Project
Inflammatory response time course, HUVEC (Wada, 2009)
Navigation
Downloads
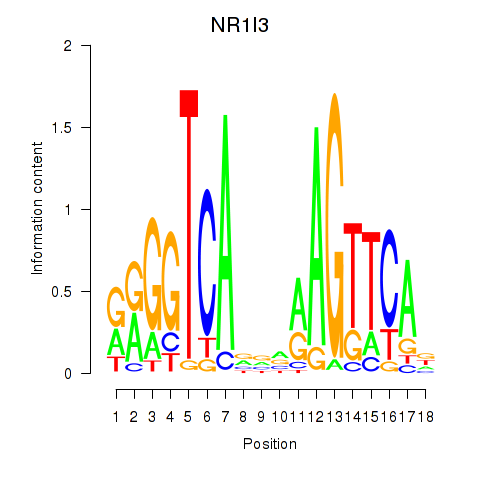
Results for NR1I3
Z-value: 0.40
Transcription factors associated with NR1I3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
NR1I3
|
ENSG00000143257.12 | NR1I3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NR1I3 | hg38_v1_chr1_-_161238085_161238111 | -0.06 | 7.9e-01 | Click! |
Activity profile of NR1I3 motif
Sorted Z-values of NR1I3 motif
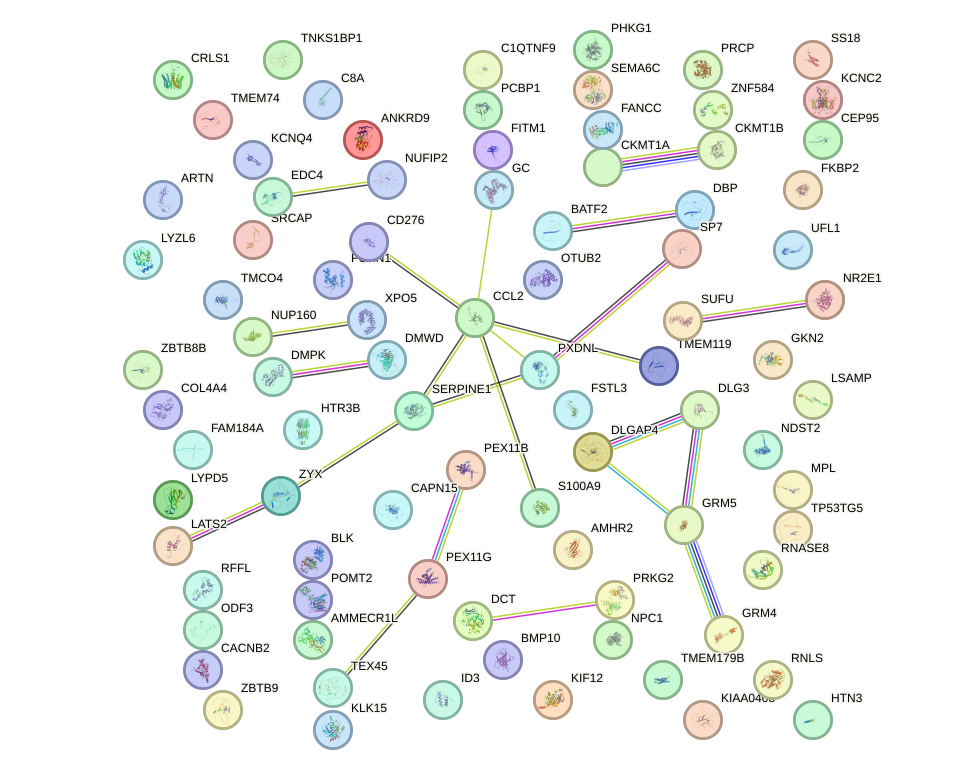
Network of associatons between targets according to the STRING database.
First level regulatory network of NR1I3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_-_205321737 | 0.53 |
ENST00000367157.6
|
NUAK2
|
NUAK family kinase 2 |
| chr17_+_34255274 | 0.51 |
ENST00000580907.5
ENST00000225831.4 |
CCL2
|
C-C motif chemokine ligand 2 |
| chr19_+_676385 | 0.51 |
ENST00000166139.9
|
FSTL3
|
follistatin like 3 |
| chr19_-_7497447 | 0.46 |
ENST00000593942.5
|
PEX11G
|
peroxisomal biogenesis factor 11 gamma |
| chr14_+_22508602 | 0.37 |
ENST00000390504.1
|
TRAJ33
|
T cell receptor alpha joining 33 |
| chr11_+_64241600 | 0.37 |
ENST00000535135.7
ENST00000652094.1 |
FKBP2
|
FKBP prolyl isomerase 2 |
| chr2_-_39120383 | 0.36 |
ENST00000395038.6
|
SOS1
|
SOS Ras/Rac guanine nucleotide exchange factor 1 |
| chr17_-_8295342 | 0.35 |
ENST00000579192.5
ENST00000577745.2 |
SLC25A35
|
solute carrier family 25 member 35 |
| chr4_+_70028452 | 0.34 |
ENST00000530128.5
ENST00000381057.3 ENST00000673563.1 |
HTN3
|
histatin 3 |
| chr12_+_53423849 | 0.34 |
ENST00000257863.9
ENST00000550311.5 ENST00000379791.7 |
AMHR2
|
anti-Mullerian hormone receptor type 2 |
| chr18_-_23586422 | 0.33 |
ENST00000269228.10
|
NPC1
|
NPC intracellular cholesterol transporter 1 |
| chr11_-_82900406 | 0.33 |
ENST00000313010.8
ENST00000393399.6 ENST00000680437.1 |
PRCP
|
prolylcarboxypeptidase |
| chr16_+_2988256 | 0.32 |
ENST00000573315.2
|
LINC00514
|
long intergenic non-protein coding RNA 514 |
| chr2_-_68952880 | 0.31 |
ENST00000481498.1
ENST00000328895.9 |
GKN2
|
gastrokine 2 |
| chr3_-_116444983 | 0.29 |
ENST00000333617.8
|
LSAMP
|
limbic system associated membrane protein |
| chr11_+_197303 | 0.28 |
ENST00000342593.6
|
ODF3
|
outer dense fiber of sperm tails 3 |
| chr19_-_45792755 | 0.28 |
ENST00000377735.7
ENST00000270223.7 |
DMWD
|
DM1 locus, WD repeat containing |
| chr16_+_30699155 | 0.27 |
ENST00000262518.9
|
SRCAP
|
Snf2 related CREBBP activator protein |
| chr7_+_143381907 | 0.26 |
ENST00000392910.6
|
ZYX
|
zyxin |
| chr2_-_68871382 | 0.26 |
ENST00000295379.2
|
BMP10
|
bone morphogenetic protein 10 |
| chr17_-_29294141 | 0.25 |
ENST00000225388.9
|
NUFIP2
|
nuclear FMR1 interacting protein 2 |
| chr15_+_84980440 | 0.25 |
ENST00000310298.8
ENST00000557957.5 |
PDE8A
|
phosphodiesterase 8A |
| chr7_+_143381561 | 0.24 |
ENST00000354434.8
|
ZYX
|
zyxin |
| chr6_+_108166015 | 0.24 |
ENST00000368986.9
|
NR2E1
|
nuclear receptor subfamily 2 group E member 1 |
| chr6_-_43528867 | 0.22 |
ENST00000455285.2
|
XPO5
|
exportin 5 |
| chr11_+_62789124 | 0.22 |
ENST00000526546.1
|
TMEM179B
|
transmembrane protein 179B |
| chr10_-_72088972 | 0.21 |
ENST00000317376.8
ENST00000412663.5 |
SPOCK2
|
SPARC (osteonectin), cwcv and kazal like domains proteoglycan 2 |
| chr14_+_21057822 | 0.21 |
ENST00000308227.2
|
RNASE8
|
ribonuclease A family member 8 |
| chr2_-_227164194 | 0.20 |
ENST00000396625.5
|
COL4A4
|
collagen type IV alpha 4 chain |
| chr11_-_57322197 | 0.19 |
ENST00000532437.1
|
TNKS1BP1
|
tankyrase 1 binding protein 1 |
| chr7_-_56092932 | 0.19 |
ENST00000446428.5
ENST00000432123.5 ENST00000297373.7 |
PHKG1
|
phosphorylase kinase catalytic subunit gamma 1 |
| chr7_+_101127095 | 0.18 |
ENST00000223095.5
|
SERPINE1
|
serpin family E member 1 |
| chr7_-_56092974 | 0.18 |
ENST00000452681.6
ENST00000537360.5 |
PHKG1
|
phosphorylase kinase catalytic subunit gamma 1 |
| chr19_-_43802438 | 0.18 |
ENST00000377950.8
|
LYPD5
|
LY6/PLAUR domain containing 5 |
| chr16_+_67873036 | 0.17 |
ENST00000358933.10
|
EDC4
|
enhancer of mRNA decapping 4 |
| chr20_+_36461303 | 0.16 |
ENST00000475894.5
|
DLGAP4
|
DLG associated protein 4 |
| chr4_-_81215185 | 0.15 |
ENST00000264399.6
|
PRKG2
|
protein kinase cGMP-dependent 2 |
| chr4_-_81214960 | 0.15 |
ENST00000395578.3
ENST00000628926.1 |
PRKG2
|
protein kinase cGMP-dependent 2 |
| chr6_+_33454543 | 0.15 |
ENST00000621915.1
ENST00000395064.3 |
ZBTB9
|
zinc finger and BTB domain containing 9 |
| chr9_+_34179005 | 0.14 |
ENST00000625521.2
ENST00000379186.8 ENST00000297661.9 ENST00000626262.2 |
UBAP1
|
ubiquitin associated protein 1 |
| chrX_-_111412162 | 0.14 |
ENST00000637570.1
ENST00000356220.8 ENST00000636035.2 ENST00000637453.1 ENST00000635795.1 |
DCX
|
doublecortin |
| chr10_-_73808616 | 0.14 |
ENST00000299641.8
|
NDST2
|
N-deacetylase and N-sulfotransferase 2 |
| chr12_-_108598036 | 0.14 |
ENST00000392806.4
ENST00000547567.1 |
TMEM119
|
transmembrane protein 119 |
| chr13_-_21061492 | 0.14 |
ENST00000382592.5
|
LATS2
|
large tumor suppressor kinase 2 |
| chr12_-_53335737 | 0.14 |
ENST00000303846.3
|
SP7
|
Sp7 transcription factor |
| chr11_-_47848539 | 0.13 |
ENST00000526870.1
|
NUP160
|
nucleoporin 160 |
| chr5_-_136193143 | 0.13 |
ENST00000607574.2
|
SMIM32
|
small integral membrane protein 32 |
| chr1_+_40817190 | 0.13 |
ENST00000443478.3
|
KCNQ4
|
potassium voltage-gated channel subfamily Q member 4 |
| chr14_+_94026314 | 0.13 |
ENST00000203664.10
ENST00000553723.1 |
OTUB2
|
OTU deubiquitinase, ubiquitin aldehyde binding 2 |
| chr19_-_45782388 | 0.13 |
ENST00000458663.6
|
DMPK
|
DM1 protein kinase |
| chr10_+_18400562 | 0.12 |
ENST00000377315.5
ENST00000650685.1 |
CACNB2
|
calcium voltage-gated channel auxiliary subunit beta 2 |
| chr8_-_51809414 | 0.12 |
ENST00000356297.5
|
PXDNL
|
peroxidasin like |
| chr14_-_77320813 | 0.11 |
ENST00000682467.1
|
POMT2
|
protein O-mannosyltransferase 2 |
| chr14_-_102509713 | 0.10 |
ENST00000286918.9
|
ANKRD9
|
ankyrin repeat domain 9 |
| chr14_-_77320855 | 0.10 |
ENST00000556394.2
ENST00000261534.9 |
POMT2
|
protein O-mannosyltransferase 2 |
| chr8_-_108787563 | 0.10 |
ENST00000297459.4
|
TMEM74
|
transmembrane protein 74 |
| chr20_+_6007245 | 0.10 |
ENST00000378868.4
|
CRLS1
|
cardiolipin synthase 1 |
| chr7_-_99784175 | 0.10 |
ENST00000651514.1
ENST00000336411.7 ENST00000415003.1 ENST00000354593.6 |
CYP3A4
|
cytochrome P450 family 3 subfamily A member 4 |
| chr3_-_116445458 | 0.10 |
ENST00000490035.7
|
LSAMP
|
limbic system associated membrane protein |
| chr10_+_102503956 | 0.10 |
ENST00000369902.8
|
SUFU
|
SUFU negative regulator of hedgehog signaling |
| chr19_-_15449920 | 0.10 |
ENST00000263381.12
ENST00000643092.1 ENST00000673675.1 |
WIZ
|
WIZ zinc finger |
| chr6_-_119149124 | 0.09 |
ENST00000368475.8
|
FAM184A
|
family with sequence similarity 184 member A |
| chr1_-_23559490 | 0.09 |
ENST00000374561.6
|
ID3
|
inhibitor of DNA binding 3, HLH protein |
| chr19_+_58408735 | 0.09 |
ENST00000599238.5
ENST00000322834.7 |
ZNF584
|
zinc finger protein 584 |
| chr19_+_58408633 | 0.09 |
ENST00000306910.9
ENST00000598901.1 ENST00000593920.5 ENST00000596281.1 |
ZNF584
|
zinc finger protein 584 |
| chr11_+_113908983 | 0.09 |
ENST00000537778.5
|
HTR3B
|
5-hydroxytryptamine receptor 3B |
| chr15_+_73684731 | 0.09 |
ENST00000560995.5
|
CD276
|
CD276 molecule |
| chr7_-_99784248 | 0.09 |
ENST00000652018.1
|
CYP3A4
|
cytochrome P450 family 3 subfamily A member 4 |
| chr6_-_127459364 | 0.09 |
ENST00000487331.2
ENST00000483725.8 |
KIAA0408
|
KIAA0408 |
| chr2_+_70087468 | 0.08 |
ENST00000303577.7
|
PCBP1
|
poly(rC) binding protein 1 |
| chr9_-_95317671 | 0.08 |
ENST00000490972.7
ENST00000647778.1 ENST00000649611.1 ENST00000289081.8 |
FANCC
|
FA complementation group C |
| chr8_+_11494367 | 0.08 |
ENST00000259089.9
ENST00000529894.1 |
BLK
|
BLK proto-oncogene, Src family tyrosine kinase |
| chr17_-_35063648 | 0.07 |
ENST00000394597.7
|
RFFL
|
ring finger and FYVE like domain containing E3 ubiquitin protein ligase |
| chr1_-_151146611 | 0.07 |
ENST00000341697.7
ENST00000368914.8 |
SEMA6C
|
semaphorin 6C |
| chr10_-_88583304 | 0.07 |
ENST00000331772.9
|
RNLS
|
renalase, FAD dependent amine oxidase |
| chr14_+_22281097 | 0.07 |
ENST00000390465.2
|
TRAV38-2DV8
|
T cell receptor alpha variable 38-2/delta variable 8 |
| chr11_-_47848313 | 0.06 |
ENST00000530326.5
ENST00000528071.5 |
NUP160
|
nucleoporin 160 |
| chr19_-_50833187 | 0.06 |
ENST00000598673.1
|
KLK15
|
kallikrein related peptidase 15 |
| chr9_-_114099275 | 0.06 |
ENST00000468460.2
ENST00000640217.2 |
KIF12
|
kinesin family member 12 |
| chr6_-_31541937 | 0.06 |
ENST00000456662.5
ENST00000431908.5 ENST00000456976.5 ENST00000428450.5 ENST00000418897.5 ENST00000396172.6 ENST00000419020.1 ENST00000428098.5 |
DDX39B
|
DExD-box helicase 39B |
| chr1_+_56854764 | 0.06 |
ENST00000361249.4
|
C8A
|
complement C8 alpha chain |
| chr16_+_527698 | 0.06 |
ENST00000219611.7
ENST00000562370.5 ENST00000568988.5 |
CAPN15
|
calpain 15 |
| chr19_+_54201122 | 0.06 |
ENST00000391753.6
ENST00000441429.1 |
RPS9
|
ribosomal protein S9 |
| chr13_+_24309565 | 0.06 |
ENST00000332018.4
|
C1QTNF9
|
C1q and TNF related 9 |
| chr1_-_151146643 | 0.06 |
ENST00000613223.1
|
SEMA6C
|
semaphorin 6C |
| chr17_+_64506953 | 0.06 |
ENST00000580188.1
ENST00000556440.7 ENST00000581056.5 |
CEP95
|
centrosomal protein 95 |
| chr1_+_32465046 | 0.06 |
ENST00000609129.2
|
ZBTB8B
|
zinc finger and BTB domain containing 8B |
| chr17_+_28506320 | 0.06 |
ENST00000579795.6
|
FOXN1
|
forkhead box N1 |
| chrX_+_70455093 | 0.06 |
ENST00000542398.1
|
DLG3
|
discs large MAGUK scaffold protein 3 |
| chr18_-_26090584 | 0.06 |
ENST00000415083.7
|
SS18
|
SS18 subunit of BAF chromatin remodeling complex |
| chr13_-_94479671 | 0.05 |
ENST00000377028.10
ENST00000446125.1 |
DCT
|
dopachrome tautomerase |
| chr11_-_64996963 | 0.05 |
ENST00000301887.9
ENST00000534177.1 |
BATF2
|
basic leucine zipper ATF-like transcription factor 2 |
| chr20_-_45378305 | 0.05 |
ENST00000372726.5
|
TP53TG5
|
TP53 target 5 |
| chr17_-_35943707 | 0.05 |
ENST00000615905.5
|
LYZL6
|
lysozyme like 6 |
| chr17_-_35943662 | 0.04 |
ENST00000618542.4
|
LYZL6
|
lysozyme like 6 |
| chr16_-_66934144 | 0.04 |
ENST00000568572.5
|
CIAO2B
|
cytosolic iron-sulfur assembly component 2B |
| chr2_-_127884860 | 0.04 |
ENST00000393001.1
|
AMMECR1L
|
AMMECR1 like |
| chr14_-_102509798 | 0.04 |
ENST00000560748.5
|
ANKRD9
|
ankyrin repeat domain 9 |
| chr14_-_77320741 | 0.04 |
ENST00000682795.1
ENST00000682247.1 |
POMT2
|
protein O-mannosyltransferase 2 |
| chr5_-_39270623 | 0.04 |
ENST00000512138.1
ENST00000646045.2 |
FYB1
|
FYN binding protein 1 |
| chr14_+_24130659 | 0.03 |
ENST00000267426.6
|
FITM1
|
fat storage inducing transmembrane protein 1 |
| chr19_+_7497544 | 0.03 |
ENST00000361664.7
|
TEX45
|
testis expressed 45 |
| chr11_-_47848467 | 0.03 |
ENST00000378460.6
|
NUP160
|
nucleoporin 160 |
| chr11_-_89065969 | 0.03 |
ENST00000305447.5
|
GRM5
|
glutamate metabotropic receptor 5 |
| chr15_+_43593054 | 0.03 |
ENST00000453782.5
ENST00000300283.10 ENST00000437924.5 |
CKMT1B
|
creatine kinase, mitochondrial 1B |
| chr12_-_75207998 | 0.03 |
ENST00000550433.5
ENST00000548513.5 |
KCNC2
|
potassium voltage-gated channel subfamily C member 2 |
| chr18_-_26091175 | 0.02 |
ENST00000579061.5
ENST00000542420.6 |
SS18
|
SS18 subunit of BAF chromatin remodeling complex |
| chr6_+_96521796 | 0.02 |
ENST00000369278.5
|
UFL1
|
UFM1 specific ligase 1 |
| chr1_+_43337803 | 0.02 |
ENST00000372470.9
ENST00000413998.7 |
MPL
|
MPL proto-oncogene, thrombopoietin receptor |
| chr1_+_153357846 | 0.02 |
ENST00000368738.4
|
S100A9
|
S100 calcium binding protein A9 |
| chr1_+_43935807 | 0.02 |
ENST00000438616.3
|
ARTN
|
artemin |
| chr19_+_54200849 | 0.02 |
ENST00000626547.2
ENST00000302907.9 ENST00000391752.5 ENST00000402367.5 ENST00000391751.7 |
RPS9
|
ribosomal protein S9 |
| chr1_-_19799872 | 0.02 |
ENST00000294543.11
|
TMCO4
|
transmembrane and coiled-coil domains 4 |
| chr1_-_145918485 | 0.02 |
ENST00000537888.1
|
PEX11B
|
peroxisomal biogenesis factor 11 beta |
| chr19_-_48637338 | 0.02 |
ENST00000601104.1
ENST00000222122.10 |
DBP
|
D-box binding PAR bZIP transcription factor |
| chr6_-_34146080 | 0.01 |
ENST00000538487.7
ENST00000374181.8 |
GRM4
|
glutamate metabotropic receptor 4 |
| chr16_-_66934362 | 0.01 |
ENST00000567511.1
ENST00000422424.7 |
CIAO2B
|
cytosolic iron-sulfur assembly component 2B |
| chr9_+_131125973 | 0.01 |
ENST00000651639.1
ENST00000451030.5 ENST00000531584.1 |
NUP214
|
nucleoporin 214 |
| chr2_+_219597838 | 0.01 |
ENST00000456909.6
|
STK11IP
|
serine/threonine kinase 11 interacting protein |
| chr1_+_160127672 | 0.01 |
ENST00000447527.1
|
ATP1A2
|
ATPase Na+/K+ transporting subunit alpha 2 |
| chr7_-_73738792 | 0.01 |
ENST00000222800.8
ENST00000458339.6 |
ABHD11
|
abhydrolase domain containing 11 |
| chr1_-_145918685 | 0.01 |
ENST00000369306.8
|
PEX11B
|
peroxisomal biogenesis factor 11 beta |
| chr4_+_37244735 | 0.01 |
ENST00000309447.6
|
NWD2
|
NACHT and WD repeat domain containing 2 |
| chr1_-_45542726 | 0.00 |
ENST00000676549.1
|
PRDX1
|
peroxiredoxin 1 |
| chr1_+_27392612 | 0.00 |
ENST00000374024.4
|
GPR3
|
G protein-coupled receptor 3 |
| chr9_+_121286586 | 0.00 |
ENST00000545652.6
|
GSN
|
gelsolin |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:2000502 | negative regulation of natural killer cell chemotaxis(GO:2000502) |
| 0.1 | 0.2 | GO:0060164 | regulation of timing of neuron differentiation(GO:0060164) |
| 0.1 | 0.3 | GO:2001226 | negative regulation of chloride transport(GO:2001226) |
| 0.1 | 0.2 | GO:0035281 | pre-miRNA export from nucleus(GO:0035281) |
| 0.1 | 0.4 | GO:1904693 | midbrain morphogenesis(GO:1904693) |
| 0.1 | 0.3 | GO:1990262 | regulation of anti-Mullerian hormone signaling pathway(GO:1902612) negative regulation of anti-Mullerian hormone signaling pathway(GO:1902613) anti-Mullerian hormone signaling pathway(GO:1990262) |
| 0.1 | 0.2 | GO:2000097 | chronological cell aging(GO:0001300) regulation of smooth muscle cell-matrix adhesion(GO:2000097) |
| 0.1 | 0.5 | GO:0044375 | regulation of peroxisome size(GO:0044375) |
| 0.0 | 0.1 | GO:0001827 | inner cell mass cell fate commitment(GO:0001827) inner cell mass cellular morphogenesis(GO:0001828) |
| 0.0 | 0.2 | GO:0009822 | alkaloid catabolic process(GO:0009822) |
| 0.0 | 0.3 | GO:0002254 | kinin cascade(GO:0002254) plasma kallikrein-kinin cascade(GO:0002353) |
| 0.0 | 0.2 | GO:0061760 | antifungal innate immune response(GO:0061760) |
| 0.0 | 0.3 | GO:0060298 | positive regulation of sarcomere organization(GO:0060298) |
| 0.0 | 0.3 | GO:1904098 | regulation of protein O-linked glycosylation(GO:1904098) positive regulation of protein O-linked glycosylation(GO:1904100) |
| 0.0 | 0.3 | GO:2000189 | positive regulation of cholesterol homeostasis(GO:2000189) |
| 0.0 | 0.5 | GO:0032926 | negative regulation of activin receptor signaling pathway(GO:0032926) |
| 0.0 | 0.1 | GO:0045903 | positive regulation of translational fidelity(GO:0045903) |
| 0.0 | 0.1 | GO:0097534 | lymphoid lineage cell migration(GO:0097534) lymphoid lineage cell migration into thymus(GO:0097535) regulation of positive thymic T cell selection(GO:1902232) |
| 0.0 | 0.1 | GO:0021775 | smoothened signaling pathway involved in ventral spinal cord interneuron specification(GO:0021775) smoothened signaling pathway involved in spinal cord motor neuron cell fate specification(GO:0021776) |
| 0.0 | 0.2 | GO:0031087 | deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
| 0.0 | 0.1 | GO:2001271 | negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 0.0 | 0.4 | GO:0035641 | locomotory exploration behavior(GO:0035641) |
| 0.0 | 0.1 | GO:1904879 | positive regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1904879) |
| 0.0 | 0.2 | GO:0032836 | glomerular basement membrane development(GO:0032836) |
| 0.0 | 0.1 | GO:0097068 | response to thyroxine(GO:0097068) response to L-phenylalanine derivative(GO:1904386) |
| 0.0 | 0.1 | GO:0030202 | heparin metabolic process(GO:0030202) heparin biosynthetic process(GO:0030210) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0042565 | RNA nuclear export complex(GO:0042565) |
| 0.0 | 0.4 | GO:0005964 | phosphorylase kinase complex(GO:0005964) |
| 0.0 | 0.2 | GO:0005587 | collagen type IV trimer(GO:0005587) |
| 0.0 | 0.2 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.0 | 0.3 | GO:0001520 | outer dense fiber(GO:0001520) |
| 0.0 | 0.5 | GO:0031231 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 0.0 | 0.2 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.1 | 0.3 | GO:0005026 | transforming growth factor beta receptor activity, type II(GO:0005026) |
| 0.1 | 0.2 | GO:0090631 | pre-miRNA transporter activity(GO:0090631) |
| 0.1 | 0.3 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 0.0 | 0.2 | GO:0030343 | vitamin D3 25-hydroxylase activity(GO:0030343) vitamin D 25-hydroxylase activity(GO:0070643) |
| 0.0 | 0.4 | GO:0004689 | phosphorylase kinase activity(GO:0004689) |
| 0.0 | 0.3 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 0.0 | 0.1 | GO:0050119 | N-acetylglucosamine deacetylase activity(GO:0050119) |
| 0.0 | 0.2 | GO:0004522 | ribonuclease A activity(GO:0004522) |
| 0.0 | 0.1 | GO:0070984 | SET domain binding(GO:0070984) |
| 0.0 | 0.5 | GO:0048185 | activin binding(GO:0048185) |
| 0.0 | 0.2 | GO:0071532 | ankyrin repeat binding(GO:0071532) |
| 0.0 | 0.3 | GO:0031433 | telethonin binding(GO:0031433) |
| 0.0 | 0.1 | GO:0022850 | serotonin-gated cation channel activity(GO:0022850) |
| 0.0 | 0.3 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.0 | 0.1 | GO:0019784 | NEDD8-specific protease activity(GO:0019784) |
| 0.0 | 0.4 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.0 | 0.1 | GO:1990932 | 5.8S rRNA binding(GO:1990932) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.0 | 0.4 | SA TRKA RECEPTOR | The TrkA receptor binds nerve growth factor to activate MAP kinase pathways and promote cell growth. |
| 0.0 | 0.5 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.0 | 0.4 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.0 | 0.3 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.0 | 0.3 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.0 | 0.5 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |