Project
Inflammatory response time course, HUVEC (Wada, 2009)
Navigation
Downloads
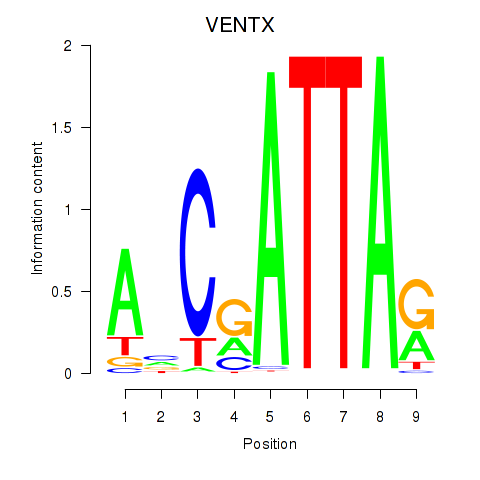
Results for VENTX
Z-value: 0.55
Transcription factors associated with VENTX
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
VENTX
|
ENSG00000151650.8 | VENTX |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| VENTX | hg38_v1_chr10_+_133237849_133237863 | 0.02 | 9.1e-01 | Click! |
Activity profile of VENTX motif
Sorted Z-values of VENTX motif
Network of associatons between targets according to the STRING database.
First level regulatory network of VENTX
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_+_85604146 | 1.81 |
ENST00000512201.5
|
ARHGAP24
|
Rho GTPase activating protein 24 |
| chr10_-_48604952 | 1.21 |
ENST00000417912.6
|
ARHGAP22
|
Rho GTPase activating protein 22 |
| chr10_-_48605032 | 1.13 |
ENST00000249601.9
|
ARHGAP22
|
Rho GTPase activating protein 22 |
| chr6_+_113857333 | 0.83 |
ENST00000612661.2
|
MARCKS
|
myristoylated alanine rich protein kinase C substrate |
| chr10_-_13707536 | 0.66 |
ENST00000632570.1
ENST00000477221.2 |
FRMD4A
|
FERM domain containing 4A |
| chr8_+_49911801 | 0.64 |
ENST00000643809.1
|
SNTG1
|
syntrophin gamma 1 |
| chr12_-_2004421 | 0.64 |
ENST00000280665.11
ENST00000535873.2 |
DCP1B
|
decapping mRNA 1B |
| chr7_-_108240049 | 0.62 |
ENST00000379022.8
|
NRCAM
|
neuronal cell adhesion molecule |
| chr5_-_124745315 | 0.57 |
ENST00000306315.9
|
ZNF608
|
zinc finger protein 608 |
| chr10_-_25062279 | 0.57 |
ENST00000615958.4
|
ENKUR
|
enkurin, TRPC channel interacting protein |
| chr4_+_168092530 | 0.57 |
ENST00000359299.8
|
ANXA10
|
annexin A10 |
| chr13_+_108629605 | 0.56 |
ENST00000457511.7
|
MYO16
|
myosin XVI |
| chr18_+_61333424 | 0.53 |
ENST00000262717.9
|
CDH20
|
cadherin 20 |
| chr1_+_244352627 | 0.46 |
ENST00000366537.5
ENST00000308105.5 |
C1orf100
|
chromosome 1 open reading frame 100 |
| chr14_+_32329256 | 0.45 |
ENST00000280979.9
|
AKAP6
|
A-kinase anchoring protein 6 |
| chr8_+_49911604 | 0.44 |
ENST00000642164.1
ENST00000644093.1 ENST00000643999.1 ENST00000647073.1 ENST00000646880.1 |
SNTG1
|
syntrophin gamma 1 |
| chr15_+_33968484 | 0.44 |
ENST00000383263.7
|
CHRM5
|
cholinergic receptor muscarinic 5 |
| chr8_+_31639291 | 0.44 |
ENST00000651149.1
ENST00000650866.1 |
NRG1
|
neuregulin 1 |
| chr8_+_31639222 | 0.43 |
ENST00000519301.6
ENST00000652698.1 |
NRG1
|
neuregulin 1 |
| chr11_-_3837858 | 0.41 |
ENST00000396979.1
|
RHOG
|
ras homolog family member G |
| chr1_+_174447944 | 0.39 |
ENST00000367685.5
|
GPR52
|
G protein-coupled receptor 52 |
| chr3_+_32817990 | 0.39 |
ENST00000383763.6
|
TRIM71
|
tripartite motif containing 71 |
| chr18_+_58341038 | 0.37 |
ENST00000679791.1
|
NEDD4L
|
NEDD4 like E3 ubiquitin protein ligase |
| chr2_-_162152404 | 0.37 |
ENST00000375497.3
|
GCG
|
glucagon |
| chr4_-_76007501 | 0.36 |
ENST00000264888.6
|
CXCL9
|
C-X-C motif chemokine ligand 9 |
| chr3_-_57199938 | 0.35 |
ENST00000473921.2
ENST00000295934.8 |
HESX1
|
HESX homeobox 1 |
| chr12_-_52434363 | 0.35 |
ENST00000252245.6
|
KRT75
|
keratin 75 |
| chr3_+_130850585 | 0.33 |
ENST00000505330.5
ENST00000504381.5 ENST00000507488.6 |
ATP2C1
|
ATPase secretory pathway Ca2+ transporting 1 |
| chr18_+_58221535 | 0.32 |
ENST00000431212.6
ENST00000586268.5 ENST00000587190.5 |
NEDD4L
|
NEDD4 like E3 ubiquitin protein ligase |
| chr1_-_94925759 | 0.32 |
ENST00000415017.1
ENST00000545882.5 |
CNN3
|
calponin 3 |
| chr1_+_40247926 | 0.31 |
ENST00000372766.4
|
TMCO2
|
transmembrane and coiled-coil domains 2 |
| chr8_+_49911396 | 0.31 |
ENST00000642720.2
|
SNTG1
|
syntrophin gamma 1 |
| chr18_-_26657401 | 0.31 |
ENST00000580191.5
|
KCTD1
|
potassium channel tetramerization domain containing 1 |
| chr22_+_38705922 | 0.30 |
ENST00000216044.10
|
GTPBP1
|
GTP binding protein 1 |
| chr7_+_18495723 | 0.28 |
ENST00000681950.1
ENST00000622668.4 ENST00000405010.7 ENST00000406451.8 ENST00000441542.7 ENST00000428307.6 ENST00000681273.1 |
HDAC9
|
histone deacetylase 9 |
| chr12_+_111034136 | 0.28 |
ENST00000261726.11
|
CUX2
|
cut like homeobox 2 |
| chr12_-_52618559 | 0.27 |
ENST00000305748.7
|
KRT73
|
keratin 73 |
| chr12_-_86256267 | 0.26 |
ENST00000620241.4
|
MGAT4C
|
MGAT4 family member C |
| chr8_-_86743626 | 0.26 |
ENST00000320005.6
|
CNGB3
|
cyclic nucleotide gated channel subunit beta 3 |
| chr14_-_26597430 | 0.26 |
ENST00000344429.9
ENST00000574031.1 ENST00000465357.6 ENST00000547619.5 |
NOVA1
|
NOVA alternative splicing regulator 1 |
| chr11_-_16356538 | 0.26 |
ENST00000683767.1
|
SOX6
|
SRY-box transcription factor 6 |
| chr4_+_70592295 | 0.26 |
ENST00000449493.2
|
AMBN
|
ameloblastin |
| chr10_+_72692125 | 0.26 |
ENST00000373053.7
ENST00000357157.10 |
MCU
|
mitochondrial calcium uniporter |
| chr3_+_171843337 | 0.25 |
ENST00000334567.9
ENST00000619900.4 ENST00000450693.1 |
TMEM212
|
transmembrane protein 212 |
| chr7_+_120988683 | 0.25 |
ENST00000340646.9
ENST00000310396.10 |
CPED1
|
cadherin like and PC-esterase domain containing 1 |
| chr16_-_74421756 | 0.24 |
ENST00000617101.4
|
CLEC18B
|
C-type lectin domain family 18 member B |
| chr11_-_36598221 | 0.23 |
ENST00000311485.8
ENST00000527033.5 ENST00000532616.1 ENST00000618712.4 |
RAG2
|
recombination activating 2 |
| chr16_-_74421392 | 0.23 |
ENST00000339953.9
ENST00000620745.1 ENST00000682950.1 |
CLEC18B
|
C-type lectin domain family 18 member B |
| chr14_+_32329341 | 0.23 |
ENST00000557354.5
ENST00000557102.1 ENST00000557272.1 |
AKAP6
|
A-kinase anchoring protein 6 |
| chr11_+_75815180 | 0.22 |
ENST00000356136.8
|
UVRAG
|
UV radiation resistance associated |
| chr19_-_49119092 | 0.21 |
ENST00000408991.4
|
C19orf73
|
chromosome 19 open reading frame 73 |
| chr19_-_45584769 | 0.21 |
ENST00000263275.5
|
OPA3
|
outer mitochondrial membrane lipid metabolism regulator OPA3 |
| chrX_+_78747705 | 0.21 |
ENST00000614823.5
ENST00000435339.3 ENST00000514744.5 |
LPAR4
|
lysophosphatidic acid receptor 4 |
| chr2_-_207167220 | 0.20 |
ENST00000421199.5
ENST00000457962.5 |
KLF7
|
Kruppel like factor 7 |
| chr6_+_12290353 | 0.20 |
ENST00000379375.6
|
EDN1
|
endothelin 1 |
| chr2_-_164842011 | 0.20 |
ENST00000409184.8
ENST00000456693.5 |
COBLL1
|
cordon-bleu WH2 repeat protein like 1 |
| chr12_+_25052732 | 0.20 |
ENST00000547044.5
|
IRAG2
|
inositol 1,4,5-triphosphate receptor associated 2 |
| chr14_+_64704380 | 0.19 |
ENST00000247226.13
ENST00000394691.7 |
PLEKHG3
|
pleckstrin homology and RhoGEF domain containing G3 |
| chr1_+_40709475 | 0.19 |
ENST00000372651.5
|
NFYC
|
nuclear transcription factor Y subunit gamma |
| chr16_+_69950907 | 0.18 |
ENST00000393701.6
ENST00000568461.5 |
CLEC18A
|
C-type lectin domain family 18 member A |
| chr12_-_119803383 | 0.18 |
ENST00000392520.2
ENST00000678677.1 ENST00000679249.1 ENST00000676849.1 |
CIT
|
citron rho-interacting serine/threonine kinase |
| chr18_+_31591869 | 0.18 |
ENST00000237014.8
|
TTR
|
transthyretin |
| chr12_-_101210232 | 0.18 |
ENST00000536262.3
|
SLC5A8
|
solute carrier family 5 member 8 |
| chr3_-_157503574 | 0.18 |
ENST00000494677.5
ENST00000468233.5 |
VEPH1
|
ventricular zone expressed PH domain containing 1 |
| chr6_-_39725335 | 0.18 |
ENST00000538893.5
|
KIF6
|
kinesin family member 6 |
| chr6_-_65707214 | 0.17 |
ENST00000370621.7
ENST00000393380.6 ENST00000503581.6 |
EYS
|
eyes shut homolog |
| chr19_-_45584810 | 0.17 |
ENST00000323060.3
|
OPA3
|
outer mitochondrial membrane lipid metabolism regulator OPA3 |
| chr2_+_44941695 | 0.17 |
ENST00000260653.5
|
SIX3
|
SIX homeobox 3 |
| chr11_+_13962676 | 0.17 |
ENST00000576479.4
|
SPON1
|
spondin 1 |
| chr4_-_103077282 | 0.17 |
ENST00000503230.5
ENST00000503818.1 |
SLC9B2
|
solute carrier family 9 member B2 |
| chr12_+_80716906 | 0.16 |
ENST00000228644.4
|
MYF5
|
myogenic factor 5 |
| chr10_+_132537814 | 0.16 |
ENST00000368593.7
|
INPP5A
|
inositol polyphosphate-5-phosphatase A |
| chr5_+_141484997 | 0.16 |
ENST00000617094.1
ENST00000306593.2 ENST00000610539.1 ENST00000618371.4 |
PCDHGC4
|
protocadherin gamma subfamily C, 4 |
| chr16_+_70173783 | 0.16 |
ENST00000541793.7
ENST00000314151.12 ENST00000565806.5 ENST00000569347.6 ENST00000536907.2 |
CLEC18C
|
C-type lectin domain family 18 member C |
| chrX_-_140505058 | 0.16 |
ENST00000370536.5
|
SOX3
|
SRY-box transcription factor 3 |
| chr14_+_64715677 | 0.16 |
ENST00000634379.2
|
PLEKHG3
|
pleckstrin homology and RhoGEF domain containing G3 |
| chr10_+_112283399 | 0.16 |
ENST00000643850.1
ENST00000646139.2 ENST00000645243.1 |
TECTB
|
tectorin beta |
| chr8_+_106726115 | 0.16 |
ENST00000521592.5
|
OXR1
|
oxidation resistance 1 |
| chr5_-_157059109 | 0.15 |
ENST00000523175.6
ENST00000522693.5 |
HAVCR1
|
hepatitis A virus cellular receptor 1 |
| chr7_+_130344810 | 0.15 |
ENST00000497503.5
ENST00000463587.5 ENST00000461828.5 ENST00000474905.6 ENST00000494311.1 ENST00000466363.6 |
CPA5
|
carboxypeptidase A5 |
| chr11_-_11353241 | 0.15 |
ENST00000528848.3
|
CSNK2A3
|
casein kinase 2 alpha 3 |
| chr12_+_69825221 | 0.15 |
ENST00000552032.7
|
MYRFL
|
myelin regulatory factor like |
| chr12_+_69825273 | 0.15 |
ENST00000547771.6
|
MYRFL
|
myelin regulatory factor like |
| chrM_+_9207 | 0.14 |
ENST00000362079.2
|
MT-CO3
|
mitochondrially encoded cytochrome c oxidase III |
| chr2_-_164842140 | 0.14 |
ENST00000496396.1
ENST00000629362.2 ENST00000445474.2 ENST00000483743.6 |
COBLL1
|
cordon-bleu WH2 repeat protein like 1 |
| chr10_-_60572599 | 0.14 |
ENST00000503366.5
|
ANK3
|
ankyrin 3 |
| chr1_-_114153863 | 0.14 |
ENST00000610222.3
ENST00000369547.6 ENST00000641643.2 |
SYT6
|
synaptotagmin 6 |
| chr12_-_119804298 | 0.14 |
ENST00000678652.1
ENST00000678494.1 |
CIT
|
citron rho-interacting serine/threonine kinase |
| chr16_+_69951180 | 0.14 |
ENST00000288040.10
ENST00000449317.6 |
CLEC18A
|
C-type lectin domain family 18 member A |
| chr4_-_83114715 | 0.14 |
ENST00000426923.2
ENST00000311507.9 ENST00000509973.5 |
PLAC8
|
placenta associated 8 |
| chr1_-_178871060 | 0.14 |
ENST00000234816.7
|
ANGPTL1
|
angiopoietin like 1 |
| chr2_+_202073282 | 0.14 |
ENST00000459709.5
|
KIAA2012
|
KIAA2012 |
| chr22_-_40533808 | 0.14 |
ENST00000422851.1
ENST00000651694.1 ENST00000652095.2 |
MRTFA
|
myocardin related transcription factor A |
| chr8_+_106726012 | 0.13 |
ENST00000449762.6
ENST00000297447.10 |
OXR1
|
oxidation resistance 1 |
| chr7_+_130344837 | 0.13 |
ENST00000485477.5
ENST00000431780.6 |
CPA5
|
carboxypeptidase A5 |
| chr2_+_202073249 | 0.13 |
ENST00000498697.3
|
KIAA2012
|
KIAA2012 |
| chr4_-_26490453 | 0.12 |
ENST00000295589.4
|
CCKAR
|
cholecystokinin A receptor |
| chr17_-_41065879 | 0.12 |
ENST00000394015.3
|
KRTAP2-4
|
keratin associated protein 2-4 |
| chr17_-_48968587 | 0.12 |
ENST00000357424.2
|
GIP
|
gastric inhibitory polypeptide |
| chr12_-_51217671 | 0.12 |
ENST00000333640.11
ENST00000550824.6 |
POU6F1
|
POU class 6 homeobox 1 |
| chr4_-_154590735 | 0.12 |
ENST00000403106.8
ENST00000622532.1 ENST00000651975.1 |
FGA
|
fibrinogen alpha chain |
| chr12_+_56080126 | 0.12 |
ENST00000411731.6
|
ERBB3
|
erb-b2 receptor tyrosine kinase 3 |
| chr3_-_9792404 | 0.11 |
ENST00000301964.7
|
TADA3
|
transcriptional adaptor 3 |
| chr6_+_167291329 | 0.10 |
ENST00000366829.2
|
UNC93A
|
unc-93 homolog A |
| chr2_+_176157293 | 0.10 |
ENST00000683222.1
|
HOXD3
|
homeobox D3 |
| chr7_-_78771265 | 0.10 |
ENST00000630991.2
ENST00000629359.2 |
MAGI2
|
membrane associated guanylate kinase, WW and PDZ domain containing 2 |
| chr7_+_54542300 | 0.10 |
ENST00000302287.7
ENST00000407838.7 |
VSTM2A
|
V-set and transmembrane domain containing 2A |
| chr6_-_138499487 | 0.10 |
ENST00000343505.9
|
NHSL1
|
NHS like 1 |
| chr4_+_168497044 | 0.10 |
ENST00000505667.6
|
PALLD
|
palladin, cytoskeletal associated protein |
| chr6_-_161274010 | 0.10 |
ENST00000366911.9
ENST00000366905.3 |
AGPAT4
|
1-acylglycerol-3-phosphate O-acyltransferase 4 |
| chr6_+_167291309 | 0.10 |
ENST00000230256.8
|
UNC93A
|
unc-93 homolog A |
| chr3_-_57226344 | 0.10 |
ENST00000495160.2
|
HESX1
|
HESX homeobox 1 |
| chr1_-_178871022 | 0.10 |
ENST00000367629.1
|
ANGPTL1
|
angiopoietin like 1 |
| chr8_-_42501224 | 0.10 |
ENST00000520262.6
ENST00000517366.1 |
SLC20A2
|
solute carrier family 20 member 2 |
| chr1_+_61952283 | 0.10 |
ENST00000307297.8
|
PATJ
|
PATJ crumbs cell polarity complex component |
| chr14_+_64986846 | 0.09 |
ENST00000246166.3
|
FNTB
|
farnesyltransferase, CAAX box, beta |
| chr6_-_161274042 | 0.09 |
ENST00000320285.9
|
AGPAT4
|
1-acylglycerol-3-phosphate O-acyltransferase 4 |
| chr7_-_78771108 | 0.09 |
ENST00000626691.2
|
MAGI2
|
membrane associated guanylate kinase, WW and PDZ domain containing 2 |
| chr7_+_54542362 | 0.09 |
ENST00000402613.4
|
VSTM2A
|
V-set and transmembrane domain containing 2A |
| chr2_-_165203870 | 0.08 |
ENST00000639244.1
ENST00000409101.7 ENST00000668657.1 |
SCN3A
|
sodium voltage-gated channel alpha subunit 3 |
| chr17_+_17782108 | 0.08 |
ENST00000395774.1
|
RAI1
|
retinoic acid induced 1 |
| chr1_+_61952036 | 0.08 |
ENST00000646453.1
ENST00000635137.1 |
PATJ
|
PATJ crumbs cell polarity complex component |
| chr4_+_168497066 | 0.07 |
ENST00000261509.10
|
PALLD
|
palladin, cytoskeletal associated protein |
| chr4_+_70592253 | 0.07 |
ENST00000322937.10
ENST00000613447.4 |
AMBN
|
ameloblastin |
| chr17_-_10469558 | 0.07 |
ENST00000255381.2
|
MYH4
|
myosin heavy chain 4 |
| chr17_+_45161864 | 0.07 |
ENST00000589230.6
ENST00000591576.5 ENST00000591070.6 ENST00000592695.1 |
HEXIM2
|
HEXIM P-TEFb complex subunit 2 |
| chr2_+_168901290 | 0.07 |
ENST00000429379.2
ENST00000375363.8 ENST00000421979.1 |
G6PC2
|
glucose-6-phosphatase catalytic subunit 2 |
| chr1_+_240123121 | 0.07 |
ENST00000681210.1
|
FMN2
|
formin 2 |
| chrX_+_106693838 | 0.07 |
ENST00000324342.7
|
RNF128
|
ring finger protein 128 |
| chr13_-_103066411 | 0.07 |
ENST00000245312.5
|
SLC10A2
|
solute carrier family 10 member 2 |
| chr6_-_49713521 | 0.06 |
ENST00000339139.5
|
CRISP2
|
cysteine rich secretory protein 2 |
| chr6_-_49713564 | 0.06 |
ENST00000616725.4
ENST00000618917.4 |
CRISP2
|
cysteine rich secretory protein 2 |
| chrX_+_136648138 | 0.06 |
ENST00000370629.7
|
CD40LG
|
CD40 ligand |
| chr12_+_55549602 | 0.06 |
ENST00000641569.1
ENST00000641851.1 |
OR6C4
|
olfactory receptor family 6 subfamily C member 4 |
| chr1_+_240123148 | 0.06 |
ENST00000681824.1
|
FMN2
|
formin 2 |
| chr7_-_28180735 | 0.06 |
ENST00000283928.10
|
JAZF1
|
JAZF zinc finger 1 |
| chr20_-_52105644 | 0.06 |
ENST00000371523.8
|
ZFP64
|
ZFP64 zinc finger protein |
| chr14_+_21841182 | 0.05 |
ENST00000390433.1
|
TRAV12-1
|
T cell receptor alpha variable 12-1 |
| chr14_-_81533800 | 0.05 |
ENST00000555824.5
ENST00000557372.1 ENST00000336735.9 |
SEL1L
|
SEL1L adaptor subunit of ERAD E3 ubiquitin ligase |
| chr18_+_78979811 | 0.05 |
ENST00000537592.7
|
SALL3
|
spalt like transcription factor 3 |
| chr10_-_54801179 | 0.05 |
ENST00000373955.5
|
PCDH15
|
protocadherin related 15 |
| chr2_-_223602284 | 0.05 |
ENST00000421386.1
ENST00000305409.3 ENST00000433889.1 |
SCG2
|
secretogranin II |
| chr6_-_132659178 | 0.05 |
ENST00000275216.3
|
TAAR1
|
trace amine associated receptor 1 |
| chr11_-_107858777 | 0.05 |
ENST00000525815.6
|
SLC35F2
|
solute carrier family 35 member F2 |
| chr3_+_9792495 | 0.04 |
ENST00000498623.6
|
ARPC4
|
actin related protein 2/3 complex subunit 4 |
| chrX_+_106920393 | 0.04 |
ENST00000336803.2
|
CLDN2
|
claudin 2 |
| chr4_+_112860981 | 0.04 |
ENST00000671704.1
|
ANK2
|
ankyrin 2 |
| chr14_+_92323154 | 0.04 |
ENST00000532405.6
ENST00000676001.1 ENST00000531433.5 |
SLC24A4
|
solute carrier family 24 member 4 |
| chr4_+_112861053 | 0.04 |
ENST00000672221.1
|
ANK2
|
ankyrin 2 |
| chr16_-_51151259 | 0.04 |
ENST00000251020.9
|
SALL1
|
spalt like transcription factor 1 |
| chr22_-_30574572 | 0.04 |
ENST00000402369.5
|
GAL3ST1
|
galactose-3-O-sulfotransferase 1 |
| chr5_-_143400716 | 0.04 |
ENST00000424646.6
ENST00000652686.1 |
NR3C1
|
nuclear receptor subfamily 3 group C member 1 |
| chr17_-_50707855 | 0.04 |
ENST00000285243.7
|
ANKRD40
|
ankyrin repeat domain 40 |
| chr11_+_46701010 | 0.04 |
ENST00000311764.3
|
ZNF408
|
zinc finger protein 408 |
| chr1_+_186296267 | 0.04 |
ENST00000533951.5
ENST00000367482.8 ENST00000635041.1 ENST00000367483.8 ENST00000367485.4 ENST00000445192.7 |
PRG4
|
proteoglycan 4 |
| chr4_+_112860912 | 0.04 |
ENST00000671951.1
|
ANK2
|
ankyrin 2 |
| chr11_-_124320197 | 0.04 |
ENST00000624618.2
|
OR8D2
|
olfactory receptor family 8 subfamily D member 2 |
| chr1_+_40709316 | 0.04 |
ENST00000372652.5
|
NFYC
|
nuclear transcription factor Y subunit gamma |
| chr10_+_24449426 | 0.04 |
ENST00000307544.10
|
KIAA1217
|
KIAA1217 |
| chr4_-_46124046 | 0.03 |
ENST00000295452.5
|
GABRG1
|
gamma-aminobutyric acid type A receptor subunit gamma1 |
| chr6_+_101398788 | 0.03 |
ENST00000369138.5
ENST00000413795.5 ENST00000358361.7 |
GRIK2
|
glutamate ionotropic receptor kainate type subunit 2 |
| chr4_+_85827891 | 0.03 |
ENST00000514229.5
|
ARHGAP24
|
Rho GTPase activating protein 24 |
| chr7_+_151048526 | 0.03 |
ENST00000349064.10
|
ASIC3
|
acid sensing ion channel subunit 3 |
| chr7_-_105269007 | 0.03 |
ENST00000357311.7
|
SRPK2
|
SRSF protein kinase 2 |
| chr7_+_148590760 | 0.02 |
ENST00000307003.3
|
C7orf33
|
chromosome 7 open reading frame 33 |
| chr16_+_14186707 | 0.02 |
ENST00000572567.5
|
MRTFB
|
myocardin related transcription factor B |
| chr9_-_23826231 | 0.02 |
ENST00000397312.7
|
ELAVL2
|
ELAV like RNA binding protein 2 |
| chr17_+_70104848 | 0.02 |
ENST00000392670.5
|
KCNJ16
|
potassium inwardly rectifying channel subfamily J member 16 |
| chrX_+_136648214 | 0.02 |
ENST00000370628.2
|
CD40LG
|
CD40 ligand |
| chr6_+_37929959 | 0.02 |
ENST00000373389.5
|
ZFAND3
|
zinc finger AN1-type containing 3 |
| chr7_+_54542393 | 0.02 |
ENST00000404951.5
|
VSTM2A
|
V-set and transmembrane domain containing 2A |
| chr5_-_131165272 | 0.01 |
ENST00000675491.1
ENST00000506908.2 |
HINT1
|
histidine triad nucleotide binding protein 1 |
| chrX_+_131058340 | 0.01 |
ENST00000276211.10
ENST00000370922.5 |
ARHGAP36
|
Rho GTPase activating protein 36 |
| chr17_-_29005913 | 0.01 |
ENST00000442608.7
ENST00000317338.17 ENST00000335960.10 |
SEZ6
|
seizure related 6 homolog |
| chr12_+_8513499 | 0.01 |
ENST00000299665.3
|
CLEC4D
|
C-type lectin domain family 4 member D |
| chr4_+_85827745 | 0.01 |
ENST00000509300.5
|
ARHGAP24
|
Rho GTPase activating protein 24 |
| chr13_-_26221703 | 0.01 |
ENST00000381570.7
ENST00000346166.7 |
RNF6
|
ring finger protein 6 |
| chrX_-_13817027 | 0.01 |
ENST00000493677.5
ENST00000355135.6 ENST00000316715.9 |
GPM6B
|
glycoprotein M6B |
| chr8_+_91249307 | 0.01 |
ENST00000309536.6
ENST00000276609.8 |
SLC26A7
|
solute carrier family 26 member 7 |
| chr12_+_25052634 | 0.00 |
ENST00000548766.5
|
IRAG2
|
inositol 1,4,5-triphosphate receptor associated 2 |
| chr2_+_119431846 | 0.00 |
ENST00000306406.5
|
TMEM37
|
transmembrane protein 37 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.7 | GO:1902261 | positive regulation of delayed rectifier potassium channel activity(GO:1902261) |
| 0.1 | 0.4 | GO:0035948 | positive regulation of gluconeogenesis by positive regulation of transcription from RNA polymerase II promoter(GO:0035948) |
| 0.1 | 0.9 | GO:0021840 | directional guidance of interneurons involved in migration from the subpallium to the cortex(GO:0021840) chemorepulsion involved in interneuron migration from the subpallium to the cortex(GO:0021842) ERBB3 signaling pathway(GO:0038129) |
| 0.1 | 0.2 | GO:0070352 | positive regulation of white fat cell proliferation(GO:0070352) |
| 0.1 | 0.2 | GO:0030185 | nitric oxide transport(GO:0030185) |
| 0.1 | 0.3 | GO:0021778 | oligodendrocyte cell fate specification(GO:0021778) oligodendrocyte cell fate commitment(GO:0021779) glial cell fate specification(GO:0021780) |
| 0.1 | 0.6 | GO:0031087 | deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
| 0.1 | 0.2 | GO:0002331 | pre-B cell allelic exclusion(GO:0002331) |
| 0.1 | 0.7 | GO:2001288 | positive regulation of caveolin-mediated endocytosis(GO:2001288) |
| 0.1 | 0.2 | GO:0014016 | proximal/distal axis specification(GO:0009946) neuroblast differentiation(GO:0014016) |
| 0.1 | 0.3 | GO:0032468 | Golgi calcium ion homeostasis(GO:0032468) |
| 0.1 | 0.4 | GO:0007197 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) |
| 0.0 | 0.5 | GO:0043950 | positive regulation of cAMP-mediated signaling(GO:0043950) |
| 0.0 | 0.6 | GO:0098870 | neuronal action potential propagation(GO:0019227) clustering of voltage-gated sodium channels(GO:0045162) action potential propagation(GO:0098870) |
| 0.0 | 0.1 | GO:0072660 | maintenance of protein location in membrane(GO:0072658) maintenance of protein location in plasma membrane(GO:0072660) positive regulation of membrane depolarization during cardiac muscle cell action potential(GO:1900827) |
| 0.0 | 0.1 | GO:0051758 | homologous chromosome movement towards spindle pole involved in homologous chromosome segregation(GO:0051758) |
| 0.0 | 0.2 | GO:0051684 | maintenance of Golgi location(GO:0051684) double-strand break repair via classical nonhomologous end joining(GO:0097680) |
| 0.0 | 0.8 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.0 | 0.4 | GO:0030916 | otic vesicle formation(GO:0030916) |
| 0.0 | 0.3 | GO:0036444 | calcium ion transmembrane import into mitochondrion(GO:0036444) |
| 0.0 | 0.3 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.0 | 0.4 | GO:0061158 | 3'-UTR-mediated mRNA destabilization(GO:0061158) |
| 0.0 | 0.1 | GO:0060478 | acrosomal vesicle exocytosis(GO:0060478) |
| 0.0 | 0.4 | GO:0048742 | regulation of skeletal muscle fiber development(GO:0048742) |
| 0.0 | 0.2 | GO:0003402 | planar cell polarity pathway involved in axis elongation(GO:0003402) |
| 0.0 | 0.2 | GO:2001206 | positive regulation of osteoclast development(GO:2001206) |
| 0.0 | 0.7 | GO:0090162 | establishment of epithelial cell polarity(GO:0090162) |
| 0.0 | 0.1 | GO:0018343 | protein farnesylation(GO:0018343) |
| 0.0 | 0.3 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.0 | 0.1 | GO:0021615 | glossopharyngeal nerve morphogenesis(GO:0021615) |
| 0.0 | 0.2 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.0 | 0.1 | GO:0036371 | protein localization to T-tubule(GO:0036371) |
| 0.0 | 0.3 | GO:0007614 | short-term memory(GO:0007614) |
| 0.0 | 0.0 | GO:0061034 | olfactory bulb mitral cell layer development(GO:0061034) |
| 0.0 | 0.1 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.8 | GO:0001674 | female germ cell nucleus(GO:0001674) |
| 0.1 | 1.4 | GO:0016013 | syntrophin complex(GO:0016013) |
| 0.1 | 0.2 | GO:0048237 | rough endoplasmic reticulum lumen(GO:0048237) |
| 0.1 | 0.7 | GO:0014701 | junctional sarcoplasmic reticulum membrane(GO:0014701) |
| 0.0 | 0.3 | GO:1990246 | uniplex complex(GO:1990246) |
| 0.0 | 0.8 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.0 | 0.1 | GO:0005965 | protein farnesyltransferase complex(GO:0005965) |
| 0.0 | 0.2 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.0 | 0.9 | GO:0030673 | axolemma(GO:0030673) |
| 0.0 | 0.3 | GO:0000177 | cytoplasmic exosome (RNase complex)(GO:0000177) |
| 0.0 | 0.2 | GO:0030897 | HOPS complex(GO:0030897) |
| 0.0 | 0.2 | GO:0036056 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.0 | 0.1 | GO:0000839 | Hrd1p ubiquitin ligase ERAD-L complex(GO:0000839) |
| 0.0 | 0.5 | GO:0097228 | sperm principal piece(GO:0097228) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0001588 | dopamine neurotransmitter receptor activity, coupled via Gs(GO:0001588) |
| 0.1 | 0.4 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 0.1 | 0.2 | GO:0010348 | lithium:proton antiporter activity(GO:0010348) |
| 0.1 | 0.4 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.1 | 0.3 | GO:0005223 | intracellular cGMP activated cation channel activity(GO:0005223) |
| 0.0 | 0.9 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 0.0 | 0.3 | GO:0030345 | extracellular matrix structural constituent conferring compression resistance(GO:0030021) structural constituent of tooth enamel(GO:0030345) |
| 0.0 | 0.7 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.0 | 0.2 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.0 | 0.6 | GO:0086080 | protein binding involved in heterotypic cell-cell adhesion(GO:0086080) |
| 0.0 | 0.2 | GO:0031708 | endothelin B receptor binding(GO:0031708) |
| 0.0 | 0.7 | GO:0019871 | sodium channel inhibitor activity(GO:0019871) |
| 0.0 | 0.1 | GO:0005174 | CD40 receptor binding(GO:0005174) |
| 0.0 | 0.2 | GO:0070699 | type II activin receptor binding(GO:0070699) |
| 0.0 | 0.2 | GO:0035727 | lysophosphatidic acid binding(GO:0035727) |
| 0.0 | 0.1 | GO:0050309 | glucose-6-phosphatase activity(GO:0004346) sugar-terminal-phosphatase activity(GO:0050309) |
| 0.0 | 0.1 | GO:0004660 | protein farnesyltransferase activity(GO:0004660) |
| 0.0 | 0.1 | GO:0015319 | sodium:inorganic phosphate symporter activity(GO:0015319) |
| 0.0 | 0.5 | GO:0030247 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.0 | 1.1 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.0 | 0.2 | GO:0042731 | inositol-polyphosphate 5-phosphatase activity(GO:0004445) PH domain binding(GO:0042731) |
| 0.0 | 0.1 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.0 | 0.4 | GO:0035198 | miRNA binding(GO:0035198) |
| 0.0 | 0.2 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.0 | 0.3 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.9 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.8 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.0 | 0.6 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.0 | 0.9 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.0 | 4.6 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.8 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 0.4 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.0 | 0.5 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.0 | 0.2 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.0 | 0.2 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |