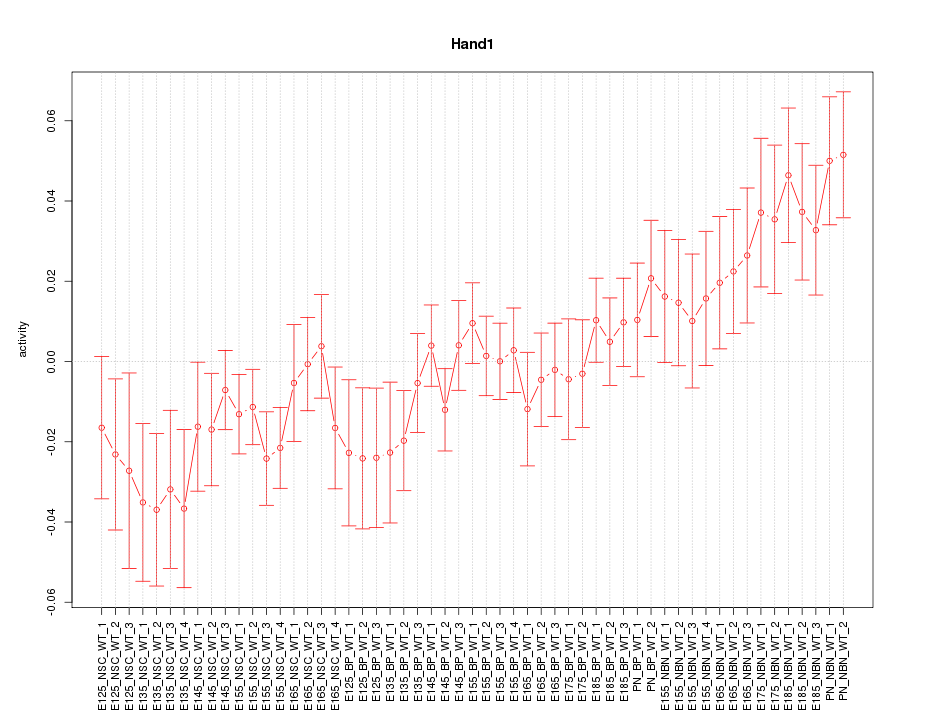
Motif ID: Hand1
Z-value: 1.382
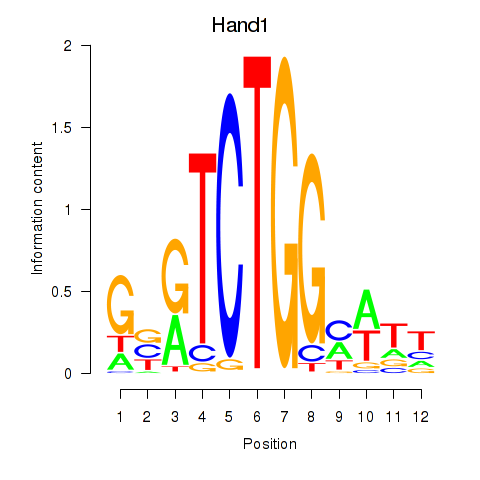
Transcription factors associated with Hand1:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| Hand1 | ENSMUSG00000037335.7 | Hand1 |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.0 | 9.1 | GO:0071873 | response to norepinephrine(GO:0071873) |
| 3.0 | 12.0 | GO:0042138 | meiotic DNA double-strand break formation(GO:0042138) |
| 2.9 | 8.6 | GO:0072378 | blood coagulation, fibrin clot formation(GO:0072378) |
| 2.2 | 6.6 | GO:0050915 | sensory perception of sour taste(GO:0050915) |
| 2.1 | 10.4 | GO:1905169 | protein localization to phagocytic vesicle(GO:1905161) regulation of protein localization to phagocytic vesicle(GO:1905169) positive regulation of protein localization to phagocytic vesicle(GO:1905171) |
| 2.0 | 6.1 | GO:0002030 | inhibitory G-protein coupled receptor phosphorylation(GO:0002030) |
| 1.9 | 5.7 | GO:1903054 | negative regulation of extracellular matrix organization(GO:1903054) |
| 1.7 | 5.2 | GO:0051715 | cytolysis in other organism(GO:0051715) |
| 1.6 | 4.9 | GO:1902256 | apoptotic process involved in outflow tract morphogenesis(GO:0003275) regulation of apoptotic process involved in outflow tract morphogenesis(GO:1902256) response to chemokine(GO:1990868) cellular response to chemokine(GO:1990869) |
| 1.6 | 4.9 | GO:0031117 | positive regulation of microtubule depolymerization(GO:0031117) |
| 1.5 | 9.2 | GO:0097646 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 1.5 | 7.6 | GO:2001025 | positive regulation of response to drug(GO:2001025) |
| 1.5 | 9.1 | GO:0046880 | regulation of follicle-stimulating hormone secretion(GO:0046880) follicle-stimulating hormone secretion(GO:0046884) |
| 1.5 | 4.5 | GO:1904339 | negative regulation of dopaminergic neuron differentiation(GO:1904339) |
| 1.3 | 6.3 | GO:0098735 | cellular response to caffeine(GO:0071313) positive regulation of the force of heart contraction(GO:0098735) |
| 1.2 | 2.5 | GO:2001023 | regulation of response to drug(GO:2001023) |
| 1.2 | 3.7 | GO:0014877 | response to inactivity(GO:0014854) response to muscle inactivity(GO:0014870) response to muscle inactivity involved in regulation of muscle adaptation(GO:0014877) response to denervation involved in regulation of muscle adaptation(GO:0014894) |
| 1.2 | 3.5 | GO:0043553 | negative regulation of phosphatidylinositol 3-kinase activity(GO:0043553) |
| 1.1 | 3.2 | GO:0030321 | transepithelial chloride transport(GO:0030321) |
| 1.0 | 2.9 | GO:0086046 | membrane depolarization during SA node cell action potential(GO:0086046) |
| 1.0 | 7.8 | GO:0097118 | neuroligin clustering involved in postsynaptic membrane assembly(GO:0097118) |
| 0.9 | 13.1 | GO:0097091 | synaptic vesicle clustering(GO:0097091) |
| 0.9 | 4.3 | GO:0032489 | regulation of Cdc42 protein signal transduction(GO:0032489) |
| 0.8 | 2.5 | GO:0002578 | negative regulation of antigen processing and presentation(GO:0002578) |
| 0.8 | 2.3 | GO:1901608 | regulation of vesicle transport along microtubule(GO:1901608) |
| 0.8 | 3.8 | GO:0072318 | clathrin coat disassembly(GO:0072318) |
| 0.8 | 3.8 | GO:2000427 | eosinophil chemotaxis(GO:0048245) T cell extravasation(GO:0072683) positive regulation of apoptotic cell clearance(GO:2000427) |
| 0.8 | 5.3 | GO:0070494 | regulation of thrombin receptor signaling pathway(GO:0070494) negative regulation of thrombin receptor signaling pathway(GO:0070495) |
| 0.7 | 6.0 | GO:0031666 | positive regulation of lipopolysaccharide-mediated signaling pathway(GO:0031666) |
| 0.7 | 6.6 | GO:0071420 | cellular response to histamine(GO:0071420) |
| 0.7 | 7.8 | GO:1900113 | negative regulation of histone H3-K9 trimethylation(GO:1900113) |
| 0.7 | 6.3 | GO:2000253 | positive regulation of feeding behavior(GO:2000253) |
| 0.7 | 2.1 | GO:0035790 | platelet-derived growth factor receptor-alpha signaling pathway(GO:0035790) |
| 0.7 | 2.1 | GO:0007525 | somatic muscle development(GO:0007525) |
| 0.7 | 4.8 | GO:0061304 | retinal blood vessel morphogenesis(GO:0061304) |
| 0.7 | 4.8 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.7 | 2.0 | GO:0071544 | diphosphoinositol polyphosphate catabolic process(GO:0071544) |
| 0.6 | 2.5 | GO:0002101 | tRNA wobble cytosine modification(GO:0002101) |
| 0.5 | 3.2 | GO:0048597 | post-embryonic camera-type eye morphogenesis(GO:0048597) |
| 0.5 | 4.1 | GO:0097369 | sodium ion import(GO:0097369) |
| 0.5 | 3.4 | GO:0033227 | dsRNA transport(GO:0033227) |
| 0.5 | 1.5 | GO:0009298 | GDP-mannose biosynthetic process(GO:0009298) |
| 0.5 | 1.4 | GO:0018199 | peptidyl-glutamine modification(GO:0018199) |
| 0.5 | 2.8 | GO:0051534 | negative regulation of NFAT protein import into nucleus(GO:0051534) |
| 0.5 | 3.2 | GO:0098703 | calcium ion import across plasma membrane(GO:0098703) calcium ion import into cell(GO:1990035) |
| 0.5 | 7.8 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.5 | 6.4 | GO:0033623 | regulation of integrin activation(GO:0033623) |
| 0.4 | 1.7 | GO:0010890 | positive regulation of sequestering of triglyceride(GO:0010890) |
| 0.4 | 1.7 | GO:0042985 | negative regulation of amyloid precursor protein biosynthetic process(GO:0042985) |
| 0.4 | 2.5 | GO:0061622 | glycolytic process through glucose-1-phosphate(GO:0061622) |
| 0.4 | 6.2 | GO:0038063 | collagen-activated tyrosine kinase receptor signaling pathway(GO:0038063) |
| 0.4 | 7.8 | GO:2000310 | regulation of N-methyl-D-aspartate selective glutamate receptor activity(GO:2000310) |
| 0.4 | 1.4 | GO:0006421 | asparaginyl-tRNA aminoacylation(GO:0006421) |
| 0.4 | 2.1 | GO:0032439 | endosome localization(GO:0032439) |
| 0.3 | 1.7 | GO:2000987 | positive regulation of fear response(GO:1903367) positive regulation of behavioral fear response(GO:2000987) |
| 0.3 | 2.7 | GO:0031642 | negative regulation of myelination(GO:0031642) |
| 0.3 | 4.9 | GO:1902260 | negative regulation of delayed rectifier potassium channel activity(GO:1902260) |
| 0.3 | 1.6 | GO:0048563 | post-embryonic organ morphogenesis(GO:0048563) |
| 0.3 | 1.6 | GO:0010724 | regulation of definitive erythrocyte differentiation(GO:0010724) |
| 0.3 | 1.3 | GO:0010796 | regulation of multivesicular body size(GO:0010796) |
| 0.3 | 1.8 | GO:0034227 | tRNA thio-modification(GO:0034227) |
| 0.3 | 9.5 | GO:0045838 | positive regulation of membrane potential(GO:0045838) |
| 0.3 | 1.4 | GO:0010991 | negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.3 | 5.8 | GO:0090036 | regulation of protein kinase C signaling(GO:0090036) |
| 0.3 | 1.5 | GO:0071955 | recycling endosome to Golgi transport(GO:0071955) |
| 0.2 | 1.2 | GO:0021631 | optic nerve morphogenesis(GO:0021631) |
| 0.2 | 4.0 | GO:0050995 | negative regulation of lipid catabolic process(GO:0050995) |
| 0.2 | 1.0 | GO:0044805 | late nucleophagy(GO:0044805) |
| 0.2 | 2.1 | GO:0035428 | hexose transmembrane transport(GO:0035428) |
| 0.2 | 4.0 | GO:0051044 | positive regulation of membrane protein ectodomain proteolysis(GO:0051044) |
| 0.2 | 0.7 | GO:0009644 | response to high light intensity(GO:0009644) |
| 0.2 | 2.6 | GO:0055003 | cardiac myofibril assembly(GO:0055003) |
| 0.2 | 2.1 | GO:0071569 | protein ufmylation(GO:0071569) |
| 0.2 | 2.0 | GO:0006686 | sphingomyelin biosynthetic process(GO:0006686) |
| 0.2 | 0.9 | GO:0006788 | heme oxidation(GO:0006788) |
| 0.2 | 5.8 | GO:0015991 | ATP hydrolysis coupled proton transport(GO:0015991) |
| 0.2 | 1.4 | GO:0098885 | modification of postsynaptic actin cytoskeleton(GO:0098885) |
| 0.2 | 2.1 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.2 | 0.4 | GO:0002925 | positive regulation of humoral immune response mediated by circulating immunoglobulin(GO:0002925) |
| 0.2 | 3.5 | GO:0006895 | Golgi to endosome transport(GO:0006895) |
| 0.2 | 1.2 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
| 0.2 | 9.4 | GO:0010765 | positive regulation of sodium ion transport(GO:0010765) |
| 0.2 | 1.5 | GO:0043312 | neutrophil degranulation(GO:0043312) |
| 0.2 | 3.4 | GO:2000251 | positive regulation of actin cytoskeleton reorganization(GO:2000251) |
| 0.2 | 0.5 | GO:0018094 | protein polyglycylation(GO:0018094) |
| 0.2 | 1.2 | GO:0034145 | positive regulation of toll-like receptor 4 signaling pathway(GO:0034145) |
| 0.2 | 1.3 | GO:0003094 | glomerular filtration(GO:0003094) |
| 0.2 | 5.5 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.2 | 1.6 | GO:0031115 | negative regulation of microtubule polymerization(GO:0031115) |
| 0.1 | 1.5 | GO:0000712 | resolution of meiotic recombination intermediates(GO:0000712) |
| 0.1 | 4.2 | GO:0016578 | histone deubiquitination(GO:0016578) |
| 0.1 | 5.4 | GO:0030199 | collagen fibril organization(GO:0030199) |
| 0.1 | 3.3 | GO:0006308 | DNA catabolic process(GO:0006308) |
| 0.1 | 20.4 | GO:0007416 | synapse assembly(GO:0007416) |
| 0.1 | 4.9 | GO:0021680 | cerebellar Purkinje cell layer development(GO:0021680) |
| 0.1 | 0.4 | GO:0009106 | lipoate metabolic process(GO:0009106) |
| 0.1 | 0.9 | GO:0034638 | phosphatidylcholine catabolic process(GO:0034638) |
| 0.1 | 2.7 | GO:0035036 | binding of sperm to zona pellucida(GO:0007339) sperm-egg recognition(GO:0035036) |
| 0.1 | 2.9 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.1 | 0.8 | GO:0097428 | protein maturation by iron-sulfur cluster transfer(GO:0097428) |
| 0.1 | 4.7 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.1 | 1.0 | GO:0090161 | Golgi ribbon formation(GO:0090161) |
| 0.1 | 7.8 | GO:0003073 | regulation of systemic arterial blood pressure(GO:0003073) |
| 0.1 | 0.9 | GO:0061734 | parkin-mediated mitophagy in response to mitochondrial depolarization(GO:0061734) |
| 0.1 | 0.4 | GO:0050812 | regulation of acetyl-CoA biosynthetic process from pyruvate(GO:0010510) regulation of acyl-CoA biosynthetic process(GO:0050812) |
| 0.1 | 1.9 | GO:0006359 | regulation of transcription from RNA polymerase III promoter(GO:0006359) |
| 0.1 | 1.1 | GO:0021869 | forebrain ventricular zone progenitor cell division(GO:0021869) |
| 0.1 | 0.6 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.1 | 0.8 | GO:0050765 | negative regulation of phagocytosis(GO:0050765) |
| 0.1 | 1.4 | GO:0060259 | regulation of feeding behavior(GO:0060259) |
| 0.1 | 0.5 | GO:0070562 | regulation of vitamin D receptor signaling pathway(GO:0070562) |
| 0.1 | 2.0 | GO:0097352 | autophagosome maturation(GO:0097352) |
| 0.1 | 1.7 | GO:0033006 | regulation of mast cell activation involved in immune response(GO:0033006) regulation of mast cell degranulation(GO:0043304) |
| 0.1 | 1.4 | GO:0042753 | positive regulation of circadian rhythm(GO:0042753) |
| 0.1 | 0.6 | GO:0006390 | transcription from mitochondrial promoter(GO:0006390) |
| 0.1 | 0.7 | GO:0031274 | positive regulation of pseudopodium assembly(GO:0031274) |
| 0.1 | 9.6 | GO:0006906 | vesicle fusion(GO:0006906) |
| 0.1 | 2.0 | GO:0046827 | positive regulation of protein export from nucleus(GO:0046827) |
| 0.1 | 2.1 | GO:0043954 | cellular component maintenance(GO:0043954) |
| 0.1 | 0.3 | GO:0032226 | positive regulation of synaptic transmission, dopaminergic(GO:0032226) |
| 0.1 | 0.7 | GO:0036159 | outer dynein arm assembly(GO:0036158) inner dynein arm assembly(GO:0036159) |
| 0.1 | 6.0 | GO:0007613 | memory(GO:0007613) |
| 0.1 | 3.6 | GO:0030071 | regulation of mitotic metaphase/anaphase transition(GO:0030071) regulation of metaphase/anaphase transition of cell cycle(GO:1902099) |
| 0.1 | 0.7 | GO:0045899 | positive regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045899) |
| 0.1 | 1.6 | GO:0090140 | regulation of mitochondrial fission(GO:0090140) |
| 0.1 | 0.5 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.1 | 1.4 | GO:2000785 | regulation of autophagosome assembly(GO:2000785) |
| 0.1 | 2.4 | GO:1904893 | negative regulation of JAK-STAT cascade(GO:0046426) negative regulation of STAT cascade(GO:1904893) |
| 0.1 | 1.8 | GO:0006829 | zinc II ion transport(GO:0006829) |
| 0.1 | 0.3 | GO:0006265 | DNA topological change(GO:0006265) |
| 0.1 | 1.2 | GO:0034308 | primary alcohol metabolic process(GO:0034308) |
| 0.1 | 9.2 | GO:1990830 | response to leukemia inhibitory factor(GO:1990823) cellular response to leukemia inhibitory factor(GO:1990830) |
| 0.1 | 1.3 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.1 | 0.4 | GO:0006451 | selenocysteine incorporation(GO:0001514) translational readthrough(GO:0006451) |
| 0.1 | 0.3 | GO:0032782 | bile acid secretion(GO:0032782) |
| 0.0 | 1.4 | GO:0009299 | mRNA transcription(GO:0009299) |
| 0.0 | 1.2 | GO:0032469 | endoplasmic reticulum calcium ion homeostasis(GO:0032469) |
| 0.0 | 3.8 | GO:0007601 | visual perception(GO:0007601) |
| 0.0 | 1.2 | GO:0010172 | embryonic body morphogenesis(GO:0010172) |
| 0.0 | 0.3 | GO:0031987 | locomotion involved in locomotory behavior(GO:0031987) |
| 0.0 | 2.5 | GO:0050885 | neuromuscular process controlling balance(GO:0050885) |
| 0.0 | 1.3 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.0 | 0.9 | GO:0015813 | L-glutamate transport(GO:0015813) |
| 0.0 | 0.6 | GO:0051220 | cytoplasmic sequestering of protein(GO:0051220) |
| 0.0 | 0.2 | GO:0071372 | cellular response to follicle-stimulating hormone stimulus(GO:0071372) |
| 0.0 | 1.4 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.0 | 2.9 | GO:0006888 | ER to Golgi vesicle-mediated transport(GO:0006888) |
| 0.0 | 1.3 | GO:0002028 | regulation of sodium ion transport(GO:0002028) |
| 0.0 | 2.3 | GO:0006998 | nuclear envelope organization(GO:0006998) |
| 0.0 | 1.9 | GO:0015908 | fatty acid transport(GO:0015908) |
| 0.0 | 3.2 | GO:0007605 | sensory perception of sound(GO:0007605) |
| 0.0 | 5.7 | GO:0031032 | actomyosin structure organization(GO:0031032) |
| 0.0 | 1.1 | GO:0043113 | receptor clustering(GO:0043113) |
| 0.0 | 0.6 | GO:0051457 | maintenance of protein location in nucleus(GO:0051457) |
| 0.0 | 1.9 | GO:0046324 | regulation of glucose import(GO:0046324) |
| 0.0 | 2.6 | GO:0043488 | regulation of mRNA stability(GO:0043488) |
| 0.0 | 0.4 | GO:0032757 | positive regulation of interleukin-8 production(GO:0032757) |
| 0.0 | 0.2 | GO:0051790 | short-chain fatty acid biosynthetic process(GO:0051790) |
| 0.0 | 2.0 | GO:0008016 | regulation of heart contraction(GO:0008016) |
| 0.0 | 2.2 | GO:0006487 | protein N-linked glycosylation(GO:0006487) |
| 0.0 | 0.4 | GO:0043457 | regulation of cellular respiration(GO:0043457) |
| 0.0 | 0.4 | GO:0030204 | chondroitin sulfate metabolic process(GO:0030204) |
| 0.0 | 0.6 | GO:0045103 | intermediate filament-based process(GO:0045103) |
| 0.0 | 0.2 | GO:0007076 | mitotic chromosome condensation(GO:0007076) |
| 0.0 | 2.6 | GO:0010506 | regulation of autophagy(GO:0010506) |
| 0.0 | 2.7 | GO:0042787 | protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0042787) |
| 0.0 | 0.6 | GO:1901799 | negative regulation of proteasomal protein catabolic process(GO:1901799) |
| 0.0 | 0.1 | GO:2000491 | positive regulation of hepatic stellate cell activation(GO:2000491) |
| 0.0 | 0.2 | GO:0044406 | adhesion of symbiont to host(GO:0044406) |
| 0.0 | 0.0 | GO:0031590 | wybutosine metabolic process(GO:0031590) wybutosine biosynthetic process(GO:0031591) |
| 0.0 | 1.3 | GO:0007050 | cell cycle arrest(GO:0007050) |
| 0.0 | 0.3 | GO:0070207 | protein homotrimerization(GO:0070207) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.0 | 9.1 | GO:0043512 | inhibin-betaglycan-ActRII complex(GO:0034673) inhibin A complex(GO:0043512) |
| 2.3 | 13.5 | GO:0044308 | axonal spine(GO:0044308) |
| 2.0 | 8.1 | GO:0099569 | presynaptic cytoskeleton(GO:0099569) |
| 1.8 | 9.2 | GO:1903440 | calcitonin family receptor complex(GO:1903439) amylin receptor complex(GO:1903440) |
| 1.8 | 9.2 | GO:0005587 | collagen type IV trimer(GO:0005587) network-forming collagen trimer(GO:0098642) collagen network(GO:0098645) basement membrane collagen trimer(GO:0098651) |
| 1.3 | 3.8 | GO:0044299 | C-fiber(GO:0044299) |
| 1.1 | 4.3 | GO:0097447 | dendritic tree(GO:0097447) |
| 1.0 | 5.0 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 1.0 | 2.9 | GO:0098855 | HCN channel complex(GO:0098855) |
| 0.9 | 7.8 | GO:0036056 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.8 | 12.5 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.6 | 6.9 | GO:0032426 | stereocilium tip(GO:0032426) |
| 0.6 | 2.5 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 0.6 | 9.1 | GO:1902711 | GABA-A receptor complex(GO:1902711) |
| 0.5 | 3.2 | GO:0097443 | sorting endosome(GO:0097443) |
| 0.5 | 3.6 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.5 | 2.5 | GO:1990357 | terminal web(GO:1990357) |
| 0.4 | 1.7 | GO:0032280 | symmetric synapse(GO:0032280) |
| 0.4 | 4.5 | GO:1990909 | Wnt signalosome(GO:1990909) |
| 0.4 | 2.7 | GO:0044352 | pinosome(GO:0044352) macropinosome(GO:0044354) |
| 0.4 | 5.5 | GO:0030122 | AP-2 adaptor complex(GO:0030122) |
| 0.4 | 4.9 | GO:0043203 | axon hillock(GO:0043203) |
| 0.3 | 4.8 | GO:0098644 | complex of collagen trimers(GO:0098644) |
| 0.3 | 3.4 | GO:0071439 | clathrin complex(GO:0071439) |
| 0.3 | 1.6 | GO:0002139 | stereocilia coupling link(GO:0002139) |
| 0.3 | 2.8 | GO:0031254 | uropod(GO:0001931) cell trailing edge(GO:0031254) |
| 0.3 | 1.4 | GO:1990130 | Iml1 complex(GO:1990130) |
| 0.3 | 2.1 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.3 | 7.8 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.2 | 13.1 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.2 | 4.2 | GO:0000124 | SAGA complex(GO:0000124) |
| 0.2 | 1.2 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.2 | 1.4 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.2 | 3.2 | GO:0005922 | connexon complex(GO:0005922) |
| 0.2 | 15.3 | GO:0031985 | Golgi cisterna(GO:0031985) |
| 0.2 | 2.4 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 0.2 | 1.3 | GO:0005828 | kinetochore microtubule(GO:0005828) |
| 0.2 | 8.5 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.2 | 1.8 | GO:0031091 | platelet alpha granule(GO:0031091) |
| 0.2 | 2.5 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.2 | 7.6 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.1 | 0.8 | GO:0031314 | extrinsic component of mitochondrial inner membrane(GO:0031314) |
| 0.1 | 2.1 | GO:0042555 | MCM complex(GO:0042555) |
| 0.1 | 1.2 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.1 | 0.4 | GO:0005967 | mitochondrial pyruvate dehydrogenase complex(GO:0005967) |
| 0.1 | 11.5 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.1 | 3.5 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.1 | 8.9 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.1 | 9.9 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.1 | 8.1 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.1 | 2.6 | GO:0005865 | striated muscle thin filament(GO:0005865) |
| 0.1 | 2.1 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.1 | 4.9 | GO:0005720 | nuclear heterochromatin(GO:0005720) |
| 0.1 | 1.0 | GO:0070971 | endoplasmic reticulum exit site(GO:0070971) |
| 0.1 | 4.4 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.1 | 0.5 | GO:0005726 | perichromatin fibrils(GO:0005726) |
| 0.1 | 2.9 | GO:0030134 | ER to Golgi transport vesicle(GO:0030134) |
| 0.1 | 0.9 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.1 | 1.9 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.1 | 0.8 | GO:0031931 | TORC1 complex(GO:0031931) |
| 0.1 | 1.4 | GO:0060170 | ciliary membrane(GO:0060170) |
| 0.1 | 1.2 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 0.1 | 0.3 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 0.1 | 6.4 | GO:0001725 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.1 | 0.7 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.1 | 0.7 | GO:0072669 | tRNA-splicing ligase complex(GO:0072669) |
| 0.1 | 2.1 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.1 | 1.6 | GO:0031514 | motile cilium(GO:0031514) |
| 0.1 | 0.6 | GO:0045298 | tubulin complex(GO:0045298) |
| 0.1 | 5.3 | GO:0036126 | sperm flagellum(GO:0036126) |
| 0.1 | 1.5 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.1 | 6.4 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.1 | 8.6 | GO:0098852 | lysosomal membrane(GO:0005765) lytic vacuole membrane(GO:0098852) |
| 0.0 | 0.2 | GO:0001939 | female pronucleus(GO:0001939) |
| 0.0 | 4.7 | GO:0055037 | recycling endosome(GO:0055037) |
| 0.0 | 6.9 | GO:0043197 | dendritic spine(GO:0043197) |
| 0.0 | 1.1 | GO:0030660 | Golgi-associated vesicle membrane(GO:0030660) |
| 0.0 | 1.3 | GO:0005697 | telomerase holoenzyme complex(GO:0005697) |
| 0.0 | 8.9 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 3.6 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.0 | 0.6 | GO:0097539 | ciliary transition fiber(GO:0097539) |
| 0.0 | 0.3 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.0 | 0.6 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.0 | 3.6 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 0.3 | GO:0000439 | core TFIIH complex(GO:0000439) |
| 0.0 | 0.3 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.0 | 2.6 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.0 | 0.2 | GO:0005832 | zona pellucida receptor complex(GO:0002199) chaperonin-containing T-complex(GO:0005832) |
| 0.0 | 0.7 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.0 | 1.2 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 1.7 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 1.1 | GO:0043679 | axon terminus(GO:0043679) |
| 0.0 | 1.0 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 5.4 | GO:0005764 | lytic vacuole(GO:0000323) lysosome(GO:0005764) |
| 0.0 | 0.3 | GO:0000153 | cytoplasmic ubiquitin ligase complex(GO:0000153) |
| 0.0 | 2.7 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.0 | 0.2 | GO:0020005 | symbiont-containing vacuole membrane(GO:0020005) |
| 0.0 | 0.6 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 0.4 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 0.7 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.0 | 2.3 | GO:0000139 | Golgi membrane(GO:0000139) |
| 0.0 | 1.6 | GO:0030141 | secretory granule(GO:0030141) |
| 0.0 | 1.1 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.0 | 1.1 | GO:0045211 | postsynaptic membrane(GO:0045211) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.0 | 12.0 | GO:0061505 | DNA topoisomerase type II (ATP-hydrolyzing) activity(GO:0003918) DNA topoisomerase II activity(GO:0061505) |
| 2.3 | 9.1 | GO:0031694 | alpha-2A adrenergic receptor binding(GO:0031694) |
| 2.1 | 8.6 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 1.8 | 9.2 | GO:0097643 | amylin receptor activity(GO:0097643) |
| 1.6 | 4.9 | GO:0004993 | G-protein coupled serotonin receptor activity(GO:0004993) |
| 1.6 | 9.5 | GO:1904315 | transmitter-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1904315) |
| 1.6 | 6.3 | GO:0086038 | calcium:sodium antiporter activity involved in regulation of cardiac muscle cell membrane potential(GO:0086038) |
| 1.4 | 5.6 | GO:0042834 | peptidoglycan binding(GO:0042834) |
| 1.3 | 5.3 | GO:0005008 | hepatocyte growth factor-activated receptor activity(GO:0005008) |
| 1.3 | 9.1 | GO:0034711 | inhibin binding(GO:0034711) |
| 1.3 | 7.8 | GO:0070699 | type II activin receptor binding(GO:0070699) |
| 1.2 | 4.7 | GO:0004720 | protein-lysine 6-oxidase activity(GO:0004720) |
| 1.2 | 3.5 | GO:0036313 | phosphatidylinositol 3-kinase catalytic subunit binding(GO:0036313) |
| 1.1 | 3.3 | GO:0003858 | 3-hydroxybutyrate dehydrogenase activity(GO:0003858) |
| 1.0 | 5.2 | GO:0038085 | vascular endothelial growth factor binding(GO:0038085) |
| 1.0 | 4.9 | GO:0003945 | N-acetyllactosamine synthase activity(GO:0003945) |
| 1.0 | 3.8 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.9 | 3.4 | GO:0051032 | nucleic acid transmembrane transporter activity(GO:0051032) RNA transmembrane transporter activity(GO:0051033) |
| 0.7 | 2.9 | GO:0005222 | intracellular cAMP activated cation channel activity(GO:0005222) |
| 0.6 | 2.6 | GO:0022851 | GABA-gated chloride ion channel activity(GO:0022851) |
| 0.6 | 2.5 | GO:0003872 | 6-phosphofructokinase activity(GO:0003872) |
| 0.5 | 6.5 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.5 | 2.7 | GO:0001032 | RNA polymerase III type 3 promoter DNA binding(GO:0001032) |
| 0.5 | 1.6 | GO:0052658 | inositol-1,4,5-trisphosphate 5-phosphatase activity(GO:0052658) |
| 0.5 | 1.6 | GO:0070139 | ubiquitin-like protein-specific endopeptidase activity(GO:0070137) SUMO-specific endopeptidase activity(GO:0070139) |
| 0.5 | 4.1 | GO:0015386 | potassium:proton antiporter activity(GO:0015386) |
| 0.5 | 5.0 | GO:0019865 | immunoglobulin binding(GO:0019865) |
| 0.5 | 1.9 | GO:0004321 | fatty-acyl-CoA synthase activity(GO:0004321) |
| 0.5 | 1.4 | GO:0004816 | asparagine-tRNA ligase activity(GO:0004816) |
| 0.5 | 3.2 | GO:0045028 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.5 | 6.4 | GO:0031005 | filamin binding(GO:0031005) |
| 0.5 | 10.4 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.4 | 7.8 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.4 | 2.0 | GO:0033188 | sphingomyelin synthase activity(GO:0033188) ceramide cholinephosphotransferase activity(GO:0047493) |
| 0.4 | 2.4 | GO:0004065 | arylsulfatase activity(GO:0004065) |
| 0.4 | 3.1 | GO:0070290 | N-acylphosphatidylethanolamine-specific phospholipase D activity(GO:0070290) |
| 0.4 | 4.8 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.4 | 2.9 | GO:0032051 | clathrin light chain binding(GO:0032051) |
| 0.3 | 2.4 | GO:0031821 | G-protein coupled serotonin receptor binding(GO:0031821) |
| 0.3 | 2.0 | GO:0098519 | nucleotide phosphatase activity, acting on free nucleotides(GO:0098519) |
| 0.3 | 2.0 | GO:0000298 | endopolyphosphatase activity(GO:0000298) diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) bis(5'-adenosyl)-hexaphosphatase activity(GO:0034431) bis(5'-adenosyl)-pentaphosphatase activity(GO:0034432) |
| 0.3 | 3.3 | GO:0016888 | endodeoxyribonuclease activity, producing 5'-phosphomonoesters(GO:0016888) |
| 0.3 | 4.9 | GO:0004033 | aldo-keto reductase (NADP) activity(GO:0004033) |
| 0.3 | 3.6 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.3 | 6.6 | GO:0005248 | voltage-gated sodium channel activity(GO:0005248) |
| 0.3 | 5.3 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.3 | 2.5 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.3 | 0.8 | GO:0019776 | Atg8 ligase activity(GO:0019776) |
| 0.3 | 6.8 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.3 | 4.5 | GO:0034236 | protein kinase A catalytic subunit binding(GO:0034236) |
| 0.2 | 1.7 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.2 | 7.8 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.2 | 0.9 | GO:0004392 | heme oxygenase (decyclizing) activity(GO:0004392) |
| 0.2 | 4.4 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.2 | 4.3 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.2 | 18.5 | GO:0035254 | glutamate receptor binding(GO:0035254) |
| 0.2 | 4.7 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.2 | 0.5 | GO:0070736 | protein-glycine ligase activity(GO:0070735) protein-glycine ligase activity, initiating(GO:0070736) |
| 0.2 | 3.5 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.2 | 2.1 | GO:0004622 | lysophospholipase activity(GO:0004622) |
| 0.2 | 0.6 | GO:0004859 | phospholipase inhibitor activity(GO:0004859) phospholipase A2 inhibitor activity(GO:0019834) |
| 0.2 | 1.4 | GO:0016755 | transferase activity, transferring amino-acyl groups(GO:0016755) |
| 0.1 | 4.9 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.1 | 1.8 | GO:0016783 | sulfurtransferase activity(GO:0016783) |
| 0.1 | 1.6 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.1 | 3.0 | GO:0005351 | sugar:proton symporter activity(GO:0005351) |
| 0.1 | 1.5 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.1 | 1.5 | GO:0016868 | intramolecular transferase activity, phosphotransferases(GO:0016868) |
| 0.1 | 1.2 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.1 | 1.4 | GO:0016500 | protein-hormone receptor activity(GO:0016500) |
| 0.1 | 2.1 | GO:0097371 | MDM2/MDM4 family protein binding(GO:0097371) |
| 0.1 | 1.2 | GO:0015271 | outward rectifier potassium channel activity(GO:0015271) |
| 0.1 | 0.4 | GO:0004740 | pyruvate dehydrogenase (acetyl-transferring) kinase activity(GO:0004740) |
| 0.1 | 2.8 | GO:0008376 | acetylgalactosaminyltransferase activity(GO:0008376) |
| 0.1 | 0.5 | GO:0034056 | estrogen response element binding(GO:0034056) |
| 0.1 | 1.0 | GO:0008556 | sodium:potassium-exchanging ATPase activity(GO:0005391) potassium-transporting ATPase activity(GO:0008556) |
| 0.1 | 6.0 | GO:0030276 | clathrin binding(GO:0030276) |
| 0.1 | 6.1 | GO:0048365 | Rac GTPase binding(GO:0048365) |
| 0.1 | 6.0 | GO:0005518 | collagen binding(GO:0005518) |
| 0.1 | 1.2 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.1 | 1.2 | GO:0036442 | hydrogen-exporting ATPase activity(GO:0036442) proton-transporting ATPase activity, rotational mechanism(GO:0046961) |
| 0.1 | 1.9 | GO:0043422 | protein kinase B binding(GO:0043422) |
| 0.1 | 1.4 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.1 | 6.2 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.1 | 4.2 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.1 | 0.9 | GO:0005314 | high-affinity glutamate transmembrane transporter activity(GO:0005314) |
| 0.1 | 1.8 | GO:0097602 | cullin family protein binding(GO:0097602) |
| 0.1 | 1.8 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.1 | 1.5 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.1 | 0.4 | GO:0015018 | galactosylgalactosylxylosylprotein 3-beta-glucuronosyltransferase activity(GO:0015018) |
| 0.1 | 2.9 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 2.2 | GO:0017048 | Rho GTPase binding(GO:0017048) |
| 0.1 | 9.7 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.1 | 1.3 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.1 | 4.5 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.1 | 0.3 | GO:0043138 | 3'-5' DNA helicase activity(GO:0043138) |
| 0.1 | 1.4 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.1 | 2.7 | GO:0030145 | manganese ion binding(GO:0030145) |
| 0.1 | 0.7 | GO:0005225 | volume-sensitive anion channel activity(GO:0005225) |
| 0.1 | 2.1 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.1 | 0.3 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.1 | 0.3 | GO:0033192 | calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.0 | 8.6 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.0 | 1.4 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.0 | 4.0 | GO:0052689 | carboxylic ester hydrolase activity(GO:0052689) |
| 0.0 | 1.2 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.0 | 5.7 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.0 | 1.1 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.0 | 0.2 | GO:0004315 | 3-oxoacyl-[acyl-carrier-protein] synthase activity(GO:0004315) |
| 0.0 | 2.1 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.0 | 2.4 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.0 | 1.7 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 0.8 | GO:0008198 | ferrous iron binding(GO:0008198) |
| 0.0 | 1.4 | GO:0045309 | protein phosphorylated amino acid binding(GO:0045309) |
| 0.0 | 1.6 | GO:0000049 | tRNA binding(GO:0000049) |
| 0.0 | 1.0 | GO:0001104 | RNA polymerase II transcription cofactor activity(GO:0001104) |
| 0.0 | 2.4 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.0 | 0.2 | GO:0005004 | GPI-linked ephrin receptor activity(GO:0005004) |
| 0.0 | 2.1 | GO:0030674 | protein binding, bridging(GO:0030674) |
| 0.0 | 12.1 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 0.0 | 1.4 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.0 | 1.0 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.0 | 0.7 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.0 | 0.2 | GO:0034237 | protein kinase A regulatory subunit binding(GO:0034237) |
| 0.0 | 2.1 | GO:0051020 | GTPase binding(GO:0051020) |
| 0.0 | 0.7 | GO:0004521 | endoribonuclease activity(GO:0004521) |
| 0.0 | 4.8 | GO:0003779 | actin binding(GO:0003779) |
| 0.0 | 1.0 | GO:0005262 | calcium channel activity(GO:0005262) |
| 0.0 | 1.5 | GO:0004386 | helicase activity(GO:0004386) |
| 0.0 | 1.2 | GO:0005539 | glycosaminoglycan binding(GO:0005539) |
| 0.0 | 0.3 | GO:1990782 | protein tyrosine kinase binding(GO:1990782) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 5.4 | PID_RHODOPSIN_PATHWAY | Visual signal transduction: Rods |
| 0.4 | 3.2 | PID_VEGF_VEGFR_PATHWAY | VEGF and VEGFR signaling network |
| 0.4 | 7.8 | ST_G_ALPHA_S_PATHWAY | G alpha s Pathway |
| 0.3 | 9.4 | PID_INTEGRIN_A9B1_PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.3 | 10.9 | NABA_PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.3 | 9.2 | NABA_BASEMENT_MEMBRANES | Genes encoding structural components of basement membranes |
| 0.2 | 2.8 | PID_INTEGRIN2_PATHWAY | Beta2 integrin cell surface interactions |
| 0.2 | 2.0 | PID_S1P_S1P4_PATHWAY | S1P4 pathway |
| 0.2 | 10.9 | PID_EPHB_FWD_PATHWAY | EPHB forward signaling |
| 0.2 | 3.8 | PID_IL23_PATHWAY | IL23-mediated signaling events |
| 0.2 | 4.5 | PID_BETA_CATENIN_DEG_PATHWAY | Degradation of beta catenin |
| 0.1 | 5.7 | PID_ARF6_PATHWAY | Arf6 signaling events |
| 0.1 | 6.1 | SIG_REGULATION_OF_THE_ACTIN_CYTOSKELETON_BY_RHO_GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.1 | 4.4 | PID_MAPK_TRK_PATHWAY | Trk receptor signaling mediated by the MAPK pathway |
| 0.1 | 3.1 | ST_GA12_PATHWAY | G alpha 12 Pathway |
| 0.1 | 2.1 | PID_PDGFRA_PATHWAY | PDGFR-alpha signaling pathway |
| 0.1 | 3.2 | PID_PTP1B_PATHWAY | Signaling events mediated by PTP1B |
| 0.1 | 3.4 | PID_HDAC_CLASSIII_PATHWAY | Signaling events mediated by HDAC Class III |
| 0.1 | 3.8 | PID_CDC42_REG_PATHWAY | Regulation of CDC42 activity |
| 0.1 | 1.3 | PID_ARF6_DOWNSTREAM_PATHWAY | Arf6 downstream pathway |
| 0.1 | 2.9 | PID_IL2_PI3K_PATHWAY | IL2 signaling events mediated by PI3K |
| 0.1 | 2.6 | PID_FAK_PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.1 | 3.7 | PID_FOXO_PATHWAY | FoxO family signaling |
| 0.1 | 3.3 | PID_HNF3B_PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 2.1 | PID_ATR_PATHWAY | ATR signaling pathway |
| 0.1 | 3.9 | PID_NOTCH_PATHWAY | Notch signaling pathway |
| 0.1 | 2.8 | PID_P53_REGULATION_PATHWAY | p53 pathway |
| 0.0 | 8.5 | NABA_SECRETED_FACTORS | Genes encoding secreted soluble factors |
| 0.0 | 0.5 | PID_IL12_STAT4_PATHWAY | IL12 signaling mediated by STAT4 |
| 0.0 | 1.9 | PID_AR_TF_PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 1.2 | PID_ECADHERIN_STABILIZATION_PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.0 | 0.4 | PID_NFKAPPAB_CANONICAL_PATHWAY | Canonical NF-kappaB pathway |
| 0.0 | 0.6 | PID_BARD1_PATHWAY | BARD1 signaling events |
| 0.0 | 0.2 | PID_EPHA_FWDPATHWAY | EPHA forward signaling |
| 0.0 | 1.0 | PID_HDAC_CLASSI_PATHWAY | Signaling events mediated by HDAC Class I |
| 0.0 | 2.0 | NABA_ECM_REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.2 | PID_FANCONI_PATHWAY | Fanconi anemia pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.3 | 9.1 | REACTOME_GLYCOPROTEIN_HORMONES | Genes involved in Glycoprotein hormones |
| 1.6 | 4.9 | REACTOME_SEROTONIN_RECEPTORS | Genes involved in Serotonin receptors |
| 1.4 | 8.6 | REACTOME_COMMON_PATHWAY | Genes involved in Common Pathway |
| 0.8 | 12.9 | REACTOME_GABA_A_RECEPTOR_ACTIVATION | Genes involved in GABA A receptor activation |
| 0.6 | 5.0 | REACTOME_INTRINSIC_PATHWAY | Genes involved in Intrinsic Pathway |
| 0.5 | 6.3 | REACTOME_PLATELET_CALCIUM_HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.4 | 9.5 | REACTOME_TRAFFICKING_OF_GLUR2_CONTAINING_AMPA_RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 0.4 | 3.2 | REACTOME_P2Y_RECEPTORS | Genes involved in P2Y receptors |
| 0.3 | 2.4 | REACTOME_THE_ACTIVATION_OF_ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.3 | 4.8 | REACTOME_CS_DS_DEGRADATION | Genes involved in CS/DS degradation |
| 0.3 | 5.0 | REACTOME_ACTIVATION_OF_RAC | Genes involved in Activation of Rac |
| 0.3 | 5.6 | REACTOME_IONOTROPIC_ACTIVITY_OF_KAINATE_RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.3 | 7.8 | REACTOME_NEPHRIN_INTERACTIONS | Genes involved in Nephrin interactions |
| 0.3 | 4.5 | REACTOME_POST_CHAPERONIN_TUBULIN_FOLDING_PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.3 | 2.7 | REACTOME_FACILITATIVE_NA_INDEPENDENT_GLUCOSE_TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.3 | 4.3 | REACTOME_SYNTHESIS_OF_SUBSTRATES_IN_N_GLYCAN_BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.3 | 3.2 | REACTOME_GAP_JUNCTION_ASSEMBLY | Genes involved in Gap junction assembly |
| 0.3 | 3.2 | REACTOME_VEGF_LIGAND_RECEPTOR_INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.3 | 4.9 | REACTOME_N_GLYCAN_ANTENNAE_ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.2 | 5.7 | REACTOME_RAS_ACTIVATION_UOPN_CA2_INFUX_THROUGH_NMDA_RECEPTOR | Genes involved in Ras activation uopn Ca2+ infux through NMDA receptor |
| 0.2 | 9.1 | REACTOME_ACTIVATED_NOTCH1_TRANSMITS_SIGNAL_TO_THE_NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.2 | 4.3 | REACTOME_SIGNALLING_TO_P38_VIA_RIT_AND_RIN | Genes involved in Signalling to p38 via RIT and RIN |
| 0.2 | 4.8 | REACTOME_INSULIN_RECEPTOR_RECYCLING | Genes involved in Insulin receptor recycling |
| 0.2 | 4.0 | REACTOME_CELL_EXTRACELLULAR_MATRIX_INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.2 | 5.7 | REACTOME_ION_TRANSPORT_BY_P_TYPE_ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.2 | 10.7 | REACTOME_GOLGI_ASSOCIATED_VESICLE_BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.2 | 3.8 | REACTOME_ACTIVATION_OF_GENES_BY_ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.1 | 3.0 | REACTOME_IMMUNOREGULATORY_INTERACTIONS_BETWEEN_A_LYMPHOID_AND_A_NON_LYMPHOID_CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.1 | 1.9 | REACTOME_ROLE_OF_DCC_IN_REGULATING_APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.1 | 8.8 | REACTOME_COLLAGEN_FORMATION | Genes involved in Collagen formation |
| 0.1 | 4.5 | REACTOME_TIGHT_JUNCTION_INTERACTIONS | Genes involved in Tight junction interactions |
| 0.1 | 1.4 | REACTOME_PROLACTIN_RECEPTOR_SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.1 | 2.1 | REACTOME_UNWINDING_OF_DNA | Genes involved in Unwinding of DNA |
| 0.1 | 4.8 | REACTOME_VOLTAGE_GATED_POTASSIUM_CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 1.8 | REACTOME_ZINC_TRANSPORTERS | Genes involved in Zinc transporters |
| 0.1 | 8.1 | REACTOME_G_ALPHA_S_SIGNALLING_EVENTS | Genes involved in G alpha (s) signalling events |
| 0.1 | 3.8 | REACTOME_ASSOCIATION_OF_TRIC_CCT_WITH_TARGET_PROTEINS_DURING_BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.1 | 3.2 | REACTOME_G_PROTEIN_ACTIVATION | Genes involved in G-protein activation |
| 0.1 | 2.9 | REACTOME_POTASSIUM_CHANNELS | Genes involved in Potassium Channels |
| 0.1 | 0.9 | REACTOME_IRON_UPTAKE_AND_TRANSPORT | Genes involved in Iron uptake and transport |
| 0.1 | 1.9 | REACTOME_RNA_POL_III_TRANSCRIPTION_INITIATION_FROM_TYPE_3_PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 3 Promoter |
| 0.1 | 2.5 | REACTOME_GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 1.3 | REACTOME_REGULATION_OF_WATER_BALANCE_BY_RENAL_AQUAPORINS | Genes involved in Regulation of Water Balance by Renal Aquaporins |
| 0.1 | 1.6 | REACTOME_SYNTHESIS_OF_PIPS_AT_THE_PLASMA_MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.1 | 6.5 | REACTOME_TOLL_RECEPTOR_CASCADES | Genes involved in Toll Receptor Cascades |
| 0.1 | 3.6 | REACTOME_G_ALPHA_I_SIGNALLING_EVENTS | Genes involved in G alpha (i) signalling events |
| 0.1 | 0.8 | REACTOME_MTORC1_MEDIATED_SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.0 | 2.0 | REACTOME_SPHINGOLIPID_DE_NOVO_BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.9 | REACTOME_FANCONI_ANEMIA_PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.0 | 3.4 | REACTOME_SIGNALING_BY_PDGF | Genes involved in Signaling by PDGF |
| 0.0 | 3.2 | REACTOME_MHC_CLASS_II_ANTIGEN_PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 0.3 | REACTOME_DARPP_32_EVENTS | Genes involved in DARPP-32 events |
| 0.0 | 0.4 | REACTOME_REGULATION_OF_PYRUVATE_DEHYDROGENASE_PDH_COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.0 | 3.1 | REACTOME_GLYCEROPHOSPHOLIPID_BIOSYNTHESIS | Genes involved in Glycerophospholipid biosynthesis |
| 0.0 | 0.5 | REACTOME_SIGNALING_BY_FGFR1_FUSION_MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.0 | 0.4 | REACTOME_METABOLISM_OF_POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.0 | 0.8 | REACTOME_SIGNALING_BY_EGFR_IN_CANCER | Genes involved in Signaling by EGFR in Cancer |
| 0.0 | 1.5 | REACTOME_FACTORS_INVOLVED_IN_MEGAKARYOCYTE_DEVELOPMENT_AND_PLATELET_PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.0 | 1.1 | REACTOME_AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 0.9 | REACTOME_RNA_POL_I_RNA_POL_III_AND_MITOCHONDRIAL_TRANSCRIPTION | Genes involved in RNA Polymerase I, RNA Polymerase III, and Mitochondrial Transcription |
| 0.0 | 0.4 | REACTOME_A_TETRASACCHARIDE_LINKER_SEQUENCE_IS_REQUIRED_FOR_GAG_SYNTHESIS | Genes involved in A tetrasaccharide linker sequence is required for GAG synthesis |
| 0.0 | 3.1 | REACTOME_ANTIGEN_PROCESSING_UBIQUITINATION_PROTEASOME_DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |