Project
Inflammatory response time course, HUVEC (Wada, 2009)
Navigation
Downloads
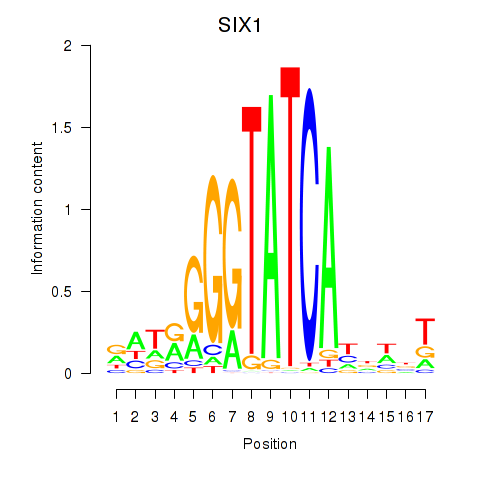
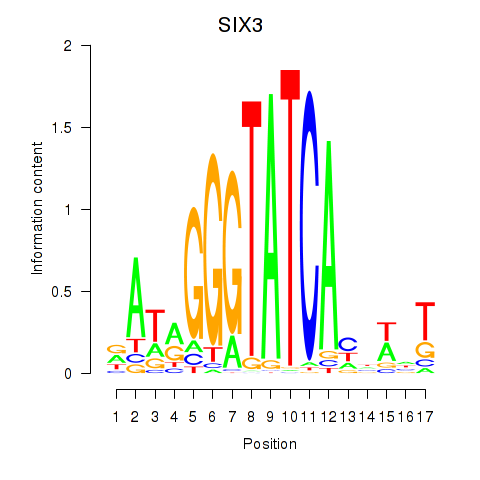
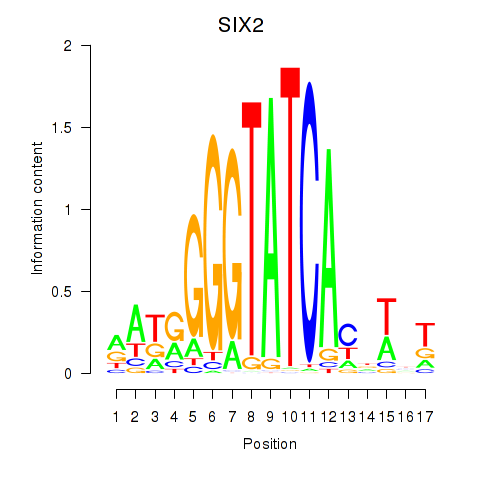
Results for SIX1_SIX3_SIX2
Z-value: 0.68
Transcription factors associated with SIX1_SIX3_SIX2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SIX1
|
ENSG00000126778.12 | SIX1 |
|
SIX3
|
ENSG00000138083.5 | SIX3 |
|
SIX2
|
ENSG00000170577.8 | SIX2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SIX2 | hg38_v1_chr2_-_45009401_45009457 | 0.20 | 3.5e-01 | Click! |
| SIX1 | hg38_v1_chr14_-_60649449_60649490 | 0.17 | 4.1e-01 | Click! |
| SIX3 | hg38_v1_chr2_+_44941695_44941708 | -0.02 | 9.2e-01 | Click! |
Activity profile of SIX1_SIX3_SIX2 motif
Sorted Z-values of SIX1_SIX3_SIX2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of SIX1_SIX3_SIX2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_-_89198868 | 3.23 |
ENST00000355754.7
|
GBP4
|
guanylate binding protein 4 |
| chr6_-_159745186 | 2.57 |
ENST00000537657.5
|
SOD2
|
superoxide dismutase 2 |
| chr5_+_126423122 | 2.45 |
ENST00000515200.5
|
GRAMD2B
|
GRAM domain containing 2B |
| chr5_+_126423363 | 2.41 |
ENST00000285689.8
|
GRAMD2B
|
GRAM domain containing 2B |
| chr5_+_126423176 | 2.40 |
ENST00000542322.5
ENST00000544396.5 |
GRAMD2B
|
GRAM domain containing 2B |
| chr6_-_154430495 | 2.05 |
ENST00000424998.3
|
CNKSR3
|
CNKSR family member 3 |
| chr3_-_128052166 | 1.88 |
ENST00000648300.1
|
MGLL
|
monoglyceride lipase |
| chr21_+_25639272 | 1.85 |
ENST00000400532.5
ENST00000312957.9 |
JAM2
|
junctional adhesion molecule 2 |
| chr21_+_25639251 | 1.70 |
ENST00000480456.6
|
JAM2
|
junctional adhesion molecule 2 |
| chr10_-_48652493 | 1.60 |
ENST00000435790.6
|
ARHGAP22
|
Rho GTPase activating protein 22 |
| chr1_-_7940825 | 1.31 |
ENST00000377507.8
|
TNFRSF9
|
TNF receptor superfamily member 9 |
| chr11_-_126062782 | 1.20 |
ENST00000531738.6
|
CDON
|
cell adhesion associated, oncogene regulated |
| chr6_-_136525961 | 1.14 |
ENST00000438100.6
|
MAP7
|
microtubule associated protein 7 |
| chr19_-_55738374 | 1.06 |
ENST00000590200.1
ENST00000332836.7 |
NLRP9
|
NLR family pyrin domain containing 9 |
| chr6_+_29100609 | 0.90 |
ENST00000377171.3
|
OR2J1
|
olfactory receptor family 2 subfamily J member 1 |
| chr15_-_55270383 | 0.89 |
ENST00000396307.6
|
RAB27A
|
RAB27A, member RAS oncogene family |
| chr1_-_13201409 | 0.88 |
ENST00000625019.3
|
PRAMEF13
|
PRAME family member 13 |
| chr11_+_34632464 | 0.87 |
ENST00000531794.5
|
EHF
|
ETS homologous factor |
| chr6_-_136526654 | 0.85 |
ENST00000611373.1
|
MAP7
|
microtubule associated protein 7 |
| chr13_-_46105009 | 0.82 |
ENST00000439329.5
ENST00000674625.1 ENST00000181383.10 |
CPB2
|
carboxypeptidase B2 |
| chr14_+_50560137 | 0.80 |
ENST00000358385.12
|
ATL1
|
atlastin GTPase 1 |
| chr6_+_4706133 | 0.75 |
ENST00000328908.9
|
CDYL
|
chromodomain Y like |
| chr3_-_11643871 | 0.71 |
ENST00000430365.7
|
VGLL4
|
vestigial like family member 4 |
| chr8_+_103298433 | 0.68 |
ENST00000522566.5
|
FZD6
|
frizzled class receptor 6 |
| chr6_-_136526472 | 0.65 |
ENST00000454590.5
ENST00000432797.6 |
MAP7
|
microtubule associated protein 7 |
| chr1_+_12857086 | 0.65 |
ENST00000240189.2
|
PRAMEF2
|
PRAME family member 2 |
| chr5_-_16742221 | 0.60 |
ENST00000505695.5
|
MYO10
|
myosin X |
| chr6_-_136526177 | 0.60 |
ENST00000617204.4
|
MAP7
|
microtubule associated protein 7 |
| chr3_+_42809439 | 0.59 |
ENST00000422265.6
ENST00000487368.4 |
ACKR2
ENSG00000273328.5
|
atypical chemokine receptor 2 novel transcript |
| chr4_+_139015751 | 0.57 |
ENST00000280614.4
|
NOCT
|
nocturnin |
| chr12_-_89526253 | 0.56 |
ENST00000547474.1
|
POC1B-GALNT4
|
POC1B-GALNT4 readthrough |
| chr20_+_36236433 | 0.56 |
ENST00000397286.7
ENST00000679519.1 ENST00000679667.1 ENST00000680933.1 ENST00000680247.1 ENST00000320849.9 ENST00000680811.1 ENST00000373932.3 |
AAR2
|
AAR2 splicing factor |
| chr18_+_58149314 | 0.56 |
ENST00000435432.6
ENST00000357895.9 ENST00000586263.5 |
NEDD4L
|
NEDD4 like E3 ubiquitin protein ligase |
| chr19_+_11346556 | 0.55 |
ENST00000587531.5
|
CCDC159
|
coiled-coil domain containing 159 |
| chr14_-_24146596 | 0.54 |
ENST00000560410.5
ENST00000216802.10 ENST00000615264.4 ENST00000630027.1 |
PSME2
|
proteasome activator subunit 2 |
| chr8_-_133060347 | 0.53 |
ENST00000427060.6
|
SLA
|
Src like adaptor |
| chr4_+_88378733 | 0.53 |
ENST00000273960.7
ENST00000380265.9 |
HERC6
|
HECT and RLD domain containing E3 ubiquitin protein ligase family member 6 |
| chr1_+_26410809 | 0.52 |
ENST00000254231.4
ENST00000326279.11 |
LIN28A
|
lin-28 homolog A |
| chr12_-_89526164 | 0.52 |
ENST00000548729.5
|
POC1B-GALNT4
|
POC1B-GALNT4 readthrough |
| chr1_+_169368175 | 0.51 |
ENST00000367808.8
ENST00000426663.1 |
BLZF1
|
basic leucine zipper nuclear factor 1 |
| chr6_+_31587002 | 0.50 |
ENST00000376090.6
|
LST1
|
leukocyte specific transcript 1 |
| chr1_-_46616804 | 0.49 |
ENST00000531769.6
ENST00000319928.8 |
MKNK1
MOB3C
|
MAPK interacting serine/threonine kinase 1 MOB kinase activator 3C |
| chr6_-_138107412 | 0.49 |
ENST00000421351.4
|
PERP
|
p53 apoptosis effector related to PMP22 |
| chr4_+_88378842 | 0.47 |
ENST00000264346.12
|
HERC6
|
HECT and RLD domain containing E3 ubiquitin protein ligase family member 6 |
| chr8_+_98117285 | 0.47 |
ENST00000401707.7
ENST00000522319.5 |
POP1
|
POP1 homolog, ribonuclease P/MRP subunit |
| chr16_+_31108294 | 0.47 |
ENST00000287507.7
ENST00000394950.7 ENST00000219794.11 ENST00000561755.1 |
BCKDK
|
branched chain keto acid dehydrogenase kinase |
| chr16_+_57976435 | 0.47 |
ENST00000290871.10
ENST00000441824.4 |
TEPP
|
testis, prostate and placenta expressed |
| chr3_+_38282294 | 0.46 |
ENST00000466887.5
ENST00000448498.6 |
SLC22A14
|
solute carrier family 22 member 14 |
| chr22_+_30409097 | 0.46 |
ENST00000439838.5
ENST00000439023.3 |
ENSG00000249590.7
|
novel SEC14-like 2 (S. cerevisiae) (SEC14L2) and mitochondrial protein 18 kDa (MTP18) protein |
| chr11_-_34511710 | 0.45 |
ENST00000620316.4
ENST00000312319.6 |
ELF5
|
E74 like ETS transcription factor 5 |
| chr1_-_51325924 | 0.43 |
ENST00000532836.5
ENST00000422925.6 |
TTC39A
|
tetratricopeptide repeat domain 39A |
| chr6_+_31587185 | 0.43 |
ENST00000376092.7
ENST00000376086.7 ENST00000303757.12 ENST00000376093.6 |
LST1
|
leukocyte specific transcript 1 |
| chr12_-_121274510 | 0.43 |
ENST00000392474.6
|
CAMKK2
|
calcium/calmodulin dependent protein kinase kinase 2 |
| chr11_+_64555684 | 0.43 |
ENST00000377585.7
|
SLC22A11
|
solute carrier family 22 member 11 |
| chr11_+_64555956 | 0.43 |
ENST00000377581.7
|
SLC22A11
|
solute carrier family 22 member 11 |
| chr11_+_64555924 | 0.43 |
ENST00000301891.9
|
SLC22A11
|
solute carrier family 22 member 11 |
| chr9_+_12693327 | 0.42 |
ENST00000388918.10
|
TYRP1
|
tyrosinase related protein 1 |
| chr3_+_42091316 | 0.41 |
ENST00000327628.10
|
TRAK1
|
trafficking kinesin protein 1 |
| chr2_+_161136901 | 0.41 |
ENST00000259075.6
ENST00000432002.5 |
TANK
|
TRAF family member associated NFKB activator |
| chr1_-_13285154 | 0.38 |
ENST00000357367.6
ENST00000614831.1 |
PRAMEF8
|
PRAME family member 8 |
| chr16_-_75495396 | 0.38 |
ENST00000332272.9
|
CHST6
|
carbohydrate sulfotransferase 6 |
| chr17_+_7561899 | 0.38 |
ENST00000321337.12
|
SENP3
|
SUMO specific peptidase 3 |
| chr4_+_70050431 | 0.38 |
ENST00000511674.5
ENST00000246896.8 |
HTN1
|
histatin 1 |
| chr1_+_12791397 | 0.38 |
ENST00000332296.7
|
PRAMEF1
|
PRAME family member 1 |
| chr2_-_89213917 | 0.38 |
ENST00000498435.1
|
IGKV1-27
|
immunoglobulin kappa variable 1-27 |
| chr3_-_157533594 | 0.37 |
ENST00000487753.5
ENST00000489602.1 ENST00000461299.5 ENST00000479987.5 |
VEPH1
|
ventricular zone expressed PH domain containing 1 |
| chr6_+_31587268 | 0.36 |
ENST00000396101.7
ENST00000490742.5 |
LST1
|
leukocyte specific transcript 1 |
| chr15_+_73994667 | 0.36 |
ENST00000395135.7
|
PML
|
PML nuclear body scaffold |
| chr6_+_31587049 | 0.36 |
ENST00000376089.6
ENST00000396112.6 |
LST1
|
leukocyte specific transcript 1 |
| chr1_-_108200849 | 0.34 |
ENST00000569674.1
|
SLC25A24
|
solute carrier family 25 member 24 |
| chr6_+_31586680 | 0.34 |
ENST00000339530.8
|
LST1
|
leukocyte specific transcript 1 |
| chr3_+_149474688 | 0.34 |
ENST00000305354.5
ENST00000465758.1 |
TM4SF4
|
transmembrane 4 L six family member 4 |
| chr11_+_55827219 | 0.33 |
ENST00000378397.1
|
OR5L2
|
olfactory receptor family 5 subfamily L member 2 |
| chr15_+_75198866 | 0.33 |
ENST00000562637.1
ENST00000360639.6 |
C15orf39
|
chromosome 15 open reading frame 39 |
| chr13_+_38349822 | 0.33 |
ENST00000379649.5
ENST00000239878.9 |
UFM1
|
ubiquitin fold modifier 1 |
| chr1_+_50106265 | 0.33 |
ENST00000357083.8
|
ELAVL4
|
ELAV like RNA binding protein 4 |
| chr3_+_128052390 | 0.33 |
ENST00000481210.5
ENST00000243253.8 |
SEC61A1
|
SEC61 translocon subunit alpha 1 |
| chr11_-_82901594 | 0.33 |
ENST00000679623.1
|
PRCP
|
prolylcarboxypeptidase |
| chr17_+_48107549 | 0.32 |
ENST00000580219.5
ENST00000452859.6 ENST00000393405.6 |
SNX11
|
sorting nexin 11 |
| chr7_-_75823355 | 0.32 |
ENST00000416943.1
|
CCL24
|
C-C motif chemokine ligand 24 |
| chr4_-_152411734 | 0.32 |
ENST00000603841.1
|
FBXW7
|
F-box and WD repeat domain containing 7 |
| chr2_+_167187364 | 0.31 |
ENST00000672671.1
|
XIRP2
|
xin actin binding repeat containing 2 |
| chr20_-_56525925 | 0.31 |
ENST00000243913.8
|
GCNT7
|
glucosaminyl (N-acetyl) transferase family member 7 |
| chr12_-_54984667 | 0.31 |
ENST00000524668.5
ENST00000533607.1 ENST00000449076.6 |
TESPA1
|
thymocyte expressed, positive selection associated 1 |
| chr5_+_70025534 | 0.30 |
ENST00000515588.1
|
SERF1B
|
small EDRK-rich factor 1B |
| chr2_+_89884740 | 0.30 |
ENST00000509129.1
|
IGKV1D-37
|
immunoglobulin kappa variable 1D-37 (non-functional) |
| chr11_-_82901623 | 0.30 |
ENST00000681637.1
ENST00000679387.1 |
PRCP
|
prolylcarboxypeptidase |
| chr5_-_132737518 | 0.30 |
ENST00000403231.6
ENST00000378735.5 ENST00000618515.4 ENST00000378746.8 |
KIF3A
|
kinesin family member 3A |
| chr11_+_70398404 | 0.30 |
ENST00000346329.7
ENST00000301843.13 ENST00000376561.7 |
CTTN
|
cortactin |
| chr2_+_167187283 | 0.30 |
ENST00000409605.1
ENST00000409273.6 |
XIRP2
|
xin actin binding repeat containing 2 |
| chr12_+_48727431 | 0.29 |
ENST00000548380.6
|
TEX49
|
testis expressed 49 |
| chr11_-_58731936 | 0.29 |
ENST00000344743.8
|
GLYAT
|
glycine-N-acyltransferase |
| chr21_+_34364003 | 0.29 |
ENST00000290310.4
|
KCNE2
|
potassium voltage-gated channel subfamily E regulatory subunit 2 |
| chr9_-_114806031 | 0.29 |
ENST00000374045.5
|
TNFSF15
|
TNF superfamily member 15 |
| chr14_+_34993240 | 0.29 |
ENST00000677647.1
|
SRP54
|
signal recognition particle 54 |
| chr19_+_48606732 | 0.29 |
ENST00000321762.3
|
SPACA4
|
sperm acrosome associated 4 |
| chr10_-_67665642 | 0.28 |
ENST00000682945.1
ENST00000330298.6 ENST00000682758.1 |
CTNNA3
|
catenin alpha 3 |
| chr2_-_89297785 | 0.28 |
ENST00000465170.1
|
IGKV1-37
|
immunoglobulin kappa variable 1-37 (non-functional) |
| chr6_-_43687774 | 0.27 |
ENST00000372133.8
ENST00000372116.5 ENST00000427312.1 |
MRPS18A
|
mitochondrial ribosomal protein S18A |
| chr14_+_58244821 | 0.27 |
ENST00000216455.9
ENST00000412908.6 ENST00000557508.5 |
PSMA3
|
proteasome 20S subunit alpha 3 |
| chrX_-_72875974 | 0.27 |
ENST00000595412.5
|
DMRTC1
|
DMRT like family C1 |
| chr12_-_89526011 | 0.26 |
ENST00000313546.8
|
POC1B
|
POC1 centriolar protein B |
| chr19_-_10928585 | 0.26 |
ENST00000590329.5
ENST00000587943.5 ENST00000586748.6 ENST00000585858.1 ENST00000586575.5 ENST00000253031.6 |
YIPF2
|
Yip1 domain family member 2 |
| chr10_+_112375196 | 0.26 |
ENST00000393081.6
|
ACSL5
|
acyl-CoA synthetase long chain family member 5 |
| chr3_+_119147375 | 0.25 |
ENST00000490594.2
|
ENSG00000251012.2
|
novel chromosome 3 open reading frame 30 (C3orf30) and uroplakin 1B (UPK1B) |
| chr20_+_833781 | 0.25 |
ENST00000381939.1
|
FAM110A
|
family with sequence similarity 110 member A |
| chr2_-_222320124 | 0.25 |
ENST00000678139.1
|
ENSG00000288658.1
|
novel protein, ortholog of Gm2102 (M. musculus) |
| chr6_+_41053194 | 0.25 |
ENST00000244669.3
|
APOBEC2
|
apolipoprotein B mRNA editing enzyme catalytic subunit 2 |
| chr3_+_111998739 | 0.24 |
ENST00000393917.6
ENST00000273368.8 |
TAGLN3
|
transgelin 3 |
| chr7_+_75882044 | 0.24 |
ENST00000428119.1
|
RHBDD2
|
rhomboid domain containing 2 |
| chr11_-_32430811 | 0.24 |
ENST00000379079.8
ENST00000530998.5 |
WT1
|
WT1 transcription factor |
| chr11_+_55262152 | 0.23 |
ENST00000417545.5
|
TRIM48
|
tripartite motif containing 48 |
| chr11_-_5343524 | 0.23 |
ENST00000300773.3
|
OR51B5
|
olfactory receptor family 51 subfamily B member 5 |
| chr3_+_111999189 | 0.23 |
ENST00000455401.6
|
TAGLN3
|
transgelin 3 |
| chr6_+_31586835 | 0.23 |
ENST00000211921.11
|
LST1
|
leukocyte specific transcript 1 |
| chr12_-_51083582 | 0.23 |
ENST00000548206.1
ENST00000546935.5 ENST00000228515.6 ENST00000548981.5 |
CSRNP2
|
cysteine and serine rich nuclear protein 2 |
| chr7_+_117611617 | 0.23 |
ENST00000468795.1
|
CFTR
|
CF transmembrane conductance regulator |
| chr3_+_111998915 | 0.22 |
ENST00000478951.6
|
TAGLN3
|
transgelin 3 |
| chr15_-_72231583 | 0.22 |
ENST00000566809.1
ENST00000567087.5 ENST00000569050.1 ENST00000568883.5 |
PKM
|
pyruvate kinase M1/2 |
| chr17_+_39628496 | 0.22 |
ENST00000394265.5
ENST00000394267.2 |
PPP1R1B
|
protein phosphatase 1 regulatory inhibitor subunit 1B |
| chr2_-_74465162 | 0.22 |
ENST00000649854.1
ENST00000650523.1 ENST00000649601.1 ENST00000448666.7 ENST00000409065.5 ENST00000414701.1 ENST00000452063.7 ENST00000649075.1 ENST00000648810.1 ENST00000462443.2 |
MOGS
|
mannosyl-oligosaccharide glucosidase |
| chrX_-_66040072 | 0.22 |
ENST00000374737.9
|
VSIG4
|
V-set and immunoglobulin domain containing 4 |
| chr19_+_57584131 | 0.22 |
ENST00000536878.6
ENST00000597219.1 ENST00000598689.1 ENST00000597850.2 |
ZIK1
|
zinc finger protein interacting with K protein 1 |
| chrX_-_66040057 | 0.22 |
ENST00000412866.2
|
VSIG4
|
V-set and immunoglobulin domain containing 4 |
| chr19_+_10928777 | 0.22 |
ENST00000270502.7
|
TIMM29
|
translocase of inner mitochondrial membrane 29 |
| chr8_-_51809414 | 0.21 |
ENST00000356297.5
|
PXDNL
|
peroxidasin like |
| chrX_-_66040107 | 0.21 |
ENST00000455586.6
|
VSIG4
|
V-set and immunoglobulin domain containing 4 |
| chr10_-_94362925 | 0.20 |
ENST00000371361.3
|
NOC3L
|
NOC3 like DNA replication regulator |
| chr3_+_178536205 | 0.20 |
ENST00000420517.6
|
KCNMB2
|
potassium calcium-activated channel subfamily M regulatory beta subunit 2 |
| chr5_-_161548124 | 0.20 |
ENST00000520240.5
ENST00000517901.5 ENST00000353437.10 |
GABRB2
|
gamma-aminobutyric acid type A receptor subunit beta2 |
| chr17_+_7627963 | 0.20 |
ENST00000575729.5
ENST00000340624.9 |
SHBG
|
sex hormone binding globulin |
| chr16_+_84768246 | 0.20 |
ENST00000569925.1
ENST00000567526.1 |
USP10
|
ubiquitin specific peptidase 10 |
| chr22_-_37844308 | 0.20 |
ENST00000411961.6
ENST00000434930.1 ENST00000215941.9 |
ANKRD54
|
ankyrin repeat domain 54 |
| chr19_-_42917837 | 0.20 |
ENST00000292125.6
ENST00000402603.8 ENST00000594375.1 ENST00000187910.7 |
PSG6
|
pregnancy specific beta-1-glycoprotein 6 |
| chr19_-_57935353 | 0.19 |
ENST00000595569.1
ENST00000599852.1 ENST00000425570.7 ENST00000601593.5 ENST00000396147.6 |
ZNF418
|
zinc finger protein 418 |
| chr1_+_169367934 | 0.19 |
ENST00000367807.7
ENST00000329281.6 ENST00000420531.1 |
BLZF1
|
basic leucine zipper nuclear factor 1 |
| chr4_-_121381007 | 0.19 |
ENST00000394427.3
|
QRFPR
|
pyroglutamylated RFamide peptide receptor |
| chr14_+_21825453 | 0.19 |
ENST00000390432.2
|
TRAV10
|
T cell receptor alpha variable 10 |
| chr19_+_2867327 | 0.19 |
ENST00000586426.5
ENST00000307635.3 |
ZNF556
|
zinc finger protein 556 |
| chr19_-_38229714 | 0.19 |
ENST00000416611.5
|
DPF1
|
double PHD fingers 1 |
| chr3_-_48188356 | 0.19 |
ENST00000351231.7
ENST00000437972.1 ENST00000302506.8 |
CDC25A
|
cell division cycle 25A |
| chr18_+_21612274 | 0.19 |
ENST00000579618.1
ENST00000300413.10 ENST00000582475.1 |
SNRPD1
|
small nuclear ribonucleoprotein D1 polypeptide |
| chr20_+_18507520 | 0.19 |
ENST00000336714.8
ENST00000646240.1 ENST00000450074.6 ENST00000262544.6 |
SEC23B
|
SEC23 homolog B, COPII coat complex component |
| chr2_-_208190001 | 0.18 |
ENST00000451346.5
ENST00000341287.9 |
C2orf80
|
chromosome 2 open reading frame 80 |
| chr19_-_1848452 | 0.17 |
ENST00000170168.9
|
REXO1
|
RNA exonuclease 1 homolog |
| chr11_-_62653285 | 0.17 |
ENST00000330574.2
|
INTS5
|
integrator complex subunit 5 |
| chr4_+_182448995 | 0.17 |
ENST00000510504.1
|
TENM3
|
teneurin transmembrane protein 3 |
| chr1_+_149475045 | 0.17 |
ENST00000651566.2
|
NBPF19
|
NBPF member 19 |
| chr6_-_35688907 | 0.17 |
ENST00000539068.5
ENST00000357266.9 |
FKBP5
|
FKBP prolyl isomerase 5 |
| chr6_-_31541937 | 0.17 |
ENST00000456662.5
ENST00000431908.5 ENST00000456976.5 ENST00000428450.5 ENST00000418897.5 ENST00000396172.6 ENST00000419020.1 ENST00000428098.5 |
DDX39B
|
DExD-box helicase 39B |
| chr10_+_133394094 | 0.17 |
ENST00000477902.6
|
MTG1
|
mitochondrial ribosome associated GTPase 1 |
| chr15_+_62561361 | 0.17 |
ENST00000561311.5
|
TLN2
|
talin 2 |
| chr15_+_81134257 | 0.17 |
ENST00000286732.5
|
CFAP161
|
cilia and flagella associated protein 161 |
| chr1_+_27726005 | 0.17 |
ENST00000530324.5
ENST00000234549.11 ENST00000373949.5 ENST00000010299.10 |
FAM76A
|
family with sequence similarity 76 member A |
| chr17_+_7012417 | 0.17 |
ENST00000548577.5
|
RNASEK
|
ribonuclease K |
| chr19_+_14440254 | 0.17 |
ENST00000342216.8
|
PKN1
|
protein kinase N1 |
| chr1_+_171512032 | 0.17 |
ENST00000426496.6
|
PRRC2C
|
proline rich coiled-coil 2C |
| chr12_-_54364467 | 0.17 |
ENST00000267015.4
ENST00000551809.1 |
GPR84
|
G protein-coupled receptor 84 |
| chr12_+_57488059 | 0.16 |
ENST00000628866.2
ENST00000262027.10 |
MARS1
|
methionyl-tRNA synthetase 1 |
| chr3_+_44712634 | 0.16 |
ENST00000449836.5
ENST00000296091.8 ENST00000436624.7 ENST00000411443.1 |
ZNF502
|
zinc finger protein 502 |
| chr11_-_62754141 | 0.16 |
ENST00000527994.1
ENST00000394807.5 ENST00000673933.1 |
ZBTB3
|
zinc finger and BTB domain containing 3 |
| chr2_-_113756604 | 0.16 |
ENST00000409342.1
|
SLC35F5
|
solute carrier family 35 member F5 |
| chr11_-_58731968 | 0.16 |
ENST00000278400.3
|
GLYAT
|
glycine-N-acyltransferase |
| chr19_+_18612848 | 0.16 |
ENST00000262817.8
|
TMEM59L
|
transmembrane protein 59 like |
| chr1_+_13254212 | 0.16 |
ENST00000622421.2
|
PRAMEF5
|
PRAME family member 5 |
| chr6_+_31586859 | 0.15 |
ENST00000433492.5
|
LST1
|
leukocyte specific transcript 1 |
| chr16_-_67833842 | 0.15 |
ENST00000566758.5
ENST00000626059.2 ENST00000564817.5 |
CENPT
|
centromere protein T |
| chr1_+_119507203 | 0.15 |
ENST00000369413.8
ENST00000528909.1 |
HSD3B1
|
hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 1 |
| chr19_-_54063905 | 0.15 |
ENST00000645936.1
ENST00000376626.5 ENST00000366170.6 ENST00000425006.3 |
VSTM1
|
V-set and transmembrane domain containing 1 |
| chr8_+_18391276 | 0.15 |
ENST00000286479.4
ENST00000520116.1 |
NAT2
|
N-acetyltransferase 2 |
| chr4_-_867162 | 0.15 |
ENST00000511980.5
ENST00000510799.1 |
GAK
|
cyclin G associated kinase |
| chr3_+_35680994 | 0.15 |
ENST00000441454.5
|
ARPP21
|
cAMP regulated phosphoprotein 21 |
| chr15_+_89088417 | 0.15 |
ENST00000569550.5
ENST00000565066.5 ENST00000565973.5 ENST00000352732.10 |
ABHD2
|
abhydrolase domain containing 2, acylglycerol lipase |
| chr19_+_11346499 | 0.15 |
ENST00000458408.6
ENST00000586451.5 ENST00000588592.5 |
CCDC159
|
coiled-coil domain containing 159 |
| chr3_-_119463606 | 0.15 |
ENST00000319172.10
ENST00000491685.5 ENST00000461654.1 |
TMEM39A
|
transmembrane protein 39A |
| chr21_-_34511243 | 0.14 |
ENST00000399284.1
|
KCNE1
|
potassium voltage-gated channel subfamily E regulatory subunit 1 |
| chr16_+_21684429 | 0.14 |
ENST00000388956.8
|
OTOA
|
otoancorin |
| chr11_+_89924064 | 0.14 |
ENST00000623787.3
|
TRIM49D2
|
tripartite motif containing 49D2 |
| chr6_-_49636832 | 0.14 |
ENST00000371175.10
ENST00000646272.1 ENST00000646939.1 ENST00000618248.3 ENST00000229810.9 ENST00000646963.1 |
RHAG
|
Rh associated glycoprotein |
| chr10_-_101843920 | 0.14 |
ENST00000358038.7
|
KCNIP2
|
potassium voltage-gated channel interacting protein 2 |
| chr19_+_46297032 | 0.14 |
ENST00000377670.9
|
HIF3A
|
hypoxia inducible factor 3 subunit alpha |
| chr1_+_156786875 | 0.14 |
ENST00000526188.5
ENST00000454659.1 |
PRCC
|
proline rich mitotic checkpoint control factor |
| chr2_-_218671975 | 0.14 |
ENST00000295704.7
|
RNF25
|
ring finger protein 25 |
| chr1_-_12947580 | 0.14 |
ENST00000376189.5
|
PRAMEF6
|
PRAME family member 6 |
| chr1_-_154178174 | 0.14 |
ENST00000302206.9
|
TPM3
|
tropomyosin 3 |
| chr10_+_46375721 | 0.13 |
ENST00000616785.1
ENST00000611655.1 ENST00000619162.5 |
ENSG00000285402.1
ANXA8L1
|
novel transcript annexin A8 like 1 |
| chr1_+_209686173 | 0.13 |
ENST00000615289.4
ENST00000367028.6 ENST00000261465.5 |
HSD11B1
|
hydroxysteroid 11-beta dehydrogenase 1 |
| chr12_-_9760893 | 0.13 |
ENST00000228434.7
ENST00000536709.1 |
CD69
|
CD69 molecule |
| chrX_+_76173010 | 0.13 |
ENST00000373357.3
ENST00000373358.8 |
PBDC1
|
polysaccharide biosynthesis domain containing 1 |
| chr7_-_44159278 | 0.13 |
ENST00000345378.7
|
GCK
|
glucokinase |
| chr15_+_80441229 | 0.13 |
ENST00000533983.5
ENST00000527771.5 ENST00000525103.1 |
ARNT2
|
aryl hydrocarbon receptor nuclear translocator 2 |
| chr19_+_36115439 | 0.13 |
ENST00000629269.2
ENST00000221855.8 ENST00000651435.1 ENST00000589996.5 ENST00000591296.1 |
TBCB
|
tubulin folding cofactor B |
| chr8_+_117135259 | 0.13 |
ENST00000519688.5
|
SLC30A8
|
solute carrier family 30 member 8 |
| chr8_-_24956604 | 0.13 |
ENST00000610854.2
|
NEFL
|
neurofilament light |
| chr16_-_4752565 | 0.13 |
ENST00000588099.1
ENST00000588942.1 |
ENSG00000266994.1
ZNF500
|
novel transcript zinc finger protein 500 |
| chr16_+_27203480 | 0.13 |
ENST00000286096.9
|
KDM8
|
lysine demethylase 8 |
| chr19_+_55947832 | 0.13 |
ENST00000291971.7
ENST00000590542.1 |
NLRP8
|
NLR family pyrin domain containing 8 |
| chr6_-_31542339 | 0.12 |
ENST00000458640.5
|
DDX39B
|
DExD-box helicase 39B |
| chr10_-_73096850 | 0.12 |
ENST00000307116.6
ENST00000373008.6 ENST00000394890.7 |
P4HA1
|
prolyl 4-hydroxylase subunit alpha 1 |
| chr19_+_57487711 | 0.12 |
ENST00000354197.8
ENST00000426954.6 ENST00000523882.5 ENST00000520540.5 ENST00000519310.1 ENST00000442920.6 ENST00000523312.1 ENST00000424930.6 |
ZNF419
|
zinc finger protein 419 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 2.6 | GO:0003069 | age-dependent response to oxidative stress(GO:0001306) age-dependent response to reactive oxygen species(GO:0001315) regulation of systemic arterial blood pressure by acetylcholine(GO:0003068) vasodilation by acetylcholine involved in regulation of systemic arterial blood pressure(GO:0003069) regulation of systemic arterial blood pressure by neurotransmitter(GO:0003070) age-dependent general metabolic decline(GO:0007571) |
| 0.2 | 1.3 | GO:2001180 | negative regulation of interleukin-10 secretion(GO:2001180) |
| 0.2 | 1.9 | GO:2000124 | regulation of endocannabinoid signaling pathway(GO:2000124) |
| 0.2 | 0.8 | GO:0003330 | regulation of extracellular matrix constituent secretion(GO:0003330) positive regulation of extracellular matrix constituent secretion(GO:0003331) |
| 0.2 | 0.7 | GO:0035880 | embryonic nail plate morphogenesis(GO:0035880) |
| 0.1 | 0.9 | GO:1903435 | positive regulation of constitutive secretory pathway(GO:1903435) |
| 0.1 | 0.5 | GO:1902159 | regulation of cyclic nucleotide-gated ion channel activity(GO:1902159) |
| 0.1 | 1.3 | GO:0015747 | urate transport(GO:0015747) |
| 0.1 | 0.5 | GO:0010587 | miRNA catabolic process(GO:0010587) |
| 0.1 | 0.4 | GO:0098957 | anterograde axonal transport of mitochondrion(GO:0098957) |
| 0.1 | 0.3 | GO:0086073 | bundle of His cell-Purkinje myocyte adhesion involved in cell communication(GO:0086073) |
| 0.1 | 1.2 | GO:0014816 | skeletal muscle satellite cell differentiation(GO:0014816) |
| 0.1 | 0.3 | GO:2000638 | regulation of SREBP signaling pathway(GO:2000638) negative regulation of SREBP signaling pathway(GO:2000639) |
| 0.1 | 0.6 | GO:0002254 | kinin cascade(GO:0002254) plasma kallikrein-kinin cascade(GO:0002353) |
| 0.1 | 0.5 | GO:0016078 | tRNA catabolic process(GO:0016078) |
| 0.1 | 0.3 | GO:0010387 | COP9 signalosome assembly(GO:0010387) |
| 0.1 | 0.3 | GO:0006617 | SRP-dependent cotranslational protein targeting to membrane, signal sequence recognition(GO:0006617) |
| 0.1 | 0.4 | GO:0061762 | CAMKK-AMPK signaling cascade(GO:0061762) |
| 0.1 | 0.4 | GO:2000158 | positive regulation of ubiquitin-specific protease activity(GO:2000158) |
| 0.1 | 0.5 | GO:0001705 | ectoderm formation(GO:0001705) ectodermal cell fate commitment(GO:0001712) |
| 0.1 | 2.1 | GO:2000651 | positive regulation of sodium ion transmembrane transporter activity(GO:2000651) |
| 0.1 | 0.2 | GO:0072299 | negative regulation of metanephric glomerulus development(GO:0072299) negative regulation of metanephric glomerular mesangial cell proliferation(GO:0072302) |
| 0.1 | 0.2 | GO:0090301 | regulation of neural crest formation(GO:0090299) negative regulation of neural crest formation(GO:0090301) negative regulation of fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:2000314) |
| 0.1 | 0.3 | GO:0039019 | pronephric nephron development(GO:0039019) |
| 0.1 | 0.3 | GO:2000418 | positive regulation of eosinophil migration(GO:2000418) |
| 0.0 | 0.2 | GO:0046833 | snRNA export from nucleus(GO:0006408) positive regulation of RNA export from nucleus(GO:0046833) |
| 0.0 | 0.4 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.0 | 0.6 | GO:2001288 | positive regulation of caveolin-mediated endocytosis(GO:2001288) |
| 0.0 | 0.3 | GO:1990564 | protein polyufmylation(GO:1990564) protein K69-linked ufmylation(GO:1990592) |
| 0.0 | 0.1 | GO:1903937 | response to acrylamide(GO:1903937) |
| 0.0 | 0.2 | GO:0035407 | histone H3-T11 phosphorylation(GO:0035407) |
| 0.0 | 0.2 | GO:0019264 | glycine biosynthetic process from serine(GO:0019264) |
| 0.0 | 0.5 | GO:0002934 | desmosome organization(GO:0002934) |
| 0.0 | 0.1 | GO:0048058 | compound eye corneal lens development(GO:0048058) |
| 0.0 | 0.1 | GO:1905224 | clathrin-coated pit assembly(GO:1905224) |
| 0.0 | 0.6 | GO:0000244 | spliceosomal tri-snRNP complex assembly(GO:0000244) |
| 0.0 | 0.5 | GO:0000290 | deadenylation-dependent decapping of nuclear-transcribed mRNA(GO:0000290) |
| 0.0 | 0.4 | GO:0045345 | positive regulation of MHC class I biosynthetic process(GO:0045345) |
| 0.0 | 0.2 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.0 | 0.1 | GO:0015670 | carbon dioxide transport(GO:0015670) |
| 0.0 | 0.4 | GO:1900378 | positive regulation of melanin biosynthetic process(GO:0048023) positive regulation of secondary metabolite biosynthetic process(GO:1900378) |
| 0.0 | 0.7 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.0 | 0.3 | GO:2000002 | negative regulation of DNA damage checkpoint(GO:2000002) |
| 0.0 | 0.1 | GO:0060557 | positive regulation of vitamin metabolic process(GO:0046136) positive regulation of vitamin D biosynthetic process(GO:0060557) positive regulation of calcidiol 1-monooxygenase activity(GO:0060559) |
| 0.0 | 0.3 | GO:0045163 | clustering of voltage-gated potassium channels(GO:0045163) |
| 0.0 | 0.0 | GO:0061110 | dense core granule biogenesis(GO:0061110) regulation of dense core granule biogenesis(GO:2000705) |
| 0.0 | 0.2 | GO:0015840 | urea transport(GO:0015840) |
| 0.0 | 0.2 | GO:0044789 | modulation by host of viral release from host cell(GO:0044789) positive regulation by host of viral release from host cell(GO:0044791) |
| 0.0 | 0.2 | GO:0042866 | pyruvate biosynthetic process(GO:0042866) |
| 0.0 | 0.4 | GO:0006544 | glycine metabolic process(GO:0006544) |
| 0.0 | 0.1 | GO:0002357 | defense response to tumor cell(GO:0002357) |
| 0.0 | 0.1 | GO:0014738 | regulation of muscle hyperplasia(GO:0014738) |
| 0.0 | 0.1 | GO:0009236 | cobalamin biosynthetic process(GO:0009236) |
| 0.0 | 0.3 | GO:0034454 | microtubule anchoring at centrosome(GO:0034454) |
| 0.0 | 0.2 | GO:0007621 | negative regulation of female receptivity(GO:0007621) |
| 0.0 | 0.1 | GO:0009732 | detection of carbohydrate stimulus(GO:0009730) detection of hexose stimulus(GO:0009732) detection of monosaccharide stimulus(GO:0034287) detection of glucose(GO:0051594) |
| 0.0 | 0.1 | GO:0008218 | bioluminescence(GO:0008218) |
| 0.0 | 0.3 | GO:0016998 | cell wall macromolecule catabolic process(GO:0016998) |
| 0.0 | 2.7 | GO:0006970 | response to osmotic stress(GO:0006970) |
| 0.0 | 0.3 | GO:0006930 | substrate-dependent cell migration, cell extension(GO:0006930) |
| 0.0 | 0.2 | GO:0018401 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) |
| 0.0 | 0.1 | GO:0036496 | regulation of translational initiation by eIF2 alpha dephosphorylation(GO:0036496) |
| 0.0 | 0.2 | GO:1902412 | regulation of mitotic cytokinesis(GO:1902412) |
| 0.0 | 0.5 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.0 | 0.0 | GO:0070409 | carbamoyl phosphate metabolic process(GO:0070408) carbamoyl phosphate biosynthetic process(GO:0070409) response to ammonia(GO:1903717) cellular response to ammonia(GO:1903718) |
| 0.0 | 0.1 | GO:1902775 | mitochondrial large ribosomal subunit assembly(GO:1902775) |
| 0.0 | 0.0 | GO:1900126 | negative regulation of hyaluronan biosynthetic process(GO:1900126) |
| 0.0 | 0.1 | GO:0030450 | regulation of complement activation, classical pathway(GO:0030450) negative regulation of complement activation, classical pathway(GO:0045959) |
| 0.0 | 0.8 | GO:0007029 | endoplasmic reticulum organization(GO:0007029) |
| 0.0 | 1.6 | GO:0050672 | negative regulation of mononuclear cell proliferation(GO:0032945) negative regulation of lymphocyte proliferation(GO:0050672) |
| 0.0 | 0.1 | GO:0030579 | ubiquitin-dependent SMAD protein catabolic process(GO:0030579) |
| 0.0 | 0.2 | GO:0015866 | ADP transport(GO:0015866) |
| 0.0 | 0.5 | GO:0009083 | branched-chain amino acid catabolic process(GO:0009083) |
| 0.0 | 0.3 | GO:0007250 | activation of NF-kappaB-inducing kinase activity(GO:0007250) |
| 0.0 | 0.3 | GO:0006704 | glucocorticoid biosynthetic process(GO:0006704) |
| 0.0 | 0.0 | GO:0006542 | glutamine biosynthetic process(GO:0006542) |
| 0.0 | 0.1 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.0 | 0.1 | GO:1902260 | negative regulation of delayed rectifier potassium channel activity(GO:1902260) |
| 0.0 | 0.7 | GO:0043001 | Golgi to plasma membrane protein transport(GO:0043001) |
| 0.0 | 0.0 | GO:0019355 | nicotinamide nucleotide biosynthetic process from aspartate(GO:0019355) 'de novo' NAD biosynthetic process from aspartate(GO:0034628) |
| 0.0 | 0.2 | GO:0007016 | cytoskeletal anchoring at plasma membrane(GO:0007016) |
| 0.0 | 0.1 | GO:0018344 | protein geranylgeranylation(GO:0018344) |
| 0.0 | 0.0 | GO:1900005 | positive regulation of serine-type endopeptidase activity(GO:1900005) positive regulation of serine-type peptidase activity(GO:1902573) |
| 0.0 | 0.0 | GO:2000397 | regulation of ubiquitin-dependent endocytosis(GO:2000395) positive regulation of ubiquitin-dependent endocytosis(GO:2000397) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.1 | 0.3 | GO:0016939 | kinesin II complex(GO:0016939) |
| 0.1 | 0.5 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.1 | 0.2 | GO:0016222 | procollagen-proline 4-dioxygenase complex(GO:0016222) |
| 0.1 | 6.2 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.1 | 0.5 | GO:0005947 | mitochondrial alpha-ketoglutarate dehydrogenase complex(GO:0005947) |
| 0.1 | 0.9 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 0.1 | 0.8 | GO:0000137 | Golgi cis cisterna(GO:0000137) |
| 0.0 | 0.2 | GO:0005846 | nuclear cap binding complex(GO:0005846) |
| 0.0 | 0.2 | GO:0031205 | endoplasmic reticulum Sec complex(GO:0031205) |
| 0.0 | 0.3 | GO:1990452 | Parkin-FBXW7-Cul1 ubiquitin ligase complex(GO:1990452) |
| 0.0 | 0.4 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.0 | 0.1 | GO:0033596 | TSC1-TSC2 complex(GO:0033596) |
| 0.0 | 0.3 | GO:0005786 | signal recognition particle, endoplasmic reticulum targeting(GO:0005786) |
| 0.0 | 0.2 | GO:0042721 | mitochondrial inner membrane protein insertion complex(GO:0042721) |
| 0.0 | 0.4 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.0 | 0.6 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.0 | 0.3 | GO:1990023 | mitotic spindle midzone(GO:1990023) |
| 0.0 | 0.2 | GO:0034715 | pICln-Sm protein complex(GO:0034715) |
| 0.0 | 0.4 | GO:0033162 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.0 | 0.2 | GO:0070552 | BRISC complex(GO:0070552) |
| 0.0 | 0.1 | GO:0005968 | Rab-protein geranylgeranyltransferase complex(GO:0005968) |
| 0.0 | 0.1 | GO:0034271 | phosphatidylinositol 3-kinase complex, class III, type I(GO:0034271) phosphatidylinositol 3-kinase complex, class III, type II(GO:0034272) |
| 0.0 | 0.2 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.0 | 0.2 | GO:0030868 | smooth endoplasmic reticulum membrane(GO:0030868) smooth endoplasmic reticulum part(GO:0097425) |
| 0.0 | 0.2 | GO:0019773 | proteasome core complex, alpha-subunit complex(GO:0019773) |
| 0.0 | 3.6 | GO:0005923 | bicellular tight junction(GO:0005923) |
| 0.0 | 0.3 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.0 | 0.1 | GO:0005687 | U4 snRNP(GO:0005687) |
| 0.0 | 0.1 | GO:0097524 | sperm plasma membrane(GO:0097524) |
| 0.0 | 0.0 | GO:0097441 | basilar dendrite(GO:0097441) |
| 0.0 | 0.1 | GO:0033063 | Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 0.0 | 0.2 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.0 | 0.2 | GO:0032039 | integrator complex(GO:0032039) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.6 | GO:0004784 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 0.1 | 2.0 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
| 0.1 | 1.3 | GO:1901702 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.1 | 0.2 | GO:0005260 | channel-conductance-controlling ATPase activity(GO:0005260) |
| 0.1 | 0.4 | GO:0047961 | glycine N-acyltransferase activity(GO:0047961) |
| 0.1 | 0.2 | GO:0031751 | D4 dopamine receptor binding(GO:0031751) |
| 0.1 | 0.4 | GO:0035800 | deubiquitinase activator activity(GO:0035800) |
| 0.1 | 0.5 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 0.1 | 0.6 | GO:0060002 | plus-end directed microfilament motor activity(GO:0060002) |
| 0.1 | 0.2 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.1 | 0.4 | GO:0001517 | N-acetylglucosamine 6-O-sulfotransferase activity(GO:0001517) |
| 0.1 | 0.2 | GO:0004060 | arylamine N-acetyltransferase activity(GO:0004060) |
| 0.0 | 0.4 | GO:0016929 | SUMO-specific protease activity(GO:0016929) |
| 0.0 | 0.7 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.0 | 0.1 | GO:0003845 | 11-beta-hydroxysteroid dehydrogenase [NAD(P)] activity(GO:0003845) |
| 0.0 | 0.3 | GO:0030942 | endoplasmic reticulum signal peptide binding(GO:0030942) |
| 0.0 | 0.2 | GO:0035402 | histone kinase activity (H3-T11 specific)(GO:0035402) |
| 0.0 | 0.2 | GO:0008732 | glycine hydroxymethyltransferase activity(GO:0004372) threonine aldolase activity(GO:0004793) L-allo-threonine aldolase activity(GO:0008732) |
| 0.0 | 0.5 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.0 | 0.3 | GO:0050816 | phosphothreonine binding(GO:0050816) |
| 0.0 | 0.2 | GO:0044729 | hemi-methylated DNA-binding(GO:0044729) |
| 0.0 | 0.5 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 0.0 | 0.2 | GO:0004769 | steroid delta-isomerase activity(GO:0004769) |
| 0.0 | 1.3 | GO:0005031 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.0 | 0.2 | GO:0004656 | procollagen-proline 4-dioxygenase activity(GO:0004656) |
| 0.0 | 0.3 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) ER retention sequence binding(GO:0046923) |
| 0.0 | 0.3 | GO:0086083 | cell adhesive protein binding involved in bundle of His cell-Purkinje myocyte communication(GO:0086083) |
| 0.0 | 0.2 | GO:1990446 | U1 snRNP binding(GO:1990446) |
| 0.0 | 0.2 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.0 | 0.9 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.0 | 0.4 | GO:1902282 | voltage-gated potassium channel activity involved in ventricular cardiac muscle cell action potential repolarization(GO:1902282) |
| 0.0 | 0.2 | GO:0005497 | androgen binding(GO:0005497) |
| 0.0 | 0.2 | GO:0004966 | galanin receptor activity(GO:0004966) |
| 0.0 | 0.6 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.0 | 0.3 | GO:0030621 | U4 snRNA binding(GO:0030621) |
| 0.0 | 0.7 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.0 | 0.5 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.0 | 0.1 | GO:0015265 | urea channel activity(GO:0015265) |
| 0.0 | 0.7 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.0 | 0.1 | GO:0086059 | voltage-gated calcium channel activity involved SA node cell action potential(GO:0086059) |
| 0.0 | 0.6 | GO:0019870 | potassium channel inhibitor activity(GO:0019870) |
| 0.0 | 0.1 | GO:0032184 | SUMO polymer binding(GO:0032184) |
| 0.0 | 0.5 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 0.4 | GO:0032183 | SUMO binding(GO:0032183) |
| 0.0 | 0.2 | GO:0004983 | neuropeptide Y receptor activity(GO:0004983) |
| 0.0 | 0.5 | GO:0015101 | organic cation transmembrane transporter activity(GO:0015101) |
| 0.0 | 0.1 | GO:0031728 | CCR3 chemokine receptor binding(GO:0031728) |
| 0.0 | 0.2 | GO:0005237 | inhibitory extracellular ligand-gated ion channel activity(GO:0005237) |
| 0.0 | 0.1 | GO:0015562 | efflux transmembrane transporter activity(GO:0015562) |
| 0.0 | 0.3 | GO:0003796 | lysozyme activity(GO:0003796) |
| 0.0 | 0.1 | GO:0004663 | Rab geranylgeranyltransferase activity(GO:0004663) |
| 0.0 | 0.2 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.0 | 0.3 | GO:0005522 | profilin binding(GO:0005522) |
| 0.0 | 0.1 | GO:0032296 | ribonuclease III activity(GO:0004525) double-stranded RNA-specific ribonuclease activity(GO:0032296) |
| 0.0 | 0.4 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.0 | 0.2 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) ADP transmembrane transporter activity(GO:0015217) |
| 0.0 | 0.1 | GO:0008865 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.0 | 0.1 | GO:0016404 | 15-hydroxyprostaglandin dehydrogenase (NAD+) activity(GO:0016404) |
| 0.0 | 0.1 | GO:0030492 | hemoglobin binding(GO:0030492) |
| 0.0 | 0.0 | GO:0004087 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 0.0 | 0.0 | GO:0004356 | glutamate-ammonia ligase activity(GO:0004356) ammonia ligase activity(GO:0016211) acid-ammonia (or amide) ligase activity(GO:0016880) |
| 0.0 | 0.3 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.0 | 0.0 | GO:0000309 | nicotinamide-nucleotide adenylyltransferase activity(GO:0000309) |
| 0.0 | 0.1 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.5 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.0 | 1.2 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.0 | 2.5 | PID FOXO PATHWAY | FoxO family signaling |
| 0.0 | 0.7 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.0 | 0.7 | PID MYC PATHWAY | C-MYC pathway |
| 0.0 | 1.3 | PID CD8 TCR DOWNSTREAM PATHWAY | Downstream signaling in naïve CD8+ T cells |
| 0.0 | 0.2 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.0 | 0.3 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.0 | 0.6 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.3 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 0.0 | 1.7 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.0 | 3.4 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.0 | 0.8 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.0 | 3.5 | REACTOME CELL SURFACE INTERACTIONS AT THE VASCULAR WALL | Genes involved in Cell surface interactions at the vascular wall |
| 0.0 | 0.2 | REACTOME P53 INDEPENDENT G1 S DNA DAMAGE CHECKPOINT | Genes involved in p53-Independent G1/S DNA damage checkpoint |
| 0.0 | 0.5 | REACTOME SPRY REGULATION OF FGF SIGNALING | Genes involved in Spry regulation of FGF signaling |
| 0.0 | 1.2 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 1.1 | REACTOME ER PHAGOSOME PATHWAY | Genes involved in ER-Phagosome pathway |
| 0.0 | 0.2 | REACTOME N GLYCAN TRIMMING IN THE ER AND CALNEXIN CALRETICULIN CYCLE | Genes involved in N-glycan trimming in the ER and Calnexin/Calreticulin cycle |
| 0.0 | 0.1 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.0 | 0.2 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.0 | 0.4 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION | Genes involved in TRAF6 mediated IRF7 activation |
| 0.0 | 0.4 | REACTOME KERATAN SULFATE BIOSYNTHESIS | Genes involved in Keratan sulfate biosynthesis |
| 0.0 | 0.2 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.0 | 0.2 | REACTOME ANDROGEN BIOSYNTHESIS | Genes involved in Androgen biosynthesis |