-_11346486 | 0.03 |
ENST00000590482.5
|
TMEM205
|
transmembrane protein 205 |
| chr13_-_83882390 | 0.03 |
ENST00000377084.3
|
SLITRK1
|
SLIT and NTRK like family member 1 |
| chr18_+_616711 | 0.03 |
ENST00000579494.1
|
CLUL1
|
clusterin like 1 |
| chr13_-_70108441 | 0.03 |
ENST00000377844.9
ENST00000545028.2 |
KLHL1
|
kelch like family member 1 |
| chr3_+_41194741 | 0.03 |
ENST00000643541.1
ENST00000426215.5 ENST00000645210.1 ENST00000646381.1 ENST00000405570.6 ENST00000642248.1 ENST00000433400.6 |
CTNNB1
|
catenin beta 1 |
| chr7_+_97732046 | 0.03 |
ENST00000350485.8
ENST00000346867.4 ENST00000319273.10 |
TAC1
|
tachykinin precursor 1 |
| chr12_-_10130082 | 0.02 |
ENST00000533022.5
|
CLEC7A
|
C-type lectin domain containing 7A |
| chr6_+_152697934 | 0.02 |
ENST00000532295.1
|
MYCT1
|
MYC target 1 |
| chr18_+_616672 | 0.02 |
ENST00000338387.11
|
CLUL1
|
clusterin like 1 |
| chr7_+_92057602 | 0.02 |
ENST00000491695.2
|
AKAP9
|
A-kinase anchoring protein 9 |
| chr17_+_46511511 | 0.02 |
ENST00000576629.5
|
LRRC37A2
|
leucine rich repeat containing 37 member A2 |
| chr12_-_10130241 | 0.02 |
ENST00000353231.9
ENST00000525605.1 |
CLEC7A
|
C-type lectin domain containing 7A |
| chr4_-_86357722 | 0.02 |
ENST00000641341.1
ENST00000642038.1 ENST00000641116.1 ENST00000641767.1 ENST00000639242.1 ENST00000638313.1 |
MAPK10
|
mitogen-activated protein kinase 10 |
| chr1_+_196888014 | 0.02 |
ENST00000367416.6
ENST00000608469.6 ENST00000251424.8 ENST00000367418.2 |
CFHR4
|
complement factor H related 4 |
| chr5_+_127649018 | 0.02 |
ENST00000379445.7
|
CTXN3
|
cortexin 3 |
| chr1_-_56819365 | 0.02 |
ENST00000343433.7
|
FYB2
|
FYN binding protein 2 |
| chr19_-_6279921 | 0.02 |
ENST00000252674.9
|
MLLT1
|
MLLT1 super elongation complex subunit |
| chr4_-_103019634 | 0.02 |
ENST00000510559.1
ENST00000296422.12 ENST00000394789.7 |
SLC9B1
|
solute carrier family 9 member B1 |
| chr10_+_92591733 | 0.02 |
ENST00000676647.1
|
KIF11
|
kinesin family member 11 |
| chr7_-_123199960 | 0.02 |
ENST00000194130.7
|
SLC13A1
|
solute carrier family 13 member 1 |
| chr17_+_7407838 | 0.02 |
ENST00000302926.7
|
NLGN2
|
neuroligin 2 |
| chrM_+_10055 | 0.02 |
ENST00000361227.2
|
MT-ND3
|
mitochondrially encoded NADH:ubiquinone oxidoreductase core subunit 3 |
| chr1_-_150235995 | 0.02 |
ENST00000436748.6
|
ANP32E
|
acidic nuclear phosphoprotein 32 family member E |
| chr7_+_30145789 | 0.01 |
ENST00000324489.5
|
MTURN
|
maturin, neural progenitor differentiation regulator homolog |
| chr7_+_107583919 | 0.01 |
ENST00000491150.5
|
BCAP29
|
B cell receptor associated protein 29 |
| chr9_+_122371014 | 0.01 |
ENST00000362012.7
|
PTGS1
|
prostaglandin-endoperoxide synthase 1 |
| chr18_-_5197241 | 0.01 |
ENST00000434239.4
|
AKAIN1
|
A-kinase anchor inhibitor 1 |
| chr9_-_21482313 | 0.00 |
ENST00000448696.4
|
IFNE
|
interferon epsilon |
| chr3_-_47475811 | 0.00 |
ENST00000265565.10
ENST00000428413.5 |
SCAP
|
SREBF chaperone |
| chr3_-_191282383 | 0.00 |
ENST00000427544.6
|
UTS2B
|
urotensin 2B |
| chr1_-_23799533 | 0.00 |
ENST00000429356.5
|
GALE
|
UDP-galactose-4-epimerase |
| chr17_-_66229380 | 0.00 |
ENST00000205948.11
|
APOH
|
apolipoprotein H |
| chr2_+_54558348 | 0.00 |
ENST00000333896.5
|
SPTBN1
|
spectrin beta, non-erythrocytic 1 |
| chr8_-_25424260 | 0.00 |
ENST00000421054.7
|
GNRH1
|
gonadotropin releasing hormone 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.4 | GO:0060005 | vestibular reflex(GO:0060005) |
| 0.1 | 0.2 | GO:1901675 | negative regulation of histone H3-K27 acetylation(GO:1901675) |
| 0.1 | 0.6 | GO:0034285 | response to sucrose(GO:0009744) response to disaccharide(GO:0034285) |
| 0.1 | 0.4 | GO:0061624 | fructose catabolic process(GO:0006001) fructose catabolic process to hydroxyacetone phosphate and glyceraldehyde-3-phosphate(GO:0061624) |
| 0.0 | 0.1 | GO:0002416 | IgG immunoglobulin transcytosis in epithelial cells mediated by FcRn immunoglobulin receptor(GO:0002416) |
| 0.0 | 0.1 | GO:0010730 | negative regulation of hydrogen peroxide biosynthetic process(GO:0010730) |
| 0.0 | 0.1 | GO:0048560 | establishment of anatomical structure orientation(GO:0048560) |
| 0.0 | 0.2 | GO:0003185 | primary heart field specification(GO:0003138) sinoatrial valve development(GO:0003172) sinoatrial valve morphogenesis(GO:0003185) |
| 0.0 | 0.1 | GO:0070105 | positive regulation of interleukin-6-mediated signaling pathway(GO:0070105) |
| 0.0 | 0.2 | GO:2000563 | positive regulation of CD4-positive, alpha-beta T cell proliferation(GO:2000563) |
| 0.0 | 0.2 | GO:0046726 | positive regulation by virus of viral protein levels in host cell(GO:0046726) |
| 0.0 | 1.1 | GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0000083) |
| 0.0 | 0.1 | GO:0032218 | riboflavin transport(GO:0032218) |
| 0.0 | 0.3 | GO:0060159 | regulation of dopamine receptor signaling pathway(GO:0060159) |
| 0.0 | 0.1 | GO:0021590 | cerebellum maturation(GO:0021590) cerebellar cortex maturation(GO:0021699) |
| 0.0 | 0.1 | GO:0060770 | negative regulation of epithelial cell proliferation involved in prostate gland development(GO:0060770) cellular response to cobalt ion(GO:0071279) |
| 0.0 | 0.8 | GO:0043486 | histone exchange(GO:0043486) |
| 0.0 | 0.1 | GO:1902572 | negative regulation of serine-type endopeptidase activity(GO:1900004) negative regulation of serine-type peptidase activity(GO:1902572) |
| 0.0 | 0.1 | GO:0040030 | nucleosome positioning(GO:0016584) regulation of molecular function, epigenetic(GO:0040030) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.4 | GO:0032426 | stereocilium tip(GO:0032426) |
| 0.0 | 0.8 | GO:0000812 | Swr1 complex(GO:0000812) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.4 | GO:0022833 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.2 | 0.6 | GO:0004948 | calcitonin receptor activity(GO:0004948) |
| 0.1 | 0.7 | GO:0042806 | fucose binding(GO:0042806) |
| 0.1 | 0.3 | GO:0050528 | acyloxyacyl hydrolase activity(GO:0050528) |
| 0.1 | 0.4 | GO:0018479 | benzaldehyde dehydrogenase (NAD+) activity(GO:0018479) |
| 0.1 | 0.2 | GO:0016524 | latrotoxin receptor activity(GO:0016524) |
| 0.0 | 0.1 | GO:0019770 | IgG receptor activity(GO:0019770) |
| 0.0 | 0.2 | GO:0102344 | 3-hydroxy-behenoyl-CoA dehydratase activity(GO:0102344) 3-hydroxy-lignoceroyl-CoA dehydratase activity(GO:0102345) |
| 0.0 | 0.1 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) |
| 0.0 | 0.2 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.0 | 0.2 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.0 | 0.4 | GO:0001011 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.0 | 0.1 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.0 | 0.1 | GO:0004991 | parathyroid hormone receptor activity(GO:0004991) |
| 0.0 | 0.1 | GO:0043035 | chromatin insulator sequence binding(GO:0043035) |
| 0.0 | 0.1 | GO:0032217 | riboflavin transporter activity(GO:0032217) |
| 0.0 | 0.1 | GO:0060002 | plus-end directed microfilament motor activity(GO:0060002) |
| 0.0 | 0.2 | GO:0003680 | AT DNA binding(GO:0003680) |
| 0.0 | 0.8 | GO:0019212 | phosphatase inhibitor activity(GO:0019212) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.0 | 0.4 | REACTOME INHIBITION OF REPLICATION INITIATION OF DAMAGED DNA BY RB1 E2F1 | Genes involved in Inhibition of replication initiation of damaged DNA by RB1/E2F1 |