Project
Inflammatory response time course, HUVEC (Wada, 2009)
Navigation
Downloads
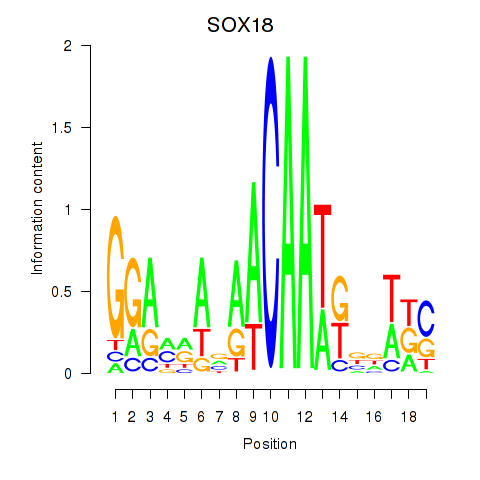
Results for SOX18
Z-value: 0.49
Transcription factors associated with SOX18
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SOX18
|
ENSG00000203883.7 | SOX18 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SOX18 | hg38_v1_chr20_-_64049631_64049646 | -0.29 | 1.7e-01 | Click! |
Activity profile of SOX18 motif
Sorted Z-values of SOX18 motif
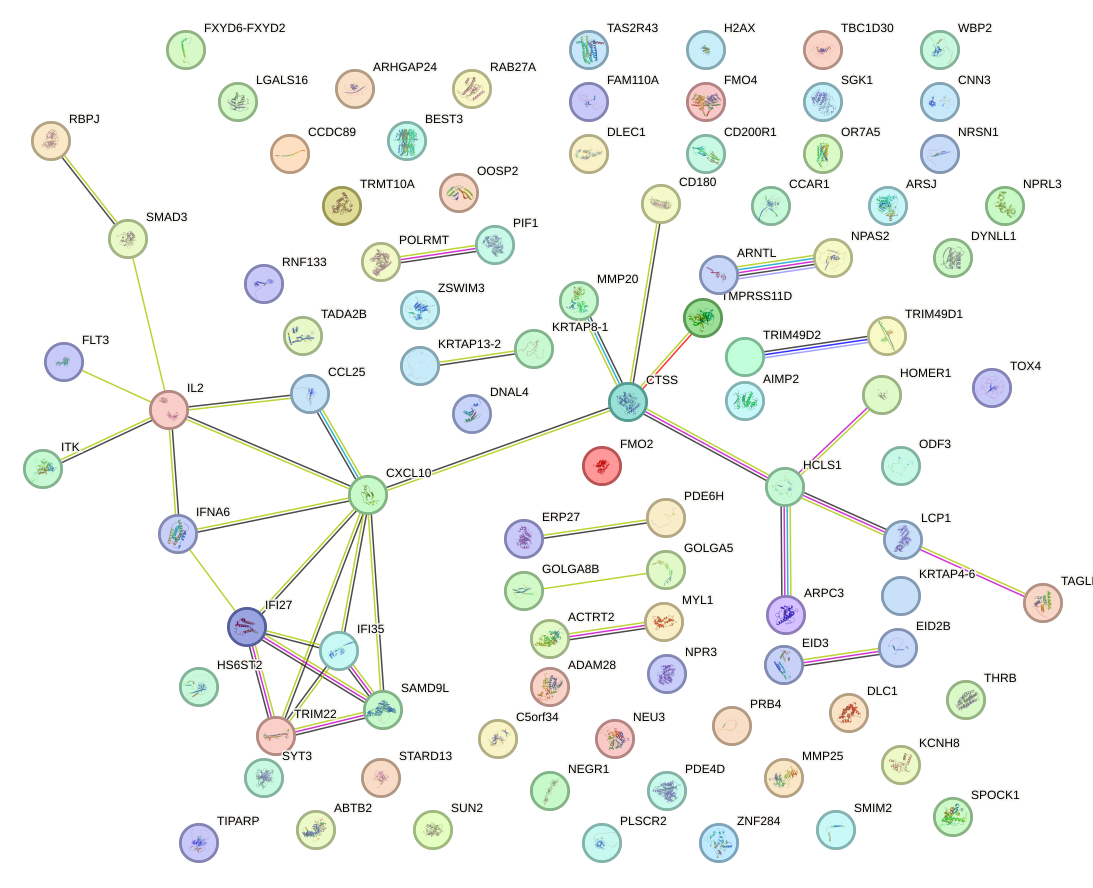
Network of associatons between targets according to the STRING database.
First level regulatory network of SOX18
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_-_76023489 | 2.09 |
ENST00000306602.3
|
CXCL10
|
C-X-C motif chemokine ligand 10 |
| chr17_+_43006740 | 1.39 |
ENST00000438323.2
ENST00000415816.7 |
IFI35
|
interferon induced protein 35 |
| chr1_-_150765735 | 1.30 |
ENST00000679898.1
ENST00000448301.7 ENST00000680664.1 ENST00000679512.1 ENST00000368985.8 ENST00000679582.1 |
CTSS
|
cathepsin S |
| chr6_-_128520358 | 1.12 |
ENST00000368215.7
ENST00000532331.5 ENST00000368213.9 ENST00000368207.7 ENST00000525459.1 ENST00000368226.9 ENST00000368210.7 |
PTPRK
|
protein tyrosine phosphatase receptor type K |
| chr11_+_5689780 | 0.74 |
ENST00000379965.8
ENST00000454828.5 |
TRIM22
|
tripartite motif containing 22 |
| chr11_-_117876719 | 0.72 |
ENST00000529335.6
ENST00000260282.8 |
FXYD6
|
FXYD domain containing ion transport regulator 6 |
| chr7_-_93148345 | 0.67 |
ENST00000437805.5
ENST00000446959.5 ENST00000439952.5 ENST00000414791.5 ENST00000446033.1 ENST00000411955.5 ENST00000318238.9 |
SAMD9L
|
sterile alpha motif domain containing 9 like |
| chr13_-_33205997 | 0.64 |
ENST00000399365.7
|
STARD13
|
StAR related lipid transfer domain containing 13 |
| chr6_-_134318097 | 0.61 |
ENST00000367858.10
ENST00000533224.1 |
SGK1
|
serum/glucocorticoid regulated kinase 1 |
| chr11_-_117876892 | 0.57 |
ENST00000539526.5
|
FXYD6
|
FXYD domain containing ion transport regulator 6 |
| chr4_-_113979635 | 0.53 |
ENST00000315366.8
|
ARSJ
|
arylsulfatase family member J |
| chr21_-_30372265 | 0.53 |
ENST00000399889.4
|
KRTAP13-2
|
keratin associated protein 13-2 |
| chr19_+_8052752 | 0.51 |
ENST00000315626.6
ENST00000253451.9 |
CCL25
|
C-C motif chemokine ligand 25 |
| chr10_+_68721012 | 0.50 |
ENST00000536391.5
ENST00000630771.2 |
CCAR1
|
cell division cycle and apoptosis regulator 1 |
| chr12_+_14973020 | 0.48 |
ENST00000266395.3
|
PDE6H
|
phosphodiesterase 6H |
| chr21_-_30813270 | 0.47 |
ENST00000329621.6
|
KRTAP8-1
|
keratin associated protein 8-1 |
| chr5_-_67196791 | 0.44 |
ENST00000256447.5
|
CD180
|
CD180 molecule |
| chr19_+_8053000 | 0.42 |
ENST00000390669.7
|
CCL25
|
C-C motif chemokine ligand 25 |
| chr1_-_247458105 | 0.41 |
ENST00000641149.1
ENST00000641527.1 |
OR2B11
|
olfactory receptor family 2 subfamily B member 11 |
| chr11_-_117876612 | 0.41 |
ENST00000584230.1
ENST00000526014.6 ENST00000584394.5 ENST00000614497.5 ENST00000532984.1 |
FXYD6
FXYD6-FXYD2
|
FXYD domain containing ion transport regulator 6 FXYD6-FXYD2 readthrough |
| chr11_-_34357994 | 0.40 |
ENST00000435224.3
|
ABTB2
|
ankyrin repeat and BTB domain containing 2 |
| chr14_+_94110728 | 0.39 |
ENST00000616764.4
ENST00000618863.1 ENST00000611954.4 ENST00000618200.4 ENST00000621160.4 ENST00000555819.5 ENST00000620396.4 ENST00000612813.4 ENST00000620066.1 |
IFI27
|
interferon alpha inducible protein 27 |
| chr19_-_14835162 | 0.38 |
ENST00000322301.5
|
OR7A5
|
olfactory receptor family 7 subfamily A member 5 |
| chr7_-_122699108 | 0.36 |
ENST00000340112.3
|
RNF133
|
ring finger protein 133 |
| chr13_-_28100556 | 0.36 |
ENST00000241453.12
|
FLT3
|
fms related receptor tyrosine kinase 3 |
| chr3_-_112975018 | 0.35 |
ENST00000471858.5
ENST00000308611.8 ENST00000295863.4 |
CD200R1
|
CD200 receptor 1 |
| chr2_-_37671633 | 0.35 |
ENST00000295324.4
|
CDC42EP3
|
CDC42 effector protein 3 |
| chr2_-_210303608 | 0.34 |
ENST00000341685.8
|
MYL1
|
myosin light chain 1 |
| chr1_+_171185293 | 0.33 |
ENST00000209929.10
|
FMO2
|
flavin containing dimethylaniline monoxygenase 2 |
| chr11_+_13277639 | 0.33 |
ENST00000527998.5
ENST00000529388.6 ENST00000401424.6 ENST00000403510.8 ENST00000403290.6 ENST00000533520.5 ENST00000389707.8 ENST00000673817.1 |
ARNTL
|
aryl hydrocarbon receptor nuclear translocator like |
| chr11_+_196761 | 0.31 |
ENST00000325113.9
|
ODF3
|
outer dense fiber of sperm tails 3 |
| chr3_+_19148500 | 0.30 |
ENST00000328405.7
|
KCNH8
|
potassium voltage-gated channel subfamily H member 8 |
| chr19_+_39655907 | 0.29 |
ENST00000392051.4
|
LGALS16
|
galectin 16 |
| chr3_+_111998739 | 0.28 |
ENST00000393917.6
ENST00000273368.8 |
TAGLN3
|
transgelin 3 |
| chr20_+_45857607 | 0.27 |
ENST00000255152.3
|
ZSWIM3
|
zinc finger SWIM-type containing 3 |
| chr13_-_51845169 | 0.27 |
ENST00000627246.3
ENST00000629372.3 |
TMEM272
|
transmembrane protein 272 |
| chr13_-_46142834 | 0.27 |
ENST00000674665.1
|
LCP1
|
lymphocyte cytosolic protein 1 |
| chr11_-_85686123 | 0.26 |
ENST00000316398.5
|
CCDC89
|
coiled-coil domain containing 89 |
| chr8_-_13276491 | 0.26 |
ENST00000512044.6
|
DLC1
|
DLC1 Rho GTPase activating protein |
| chr22_-_38794111 | 0.26 |
ENST00000406622.5
ENST00000216068.9 ENST00000406199.3 |
SUN2
DNAL4
|
Sad1 and UNC84 domain containing 2 dynein axonemal light chain 4 |
| chr2_+_100974849 | 0.26 |
ENST00000450763.1
|
NPAS2
|
neuronal PAS domain protein 2 |
| chr19_-_633500 | 0.25 |
ENST00000588649.7
|
POLRMT
|
RNA polymerase mitochondrial |
| chr11_+_89926762 | 0.25 |
ENST00000526396.3
|
TRIM49D2
|
tripartite motif containing 49D2 |
| chr12_-_11092313 | 0.25 |
ENST00000531678.1
|
TAS2R43
|
taste 2 receptor member 43 |
| chr3_+_111998915 | 0.24 |
ENST00000478951.6
|
TAGLN3
|
transgelin 3 |
| chr11_+_74988896 | 0.24 |
ENST00000526068.1
ENST00000294064.9 ENST00000532963.1 ENST00000531619.1 ENST00000534628.1 |
NEU3
|
neuraminidase 3 |
| chr19_-_56836047 | 0.23 |
ENST00000601070.5
|
ZIM2
|
zinc finger imprinted 2 |
| chr15_-_64825608 | 0.23 |
ENST00000559239.2
ENST00000268043.8 ENST00000333425.10 |
PIF1
|
PIF1 5'-to-3' DNA helicase |
| chr3_-_146460440 | 0.22 |
ENST00000610787.5
|
PLSCR2
|
phospholipid scramblase 2 |
| chr12_-_14938508 | 0.21 |
ENST00000266397.7
|
ERP27
|
endoplasmic reticulum protein 27 |
| chr1_-_94927079 | 0.21 |
ENST00000370206.9
ENST00000394202.8 |
CNN3
|
calponin 3 |
| chr6_+_24126186 | 0.20 |
ENST00000378478.5
ENST00000378491.9 ENST00000378477.2 |
NRSN1
|
neurensin 1 |
| chr5_-_137499293 | 0.20 |
ENST00000510689.5
ENST00000394945.6 |
SPOCK1
|
SPARC (osteonectin), cwcv and kazal like domains proteoglycan 1 |
| chr4_-_122456725 | 0.20 |
ENST00000226730.5
|
IL2
|
interleukin 2 |
| chr3_+_156674579 | 0.20 |
ENST00000295924.12
|
TIPARP
|
TCDD inducible poly(ADP-ribose) polymerase |
| chr12_+_104303726 | 0.19 |
ENST00000527879.2
|
EID3
|
EP300 interacting inhibitor of differentiation 3 |
| chr4_+_26321192 | 0.19 |
ENST00000681484.1
|
RBPJ
|
recombination signal binding protein for immunoglobulin kappa J region |
| chr15_+_67125707 | 0.19 |
ENST00000540846.6
|
SMAD3
|
SMAD family member 3 |
| chr4_+_26321365 | 0.19 |
ENST00000505958.6
|
RBPJ
|
recombination signal binding protein for immunoglobulin kappa J region |
| chr4_-_99563668 | 0.19 |
ENST00000273962.7
|
TRMT10A
|
tRNA methyltransferase 10A |
| chr13_-_46182136 | 0.19 |
ENST00000323076.7
|
LCP1
|
lymphocyte cytosolic protein 1 |
| chr1_+_3021466 | 0.19 |
ENST00000378404.4
|
ACTRT2
|
actin related protein T2 |
| chr8_+_24294044 | 0.19 |
ENST00000265769.9
|
ADAM28
|
ADAM metallopeptidase domain 28 |
| chr12_-_110445540 | 0.18 |
ENST00000547365.1
|
ARPC3
|
actin related protein 2/3 complex subunit 3 |
| chr7_+_6009222 | 0.18 |
ENST00000400479.6
ENST00000223029.8 ENST00000395236.2 |
AIMP2
|
aminoacyl tRNA synthetase complex interacting multifunctional protein 2 |
| chr5_-_59356962 | 0.18 |
ENST00000405755.6
|
PDE4D
|
phosphodiesterase 4D |
| chr19_+_44072142 | 0.18 |
ENST00000421176.4
|
ZNF284
|
zinc finger protein 284 |
| chr3_+_57756230 | 0.18 |
ENST00000295951.7
ENST00000659705.1 ENST00000671191.1 |
SLMAP
|
sarcolemma associated protein |
| chr2_-_37672178 | 0.17 |
ENST00000457889.1
|
CDC42EP3
|
CDC42 effector protein 3 |
| chr11_-_102625332 | 0.17 |
ENST00000260228.3
|
MMP20
|
matrix metallopeptidase 20 |
| chr4_+_7043315 | 0.16 |
ENST00000310074.8
ENST00000512388.1 |
TADA2B
|
transcriptional adaptor 2B |
| chr3_+_127193131 | 0.16 |
ENST00000624688.2
|
C3orf56
|
chromosome 3 open reading frame 56 |
| chr15_+_21651844 | 0.16 |
ENST00000623441.1
|
OR4N4C
|
olfactory receptor family 4 subfamily N member 4C |
| chrX_-_111412162 | 0.16 |
ENST00000637570.1
ENST00000356220.8 ENST00000636035.2 ENST00000637453.1 ENST00000635795.1 |
DCX
|
doublecortin |
| chr14_+_92794297 | 0.16 |
ENST00000163416.7
|
GOLGA5
|
golgin A5 |
| chr11_+_60040402 | 0.15 |
ENST00000278855.7
ENST00000532905.1 |
OOSP2
|
oocyte secreted protein 2 |
| chr5_-_79512794 | 0.15 |
ENST00000282260.10
ENST00000508576.5 ENST00000535690.1 |
HOMER1
|
homer scaffold protein 1 |
| chr20_+_841238 | 0.15 |
ENST00000541082.2
|
FAM110A
|
family with sequence similarity 110 member A |
| chr7_+_142111739 | 0.15 |
ENST00000550469.6
ENST00000477922.3 |
MGAM2
|
maltase-glucoamylase 2 (putative) |
| chr19_-_50639827 | 0.15 |
ENST00000593901.5
ENST00000600079.6 |
SYT3
|
synaptotagmin 3 |
| chr7_+_142511614 | 0.15 |
ENST00000390377.1
|
TRBV7-7
|
T cell receptor beta variable 7-7 |
| chr4_-_67883987 | 0.15 |
ENST00000283916.11
|
TMPRSS11D
|
transmembrane serine protease 11D |
| chr15_-_80252205 | 0.15 |
ENST00000560778.3
|
CTXND1
|
cortexin domain containing 1 |
| chr3_-_24495532 | 0.15 |
ENST00000643772.1
ENST00000642307.1 ENST00000645139.1 |
THRB
|
thyroid hormone receptor beta |
| chr9_-_21351378 | 0.15 |
ENST00000380210.1
|
IFNA6
|
interferon alpha 6 |
| chr12_-_69699353 | 0.15 |
ENST00000331471.8
|
BEST3
|
bestrophin 3 |
| chr3_-_121660892 | 0.15 |
ENST00000428394.6
ENST00000314583.8 |
HCLS1
|
hematopoietic cell-specific Lyn substrate 1 |
| chr12_-_69699331 | 0.15 |
ENST00000548658.1
ENST00000476098.5 |
BEST3
|
bestrophin 3 |
| chr5_-_43515128 | 0.14 |
ENST00000306862.7
|
C5orf34
|
chromosome 5 open reading frame 34 |
| chr11_-_119095456 | 0.14 |
ENST00000530167.1
|
H2AX
|
H2A.X variant histone |
| chr17_-_75855204 | 0.14 |
ENST00000589642.5
ENST00000593002.1 ENST00000590221.5 ENST00000587374.5 ENST00000585462.5 ENST00000254806.8 ENST00000433525.6 ENST00000626827.2 |
WBP2
|
WW domain binding protein 2 |
| chr11_+_124184244 | 0.14 |
ENST00000641546.1
|
OR10D3
|
olfactory receptor family 10 subfamily D member 3 |
| chr4_+_85604146 | 0.14 |
ENST00000512201.5
|
ARHGAP24
|
Rho GTPase activating protein 24 |
| chr13_-_44161257 | 0.14 |
ENST00000400419.2
|
SMIM2
|
small integral membrane protein 2 |
| chr17_-_41140487 | 0.14 |
ENST00000345847.4
|
KRTAP4-6
|
keratin associated protein 4-6 |
| chr5_+_32711723 | 0.14 |
ENST00000415167.2
|
NPR3
|
natriuretic peptide receptor 3 |
| chr12_-_11310420 | 0.14 |
ENST00000621732.4
ENST00000445719.2 ENST00000279575.7 |
PRB4
|
proline rich protein BstNI subfamily 4 |
| chr5_+_157180816 | 0.14 |
ENST00000422843.8
|
ITK
|
IL2 inducible T cell kinase |
| chr1_-_72100930 | 0.14 |
ENST00000306821.3
|
NEGR1
|
neuronal growth regulator 1 |
| chrX_-_132961390 | 0.13 |
ENST00000370836.6
ENST00000521489.5 |
HS6ST2
|
heparan sulfate 6-O-sulfotransferase 2 |
| chr15_-_55249029 | 0.13 |
ENST00000566877.5
|
RAB27A
|
RAB27A, member RAS oncogene family |
| chr12_+_64824602 | 0.13 |
ENST00000539867.6
ENST00000544457.1 ENST00000539120.1 |
TBC1D30
|
TBC1 domain family member 30 |
| chr12_-_95996302 | 0.13 |
ENST00000261208.8
ENST00000538703.5 ENST00000541929.5 |
HAL
|
histidine ammonia-lyase |
| chr18_-_5396265 | 0.13 |
ENST00000579951.2
|
EPB41L3
|
erythrocyte membrane protein band 4.1 like 3 |
| chr19_+_36111151 | 0.13 |
ENST00000633214.1
ENST00000585332.3 |
OVOL3
|
ovo like zinc finger 3 |
| chr9_-_134917872 | 0.12 |
ENST00000616356.4
ENST00000371806.4 |
FCN1
|
ficolin 1 |
| chr5_+_159263282 | 0.12 |
ENST00000296786.8
|
UBLCP1
|
ubiquitin like domain containing CTD phosphatase 1 |
| chr7_-_149028452 | 0.12 |
ENST00000413966.1
ENST00000652332.1 |
PDIA4
|
protein disulfide isomerase family A member 4 |
| chr2_+_201260510 | 0.12 |
ENST00000673742.1
|
CASP8
|
caspase 8 |
| chr2_+_201260496 | 0.12 |
ENST00000323492.11
|
CASP8
|
caspase 8 |
| chr15_+_22094522 | 0.12 |
ENST00000328795.5
|
OR4N4
|
olfactory receptor family 4 subfamily N member 4 |
| chr7_-_149028651 | 0.12 |
ENST00000286091.9
|
PDIA4
|
protein disulfide isomerase family A member 4 |
| chr3_-_36993103 | 0.12 |
ENST00000322716.8
|
EPM2AIP1
|
EPM2A interacting protein 1 |
| chrX_+_120362079 | 0.12 |
ENST00000539306.5
ENST00000218008.8 ENST00000361319.3 |
ATP1B4
|
ATPase Na+/K+ transporting family member beta 4 |
| chr14_-_106675544 | 0.12 |
ENST00000390632.2
|
IGHV3-66
|
immunoglobulin heavy variable 3-66 |
| chrX_+_30243715 | 0.12 |
ENST00000378981.8
ENST00000397550.6 |
MAGEB1
|
MAGE family member B1 |
| chr3_-_71064915 | 0.12 |
ENST00000614176.5
ENST00000485326.7 |
FOXP1
|
forkhead box P1 |
| chr11_-_118176576 | 0.12 |
ENST00000278947.6
|
SCN2B
|
sodium voltage-gated channel beta subunit 2 |
| chr1_-_11858935 | 0.12 |
ENST00000376468.4
|
NPPB
|
natriuretic peptide B |
| chr12_-_11395556 | 0.11 |
ENST00000565533.1
ENST00000389362.6 ENST00000546254.3 |
PRB2
PRB1
|
proline rich protein BstNI subfamily 2 proline rich protein BstNI subfamily 1 |
| chrX_+_120604084 | 0.11 |
ENST00000371317.10
|
MCTS1
|
MCTS1 re-initiation and release factor |
| chr1_+_13060769 | 0.11 |
ENST00000617807.3
|
HNRNPCL3
|
heterogeneous nuclear ribonucleoprotein C like 3 |
| chr7_+_142598016 | 0.11 |
ENST00000620773.1
|
TRBV16
|
T cell receptor beta variable 16 |
| chr3_-_71064964 | 0.11 |
ENST00000650387.1
|
FOXP1
|
forkhead box P1 |
| chr12_-_49605634 | 0.11 |
ENST00000257894.2
|
FAM186B
|
family with sequence similarity 186 member B |
| chr14_+_21768482 | 0.11 |
ENST00000390428.3
|
TRAV6
|
T cell receptor alpha variable 6 |
| chr19_+_48271327 | 0.11 |
ENST00000594024.5
ENST00000595408.5 ENST00000315849.5 |
ZNF114
|
zinc finger protein 114 |
| chr5_-_142644201 | 0.11 |
ENST00000394496.2
ENST00000610990.4 |
FGF1
|
fibroblast growth factor 1 |
| chr8_+_24294107 | 0.11 |
ENST00000437154.6
|
ADAM28
|
ADAM metallopeptidase domain 28 |
| chr12_+_32679200 | 0.11 |
ENST00000452533.6
ENST00000414834.6 |
DNM1L
|
dynamin 1 like |
| chr8_-_85341705 | 0.11 |
ENST00000517618.5
|
CA1
|
carbonic anhydrase 1 |
| chr4_+_109815503 | 0.11 |
ENST00000394631.7
|
GAR1
|
GAR1 ribonucleoprotein |
| chr11_+_65181920 | 0.10 |
ENST00000279247.11
ENST00000532285.1 ENST00000534373.5 |
CAPN1
|
calpain 1 |
| chr4_+_3441960 | 0.10 |
ENST00000382774.8
ENST00000511533.1 |
HGFAC
|
HGF activator |
| chr1_-_153540694 | 0.10 |
ENST00000368717.2
|
S100A5
|
S100 calcium binding protein A5 |
| chr14_+_35046238 | 0.10 |
ENST00000280987.9
|
FAM177A1
|
family with sequence similarity 177 member A1 |
| chr15_-_42491105 | 0.10 |
ENST00000565380.5
ENST00000564754.7 |
ZNF106
|
zinc finger protein 106 |
| chr8_-_21812320 | 0.10 |
ENST00000517328.5
|
GFRA2
|
GDNF family receptor alpha 2 |
| chr17_-_17281232 | 0.10 |
ENST00000417352.5
ENST00000268717.10 |
COPS3
|
COP9 signalosome subunit 3 |
| chr8_+_117135259 | 0.10 |
ENST00000519688.5
|
SLC30A8
|
solute carrier family 30 member 8 |
| chr12_-_2876986 | 0.10 |
ENST00000342628.6
ENST00000361953.7 |
FOXM1
|
forkhead box M1 |
| chr3_-_112974912 | 0.10 |
ENST00000440122.6
ENST00000490004.1 |
CD200R1
|
CD200 receptor 1 |
| chr1_-_165445220 | 0.10 |
ENST00000619224.1
|
RXRG
|
retinoid X receptor gamma |
| chr1_+_179954740 | 0.10 |
ENST00000491495.2
ENST00000367607.8 |
CEP350
|
centrosomal protein 350 |
| chr5_+_44808930 | 0.09 |
ENST00000507110.6
|
MRPS30
|
mitochondrial ribosomal protein S30 |
| chr1_+_13303539 | 0.09 |
ENST00000437300.2
|
PRAMEF33
|
PRAME family member 33 |
| chr1_-_248645278 | 0.09 |
ENST00000641268.1
|
OR2T35
|
olfactory receptor family 2 subfamily T member 35 |
| chr16_-_28356654 | 0.09 |
ENST00000533640.5
|
NPIPB6
|
nuclear pore complex interacting protein family member B6 |
| chr2_+_53767772 | 0.09 |
ENST00000295304.5
|
CHAC2
|
ChaC glutathione specific gamma-glutamylcyclotransferase 2 |
| chr12_+_69359673 | 0.09 |
ENST00000548020.5
ENST00000549685.5 ENST00000247843.7 ENST00000552955.1 |
YEATS4
|
YEATS domain containing 4 |
| chr8_-_86743626 | 0.09 |
ENST00000320005.6
|
CNGB3
|
cyclic nucleotide gated channel subunit beta 3 |
| chr2_+_3658193 | 0.09 |
ENST00000252505.4
|
ALLC
|
allantoicase |
| chr6_-_169250825 | 0.09 |
ENST00000676869.1
ENST00000676760.1 |
THBS2
|
thrombospondin 2 |
| chr1_-_156248084 | 0.09 |
ENST00000652405.1
ENST00000335852.6 ENST00000540423.5 ENST00000612424.4 ENST00000613336.4 ENST00000623241.3 |
PAQR6
|
progestin and adipoQ receptor family member 6 |
| chr17_+_28371656 | 0.09 |
ENST00000585482.6
|
SARM1
|
sterile alpha and TIR motif containing 1 |
| chr12_-_262828 | 0.09 |
ENST00000343164.9
ENST00000436453.1 ENST00000445055.6 ENST00000546319.5 |
SLC6A13
|
solute carrier family 6 member 13 |
| chr7_-_135977308 | 0.09 |
ENST00000435723.1
ENST00000393085.4 |
MTPN
|
myotrophin |
| chr1_+_153031195 | 0.09 |
ENST00000307098.5
|
SPRR1B
|
small proline rich protein 1B |
| chr22_+_42553862 | 0.09 |
ENST00000340239.8
ENST00000327678.10 |
SERHL2
|
serine hydrolase like 2 |
| chr16_+_33827140 | 0.08 |
ENST00000562905.2
|
IGHV3OR16-13
|
immunoglobulin heavy variable 3/OR16-13 (non-functional) |
| chr15_-_40874216 | 0.08 |
ENST00000220507.5
|
RHOV
|
ras homolog family member V |
| chr11_-_105023136 | 0.08 |
ENST00000526056.5
ENST00000531367.5 ENST00000456094.1 ENST00000444749.6 ENST00000393141.6 ENST00000418434.5 ENST00000260315.8 |
CASP5
|
caspase 5 |
| chr19_-_19733091 | 0.08 |
ENST00000344099.4
|
ZNF14
|
zinc finger protein 14 |
| chr10_+_84424919 | 0.08 |
ENST00000543283.2
ENST00000494586.5 |
CCSER2
|
coiled-coil serine rich protein 2 |
| chr12_+_69239592 | 0.08 |
ENST00000456847.7
ENST00000266679.8 |
CPSF6
|
cleavage and polyadenylation specific factor 6 |
| chr10_-_104232301 | 0.08 |
ENST00000369720.6
ENST00000369719.2 ENST00000278064.7 ENST00000357060.8 |
CFAP43
|
cilia and flagella associated protein 43 |
| chr18_+_35041387 | 0.08 |
ENST00000538170.6
ENST00000300249.10 ENST00000588910.5 |
MAPRE2
|
microtubule associated protein RP/EB family member 2 |
| chr13_-_83882390 | 0.08 |
ENST00000377084.3
|
SLITRK1
|
SLIT and NTRK like family member 1 |
| chrX_-_47145035 | 0.08 |
ENST00000276062.8
|
NDUFB11
|
NADH:ubiquinone oxidoreductase subunit B11 |
| chr3_-_151461011 | 0.08 |
ENST00000282466.4
|
IGSF10
|
immunoglobulin superfamily member 10 |
| chr20_-_37095985 | 0.08 |
ENST00000344359.7
ENST00000373664.8 |
RBL1
|
RB transcriptional corepressor like 1 |
| chr12_-_2877113 | 0.08 |
ENST00000627656.2
ENST00000359843.8 |
FOXM1
|
forkhead box M1 |
| chr2_+_105337515 | 0.08 |
ENST00000410049.1
ENST00000258457.7 |
C2orf49
|
chromosome 2 open reading frame 49 |
| chr19_+_7049321 | 0.08 |
ENST00000381393.3
|
MBD3L2
|
methyl-CpG binding domain protein 3 like 2 |
| chr11_+_65182413 | 0.08 |
ENST00000531068.5
ENST00000527699.5 ENST00000533909.5 ENST00000527323.5 |
CAPN1
|
calpain 1 |
| chrX_-_49184789 | 0.08 |
ENST00000453382.5
ENST00000432913.5 |
PRICKLE3
|
prickle planar cell polarity protein 3 |
| chr1_-_12898270 | 0.08 |
ENST00000235347.4
|
PRAMEF10
|
PRAME family member 10 |
| chr5_-_138213308 | 0.08 |
ENST00000394886.7
ENST00000505120.1 ENST00000394884.7 |
CDC23
|
cell division cycle 23 |
| chr19_-_35745391 | 0.07 |
ENST00000378975.8
ENST00000587886.2 ENST00000412391.6 ENST00000292879.9 |
U2AF1L4
|
U2 small nuclear RNA auxiliary factor 1 like 4 |
| chr1_+_160081529 | 0.07 |
ENST00000368088.4
|
KCNJ9
|
potassium inwardly rectifying channel subfamily J member 9 |
| chr12_-_69699285 | 0.07 |
ENST00000553096.5
ENST00000330891.10 |
BEST3
|
bestrophin 3 |
| chr1_-_157777124 | 0.07 |
ENST00000361516.8
ENST00000368181.4 |
FCRL2
|
Fc receptor like 2 |
| chr12_+_69239560 | 0.07 |
ENST00000435070.7
|
CPSF6
|
cleavage and polyadenylation specific factor 6 |
| chrX_+_47193796 | 0.07 |
ENST00000442035.5
ENST00000335972.11 ENST00000457753.5 |
UBA1
|
ubiquitin like modifier activating enzyme 1 |
| chr5_-_98928992 | 0.07 |
ENST00000614616.5
|
CHD1
|
chromodomain helicase DNA binding protein 1 |
| chr6_-_31158073 | 0.07 |
ENST00000507751.5
ENST00000448162.6 ENST00000502557.5 ENST00000503420.5 ENST00000507892.1 ENST00000507226.1 ENST00000513222.1 ENST00000503934.5 ENST00000396263.6 ENST00000508683.5 ENST00000428174.1 ENST00000448141.6 ENST00000507829.5 ENST00000455279.6 ENST00000376266.9 |
CCHCR1
|
coiled-coil alpha-helical rod protein 1 |
| chr1_-_156248038 | 0.07 |
ENST00000470198.5
ENST00000292291.10 ENST00000356983.7 |
PAQR6
|
progestin and adipoQ receptor family member 6 |
| chr9_+_35791570 | 0.07 |
ENST00000342694.7
|
NPR2
|
natriuretic peptide receptor 2 |
| chr2_+_44275457 | 0.07 |
ENST00000611973.4
ENST00000409387.5 |
SLC3A1
|
solute carrier family 3 member 1 |
| chr1_+_201780490 | 0.07 |
ENST00000430015.5
|
NAV1
|
neuron navigator 1 |
| chr15_-_79090760 | 0.07 |
ENST00000419573.7
ENST00000558480.7 |
RASGRF1
|
Ras protein specific guanine nucleotide releasing factor 1 |
| chr7_-_93574721 | 0.07 |
ENST00000426151.7
ENST00000649521.1 |
CALCR
|
calcitonin receptor |
| chr2_+_79025696 | 0.07 |
ENST00000272324.10
|
REG3G
|
regenerating family member 3 gamma |
| chr9_-_125484490 | 0.07 |
ENST00000444226.1
|
MAPKAP1
|
MAPK associated protein 1 |
| chr11_+_124183219 | 0.07 |
ENST00000641351.2
|
OR10D3
|
olfactory receptor family 10 subfamily D member 3 |
| chr1_-_156248013 | 0.06 |
ENST00000368270.2
|
PAQR6
|
progestin and adipoQ receptor family member 6 |
| chr14_-_106811131 | 0.06 |
ENST00000424969.2
|
IGHV3-74
|
immunoglobulin heavy variable 3-74 |
| chr8_+_54616078 | 0.06 |
ENST00000220676.2
|
RP1
|
RP1 axonemal microtubule associated |
| chr8_-_85341659 | 0.06 |
ENST00000522389.5
|
CA1
|
carbonic anhydrase 1 |
| chr9_-_96302142 | 0.06 |
ENST00000648799.1
|
HSD17B3
|
hydroxysteroid 17-beta dehydrogenase 3 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.3 | GO:0034769 | basement membrane disassembly(GO:0034769) |
| 0.2 | 2.1 | GO:0043950 | positive regulation of cAMP-mediated signaling(GO:0043950) |
| 0.1 | 0.9 | GO:1903237 | negative regulation of leukocyte tethering or rolling(GO:1903237) |
| 0.1 | 0.6 | GO:0097498 | endothelial tube lumen extension(GO:0097498) |
| 0.1 | 0.3 | GO:0006391 | transcription initiation from mitochondrial promoter(GO:0006391) |
| 0.1 | 0.3 | GO:0090403 | oxidative stress-induced premature senescence(GO:0090403) |
| 0.1 | 0.6 | GO:0070294 | renal sodium ion absorption(GO:0070294) |
| 0.1 | 0.4 | GO:0002322 | B cell proliferation involved in immune response(GO:0002322) |
| 0.1 | 0.2 | GO:0039513 | modulation by virus of host molecular function(GO:0039506) suppression by virus of host molecular function(GO:0039507) suppression by virus of host catalytic activity(GO:0039513) modulation by virus of host catalytic activity(GO:0039516) suppression by virus of host cysteine-type endopeptidase activity involved in apoptotic process(GO:0039650) negative regulation by symbiont of host catalytic activity(GO:0052053) negative regulation by symbiont of host molecular function(GO:0052056) modulation by symbiont of host catalytic activity(GO:0052148) |
| 0.1 | 0.7 | GO:0034058 | endosomal vesicle fusion(GO:0034058) |
| 0.1 | 1.1 | GO:0010839 | negative regulation of keratinocyte proliferation(GO:0010839) |
| 0.1 | 0.4 | GO:1901189 | positive regulation of ephrin receptor signaling pathway(GO:1901189) positive regulation of canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:1901297) positive regulation of canonical Wnt signaling pathway involved in heart development(GO:1905068) |
| 0.0 | 0.2 | GO:1905150 | regulation of voltage-gated sodium channel activity(GO:1905150) |
| 0.0 | 0.4 | GO:0002318 | myeloid progenitor cell differentiation(GO:0002318) |
| 0.0 | 0.1 | GO:0002752 | cell surface pattern recognition receptor signaling pathway(GO:0002752) |
| 0.0 | 0.3 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 0.0 | 0.2 | GO:0046013 | regulation of T cell homeostatic proliferation(GO:0046013) |
| 0.0 | 0.2 | GO:0007206 | phospholipase C-activating G-protein coupled glutamate receptor signaling pathway(GO:0007206) |
| 0.0 | 0.2 | GO:1903630 | regulation of aminoacyl-tRNA ligase activity(GO:1903630) |
| 0.0 | 0.1 | GO:0090149 | synaptic vesicle recycling via endosome(GO:0036466) mitochondrial membrane fission(GO:0090149) |
| 0.0 | 0.4 | GO:0090292 | nuclear matrix anchoring at nuclear membrane(GO:0090292) |
| 0.0 | 0.5 | GO:0031274 | positive regulation of pseudopodium assembly(GO:0031274) |
| 0.0 | 0.1 | GO:0002188 | translation reinitiation(GO:0002188) |
| 0.0 | 0.2 | GO:0061767 | negative regulation of lung blood pressure(GO:0061767) |
| 0.0 | 0.2 | GO:0044806 | G-quadruplex DNA unwinding(GO:0044806) |
| 0.0 | 0.3 | GO:0000160 | phosphorelay signal transduction system(GO:0000160) |
| 0.0 | 0.1 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 0.0 | 0.0 | GO:0045917 | positive regulation of complement activation(GO:0045917) positive regulation of protein activation cascade(GO:2000259) |
| 0.0 | 0.1 | GO:0000454 | snoRNA guided rRNA pseudouridine synthesis(GO:0000454) |
| 0.0 | 0.1 | GO:0043606 | histidine catabolic process to glutamate and formamide(GO:0019556) histidine catabolic process to glutamate and formate(GO:0019557) formamide metabolic process(GO:0043606) |
| 0.0 | 0.1 | GO:1900194 | negative regulation of oocyte maturation(GO:1900194) |
| 0.0 | 0.2 | GO:0006689 | ganglioside catabolic process(GO:0006689) |
| 0.0 | 0.1 | GO:0071442 | positive regulation of histone H3-K14 acetylation(GO:0071442) |
| 0.0 | 0.5 | GO:0071803 | positive regulation of podosome assembly(GO:0071803) |
| 0.0 | 0.1 | GO:0043605 | cellular amide catabolic process(GO:0043605) |
| 0.0 | 0.1 | GO:1903435 | positive regulation of constitutive secretory pathway(GO:1903435) |
| 0.0 | 0.2 | GO:0007619 | courtship behavior(GO:0007619) female courtship behavior(GO:0008050) |
| 0.0 | 0.1 | GO:0044278 | cell wall disruption in other organism(GO:0044278) |
| 0.0 | 1.8 | GO:2000649 | regulation of sodium ion transmembrane transporter activity(GO:2000649) |
| 0.0 | 0.2 | GO:0086024 | adrenergic receptor signaling pathway involved in positive regulation of heart rate(GO:0086024) |
| 0.0 | 0.3 | GO:0070234 | positive regulation of T cell apoptotic process(GO:0070234) |
| 0.0 | 0.3 | GO:1900119 | positive regulation of execution phase of apoptosis(GO:1900119) |
| 0.0 | 0.3 | GO:0051775 | response to redox state(GO:0051775) |
| 0.0 | 0.0 | GO:0030185 | nitric oxide transport(GO:0030185) |
| 0.0 | 0.1 | GO:2000467 | positive regulation of glycogen (starch) synthase activity(GO:2000467) |
| 0.0 | 0.3 | GO:1901509 | regulation of endothelial tube morphogenesis(GO:1901509) |
| 0.0 | 0.1 | GO:0051771 | negative regulation of nitric-oxide synthase biosynthetic process(GO:0051771) |
| 0.0 | 0.1 | GO:0002175 | protein localization to paranode region of axon(GO:0002175) |
| 0.0 | 0.2 | GO:0060056 | mammary gland involution(GO:0060056) |
| 0.0 | 0.1 | GO:0035234 | ectopic germ cell programmed cell death(GO:0035234) |
| 0.0 | 0.1 | GO:0030950 | establishment or maintenance of actin cytoskeleton polarity(GO:0030950) |
| 0.0 | 2.0 | GO:0071357 | type I interferon signaling pathway(GO:0060337) cellular response to type I interferon(GO:0071357) |
| 0.0 | 0.1 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.0 | 0.1 | GO:0001865 | NK T cell differentiation(GO:0001865) |
| 0.0 | 0.1 | GO:0034127 | regulation of MyD88-independent toll-like receptor signaling pathway(GO:0034127) negative regulation of MyD88-independent toll-like receptor signaling pathway(GO:0034128) |
| 0.0 | 0.5 | GO:0045745 | positive regulation of G-protein coupled receptor protein signaling pathway(GO:0045745) |
| 0.0 | 0.1 | GO:0030854 | positive regulation of granulocyte differentiation(GO:0030854) |
| 0.0 | 0.2 | GO:0070170 | regulation of tooth mineralization(GO:0070170) |
| 0.0 | 0.2 | GO:2000781 | positive regulation of double-strand break repair(GO:2000781) |
| 0.0 | 0.1 | GO:0033141 | positive regulation of peptidyl-serine phosphorylation of STAT protein(GO:0033141) |
| 0.0 | 0.2 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.0 | 0.1 | GO:0060467 | negative regulation of fertilization(GO:0060467) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.3 | GO:0036021 | endolysosome lumen(GO:0036021) |
| 0.0 | 0.4 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.0 | 0.2 | GO:0017101 | aminoacyl-tRNA synthetase multienzyme complex(GO:0017101) |
| 0.0 | 0.2 | GO:0030915 | Smc5-Smc6 complex(GO:0030915) |
| 0.0 | 0.2 | GO:0071144 | SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.0 | 0.3 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.0 | 0.4 | GO:0034992 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.0 | 0.3 | GO:0001520 | outer dense fiber(GO:0001520) |
| 0.0 | 0.1 | GO:0005953 | CAAX-protein geranylgeranyltransferase complex(GO:0005953) |
| 0.0 | 0.2 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.0 | 0.1 | GO:0072558 | NLRP1 inflammasome complex(GO:0072558) |
| 0.0 | 0.1 | GO:0090661 | box H/ACA scaRNP complex(GO:0072589) box H/ACA telomerase RNP complex(GO:0090661) |
| 0.0 | 0.5 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.0 | 0.1 | GO:0030896 | checkpoint clamp complex(GO:0030896) |
| 0.0 | 0.0 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.0 | 1.4 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.0 | 0.2 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) |
| 0.0 | 0.1 | GO:0089701 | U2AF(GO:0089701) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 2.1 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 0.2 | 0.9 | GO:0031735 | CCR10 chemokine receptor binding(GO:0031735) |
| 0.2 | 0.5 | GO:0005519 | cytoskeletal regulatory protein binding(GO:0005519) |
| 0.1 | 0.2 | GO:0017116 | single-stranded DNA-dependent ATP-dependent DNA helicase activity(GO:0017116) |
| 0.1 | 0.3 | GO:0030395 | lactose binding(GO:0030395) |
| 0.1 | 0.2 | GO:0004339 | glucan 1,4-alpha-glucosidase activity(GO:0004339) |
| 0.0 | 0.2 | GO:0052794 | exo-alpha-(2->3)-sialidase activity(GO:0052794) exo-alpha-(2->6)-sialidase activity(GO:0052795) exo-alpha-(2->8)-sialidase activity(GO:0052796) |
| 0.0 | 0.3 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.0 | 0.1 | GO:0017095 | heparan sulfate 6-O-sulfotransferase activity(GO:0017095) |
| 0.0 | 1.1 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 0.0 | 0.2 | GO:0016941 | natriuretic peptide receptor activity(GO:0016941) |
| 0.0 | 0.2 | GO:0097199 | cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:0097199) |
| 0.0 | 0.6 | GO:0004065 | arylsulfatase activity(GO:0004065) |
| 0.0 | 0.2 | GO:0043208 | glycosphingolipid binding(GO:0043208) |
| 0.0 | 2.4 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.0 | 0.2 | GO:0031962 | mineralocorticoid receptor binding(GO:0031962) |
| 0.0 | 0.3 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.0 | 0.4 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.0 | 1.3 | GO:0043394 | proteoglycan binding(GO:0043394) |
| 0.0 | 0.2 | GO:0009019 | tRNA (guanine-N1-)-methyltransferase activity(GO:0009019) |
| 0.0 | 0.1 | GO:0004886 | 9-cis retinoic acid receptor activity(GO:0004886) |
| 0.0 | 0.1 | GO:0003839 | gamma-glutamylcyclotransferase activity(GO:0003839) |
| 0.0 | 0.7 | GO:0005521 | lamin binding(GO:0005521) |
| 0.0 | 0.4 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.0 | 0.3 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.0 | 0.1 | GO:0016841 | ammonia-lyase activity(GO:0016841) |
| 0.0 | 0.1 | GO:0004662 | CAAX-protein geranylgeranyltransferase activity(GO:0004662) |
| 0.0 | 0.1 | GO:0005223 | intracellular cGMP activated cation channel activity(GO:0005223) |
| 0.0 | 0.1 | GO:0004948 | calcitonin receptor activity(GO:0004948) |
| 0.0 | 0.2 | GO:0004064 | arylesterase activity(GO:0004064) |
| 0.0 | 0.0 | GO:0004345 | glucose-6-phosphate dehydrogenase activity(GO:0004345) |
| 0.0 | 0.1 | GO:0004447 | iodide peroxidase activity(GO:0004447) |
| 0.0 | 0.1 | GO:0016167 | glial cell-derived neurotrophic factor receptor activity(GO:0016167) |
| 0.0 | 0.5 | GO:0047555 | cGMP binding(GO:0030553) 3',5'-cyclic-GMP phosphodiesterase activity(GO:0047555) |
| 0.0 | 0.1 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.0 | 0.1 | GO:0034513 | box H/ACA snoRNA binding(GO:0034513) |
| 0.0 | 0.1 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.0 | 0.2 | GO:0015037 | peptide disulfide oxidoreductase activity(GO:0015037) |
| 0.0 | 0.1 | GO:0005332 | gamma-aminobutyric acid:sodium symporter activity(GO:0005332) |
| 0.0 | 0.1 | GO:0030628 | pre-mRNA 3'-splice site binding(GO:0030628) |
| 0.0 | 0.2 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.0 | 0.2 | GO:0070324 | thyroid hormone binding(GO:0070324) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.0 | 2.2 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 0.0 | 0.6 | PID CONE PATHWAY | Visual signal transduction: Cones |
| 0.0 | 1.3 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.0 | 0.7 | PID PI3KCI PATHWAY | Class I PI3K signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.3 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 0.0 | 0.6 | REACTOME THE ACTIVATION OF ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.0 | 2.2 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 0.2 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.0 | 2.0 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 0.6 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.0 | 0.4 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.0 | 0.2 | REACTOME NFKB ACTIVATION THROUGH FADD RIP1 PATHWAY MEDIATED BY CASPASE 8 AND10 | Genes involved in NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 |