Project
Inflammatory response time course, HUVEC (Wada, 2009)
Navigation
Downloads
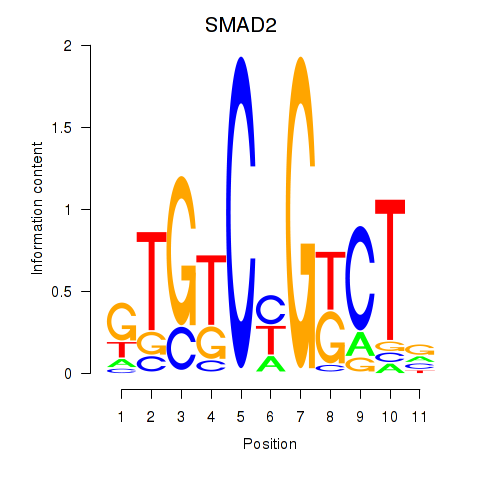
Results for SMAD2
Z-value: 0.63
Transcription factors associated with SMAD2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SMAD2
|
ENSG00000175387.16 | SMAD2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SMAD2 | hg38_v1_chr18_-_47931107_47931146 | -0.30 | 1.4e-01 | Click! |
Activity profile of SMAD2 motif
Sorted Z-values of SMAD2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of SMAD2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_99646025 | 4.26 |
ENST00000263174.9
ENST00000605497.5 ENST00000615664.1 |
PALMD
|
palmdelphin |
| chr12_+_53050179 | 4.04 |
ENST00000546602.5
ENST00000552570.5 ENST00000549700.5 |
TNS2
|
tensin 2 |
| chr18_+_23135452 | 2.34 |
ENST00000580153.5
ENST00000256925.12 |
CABLES1
|
Cdk5 and Abl enzyme substrate 1 |
| chr12_+_53050014 | 2.06 |
ENST00000314250.11
|
TNS2
|
tensin 2 |
| chr11_-_33892010 | 1.57 |
ENST00000257818.3
|
LMO2
|
LIM domain only 2 |
| chr21_+_44600597 | 1.43 |
ENST00000609664.2
|
KRTAP10-7
|
keratin associated protein 10-7 |
| chr1_+_185734362 | 1.37 |
ENST00000271588.9
|
HMCN1
|
hemicentin 1 |
| chr10_-_124093582 | 1.27 |
ENST00000462406.1
ENST00000435907.6 |
CHST15
|
carbohydrate sulfotransferase 15 |
| chr8_+_94895763 | 1.16 |
ENST00000523378.5
|
NDUFAF6
|
NADH:ubiquinone oxidoreductase complex assembly factor 6 |
| chr22_+_19718390 | 1.11 |
ENST00000383045.7
ENST00000438754.6 |
SEPTIN5
|
septin 5 |
| chrX_-_63754664 | 0.86 |
ENST00000677315.1
ENST00000636392.1 ENST00000637040.1 ENST00000637178.1 ENST00000637557.1 ENST00000636048.1 ENST00000638021.1 ENST00000672513.1 |
ENSG00000288661.1
ARHGEF9
|
novel protein Cdc42 guanine nucleotide exchange factor 9 |
| chr3_+_194685874 | 0.77 |
ENST00000329759.6
|
FAM43A
|
family with sequence similarity 43 member A |
| chr21_+_44573724 | 0.77 |
ENST00000622352.3
ENST00000400374.4 ENST00000616689.2 |
KRTAP10-4
|
keratin associated protein 10-4 |
| chr11_+_68312542 | 0.77 |
ENST00000294304.12
|
LRP5
|
LDL receptor related protein 5 |
| chr20_+_32010429 | 0.77 |
ENST00000452892.3
ENST00000262659.12 |
CCM2L
|
CCM2 like scaffold protein |
| chr11_+_59755427 | 0.76 |
ENST00000529177.5
|
STX3
|
syntaxin 3 |
| chr20_-_23086316 | 0.73 |
ENST00000246006.5
|
CD93
|
CD93 molecule |
| chr4_-_113761441 | 0.71 |
ENST00000394524.7
|
CAMK2D
|
calcium/calmodulin dependent protein kinase II delta |
| chr17_+_7888783 | 0.69 |
ENST00000330494.12
ENST00000358181.8 |
CHD3
|
chromodomain helicase DNA binding protein 3 |
| chr19_+_735026 | 0.69 |
ENST00000592155.5
ENST00000590161.2 |
PALM
|
paralemmin |
| chr5_-_88731827 | 0.68 |
ENST00000627170.2
|
MEF2C
|
myocyte enhancer factor 2C |
| chr3_+_52495330 | 0.66 |
ENST00000321725.10
|
STAB1
|
stabilin 1 |
| chr22_+_31122923 | 0.61 |
ENST00000620191.4
ENST00000412277.6 ENST00000412985.5 ENST00000331075.10 ENST00000420017.5 ENST00000400294.6 ENST00000405300.5 ENST00000404390.7 |
INPP5J
|
inositol polyphosphate-5-phosphatase J |
| chr1_-_58784035 | 0.58 |
ENST00000371222.4
|
JUN
|
Jun proto-oncogene, AP-1 transcription factor subunit |
| chr6_+_32164586 | 0.57 |
ENST00000333845.11
ENST00000395512.5 ENST00000432129.1 |
EGFL8
|
EGF like domain multiple 8 |
| chr4_-_113761724 | 0.57 |
ENST00000511664.6
|
CAMK2D
|
calcium/calmodulin dependent protein kinase II delta |
| chr15_+_96332432 | 0.57 |
ENST00000559679.1
ENST00000394171.6 |
NR2F2
|
nuclear receptor subfamily 2 group F member 2 |
| chr1_+_2050387 | 0.55 |
ENST00000378567.8
|
PRKCZ
|
protein kinase C zeta |
| chr22_+_50089879 | 0.54 |
ENST00000545383.5
ENST00000262794.10 |
MOV10L1
|
Mov10 like RISC complex RNA helicase 1 |
| chr3_+_111674654 | 0.53 |
ENST00000636933.1
ENST00000393934.7 ENST00000477665.2 |
PLCXD2
|
phosphatidylinositol specific phospholipase C X domain containing 2 |
| chr1_-_94121105 | 0.53 |
ENST00000649773.1
ENST00000370225.4 |
ABCA4
|
ATP binding cassette subfamily A member 4 |
| chr7_-_130441136 | 0.53 |
ENST00000675596.1
ENST00000676312.1 |
CEP41
|
centrosomal protein 41 |
| chr5_-_74640575 | 0.52 |
ENST00000651128.1
|
ENC1
|
ectodermal-neural cortex 1 |
| chrX_+_153494970 | 0.51 |
ENST00000331595.9
ENST00000431891.1 |
BGN
|
biglycan |
| chr3_+_124094663 | 0.50 |
ENST00000460856.5
ENST00000240874.7 |
KALRN
|
kalirin RhoGEF kinase |
| chr9_+_125747345 | 0.49 |
ENST00000342287.9
ENST00000373489.10 ENST00000373487.8 |
PBX3
|
PBX homeobox 3 |
| chr1_+_37474572 | 0.47 |
ENST00000373087.7
|
ZC3H12A
|
zinc finger CCCH-type containing 12A |
| chr19_+_49591170 | 0.47 |
ENST00000418929.7
|
PRR12
|
proline rich 12 |
| chr1_+_26159071 | 0.46 |
ENST00000374268.5
|
FAM110D
|
family with sequence similarity 110 member D |
| chr22_+_22822658 | 0.44 |
ENST00000620395.2
|
IGLV2-8
|
immunoglobulin lambda variable 2-8 |
| chr4_-_113761068 | 0.41 |
ENST00000342666.9
ENST00000514328.5 ENST00000515496.5 ENST00000508738.5 ENST00000379773.6 |
CAMK2D
|
calcium/calmodulin dependent protein kinase II delta |
| chr11_-_5441514 | 0.41 |
ENST00000380211.1
|
OR51I1
|
olfactory receptor family 51 subfamily I member 1 |
| chr19_+_48624952 | 0.41 |
ENST00000599748.5
ENST00000599029.2 |
SPHK2
|
sphingosine kinase 2 |
| chr2_-_240682879 | 0.40 |
ENST00000407834.4
ENST00000621682.4 |
AQP12B
|
aquaporin 12B |
| chr11_+_66857056 | 0.40 |
ENST00000309602.5
ENST00000393952.3 |
LRFN4
|
leucine rich repeat and fibronectin type III domain containing 4 |
| chr21_+_44697172 | 0.40 |
ENST00000400365.3
|
KRTAP10-12
|
keratin associated protein 10-12 |
| chr1_-_206003385 | 0.40 |
ENST00000617070.5
|
RAB7B
|
RAB7B, member RAS oncogene family |
| chr19_+_49513154 | 0.40 |
ENST00000426395.7
ENST00000600273.5 ENST00000599988.5 |
FCGRT
|
Fc fragment of IgG receptor and transporter |
| chr22_-_38455199 | 0.39 |
ENST00000303592.3
|
KCNJ4
|
potassium inwardly rectifying channel subfamily J member 4 |
| chr22_+_37560472 | 0.39 |
ENST00000430687.1
ENST00000249014.5 |
CDC42EP1
|
CDC42 effector protein 1 |
| chr14_-_104978360 | 0.39 |
ENST00000333244.6
|
AHNAK2
|
AHNAK nucleoprotein 2 |
| chr22_-_20016807 | 0.38 |
ENST00000263207.8
|
ARVCF
|
ARVCF delta catenin family member |
| chr17_-_78925376 | 0.38 |
ENST00000262768.11
|
TIMP2
|
TIMP metallopeptidase inhibitor 2 |
| chr3_+_124094696 | 0.38 |
ENST00000360013.7
ENST00000684186.1 ENST00000684276.1 |
KALRN
|
kalirin RhoGEF kinase |
| chr1_+_43946905 | 0.37 |
ENST00000372343.8
|
IPO13
|
importin 13 |
| chr20_+_58651785 | 0.36 |
ENST00000358029.8
|
STX16
|
syntaxin 16 |
| chr12_-_12266769 | 0.36 |
ENST00000543091.1
|
LRP6
|
LDL receptor related protein 6 |
| chr16_+_55479188 | 0.36 |
ENST00000219070.9
|
MMP2
|
matrix metallopeptidase 2 |
| chr19_-_41353904 | 0.35 |
ENST00000221930.6
|
TGFB1
|
transforming growth factor beta 1 |
| chr11_+_9573633 | 0.35 |
ENST00000450114.7
|
WEE1
|
WEE1 G2 checkpoint kinase |
| chr5_-_151686908 | 0.35 |
ENST00000231061.9
|
SPARC
|
secreted protein acidic and cysteine rich |
| chr9_+_122519141 | 0.31 |
ENST00000340750.1
|
OR1J4
|
olfactory receptor family 1 subfamily J member 4 |
| chr1_-_44031352 | 0.31 |
ENST00000372306.7
ENST00000475075.6 |
SLC6A9
|
solute carrier family 6 member 9 |
| chr1_-_234479131 | 0.30 |
ENST00000040877.2
|
TARBP1
|
TAR (HIV-1) RNA binding protein 1 |
| chr12_-_55712402 | 0.30 |
ENST00000452168.6
|
ITGA7
|
integrin subunit alpha 7 |
| chr17_-_41424583 | 0.29 |
ENST00000225550.4
|
KRT37
|
keratin 37 |
| chr5_+_43603163 | 0.29 |
ENST00000660752.1
ENST00000654405.1 ENST00000344920.9 ENST00000657172.1 ENST00000512996.6 ENST00000671668.1 |
NNT
|
nicotinamide nucleotide transhydrogenase |
| chr6_+_43132472 | 0.29 |
ENST00000489707.5
|
PTK7
|
protein tyrosine kinase 7 (inactive) |
| chr12_+_103930600 | 0.29 |
ENST00000680316.1
ENST00000679861.1 |
HSP90B1
|
heat shock protein 90 beta family member 1 |
| chr1_-_154961720 | 0.29 |
ENST00000368457.3
|
PYGO2
|
pygopus family PHD finger 2 |
| chr2_-_20225433 | 0.28 |
ENST00000381150.5
|
SDC1
|
syndecan 1 |
| chr1_-_2801717 | 0.28 |
ENST00000401095.8
|
TTC34
|
tetratricopeptide repeat domain 34 |
| chrX_+_153764178 | 0.28 |
ENST00000538966.5
|
PLXNB3
|
plexin B3 |
| chr13_-_41961078 | 0.28 |
ENST00000379310.8
ENST00000281496.6 |
VWA8
|
von Willebrand factor A domain containing 8 |
| chrX_-_107775740 | 0.27 |
ENST00000372383.9
|
TSC22D3
|
TSC22 domain family member 3 |
| chrX_+_153764233 | 0.27 |
ENST00000361971.10
|
PLXNB3
|
plexin B3 |
| chr21_-_46285608 | 0.27 |
ENST00000291688.6
|
MCM3AP
|
minichromosome maintenance complex component 3 associated protein |
| chr16_+_53130921 | 0.27 |
ENST00000564845.5
|
CHD9
|
chromodomain helicase DNA binding protein 9 |
| chr3_-_195876635 | 0.26 |
ENST00000672669.1
ENST00000672886.1 ENST00000672098.1 ENST00000671767.1 ENST00000672548.1 |
TNK2
|
tyrosine kinase non receptor 2 |
| chr9_+_122264603 | 0.26 |
ENST00000297908.7
|
MRRF
|
mitochondrial ribosome recycling factor |
| chr6_-_159045010 | 0.25 |
ENST00000338313.5
|
TAGAP
|
T cell activation RhoGTPase activating protein |
| chr22_+_22327298 | 0.24 |
ENST00000390291.2
|
IGLV1-50
|
immunoglobulin lambda variable 1-50 (non-functional) |
| chr14_-_106038355 | 0.23 |
ENST00000390597.3
|
IGHV2-5
|
immunoglobulin heavy variable 2-5 |
| chr6_+_34514853 | 0.22 |
ENST00000538621.2
|
PACSIN1
|
protein kinase C and casein kinase substrate in neurons 1 |
| chr19_-_49362376 | 0.22 |
ENST00000601519.5
ENST00000593945.6 ENST00000539846.5 ENST00000596757.1 ENST00000311227.6 |
TEAD2
|
TEA domain transcription factor 2 |
| chr11_-_75669028 | 0.22 |
ENST00000304771.8
|
MAP6
|
microtubule associated protein 6 |
| chr6_-_27873525 | 0.22 |
ENST00000618305.2
|
H4C13
|
H4 clustered histone 13 |
| chrX_-_107775951 | 0.22 |
ENST00000315660.8
ENST00000372384.6 ENST00000502650.1 ENST00000506724.1 |
TSC22D3
|
TSC22 domain family member 3 |
| chr19_-_55069488 | 0.22 |
ENST00000396247.7
ENST00000586945.5 ENST00000587026.1 |
RDH13
|
retinol dehydrogenase 13 |
| chr17_+_68205453 | 0.21 |
ENST00000674770.2
|
AMZ2
|
archaelysin family metallopeptidase 2 |
| chr12_+_6927946 | 0.21 |
ENST00000396684.3
|
ATN1
|
atrophin 1 |
| chr1_-_1307861 | 0.21 |
ENST00000354700.10
|
ACAP3
|
ArfGAP with coiled-coil, ankyrin repeat and PH domains 3 |
| chr3_-_195442977 | 0.21 |
ENST00000326793.11
|
ACAP2
|
ArfGAP with coiled-coil, ankyrin repeat and PH domains 2 |
| chr17_+_75516514 | 0.21 |
ENST00000333213.11
ENST00000545228.3 ENST00000680999.1 |
TSEN54
|
tRNA splicing endonuclease subunit 54 |
| chr1_-_151146611 | 0.21 |
ENST00000341697.7
ENST00000368914.8 |
SEMA6C
|
semaphorin 6C |
| chr10_+_103555124 | 0.20 |
ENST00000437579.1
|
NEURL1
|
neuralized E3 ubiquitin protein ligase 1 |
| chr5_+_175871670 | 0.20 |
ENST00000514150.5
|
CPLX2
|
complexin 2 |
| chr7_-_10940123 | 0.20 |
ENST00000339600.6
|
NDUFA4
|
NDUFA4 mitochondrial complex associated |
| chr6_-_56954747 | 0.20 |
ENST00000680361.1
|
DST
|
dystonin |
| chr7_-_150955796 | 0.19 |
ENST00000330883.9
|
KCNH2
|
potassium voltage-gated channel subfamily H member 2 |
| chr4_-_39638893 | 0.19 |
ENST00000511809.5
ENST00000505729.1 |
SMIM14
|
small integral membrane protein 14 |
| chr1_-_151146643 | 0.19 |
ENST00000613223.1
|
SEMA6C
|
semaphorin 6C |
| chr12_-_57016517 | 0.19 |
ENST00000441881.5
ENST00000458521.7 |
TAC3
|
tachykinin precursor 3 |
| chrX_-_135098695 | 0.18 |
ENST00000433425.4
|
SMIM10L2B
|
small integral membrane protein 10 like 2B |
| chr7_-_105876575 | 0.18 |
ENST00000318724.8
ENST00000419735.8 |
ATXN7L1
|
ataxin 7 like 1 |
| chr19_+_48469354 | 0.18 |
ENST00000452733.7
ENST00000641098.1 |
CYTH2
|
cytohesin 2 |
| chr19_+_35936360 | 0.17 |
ENST00000246529.4
|
LRFN3
|
leucine rich repeat and fibronectin type III domain containing 3 |
| chr1_-_206946448 | 0.17 |
ENST00000356495.5
|
PIGR
|
polymeric immunoglobulin receptor |
| chr20_+_1895365 | 0.17 |
ENST00000358771.5
|
SIRPA
|
signal regulatory protein alpha |
| chr16_+_90022600 | 0.16 |
ENST00000620723.4
ENST00000268699.9 |
GAS8
|
growth arrest specific 8 |
| chr1_-_206003442 | 0.16 |
ENST00000623893.1
|
RAB7B
|
RAB7B, member RAS oncogene family |
| chr3_-_168095885 | 0.16 |
ENST00000470487.6
|
GOLIM4
|
golgi integral membrane protein 4 |
| chrX_-_30577759 | 0.16 |
ENST00000378962.4
|
TASL
|
TLR adaptor interacting with endolysosomal SLC15A4 |
| chr16_+_57668299 | 0.16 |
ENST00000333493.9
ENST00000450388.7 |
ADGRG3
|
adhesion G protein-coupled receptor G3 |
| chr3_-_51903341 | 0.16 |
ENST00000310914.10
|
IQCF1
|
IQ motif containing F1 |
| chr5_-_43412323 | 0.16 |
ENST00000361115.4
|
CCL28
|
C-C motif chemokine ligand 28 |
| chr4_-_39638846 | 0.16 |
ENST00000295958.10
|
SMIM14
|
small integral membrane protein 14 |
| chr19_+_36140059 | 0.15 |
ENST00000246533.8
ENST00000587718.5 ENST00000592483.5 ENST00000590874.5 ENST00000588815.5 |
CAPNS1
|
calpain small subunit 1 |
| chr6_+_32970888 | 0.15 |
ENST00000374831.8
|
BRD2
|
bromodomain containing 2 |
| chr9_-_133131109 | 0.15 |
ENST00000372062.8
|
RALGDS
|
ral guanine nucleotide dissociation stimulator |
| chrX_+_102651366 | 0.14 |
ENST00000415986.5
ENST00000444152.5 ENST00000361600.9 |
GPRASP1
|
G protein-coupled receptor associated sorting protein 1 |
| chr1_+_36306359 | 0.14 |
ENST00000453908.8
|
SH3D21
|
SH3 domain containing 21 |
| chr20_+_64091954 | 0.14 |
ENST00000672146.2
|
OPRL1
|
opioid related nociceptin receptor 1 |
| chr16_+_24729641 | 0.14 |
ENST00000395799.8
|
TNRC6A
|
trinucleotide repeat containing adaptor 6A |
| chr11_-_66317037 | 0.14 |
ENST00000311330.4
|
CD248
|
CD248 molecule |
| chr5_+_64768921 | 0.14 |
ENST00000381070.8
ENST00000508024.1 |
CWC27
|
CWC27 spliceosome associated cyclophilin |
| chr7_-_99408548 | 0.13 |
ENST00000626285.1
ENST00000350498.8 |
PDAP1
|
PDGFA associated protein 1 |
| chr2_-_227164194 | 0.13 |
ENST00000396625.5
|
COL4A4
|
collagen type IV alpha 4 chain |
| chr19_-_51458448 | 0.13 |
ENST00000430817.5
ENST00000321424.7 ENST00000340550.5 |
SIGLEC8
|
sialic acid binding Ig like lectin 8 |
| chr6_-_143450662 | 0.13 |
ENST00000237283.9
|
ADAT2
|
adenosine deaminase tRNA specific 2 |
| chr7_-_102616692 | 0.13 |
ENST00000521076.5
ENST00000462172.5 ENST00000522801.5 ENST00000262940.12 ENST00000449970.6 |
RASA4
|
RAS p21 protein activator 4 |
| chr19_-_14114156 | 0.13 |
ENST00000589994.6
|
PRKACA
|
protein kinase cAMP-activated catalytic subunit alpha |
| chr12_+_109900518 | 0.13 |
ENST00000312777.9
ENST00000536408.2 |
TCHP
|
trichoplein keratin filament binding |
| chr9_+_122510802 | 0.12 |
ENST00000335302.5
|
OR1J2
|
olfactory receptor family 1 subfamily J member 2 |
| chr18_+_13218195 | 0.12 |
ENST00000679167.1
|
LDLRAD4
|
low density lipoprotein receptor class A domain containing 4 |
| chr3_-_88059042 | 0.12 |
ENST00000309534.10
|
CGGBP1
|
CGG triplet repeat binding protein 1 |
| chr7_-_102517755 | 0.12 |
ENST00000306682.6
ENST00000465829.6 ENST00000541662.5 |
RASA4B
|
RAS p21 protein activator 4B |
| chr3_-_88058928 | 0.12 |
ENST00000482016.6
|
CGGBP1
|
CGG triplet repeat binding protein 1 |
| chr9_+_79572715 | 0.12 |
ENST00000265284.10
|
TLE4
|
TLE family member 4, transcriptional corepressor |
| chr1_+_15456727 | 0.11 |
ENST00000359621.5
|
CELA2A
|
chymotrypsin like elastase 2A |
| chr17_+_7407838 | 0.11 |
ENST00000302926.7
|
NLGN2
|
neuroligin 2 |
| chr22_+_44677044 | 0.11 |
ENST00000006251.11
|
PRR5
|
proline rich 5 |
| chr5_+_175871570 | 0.11 |
ENST00000512824.5
ENST00000393745.8 |
CPLX2
|
complexin 2 |
| chr11_-_117295485 | 0.11 |
ENST00000680971.1
|
BACE1
|
beta-secretase 1 |
| chr11_+_75562056 | 0.11 |
ENST00000533603.5
|
SERPINH1
|
serpin family H member 1 |
| chr4_+_37244735 | 0.11 |
ENST00000309447.6
|
NWD2
|
NACHT and WD repeat domain containing 2 |
| chr19_+_16888991 | 0.11 |
ENST00000248076.4
|
F2RL3
|
F2R like thrombin or trypsin receptor 3 |
| chr1_+_36307090 | 0.10 |
ENST00000505871.7
|
SH3D21
|
SH3 domain containing 21 |
| chr17_-_42609356 | 0.10 |
ENST00000309428.10
|
RETREG3
|
reticulophagy regulator family member 3 |
| chr5_+_134526176 | 0.10 |
ENST00000681820.1
ENST00000512386.6 ENST00000612830.2 |
JADE2
|
jade family PHD finger 2 |
| chr22_+_44677077 | 0.10 |
ENST00000403581.5
|
PRR5
|
proline rich 5 |
| chr16_+_28878382 | 0.10 |
ENST00000357084.7
|
ATP2A1
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 1 |
| chr7_-_105876477 | 0.10 |
ENST00000478915.1
|
ATXN7L1
|
ataxin 7 like 1 |
| chr1_+_159171607 | 0.09 |
ENST00000368124.8
ENST00000368125.9 ENST00000416746.1 |
CADM3
|
cell adhesion molecule 3 |
| chr1_+_32362537 | 0.09 |
ENST00000373534.4
|
TSSK3
|
testis specific serine kinase 3 |
| chr17_-_10114546 | 0.09 |
ENST00000323816.8
|
GAS7
|
growth arrest specific 7 |
| chr9_+_79573162 | 0.09 |
ENST00000425506.5
|
TLE4
|
TLE family member 4, transcriptional corepressor |
| chr6_+_39792298 | 0.09 |
ENST00000633794.1
ENST00000274867.9 |
DAAM2
|
dishevelled associated activator of morphogenesis 2 |
| chr16_-_4937064 | 0.08 |
ENST00000590782.6
ENST00000345988.7 |
PPL
|
periplakin |
| chr8_-_143617457 | 0.08 |
ENST00000529048.5
ENST00000529064.5 |
GFUS
|
GDP-L-fucose synthase |
| chr5_+_180494344 | 0.08 |
ENST00000261951.9
|
CNOT6
|
CCR4-NOT transcription complex subunit 6 |
| chr16_+_28878480 | 0.08 |
ENST00000395503.9
|
ATP2A1
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 1 |
| chr1_-_41242106 | 0.07 |
ENST00000337495.9
ENST00000372597.5 ENST00000372596.5 |
SCMH1
|
Scm polycomb group protein homolog 1 |
| chr19_-_35135011 | 0.07 |
ENST00000310123.8
|
LGI4
|
leucine rich repeat LGI family member 4 |
| chr10_+_18261080 | 0.07 |
ENST00000377319.8
ENST00000377331.8 |
CACNB2
|
calcium voltage-gated channel auxiliary subunit beta 2 |
| chr6_+_39792993 | 0.07 |
ENST00000538976.5
|
DAAM2
|
dishevelled associated activator of morphogenesis 2 |
| chr16_+_2033264 | 0.07 |
ENST00000565855.5
ENST00000566198.1 |
SLC9A3R2
|
SLC9A3 regulator 2 |
| chr7_+_102363874 | 0.07 |
ENST00000496391.5
|
PRKRIP1
|
PRKR interacting protein 1 |
| chr7_-_155809072 | 0.07 |
ENST00000430104.5
|
SHH
|
sonic hedgehog signaling molecule |
| chr5_+_134526100 | 0.07 |
ENST00000395003.5
|
JADE2
|
jade family PHD finger 2 |
| chr11_+_14643782 | 0.07 |
ENST00000282096.9
|
PDE3B
|
phosphodiesterase 3B |
| chr9_+_101028721 | 0.06 |
ENST00000374874.8
|
PLPPR1
|
phospholipid phosphatase related 1 |
| chr21_-_33479914 | 0.06 |
ENST00000542230.7
|
TMEM50B
|
transmembrane protein 50B |
| chr8_-_144416481 | 0.06 |
ENST00000276833.9
|
SLC39A4
|
solute carrier family 39 member 4 |
| chr9_-_98708856 | 0.06 |
ENST00000259455.4
|
GABBR2
|
gamma-aminobutyric acid type B receptor subunit 2 |
| chr11_+_14643826 | 0.06 |
ENST00000455098.2
|
PDE3B
|
phosphodiesterase 3B |
| chr18_+_3448456 | 0.05 |
ENST00000549780.5
|
TGIF1
|
TGFB induced factor homeobox 1 |
| chrX_+_48802156 | 0.05 |
ENST00000643374.1
ENST00000644068.1 ENST00000441703.6 ENST00000643934.1 ENST00000489352.5 |
HDAC6
|
histone deacetylase 6 |
| chr22_+_24495242 | 0.05 |
ENST00000382760.2
ENST00000326010.10 |
UPB1
|
beta-ureidopropionase 1 |
| chr6_+_39793008 | 0.05 |
ENST00000398904.6
|
DAAM2
|
dishevelled associated activator of morphogenesis 2 |
| chr12_-_6689450 | 0.05 |
ENST00000355772.8
ENST00000417772.7 ENST00000319770.7 ENST00000396801.7 |
ZNF384
|
zinc finger protein 384 |
| chr3_-_9792404 | 0.05 |
ENST00000301964.7
|
TADA3
|
transcriptional adaptor 3 |
| chr18_-_3013114 | 0.04 |
ENST00000677752.1
|
LPIN2
|
lipin 2 |
| chr14_+_35826298 | 0.04 |
ENST00000216807.12
|
BRMS1L
|
BRMS1 like transcriptional repressor |
| chr8_-_94895195 | 0.04 |
ENST00000308108.9
ENST00000396133.7 |
CCNE2
|
cyclin E2 |
| chr17_-_15262537 | 0.04 |
ENST00000395936.7
ENST00000675819.1 ENST00000674707.1 ENST00000675854.1 ENST00000426385.4 ENST00000395938.7 ENST00000612492.5 ENST00000675808.1 |
PMP22
|
peripheral myelin protein 22 |
| chr22_-_43891995 | 0.04 |
ENST00000438734.1
ENST00000597664.6 ENST00000216177.8 ENST00000593866.5 ENST00000381198.6 |
PNPLA5
|
patatin like phospholipase domain containing 5 |
| chr22_+_37639660 | 0.04 |
ENST00000649765.2
ENST00000451997.6 |
SH3BP1
ENSG00000285304.1
|
SH3 domain binding protein 1 novel protein |
| chr9_-_133356448 | 0.04 |
ENST00000615505.4
ENST00000371974.8 |
SURF1
|
SURF1 cytochrome c oxidase assembly factor |
| chr16_-_4801301 | 0.04 |
ENST00000586504.5
ENST00000649556.1 |
ROGDI
ENSG00000285952.1
|
rogdi atypical leucine zipper novel transcript |
| chr12_-_6689244 | 0.03 |
ENST00000361959.7
ENST00000436774.6 ENST00000544482.1 |
ZNF384
|
zinc finger protein 384 |
| chrX_+_21940693 | 0.03 |
ENST00000404933.7
ENST00000379404.5 |
SMS
|
spermine synthase |
| chr22_-_24245059 | 0.03 |
ENST00000398292.3
ENST00000263112.11 ENST00000327365.10 ENST00000424217.1 |
GGT5
|
gamma-glutamyltransferase 5 |
| chr12_-_6689359 | 0.03 |
ENST00000683879.1
|
ZNF384
|
zinc finger protein 384 |
| chr19_+_11355491 | 0.03 |
ENST00000591608.1
|
PLPPR2
|
phospholipid phosphatase related 2 |
| chr8_-_143617498 | 0.03 |
ENST00000425753.7
|
GFUS
|
GDP-L-fucose synthase |
| chr20_+_45469745 | 0.03 |
ENST00000372676.8
ENST00000217425.9 ENST00000339946.7 |
WFDC2
|
WAP four-disulfide core domain 2 |
| chr19_+_11355386 | 0.03 |
ENST00000251473.9
ENST00000591329.5 ENST00000586380.5 |
PLPPR2
|
phospholipid phosphatase related 2 |
| chr6_-_159044980 | 0.03 |
ENST00000367066.8
|
TAGAP
|
T cell activation RhoGTPase activating protein |
| chr5_-_64768619 | 0.02 |
ENST00000513458.9
|
SREK1IP1
|
SREK1 interacting protein 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.8 | GO:0090271 | positive regulation of fibroblast growth factor production(GO:0090271) |
| 0.2 | 0.6 | GO:0051365 | leading edge cell differentiation(GO:0035026) cellular response to potassium ion starvation(GO:0051365) |
| 0.2 | 6.1 | GO:0014850 | response to muscle activity(GO:0014850) |
| 0.2 | 0.8 | GO:0061304 | retinal blood vessel morphogenesis(GO:0061304) |
| 0.1 | 0.6 | GO:0009956 | radial pattern formation(GO:0009956) |
| 0.1 | 0.6 | GO:0034164 | negative regulation of toll-like receptor 9 signaling pathway(GO:0034164) |
| 0.1 | 0.4 | GO:0002416 | IgG immunoglobulin transcytosis in epithelial cells mediated by FcRn immunoglobulin receptor(GO:0002416) |
| 0.1 | 0.4 | GO:0044335 | canonical Wnt signaling pathway involved in neural crest cell differentiation(GO:0044335) |
| 0.1 | 0.5 | GO:1904637 | response to ionomycin(GO:1904636) cellular response to ionomycin(GO:1904637) |
| 0.1 | 0.4 | GO:0006669 | sphinganine-1-phosphate biosynthetic process(GO:0006669) |
| 0.1 | 1.7 | GO:1901725 | regulation of histone deacetylase activity(GO:1901725) |
| 0.1 | 0.3 | GO:0061536 | glycine secretion(GO:0061536) glycine secretion, neurotransmission(GO:0061537) |
| 0.1 | 0.7 | GO:0060160 | negative regulation of dopamine receptor signaling pathway(GO:0060160) |
| 0.1 | 0.3 | GO:0031247 | actin rod assembly(GO:0031247) |
| 0.1 | 0.5 | GO:2001181 | positive regulation of interleukin-10 secretion(GO:2001181) |
| 0.1 | 0.4 | GO:0052553 | induction by symbiont of host defense response(GO:0044416) induction of host immune response by virus(GO:0046730) active induction of host immune response by virus(GO:0046732) modulation by symbiont of host defense response(GO:0052031) induction by organism of defense response of other organism involved in symbiotic interaction(GO:0052251) modulation by organism of defense response of other organism involved in symbiotic interaction(GO:0052255) positive regulation by symbiont of host defense response(GO:0052509) positive regulation by organism of defense response of other organism involved in symbiotic interaction(GO:0052510) modulation by organism of immune response of other organism involved in symbiotic interaction(GO:0052552) modulation by symbiont of host immune response(GO:0052553) modulation by virus of host immune response(GO:0075528) |
| 0.1 | 0.9 | GO:0060125 | habituation(GO:0046959) negative regulation of growth hormone secretion(GO:0060125) |
| 0.1 | 0.7 | GO:0003185 | primary heart field specification(GO:0003138) sinoatrial valve development(GO:0003172) sinoatrial valve morphogenesis(GO:0003185) |
| 0.1 | 0.5 | GO:0007386 | compartment pattern specification(GO:0007386) |
| 0.1 | 0.8 | GO:0098967 | exocytic insertion of neurotransmitter receptor to plasma membrane(GO:0098881) exocytic insertion of neurotransmitter receptor to postsynaptic membrane(GO:0098967) |
| 0.1 | 0.3 | GO:0000451 | rRNA 2'-O-methylation(GO:0000451) |
| 0.1 | 0.3 | GO:0048627 | myoblast development(GO:0048627) |
| 0.1 | 1.6 | GO:0042789 | mRNA transcription from RNA polymerase II promoter(GO:0042789) |
| 0.1 | 0.2 | GO:0003064 | regulation of heart rate by hormone(GO:0003064) negative regulation of potassium ion export(GO:1902303) |
| 0.1 | 0.4 | GO:0032487 | regulation of Rap protein signal transduction(GO:0032487) |
| 0.1 | 0.2 | GO:0031448 | regulation of fast-twitch skeletal muscle fiber contraction(GO:0031446) positive regulation of fast-twitch skeletal muscle fiber contraction(GO:0031448) |
| 0.1 | 0.4 | GO:0033591 | response to L-ascorbic acid(GO:0033591) |
| 0.1 | 0.3 | GO:0006740 | NADPH regeneration(GO:0006740) |
| 0.1 | 0.2 | GO:0002415 | immunoglobulin transcytosis in epithelial cells mediated by polymeric immunoglobulin receptor(GO:0002415) |
| 0.1 | 0.5 | GO:0070236 | negative regulation of activation-induced cell death of T cells(GO:0070236) |
| 0.0 | 0.5 | GO:0010499 | proteasomal ubiquitin-independent protein catabolic process(GO:0010499) |
| 0.0 | 0.6 | GO:0010593 | negative regulation of lamellipodium assembly(GO:0010593) |
| 0.0 | 0.4 | GO:0036072 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 0.0 | 0.5 | GO:0018095 | protein polyglutamylation(GO:0018095) |
| 0.0 | 0.2 | GO:0000379 | tRNA-type intron splice site recognition and cleavage(GO:0000379) |
| 0.0 | 0.2 | GO:0060474 | positive regulation of sperm motility involved in capacitation(GO:0060474) |
| 0.0 | 1.8 | GO:0030206 | chondroitin sulfate biosynthetic process(GO:0030206) |
| 0.0 | 0.2 | GO:0009644 | response to high light intensity(GO:0009644) |
| 0.0 | 0.1 | GO:0033092 | positive regulation of immature T cell proliferation in thymus(GO:0033092) |
| 0.0 | 0.2 | GO:0032792 | negative regulation of CREB transcription factor activity(GO:0032792) |
| 0.0 | 0.1 | GO:1904059 | regulation of locomotor rhythm(GO:1904059) |
| 0.0 | 0.2 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.0 | 0.5 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 0.0 | 0.6 | GO:0031115 | negative regulation of microtubule polymerization(GO:0031115) |
| 0.0 | 0.3 | GO:0045198 | establishment of epithelial cell apical/basal polarity(GO:0045198) |
| 0.0 | 0.1 | GO:0070846 | misfolded protein transport(GO:0070843) polyubiquitinated protein transport(GO:0070844) polyubiquitinated misfolded protein transport(GO:0070845) Hsp90 deacetylation(GO:0070846) |
| 0.0 | 0.3 | GO:0032790 | ribosome disassembly(GO:0032790) |
| 0.0 | 0.4 | GO:0031274 | positive regulation of pseudopodium assembly(GO:0031274) |
| 0.0 | 0.2 | GO:0048170 | positive regulation of long-term neuronal synaptic plasticity(GO:0048170) |
| 0.0 | 0.2 | GO:1903237 | negative regulation of leukocyte tethering or rolling(GO:1903237) |
| 0.0 | 0.3 | GO:0031915 | positive regulation of synaptic plasticity(GO:0031915) |
| 0.0 | 0.1 | GO:1904862 | inhibitory synapse assembly(GO:1904862) |
| 0.0 | 0.4 | GO:0001778 | plasma membrane repair(GO:0001778) |
| 0.0 | 0.4 | GO:0090161 | Golgi ribbon formation(GO:0090161) |
| 0.0 | 0.2 | GO:0007217 | tachykinin receptor signaling pathway(GO:0007217) |
| 0.0 | 0.1 | GO:0033629 | negative regulation of cell adhesion mediated by integrin(GO:0033629) |
| 0.0 | 4.6 | GO:0008360 | regulation of cell shape(GO:0008360) |
| 0.0 | 0.0 | GO:0019483 | beta-alanine biosynthetic process(GO:0019483) |
| 0.0 | 0.1 | GO:0070966 | nuclear-transcribed mRNA catabolic process, no-go decay(GO:0070966) |
| 0.0 | 0.1 | GO:0034224 | cellular response to zinc ion starvation(GO:0034224) |
| 0.0 | 2.9 | GO:0031424 | keratinization(GO:0031424) |
| 0.0 | 0.5 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.0 | 0.2 | GO:0038203 | TORC2 signaling(GO:0038203) |
| 0.0 | 0.3 | GO:0035563 | positive regulation of chromatin binding(GO:0035563) |
| 0.0 | 1.2 | GO:0032981 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.0 | 0.2 | GO:0008090 | retrograde axonal transport(GO:0008090) |
| 0.0 | 0.1 | GO:0002097 | tRNA wobble base modification(GO:0002097) |
| 0.0 | 0.2 | GO:0030903 | notochord development(GO:0030903) |
| 0.0 | 0.1 | GO:0003433 | chondrocyte development involved in endochondral bone morphogenesis(GO:0003433) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.1 | GO:1990851 | Wnt-Frizzled-LRP5/6 complex(GO:1990851) |
| 0.2 | 0.8 | GO:1990796 | photoreceptor cell terminal bouton(GO:1990796) |
| 0.1 | 0.6 | GO:0035976 | AP1 complex(GO:0035976) |
| 0.1 | 0.5 | GO:0071546 | pi-body(GO:0071546) |
| 0.1 | 0.2 | GO:0071149 | TEAD-2-YAP complex(GO:0071149) |
| 0.1 | 0.5 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.1 | 0.7 | GO:0045179 | apical cortex(GO:0045179) |
| 0.0 | 0.3 | GO:0070554 | synaptobrevin 2-SNAP-25-syntaxin-3-complexin complex(GO:0070554) |
| 0.0 | 0.2 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.0 | 0.3 | GO:0070436 | Grb2-EGFR complex(GO:0070436) |
| 0.0 | 0.1 | GO:0070931 | Golgi-associated vesicle lumen(GO:0070931) |
| 0.0 | 0.2 | GO:0000214 | tRNA-intron endonuclease complex(GO:0000214) |
| 0.0 | 0.2 | GO:0031673 | H zone(GO:0031673) |
| 0.0 | 0.7 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.0 | 2.2 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 1.9 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.0 | 0.6 | GO:0071682 | endocytic vesicle lumen(GO:0071682) |
| 0.0 | 0.7 | GO:0090545 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.0 | 1.4 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 0.5 | GO:0097381 | photoreceptor disc membrane(GO:0097381) |
| 0.0 | 4.2 | GO:0043197 | dendritic spine(GO:0043197) |
| 0.0 | 0.6 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.0 | 0.1 | GO:0005587 | collagen type IV trimer(GO:0005587) |
| 0.0 | 0.0 | GO:0097135 | cyclin E2-CDK2 complex(GO:0097135) |
| 0.0 | 0.1 | GO:0038039 | G-protein coupled receptor heterodimeric complex(GO:0038039) |
| 0.0 | 0.2 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.1 | GO:1904928 | coreceptor activity involved in canonical Wnt signaling pathway(GO:1904928) |
| 0.2 | 1.3 | GO:0050659 | N-acetylgalactosamine 4-sulfate 6-O-sulfotransferase activity(GO:0050659) |
| 0.2 | 0.5 | GO:0090555 | phosphatidylethanolamine-translocating ATPase activity(GO:0090555) |
| 0.1 | 0.7 | GO:0031750 | D3 dopamine receptor binding(GO:0031750) |
| 0.1 | 0.3 | GO:0070039 | rRNA (guanosine-2'-O-)-methyltransferase activity(GO:0070039) |
| 0.1 | 0.4 | GO:0019770 | IgG receptor activity(GO:0019770) |
| 0.1 | 0.3 | GO:0016652 | NAD(P)+ transhydrogenase activity(GO:0008746) oxidoreductase activity, acting on NAD(P)H, NAD(P) as acceptor(GO:0016652) |
| 0.1 | 0.7 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.1 | 0.6 | GO:0051022 | GDP-dissociation inhibitor binding(GO:0051021) Rho GDP-dissociation inhibitor binding(GO:0051022) |
| 0.1 | 0.3 | GO:0015375 | glycine:sodium symporter activity(GO:0015375) |
| 0.1 | 1.7 | GO:0019871 | sodium channel inhibitor activity(GO:0019871) |
| 0.1 | 0.6 | GO:0034485 | phosphatidylinositol-3,4,5-trisphosphate 5-phosphatase activity(GO:0034485) |
| 0.1 | 0.4 | GO:0034714 | type III transforming growth factor beta receptor binding(GO:0034714) |
| 0.1 | 0.4 | GO:0017050 | sphinganine kinase activity(GO:0008481) D-erythro-sphingosine kinase activity(GO:0017050) |
| 0.1 | 0.5 | GO:0043426 | MRF binding(GO:0043426) |
| 0.0 | 0.8 | GO:0050544 | arachidonic acid binding(GO:0050544) |
| 0.0 | 0.2 | GO:0019763 | immunoglobulin receptor activity(GO:0019763) |
| 0.0 | 0.1 | GO:0004119 | cGMP-inhibited cyclic-nucleotide phosphodiesterase activity(GO:0004119) |
| 0.0 | 1.1 | GO:0035198 | miRNA binding(GO:0035198) |
| 0.0 | 0.3 | GO:0071936 | coreceptor activity involved in Wnt signaling pathway(GO:0071936) |
| 0.0 | 0.1 | GO:0001626 | nociceptin receptor activity(GO:0001626) |
| 0.0 | 0.3 | GO:0046790 | virion binding(GO:0046790) |
| 0.0 | 2.2 | GO:0001102 | RNA polymerase II activating transcription factor binding(GO:0001102) |
| 0.0 | 0.1 | GO:0008798 | beta-aspartyl-peptidase activity(GO:0008798) |
| 0.0 | 0.3 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.0 | 0.7 | GO:0030169 | low-density lipoprotein particle binding(GO:0030169) |
| 0.0 | 0.2 | GO:0000213 | tRNA-intron endonuclease activity(GO:0000213) |
| 0.0 | 0.5 | GO:0043560 | insulin receptor substrate binding(GO:0043560) |
| 0.0 | 0.9 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.0 | 0.2 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.0 | 5.1 | GO:0004721 | phosphoprotein phosphatase activity(GO:0004721) |
| 0.0 | 0.2 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.0 | 0.4 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.1 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.0 | 0.6 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.0 | 0.4 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.0 | 0.1 | GO:0042903 | tubulin deacetylase activity(GO:0042903) |
| 0.0 | 0.2 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.0 | 0.7 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.0 | 0.1 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.0 | 0.1 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 0.0 | 0.2 | GO:0001134 | transcription factor activity, transcription factor recruiting(GO:0001134) |
| 0.0 | 0.3 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.0 | 0.5 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.7 | ST WNT CA2 CYCLIC GMP PATHWAY | Wnt/Ca2+/cyclic GMP signaling. |
| 0.0 | 1.1 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.0 | 0.9 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.0 | 2.4 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.0 | 0.4 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.0 | 1.5 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 0.6 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.0 | 0.5 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 0.8 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.0 | 0.5 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.0 | 0.9 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 0.4 | PID S1P META PATHWAY | Sphingosine 1-phosphate (S1P) pathway |
| 0.0 | 0.7 | PID MAPK TRK PATHWAY | Trk receptor signaling mediated by the MAPK pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.8 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |
| 0.0 | 1.7 | REACTOME UNBLOCKING OF NMDA RECEPTOR GLUTAMATE BINDING AND ACTIVATION | Genes involved in Unblocking of NMDA receptor, glutamate binding and activation |
| 0.0 | 1.1 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.0 | 0.9 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.0 | 0.9 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.0 | 0.6 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.0 | 0.3 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.0 | 0.6 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 0.4 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.0 | 0.7 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.0 | 0.7 | REACTOME ERK MAPK TARGETS | Genes involved in ERK/MAPK targets |
| 0.0 | 2.3 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.0 | 0.6 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.0 | 0.3 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 0.1 | REACTOME P38MAPK EVENTS | Genes involved in p38MAPK events |
| 0.0 | 0.9 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.0 | 0.4 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.3 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |