Project
Inflammatory response time course, HUVEC (Wada, 2009)
Navigation
Downloads
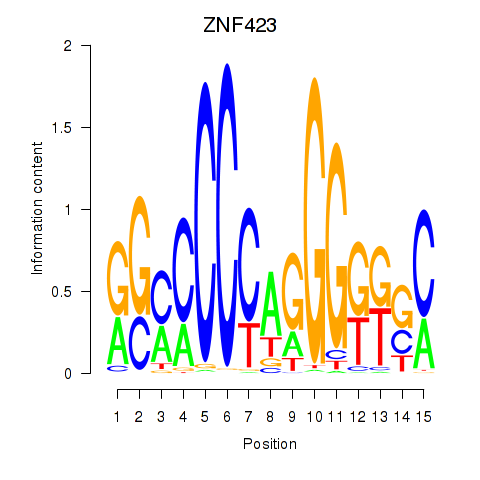
Results for ZNF423
Z-value: 0.99
Transcription factors associated with ZNF423
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ZNF423
|
ENSG00000102935.12 | ZNF423 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ZNF423 | hg38_v1_chr16_-_49856105_49856119 | -0.66 | 3.0e-04 | Click! |
Activity profile of ZNF423 motif
Sorted Z-values of ZNF423 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ZNF423
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr16_-_11587162 | 4.96 |
ENST00000570904.5
ENST00000574701.5 |
LITAF
|
lipopolysaccharide induced TNF factor |
| chr17_+_79024142 | 4.00 |
ENST00000579760.6
ENST00000339142.6 |
C1QTNF1
|
C1q and TNF related 1 |
| chr17_+_79024243 | 3.97 |
ENST00000311661.4
|
C1QTNF1
|
C1q and TNF related 1 |
| chr16_-_11587450 | 3.41 |
ENST00000571688.5
|
LITAF
|
lipopolysaccharide induced TNF factor |
| chr12_-_49707368 | 3.29 |
ENST00000352151.9
ENST00000335154.10 |
FMNL3
|
formin like 3 |
| chr12_-_57129001 | 2.85 |
ENST00000556155.5
|
STAT6
|
signal transducer and activator of transcription 6 |
| chr12_-_49707220 | 2.62 |
ENST00000550488.5
|
FMNL3
|
formin like 3 |
| chr4_-_119628007 | 2.50 |
ENST00000420633.1
ENST00000394439.5 |
PDE5A
|
phosphodiesterase 5A |
| chr8_-_79767843 | 2.40 |
ENST00000337919.9
|
HEY1
|
hes related family bHLH transcription factor with YRPW motif 1 |
| chr8_-_79767462 | 2.37 |
ENST00000674295.1
ENST00000518733.1 ENST00000674418.1 ENST00000674358.1 ENST00000354724.8 |
HEY1
|
hes related family bHLH transcription factor with YRPW motif 1 |
| chr5_+_136049513 | 2.08 |
ENST00000514554.5
|
TGFBI
|
transforming growth factor beta induced |
| chr7_-_19145306 | 1.94 |
ENST00000275461.3
|
FERD3L
|
Fer3 like bHLH transcription factor |
| chr1_-_203086001 | 1.93 |
ENST00000241651.5
|
MYOG
|
myogenin |
| chr16_+_2988256 | 1.82 |
ENST00000573315.2
|
LINC00514
|
long intergenic non-protein coding RNA 514 |
| chr5_-_139482341 | 1.79 |
ENST00000651699.1
|
STING1
|
stimulator of interferon response cGAMP interactor 1 |
| chr21_-_17612842 | 1.64 |
ENST00000339775.10
ENST00000348354.7 |
BTG3
|
BTG anti-proliferation factor 3 |
| chr10_-_119536533 | 1.60 |
ENST00000392865.5
|
RGS10
|
regulator of G protein signaling 10 |
| chr16_+_30664334 | 1.51 |
ENST00000287468.5
|
FBRS
|
fibrosin |
| chr2_+_27496830 | 1.49 |
ENST00000264717.7
|
GCKR
|
glucokinase regulator |
| chr10_-_98268186 | 1.43 |
ENST00000260702.4
|
LOXL4
|
lysyl oxidase like 4 |
| chr19_+_45079195 | 1.40 |
ENST00000591607.1
ENST00000591747.5 ENST00000270257.9 ENST00000391951.2 ENST00000587566.5 |
GEMIN7
MARK4
|
gem nuclear organelle associated protein 7 microtubule affinity regulating kinase 4 |
| chr11_-_61891381 | 1.34 |
ENST00000525588.5
|
FADS3
|
fatty acid desaturase 3 |
| chr22_-_37427433 | 1.30 |
ENST00000452946.1
ENST00000402918.7 |
ELFN2
ELFN2
|
extracellular leucine rich repeat and fibronectin type III domain containing 2 extracellular leucine rich repeat and fibronectin type III domain containing 2 |
| chr7_-_23014074 | 1.27 |
ENST00000409763.1
ENST00000679826.1 ENST00000409923.5 ENST00000681766.1 |
FAM126A
|
family with sequence similarity 126 member A |
| chr19_+_14433284 | 1.24 |
ENST00000242783.11
|
PKN1
|
protein kinase N1 |
| chr4_-_119627631 | 1.22 |
ENST00000264805.9
|
PDE5A
|
phosphodiesterase 5A |
| chr5_+_53480619 | 1.15 |
ENST00000396947.7
ENST00000256759.8 |
FST
|
follistatin |
| chr11_-_61891534 | 1.13 |
ENST00000278829.7
|
FADS3
|
fatty acid desaturase 3 |
| chr6_+_11093753 | 1.13 |
ENST00000416247.4
|
SMIM13
|
small integral membrane protein 13 |
| chr7_-_23014099 | 1.12 |
ENST00000432176.7
ENST00000440481.6 |
FAM126A
|
family with sequence similarity 126 member A |
| chr16_+_27313879 | 1.05 |
ENST00000562142.5
ENST00000561742.5 ENST00000543915.6 ENST00000395762.7 ENST00000563002.5 |
IL4R
|
interleukin 4 receptor |
| chrX_+_30247139 | 0.99 |
ENST00000397548.4
|
MAGEB1
|
MAGE family member B1 |
| chr20_+_33993904 | 0.98 |
ENST00000246194.8
|
RALY
|
RALY heterogeneous nuclear ribonucleoprotein |
| chr1_+_1512137 | 0.97 |
ENST00000378756.8
ENST00000378755.9 |
ATAD3A
|
ATPase family AAA domain containing 3A |
| chr22_-_38088915 | 0.96 |
ENST00000428572.1
|
BAIAP2L2
|
BAR/IMD domain containing adaptor protein 2 like 2 |
| chr6_+_109440695 | 0.94 |
ENST00000258052.8
|
SMPD2
|
sphingomyelin phosphodiesterase 2 |
| chr9_+_109780292 | 0.93 |
ENST00000374530.7
|
PALM2AKAP2
|
PALM2 and AKAP2 fusion |
| chr15_-_75368578 | 0.91 |
ENST00000569482.5
ENST00000565683.5 ENST00000561615.1 ENST00000563622.5 ENST00000568374.5 ENST00000267978.10 ENST00000566256.5 |
MAN2C1
|
mannosidase alpha class 2C member 1 |
| chr19_-_49867251 | 0.86 |
ENST00000631020.2
ENST00000596014.5 ENST00000636994.1 |
PNKP
|
polynucleotide kinase 3'-phosphatase |
| chr1_+_1471751 | 0.84 |
ENST00000673477.1
ENST00000308647.8 |
ATAD3B
|
ATPase family AAA domain containing 3B |
| chr5_+_171309239 | 0.81 |
ENST00000296921.6
|
TLX3
|
T cell leukemia homeobox 3 |
| chr12_+_52079700 | 0.80 |
ENST00000546390.2
|
SMIM41
|
small integral membrane protein 41 |
| chr19_+_35533436 | 0.79 |
ENST00000222286.9
|
GAPDHS
|
glyceraldehyde-3-phosphate dehydrogenase, spermatogenic |
| chr9_+_132162045 | 0.77 |
ENST00000393229.4
|
NTNG2
|
netrin G2 |
| chr1_+_26993684 | 0.76 |
ENST00000522111.3
|
TRNP1
|
TMF1 regulated nuclear protein 1 |
| chr11_-_61295289 | 0.76 |
ENST00000335613.10
|
VWCE
|
von Willebrand factor C and EGF domains |
| chr8_+_30384511 | 0.76 |
ENST00000339877.8
ENST00000320203.8 ENST00000287771.9 ENST00000397323.9 |
RBPMS
|
RNA binding protein, mRNA processing factor |
| chr7_-_100894233 | 0.71 |
ENST00000426415.5
ENST00000430554.1 ENST00000412389.5 |
ACHE
|
acetylcholinesterase (Cartwright blood group) |
| chr19_-_10339610 | 0.69 |
ENST00000589261.5
ENST00000160262.10 ENST00000590569.1 ENST00000589580.1 ENST00000589249.1 |
ICAM3
|
intercellular adhesion molecule 3 |
| chr7_-_103989649 | 0.68 |
ENST00000428762.6
|
RELN
|
reelin |
| chr22_-_42649332 | 0.67 |
ENST00000352397.10
|
CYB5R3
|
cytochrome b5 reductase 3 |
| chr8_+_22165358 | 0.67 |
ENST00000306349.13
ENST00000306385.10 |
BMP1
|
bone morphogenetic protein 1 |
| chr11_-_30586866 | 0.66 |
ENST00000528686.2
|
MPPED2
|
metallophosphoesterase domain containing 2 |
| chr1_-_6180265 | 0.64 |
ENST00000262450.8
|
CHD5
|
chromodomain helicase DNA binding protein 5 |
| chrX_+_137566119 | 0.64 |
ENST00000287538.10
|
ZIC3
|
Zic family member 3 |
| chr16_-_70685791 | 0.60 |
ENST00000616026.4
|
MTSS2
|
MTSS I-BAR domain containing 2 |
| chr17_+_7438267 | 0.58 |
ENST00000575235.5
|
FGF11
|
fibroblast growth factor 11 |
| chr10_+_84173793 | 0.56 |
ENST00000372126.4
|
C10orf99
|
chromosome 10 open reading frame 99 |
| chr8_+_22565655 | 0.55 |
ENST00000523965.5
|
SORBS3
|
sorbin and SH3 domain containing 3 |
| chr22_+_35257452 | 0.54 |
ENST00000420166.5
ENST00000216106.6 ENST00000455359.5 |
HMGXB4
|
HMG-box containing 4 |
| chr5_+_169583636 | 0.54 |
ENST00000506574.5
ENST00000515224.5 ENST00000629457.2 ENST00000508247.5 ENST00000265295.9 ENST00000513941.5 |
SPDL1
|
spindle apparatus coiled-coil protein 1 |
| chr6_+_33621313 | 0.53 |
ENST00000605930.3
|
ITPR3
|
inositol 1,4,5-trisphosphate receptor type 3 |
| chr8_+_22165140 | 0.52 |
ENST00000397814.7
ENST00000354870.5 |
BMP1
|
bone morphogenetic protein 1 |
| chr9_+_113876282 | 0.52 |
ENST00000374126.9
ENST00000615615.4 ENST00000288466.11 |
ZNF618
|
zinc finger protein 618 |
| chr6_-_35512882 | 0.51 |
ENST00000229771.11
|
TULP1
|
TUB like protein 1 |
| chr12_+_57128656 | 0.51 |
ENST00000338962.8
|
LRP1
|
LDL receptor related protein 1 |
| chr19_+_4279247 | 0.50 |
ENST00000543264.7
|
SHD
|
Src homology 2 domain containing transforming protein D |
| chr18_-_5543988 | 0.50 |
ENST00000341928.7
|
EPB41L3
|
erythrocyte membrane protein band 4.1 like 3 |
| chr6_-_35512863 | 0.49 |
ENST00000428978.1
ENST00000614066.4 ENST00000322263.8 |
TULP1
|
TUB like protein 1 |
| chr3_-_196082078 | 0.49 |
ENST00000360110.9
ENST00000392396.7 ENST00000420415.5 |
TFRC
|
transferrin receptor |
| chr6_-_31112566 | 0.48 |
ENST00000259870.4
|
C6orf15
|
chromosome 6 open reading frame 15 |
| chr20_+_44885679 | 0.48 |
ENST00000353703.9
ENST00000372839.7 ENST00000428262.1 ENST00000445830.1 |
YWHAB
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein beta |
| chr10_+_123154768 | 0.47 |
ENST00000407911.2
|
BUB3
|
BUB3 mitotic checkpoint protein |
| chr12_-_54385727 | 0.46 |
ENST00000551109.5
ENST00000546970.5 |
ZNF385A
|
zinc finger protein 385A |
| chr2_-_120223371 | 0.42 |
ENST00000426077.3
|
TMEM185B
|
transmembrane protein 185B |
| chr20_+_33993646 | 0.42 |
ENST00000375114.7
ENST00000448364.5 |
RALY
|
RALY heterogeneous nuclear ribonucleoprotein |
| chr19_-_11481044 | 0.42 |
ENST00000359227.8
|
ELAVL3
|
ELAV like RNA binding protein 3 |
| chr16_+_3046552 | 0.41 |
ENST00000336577.9
|
MMP25
|
matrix metallopeptidase 25 |
| chr19_-_42242526 | 0.41 |
ENST00000222330.8
ENST00000676537.1 |
GSK3A
|
glycogen synthase kinase 3 alpha |
| chr4_-_20984011 | 0.40 |
ENST00000382149.9
|
KCNIP4
|
potassium voltage-gated channel interacting protein 4 |
| chr7_+_20330893 | 0.39 |
ENST00000222573.5
|
ITGB8
|
integrin subunit beta 8 |
| chr20_+_31475278 | 0.39 |
ENST00000201979.3
|
REM1
|
RRAD and GEM like GTPase 1 |
| chr10_+_123154414 | 0.39 |
ENST00000368858.9
|
BUB3
|
BUB3 mitotic checkpoint protein |
| chr11_-_30586344 | 0.38 |
ENST00000358117.10
|
MPPED2
|
metallophosphoesterase domain containing 2 |
| chr19_-_55599493 | 0.37 |
ENST00000221665.5
ENST00000592585.1 |
FIZ1
|
FLT3 interacting zinc finger 1 |
| chr10_+_123154364 | 0.36 |
ENST00000368859.6
ENST00000368865.9 |
BUB3
|
BUB3 mitotic checkpoint protein |
| chr20_+_23035312 | 0.36 |
ENST00000255008.5
|
SSTR4
|
somatostatin receptor 4 |
| chr17_-_44376169 | 0.36 |
ENST00000587295.5
|
ITGA2B
|
integrin subunit alpha 2b |
| chr10_+_133527355 | 0.35 |
ENST00000252945.8
ENST00000421586.5 ENST00000418356.1 |
CYP2E1
|
cytochrome P450 family 2 subfamily E member 1 |
| chr5_-_176630517 | 0.35 |
ENST00000393693.7
ENST00000614675.4 |
SNCB
|
synuclein beta |
| chr11_-_107858777 | 0.35 |
ENST00000525815.6
|
SLC35F2
|
solute carrier family 35 member F2 |
| chr19_-_15200902 | 0.34 |
ENST00000601011.1
ENST00000263388.7 |
NOTCH3
|
notch receptor 3 |
| chr2_+_170929198 | 0.34 |
ENST00000234160.5
|
GORASP2
|
golgi reassembly stacking protein 2 |
| chr2_+_227616998 | 0.34 |
ENST00000641801.1
|
SCYGR4
|
small cysteine and glycine repeat containing 4 |
| chr6_-_90586883 | 0.34 |
ENST00000369325.7
ENST00000369327.7 |
MAP3K7
|
mitogen-activated protein kinase kinase kinase 7 |
| chr16_+_283157 | 0.33 |
ENST00000219406.11
ENST00000404312.5 ENST00000456379.1 |
PDIA2
|
protein disulfide isomerase family A member 2 |
| chr19_-_38224215 | 0.33 |
ENST00000355526.10
ENST00000420980.7 ENST00000614244.4 |
DPF1
|
double PHD fingers 1 |
| chr5_-_176630364 | 0.32 |
ENST00000310112.7
|
SNCB
|
synuclein beta |
| chr1_-_228109255 | 0.32 |
ENST00000483159.1
ENST00000366744.5 ENST00000348259.9 ENST00000495434.1 ENST00000295008.8 ENST00000366731.9 ENST00000366746.7 ENST00000366747.7 ENST00000336520.8 ENST00000391867.8 ENST00000457264.5 ENST00000464148.2 ENST00000336300.9 ENST00000430433.5 |
MRPL55
|
mitochondrial ribosomal protein L55 |
| chr19_+_4279285 | 0.31 |
ENST00000599689.1
|
SHD
|
Src homology 2 domain containing transforming protein D |
| chr9_+_98807619 | 0.31 |
ENST00000375011.4
|
GALNT12
|
polypeptide N-acetylgalactosaminyltransferase 12 |
| chr6_+_30163188 | 0.31 |
ENST00000619857.4
|
TRIM15
|
tripartite motif containing 15 |
| chr11_-_44950151 | 0.30 |
ENST00000533940.5
ENST00000533937.1 |
TP53I11
|
tumor protein p53 inducible protein 11 |
| chr7_-_5530581 | 0.30 |
ENST00000676397.1
ENST00000645576.1 ENST00000674681.1 ENST00000646664.1 ENST00000493945.6 ENST00000676319.1 ENST00000473257.3 ENST00000647275.1 ENST00000432588.6 |
ACTB
|
actin beta |
| chr10_-_73495966 | 0.28 |
ENST00000342558.3
ENST00000360663.10 ENST00000394828.6 ENST00000394829.6 |
PPP3CB
|
protein phosphatase 3 catalytic subunit beta |
| chr19_-_4066892 | 0.28 |
ENST00000322357.9
|
ZBTB7A
|
zinc finger and BTB domain containing 7A |
| chr19_+_48393657 | 0.28 |
ENST00000263269.4
|
GRIN2D
|
glutamate ionotropic receptor NMDA type subunit 2D |
| chr6_-_90587018 | 0.26 |
ENST00000369332.7
ENST00000369329.8 |
MAP3K7
|
mitogen-activated protein kinase kinase kinase 7 |
| chr7_+_29194757 | 0.25 |
ENST00000222792.11
|
CHN2
|
chimerin 2 |
| chr9_+_128787331 | 0.25 |
ENST00000223865.8
|
TBC1D13
|
TBC1 domain family member 13 |
| chr7_+_151051201 | 0.25 |
ENST00000490540.1
|
ASIC3
|
acid sensing ion channel subunit 3 |
| chr8_+_22565236 | 0.24 |
ENST00000523900.5
|
SORBS3
|
sorbin and SH3 domain containing 3 |
| chr20_-_31475125 | 0.23 |
ENST00000317676.3
|
DEFB124
|
defensin beta 124 |
| chr2_-_206086057 | 0.23 |
ENST00000403263.6
|
INO80D
|
INO80 complex subunit D |
| chr6_+_30163541 | 0.23 |
ENST00000376694.9
|
TRIM15
|
tripartite motif containing 15 |
| chr1_+_26317950 | 0.22 |
ENST00000374213.3
|
CD52
|
CD52 molecule |
| chr1_-_101996919 | 0.22 |
ENST00000370103.9
|
OLFM3
|
olfactomedin 3 |
| chr17_+_6444441 | 0.22 |
ENST00000250056.12
ENST00000571373.5 ENST00000570337.6 ENST00000572595.6 ENST00000572447.6 ENST00000576056.5 |
PIMREG
|
PICALM interacting mitotic regulator |
| chrX_+_49171889 | 0.22 |
ENST00000376327.6
|
PLP2
|
proteolipid protein 2 |
| chr1_-_32901330 | 0.21 |
ENST00000329151.5
ENST00000373463.8 |
TMEM54
|
transmembrane protein 54 |
| chr7_-_100158679 | 0.21 |
ENST00000456769.5
ENST00000316937.8 |
TRAPPC14
|
trafficking protein particle complex 14 |
| chr12_+_109713817 | 0.20 |
ENST00000538780.2
|
FAM222A
|
family with sequence similarity 222 member A |
| chr9_-_128724088 | 0.20 |
ENST00000406904.2
ENST00000452105.5 ENST00000372667.9 ENST00000372663.9 |
ZDHHC12
|
zinc finger DHHC-type palmitoyltransferase 12 |
| chr11_-_1036706 | 0.20 |
ENST00000421673.7
|
MUC6
|
mucin 6, oligomeric mucus/gel-forming |
| chr9_-_37904085 | 0.20 |
ENST00000377716.6
ENST00000242275.7 |
SLC25A51
|
solute carrier family 25 member 51 |
| chr16_+_2964216 | 0.19 |
ENST00000572045.5
ENST00000571007.5 ENST00000575885.5 ENST00000303746.10 ENST00000319500.10 |
KREMEN2
|
kringle containing transmembrane protein 2 |
| chr11_-_44950867 | 0.18 |
ENST00000528290.5
ENST00000525680.6 ENST00000530035.5 ENST00000527685.5 |
TP53I11
|
tumor protein p53 inducible protein 11 |
| chr11_-_63015831 | 0.17 |
ENST00000430500.6
ENST00000336232.7 |
SLC22A8
|
solute carrier family 22 member 8 |
| chr1_-_165445220 | 0.17 |
ENST00000619224.1
|
RXRG
|
retinoid X receptor gamma |
| chrX_+_49171918 | 0.15 |
ENST00000376322.7
|
PLP2
|
proteolipid protein 2 |
| chr19_+_45251363 | 0.14 |
ENST00000620044.4
|
MARK4
|
microtubule affinity regulating kinase 4 |
| chr9_-_127980976 | 0.14 |
ENST00000373095.6
|
FAM102A
|
family with sequence similarity 102 member A |
| chr13_+_51222391 | 0.14 |
ENST00000322475.13
|
FAM124A
|
family with sequence similarity 124 member A |
| chr11_-_44950839 | 0.14 |
ENST00000395648.7
ENST00000531928.6 |
TP53I11
|
tumor protein p53 inducible protein 11 |
| chr19_+_45251249 | 0.14 |
ENST00000262891.9
ENST00000300843.8 |
MARK4
|
microtubule affinity regulating kinase 4 |
| chr1_-_165445088 | 0.13 |
ENST00000359842.10
|
RXRG
|
retinoid X receptor gamma |
| chr10_+_125896549 | 0.11 |
ENST00000368693.6
|
FANK1
|
fibronectin type III and ankyrin repeat domains 1 |
| chr1_-_155207886 | 0.10 |
ENST00000368378.7
ENST00000541990.5 ENST00000457183.6 ENST00000541576.5 |
THBS3
|
thrombospondin 3 |
| chr17_-_81239025 | 0.09 |
ENST00000637944.2
|
TEPSIN
|
TEPSIN adaptor related protein complex 4 accessory protein |
| chr6_-_109440504 | 0.09 |
ENST00000520723.5
ENST00000518648.1 ENST00000417394.6 ENST00000521072.7 |
PPIL6
|
peptidylprolyl isomerase like 6 |
| chr3_+_184338826 | 0.09 |
ENST00000453072.5
|
FAM131A
|
family with sequence similarity 131 member A |
| chr3_+_138349209 | 0.08 |
ENST00000474559.1
|
MRAS
|
muscle RAS oncogene homolog |
| chr7_+_20330678 | 0.07 |
ENST00000537992.5
|
ITGB8
|
integrin subunit beta 8 |
| chr20_-_25082131 | 0.04 |
ENST00000429762.7
ENST00000444511.6 ENST00000376707.4 ENST00000376709.9 |
VSX1
|
visual system homeobox 1 |
| chr14_-_93115812 | 0.03 |
ENST00000553452.5
|
ITPK1
|
inositol-tetrakisphosphate 1-kinase |
| chr9_+_136982993 | 0.02 |
ENST00000408973.3
|
LCNL1
|
lipocalin like 1 |
| chr17_+_2337622 | 0.02 |
ENST00000574563.5
|
SGSM2
|
small G protein signaling modulator 2 |
| chr9_-_127897399 | 0.02 |
ENST00000542456.5
ENST00000373141.5 |
ST6GALNAC6
|
ST6 N-acetylgalactosaminide alpha-2,6-sialyltransferase 6 |
| chr15_-_41230697 | 0.02 |
ENST00000314992.9
ENST00000558396.1 ENST00000458580.7 |
EXD1
|
exonuclease 3'-5' domain containing 1 |
| chr9_-_96619378 | 0.01 |
ENST00000375240.7
ENST00000463569.5 |
CDC14B
|
cell division cycle 14B |
| chrX_+_106920393 | 0.00 |
ENST00000336803.2
|
CLDN2
|
claudin 2 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 4.8 | GO:0061027 | umbilical cord morphogenesis(GO:0036304) umbilical cord development(GO:0061027) |
| 1.1 | 8.0 | GO:2000857 | positive regulation of mineralocorticoid secretion(GO:2000857) positive regulation of aldosterone secretion(GO:2000860) |
| 0.7 | 2.9 | GO:0002296 | T-helper 1 cell lineage commitment(GO:0002296) |
| 0.6 | 1.9 | GO:0014873 | response to muscle activity involved in regulation of muscle adaptation(GO:0014873) |
| 0.5 | 3.7 | GO:0060282 | positive regulation of oocyte development(GO:0060282) |
| 0.3 | 1.2 | GO:0035407 | histone H3-T11 phosphorylation(GO:0035407) |
| 0.3 | 1.1 | GO:0045626 | negative regulation of T-helper 1 cell differentiation(GO:0045626) response to odorant(GO:1990834) |
| 0.2 | 1.9 | GO:0033504 | floor plate development(GO:0033504) |
| 0.2 | 0.7 | GO:0045212 | negative regulation of synaptic transmission, cholinergic(GO:0032223) neurotransmitter receptor biosynthetic process(GO:0045212) |
| 0.2 | 0.7 | GO:0097477 | lateral motor column neuron migration(GO:0097477) |
| 0.2 | 8.4 | GO:0042347 | negative regulation of NF-kappaB import into nucleus(GO:0042347) |
| 0.2 | 1.5 | GO:1903300 | negative regulation of glucokinase activity(GO:0033132) negative regulation of hexokinase activity(GO:1903300) |
| 0.2 | 0.5 | GO:1901253 | negative regulation of intracellular transport of viral material(GO:1901253) |
| 0.2 | 0.9 | GO:0010836 | negative regulation of protein ADP-ribosylation(GO:0010836) |
| 0.1 | 0.5 | GO:1905167 | positive regulation of lysosomal protein catabolic process(GO:1905167) |
| 0.1 | 0.9 | GO:0006013 | mannose metabolic process(GO:0006013) |
| 0.1 | 0.5 | GO:0010609 | mRNA localization resulting in posttranscriptional regulation of gene expression(GO:0010609) platelet alpha granule organization(GO:0070889) regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902162) positive regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902164) |
| 0.1 | 1.0 | GO:0097500 | receptor localization to nonmotile primary cilium(GO:0097500) |
| 0.1 | 0.6 | GO:0098532 | histone H3-K27 trimethylation(GO:0098532) |
| 0.1 | 0.4 | GO:0006258 | UDP-glucose catabolic process(GO:0006258) negative regulation of type B pancreatic cell development(GO:2000077) |
| 0.1 | 0.3 | GO:0098974 | postsynaptic actin cytoskeleton organization(GO:0098974) |
| 0.1 | 0.4 | GO:1901842 | negative regulation of high voltage-gated calcium channel activity(GO:1901842) |
| 0.1 | 0.5 | GO:0050917 | sensory perception of sweet taste(GO:0050916) sensory perception of umami taste(GO:0050917) |
| 0.1 | 0.6 | GO:0035469 | determination of pancreatic left/right asymmetry(GO:0035469) |
| 0.1 | 0.4 | GO:0038170 | somatostatin receptor signaling pathway(GO:0038169) somatostatin signaling pathway(GO:0038170) |
| 0.1 | 0.4 | GO:0010193 | response to ozone(GO:0010193) |
| 0.1 | 1.2 | GO:0032926 | negative regulation of activin receptor signaling pathway(GO:0032926) positive regulation of hair follicle development(GO:0051798) |
| 0.1 | 1.6 | GO:0007213 | G-protein coupled acetylcholine receptor signaling pathway(GO:0007213) |
| 0.1 | 0.5 | GO:0002175 | protein localization to paranode region of axon(GO:0002175) |
| 0.0 | 0.2 | GO:0015891 | iron chelate transport(GO:0015688) siderophore transport(GO:0015891) |
| 0.0 | 0.3 | GO:0072103 | glomerulus vasculature morphogenesis(GO:0072103) glomerular capillary formation(GO:0072104) |
| 0.0 | 0.9 | GO:2000304 | positive regulation of sphingolipid biosynthetic process(GO:0090154) positive regulation of ceramide biosynthetic process(GO:2000304) |
| 0.0 | 1.2 | GO:0034380 | high-density lipoprotein particle assembly(GO:0034380) |
| 0.0 | 0.8 | GO:0002087 | regulation of respiratory gaseous exchange by neurological system process(GO:0002087) |
| 0.0 | 0.6 | GO:0002726 | positive regulation of T cell cytokine production(GO:0002726) |
| 0.0 | 1.0 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.0 | 0.3 | GO:0001915 | negative regulation of T cell mediated cytotoxicity(GO:0001915) |
| 0.0 | 1.4 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.0 | 1.2 | GO:0008608 | attachment of spindle microtubules to kinetochore(GO:0008608) |
| 0.0 | 0.3 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 0.0 | 0.5 | GO:0034501 | protein localization to kinetochore(GO:0034501) |
| 0.0 | 0.7 | GO:0019852 | L-ascorbic acid metabolic process(GO:0019852) |
| 0.0 | 0.5 | GO:0046852 | positive regulation of bone resorption(GO:0045780) positive regulation of bone remodeling(GO:0046852) |
| 0.0 | 2.4 | GO:0006636 | unsaturated fatty acid biosynthetic process(GO:0006636) |
| 0.0 | 0.4 | GO:0060022 | hard palate development(GO:0060022) |
| 0.0 | 5.9 | GO:0008360 | regulation of cell shape(GO:0008360) |
| 0.0 | 0.6 | GO:0045821 | positive regulation of glycolytic process(GO:0045821) |
| 0.0 | 0.5 | GO:0001573 | ganglioside metabolic process(GO:0001573) |
| 0.0 | 2.4 | GO:0046854 | phosphatidylinositol phosphorylation(GO:0046854) |
| 0.0 | 0.5 | GO:0051220 | cytoplasmic sequestering of protein(GO:0051220) |
| 0.0 | 0.2 | GO:2000188 | regulation of cholesterol homeostasis(GO:2000188) |
| 0.0 | 0.8 | GO:0051496 | positive regulation of stress fiber assembly(GO:0051496) |
| 0.0 | 0.7 | GO:0042417 | dopamine metabolic process(GO:0042417) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 8.4 | GO:0098560 | cytoplasmic side of early endosome membrane(GO:0098559) cytoplasmic side of late endosome membrane(GO:0098560) |
| 0.4 | 1.2 | GO:1990298 | mitotic checkpoint complex(GO:0033597) bub1-bub3 complex(GO:1990298) |
| 0.2 | 0.5 | GO:0034686 | integrin alphav-beta8 complex(GO:0034686) |
| 0.1 | 1.4 | GO:0097504 | Gemini of coiled bodies(GO:0097504) |
| 0.1 | 8.0 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.1 | 1.0 | GO:0071439 | clathrin complex(GO:0071439) |
| 0.1 | 0.7 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.1 | 0.5 | GO:0016589 | NURF complex(GO:0016589) |
| 0.0 | 0.7 | GO:0005833 | hemoglobin complex(GO:0005833) |
| 0.0 | 0.6 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.0 | 0.5 | GO:1990712 | HFE-transferrin receptor complex(GO:1990712) |
| 0.0 | 0.5 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
| 0.0 | 0.8 | GO:0005685 | U1 snRNP(GO:0005685) |
| 0.0 | 0.5 | GO:0031095 | platelet dense tubular network membrane(GO:0031095) |
| 0.0 | 0.3 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.0 | 0.5 | GO:0033270 | paranode region of axon(GO:0033270) |
| 0.0 | 0.8 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.0 | 0.4 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.0 | 0.3 | GO:0097433 | dense body(GO:0097433) |
| 0.0 | 0.6 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.0 | 1.0 | GO:0001917 | photoreceptor inner segment(GO:0001917) |
| 0.0 | 1.0 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 0.8 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.0 | 1.2 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 1.4 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 4.8 | GO:0035939 | microsatellite binding(GO:0035939) |
| 0.4 | 1.5 | GO:0070095 | fructose-6-phosphate binding(GO:0070095) |
| 0.3 | 1.2 | GO:0035402 | histone kinase activity (H3-T11 specific)(GO:0035402) |
| 0.2 | 0.9 | GO:0051734 | ATP-dependent polydeoxyribonucleotide 5'-hydroxyl-kinase activity(GO:0046404) polydeoxyribonucleotide kinase activity(GO:0051733) ATP-dependent polynucleotide kinase activity(GO:0051734) |
| 0.2 | 5.9 | GO:0032794 | GTPase activating protein binding(GO:0032794) |
| 0.2 | 0.8 | GO:0043891 | glyceraldehyde-3-phosphate dehydrogenase (NAD+) (phosphorylating) activity(GO:0004365) glyceraldehyde-3-phosphate dehydrogenase (NAD(P)+) (phosphorylating) activity(GO:0043891) |
| 0.2 | 0.7 | GO:0003990 | acetylcholinesterase activity(GO:0003990) |
| 0.2 | 0.5 | GO:0004998 | transferrin receptor activity(GO:0004998) |
| 0.2 | 0.6 | GO:0061628 | H3K27me3 modified histone binding(GO:0061628) |
| 0.2 | 0.5 | GO:0070052 | collagen V binding(GO:0070052) |
| 0.2 | 0.8 | GO:0035614 | snRNA stem-loop binding(GO:0035614) |
| 0.1 | 8.4 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.1 | 0.7 | GO:0070326 | very-low-density lipoprotein particle receptor binding(GO:0070326) |
| 0.1 | 0.5 | GO:0000822 | inositol hexakisphosphate binding(GO:0000822) |
| 0.1 | 0.5 | GO:0030226 | apolipoprotein receptor activity(GO:0030226) |
| 0.1 | 3.7 | GO:0047555 | 3',5'-cyclic-GMP phosphodiesterase activity(GO:0047555) |
| 0.1 | 0.5 | GO:0043515 | kinetochore binding(GO:0043515) |
| 0.1 | 0.3 | GO:0098918 | structural constituent of synapse(GO:0098918) structural constituent of postsynaptic actin cytoskeleton(GO:0098973) |
| 0.1 | 0.5 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.1 | 0.4 | GO:0070051 | fibrinogen binding(GO:0070051) |
| 0.1 | 0.9 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.1 | 0.3 | GO:0004886 | 9-cis retinoic acid receptor activity(GO:0004886) |
| 0.1 | 0.4 | GO:0004994 | somatostatin receptor activity(GO:0004994) |
| 0.1 | 0.5 | GO:0050815 | phosphoserine binding(GO:0050815) |
| 0.1 | 8.7 | GO:0005518 | collagen binding(GO:0005518) |
| 0.1 | 0.7 | GO:0004128 | cytochrome-b5 reductase activity, acting on NAD(P)H(GO:0004128) |
| 0.1 | 0.9 | GO:0004559 | alpha-mannosidase activity(GO:0004559) |
| 0.1 | 1.4 | GO:0016641 | oxidoreductase activity, acting on the CH-NH2 group of donors, oxygen as acceptor(GO:0016641) |
| 0.1 | 1.2 | GO:0048185 | activin binding(GO:0048185) |
| 0.1 | 1.6 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 0.7 | GO:1903136 | cuprous ion binding(GO:1903136) |
| 0.0 | 1.7 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.0 | 0.3 | GO:0033192 | calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.0 | 0.8 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.0 | 1.9 | GO:0070888 | E-box binding(GO:0070888) |
| 0.0 | 0.3 | GO:0004972 | NMDA glutamate receptor activity(GO:0004972) |
| 0.0 | 0.3 | GO:0031545 | peptidyl-proline 4-dioxygenase activity(GO:0031545) |
| 0.0 | 1.3 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.0 | 0.3 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.0 | 0.4 | GO:0034236 | protein kinase A catalytic subunit binding(GO:0034236) |
| 0.0 | 1.7 | GO:0000979 | RNA polymerase II core promoter sequence-specific DNA binding(GO:0000979) |
| 0.0 | 0.4 | GO:0016712 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen(GO:0016712) |
| 0.0 | 1.0 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.0 | 1.6 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.1 | ST IL 13 PATHWAY | Interleukin 13 (IL-13) Pathway |
| 0.1 | 2.9 | ST INTERLEUKIN 4 PATHWAY | Interleukin 4 (IL-4) Pathway |
| 0.1 | 4.8 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.1 | 1.2 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.1 | 2.8 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.0 | 9.5 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 0.9 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.0 | 0.6 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.0 | 2.2 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.0 | 0.5 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.0 | 1.2 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 1.1 | SIG BCR SIGNALING PATHWAY | Members of the BCR signaling pathway |
| 0.0 | 2.1 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 0.4 | PID INTEGRIN CS PATHWAY | Integrin family cell surface interactions |
| 0.0 | 0.9 | PID TNF PATHWAY | TNF receptor signaling pathway |
| 0.0 | 0.5 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.0 | 0.8 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 0.3 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.0 | 0.7 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.7 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.1 | 0.3 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.1 | 0.6 | REACTOME IRAK2 MEDIATED ACTIVATION OF TAK1 COMPLEX UPON TLR7 8 OR 9 STIMULATION | Genes involved in IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation |
| 0.1 | 0.5 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.1 | 4.8 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.1 | 1.4 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.0 | 1.2 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.0 | 2.1 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 1.5 | REACTOME REGULATION OF GLUCOKINASE BY GLUCOKINASE REGULATORY PROTEIN | Genes involved in Regulation of Glucokinase by Glucokinase Regulatory Protein |
| 0.0 | 0.5 | REACTOME RAF MAP KINASE CASCADE | Genes involved in RAF/MAP kinase cascade |
| 0.0 | 1.2 | REACTOME INHIBITION OF THE PROTEOLYTIC ACTIVITY OF APC C REQUIRED FOR THE ONSET OF ANAPHASE BY MITOTIC SPINDLE CHECKPOINT COMPONENTS | Genes involved in Inhibition of the proteolytic activity of APC/C required for the onset of anaphase by mitotic spindle checkpoint components |
| 0.0 | 0.3 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.0 | 1.9 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 3.3 | REACTOME DOWNSTREAM SIGNAL TRANSDUCTION | Genes involved in Downstream signal transduction |
| 0.0 | 0.7 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.0 | 0.4 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 0.0 | 0.8 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.0 | 0.4 | REACTOME GRB2 SOS PROVIDES LINKAGE TO MAPK SIGNALING FOR INTERGRINS | Genes involved in GRB2:SOS provides linkage to MAPK signaling for Intergrins |
| 0.0 | 0.9 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.0 | 0.5 | REACTOME TRANSFERRIN ENDOCYTOSIS AND RECYCLING | Genes involved in Transferrin endocytosis and recycling |
| 0.0 | 0.8 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.0 | 0.4 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |