Project
Inflammatory response time course, HUVEC (Wada, 2009)
Navigation
Downloads
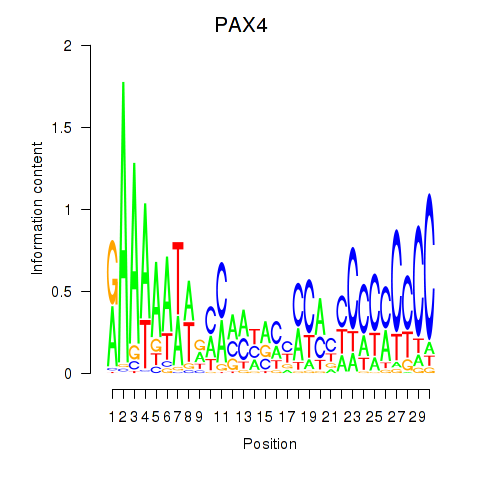
Results for PAX4
Z-value: 0.77
Transcription factors associated with PAX4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
PAX4
|
ENSG00000106331.17 | PAX4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| PAX4 | hg38_v1_chr7_-_127618137_127618145 | 0.12 | 5.8e-01 | Click! |
Activity profile of PAX4 motif
Sorted Z-values of PAX4 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of PAX4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_+_102355881 | 4.36 |
ENST00000409599.5
|
IL18R1
|
interleukin 18 receptor 1 |
| chr2_+_102355750 | 4.36 |
ENST00000233957.7
|
IL18R1
|
interleukin 18 receptor 1 |
| chr9_-_120926752 | 3.47 |
ENST00000373887.8
|
TRAF1
|
TNF receptor associated factor 1 |
| chr9_-_120914549 | 2.97 |
ENST00000546084.5
|
TRAF1
|
TNF receptor associated factor 1 |
| chr8_+_32548303 | 1.96 |
ENST00000650967.1
|
NRG1
|
neuregulin 1 |
| chr3_+_112086335 | 1.86 |
ENST00000431717.6
ENST00000480282.5 |
C3orf52
|
chromosome 3 open reading frame 52 |
| chr14_-_91244669 | 1.85 |
ENST00000650645.1
|
GPR68
|
G protein-coupled receptor 68 |
| chr8_+_32548210 | 1.74 |
ENST00000523079.5
ENST00000650919.1 |
NRG1
|
neuregulin 1 |
| chr3_-_123620496 | 1.70 |
ENST00000578202.1
|
MYLK
|
myosin light chain kinase |
| chr3_+_112086364 | 1.66 |
ENST00000264848.10
|
C3orf52
|
chromosome 3 open reading frame 52 |
| chr3_-_79767987 | 1.66 |
ENST00000464233.6
|
ROBO1
|
roundabout guidance receptor 1 |
| chr3_-_123620571 | 1.65 |
ENST00000583087.5
|
MYLK
|
myosin light chain kinase |
| chr7_-_22220226 | 1.60 |
ENST00000420196.5
|
RAPGEF5
|
Rap guanine nucleotide exchange factor 5 |
| chr16_-_28610032 | 1.59 |
ENST00000567512.1
|
SULT1A1
|
sulfotransferase family 1A member 1 |
| chr6_+_31586124 | 1.49 |
ENST00000418507.6
ENST00000376096.5 ENST00000376099.5 ENST00000376110.7 |
LST1
|
leukocyte specific transcript 1 |
| chr6_-_32838727 | 1.46 |
ENST00000652259.1
ENST00000374897.4 ENST00000620123.4 ENST00000452392.2 |
TAP2
ENSG00000250264.1
|
transporter 2, ATP binding cassette subfamily B member novel protein, TAP2-HLA-DOB readthrough |
| chr6_+_31586269 | 1.28 |
ENST00000438075.7
|
LST1
|
leukocyte specific transcript 1 |
| chr12_+_78036248 | 1.11 |
ENST00000644176.1
|
NAV3
|
neuron navigator 3 |
| chr10_+_6144883 | 0.98 |
ENST00000379789.8
|
PFKFB3
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 3 |
| chr16_+_50696999 | 0.98 |
ENST00000300589.6
|
NOD2
|
nucleotide binding oligomerization domain containing 2 |
| chr6_+_31586680 | 0.97 |
ENST00000339530.8
|
LST1
|
leukocyte specific transcript 1 |
| chr7_+_26152188 | 0.93 |
ENST00000056233.4
|
NFE2L3
|
nuclear factor, erythroid 2 like 3 |
| chr4_-_112285892 | 0.91 |
ENST00000361717.4
|
TIFA
|
TRAF interacting protein with forkhead associated domain |
| chr7_-_16833411 | 0.90 |
ENST00000412973.1
|
AGR2
|
anterior gradient 2, protein disulphide isomerase family member |
| chr5_-_143403297 | 0.86 |
ENST00000415690.6
|
NR3C1
|
nuclear receptor subfamily 3 group C member 1 |
| chr6_-_25042003 | 0.86 |
ENST00000510784.8
|
RIPOR2
|
RHO family interacting cell polarization regulator 2 |
| chr3_-_71583713 | 0.85 |
ENST00000649528.3
ENST00000471386.3 ENST00000493089.7 |
FOXP1
|
forkhead box P1 |
| chr21_+_42513834 | 0.82 |
ENST00000352133.3
|
SLC37A1
|
solute carrier family 37 member 1 |
| chr1_-_204494752 | 0.81 |
ENST00000684373.1
|
PIK3C2B
|
phosphatidylinositol-4-phosphate 3-kinase catalytic subunit type 2 beta |
| chr2_+_202073282 | 0.81 |
ENST00000459709.5
|
KIAA2012
|
KIAA2012 |
| chr2_+_202073249 | 0.80 |
ENST00000498697.3
|
KIAA2012
|
KIAA2012 |
| chr8_+_32548267 | 0.80 |
ENST00000356819.7
|
NRG1
|
neuregulin 1 |
| chr5_-_133968578 | 0.79 |
ENST00000231512.5
|
C5orf15
|
chromosome 5 open reading frame 15 |
| chr6_+_31586859 | 0.75 |
ENST00000433492.5
|
LST1
|
leukocyte specific transcript 1 |
| chr5_-_16916400 | 0.73 |
ENST00000513882.5
|
MYO10
|
myosin X |
| chr17_-_79009731 | 0.72 |
ENST00000392446.10
ENST00000590370.5 ENST00000591625.5 |
CANT1
|
calcium activated nucleotidase 1 |
| chr5_-_143403611 | 0.71 |
ENST00000394464.7
ENST00000231509.7 |
NR3C1
|
nuclear receptor subfamily 3 group C member 1 |
| chr11_-_18588699 | 0.70 |
ENST00000379387.8
ENST00000541984.5 ENST00000396197.8 ENST00000543987.5 |
UEVLD
|
UEV and lactate/malate dehyrogenase domains |
| chr6_+_31586835 | 0.70 |
ENST00000211921.11
|
LST1
|
leukocyte specific transcript 1 |
| chr17_-_79009778 | 0.70 |
ENST00000591773.5
ENST00000588611.5 ENST00000586916.6 ENST00000592033.5 ENST00000588075.5 ENST00000302345.6 ENST00000591811.1 |
CANT1
|
calcium activated nucleotidase 1 |
| chr18_-_5396265 | 0.70 |
ENST00000579951.2
|
EPB41L3
|
erythrocyte membrane protein band 4.1 like 3 |
| chr5_-_59586393 | 0.68 |
ENST00000505453.1
ENST00000360047.9 |
PDE4D
|
phosphodiesterase 4D |
| chr8_-_673547 | 0.65 |
ENST00000522893.1
|
ERICH1
|
glutamate rich 1 |
| chr1_+_152908538 | 0.64 |
ENST00000368764.4
|
IVL
|
involucrin |
| chr6_+_31547560 | 0.64 |
ENST00000376148.9
ENST00000376145.8 |
NFKBIL1
|
NFKB inhibitor like 1 |
| chr2_-_24328113 | 0.63 |
ENST00000622089.4
|
ITSN2
|
intersectin 2 |
| chr3_+_119579676 | 0.61 |
ENST00000357003.7
|
ADPRH
|
ADP-ribosylarginine hydrolase |
| chr5_+_144205250 | 0.60 |
ENST00000507359.3
|
KCTD16
|
potassium channel tetramerization domain containing 16 |
| chr14_-_68793055 | 0.60 |
ENST00000439696.3
|
ZFP36L1
|
ZFP36 ring finger protein like 1 |
| chr17_+_1771688 | 0.60 |
ENST00000572048.1
ENST00000573763.1 |
SERPINF1
|
serpin family F member 1 |
| chrX_+_110944285 | 0.59 |
ENST00000425146.5
ENST00000446737.5 |
PAK3
|
p21 (RAC1) activated kinase 3 |
| chr5_-_147081428 | 0.59 |
ENST00000394413.7
|
PPP2R2B
|
protein phosphatase 2 regulatory subunit Bbeta |
| chr3_-_142448060 | 0.55 |
ENST00000264951.8
|
XRN1
|
5'-3' exoribonuclease 1 |
| chr12_-_108827384 | 0.54 |
ENST00000326470.9
|
SSH1
|
slingshot protein phosphatase 1 |
| chr1_-_228406761 | 0.54 |
ENST00000366699.3
ENST00000284551.11 |
TRIM11
|
tripartite motif containing 11 |
| chr3_+_130850585 | 0.53 |
ENST00000505330.5
ENST00000504381.5 ENST00000507488.6 |
ATP2C1
|
ATPase secretory pathway Ca2+ transporting 1 |
| chr1_+_15153698 | 0.53 |
ENST00000400796.7
ENST00000376008.3 ENST00000434578.6 |
TMEM51
|
transmembrane protein 51 |
| chr16_-_28597042 | 0.52 |
ENST00000533150.5
ENST00000335715.9 |
SULT1A2
|
sulfotransferase family 1A member 2 |
| chr9_+_89605004 | 0.52 |
ENST00000252506.11
ENST00000375769.1 |
GADD45G
|
growth arrest and DNA damage inducible gamma |
| chr16_-_28609976 | 0.52 |
ENST00000566189.5
|
SULT1A1
|
sulfotransferase family 1A member 1 |
| chr10_+_112375196 | 0.51 |
ENST00000393081.6
|
ACSL5
|
acyl-CoA synthetase long chain family member 5 |
| chr3_-_142448028 | 0.51 |
ENST00000392981.7
|
XRN1
|
5'-3' exoribonuclease 1 |
| chr10_+_84452208 | 0.51 |
ENST00000480006.1
|
CCSER2
|
coiled-coil serine rich protein 2 |
| chr15_-_40108861 | 0.50 |
ENST00000354670.9
ENST00000559701.5 ENST00000557870.1 ENST00000558774.5 |
BMF
|
Bcl2 modifying factor |
| chr1_-_247760556 | 0.50 |
ENST00000641256.1
|
OR1C1
|
olfactory receptor family 1 subfamily C member 1 |
| chr7_-_138002017 | 0.49 |
ENST00000452463.5
ENST00000456390.5 ENST00000330387.11 |
CREB3L2
|
cAMP responsive element binding protein 3 like 2 |
| chr12_+_121743623 | 0.49 |
ENST00000541467.1
|
TMEM120B
|
transmembrane protein 120B |
| chr12_-_13095664 | 0.49 |
ENST00000337630.10
ENST00000545699.1 |
GSG1
|
germ cell associated 1 |
| chr8_-_133297092 | 0.48 |
ENST00000522890.5
ENST00000675983.1 ENST00000518176.5 ENST00000323851.13 ENST00000522476.5 ENST00000518066.5 ENST00000521544.5 ENST00000674605.1 ENST00000518480.5 ENST00000523892.5 |
NDRG1
|
N-myc downstream regulated 1 |
| chr10_+_72893572 | 0.48 |
ENST00000622652.1
|
OIT3
|
oncoprotein induced transcript 3 |
| chr11_+_72114840 | 0.47 |
ENST00000622388.4
|
FOLR3
|
folate receptor gamma |
| chr11_-_35419213 | 0.46 |
ENST00000642171.1
ENST00000644050.1 ENST00000643134.1 |
SLC1A2
|
solute carrier family 1 member 2 |
| chr12_+_64497968 | 0.46 |
ENST00000676593.1
ENST00000677093.1 |
TBK1
ENSG00000288665.1
|
TANK binding kinase 1 novel transcript |
| chr6_+_29301701 | 0.46 |
ENST00000641895.1
|
OR14J1
|
olfactory receptor family 14 subfamily J member 1 |
| chr19_+_14433284 | 0.43 |
ENST00000242783.11
|
PKN1
|
protein kinase N1 |
| chr16_-_28609992 | 0.42 |
ENST00000314752.11
|
SULT1A1
|
sulfotransferase family 1A member 1 |
| chr7_+_100884974 | 0.42 |
ENST00000448764.5
|
SRRT
|
serrate, RNA effector molecule |
| chr7_-_138001794 | 0.42 |
ENST00000616381.4
ENST00000620715.4 |
CREB3L2
|
cAMP responsive element binding protein 3 like 2 |
| chrX_+_47145240 | 0.41 |
ENST00000628161.2
ENST00000345781.10 |
RBM10
|
RNA binding motif protein 10 |
| chr11_+_62123991 | 0.41 |
ENST00000533896.5
ENST00000278849.4 ENST00000394818.8 |
INCENP
|
inner centromere protein |
| chr11_-_35420050 | 0.41 |
ENST00000395753.6
ENST00000395750.6 ENST00000645634.1 |
SLC1A2
|
solute carrier family 1 member 2 |
| chr11_-_35420017 | 0.40 |
ENST00000643000.1
ENST00000646099.1 ENST00000647372.1 ENST00000642578.1 |
SLC1A2
|
solute carrier family 1 member 2 |
| chr11_-_35419098 | 0.40 |
ENST00000606205.6
ENST00000645303.1 |
SLC1A2
|
solute carrier family 1 member 2 |
| chr12_-_44875647 | 0.40 |
ENST00000395487.6
|
NELL2
|
neural EGFL like 2 |
| chr15_+_63048436 | 0.39 |
ENST00000334895.10
ENST00000404484.9 ENST00000558910.3 ENST00000317516.12 |
TPM1
|
tropomyosin 1 |
| chr8_+_17577179 | 0.39 |
ENST00000251630.11
|
PDGFRL
|
platelet derived growth factor receptor like |
| chr17_-_42964437 | 0.38 |
ENST00000427569.7
ENST00000430739.5 |
AARSD1
|
alanyl-tRNA synthetase domain containing 1 |
| chrM_+_5824 | 0.37 |
ENST00000361624.2
|
MT-CO1
|
mitochondrially encoded cytochrome c oxidase I |
| chr10_-_68332878 | 0.37 |
ENST00000309049.8
|
PBLD
|
phenazine biosynthesis like protein domain containing |
| chr1_+_172420681 | 0.37 |
ENST00000367727.9
|
C1orf105
|
chromosome 1 open reading frame 105 |
| chr18_-_3874270 | 0.37 |
ENST00000400149.7
ENST00000400155.5 ENST00000400150.7 |
DLGAP1
|
DLG associated protein 1 |
| chrX_+_47145200 | 0.37 |
ENST00000377604.8
|
RBM10
|
RNA binding motif protein 10 |
| chr22_-_17219424 | 0.36 |
ENST00000649540.1
ENST00000399837.8 ENST00000543038.1 |
ADA2
|
adenosine deaminase 2 |
| chr19_-_10309783 | 0.36 |
ENST00000403352.1
ENST00000403903.7 |
ZGLP1
|
zinc finger GATA like protein 1 |
| chr6_+_10694916 | 0.36 |
ENST00000379568.4
|
PAK1IP1
|
PAK1 interacting protein 1 |
| chr8_+_142700465 | 0.34 |
ENST00000522591.1
|
LY6K
|
lymphocyte antigen 6 family member K |
| chr9_-_83267230 | 0.34 |
ENST00000328788.5
|
FRMD3
|
FERM domain containing 3 |
| chr22_-_37109703 | 0.34 |
ENST00000406856.7
ENST00000676104.1 |
TMPRSS6
|
transmembrane serine protease 6 |
| chr3_-_21751189 | 0.34 |
ENST00000281523.8
|
ZNF385D
|
zinc finger protein 385D |
| chr10_+_113126670 | 0.33 |
ENST00000369389.6
|
TCF7L2
|
transcription factor 7 like 2 |
| chr11_-_114595777 | 0.33 |
ENST00000375478.4
|
NXPE4
|
neurexophilin and PC-esterase domain family member 4 |
| chr5_+_168486462 | 0.33 |
ENST00000231572.8
ENST00000626454.1 |
RARS1
|
arginyl-tRNA synthetase 1 |
| chr19_+_49119531 | 0.33 |
ENST00000334186.9
|
PPFIA3
|
PTPRF interacting protein alpha 3 |
| chr4_-_175812746 | 0.33 |
ENST00000393658.6
|
GPM6A
|
glycoprotein M6A |
| chr13_+_36674013 | 0.33 |
ENST00000315190.4
|
SERTM1
|
serine rich and transmembrane domain containing 1 |
| chr8_+_142700095 | 0.33 |
ENST00000292430.10
ENST00000518841.5 ENST00000519387.1 |
LY6K
|
lymphocyte antigen 6 family member K |
| chr18_-_3874247 | 0.32 |
ENST00000581699.5
|
DLGAP1
|
DLG associated protein 1 |
| chr22_-_17219571 | 0.32 |
ENST00000610390.4
|
ADA2
|
adenosine deaminase 2 |
| chr17_+_45221993 | 0.32 |
ENST00000328118.7
|
FMNL1
|
formin like 1 |
| chr5_-_169980474 | 0.31 |
ENST00000377365.4
|
INSYN2B
|
inhibitory synaptic factor family member 2B |
| chr19_-_49451793 | 0.31 |
ENST00000262265.10
|
PIH1D1
|
PIH1 domain containing 1 |
| chr15_+_80152772 | 0.30 |
ENST00000407106.5
ENST00000537726.5 ENST00000558767.6 ENST00000261755.9 |
FAH
|
fumarylacetoacetate hydrolase |
| chr11_-_114595750 | 0.30 |
ENST00000424261.6
|
NXPE4
|
neurexophilin and PC-esterase domain family member 4 |
| chr6_+_55174508 | 0.29 |
ENST00000370862.4
|
HCRTR2
|
hypocretin receptor 2 |
| chr4_-_170090153 | 0.29 |
ENST00000509167.5
ENST00000353187.6 ENST00000507375.5 ENST00000515480.5 |
AADAT
|
aminoadipate aminotransferase |
| chr17_-_17577440 | 0.29 |
ENST00000395782.5
|
PEMT
|
phosphatidylethanolamine N-methyltransferase |
| chr19_+_38289138 | 0.28 |
ENST00000590738.1
ENST00000587519.4 ENST00000591889.2 |
SPINT2
ENSG00000267748.4
|
serine peptidase inhibitor, Kunitz type 2 novel protein |
| chr20_-_53593829 | 0.28 |
ENST00000371471.7
|
ZNF217
|
zinc finger protein 217 |
| chr9_-_5185628 | 0.28 |
ENST00000381641.4
ENST00000649639.1 |
INSL6
|
insulin like 6 |
| chr12_-_13095798 | 0.27 |
ENST00000396302.7
|
GSG1
|
germ cell associated 1 |
| chr6_+_110982028 | 0.27 |
ENST00000441448.7
|
RPF2
|
ribosome production factor 2 homolog |
| chr4_+_164754116 | 0.27 |
ENST00000507311.1
|
SMIM31
|
small integral membrane protein 31 |
| chrX_-_75785205 | 0.27 |
ENST00000373359.4
|
MAGEE2
|
MAGE family member E2 |
| chrX_-_103688090 | 0.26 |
ENST00000433176.6
|
MORF4L2
|
mortality factor 4 like 2 |
| chr2_+_104854104 | 0.26 |
ENST00000361360.4
|
POU3F3
|
POU class 3 homeobox 3 |
| chr2_-_216694794 | 0.26 |
ENST00000449583.1
|
IGFBP5
|
insulin like growth factor binding protein 5 |
| chr7_+_73433761 | 0.25 |
ENST00000344575.5
|
FZD9
|
frizzled class receptor 9 |
| chr8_-_38996466 | 0.25 |
ENST00000456845.6
ENST00000456397.7 ENST00000397070.6 ENST00000517872.1 |
TM2D2
|
TM2 domain containing 2 |
| chr17_+_16381083 | 0.25 |
ENST00000535788.1
ENST00000302182.8 |
UBB
|
ubiquitin B |
| chr12_-_13095628 | 0.25 |
ENST00000457134.6
ENST00000537302.5 |
GSG1
|
germ cell associated 1 |
| chr1_+_151060357 | 0.24 |
ENST00000368921.5
|
MLLT11
|
MLLT11 transcription factor 7 cofactor |
| chr3_-_142448004 | 0.24 |
ENST00000463916.5
|
XRN1
|
5'-3' exoribonuclease 1 |
| chrX_-_103688033 | 0.24 |
ENST00000434230.5
ENST00000418819.5 ENST00000360458.5 |
MORF4L2
|
mortality factor 4 like 2 |
| chr11_-_65606959 | 0.24 |
ENST00000532507.5
|
MAP3K11
|
mitogen-activated protein kinase kinase kinase 11 |
| chr8_+_68330923 | 0.23 |
ENST00000518698.6
|
C8orf34
|
chromosome 8 open reading frame 34 |
| chr10_+_113129285 | 0.23 |
ENST00000637574.1
|
TCF7L2
|
transcription factor 7 like 2 |
| chr11_-_62601223 | 0.23 |
ENST00000527204.5
|
MTA2
|
metastasis associated 1 family member 2 |
| chr2_+_104853259 | 0.22 |
ENST00000598623.1
ENST00000653688.1 ENST00000662784.1 ENST00000666977.1 ENST00000674056.1 |
ENSG00000269707.2
POU3F3
|
novel transcript, sense overlapping POU3F3 POU class 3 homeobox 3 |
| chr16_-_3024230 | 0.22 |
ENST00000572355.5
ENST00000574980.5 ENST00000354679.3 ENST00000573842.1 |
HCFC1R1
|
host cell factor C1 regulator 1 |
| chr15_+_80152978 | 0.22 |
ENST00000561421.6
ENST00000684363.1 |
FAH
|
fumarylacetoacetate hydrolase |
| chr12_-_113136224 | 0.21 |
ENST00000546530.5
ENST00000261729.9 |
RASAL1
|
RAS protein activator like 1 |
| chr19_+_10013468 | 0.21 |
ENST00000591589.3
|
RDH8
|
retinol dehydrogenase 8 |
| chr5_-_32444722 | 0.21 |
ENST00000265069.13
|
ZFR
|
zinc finger RNA binding protein |
| chr20_-_56005466 | 0.21 |
ENST00000064571.3
|
CBLN4
|
cerebellin 4 precursor |
| chr13_-_46897021 | 0.20 |
ENST00000542664.4
ENST00000543956.5 |
HTR2A
|
5-hydroxytryptamine receptor 2A |
| chr2_-_131093378 | 0.20 |
ENST00000409185.5
ENST00000389915.4 |
FAM168B
|
family with sequence similarity 168 member B |
| chr1_-_244862381 | 0.20 |
ENST00000640001.1
ENST00000639628.1 |
HNRNPU
|
heterogeneous nuclear ribonucleoprotein U |
| chr11_-_35419899 | 0.19 |
ENST00000646847.1
ENST00000449068.2 ENST00000643401.1 ENST00000645966.1 ENST00000647104.1 |
SLC1A2
|
solute carrier family 1 member 2 |
| chr2_+_203014842 | 0.19 |
ENST00000683969.1
ENST00000449802.5 |
NBEAL1
|
neurobeachin like 1 |
| chr9_+_127803208 | 0.19 |
ENST00000373225.7
ENST00000431857.5 |
FPGS
|
folylpolyglutamate synthase |
| chr17_+_74274229 | 0.19 |
ENST00000311014.11
|
DNAI2
|
dynein axonemal intermediate chain 2 |
| chr7_-_127252919 | 0.19 |
ENST00000339582.7
|
GRM8
|
glutamate metabotropic receptor 8 |
| chr8_+_7495892 | 0.19 |
ENST00000355602.3
|
DEFB107B
|
defensin beta 107B |
| chr1_+_206440061 | 0.19 |
ENST00000604925.5
|
SRGAP2
|
SLIT-ROBO Rho GTPase activating protein 2 |
| chr19_+_49388243 | 0.19 |
ENST00000447857.8
|
KASH5
|
KASH domain containing 5 |
| chr12_-_121039204 | 0.19 |
ENST00000620239.5
|
OASL
|
2'-5'-oligoadenylate synthetase like |
| chr8_+_103880412 | 0.18 |
ENST00000436393.6
|
RIMS2
|
regulating synaptic membrane exocytosis 2 |
| chr9_+_19049385 | 0.18 |
ENST00000380527.3
|
RRAGA
|
Ras related GTP binding A |
| chr19_-_6333603 | 0.18 |
ENST00000301452.5
|
ACER1
|
alkaline ceramidase 1 |
| chr22_-_16825404 | 0.17 |
ENST00000684488.1
|
XKR3
|
XK related 3 |
| chr21_-_32279012 | 0.17 |
ENST00000290130.4
|
MIS18A
|
MIS18 kinetochore protein A |
| chr12_-_2876986 | 0.17 |
ENST00000342628.6
ENST00000361953.7 |
FOXM1
|
forkhead box M1 |
| chr16_-_84239750 | 0.17 |
ENST00000568181.1
|
KCNG4
|
potassium voltage-gated channel modifier subfamily G member 4 |
| chr12_-_121039236 | 0.16 |
ENST00000257570.9
|
OASL
|
2'-5'-oligoadenylate synthetase like |
| chr6_-_116668751 | 0.16 |
ENST00000368576.8
ENST00000368573.5 |
ZUP1
|
zinc finger containing ubiquitin peptidase 1 |
| chr3_-_9880250 | 0.16 |
ENST00000423850.5
ENST00000336832.7 ENST00000675828.1 ENST00000618572.4 ENST00000455015.6 |
CIDEC
|
cell death inducing DFFA like effector c |
| chr5_+_147878703 | 0.16 |
ENST00000296694.5
|
SCGB3A2
|
secretoglobin family 3A member 2 |
| chr8_-_144428502 | 0.15 |
ENST00000531032.5
ENST00000530790.5 ENST00000292510.6 ENST00000533806.5 ENST00000377348.6 |
VPS28
|
VPS28 subunit of ESCRT-I |
| chr17_-_76053639 | 0.15 |
ENST00000602720.5
|
SRP68
|
signal recognition particle 68 |
| chr11_-_111871530 | 0.15 |
ENST00000614444.4
ENST00000616540.5 |
ALG9
|
ALG9 alpha-1,2-mannosyltransferase |
| chr11_+_120236635 | 0.15 |
ENST00000260264.8
|
POU2F3
|
POU class 2 homeobox 3 |
| chr19_-_50833187 | 0.14 |
ENST00000598673.1
|
KLK15
|
kallikrein related peptidase 15 |
| chr10_-_24723185 | 0.14 |
ENST00000376410.7
ENST00000446003.5 |
ARHGAP21
|
Rho GTPase activating protein 21 |
| chr6_+_31615215 | 0.14 |
ENST00000337917.11
ENST00000376059.8 |
AIF1
|
allograft inflammatory factor 1 |
| chr2_-_169694367 | 0.14 |
ENST00000447353.6
|
CCDC173
|
coiled-coil domain containing 173 |
| chr19_+_14941489 | 0.14 |
ENST00000248072.3
|
OR7C2
|
olfactory receptor family 7 subfamily C member 2 |
| chr22_-_39244969 | 0.14 |
ENST00000331163.11
|
PDGFB
|
platelet derived growth factor subunit B |
| chr8_-_7815716 | 0.14 |
ENST00000335021.2
|
DEFB107A
|
defensin beta 107A |
| chr11_-_31369728 | 0.14 |
ENST00000684477.1
ENST00000452803.1 |
DCDC1
|
doublecortin domain containing 1 |
| chr8_+_41490396 | 0.13 |
ENST00000518270.5
ENST00000520817.5 |
GOLGA7
|
golgin A7 |
| chr17_+_45221854 | 0.13 |
ENST00000331495.8
|
FMNL1
|
formin like 1 |
| chr22_-_31292445 | 0.13 |
ENST00000402249.7
ENST00000215912.10 ENST00000443175.1 ENST00000441972.5 |
PIK3IP1
|
phosphoinositide-3-kinase interacting protein 1 |
| chr2_+_86942118 | 0.13 |
ENST00000641458.2
|
RGPD1
|
RANBP2 like and GRIP domain containing 1 |
| chr12_+_116559381 | 0.13 |
ENST00000556529.4
|
MAP1LC3B2
|
microtubule associated protein 1 light chain 3 beta 2 |
| chr10_-_69573416 | 0.13 |
ENST00000242462.5
|
NEUROG3
|
neurogenin 3 |
| chr1_-_151346806 | 0.13 |
ENST00000392746.7
|
RFX5
|
regulatory factor X5 |
| chr12_-_121039156 | 0.13 |
ENST00000339275.10
|
OASL
|
2'-5'-oligoadenylate synthetase like |
| chr7_-_23470469 | 0.13 |
ENST00000258729.8
|
IGF2BP3
|
insulin like growth factor 2 mRNA binding protein 3 |
| chr3_+_35641421 | 0.12 |
ENST00000449196.5
|
ARPP21
|
cAMP regulated phosphoprotein 21 |
| chr1_-_109075944 | 0.12 |
ENST00000338366.6
|
TAF13
|
TATA-box binding protein associated factor 13 |
| chr17_+_7281711 | 0.12 |
ENST00000317370.13
ENST00000571308.5 |
SLC2A4
|
solute carrier family 2 member 4 |
| chr11_-_59212869 | 0.12 |
ENST00000361050.4
|
MPEG1
|
macrophage expressed 1 |
| chrX_-_132489842 | 0.12 |
ENST00000436215.5
|
MBNL3
|
muscleblind like splicing regulator 3 |
| chr7_+_66921217 | 0.12 |
ENST00000341567.8
ENST00000607045.5 |
TMEM248
|
transmembrane protein 248 |
| chrX_+_66164340 | 0.11 |
ENST00000441993.7
ENST00000419594.6 ENST00000425114.2 |
HEPH
|
hephaestin |
| chrX_+_77447387 | 0.11 |
ENST00000439435.3
|
FGF16
|
fibroblast growth factor 16 |
| chr1_+_26317950 | 0.11 |
ENST00000374213.3
|
CD52
|
CD52 molecule |
| chr1_+_162069768 | 0.11 |
ENST00000530878.5
|
NOS1AP
|
nitric oxide synthase 1 adaptor protein |
| chr12_+_123458092 | 0.11 |
ENST00000526639.3
|
SNRNP35
|
small nuclear ribonucleoprotein U11/U12 subunit 35 |
| chr16_+_69924984 | 0.11 |
ENST00000568684.1
|
WWP2
|
WW domain containing E3 ubiquitin protein ligase 2 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 8.9 | GO:0035655 | interleukin-18-mediated signaling pathway(GO:0035655) |
| 0.5 | 1.5 | GO:0002428 | antigen processing and presentation of peptide antigen via MHC class Ib(GO:0002428) cytosol to ER transport(GO:0046967) |
| 0.5 | 3.4 | GO:0060414 | aorta smooth muscle tissue morphogenesis(GO:0060414) |
| 0.4 | 1.7 | GO:0021827 | cerebral cortex tangential migration using cell-cell interactions(GO:0021823) postnatal olfactory bulb interneuron migration(GO:0021827) chemorepulsion involved in postnatal olfactory bulb interneuron migration(GO:0021836) negative regulation of negative chemotaxis(GO:0050925) |
| 0.3 | 4.5 | GO:0021840 | directional guidance of interneurons involved in migration from the subpallium to the cortex(GO:0021840) chemorepulsion involved in interneuron migration from the subpallium to the cortex(GO:0021842) ERBB3 signaling pathway(GO:0038129) |
| 0.3 | 1.3 | GO:0071409 | cellular response to cycloheximide(GO:0071409) |
| 0.2 | 1.0 | GO:0032499 | detection of peptidoglycan(GO:0032499) activation of MAPK activity involved in innate immune response(GO:0035419) regulation of prostaglandin-endoperoxide synthase activity(GO:0060584) positive regulation of prostaglandin-endoperoxide synthase activity(GO:0060585) |
| 0.2 | 1.8 | GO:2001206 | positive regulation of osteoclast development(GO:2001206) |
| 0.2 | 0.6 | GO:1904580 | regulation of intracellular mRNA localization(GO:1904580) positive regulation of intracellular mRNA localization(GO:1904582) |
| 0.2 | 1.6 | GO:0043402 | glucocorticoid mediated signaling pathway(GO:0043402) |
| 0.2 | 0.9 | GO:0060480 | lung goblet cell differentiation(GO:0060480) |
| 0.2 | 0.5 | GO:1904717 | excitatory chemical synaptic transmission(GO:0098976) regulation of AMPA glutamate receptor clustering(GO:1904717) positive regulation of AMPA glutamate receptor clustering(GO:1904719) |
| 0.2 | 0.9 | GO:1903904 | negative regulation of establishment of T cell polarity(GO:1903904) negative regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001107) |
| 0.2 | 0.5 | GO:0072023 | thick ascending limb development(GO:0072023) metanephric thick ascending limb development(GO:0072233) |
| 0.2 | 0.6 | GO:0018199 | peptidyl-glutamine modification(GO:0018199) |
| 0.1 | 1.0 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.1 | 1.9 | GO:0070779 | D-aspartate transport(GO:0070777) D-aspartate import(GO:0070779) |
| 0.1 | 3.1 | GO:0006068 | ethanol catabolic process(GO:0006068) |
| 0.1 | 0.9 | GO:0015712 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.1 | 0.3 | GO:1904204 | skeletal muscle hypertrophy(GO:0014734) regulation of skeletal muscle hypertrophy(GO:1904204) |
| 0.1 | 0.3 | GO:0060392 | negative regulation of SMAD protein import into nucleus(GO:0060392) |
| 0.1 | 0.4 | GO:0035407 | histone H3-T11 phosphorylation(GO:0035407) |
| 0.1 | 0.3 | GO:1900110 | negative regulation of histone H3-K9 dimethylation(GO:1900110) regulation of TORC2 signaling(GO:1903939) |
| 0.1 | 0.6 | GO:0071279 | cellular response to cobalt ion(GO:0071279) |
| 0.1 | 0.7 | GO:0046103 | adenosine catabolic process(GO:0006154) inosine biosynthetic process(GO:0046103) |
| 0.1 | 0.5 | GO:0044565 | dendritic cell proliferation(GO:0044565) |
| 0.1 | 0.5 | GO:0032468 | Golgi calcium ion homeostasis(GO:0032468) |
| 0.1 | 0.3 | GO:1904397 | negative regulation of neuromuscular junction development(GO:1904397) |
| 0.1 | 0.4 | GO:0003065 | positive regulation of heart rate by epinephrine(GO:0003065) |
| 0.1 | 0.7 | GO:0002175 | protein localization to paranode region of axon(GO:0002175) |
| 0.1 | 0.6 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
| 0.1 | 0.3 | GO:0060671 | epithelial cell differentiation involved in embryonic placenta development(GO:0060671) epithelial cell morphogenesis involved in placental branching(GO:0060672) |
| 0.1 | 0.7 | GO:0086024 | adrenergic receptor signaling pathway involved in positive regulation of heart rate(GO:0086024) |
| 0.1 | 0.2 | GO:0003363 | lamellipodium assembly involved in ameboidal cell migration(GO:0003363) |
| 0.1 | 0.3 | GO:0033512 | L-lysine catabolic process to acetyl-CoA via saccharopine(GO:0033512) |
| 0.1 | 6.4 | GO:0010803 | regulation of tumor necrosis factor-mediated signaling pathway(GO:0010803) |
| 0.1 | 0.2 | GO:1905026 | positive regulation of voltage-gated potassium channel activity involved in ventricular cardiac muscle cell action potential repolarization(GO:1903762) positive regulation of ventricular cardiac muscle cell action potential(GO:1903947) positive regulation of membrane repolarization during ventricular cardiac muscle cell action potential(GO:1905026) positive regulation of membrane repolarization during cardiac muscle cell action potential(GO:1905033) |
| 0.1 | 0.5 | GO:0006572 | tyrosine catabolic process(GO:0006572) |
| 0.1 | 0.2 | GO:2000395 | regulation of ubiquitin-dependent endocytosis(GO:2000395) positive regulation of ubiquitin-dependent endocytosis(GO:2000397) |
| 0.0 | 0.8 | GO:0072619 | interleukin-21 production(GO:0032625) interleukin-21 secretion(GO:0072619) |
| 0.0 | 0.3 | GO:0010840 | regulation of circadian sleep/wake cycle, wakefulness(GO:0010840) circadian sleep/wake cycle, wakefulness(GO:0042746) |
| 0.0 | 0.5 | GO:0046598 | positive regulation of viral entry into host cell(GO:0046598) |
| 0.0 | 0.2 | GO:0046900 | tetrahydrofolylpolyglutamate metabolic process(GO:0046900) |
| 0.0 | 0.6 | GO:0048625 | myoblast fate commitment(GO:0048625) |
| 0.0 | 0.5 | GO:0032287 | peripheral nervous system myelin maintenance(GO:0032287) |
| 0.0 | 0.2 | GO:0010446 | response to alkaline pH(GO:0010446) |
| 0.0 | 0.2 | GO:0010513 | positive regulation of phosphatidylinositol biosynthetic process(GO:0010513) |
| 0.0 | 0.6 | GO:0031665 | negative regulation of lipopolysaccharide-mediated signaling pathway(GO:0031665) |
| 0.0 | 0.1 | GO:0014738 | regulation of muscle hyperplasia(GO:0014738) |
| 0.0 | 0.3 | GO:0006686 | sphingomyelin biosynthetic process(GO:0006686) positive regulation of lipoprotein metabolic process(GO:0050747) |
| 0.0 | 3.9 | GO:0032945 | negative regulation of mononuclear cell proliferation(GO:0032945) negative regulation of lymphocyte proliferation(GO:0050672) |
| 0.0 | 0.2 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.0 | 0.4 | GO:0070050 | neuron cellular homeostasis(GO:0070050) |
| 0.0 | 0.4 | GO:0035791 | platelet-derived growth factor receptor-beta signaling pathway(GO:0035791) |
| 0.0 | 0.5 | GO:0015884 | folic acid transport(GO:0015884) |
| 0.0 | 0.3 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 0.0 | 0.5 | GO:0032464 | positive regulation of protein homooligomerization(GO:0032464) |
| 0.0 | 1.0 | GO:1903861 | positive regulation of dendrite extension(GO:1903861) |
| 0.0 | 0.4 | GO:0031053 | primary miRNA processing(GO:0031053) |
| 0.0 | 0.4 | GO:0006450 | regulation of translational fidelity(GO:0006450) |
| 0.0 | 0.2 | GO:1904263 | positive regulation of TORC1 signaling(GO:1904263) |
| 0.0 | 0.6 | GO:0070262 | peptidyl-serine dephosphorylation(GO:0070262) |
| 0.0 | 0.2 | GO:0007256 | activation of JNKK activity(GO:0007256) |
| 0.0 | 0.1 | GO:0051684 | maintenance of Golgi location(GO:0051684) |
| 0.0 | 0.5 | GO:1900745 | positive regulation of p38MAPK cascade(GO:1900745) |
| 0.0 | 0.6 | GO:0010763 | positive regulation of fibroblast migration(GO:0010763) |
| 0.0 | 0.3 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.0 | 0.1 | GO:0051801 | cytolysis in other organism involved in symbiotic interaction(GO:0051801) neutrophil mediated killing of gram-negative bacterium(GO:0070945) positive regulation of phospholipase C-activating G-protein coupled receptor signaling pathway(GO:1900738) |
| 0.0 | 0.5 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.0 | 0.3 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.0 | 0.1 | GO:1904849 | positive regulation of fat cell proliferation(GO:0070346) positive regulation of cell chemotaxis to fibroblast growth factor(GO:1904849) positive regulation of endothelial cell chemotaxis to fibroblast growth factor(GO:2000546) |
| 0.0 | 1.1 | GO:0051489 | regulation of filopodium assembly(GO:0051489) |
| 0.0 | 0.4 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.0 | 0.0 | GO:0002503 | peptide antigen assembly with MHC class II protein complex(GO:0002503) |
| 0.0 | 0.2 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.0 | 0.2 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.0 | 0.7 | GO:0007339 | binding of sperm to zona pellucida(GO:0007339) |
| 0.0 | 0.5 | GO:0035338 | long-chain fatty-acyl-CoA biosynthetic process(GO:0035338) |
| 0.0 | 0.2 | GO:0051901 | positive regulation of mitochondrial depolarization(GO:0051901) |
| 0.0 | 0.2 | GO:0070934 | CRD-mediated mRNA stabilization(GO:0070934) |
| 0.0 | 0.3 | GO:0036158 | outer dynein arm assembly(GO:0036158) |
| 0.0 | 1.1 | GO:0030166 | proteoglycan biosynthetic process(GO:0030166) |
| 0.0 | 0.2 | GO:0007196 | adenylate cyclase-inhibiting G-protein coupled glutamate receptor signaling pathway(GO:0007196) |
| 0.0 | 0.2 | GO:2000781 | positive regulation of double-strand break repair(GO:2000781) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.5 | GO:0042825 | TAP complex(GO:0042825) |
| 0.1 | 7.1 | GO:0030673 | axolemma(GO:0030673) |
| 0.1 | 0.3 | GO:1990742 | microvesicle(GO:1990742) |
| 0.1 | 0.9 | GO:0060171 | stereocilium membrane(GO:0060171) |
| 0.1 | 0.4 | GO:0000801 | central element(GO:0000801) |
| 0.0 | 0.6 | GO:0043203 | axon hillock(GO:0043203) |
| 0.0 | 0.8 | GO:0032059 | bleb(GO:0032059) |
| 0.0 | 4.3 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 0.6 | GO:0070369 | beta-catenin-TCF7L2 complex(GO:0070369) |
| 0.0 | 0.2 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.0 | 0.3 | GO:0042567 | insulin-like growth factor ternary complex(GO:0042567) |
| 0.0 | 0.3 | GO:0070761 | pre-snoRNP complex(GO:0070761) |
| 0.0 | 0.7 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.0 | 0.2 | GO:1990131 | Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.0 | 0.4 | GO:0045275 | mitochondrial respiratory chain complex III(GO:0005750) respiratory chain complex III(GO:0045275) |
| 0.0 | 0.7 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.0 | 0.2 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.0 | 0.3 | GO:0036157 | outer dynein arm(GO:0036157) |
| 0.0 | 2.1 | GO:1904724 | tertiary granule lumen(GO:1904724) |
| 0.0 | 1.1 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.0 | 0.2 | GO:0044327 | dendritic spine head(GO:0044327) |
| 0.0 | 1.0 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 0.2 | GO:0005786 | signal recognition particle, endoplasmic reticulum targeting(GO:0005786) |
| 0.0 | 0.5 | GO:1902562 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.0 | 0.1 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.0 | 0.6 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.0 | 0.3 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.0 | 0.1 | GO:0000243 | commitment complex(GO:0000243) |
| 0.0 | 0.2 | GO:0070937 | CRD-mediated mRNA stability complex(GO:0070937) |
| 0.0 | 1.9 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 0.6 | GO:0001533 | cornified envelope(GO:0001533) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.9 | 8.7 | GO:0042008 | interleukin-18 receptor activity(GO:0042008) |
| 0.5 | 3.1 | GO:0047894 | flavonol 3-sulfotransferase activity(GO:0047894) |
| 0.5 | 1.4 | GO:0004382 | guanosine-diphosphatase activity(GO:0004382) |
| 0.4 | 3.4 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.4 | 6.4 | GO:0031996 | thioesterase binding(GO:0031996) |
| 0.3 | 1.6 | GO:0038051 | glucocorticoid receptor activity(GO:0004883) glucocorticoid-activated RNA polymerase II transcription factor binding transcription factor activity(GO:0038051) |
| 0.3 | 1.5 | GO:0015433 | peptide antigen-transporting ATPase activity(GO:0015433) |
| 0.2 | 4.5 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 0.2 | 1.3 | GO:0004534 | 5'-3' exoribonuclease activity(GO:0004534) |
| 0.2 | 0.9 | GO:0015315 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 0.2 | 1.7 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.2 | 1.0 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) |
| 0.1 | 1.9 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 0.1 | 0.4 | GO:0005017 | platelet-derived growth factor-activated receptor activity(GO:0005017) |
| 0.1 | 0.6 | GO:0003875 | ADP-ribosylarginine hydrolase activity(GO:0003875) |
| 0.1 | 0.4 | GO:0035402 | histone kinase activity (H3-T11 specific)(GO:0035402) |
| 0.1 | 0.5 | GO:0016822 | hydrolase activity, acting on acid carbon-carbon bonds(GO:0016822) hydrolase activity, acting on acid carbon-carbon bonds, in ketonic substances(GO:0016823) |
| 0.1 | 0.3 | GO:0080101 | phosphatidyl-N-methylethanolamine N-methyltransferase activity(GO:0000773) phosphatidylethanolamine N-methyltransferase activity(GO:0004608) phosphatidyl-N-dimethylethanolamine N-methyltransferase activity(GO:0080101) |
| 0.1 | 0.8 | GO:0001727 | lipid kinase activity(GO:0001727) |
| 0.1 | 0.7 | GO:0060002 | plus-end directed microfilament motor activity(GO:0060002) |
| 0.1 | 1.0 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.1 | 0.7 | GO:0031685 | adenosine receptor binding(GO:0031685) |
| 0.1 | 1.0 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.0 | 0.2 | GO:0004706 | JUN kinase kinase kinase activity(GO:0004706) |
| 0.0 | 0.3 | GO:0036137 | kynurenine-oxoglutarate transaminase activity(GO:0016212) kynurenine aminotransferase activity(GO:0036137) |
| 0.0 | 0.4 | GO:0002161 | aminoacyl-tRNA editing activity(GO:0002161) |
| 0.0 | 0.5 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 0.0 | 1.5 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.0 | 0.3 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.0 | 0.7 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.0 | 0.4 | GO:0005522 | profilin binding(GO:0005522) |
| 0.0 | 0.6 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.0 | 0.7 | GO:0031698 | beta-2 adrenergic receptor binding(GO:0031698) |
| 0.0 | 0.2 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.0 | 0.5 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.0 | 0.5 | GO:0005542 | folic acid binding(GO:0005542) |
| 0.0 | 0.2 | GO:0005047 | signal recognition particle binding(GO:0005047) endoplasmic reticulum signal peptide binding(GO:0030942) |
| 0.0 | 0.3 | GO:0045504 | dynein heavy chain binding(GO:0045504) |
| 0.0 | 0.3 | GO:0008097 | 5S rRNA binding(GO:0008097) |
| 0.0 | 0.6 | GO:0005549 | odorant binding(GO:0005549) |
| 0.0 | 0.2 | GO:0051378 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.0 | 0.1 | GO:0001165 | RNA polymerase I upstream control element sequence-specific DNA binding(GO:0001165) |
| 0.0 | 0.2 | GO:0004303 | estradiol 17-beta-dehydrogenase activity(GO:0004303) |
| 0.0 | 0.5 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.0 | 0.6 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.0 | 0.5 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.0 | 0.1 | GO:0038052 | RNA polymerase II transcription factor activity, estrogen-activated sequence-specific DNA binding(GO:0038052) |
| 0.0 | 0.1 | GO:0016724 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.0 | 0.9 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 1.0 | GO:0070063 | RNA polymerase binding(GO:0070063) |
| 0.0 | 0.1 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) platelet-derived growth factor binding(GO:0048407) |
| 0.0 | 0.4 | GO:0015002 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 8.8 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.1 | 4.5 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.1 | 6.4 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.1 | 3.9 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 0.4 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 1.7 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 0.6 | SIG CD40PATHWAYMAP | Genes related to CD40 signaling |
| 0.0 | 0.5 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.0 | 0.5 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.0 | 0.7 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 0.2 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.0 | 1.1 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.0 | 0.8 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 4.5 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.1 | 3.1 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.1 | 1.9 | REACTOME DESTABILIZATION OF MRNA BY BRF1 | Genes involved in Destabilization of mRNA by Butyrate Response Factor 1 (BRF1) |
| 0.1 | 1.7 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.1 | 3.2 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.1 | 1.5 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.0 | 1.0 | REACTOME ACTIVATED TAK1 MEDIATES P38 MAPK ACTIVATION | Genes involved in activated TAK1 mediates p38 MAPK activation |
| 0.0 | 1.6 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.0 | 0.3 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.0 | 0.5 | REACTOME ACTIVATION OF IRF3 IRF7 MEDIATED BY TBK1 IKK EPSILON | Genes involved in Activation of IRF3/IRF7 mediated by TBK1/IKK epsilon |
| 0.0 | 0.5 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.0 | 1.9 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.0 | 0.4 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.0 | 0.2 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 0.0 | 0.5 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 0.7 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.0 | 0.2 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.0 | 0.7 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.0 | 0.8 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 0.7 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 0.3 | REACTOME BETA DEFENSINS | Genes involved in Beta defensins |