Project
Inflammatory response time course, HUVEC (Wada, 2009)
Navigation
Downloads
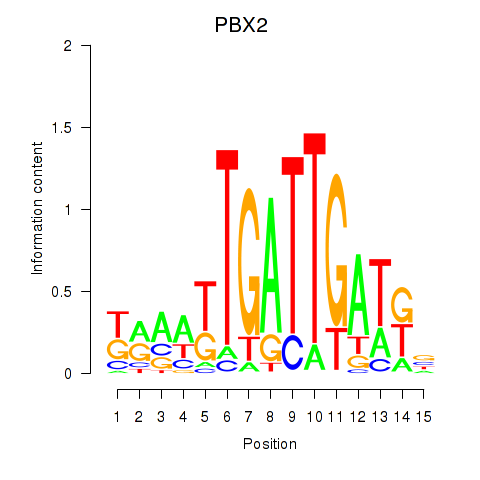
Results for PBX2
Z-value: 0.25
Transcription factors associated with PBX2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
PBX2
|
ENSG00000204304.12 | PBX2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| PBX2 | hg38_v1_chr6_-_32190170_32190214 | 0.46 | 2.0e-02 | Click! |
Activity profile of PBX2 motif
Sorted Z-values of PBX2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of PBX2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_+_12716893 | 0.81 |
ENST00000381465.2
ENST00000155926.9 |
TRIB2
|
tribbles pseudokinase 2 |
| chr8_-_6563409 | 0.45 |
ENST00000325203.9
|
ANGPT2
|
angiopoietin 2 |
| chr3_-_129440007 | 0.43 |
ENST00000503197.5
ENST00000249910.5 ENST00000507208.1 ENST00000393278.6 |
MBD4
|
methyl-CpG binding domain 4, DNA glycosylase |
| chr3_-_129439925 | 0.38 |
ENST00000429544.7
|
MBD4
|
methyl-CpG binding domain 4, DNA glycosylase |
| chr10_-_44978789 | 0.36 |
ENST00000448778.1
ENST00000298295.4 |
DEPP1
|
DEPP1 autophagy regulator |
| chr18_-_75208417 | 0.35 |
ENST00000581620.1
ENST00000582437.1 |
ZADH2
|
zinc binding alcohol dehydrogenase domain containing 2 |
| chr17_+_7855055 | 0.35 |
ENST00000574668.1
ENST00000301599.7 |
TMEM88
|
transmembrane protein 88 |
| chr11_-_104898670 | 0.21 |
ENST00000422698.6
|
CASP12
|
caspase 12 (gene/pseudogene) |
| chr14_+_22516273 | 0.19 |
ENST00000390510.1
|
TRAJ27
|
T cell receptor alpha joining 27 |
| chr3_+_129440082 | 0.18 |
ENST00000347300.6
ENST00000296266.7 |
IFT122
|
intraflagellar transport 122 |
| chr2_+_33436304 | 0.17 |
ENST00000402538.7
|
RASGRP3
|
RAS guanyl releasing protein 3 |
| chr2_+_20447065 | 0.17 |
ENST00000272233.6
|
RHOB
|
ras homolog family member B |
| chr7_-_33100886 | 0.16 |
ENST00000448915.1
|
RP9
|
RP9 pre-mRNA splicing factor |
| chr3_+_129440196 | 0.16 |
ENST00000507564.5
ENST00000348417.7 ENST00000504021.5 ENST00000349441.6 |
IFT122
|
intraflagellar transport 122 |
| chr14_-_23155302 | 0.15 |
ENST00000529705.6
|
SLC7A8
|
solute carrier family 7 member 8 |
| chr2_-_264024 | 0.12 |
ENST00000403712.6
ENST00000356150.10 ENST00000626873.2 ENST00000405430.5 |
SH3YL1
|
SH3 and SYLF domain containing 1 |
| chr9_-_20382461 | 0.11 |
ENST00000380321.5
ENST00000629733.3 |
MLLT3
|
MLLT3 super elongation complex subunit |
| chr18_-_75209126 | 0.10 |
ENST00000322342.4
|
ZADH2
|
zinc binding alcohol dehydrogenase domain containing 2 |
| chr2_+_233693659 | 0.10 |
ENST00000406651.1
|
UGT1A6
|
UDP glucuronosyltransferase family 1 member A6 |
| chr12_-_27971970 | 0.10 |
ENST00000395872.5
ENST00000201015.8 |
PTHLH
|
parathyroid hormone like hormone |
| chr6_+_26045374 | 0.09 |
ENST00000612966.3
|
H3C3
|
H3 clustered histone 3 |
| chr10_+_122560639 | 0.08 |
ENST00000344338.7
ENST00000330163.8 ENST00000652446.2 ENST00000666315.1 ENST00000368955.7 ENST00000368909.7 ENST00000368956.6 ENST00000619379.1 |
DMBT1
|
deleted in malignant brain tumors 1 |
| chr10_+_122560751 | 0.08 |
ENST00000338354.10
ENST00000664692.1 ENST00000653442.1 ENST00000664974.1 |
DMBT1
|
deleted in malignant brain tumors 1 |
| chr10_+_122560679 | 0.07 |
ENST00000657942.1
|
DMBT1
|
deleted in malignant brain tumors 1 |
| chr14_+_58298497 | 0.06 |
ENST00000348476.7
ENST00000355431.8 ENST00000395168.7 |
ARID4A
|
AT-rich interaction domain 4A |
| chr13_-_106568107 | 0.06 |
ENST00000400198.8
|
ARGLU1
|
arginine and glutamate rich 1 |
| chr5_+_141392616 | 0.05 |
ENST00000398604.3
|
PCDHGA8
|
protocadherin gamma subfamily A, 8 |
| chr7_-_32490361 | 0.05 |
ENST00000410044.5
ENST00000450169.7 ENST00000409987.5 ENST00000409782.5 |
LSM5
|
LSM5 homolog, U6 small nuclear RNA and mRNA degradation associated |
| chr16_+_1078781 | 0.05 |
ENST00000293897.5
|
SSTR5
|
somatostatin receptor 5 |
| chr17_+_8288637 | 0.04 |
ENST00000407006.8
ENST00000226105.11 ENST00000580434.5 ENST00000439238.3 |
RANGRF
|
RAN guanine nucleotide release factor |
| chrX_+_120604199 | 0.04 |
ENST00000371315.3
|
MCTS1
|
MCTS1 re-initiation and release factor |
| chr1_+_206865620 | 0.04 |
ENST00000367098.6
|
IL20
|
interleukin 20 |
| chr13_+_108269629 | 0.04 |
ENST00000430559.5
ENST00000375887.9 |
TNFSF13B
|
TNF superfamily member 13b |
| chr6_+_159790448 | 0.04 |
ENST00000367034.5
|
MRPL18
|
mitochondrial ribosomal protein L18 |
| chr11_-_55936400 | 0.03 |
ENST00000301532.3
|
OR5I1
|
olfactory receptor family 5 subfamily I member 1 |
| chrX_+_1615049 | 0.03 |
ENST00000381241.9
|
ASMT
|
acetylserotonin O-methyltransferase |
| chr3_+_189171948 | 0.02 |
ENST00000345063.8
|
TPRG1
|
tumor protein p63 regulated 1 |
| chr4_-_117085541 | 0.02 |
ENST00000310754.5
|
TRAM1L1
|
translocation associated membrane protein 1 like 1 |
| chr14_+_64388296 | 0.02 |
ENST00000554739.5
ENST00000554768.6 ENST00000652179.1 ENST00000652337.1 ENST00000557370.3 |
MTHFD1
|
methylenetetrahydrofolate dehydrogenase, cyclohydrolase and formyltetrahydrofolate synthetase 1 |
| chr17_-_30824665 | 0.02 |
ENST00000324238.7
|
CRLF3
|
cytokine receptor like factor 3 |
| chr3_+_120908072 | 0.02 |
ENST00000273666.10
ENST00000471454.6 ENST00000472879.5 ENST00000497029.5 ENST00000492541.5 |
STXBP5L
|
syntaxin binding protein 5 like |
| chr2_+_112275588 | 0.02 |
ENST00000409871.6
ENST00000343936.4 |
ZC3H6
|
zinc finger CCCH-type containing 6 |
| chr10_+_68109433 | 0.02 |
ENST00000613327.4
ENST00000358913.10 ENST00000373675.3 |
MYPN
|
myopalladin |
| chr3_-_52830664 | 0.02 |
ENST00000266041.9
ENST00000406595.5 ENST00000485816.5 |
ITIH4
|
inter-alpha-trypsin inhibitor heavy chain 4 |
| chr3_+_160225409 | 0.01 |
ENST00000326474.5
|
C3orf80
|
chromosome 3 open reading frame 80 |
| chrX_+_120604084 | 0.01 |
ENST00000371317.10
|
MCTS1
|
MCTS1 re-initiation and release factor |
| chr7_-_151519891 | 0.00 |
ENST00000262187.10
|
RHEB
|
Ras homolog, mTORC1 binding |
| chr1_+_101237009 | 0.00 |
ENST00000305352.7
|
S1PR1
|
sphingosine-1-phosphate receptor 1 |
| chrX_+_46837034 | 0.00 |
ENST00000218340.4
|
RP2
|
RP2 activator of ARL3 GTPase |
| chr2_+_203867943 | 0.00 |
ENST00000295854.10
ENST00000487393.1 ENST00000472206.1 |
CTLA4
|
cytotoxic T-lymphocyte associated protein 4 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.8 | GO:0045074 | interleukin-10 biosynthetic process(GO:0042091) regulation of interleukin-10 biosynthetic process(GO:0045074) |
| 0.2 | 0.5 | GO:0050928 | negative regulation of positive chemotaxis(GO:0050928) |
| 0.1 | 0.3 | GO:0035720 | intraciliary anterograde transport(GO:0035720) |
| 0.0 | 0.8 | GO:0045008 | depyrimidination(GO:0045008) |
| 0.0 | 0.2 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.0 | 0.0 | GO:0098905 | regulation of bundle of His cell action potential(GO:0098905) |
| 0.0 | 0.1 | GO:0002188 | translation reinitiation(GO:0002188) |
| 0.0 | 0.0 | GO:0035928 | rRNA import into mitochondrion(GO:0035928) |
| 0.0 | 0.1 | GO:0097368 | establishment of Sertoli cell barrier(GO:0097368) |
| 0.0 | 0.0 | GO:0030186 | melatonin metabolic process(GO:0030186) melatonin biosynthetic process(GO:0030187) |
| 0.0 | 0.2 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.0 | 0.0 | GO:0002636 | positive regulation of germinal center formation(GO:0002636) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | GO:0072557 | IPAF inflammasome complex(GO:0072557) AIM2 inflammasome complex(GO:0097169) |
| 0.0 | 0.3 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.0 | 0.2 | GO:0042589 | zymogen granule membrane(GO:0042589) |
| 0.0 | 0.2 | GO:0005785 | signal recognition particle receptor complex(GO:0005785) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.8 | GO:0008263 | pyrimidine-specific mismatch base pair DNA N-glycosylase activity(GO:0008263) |
| 0.1 | 0.2 | GO:0035375 | zymogen binding(GO:0035375) |
| 0.1 | 0.5 | GO:0047522 | 13-prostaglandin reductase activity(GO:0036132) 15-oxoprostaglandin 13-oxidase activity(GO:0047522) |
| 0.0 | 0.8 | GO:0055106 | ubiquitin-protein transferase regulator activity(GO:0055106) |
| 0.0 | 0.2 | GO:0019534 | toxin transporter activity(GO:0019534) |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.8 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 0.0 | 0.5 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |