Project
Inflammatory response time course, HUVEC (Wada, 2009)
Navigation
Downloads
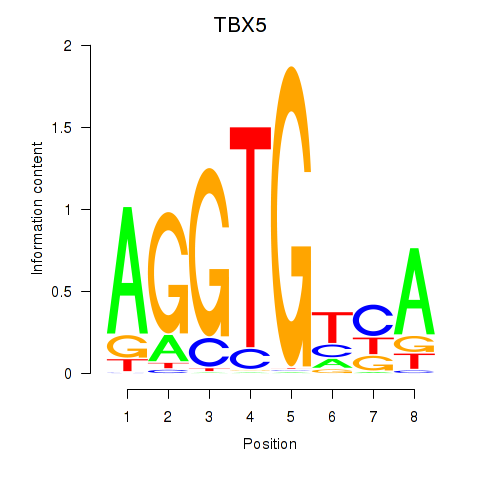
Results for TBX5
Z-value: 0.65
Transcription factors associated with TBX5
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
TBX5
|
ENSG00000089225.20 | TBX5 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| TBX5 | hg38_v1_chr12_-_114406133_114406150 | 0.13 | 5.5e-01 | Click! |
Activity profile of TBX5 motif
Sorted Z-values of TBX5 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of TBX5
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_-_12616762 | 1.47 |
ENST00000464917.5
|
DHRS3
|
dehydrogenase/reductase 3 |
| chr15_+_88638947 | 1.39 |
ENST00000559876.2
|
ISG20
|
interferon stimulated exonuclease gene 20 |
| chr11_-_72674394 | 1.32 |
ENST00000418754.6
ENST00000334456.10 ENST00000542969.2 |
PDE2A
|
phosphodiesterase 2A |
| chr15_+_88639009 | 1.26 |
ENST00000306072.10
|
ISG20
|
interferon stimulated exonuclease gene 20 |
| chr15_-_55289756 | 1.22 |
ENST00000336787.6
|
RAB27A
|
RAB27A, member RAS oncogene family |
| chr8_+_32548303 | 1.11 |
ENST00000650967.1
|
NRG1
|
neuregulin 1 |
| chr11_-_126062782 | 1.07 |
ENST00000531738.6
|
CDON
|
cell adhesion associated, oncogene regulated |
| chr1_-_153549238 | 1.00 |
ENST00000368713.8
|
S100A3
|
S100 calcium binding protein A3 |
| chr2_-_162318129 | 0.98 |
ENST00000679938.1
|
IFIH1
|
interferon induced with helicase C domain 1 |
| chr17_+_79024142 | 0.94 |
ENST00000579760.6
ENST00000339142.6 |
C1QTNF1
|
C1q and TNF related 1 |
| chr17_+_79024243 | 0.93 |
ENST00000311661.4
|
C1QTNF1
|
C1q and TNF related 1 |
| chr8_+_32548210 | 0.92 |
ENST00000523079.5
ENST00000650919.1 |
NRG1
|
neuregulin 1 |
| chr21_+_42219123 | 0.88 |
ENST00000398449.8
|
ABCG1
|
ATP binding cassette subfamily G member 1 |
| chr2_-_2326378 | 0.88 |
ENST00000647618.1
|
MYT1L
|
myelin transcription factor 1 like |
| chr21_+_42219111 | 0.88 |
ENST00000450121.5
ENST00000361802.6 |
ABCG1
|
ATP binding cassette subfamily G member 1 |
| chr1_-_120051714 | 0.74 |
ENST00000579475.7
|
NOTCH2
|
notch receptor 2 |
| chr17_+_79025612 | 0.72 |
ENST00000392445.6
|
C1QTNF1
|
C1q and TNF related 1 |
| chr2_-_175005357 | 0.70 |
ENST00000409156.7
ENST00000444573.2 ENST00000409900.9 |
CHN1
|
chimerin 1 |
| chr5_+_114362043 | 0.70 |
ENST00000673685.1
|
KCNN2
|
potassium calcium-activated channel subfamily N member 2 |
| chr5_+_114362286 | 0.69 |
ENST00000610748.4
ENST00000264773.7 |
KCNN2
|
potassium calcium-activated channel subfamily N member 2 |
| chr21_+_42513834 | 0.68 |
ENST00000352133.3
|
SLC37A1
|
solute carrier family 37 member 1 |
| chr10_+_13100075 | 0.64 |
ENST00000378747.8
ENST00000378757.6 ENST00000378752.7 ENST00000378748.7 |
OPTN
|
optineurin |
| chr17_+_17042433 | 0.62 |
ENST00000651222.2
|
MPRIP
|
myosin phosphatase Rho interacting protein |
| chr16_+_56989479 | 0.62 |
ENST00000262510.10
|
NLRC5
|
NLR family CARD domain containing 5 |
| chr8_+_122781621 | 0.61 |
ENST00000314393.6
|
ZHX2
|
zinc fingers and homeoboxes 2 |
| chr11_-_16356538 | 0.61 |
ENST00000683767.1
|
SOX6
|
SRY-box transcription factor 6 |
| chr11_+_35176611 | 0.55 |
ENST00000279452.10
|
CD44
|
CD44 molecule (Indian blood group) |
| chr7_-_80919017 | 0.55 |
ENST00000265361.8
|
SEMA3C
|
semaphorin 3C |
| chr17_+_76376581 | 0.54 |
ENST00000591651.5
ENST00000545180.5 |
SPHK1
|
sphingosine kinase 1 |
| chr2_+_240625237 | 0.52 |
ENST00000407714.1
|
GPR35
|
G protein-coupled receptor 35 |
| chr15_-_58065703 | 0.49 |
ENST00000249750.9
|
ALDH1A2
|
aldehyde dehydrogenase 1 family member A2 |
| chr11_+_35176696 | 0.48 |
ENST00000528455.5
|
CD44
|
CD44 molecule (Indian blood group) |
| chr15_-_58065734 | 0.48 |
ENST00000347587.7
|
ALDH1A2
|
aldehyde dehydrogenase 1 family member A2 |
| chr2_-_168913277 | 0.47 |
ENST00000451987.5
|
SPC25
|
SPC25 component of NDC80 kinetochore complex |
| chr5_-_67196791 | 0.46 |
ENST00000256447.5
|
CD180
|
CD180 molecule |
| chr13_+_24270681 | 0.46 |
ENST00000343003.10
ENST00000399949.6 |
SPATA13
|
spermatogenesis associated 13 |
| chr16_+_2537997 | 0.46 |
ENST00000441549.7
ENST00000268673.11 ENST00000342085.9 ENST00000389224.7 |
PDPK1
|
3-phosphoinositide dependent protein kinase 1 |
| chr15_-_58065870 | 0.46 |
ENST00000537372.5
|
ALDH1A2
|
aldehyde dehydrogenase 1 family member A2 |
| chrX_-_11665908 | 0.46 |
ENST00000337414.9
|
ARHGAP6
|
Rho GTPase activating protein 6 |
| chrX_-_11427725 | 0.45 |
ENST00000380736.5
|
ARHGAP6
|
Rho GTPase activating protein 6 |
| chr5_+_55024250 | 0.44 |
ENST00000231009.3
|
GZMK
|
granzyme K |
| chrX_-_84502442 | 0.43 |
ENST00000297977.9
ENST00000506585.6 ENST00000373177.3 ENST00000449553.2 |
HDX
|
highly divergent homeobox |
| chr10_+_58269132 | 0.43 |
ENST00000333926.6
|
CISD1
|
CDGSH iron sulfur domain 1 |
| chr1_-_10982037 | 0.41 |
ENST00000377004.8
|
C1orf127
|
chromosome 1 open reading frame 127 |
| chr19_-_38617912 | 0.41 |
ENST00000591517.5
|
MAP4K1
|
mitogen-activated protein kinase kinase kinase kinase 1 |
| chr9_+_137230757 | 0.41 |
ENST00000673865.1
ENST00000538474.5 ENST00000673835.1 ENST00000673953.1 ENST00000361134.2 |
SLC34A3
|
solute carrier family 34 member 3 |
| chr4_+_139301478 | 0.40 |
ENST00000296543.10
ENST00000398947.1 |
NAA15
|
N-alpha-acetyltransferase 15, NatA auxiliary subunit |
| chr1_-_72282457 | 0.40 |
ENST00000357731.10
|
NEGR1
|
neuronal growth regulator 1 |
| chr2_-_144516154 | 0.39 |
ENST00000637304.1
|
ZEB2
|
zinc finger E-box binding homeobox 2 |
| chr19_-_54360949 | 0.39 |
ENST00000622064.1
|
LAIR1
|
leukocyte associated immunoglobulin like receptor 1 |
| chr1_+_26410809 | 0.39 |
ENST00000254231.4
ENST00000326279.11 |
LIN28A
|
lin-28 homolog A |
| chr19_-_38617928 | 0.39 |
ENST00000396857.7
ENST00000586296.5 |
MAP4K1
|
mitogen-activated protein kinase kinase kinase kinase 1 |
| chr11_+_35176575 | 0.38 |
ENST00000526000.6
|
CD44
|
CD44 molecule (Indian blood group) |
| chr19_+_7669080 | 0.38 |
ENST00000629642.1
|
RETN
|
resistin |
| chr14_+_22281097 | 0.38 |
ENST00000390465.2
|
TRAV38-2DV8
|
T cell receptor alpha variable 38-2/delta variable 8 |
| chr6_+_45422564 | 0.38 |
ENST00000625924.1
|
RUNX2
|
RUNX family transcription factor 2 |
| chr3_-_183162726 | 0.37 |
ENST00000265598.8
|
LAMP3
|
lysosomal associated membrane protein 3 |
| chr11_+_35176639 | 0.37 |
ENST00000527889.6
|
CD44
|
CD44 molecule (Indian blood group) |
| chr16_-_69351778 | 0.36 |
ENST00000288025.4
|
TMED6
|
transmembrane p24 trafficking protein 6 |
| chr19_+_35138993 | 0.35 |
ENST00000612146.4
ENST00000589209.5 |
FXYD1
|
FXYD domain containing ion transport regulator 1 |
| chr8_+_32548267 | 0.35 |
ENST00000356819.7
|
NRG1
|
neuregulin 1 |
| chr16_+_2988256 | 0.35 |
ENST00000573315.2
|
LINC00514
|
long intergenic non-protein coding RNA 514 |
| chr7_-_27095972 | 0.35 |
ENST00000355633.5
ENST00000643460.2 |
HOXA1
|
homeobox A1 |
| chr10_-_101229449 | 0.34 |
ENST00000370193.4
|
LBX1
|
ladybird homeobox 1 |
| chr6_+_45422485 | 0.34 |
ENST00000359524.7
|
RUNX2
|
RUNX family transcription factor 2 |
| chr11_-_45907265 | 0.33 |
ENST00000449465.2
|
C11orf94
|
chromosome 11 open reading frame 94 |
| chr22_-_31489528 | 0.33 |
ENST00000397525.5
|
EIF4ENIF1
|
eukaryotic translation initiation factor 4E nuclear import factor 1 |
| chr6_+_30327259 | 0.32 |
ENST00000376659.9
ENST00000428555.1 |
TRIM39
|
tripartite motif containing 39 |
| chr17_+_36103819 | 0.32 |
ENST00000615863.2
ENST00000621626.1 |
CCL4
|
C-C motif chemokine ligand 4 |
| chr4_+_8580387 | 0.32 |
ENST00000382487.5
|
GPR78
|
G protein-coupled receptor 78 |
| chr2_+_68734773 | 0.32 |
ENST00000409202.8
|
ARHGAP25
|
Rho GTPase activating protein 25 |
| chr19_-_47231191 | 0.32 |
ENST00000439096.3
|
BBC3
|
BCL2 binding component 3 |
| chr1_-_156816841 | 0.32 |
ENST00000368199.8
ENST00000392306.2 |
SH2D2A
|
SH2 domain containing 2A |
| chr3_-_39192584 | 0.31 |
ENST00000340369.4
ENST00000421646.1 ENST00000396251.1 |
XIRP1
|
xin actin binding repeat containing 1 |
| chr1_-_19923617 | 0.31 |
ENST00000375116.3
|
PLA2G2E
|
phospholipase A2 group IIE |
| chr4_+_80197493 | 0.30 |
ENST00000415738.3
|
PRDM8
|
PR/SET domain 8 |
| chr2_+_68734861 | 0.30 |
ENST00000467265.5
|
ARHGAP25
|
Rho GTPase activating protein 25 |
| chr1_-_156816738 | 0.29 |
ENST00000368198.7
|
SH2D2A
|
SH2 domain containing 2A |
| chr3_-_134029914 | 0.29 |
ENST00000493729.5
ENST00000310926.11 |
SLCO2A1
|
solute carrier organic anion transporter family member 2A1 |
| chr14_+_24310170 | 0.29 |
ENST00000530080.1
|
LTB4R2
|
leukotriene B4 receptor 2 |
| chr8_+_31639291 | 0.28 |
ENST00000651149.1
ENST00000650866.1 |
NRG1
|
neuregulin 1 |
| chr14_-_50830479 | 0.28 |
ENST00000382043.8
|
NIN
|
ninein |
| chr6_+_30326835 | 0.28 |
ENST00000440271.5
ENST00000376656.8 ENST00000396551.7 ENST00000428728.5 ENST00000396548.5 ENST00000428404.5 |
TRIM39
|
tripartite motif containing 39 |
| chr7_-_76626127 | 0.28 |
ENST00000454397.1
|
POMZP3
|
POM121 and ZP3 fusion |
| chr11_-_123654581 | 0.28 |
ENST00000392770.6
ENST00000530277.5 ENST00000299333.8 |
SCN3B
|
sodium voltage-gated channel beta subunit 3 |
| chr3_-_24165548 | 0.28 |
ENST00000280696.9
|
THRB
|
thyroid hormone receptor beta |
| chr8_+_31639222 | 0.28 |
ENST00000519301.6
ENST00000652698.1 |
NRG1
|
neuregulin 1 |
| chr20_+_836052 | 0.27 |
ENST00000246100.3
|
FAM110A
|
family with sequence similarity 110 member A |
| chr2_+_209771972 | 0.27 |
ENST00000439458.5
ENST00000272845.10 |
UNC80
|
unc-80 homolog, NALCN channel complex subunit |
| chr2_+_209771814 | 0.27 |
ENST00000673951.1
ENST00000673920.1 |
UNC80
|
unc-80 homolog, NALCN channel complex subunit |
| chr14_+_21057822 | 0.27 |
ENST00000308227.2
|
RNASE8
|
ribonuclease A family member 8 |
| chr12_+_107318395 | 0.27 |
ENST00000420571.6
ENST00000280758.10 |
BTBD11
|
BTB domain containing 11 |
| chr17_-_35089212 | 0.27 |
ENST00000584655.5
ENST00000447669.6 ENST00000315249.11 |
RFFL
|
ring finger and FYVE like domain containing E3 ubiquitin protein ligase |
| chr8_+_1858750 | 0.26 |
ENST00000398560.2
|
ARHGEF10
|
Rho guanine nucleotide exchange factor 10 |
| chr18_+_61333424 | 0.26 |
ENST00000262717.9
|
CDH20
|
cadherin 20 |
| chrX_+_70423031 | 0.26 |
ENST00000453994.6
ENST00000538649.5 ENST00000536730.5 |
GDPD2
|
glycerophosphodiester phosphodiesterase domain containing 2 |
| chr2_+_201129318 | 0.26 |
ENST00000417748.1
|
CFLAR
|
CASP8 and FADD like apoptosis regulator |
| chr1_-_150010675 | 0.25 |
ENST00000417191.2
ENST00000581312.6 |
OTUD7B
|
OTU deubiquitinase 7B |
| chrX_+_70423301 | 0.25 |
ENST00000374382.4
|
GDPD2
|
glycerophosphodiester phosphodiesterase domain containing 2 |
| chr12_+_18738102 | 0.25 |
ENST00000317658.5
|
CAPZA3
|
capping actin protein of muscle Z-line subunit alpha 3 |
| chr7_+_97005538 | 0.25 |
ENST00000518156.3
|
DLX6
|
distal-less homeobox 6 |
| chr20_+_49982969 | 0.25 |
ENST00000244050.3
|
SNAI1
|
snail family transcriptional repressor 1 |
| chr1_-_206921867 | 0.25 |
ENST00000628511.2
ENST00000367091.8 |
FCMR
|
Fc fragment of IgM receptor |
| chr15_+_41621492 | 0.25 |
ENST00000570161.6
|
MGA
|
MAX dimerization protein MGA |
| chr16_-_50681328 | 0.25 |
ENST00000300590.7
|
SNX20
|
sorting nexin 20 |
| chr11_-_34513785 | 0.25 |
ENST00000257832.7
ENST00000429939.6 |
ELF5
|
E74 like ETS transcription factor 5 |
| chr11_-_41459592 | 0.25 |
ENST00000528697.6
ENST00000530763.5 |
LRRC4C
|
leucine rich repeat containing 4C |
| chr4_-_156971769 | 0.24 |
ENST00000502773.6
|
PDGFC
|
platelet derived growth factor C |
| chr18_+_34493289 | 0.24 |
ENST00000682923.1
ENST00000596745.5 ENST00000283365.14 ENST00000315456.10 ENST00000598774.6 ENST00000684266.1 ENST00000683092.1 ENST00000683379.1 ENST00000684359.1 |
DTNA
|
dystrobrevin alpha |
| chr15_+_36702009 | 0.24 |
ENST00000562489.1
|
CDIN1
|
CDAN1 interacting nuclease 1 |
| chr11_+_131911396 | 0.24 |
ENST00000425719.6
ENST00000374784.5 |
NTM
|
neurotrimin |
| chr3_+_9649433 | 0.24 |
ENST00000353332.9
ENST00000420925.5 ENST00000296003.9 ENST00000351233.9 |
MTMR14
|
myotubularin related protein 14 |
| chr17_+_3207539 | 0.24 |
ENST00000641322.1
ENST00000641732.2 |
OR1A1
|
olfactory receptor family 1 subfamily A member 1 |
| chr16_-_4345904 | 0.24 |
ENST00000571941.5
|
PAM16
|
presequence translocase associated motor 16 |
| chr17_-_76537699 | 0.24 |
ENST00000293230.10
|
CYGB
|
cytoglobin |
| chr17_+_58192723 | 0.23 |
ENST00000225371.6
|
EPX
|
eosinophil peroxidase |
| chr6_+_34236865 | 0.23 |
ENST00000674029.1
ENST00000447654.5 ENST00000347617.10 ENST00000401473.7 ENST00000311487.9 |
HMGA1
|
high mobility group AT-hook 1 |
| chr12_-_52834307 | 0.23 |
ENST00000330553.6
|
KRT79
|
keratin 79 |
| chr5_+_140107777 | 0.23 |
ENST00000505703.2
ENST00000651386.1 |
PURA
|
purine rich element binding protein A |
| chr9_+_4662281 | 0.23 |
ENST00000381883.5
|
PLPP6
|
phospholipid phosphatase 6 |
| chr18_+_34493428 | 0.22 |
ENST00000682483.1
|
DTNA
|
dystrobrevin alpha |
| chrX_-_133415478 | 0.22 |
ENST00000370828.4
|
GPC4
|
glypican 4 |
| chr19_+_38304105 | 0.22 |
ENST00000588605.5
ENST00000301246.10 |
C19orf33
|
chromosome 19 open reading frame 33 |
| chr20_+_31819302 | 0.22 |
ENST00000375994.6
|
MYLK2
|
myosin light chain kinase 2 |
| chrX_+_96684712 | 0.22 |
ENST00000373049.8
|
DIAPH2
|
diaphanous related formin 2 |
| chrX_-_103832204 | 0.22 |
ENST00000674363.1
ENST00000674162.1 ENST00000674338.1 ENST00000674274.1 ENST00000674271.1 ENST00000674265.1 ENST00000674212.1 ENST00000674255.1 ENST00000674342.1 ENST00000674430.1 ENST00000243298.3 |
ENSG00000288597.1
RAB9B
|
novel transcript RAB9B, member RAS oncogene family |
| chr19_+_1000419 | 0.22 |
ENST00000234389.3
|
GRIN3B
|
glutamate ionotropic receptor NMDA type subunit 3B |
| chr6_+_25652201 | 0.22 |
ENST00000612225.4
ENST00000377961.3 |
SCGN
|
secretagogin, EF-hand calcium binding protein |
| chr1_+_148889403 | 0.22 |
ENST00000464103.5
ENST00000534536.5 ENST00000369356.8 ENST00000369354.7 ENST00000369347.8 ENST00000369349.7 ENST00000369351.7 |
PDE4DIP
|
phosphodiesterase 4D interacting protein |
| chr16_+_57186281 | 0.22 |
ENST00000564435.5
ENST00000562959.1 ENST00000568505.6 ENST00000394420.9 ENST00000537866.5 |
RSPRY1
|
ring finger and SPRY domain containing 1 |
| chr12_-_116276759 | 0.22 |
ENST00000548743.2
|
MED13L
|
mediator complex subunit 13L |
| chr18_+_34493386 | 0.22 |
ENST00000679936.1
|
DTNA
|
dystrobrevin alpha |
| chr5_+_141489150 | 0.22 |
ENST00000610789.1
|
PCDHGC5
|
protocadherin gamma subfamily C, 5 |
| chr2_-_223837484 | 0.21 |
ENST00000446015.6
ENST00000409375.1 |
AP1S3
|
adaptor related protein complex 1 subunit sigma 3 |
| chr6_+_25652272 | 0.21 |
ENST00000334979.6
|
SCGN
|
secretagogin, EF-hand calcium binding protein |
| chr12_+_30926716 | 0.21 |
ENST00000546076.6
ENST00000535215.5 ENST00000261177.10 |
TSPAN11
|
tetraspanin 11 |
| chr17_-_7394514 | 0.21 |
ENST00000571802.1
ENST00000619711.5 ENST00000576201.5 ENST00000573213.1 ENST00000324822.15 |
PLSCR3
|
phospholipid scramblase 3 |
| chr20_+_31819348 | 0.21 |
ENST00000375985.5
|
MYLK2
|
myosin light chain kinase 2 |
| chr2_-_118014035 | 0.21 |
ENST00000376300.7
|
CCDC93
|
coiled-coil domain containing 93 |
| chr15_+_41621134 | 0.20 |
ENST00000566718.6
|
MGA
|
MAX dimerization protein MGA |
| chr1_+_202462730 | 0.20 |
ENST00000290419.9
ENST00000491336.5 |
PPP1R12B
|
protein phosphatase 1 regulatory subunit 12B |
| chr14_-_23408265 | 0.20 |
ENST00000405093.9
|
MYH6
|
myosin heavy chain 6 |
| chr15_+_81296913 | 0.20 |
ENST00000394652.6
|
IL16
|
interleukin 16 |
| chr5_+_83471668 | 0.20 |
ENST00000342785.8
ENST00000343200.9 |
VCAN
|
versican |
| chr17_-_7208325 | 0.20 |
ENST00000650120.1
ENST00000648760.1 |
DLG4
|
discs large MAGUK scaffold protein 4 |
| chr1_-_41918666 | 0.20 |
ENST00000372584.5
|
HIVEP3
|
HIVEP zinc finger 3 |
| chr15_+_88803468 | 0.20 |
ENST00000558207.5
|
ACAN
|
aggrecan |
| chrX_+_103607906 | 0.20 |
ENST00000243286.7
ENST00000372627.10 |
TCEAL3
|
transcription elongation factor A like 3 |
| chr17_-_7394240 | 0.20 |
ENST00000576362.5
ENST00000571078.5 |
PLSCR3
|
phospholipid scramblase 3 |
| chr1_-_205455954 | 0.20 |
ENST00000495594.2
ENST00000629624.2 |
LEMD1
BLACAT1
|
LEM domain containing 1 bladder cancer associated transcript 1 |
| chr6_-_46015812 | 0.20 |
ENST00000544153.3
ENST00000339561.12 |
CLIC5
|
chloride intracellular channel 5 |
| chr1_-_241357085 | 0.19 |
ENST00000366564.5
|
RGS7
|
regulator of G protein signaling 7 |
| chr2_-_118014125 | 0.19 |
ENST00000319432.9
|
CCDC93
|
coiled-coil domain containing 93 |
| chr11_-_74949114 | 0.19 |
ENST00000527087.5
|
XRRA1
|
X-ray radiation resistance associated 1 |
| chr19_+_35748549 | 0.19 |
ENST00000301159.14
|
LIN37
|
lin-37 DREAM MuvB core complex component |
| chr1_+_46671821 | 0.19 |
ENST00000334122.5
ENST00000415500.1 |
TEX38
|
testis expressed 38 |
| chr19_-_10309783 | 0.19 |
ENST00000403352.1
ENST00000403903.7 |
ZGLP1
|
zinc finger GATA like protein 1 |
| chr1_-_183653307 | 0.19 |
ENST00000308641.6
|
APOBEC4
|
apolipoprotein B mRNA editing enzyme catalytic polypeptide like 4 |
| chr22_-_21629991 | 0.18 |
ENST00000292778.11
ENST00000398873.4 |
YDJC
|
YdjC chitooligosaccharide deacetylase homolog |
| chr2_+_120346130 | 0.18 |
ENST00000295228.4
|
INHBB
|
inhibin subunit beta B |
| chr17_-_3916455 | 0.18 |
ENST00000225538.4
|
P2RX1
|
purinergic receptor P2X 1 |
| chr11_-_95910665 | 0.18 |
ENST00000674610.1
ENST00000675660.1 |
MTMR2
|
myotubularin related protein 2 |
| chr12_+_56118241 | 0.18 |
ENST00000551790.5
ENST00000552345.1 ENST00000257940.7 ENST00000551880.1 |
ESYT1
ZC3H10
|
extended synaptotagmin 1 zinc finger CCCH-type containing 10 |
| chr12_+_1629197 | 0.18 |
ENST00000397196.7
|
WNT5B
|
Wnt family member 5B |
| chrX_+_96684638 | 0.18 |
ENST00000355827.8
ENST00000373061.7 |
DIAPH2
|
diaphanous related formin 2 |
| chr12_-_102478539 | 0.18 |
ENST00000424202.6
|
IGF1
|
insulin like growth factor 1 |
| chr1_-_241357225 | 0.18 |
ENST00000366565.5
|
RGS7
|
regulator of G protein signaling 7 |
| chr1_-_241357171 | 0.18 |
ENST00000440928.6
|
RGS7
|
regulator of G protein signaling 7 |
| chr8_+_103819244 | 0.17 |
ENST00000262231.14
ENST00000507740.5 ENST00000408894.6 |
RIMS2
|
regulating synaptic membrane exocytosis 2 |
| chr12_-_52434363 | 0.17 |
ENST00000252245.6
|
KRT75
|
keratin 75 |
| chr17_+_32487686 | 0.17 |
ENST00000584792.5
|
CDK5R1
|
cyclin dependent kinase 5 regulatory subunit 1 |
| chr19_+_48364361 | 0.17 |
ENST00000344846.7
|
SYNGR4
|
synaptogyrin 4 |
| chr12_-_120904337 | 0.17 |
ENST00000353487.7
|
SPPL3
|
signal peptide peptidase like 3 |
| chr11_-_74949079 | 0.17 |
ENST00000528219.5
ENST00000684022.1 ENST00000531852.5 |
XRRA1
|
X-ray radiation resistance associated 1 |
| chr11_+_450255 | 0.17 |
ENST00000308020.6
|
PTDSS2
|
phosphatidylserine synthase 2 |
| chr12_+_53252085 | 0.17 |
ENST00000329548.5
|
MFSD5
|
major facilitator superfamily domain containing 5 |
| chr17_+_55266216 | 0.17 |
ENST00000573945.5
|
HLF
|
HLF transcription factor, PAR bZIP family member |
| chr11_-_40293102 | 0.17 |
ENST00000527150.5
|
LRRC4C
|
leucine rich repeat containing 4C |
| chr3_+_167735704 | 0.17 |
ENST00000446050.7
ENST00000295777.9 ENST00000472747.2 |
SERPINI1
|
serpin family I member 1 |
| chr1_-_1959974 | 0.16 |
ENST00000642590.1
|
CFAP74
|
cilia and flagella associated protein 74 |
| chr17_-_15566276 | 0.16 |
ENST00000630868.1
|
CDRT1
|
CMT1A duplicated region transcript 1 |
| chr2_-_24972032 | 0.16 |
ENST00000534855.5
|
DNAJC27
|
DnaJ heat shock protein family (Hsp40) member C27 |
| chr1_-_37034492 | 0.16 |
ENST00000373091.8
|
GRIK3
|
glutamate ionotropic receptor kainate type subunit 3 |
| chr5_+_132673983 | 0.16 |
ENST00000622422.1
ENST00000231449.7 ENST00000350025.2 |
IL4
|
interleukin 4 |
| chr11_-_74949142 | 0.16 |
ENST00000321448.12
ENST00000340360.10 |
XRRA1
|
X-ray radiation resistance associated 1 |
| chr11_-_132943671 | 0.16 |
ENST00000331898.11
|
OPCML
|
opioid binding protein/cell adhesion molecule like |
| chr3_-_182985926 | 0.16 |
ENST00000487822.5
ENST00000460412.6 ENST00000469954.5 |
DCUN1D1
|
defective in cullin neddylation 1 domain containing 1 |
| chr11_-_45917867 | 0.16 |
ENST00000378750.10
|
PEX16
|
peroxisomal biogenesis factor 16 |
| chr17_+_9162935 | 0.16 |
ENST00000436734.1
|
NTN1
|
netrin 1 |
| chr8_+_17027230 | 0.16 |
ENST00000318063.10
|
MICU3
|
mitochondrial calcium uptake family member 3 |
| chr11_-_45918014 | 0.16 |
ENST00000525192.5
|
PEX16
|
peroxisomal biogenesis factor 16 |
| chr9_+_122375286 | 0.16 |
ENST00000373698.7
|
PTGS1
|
prostaglandin-endoperoxide synthase 1 |
| chr19_-_49441508 | 0.16 |
ENST00000221485.8
|
SLC17A7
|
solute carrier family 17 member 7 |
| chr19_+_41443130 | 0.16 |
ENST00000378187.3
|
ERICH4
|
glutamate rich 4 |
| chr5_+_126423122 | 0.16 |
ENST00000515200.5
|
GRAMD2B
|
GRAM domain containing 2B |
| chr12_-_48957365 | 0.16 |
ENST00000398092.4
ENST00000539611.1 |
ENSG00000272822.2
ARF3
|
novel protein ADP ribosylation factor 3 |
| chr12_+_104064520 | 0.16 |
ENST00000229330.9
|
HCFC2
|
host cell factor C2 |
| chr2_-_223837553 | 0.16 |
ENST00000396654.7
ENST00000396653.2 ENST00000443700.5 |
AP1S3
|
adaptor related protein complex 1 subunit sigma 3 |
| chr5_-_115180037 | 0.16 |
ENST00000514154.1
ENST00000282369.7 |
TRIM36
|
tripartite motif containing 36 |
| chr12_-_18738006 | 0.16 |
ENST00000266505.12
ENST00000543242.5 ENST00000539072.5 ENST00000541966.5 ENST00000648272.1 |
PLCZ1
|
phospholipase C zeta 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.6 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 0.6 | 1.8 | GO:0009720 | detection of hormone stimulus(GO:0009720) |
| 0.4 | 2.6 | GO:2000857 | positive regulation of mineralocorticoid secretion(GO:2000857) positive regulation of aldosterone secretion(GO:2000860) |
| 0.3 | 1.3 | GO:0033159 | negative regulation of protein import into nucleus, translocation(GO:0033159) |
| 0.2 | 2.9 | GO:0038129 | directional guidance of interneurons involved in migration from the subpallium to the cortex(GO:0021840) chemorepulsion involved in interneuron migration from the subpallium to the cortex(GO:0021842) ERBB3 signaling pathway(GO:0038129) |
| 0.2 | 1.2 | GO:1903435 | positive regulation of constitutive secretory pathway(GO:1903435) |
| 0.2 | 1.0 | GO:0034343 | type III interferon production(GO:0034343) regulation of type III interferon production(GO:0034344) |
| 0.2 | 0.5 | GO:0046521 | sphingoid catabolic process(GO:0046521) |
| 0.2 | 1.4 | GO:0042905 | 9-cis-retinoic acid biosynthetic process(GO:0042904) 9-cis-retinoic acid metabolic process(GO:0042905) |
| 0.1 | 1.8 | GO:1900623 | regulation of monocyte aggregation(GO:1900623) positive regulation of monocyte aggregation(GO:1900625) |
| 0.1 | 0.5 | GO:2000196 | positive regulation of female gonad development(GO:2000196) |
| 0.1 | 0.6 | GO:0021779 | oligodendrocyte cell fate specification(GO:0021778) oligodendrocyte cell fate commitment(GO:0021779) glial cell fate specification(GO:0021780) |
| 0.1 | 0.5 | GO:0032581 | ER-dependent peroxisome organization(GO:0032581) |
| 0.1 | 0.5 | GO:0003350 | pulmonary myocardium development(GO:0003350) |
| 0.1 | 0.6 | GO:0060332 | positive regulation of response to interferon-gamma(GO:0060332) positive regulation of interferon-gamma-mediated signaling pathway(GO:0060335) |
| 0.1 | 1.4 | GO:0098914 | membrane repolarization during atrial cardiac muscle cell action potential(GO:0098914) |
| 0.1 | 1.5 | GO:0048387 | negative regulation of retinoic acid receptor signaling pathway(GO:0048387) |
| 0.1 | 0.3 | GO:0090222 | centrosome-templated microtubule nucleation(GO:0090222) |
| 0.1 | 0.6 | GO:0001920 | negative regulation of receptor recycling(GO:0001920) |
| 0.1 | 0.7 | GO:0015760 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.1 | 0.7 | GO:0002051 | osteoblast fate commitment(GO:0002051) |
| 0.1 | 0.4 | GO:0010587 | miRNA catabolic process(GO:0010587) |
| 0.1 | 0.5 | GO:0051970 | negative regulation of transmission of nerve impulse(GO:0051970) |
| 0.1 | 1.1 | GO:0060059 | embryonic retina morphogenesis in camera-type eye(GO:0060059) |
| 0.1 | 0.3 | GO:0035803 | egg coat formation(GO:0035803) |
| 0.1 | 0.5 | GO:0002322 | B cell proliferation involved in immune response(GO:0002322) |
| 0.1 | 0.2 | GO:1904328 | regulation of myofibroblast contraction(GO:1904328) myofibroblast contraction(GO:1990764) |
| 0.1 | 0.4 | GO:2000503 | positive regulation of natural killer cell chemotaxis(GO:2000503) |
| 0.1 | 0.4 | GO:0032971 | regulation of muscle filament sliding(GO:0032971) |
| 0.1 | 0.2 | GO:0060279 | positive regulation of ovulation(GO:0060279) |
| 0.1 | 0.4 | GO:0010734 | protein glutathionylation(GO:0010731) regulation of protein glutathionylation(GO:0010732) negative regulation of protein glutathionylation(GO:0010734) |
| 0.1 | 0.2 | GO:0002215 | defense response to nematode(GO:0002215) |
| 0.1 | 0.2 | GO:0021718 | superior olivary nucleus development(GO:0021718) superior olivary nucleus maturation(GO:0021722) |
| 0.1 | 0.2 | GO:0006659 | phosphatidylserine biosynthetic process(GO:0006659) |
| 0.1 | 0.2 | GO:0007314 | oocyte construction(GO:0007308) oocyte axis specification(GO:0007309) oocyte anterior/posterior axis specification(GO:0007314) pole plasm assembly(GO:0007315) maternal determination of anterior/posterior axis, embryo(GO:0008358) P granule organization(GO:0030719) |
| 0.1 | 0.1 | GO:0097325 | melanocyte proliferation(GO:0097325) |
| 0.1 | 0.3 | GO:0061760 | antifungal innate immune response(GO:0061760) |
| 0.1 | 0.3 | GO:2001271 | negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 0.1 | 0.2 | GO:0042137 | sequestering of neurotransmitter(GO:0042137) |
| 0.0 | 1.0 | GO:0061314 | Notch signaling involved in heart development(GO:0061314) |
| 0.0 | 0.4 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.0 | 0.2 | GO:0090402 | oncogene-induced cell senescence(GO:0090402) |
| 0.0 | 0.4 | GO:0072502 | cellular phosphate ion homeostasis(GO:0030643) cellular trivalent inorganic anion homeostasis(GO:0072502) |
| 0.0 | 0.4 | GO:0017196 | N-terminal peptidyl-methionine acetylation(GO:0017196) |
| 0.0 | 0.2 | GO:0035519 | protein K29-linked ubiquitination(GO:0035519) |
| 0.0 | 0.1 | GO:1903719 | regulation of I-kappaB phosphorylation(GO:1903719) positive regulation of I-kappaB phosphorylation(GO:1903721) |
| 0.0 | 0.3 | GO:0007621 | negative regulation of female receptivity(GO:0007621) |
| 0.0 | 0.3 | GO:0019255 | UDP-glucuronate biosynthetic process(GO:0006065) glucose 1-phosphate metabolic process(GO:0019255) |
| 0.0 | 0.3 | GO:0071947 | protein deubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0071947) |
| 0.0 | 0.3 | GO:2000490 | negative regulation of hepatic stellate cell activation(GO:2000490) |
| 0.0 | 0.2 | GO:0002296 | T-helper 1 cell lineage commitment(GO:0002296) |
| 0.0 | 0.1 | GO:1902303 | regulation of heart rate by hormone(GO:0003064) negative regulation of potassium ion export(GO:1902303) |
| 0.0 | 0.1 | GO:0061537 | glycine secretion(GO:0061536) glycine secretion, neurotransmission(GO:0061537) |
| 0.0 | 0.1 | GO:0051039 | histone displacement(GO:0001207) positive regulation of transcription involved in meiotic cell cycle(GO:0051039) |
| 0.0 | 0.1 | GO:1901877 | regulation of calcium ion binding(GO:1901876) negative regulation of calcium ion binding(GO:1901877) |
| 0.0 | 0.1 | GO:0015855 | pyrimidine nucleobase transport(GO:0015855) purine nucleobase transmembrane transport(GO:1904823) |
| 0.0 | 0.2 | GO:0002442 | serotonin production involved in inflammatory response(GO:0002351) serotonin secretion involved in inflammatory response(GO:0002442) serotonin secretion by platelet(GO:0002554) |
| 0.0 | 0.1 | GO:0044333 | Wnt signaling pathway involved in digestive tract morphogenesis(GO:0044333) |
| 0.0 | 0.4 | GO:1903944 | regulation of hepatocyte apoptotic process(GO:1903943) negative regulation of hepatocyte apoptotic process(GO:1903944) |
| 0.0 | 0.1 | GO:0033861 | negative regulation of NAD(P)H oxidase activity(GO:0033861) |
| 0.0 | 0.2 | GO:1904073 | regulation of trophectodermal cell proliferation(GO:1904073) positive regulation of trophectodermal cell proliferation(GO:1904075) |
| 0.0 | 0.2 | GO:0045204 | MAPK export from nucleus(GO:0045204) |
| 0.0 | 0.2 | GO:0001712 | ectoderm formation(GO:0001705) ectodermal cell fate commitment(GO:0001712) |
| 0.0 | 0.3 | GO:0060373 | regulation of ventricular cardiac muscle cell membrane depolarization(GO:0060373) |
| 0.0 | 0.2 | GO:1902626 | assembly of large subunit precursor of preribosome(GO:1902626) |
| 0.0 | 0.3 | GO:0061737 | leukotriene signaling pathway(GO:0061737) |
| 0.0 | 0.2 | GO:0003228 | atrial cardiac muscle tissue development(GO:0003228) atrial cardiac muscle tissue morphogenesis(GO:0055009) |
| 0.0 | 0.0 | GO:1903937 | response to acrylamide(GO:1903937) |
| 0.0 | 0.3 | GO:0048752 | semicircular canal morphogenesis(GO:0048752) |
| 0.0 | 0.2 | GO:0007343 | egg activation(GO:0007343) |
| 0.0 | 0.0 | GO:0010847 | regulation of chromatin assembly(GO:0010847) |
| 0.0 | 0.2 | GO:0060370 | susceptibility to T cell mediated cytotoxicity(GO:0060370) |
| 0.0 | 0.2 | GO:0046340 | diacylglycerol catabolic process(GO:0046340) |
| 0.0 | 0.1 | GO:0007206 | phospholipase C-activating G-protein coupled glutamate receptor signaling pathway(GO:0007206) |
| 0.0 | 0.3 | GO:0060753 | regulation of mast cell chemotaxis(GO:0060753) |
| 0.0 | 0.6 | GO:2000059 | negative regulation of protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:2000059) |
| 0.0 | 0.3 | GO:0048664 | neuron fate determination(GO:0048664) |
| 0.0 | 0.1 | GO:0001994 | norepinephrine-epinephrine vasoconstriction involved in regulation of systemic arterial blood pressure(GO:0001994) |
| 0.0 | 0.1 | GO:0001897 | cytolysis by symbiont of host cells(GO:0001897) |
| 0.0 | 0.3 | GO:0030091 | protein repair(GO:0030091) |
| 0.0 | 0.1 | GO:0002439 | chronic inflammatory response to antigenic stimulus(GO:0002439) |
| 0.0 | 0.1 | GO:1904397 | positive regulation of skeletal muscle acetylcholine-gated channel clustering(GO:1904395) negative regulation of neuromuscular junction development(GO:1904397) |
| 0.0 | 0.3 | GO:0031642 | negative regulation of myelination(GO:0031642) |
| 0.0 | 0.8 | GO:0000185 | activation of MAPKKK activity(GO:0000185) |
| 0.0 | 0.5 | GO:0017121 | phospholipid scrambling(GO:0017121) |
| 0.0 | 0.5 | GO:0090527 | actin filament reorganization(GO:0090527) |
| 0.0 | 0.1 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 0.0 | 0.2 | GO:0061669 | spontaneous neurotransmitter secretion(GO:0061669) spontaneous synaptic transmission(GO:0098814) |
| 0.0 | 0.3 | GO:0050930 | induction of positive chemotaxis(GO:0050930) |
| 0.0 | 0.5 | GO:0010667 | negative regulation of cardiac muscle cell apoptotic process(GO:0010667) |
| 0.0 | 0.1 | GO:0007518 | myoblast fate determination(GO:0007518) |
| 0.0 | 0.1 | GO:0030221 | basophil differentiation(GO:0030221) |
| 0.0 | 0.3 | GO:0042340 | keratan sulfate catabolic process(GO:0042340) |
| 0.0 | 0.2 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.0 | 0.1 | GO:0042335 | cuticle development(GO:0042335) |
| 0.0 | 0.1 | GO:0050882 | voluntary musculoskeletal movement(GO:0050882) |
| 0.0 | 0.1 | GO:1902896 | terminal web assembly(GO:1902896) |
| 0.0 | 0.6 | GO:0051895 | negative regulation of focal adhesion assembly(GO:0051895) |
| 0.0 | 0.2 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
| 0.0 | 0.2 | GO:0098735 | positive regulation of the force of heart contraction(GO:0098735) |
| 0.0 | 0.3 | GO:0034374 | low-density lipoprotein particle remodeling(GO:0034374) |
| 0.0 | 0.3 | GO:0016188 | synaptic vesicle maturation(GO:0016188) |
| 0.0 | 0.1 | GO:0001661 | conditioned taste aversion(GO:0001661) |
| 0.0 | 0.0 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.0 | 0.1 | GO:0048625 | myoblast fate commitment(GO:0048625) |
| 0.0 | 0.1 | GO:0035521 | monoubiquitinated histone deubiquitination(GO:0035521) monoubiquitinated histone H2A deubiquitination(GO:0035522) |
| 0.0 | 0.5 | GO:0048665 | neuron fate specification(GO:0048665) |
| 0.0 | 0.0 | GO:0016479 | negative regulation of transcription from RNA polymerase I promoter(GO:0016479) |
| 0.0 | 0.7 | GO:0008045 | motor neuron axon guidance(GO:0008045) |
| 0.0 | 0.0 | GO:0044828 | negative regulation by host of viral genome replication(GO:0044828) |
| 0.0 | 0.2 | GO:0007171 | activation of transmembrane receptor protein tyrosine kinase activity(GO:0007171) |
| 0.0 | 0.1 | GO:0070236 | negative regulation of activation-induced cell death of T cells(GO:0070236) |
| 0.0 | 0.1 | GO:0042427 | serotonin biosynthetic process(GO:0042427) |
| 0.0 | 0.2 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade(GO:0070886) |
| 0.0 | 0.3 | GO:0015732 | prostaglandin transport(GO:0015732) |
| 0.0 | 0.7 | GO:0007274 | neuromuscular synaptic transmission(GO:0007274) |
| 0.0 | 0.1 | GO:1901165 | positive regulation of trophoblast cell migration(GO:1901165) |
| 0.0 | 0.0 | GO:0031508 | pericentric heterochromatin assembly(GO:0031508) |
| 0.0 | 0.2 | GO:0070307 | lens fiber cell development(GO:0070307) |
| 0.0 | 0.2 | GO:0060088 | auditory receptor cell stereocilium organization(GO:0060088) |
| 0.0 | 0.4 | GO:0043457 | regulation of cellular respiration(GO:0043457) |
| 0.0 | 0.2 | GO:0007196 | adenylate cyclase-inhibiting G-protein coupled glutamate receptor signaling pathway(GO:0007196) |
| 0.0 | 0.1 | GO:0006432 | phenylalanyl-tRNA aminoacylation(GO:0006432) |
| 0.0 | 0.0 | GO:0072434 | signal transduction involved in G2 DNA damage checkpoint(GO:0072425) signal transduction involved in mitotic G2 DNA damage checkpoint(GO:0072434) |
| 0.0 | 0.2 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) negative regulation of ATPase activity(GO:0032780) |
| 0.0 | 0.1 | GO:0070050 | neuron cellular homeostasis(GO:0070050) |
| 0.0 | 0.7 | GO:0010165 | response to X-ray(GO:0010165) |
| 0.0 | 0.1 | GO:0010482 | epidermal cell division(GO:0010481) regulation of epidermal cell division(GO:0010482) |
| 0.0 | 0.1 | GO:0008612 | peptidyl-lysine modification to peptidyl-hypusine(GO:0008612) |
| 0.0 | 0.1 | GO:0038042 | positive regulation of protein kinase A signaling(GO:0010739) dimeric G-protein coupled receptor signaling pathway(GO:0038042) calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.0 | 0.4 | GO:0035455 | response to interferon-alpha(GO:0035455) |
| 0.0 | 0.1 | GO:0048003 | antigen processing and presentation of lipid antigen via MHC class Ib(GO:0048003) antigen processing and presentation, exogenous lipid antigen via MHC class Ib(GO:0048007) |
| 0.0 | 0.2 | GO:0006600 | creatine metabolic process(GO:0006600) |
| 0.0 | 0.9 | GO:0046847 | filopodium assembly(GO:0046847) |
| 0.0 | 0.1 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.0 | 0.2 | GO:0045116 | protein neddylation(GO:0045116) |
| 0.0 | 0.2 | GO:0019371 | cyclooxygenase pathway(GO:0019371) |
| 0.0 | 0.1 | GO:0032364 | oxygen homeostasis(GO:0032364) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.8 | GO:0035692 | macrophage migration inhibitory factor receptor complex(GO:0035692) |
| 0.1 | 1.2 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 0.1 | 0.5 | GO:0031262 | Ndc80 complex(GO:0031262) |
| 0.1 | 1.5 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.1 | 2.9 | GO:0030673 | axolemma(GO:0030673) |
| 0.1 | 0.4 | GO:0031415 | NatA complex(GO:0031415) |
| 0.0 | 0.2 | GO:0001405 | presequence translocase-associated import motor(GO:0001405) |
| 0.0 | 0.3 | GO:0032777 | Piccolo NuA4 histone acetyltransferase complex(GO:0032777) |
| 0.0 | 0.2 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 0.0 | 2.6 | GO:0015030 | Cajal body(GO:0015030) |
| 0.0 | 0.1 | GO:0098592 | cytoplasmic side of apical plasma membrane(GO:0098592) |
| 0.0 | 0.1 | GO:0097454 | Schwann cell microvillus(GO:0097454) |
| 0.0 | 0.2 | GO:0035985 | senescence-associated heterochromatin focus(GO:0035985) |
| 0.0 | 0.1 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.0 | 0.3 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.0 | 0.1 | GO:0030934 | anchoring collagen complex(GO:0030934) |
| 0.0 | 0.2 | GO:0035867 | alphav-beta3 integrin-IGF-1-IGF1R complex(GO:0035867) |
| 0.0 | 2.6 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 0.2 | GO:0060203 | clathrin-sculpted glutamate transport vesicle(GO:0060199) clathrin-sculpted glutamate transport vesicle membrane(GO:0060203) |
| 0.0 | 0.2 | GO:0071546 | pi-body(GO:0071546) |
| 0.0 | 0.1 | GO:0035976 | AP1 complex(GO:0035976) |
| 0.0 | 0.4 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) |
| 0.0 | 0.2 | GO:0097197 | tetraspanin-enriched microdomain(GO:0097197) |
| 0.0 | 0.1 | GO:0009328 | phenylalanine-tRNA ligase complex(GO:0009328) |
| 0.0 | 0.2 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 0.0 | 0.1 | GO:1990357 | terminal web(GO:1990357) |
| 0.0 | 0.4 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.0 | 0.1 | GO:0035517 | PR-DUB complex(GO:0035517) |
| 0.0 | 0.0 | GO:0070195 | growth hormone receptor complex(GO:0070195) |
| 0.0 | 0.2 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.0 | 0.5 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 0.0 | 0.2 | GO:0098839 | postsynaptic density membrane(GO:0098839) |
| 0.0 | 0.1 | GO:1903440 | calcitonin family receptor complex(GO:1903439) amylin receptor complex(GO:1903440) |
| 0.0 | 1.7 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.0 | 2.6 | GO:0055037 | recycling endosome(GO:0055037) |
| 0.0 | 0.2 | GO:0005883 | neurofilament(GO:0005883) |
| 0.0 | 0.2 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.0 | 0.2 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.0 | 0.2 | GO:0031088 | platelet dense granule membrane(GO:0031088) |
| 0.0 | 0.1 | GO:0070557 | PCNA-p21 complex(GO:0070557) |
| 0.0 | 0.1 | GO:0070369 | beta-catenin-TCF7L2 complex(GO:0070369) |
| 0.0 | 0.5 | GO:0044665 | MLL1/2 complex(GO:0044665) MLL1 complex(GO:0071339) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 2.6 | GO:0008859 | exoribonuclease II activity(GO:0008859) |
| 0.4 | 1.7 | GO:0034041 | sterol-transporting ATPase activity(GO:0034041) |
| 0.2 | 2.9 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 0.1 | 1.4 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.1 | 0.7 | GO:0015526 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 0.1 | 1.3 | GO:0004118 | cGMP-stimulated cyclic-nucleotide phosphodiesterase activity(GO:0004118) TPR domain binding(GO:0030911) |
| 0.1 | 1.5 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.1 | 1.4 | GO:0004028 | 3-chloroallyl aldehyde dehydrogenase activity(GO:0004028) |
| 0.1 | 0.7 | GO:0038049 | transcription factor activity, ligand-activated RNA polymerase II transcription factor binding(GO:0038049) |
| 0.1 | 0.5 | GO:0008481 | sphinganine kinase activity(GO:0008481) D-erythro-sphingosine kinase activity(GO:0017050) |
| 0.1 | 0.5 | GO:0008889 | glycerophosphodiester phosphodiesterase activity(GO:0008889) |
| 0.1 | 0.3 | GO:0047888 | fatty acid peroxidase activity(GO:0047888) |
| 0.1 | 0.3 | GO:0001632 | leukotriene B4 receptor activity(GO:0001632) |
| 0.1 | 0.3 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) |
| 0.1 | 0.3 | GO:0051748 | UTP:glucose-1-phosphate uridylyltransferase activity(GO:0003983) UTP-monosaccharide-1-phosphate uridylyltransferase activity(GO:0051748) |
| 0.1 | 0.2 | GO:0032422 | purine-rich negative regulatory element binding(GO:0032422) |
| 0.1 | 0.2 | GO:0016534 | cyclin-dependent protein kinase 5 activator activity(GO:0016534) |
| 0.1 | 0.6 | GO:0015321 | sodium-dependent phosphate transmembrane transporter activity(GO:0015321) |
| 0.1 | 2.0 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.0 | 0.1 | GO:0015389 | pyrimidine- and adenine-specific:sodium symporter activity(GO:0015389) |
| 0.0 | 0.4 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.0 | 0.6 | GO:0016494 | C-X-C chemokine receptor activity(GO:0016494) |
| 0.0 | 0.3 | GO:0004522 | ribonuclease A activity(GO:0004522) |
| 0.0 | 1.3 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.0 | 0.2 | GO:0017018 | myosin phosphatase activity(GO:0017018) |
| 0.0 | 0.2 | GO:0005136 | interleukin-4 receptor binding(GO:0005136) |
| 0.0 | 0.1 | GO:0033878 | hormone-sensitive lipase activity(GO:0033878) |
| 0.0 | 0.1 | GO:0004937 | alpha1-adrenergic receptor activity(GO:0004937) |
| 0.0 | 0.8 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.0 | 0.6 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 0.3 | GO:0033743 | peptide-methionine (R)-S-oxide reductase activity(GO:0033743) |
| 0.0 | 0.2 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) |
| 0.0 | 0.4 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.0 | 0.5 | GO:0016004 | phospholipase activator activity(GO:0016004) |
| 0.0 | 0.1 | GO:0015375 | glycine:sodium symporter activity(GO:0015375) |
| 0.0 | 0.3 | GO:0032190 | acrosin binding(GO:0032190) |
| 0.0 | 0.2 | GO:1904929 | coreceptor activity involved in Wnt signaling pathway, planar cell polarity pathway(GO:1904929) |
| 0.0 | 0.3 | GO:0042609 | CD4 receptor binding(GO:0042609) |
| 0.0 | 0.2 | GO:0001640 | adenylate cyclase inhibiting G-protein coupled glutamate receptor activity(GO:0001640) G-protein coupled glutamate receptor activity(GO:0098988) |
| 0.0 | 0.1 | GO:0004948 | calcitonin receptor activity(GO:0004948) |
| 0.0 | 0.6 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.0 | 0.2 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.0 | 0.2 | GO:0031811 | G-protein coupled nucleotide receptor binding(GO:0031811) P2Y1 nucleotide receptor binding(GO:0031812) |
| 0.0 | 0.8 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.0 | 0.4 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 0.0 | 0.2 | GO:0042500 | aspartic endopeptidase activity, intramembrane cleaving(GO:0042500) |
| 0.0 | 0.1 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.0 | 0.2 | GO:0099580 | ion antiporter activity involved in regulation of postsynaptic membrane potential(GO:0099580) |
| 0.0 | 0.2 | GO:0034711 | inhibin binding(GO:0034711) |
| 0.0 | 0.1 | GO:0031751 | D4 dopamine receptor binding(GO:0031751) |
| 0.0 | 2.5 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.2 | GO:0086006 | voltage-gated sodium channel activity involved in cardiac muscle cell action potential(GO:0086006) |
| 0.0 | 0.2 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.0 | 0.1 | GO:0008401 | retinoic acid 4-hydroxylase activity(GO:0008401) |
| 0.0 | 0.1 | GO:0005220 | inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0005220) |
| 0.0 | 0.1 | GO:0008426 | protein kinase C inhibitor activity(GO:0008426) |
| 0.0 | 0.3 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.0 | 0.4 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.0 | 0.1 | GO:0035373 | chondroitin sulfate proteoglycan binding(GO:0035373) |
| 0.0 | 0.2 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.0 | 0.1 | GO:0036134 | thromboxane-A synthase activity(GO:0004796) 12-hydroxyheptadecatrienoic acid synthase activity(GO:0036134) |
| 0.0 | 0.2 | GO:0017169 | CDP-alcohol phosphatidyltransferase activity(GO:0017169) |
| 0.0 | 0.2 | GO:0004931 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.0 | 2.0 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.0 | 0.2 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.0 | 0.1 | GO:0004013 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.0 | 0.1 | GO:0016749 | 5-aminolevulinate synthase activity(GO:0003870) N-succinyltransferase activity(GO:0016749) |
| 0.0 | 0.2 | GO:0008517 | folic acid transporter activity(GO:0008517) |
| 0.0 | 0.1 | GO:0008142 | oxysterol binding(GO:0008142) |
| 0.0 | 0.3 | GO:0015245 | fatty acid transporter activity(GO:0015245) |
| 0.0 | 0.1 | GO:0016404 | 15-hydroxyprostaglandin dehydrogenase (NAD+) activity(GO:0016404) |
| 0.0 | 0.1 | GO:0060002 | plus-end directed microfilament motor activity(GO:0060002) |
| 0.0 | 0.0 | GO:0030550 | acetylcholine receptor inhibitor activity(GO:0030550) |
| 0.0 | 0.1 | GO:0043426 | MRF binding(GO:0043426) |
| 0.0 | 0.4 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.0 | 0.4 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.0 | 0.2 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 0.0 | 0.1 | GO:0099583 | neurotransmitter receptor activity involved in regulation of postsynaptic cytosolic calcium ion concentration(GO:0099583) |
| 0.0 | 0.1 | GO:0030883 | lipid antigen binding(GO:0030882) endogenous lipid antigen binding(GO:0030883) exogenous lipid antigen binding(GO:0030884) |
| 0.0 | 0.2 | GO:0016594 | glycine binding(GO:0016594) |
| 0.0 | 0.2 | GO:0005161 | platelet-derived growth factor receptor binding(GO:0005161) |
| 0.0 | 0.1 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
| 0.0 | 0.1 | GO:0004826 | phenylalanine-tRNA ligase activity(GO:0004826) |
| 0.0 | 0.1 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.0 | 0.0 | GO:0061609 | fructose-1-phosphate aldolase activity(GO:0061609) |
| 0.0 | 0.5 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.0 | 0.0 | GO:0001156 | TFIIIC-class transcription factor binding(GO:0001156) |
| 0.0 | 0.0 | GO:0005148 | prolactin receptor binding(GO:0005148) |
| 0.0 | 0.1 | GO:0019912 | cyclin-dependent protein kinase activating kinase activity(GO:0019912) |
| 0.0 | 0.1 | GO:0031014 | troponin T binding(GO:0031014) |
| 0.0 | 0.1 | GO:0005280 | hydrogen:amino acid symporter activity(GO:0005280) |
| 0.0 | 0.1 | GO:0046920 | alpha-(1->3)-fucosyltransferase activity(GO:0046920) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.9 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 1.8 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.0 | 0.8 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.0 | 1.0 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.0 | 2.0 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.0 | 0.3 | PID TNF PATHWAY | TNF receptor signaling pathway |
| 0.0 | 0.8 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.0 | 1.2 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.0 | 0.3 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.0 | 0.3 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.0 | 1.2 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.0 | 0.3 | PID S1P S1P1 PATHWAY | S1P1 pathway |
| 0.0 | 0.8 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.0 | 0.5 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.0 | 0.1 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.0 | 2.6 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 0.6 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.9 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.1 | 1.8 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.1 | 1.5 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.0 | 1.0 | REACTOME NFKB ACTIVATION THROUGH FADD RIP1 PATHWAY MEDIATED BY CASPASE 8 AND10 | Genes involved in NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 |
| 0.0 | 1.3 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.0 | 0.7 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.0 | 1.3 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.0 | 0.3 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.0 | 0.2 | REACTOME APOBEC3G MEDIATED RESISTANCE TO HIV1 INFECTION | Genes involved in APOBEC3G mediated resistance to HIV-1 infection |
| 0.0 | 0.2 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.0 | 0.3 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.0 | 0.7 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.0 | 0.2 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.0 | 0.7 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.4 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.0 | 0.3 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.0 | 0.2 | REACTOME GLYCOPROTEIN HORMONES | Genes involved in Glycoprotein hormones |
| 0.0 | 0.3 | REACTOME SYNTHESIS OF PIPS AT THE GOLGI MEMBRANE | Genes involved in Synthesis of PIPs at the Golgi membrane |
| 0.0 | 0.5 | REACTOME G BETA GAMMA SIGNALLING THROUGH PI3KGAMMA | Genes involved in G beta:gamma signalling through PI3Kgamma |
| 0.0 | 1.6 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 0.2 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.0 | 0.3 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.0 | 0.1 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.0 | 0.3 | REACTOME EICOSANOID LIGAND BINDING RECEPTORS | Genes involved in Eicosanoid ligand-binding receptors |
| 0.0 | 0.3 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.0 | 0.3 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.0 | 0.5 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |