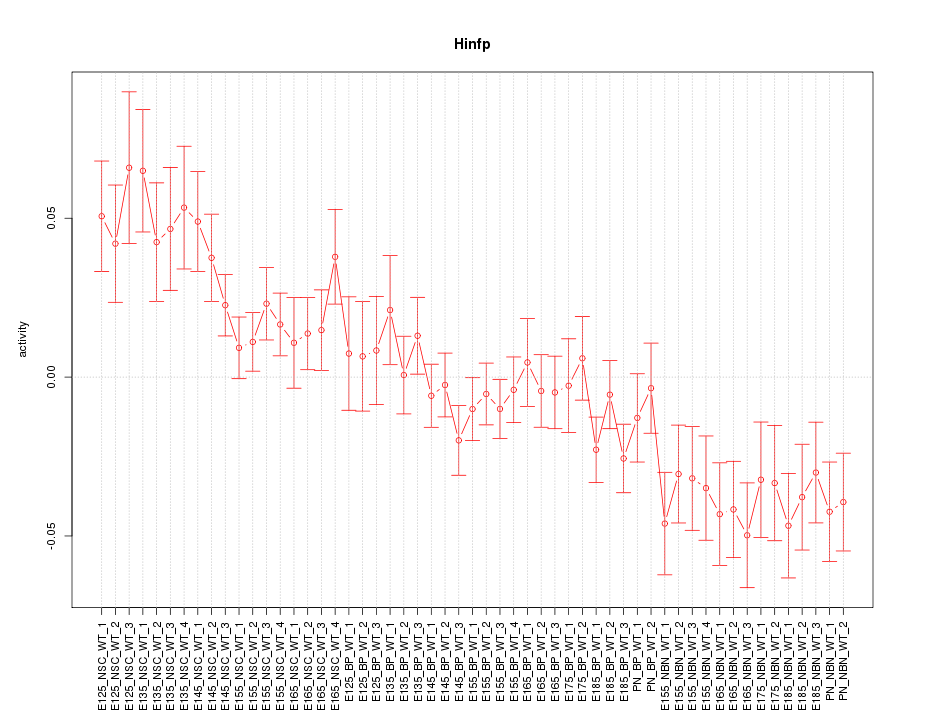
Motif ID: Hinfp
Z-value: 1.890
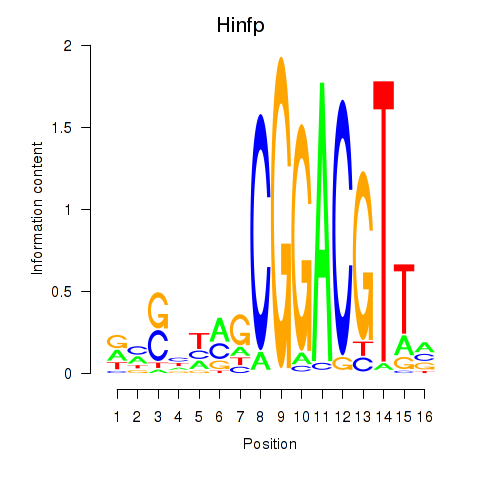
Transcription factors associated with Hinfp:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| Hinfp | ENSMUSG00000032119.4 | Hinfp |
Activity-expression correlation:
| Gene Symbol | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Hinfp | mm10_v2_chr9_-_44305595_44305688 | -0.32 | 1.6e-02 | Click! |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.7 | 23.1 | GO:0002302 | CD8-positive, alpha-beta T cell differentiation involved in immune response(GO:0002302) |
| 4.5 | 13.4 | GO:0030421 | defecation(GO:0030421) |
| 4.4 | 17.5 | GO:2000313 | fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:0060825) regulation of fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:2000313) |
| 4.0 | 31.8 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 3.8 | 11.4 | GO:0035519 | protein K29-linked ubiquitination(GO:0035519) |
| 3.7 | 18.7 | GO:0007089 | traversing start control point of mitotic cell cycle(GO:0007089) |
| 3.1 | 12.4 | GO:0040038 | polar body extrusion after meiotic divisions(GO:0040038) |
| 3.0 | 12.1 | GO:0006014 | D-ribose metabolic process(GO:0006014) |
| 2.8 | 8.4 | GO:0070315 | G1 to G0 transition involved in cell differentiation(GO:0070315) |
| 2.8 | 11.1 | GO:0030997 | regulation of centriole-centriole cohesion(GO:0030997) |
| 2.7 | 8.1 | GO:0001757 | somite specification(GO:0001757) arterial endothelial cell fate commitment(GO:0060844) blood vessel endothelial cell fate commitment(GO:0060846) endothelial cell fate specification(GO:0060847) Notch signaling pathway involved in arterial endothelial cell fate commitment(GO:0060853) blood vessel endothelial cell fate specification(GO:0097101) |
| 2.5 | 17.8 | GO:0009449 | gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 2.1 | 10.6 | GO:0051661 | maintenance of centrosome location(GO:0051661) |
| 2.1 | 8.3 | GO:0051316 | attachment of spindle microtubules to kinetochore involved in meiotic chromosome segregation(GO:0051316) |
| 2.0 | 11.8 | GO:0045743 | positive regulation of fibroblast growth factor receptor signaling pathway(GO:0045743) |
| 1.8 | 7.0 | GO:0070537 | histone H2A K63-linked deubiquitination(GO:0070537) |
| 1.6 | 4.8 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 1.5 | 4.6 | GO:0072734 | response to staurosporine(GO:0072733) cellular response to staurosporine(GO:0072734) |
| 1.5 | 16.5 | GO:0060539 | diaphragm development(GO:0060539) |
| 1.5 | 4.4 | GO:0014738 | regulation of muscle hyperplasia(GO:0014738) muscle hyperplasia(GO:0014900) |
| 1.5 | 4.4 | GO:0019343 | cysteine biosynthetic process via cystathionine(GO:0019343) cysteine biosynthetic process(GO:0019344) |
| 1.4 | 4.1 | GO:0071579 | phosphatidylinositol-3-phosphate biosynthetic process(GO:0036092) regulation of zinc ion transport(GO:0071579) |
| 1.4 | 5.4 | GO:0001920 | negative regulation of receptor recycling(GO:0001920) negative regulation of low-density lipoprotein particle clearance(GO:0010989) positive regulation of low-density lipoprotein particle receptor catabolic process(GO:0032805) |
| 1.4 | 8.1 | GO:0030644 | cellular chloride ion homeostasis(GO:0030644) |
| 1.3 | 5.4 | GO:0046552 | eye photoreceptor cell fate commitment(GO:0042706) photoreceptor cell fate commitment(GO:0046552) |
| 1.3 | 5.3 | GO:0044340 | canonical Wnt signaling pathway involved in regulation of cell proliferation(GO:0044340) |
| 1.3 | 6.4 | GO:0019659 | fermentation(GO:0006113) lactate biosynthetic process from pyruvate(GO:0019244) glucose catabolic process to lactate(GO:0019659) glycolytic fermentation(GO:0019660) glucose catabolic process to lactate via pyruvate(GO:0019661) |
| 1.2 | 3.7 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 1.2 | 20.9 | GO:0030953 | astral microtubule organization(GO:0030953) |
| 1.1 | 13.3 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 1.1 | 11.8 | GO:0042487 | regulation of odontogenesis of dentin-containing tooth(GO:0042487) |
| 1.0 | 6.0 | GO:1903755 | regulation of SUMO transferase activity(GO:1903182) positive regulation of SUMO transferase activity(GO:1903755) |
| 1.0 | 2.9 | GO:1904760 | myofibroblast differentiation(GO:0036446) regulation of myofibroblast differentiation(GO:1904760) |
| 0.9 | 2.7 | GO:0001827 | inner cell mass cell fate commitment(GO:0001827) |
| 0.9 | 4.3 | GO:0031860 | telomeric 3' overhang formation(GO:0031860) |
| 0.8 | 3.4 | GO:0072272 | proximal/distal pattern formation involved in metanephric nephron development(GO:0072272) |
| 0.8 | 19.1 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.8 | 4.1 | GO:0006287 | base-excision repair, gap-filling(GO:0006287) |
| 0.8 | 4.6 | GO:0003383 | apical constriction(GO:0003383) |
| 0.8 | 10.1 | GO:0019985 | translesion synthesis(GO:0019985) |
| 0.7 | 3.0 | GO:0010700 | negative regulation of norepinephrine secretion(GO:0010700) |
| 0.7 | 2.2 | GO:0034241 | positive regulation of macrophage fusion(GO:0034241) |
| 0.7 | 2.8 | GO:1902268 | negative regulation of polyamine transmembrane transport(GO:1902268) |
| 0.7 | 2.7 | GO:0006776 | vitamin A metabolic process(GO:0006776) positive regulation of skeletal muscle tissue regeneration(GO:0043415) |
| 0.7 | 4.6 | GO:0032534 | regulation of microvillus assembly(GO:0032534) |
| 0.6 | 7.7 | GO:0007042 | lysosomal lumen acidification(GO:0007042) |
| 0.6 | 5.7 | GO:0001842 | neural fold formation(GO:0001842) |
| 0.6 | 2.4 | GO:0010510 | regulation of acetyl-CoA biosynthetic process from pyruvate(GO:0010510) |
| 0.6 | 10.3 | GO:0051451 | myoblast migration(GO:0051451) |
| 0.6 | 2.4 | GO:0072137 | condensed mesenchymal cell proliferation(GO:0072137) |
| 0.6 | 5.2 | GO:0031547 | brain-derived neurotrophic factor receptor signaling pathway(GO:0031547) |
| 0.6 | 2.9 | GO:0034196 | acylglycerol transport(GO:0034196) triglyceride transport(GO:0034197) |
| 0.6 | 2.2 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 0.5 | 1.6 | GO:0000966 | RNA 5'-end processing(GO:0000966) |
| 0.5 | 3.3 | GO:0070345 | negative regulation of fat cell proliferation(GO:0070345) |
| 0.5 | 1.5 | GO:0045658 | eosinophil differentiation(GO:0030222) regulation of neutrophil differentiation(GO:0045658) negative regulation of neutrophil differentiation(GO:0045659) |
| 0.5 | 2.5 | GO:0035720 | intraciliary anterograde transport(GO:0035720) |
| 0.5 | 4.5 | GO:0006071 | glycerol metabolic process(GO:0006071) |
| 0.5 | 5.8 | GO:0075522 | IRES-dependent viral translational initiation(GO:0075522) |
| 0.5 | 1.4 | GO:0021998 | neural plate mediolateral regionalization(GO:0021998) mesoderm structural organization(GO:0048338) paraxial mesoderm structural organization(GO:0048352) regulation of cardiac ventricle development(GO:1904412) positive regulation of cardiac ventricle development(GO:1904414) |
| 0.4 | 1.3 | GO:2000393 | negative regulation of lamellipodium morphogenesis(GO:2000393) |
| 0.4 | 1.3 | GO:0010757 | negative regulation of plasminogen activation(GO:0010757) |
| 0.4 | 4.8 | GO:1901250 | regulation of lung goblet cell differentiation(GO:1901249) negative regulation of lung goblet cell differentiation(GO:1901250) |
| 0.4 | 4.3 | GO:0031119 | tRNA pseudouridine synthesis(GO:0031119) |
| 0.4 | 3.9 | GO:0050882 | voluntary musculoskeletal movement(GO:0050882) |
| 0.4 | 5.0 | GO:0072189 | ureter development(GO:0072189) |
| 0.4 | 3.6 | GO:2000051 | negative regulation of non-canonical Wnt signaling pathway(GO:2000051) |
| 0.4 | 3.1 | GO:0019885 | antigen processing and presentation of endogenous peptide antigen via MHC class I(GO:0019885) |
| 0.4 | 1.9 | GO:0090669 | snoRNA guided rRNA pseudouridine synthesis(GO:0000454) telomerase RNA stabilization(GO:0090669) |
| 0.4 | 19.5 | GO:0045737 | positive regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0045737) |
| 0.4 | 1.1 | GO:0033600 | negative regulation of mammary gland epithelial cell proliferation(GO:0033600) |
| 0.3 | 1.0 | GO:0009838 | abscission(GO:0009838) |
| 0.3 | 6.2 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.3 | 1.7 | GO:0040032 | post-embryonic body morphogenesis(GO:0040032) |
| 0.3 | 4.1 | GO:0002227 | innate immune response in mucosa(GO:0002227) |
| 0.3 | 2.8 | GO:0007000 | nucleolus organization(GO:0007000) |
| 0.3 | 7.8 | GO:0006298 | mismatch repair(GO:0006298) |
| 0.3 | 1.9 | GO:0003065 | positive regulation of heart rate by epinephrine(GO:0003065) |
| 0.3 | 1.9 | GO:0000447 | endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000447) |
| 0.3 | 5.3 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 0.3 | 1.2 | GO:1902626 | assembly of large subunit precursor of preribosome(GO:1902626) |
| 0.3 | 9.1 | GO:0034723 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.3 | 2.0 | GO:0042590 | antigen processing and presentation of exogenous peptide antigen via MHC class I(GO:0042590) |
| 0.3 | 3.6 | GO:0006301 | postreplication repair(GO:0006301) |
| 0.3 | 1.1 | GO:1903347 | negative regulation of bicellular tight junction assembly(GO:1903347) |
| 0.3 | 4.6 | GO:0040034 | regulation of development, heterochronic(GO:0040034) |
| 0.3 | 9.2 | GO:0001709 | cell fate determination(GO:0001709) |
| 0.3 | 1.3 | GO:0050651 | dermatan sulfate proteoglycan biosynthetic process(GO:0050651) |
| 0.3 | 3.9 | GO:0036297 | interstrand cross-link repair(GO:0036297) |
| 0.3 | 15.3 | GO:0006635 | fatty acid beta-oxidation(GO:0006635) |
| 0.3 | 0.8 | GO:0071921 | establishment of sister chromatid cohesion(GO:0034085) cohesin loading(GO:0071921) regulation of cohesin loading(GO:0071922) |
| 0.3 | 3.0 | GO:0002467 | germinal center formation(GO:0002467) |
| 0.2 | 6.9 | GO:0010960 | magnesium ion homeostasis(GO:0010960) |
| 0.2 | 2.4 | GO:0033314 | mitotic DNA replication checkpoint(GO:0033314) |
| 0.2 | 11.0 | GO:0070303 | negative regulation of stress-activated MAPK cascade(GO:0032873) negative regulation of stress-activated protein kinase signaling cascade(GO:0070303) |
| 0.2 | 3.5 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.2 | 2.3 | GO:1902083 | negative regulation of peptidyl-cysteine S-nitrosylation(GO:1902083) |
| 0.2 | 4.6 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.2 | 1.4 | GO:0036066 | protein O-linked fucosylation(GO:0036066) |
| 0.2 | 1.7 | GO:0006265 | DNA topological change(GO:0006265) |
| 0.2 | 0.8 | GO:0032790 | ribosome disassembly(GO:0032790) |
| 0.2 | 0.8 | GO:0035470 | positive regulation of vascular wound healing(GO:0035470) regulation of vascular wound healing(GO:0061043) |
| 0.2 | 0.8 | GO:0032417 | positive regulation of sodium:proton antiporter activity(GO:0032417) negative regulation of calcineurin-NFAT signaling cascade(GO:0070885) |
| 0.2 | 8.2 | GO:0032728 | positive regulation of interferon-beta production(GO:0032728) |
| 0.2 | 2.2 | GO:0046784 | viral mRNA export from host cell nucleus(GO:0046784) |
| 0.2 | 4.2 | GO:0032897 | negative regulation of viral transcription(GO:0032897) |
| 0.2 | 2.2 | GO:0043982 | histone H4-K5 acetylation(GO:0043981) histone H4-K8 acetylation(GO:0043982) |
| 0.2 | 1.8 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 0.2 | 32.7 | GO:0043524 | negative regulation of neuron apoptotic process(GO:0043524) |
| 0.2 | 1.6 | GO:1900262 | regulation of DNA-directed DNA polymerase activity(GO:1900262) positive regulation of DNA-directed DNA polymerase activity(GO:1900264) |
| 0.2 | 2.2 | GO:0014823 | response to activity(GO:0014823) |
| 0.2 | 1.8 | GO:1990173 | protein localization to nuclear body(GO:1903405) positive regulation of establishment of protein localization to telomere(GO:1904851) protein localization to Cajal body(GO:1904867) regulation of protein localization to Cajal body(GO:1904869) positive regulation of protein localization to Cajal body(GO:1904871) protein localization to nucleoplasm(GO:1990173) |
| 0.2 | 0.8 | GO:0014886 | transition between slow and fast fiber(GO:0014886) |
| 0.1 | 0.4 | GO:0032201 | telomere maintenance via semi-conservative replication(GO:0032201) |
| 0.1 | 1.7 | GO:0045899 | positive regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045899) |
| 0.1 | 2.2 | GO:0043162 | ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043162) |
| 0.1 | 8.7 | GO:0048286 | lung alveolus development(GO:0048286) |
| 0.1 | 8.6 | GO:0035914 | skeletal muscle cell differentiation(GO:0035914) |
| 0.1 | 6.9 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.1 | 2.0 | GO:0045647 | negative regulation of erythrocyte differentiation(GO:0045647) |
| 0.1 | 0.4 | GO:0031590 | wybutosine metabolic process(GO:0031590) wybutosine biosynthetic process(GO:0031591) |
| 0.1 | 5.9 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.1 | 2.0 | GO:0042407 | cristae formation(GO:0042407) |
| 0.1 | 4.4 | GO:0000002 | mitochondrial genome maintenance(GO:0000002) |
| 0.1 | 4.8 | GO:0035411 | catenin import into nucleus(GO:0035411) |
| 0.1 | 1.3 | GO:0008063 | Toll signaling pathway(GO:0008063) |
| 0.1 | 1.8 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 0.1 | 0.4 | GO:1902953 | positive regulation of ER to Golgi vesicle-mediated transport(GO:1902953) |
| 0.1 | 1.0 | GO:0032876 | negative regulation of DNA endoreduplication(GO:0032876) modulation by host of viral release from host cell(GO:0044789) positive regulation by host of viral release from host cell(GO:0044791) |
| 0.1 | 0.7 | GO:0090070 | positive regulation of ribosome biogenesis(GO:0090070) positive regulation of rRNA processing(GO:2000234) |
| 0.1 | 1.5 | GO:0014912 | negative regulation of smooth muscle cell migration(GO:0014912) |
| 0.1 | 1.5 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.1 | 3.0 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.1 | 3.1 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.1 | 0.7 | GO:1903003 | positive regulation of protein deubiquitination(GO:1903003) |
| 0.1 | 0.4 | GO:0009405 | pathogenesis(GO:0009405) |
| 0.1 | 7.2 | GO:0032414 | positive regulation of ion transmembrane transporter activity(GO:0032414) |
| 0.1 | 4.0 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.1 | 7.8 | GO:0008203 | cholesterol metabolic process(GO:0008203) |
| 0.1 | 1.5 | GO:0048240 | sperm capacitation(GO:0048240) |
| 0.1 | 0.2 | GO:0006404 | RNA import into nucleus(GO:0006404) |
| 0.1 | 1.6 | GO:0060765 | regulation of androgen receptor signaling pathway(GO:0060765) |
| 0.1 | 3.1 | GO:0097031 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.1 | 2.1 | GO:0048025 | negative regulation of mRNA splicing, via spliceosome(GO:0048025) |
| 0.1 | 1.8 | GO:0071391 | cellular response to estrogen stimulus(GO:0071391) |
| 0.1 | 0.5 | GO:0098789 | pre-mRNA cleavage required for polyadenylation(GO:0098789) |
| 0.1 | 0.7 | GO:0014067 | negative regulation of phosphatidylinositol 3-kinase signaling(GO:0014067) |
| 0.1 | 2.5 | GO:0035315 | hair cell differentiation(GO:0035315) auditory receptor cell differentiation(GO:0042491) |
| 0.1 | 4.6 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 0.1 | 5.9 | GO:0006413 | translational initiation(GO:0006413) |
| 0.1 | 0.9 | GO:0035563 | positive regulation of chromatin binding(GO:0035563) |
| 0.1 | 2.0 | GO:0048599 | oocyte development(GO:0048599) |
| 0.1 | 0.5 | GO:0021554 | optic nerve development(GO:0021554) |
| 0.1 | 1.1 | GO:0034453 | microtubule anchoring(GO:0034453) |
| 0.1 | 1.6 | GO:0061099 | negative regulation of protein tyrosine kinase activity(GO:0061099) |
| 0.1 | 1.6 | GO:0000079 | regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0000079) |
| 0.1 | 0.8 | GO:0006744 | ubiquinone metabolic process(GO:0006743) ubiquinone biosynthetic process(GO:0006744) |
| 0.1 | 0.8 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.1 | 0.1 | GO:0002238 | response to molecule of fungal origin(GO:0002238) serotonin secretion by platelet(GO:0002554) beta selection(GO:0043366) |
| 0.1 | 0.3 | GO:0033140 | negative regulation of peptidyl-serine phosphorylation of STAT protein(GO:0033140) |
| 0.1 | 2.4 | GO:0034394 | protein localization to cell surface(GO:0034394) |
| 0.1 | 0.3 | GO:0008635 | activation of cysteine-type endopeptidase activity involved in apoptotic process by cytochrome c(GO:0008635) |
| 0.0 | 2.9 | GO:0001824 | blastocyst development(GO:0001824) |
| 0.0 | 1.1 | GO:0030970 | retrograde protein transport, ER to cytosol(GO:0030970) |
| 0.0 | 2.0 | GO:0043966 | histone H3 acetylation(GO:0043966) |
| 0.0 | 5.0 | GO:0048864 | stem cell development(GO:0048864) |
| 0.0 | 7.4 | GO:0016485 | protein processing(GO:0016485) |
| 0.0 | 1.2 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.0 | 2.1 | GO:0018105 | peptidyl-serine phosphorylation(GO:0018105) |
| 0.0 | 2.5 | GO:0032436 | positive regulation of proteasomal ubiquitin-dependent protein catabolic process(GO:0032436) |
| 0.0 | 1.0 | GO:0006383 | transcription from RNA polymerase III promoter(GO:0006383) |
| 0.0 | 0.2 | GO:0052428 | modulation of molecular function in other organism(GO:0044359) negative regulation of molecular function in other organism(GO:0044362) negative regulation of molecular function in other organism involved in symbiotic interaction(GO:0052204) modulation of molecular function in other organism involved in symbiotic interaction(GO:0052205) negative regulation by host of symbiont molecular function(GO:0052405) modification by host of symbiont molecular function(GO:0052428) |
| 0.0 | 2.3 | GO:0007338 | single fertilization(GO:0007338) |
| 0.0 | 0.2 | GO:0051533 | positive regulation of NFAT protein import into nucleus(GO:0051533) |
| 0.0 | 0.9 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.0 | 0.1 | GO:1903588 | negative regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903588) |
| 0.0 | 2.4 | GO:0006342 | chromatin silencing(GO:0006342) negative regulation of gene expression, epigenetic(GO:0045814) |
| 0.0 | 0.7 | GO:0006760 | folic acid-containing compound metabolic process(GO:0006760) |
| 0.0 | 0.6 | GO:0051193 | regulation of cofactor metabolic process(GO:0051193) |
| 0.0 | 0.7 | GO:0006399 | tRNA metabolic process(GO:0006399) |
| 0.0 | 0.4 | GO:0030520 | intracellular estrogen receptor signaling pathway(GO:0030520) |
| 0.0 | 0.5 | GO:0000154 | rRNA modification(GO:0000154) |
| 0.0 | 1.5 | GO:0007050 | cell cycle arrest(GO:0007050) |
| 0.0 | 1.7 | GO:0006302 | double-strand break repair(GO:0006302) |
| 0.0 | 0.9 | GO:1904893 | negative regulation of JAK-STAT cascade(GO:0046426) negative regulation of STAT cascade(GO:1904893) |
| 0.0 | 0.9 | GO:0006986 | response to unfolded protein(GO:0006986) |
| 0.0 | 0.3 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.2 | 15.6 | GO:0036125 | mitochondrial fatty acid beta-oxidation multienzyme complex(GO:0016507) fatty acid beta-oxidation multienzyme complex(GO:0036125) |
| 3.9 | 39.3 | GO:0000796 | condensin complex(GO:0000796) |
| 3.1 | 12.4 | GO:0000942 | condensed nuclear chromosome outer kinetochore(GO:0000942) |
| 2.1 | 10.6 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 2.1 | 8.3 | GO:0000939 | nuclear MIS12/MIND complex(GO:0000818) condensed chromosome inner kinetochore(GO:0000939) |
| 1.8 | 5.4 | GO:1990667 | PCSK9-AnxA2 complex(GO:1990667) |
| 1.8 | 7.0 | GO:0070552 | BRISC complex(GO:0070552) |
| 1.6 | 4.8 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 1.5 | 10.3 | GO:0036195 | muscle cell projection(GO:0036194) muscle cell projection membrane(GO:0036195) |
| 1.4 | 10.1 | GO:0031465 | Cul4B-RING E3 ubiquitin ligase complex(GO:0031465) |
| 1.4 | 5.7 | GO:0090537 | CERF complex(GO:0090537) |
| 1.2 | 3.7 | GO:0097543 | ciliary inversin compartment(GO:0097543) |
| 1.0 | 4.1 | GO:0045251 | mitochondrial electron transfer flavoprotein complex(GO:0017133) electron transfer flavoprotein complex(GO:0045251) |
| 1.0 | 6.0 | GO:1990356 | sumoylated E2 ligase complex(GO:1990356) |
| 0.8 | 4.8 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.7 | 4.3 | GO:0030870 | Mre11 complex(GO:0030870) |
| 0.6 | 3.6 | GO:0042405 | nuclear inclusion body(GO:0042405) |
| 0.6 | 4.6 | GO:0033269 | internode region of axon(GO:0033269) |
| 0.6 | 8.3 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.5 | 7.1 | GO:0031011 | Ino80 complex(GO:0031011) |
| 0.5 | 6.4 | GO:0035686 | sperm fibrous sheath(GO:0035686) |
| 0.4 | 3.0 | GO:0033256 | I-kappaB/NF-kappaB complex(GO:0033256) |
| 0.4 | 8.6 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.4 | 2.0 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 0.4 | 8.2 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.4 | 21.4 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.3 | 12.3 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.3 | 7.9 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.3 | 4.4 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
| 0.3 | 4.6 | GO:0034719 | SMN-Sm protein complex(GO:0034719) |
| 0.3 | 23.4 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.3 | 17.0 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.3 | 1.8 | GO:0033061 | DNA recombinase mediator complex(GO:0033061) |
| 0.3 | 1.0 | GO:1990452 | Parkin-FBXW7-Cul1 ubiquitin ligase complex(GO:1990452) |
| 0.3 | 1.8 | GO:0005655 | nucleolar ribonuclease P complex(GO:0005655) |
| 0.2 | 1.0 | GO:0034991 | nuclear meiotic cohesin complex(GO:0034991) |
| 0.2 | 1.1 | GO:0000839 | Hrd1p ubiquitin ligase ERAD-L complex(GO:0000839) |
| 0.2 | 15.7 | GO:0000794 | condensed nuclear chromosome(GO:0000794) |
| 0.2 | 0.7 | GO:0071001 | U4/U6 snRNP(GO:0071001) |
| 0.2 | 2.2 | GO:0000347 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.2 | 12.9 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.2 | 6.2 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.2 | 9.7 | GO:0016235 | aggresome(GO:0016235) |
| 0.2 | 1.7 | GO:0061617 | MICOS complex(GO:0061617) |
| 0.2 | 4.1 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.2 | 8.8 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.2 | 1.6 | GO:0031390 | Ctf18 RFC-like complex(GO:0031390) |
| 0.2 | 1.7 | GO:0031595 | nuclear proteasome complex(GO:0031595) |
| 0.2 | 3.8 | GO:0031305 | integral component of mitochondrial inner membrane(GO:0031305) |
| 0.2 | 2.9 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.2 | 15.4 | GO:0044291 | cell-cell contact zone(GO:0044291) |
| 0.2 | 2.2 | GO:0016010 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.2 | 2.2 | GO:0031233 | intrinsic component of external side of plasma membrane(GO:0031233) |
| 0.2 | 1.8 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.2 | 8.7 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.1 | 1.0 | GO:0090543 | Flemming body(GO:0090543) |
| 0.1 | 1.9 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 0.1 | 2.4 | GO:0005732 | small nucleolar ribonucleoprotein complex(GO:0005732) |
| 0.1 | 7.2 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.1 | 1.0 | GO:0000127 | transcription factor TFIIIC complex(GO:0000127) |
| 0.1 | 4.1 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.1 | 3.2 | GO:0032040 | small-subunit processome(GO:0032040) |
| 0.1 | 2.9 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.1 | 8.2 | GO:0005844 | polysome(GO:0005844) |
| 0.1 | 0.4 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 0.1 | 4.4 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.1 | 0.7 | GO:0016589 | NURF complex(GO:0016589) |
| 0.1 | 1.3 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.1 | 1.1 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.1 | 2.2 | GO:0071339 | MLL1/2 complex(GO:0044665) MLL1 complex(GO:0071339) |
| 0.1 | 3.4 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.1 | 1.2 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.1 | 1.6 | GO:0005849 | mRNA cleavage factor complex(GO:0005849) |
| 0.1 | 3.1 | GO:0045271 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.1 | 5.7 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 1.7 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 1.5 | GO:0031514 | motile cilium(GO:0031514) |
| 0.0 | 2.1 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 0.6 | GO:0030126 | COPI vesicle coat(GO:0030126) |
| 0.0 | 2.8 | GO:0031985 | Golgi cisterna(GO:0031985) |
| 0.0 | 1.3 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.0 | 0.1 | GO:0000931 | gamma-tubulin large complex(GO:0000931) gamma-tubulin ring complex(GO:0008274) |
| 0.0 | 1.3 | GO:0099738 | cell cortex region(GO:0099738) |
| 0.0 | 1.2 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.0 | 0.5 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.0 | 0.7 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.0 | 0.1 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 0.0 | 4.3 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 1.6 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.0 | 25.7 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 2.3 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.0 | 0.8 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.0 | 1.3 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.0 | 1.1 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.0 | 1.4 | GO:0005901 | caveola(GO:0005901) |
| 0.0 | 10.6 | GO:0005815 | microtubule organizing center(GO:0005815) |
| 0.0 | 5.4 | GO:0000790 | nuclear chromatin(GO:0000790) |
| 0.0 | 1.8 | GO:0043197 | dendritic spine(GO:0043197) |
| 0.0 | 13.2 | GO:0005829 | cytosol(GO:0005829) |
| 0.0 | 0.2 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.0 | 0.7 | GO:0097014 | axoneme(GO:0005930) ciliary plasm(GO:0097014) |
| 0.0 | 3.1 | GO:0005667 | transcription factor complex(GO:0005667) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.4 | 21.7 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 4.1 | 16.4 | GO:0016508 | long-chain-enoyl-CoA hydratase activity(GO:0016508) |
| 3.8 | 11.3 | GO:0044715 | 8-oxo-dGDP phosphatase activity(GO:0044715) |
| 3.3 | 30.1 | GO:0031730 | CCR5 chemokine receptor binding(GO:0031730) |
| 2.3 | 16.0 | GO:0004351 | glutamate decarboxylase activity(GO:0004351) |
| 2.1 | 12.4 | GO:0010997 | anaphase-promoting complex binding(GO:0010997) |
| 2.0 | 8.1 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 2.0 | 8.1 | GO:0008802 | betaine-aldehyde dehydrogenase activity(GO:0008802) |
| 1.8 | 16.5 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 1.8 | 5.4 | GO:0034190 | very-low-density lipoprotein particle binding(GO:0034189) apolipoprotein receptor binding(GO:0034190) |
| 1.8 | 7.2 | GO:0051378 | serotonin binding(GO:0051378) |
| 1.6 | 6.4 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 1.5 | 13.3 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 1.5 | 4.4 | GO:0098809 | nitrite reductase activity(GO:0098809) |
| 1.1 | 4.4 | GO:0001010 | transcription factor activity, sequence-specific DNA binding transcription factor recruiting(GO:0001010) |
| 1.0 | 6.1 | GO:0001069 | regulatory region RNA binding(GO:0001069) |
| 1.0 | 6.0 | GO:0061656 | SUMO conjugating enzyme activity(GO:0061656) |
| 1.0 | 3.0 | GO:0031686 | A1 adenosine receptor binding(GO:0031686) |
| 0.9 | 2.7 | GO:0016501 | prostacyclin receptor activity(GO:0016501) |
| 0.9 | 6.9 | GO:0009019 | tRNA (guanine-N1-)-methyltransferase activity(GO:0009019) |
| 0.8 | 5.6 | GO:0034056 | estrogen response element binding(GO:0034056) |
| 0.7 | 2.1 | GO:0017099 | very-long-chain-acyl-CoA dehydrogenase activity(GO:0017099) |
| 0.7 | 7.8 | GO:0008525 | phosphatidylcholine transporter activity(GO:0008525) |
| 0.7 | 2.8 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
| 0.6 | 19.1 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.6 | 2.4 | GO:0004740 | pyruvate dehydrogenase (acetyl-transferring) kinase activity(GO:0004740) |
| 0.6 | 4.6 | GO:0070290 | N-acylphosphatidylethanolamine-specific phospholipase D activity(GO:0070290) |
| 0.5 | 13.1 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.5 | 3.6 | GO:0000403 | Y-form DNA binding(GO:0000403) |
| 0.5 | 8.1 | GO:0030957 | Tat protein binding(GO:0030957) |
| 0.5 | 3.9 | GO:0046790 | virion binding(GO:0046790) |
| 0.4 | 1.7 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 0.4 | 4.1 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.4 | 4.1 | GO:0051575 | 5'-deoxyribose-5-phosphate lyase activity(GO:0051575) |
| 0.4 | 8.2 | GO:0001056 | RNA polymerase III activity(GO:0001056) |
| 0.4 | 4.4 | GO:0017176 | phosphatidylinositol N-acetylglucosaminyltransferase activity(GO:0017176) |
| 0.4 | 23.8 | GO:0001102 | RNA polymerase II activating transcription factor binding(GO:0001102) |
| 0.4 | 3.1 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 0.4 | 5.6 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.4 | 4.0 | GO:0003689 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.4 | 1.4 | GO:0098821 | BMP receptor activity(GO:0098821) |
| 0.4 | 0.7 | GO:0016314 | phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase activity(GO:0016314) |
| 0.4 | 38.6 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.3 | 1.8 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 0.3 | 1.1 | GO:0030292 | protein tyrosine kinase inhibitor activity(GO:0030292) |
| 0.3 | 0.8 | GO:0016494 | C-X-C chemokine receptor activity(GO:0016494) |
| 0.3 | 1.3 | GO:0072542 | protein phosphatase activator activity(GO:0072542) |
| 0.3 | 2.3 | GO:0097153 | cysteine-type endopeptidase activity involved in apoptotic process(GO:0097153) cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.3 | 4.3 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.2 | 1.7 | GO:0036402 | proteasome-activating ATPase activity(GO:0036402) |
| 0.2 | 1.0 | GO:0036033 | mediator complex binding(GO:0036033) |
| 0.2 | 2.2 | GO:0043995 | histone acetyltransferase activity (H4-K5 specific)(GO:0043995) histone acetyltransferase activity (H4-K8 specific)(GO:0043996) histone acetyltransferase activity (H4-K16 specific)(GO:0046972) |
| 0.2 | 11.4 | GO:0061650 | ubiquitin-like protein conjugating enzyme activity(GO:0061650) |
| 0.2 | 7.5 | GO:0043425 | bHLH transcription factor binding(GO:0043425) |
| 0.2 | 17.5 | GO:0019955 | cytokine binding(GO:0019955) |
| 0.2 | 2.0 | GO:1990446 | U1 snRNP binding(GO:1990446) |
| 0.2 | 3.9 | GO:0097371 | MDM2/MDM4 family protein binding(GO:0097371) |
| 0.2 | 0.7 | GO:0004140 | dephospho-CoA kinase activity(GO:0004140) |
| 0.2 | 9.3 | GO:0004407 | histone deacetylase activity(GO:0004407) |
| 0.2 | 15.9 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.2 | 1.8 | GO:1990381 | ubiquitin-specific protease binding(GO:1990381) |
| 0.2 | 3.6 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.2 | 4.3 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.2 | 1.9 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.2 | 3.1 | GO:0004659 | prenyltransferase activity(GO:0004659) |
| 0.2 | 7.1 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.2 | 2.8 | GO:0008649 | rRNA methyltransferase activity(GO:0008649) |
| 0.2 | 3.8 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 0.2 | 1.4 | GO:0008417 | fucosyltransferase activity(GO:0008417) |
| 0.2 | 2.4 | GO:0002161 | aminoacyl-tRNA editing activity(GO:0002161) |
| 0.2 | 1.8 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 0.2 | 11.4 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.2 | 1.9 | GO:0070034 | telomerase RNA binding(GO:0070034) |
| 0.2 | 12.9 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.1 | 1.6 | GO:0042162 | telomeric DNA binding(GO:0042162) |
| 0.1 | 0.9 | GO:1990932 | 5.8S rRNA binding(GO:1990932) |
| 0.1 | 1.9 | GO:0008097 | 5S rRNA binding(GO:0008097) |
| 0.1 | 1.9 | GO:0019213 | deacetylase activity(GO:0019213) |
| 0.1 | 2.9 | GO:0034185 | apolipoprotein binding(GO:0034185) |
| 0.1 | 13.4 | GO:0008083 | growth factor activity(GO:0008083) |
| 0.1 | 5.3 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.1 | 9.5 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.1 | 2.2 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.1 | 6.0 | GO:0031593 | polyubiquitin binding(GO:0031593) |
| 0.1 | 1.5 | GO:0055106 | ubiquitin-protein transferase regulator activity(GO:0055106) |
| 0.1 | 7.3 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.1 | 3.1 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.1 | 1.0 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.1 | 1.3 | GO:0031996 | thioesterase binding(GO:0031996) |
| 0.1 | 3.0 | GO:0019707 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.1 | 6.9 | GO:0008026 | RNA helicase activity(GO:0003724) ATP-dependent RNA helicase activity(GO:0004004) ATP-dependent helicase activity(GO:0008026) RNA-dependent ATPase activity(GO:0008186) purine NTP-dependent helicase activity(GO:0070035) |
| 0.1 | 0.4 | GO:0003880 | protein C-terminal carboxyl O-methyltransferase activity(GO:0003880) |
| 0.1 | 1.8 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.1 | 0.8 | GO:0016861 | intramolecular oxidoreductase activity, interconverting aldoses and ketoses(GO:0016861) |
| 0.1 | 2.0 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.1 | 2.4 | GO:0008236 | serine-type peptidase activity(GO:0008236) |
| 0.1 | 1.5 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.1 | 20.9 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.1 | 0.7 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.1 | 10.0 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.1 | 31.5 | GO:0004674 | protein serine/threonine kinase activity(GO:0004674) |
| 0.1 | 3.0 | GO:0009055 | electron carrier activity(GO:0009055) |
| 0.1 | 1.9 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.1 | 0.7 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.0 | 0.9 | GO:0046965 | retinoid X receptor binding(GO:0046965) |
| 0.0 | 0.4 | GO:0008432 | JUN kinase binding(GO:0008432) |
| 0.0 | 5.4 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.0 | 8.2 | GO:0042393 | histone binding(GO:0042393) |
| 0.0 | 8.9 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.0 | 1.8 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.0 | 0.5 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.0 | 3.8 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.0 | 8.5 | GO:0061630 | ubiquitin protein ligase activity(GO:0061630) |
| 0.0 | 0.8 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
| 0.0 | 1.3 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.0 | 1.5 | GO:0043130 | ubiquitin binding(GO:0043130) |
| 0.0 | 1.4 | GO:0020037 | heme binding(GO:0020037) |
| 0.0 | 1.5 | GO:0004860 | protein kinase inhibitor activity(GO:0004860) |
| 0.0 | 2.6 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 0.0 | 1.0 | GO:0045296 | cadherin binding(GO:0045296) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 23.5 | PID_SMAD2_3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.6 | 22.7 | PID_AURORA_A_PATHWAY | Aurora A signaling |
| 0.6 | 5.4 | ST_TYPE_I_INTERFERON_PATHWAY | Type I Interferon (alpha/beta IFN) Pathway |
| 0.6 | 23.5 | PID_FOXM1_PATHWAY | FOXM1 transcription factor network |
| 0.6 | 26.8 | PID_AURORA_B_PATHWAY | Aurora B signaling |
| 0.5 | 26.1 | PID_IL12_2PATHWAY | IL12-mediated signaling events |
| 0.4 | 14.2 | PID_SYNDECAN_4_PATHWAY | Syndecan-4-mediated signaling events |
| 0.4 | 18.9 | PID_BMP_PATHWAY | BMP receptor signaling |
| 0.4 | 6.3 | PID_BETA_CATENIN_DEG_PATHWAY | Degradation of beta catenin |
| 0.3 | 15.1 | PID_FANCONI_PATHWAY | Fanconi anemia pathway |
| 0.2 | 6.2 | PID_MYC_PATHWAY | C-MYC pathway |
| 0.2 | 10.4 | PID_PS1_PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.2 | 5.8 | PID_CD40_PATHWAY | CD40/CD40L signaling |
| 0.2 | 4.6 | PID_ARF6_DOWNSTREAM_PATHWAY | Arf6 downstream pathway |
| 0.2 | 4.8 | PID_DELTA_NP63_PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.2 | 4.1 | PID_DNA_PK_PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.1 | 6.4 | PID_TELOMERASE_PATHWAY | Regulation of Telomerase |
| 0.1 | 5.0 | PID_HNF3A_PATHWAY | FOXA1 transcription factor network |
| 0.1 | 9.5 | PID_MYC_ACTIV_PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.1 | 7.1 | PID_CMYB_PATHWAY | C-MYB transcription factor network |
| 0.1 | 1.1 | PID_THROMBIN_PAR4_PATHWAY | PAR4-mediated thrombin signaling events |
| 0.1 | 11.8 | NABA_SECRETED_FACTORS | Genes encoding secreted soluble factors |
| 0.1 | 4.3 | SIG_INSULIN_RECEPTOR_PATHWAY_IN_CARDIAC_MYOCYTES | Genes related to the insulin receptor pathway |
| 0.1 | 1.0 | PID_ATM_PATHWAY | ATM pathway |
| 0.1 | 2.1 | PID_HNF3B_PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 4.4 | PID_BETA_CATENIN_NUC_PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 2.8 | PID_MTOR_4PATHWAY | mTOR signaling pathway |
| 0.0 | 2.2 | PID_ERA_GENOMIC_PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.0 | 0.1 | PID_NFKAPPAB_ATYPICAL_PATHWAY | Atypical NF-kappaB pathway |
| 0.0 | 0.7 | PID_AVB3_INTEGRIN_PATHWAY | Integrins in angiogenesis |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 17.7 | REACTOME_MITOCHONDRIAL_FATTY_ACID_BETA_OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.8 | 3.3 | REACTOME_CDC6_ASSOCIATION_WITH_THE_ORC_ORIGIN_COMPLEX | Genes involved in CDC6 association with the ORC:origin complex |
| 0.8 | 7.8 | REACTOME_REMOVAL_OF_THE_FLAP_INTERMEDIATE_FROM_THE_C_STRAND | Genes involved in Removal of the Flap Intermediate from the C-strand |
| 0.7 | 12.4 | REACTOME_CYCLIN_A_B1_ASSOCIATED_EVENTS_DURING_G2_M_TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.7 | 16.0 | REACTOME_GABA_SYNTHESIS_RELEASE_REUPTAKE_AND_DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.6 | 18.9 | REACTOME_SIGNALING_BY_BMP | Genes involved in Signaling by BMP |
| 0.6 | 21.2 | REACTOME_ACTIVATION_OF_ATR_IN_RESPONSE_TO_REPLICATION_STRESS | Genes involved in Activation of ATR in response to replication stress |
| 0.6 | 8.1 | REACTOME_RECEPTOR_LIGAND_BINDING_INITIATES_THE_SECOND_PROTEOLYTIC_CLEAVAGE_OF_NOTCH_RECEPTOR | Genes involved in Receptor-ligand binding initiates the second proteolytic cleavage of Notch receptor |
| 0.5 | 11.1 | REACTOME_APC_CDC20_MEDIATED_DEGRADATION_OF_NEK2A | Genes involved in APC-Cdc20 mediated degradation of Nek2A |
| 0.4 | 4.3 | REACTOME_HOMOLOGOUS_RECOMBINATION_REPAIR_OF_REPLICATION_INDEPENDENT_DOUBLE_STRAND_BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.4 | 8.2 | REACTOME_RNA_POL_III_CHAIN_ELONGATION | Genes involved in RNA Polymerase III Chain Elongation |
| 0.4 | 7.2 | REACTOME_LIGAND_GATED_ION_CHANNEL_TRANSPORT | Genes involved in Ligand-gated ion channel transport |
| 0.3 | 6.7 | REACTOME_FANCONI_ANEMIA_PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.3 | 3.0 | REACTOME_P2Y_RECEPTORS | Genes involved in P2Y receptors |
| 0.3 | 7.8 | REACTOME_SYNTHESIS_OF_GLYCOSYLPHOSPHATIDYLINOSITOL_GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.3 | 4.1 | REACTOME_SYNTHESIS_OF_PIPS_AT_THE_EARLY_ENDOSOME_MEMBRANE | Genes involved in Synthesis of PIPs at the early endosome membrane |
| 0.3 | 3.2 | REACTOME_SLBP_DEPENDENT_PROCESSING_OF_REPLICATION_DEPENDENT_HISTONE_PRE_MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.3 | 8.0 | REACTOME_SIGNALING_BY_HIPPO | Genes involved in Signaling by Hippo |
| 0.3 | 1.0 | REACTOME_SCF_BETA_TRCP_MEDIATED_DEGRADATION_OF_EMI1 | Genes involved in SCF-beta-TrCP mediated degradation of Emi1 |
| 0.2 | 4.6 | REACTOME_ANTIGEN_PRESENTATION_FOLDING_ASSEMBLY_AND_PEPTIDE_LOADING_OF_CLASS_I_MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.2 | 5.1 | REACTOME_PYRUVATE_METABOLISM | Genes involved in Pyruvate metabolism |
| 0.2 | 2.9 | REACTOME_CHYLOMICRON_MEDIATED_LIPID_TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.2 | 4.6 | REACTOME_CASPASE_MEDIATED_CLEAVAGE_OF_CYTOSKELETAL_PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.2 | 15.8 | REACTOME_ANTIVIRAL_MECHANISM_BY_IFN_STIMULATED_GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.2 | 4.4 | REACTOME_SULFUR_AMINO_ACID_METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.2 | 11.1 | REACTOME_MEIOTIC_SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.2 | 1.3 | REACTOME_ACTIVATION_OF_IRF3_IRF7_MEDIATED_BY_TBK1_IKK_EPSILON | Genes involved in Activation of IRF3/IRF7 mediated by TBK1/IKK epsilon |
| 0.1 | 4.6 | REACTOME_TIGHT_JUNCTION_INTERACTIONS | Genes involved in Tight junction interactions |
| 0.1 | 4.6 | REACTOME_SYNTHESIS_OF_PA | Genes involved in Synthesis of PA |
| 0.1 | 3.0 | REACTOME_RIP_MEDIATED_NFKB_ACTIVATION_VIA_DAI | Genes involved in RIP-mediated NFkB activation via DAI |
| 0.1 | 21.6 | REACTOME_METABOLISM_OF_AMINO_ACIDS_AND_DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.1 | 1.7 | REACTOME_SYNTHESIS_SECRETION_AND_DEACYLATION_OF_GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.1 | 4.8 | REACTOME_NCAM1_INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.1 | 7.3 | REACTOME_MITOTIC_PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.1 | 4.6 | REACTOME_MITOCHONDRIAL_PROTEIN_IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.1 | 6.0 | REACTOME_RESPIRATORY_ELECTRON_TRANSPORT | Genes involved in Respiratory electron transport |
| 0.1 | 1.8 | REACTOME_FORMATION_OF_TUBULIN_FOLDING_INTERMEDIATES_BY_CCT_TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.1 | 3.0 | REACTOME_MRNA_SPLICING_MINOR_PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.1 | 1.9 | REACTOME_STRIATED_MUSCLE_CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.1 | 2.2 | REACTOME_SIGNALING_BY_FGFR1_FUSION_MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.1 | 1.7 | REACTOME_MEIOSIS | Genes involved in Meiosis |
| 0.1 | 0.9 | REACTOME_METABOLISM_OF_NON_CODING_RNA | Genes involved in Metabolism of non-coding RNA |
| 0.1 | 0.8 | REACTOME_VITAMIN_B5_PANTOTHENATE_METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.0 | 1.7 | REACTOME_PROSTACYCLIN_SIGNALLING_THROUGH_PROSTACYCLIN_RECEPTOR | Genes involved in Prostacyclin signalling through prostacyclin receptor |
| 0.0 | 0.6 | REACTOME_COPI_MEDIATED_TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.0 | 1.3 | REACTOME_A_TETRASACCHARIDE_LINKER_SEQUENCE_IS_REQUIRED_FOR_GAG_SYNTHESIS | Genes involved in A tetrasaccharide linker sequence is required for GAG synthesis |
| 0.0 | 0.8 | REACTOME_EARLY_PHASE_OF_HIV_LIFE_CYCLE | Genes involved in Early Phase of HIV Life Cycle |
| 0.0 | 3.8 | REACTOME_MRNA_SPLICING | Genes involved in mRNA Splicing |
| 0.0 | 2.7 | REACTOME_NUCLEAR_RECEPTOR_TRANSCRIPTION_PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 1.3 | REACTOME_LYSOSOME_VESICLE_BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
| 0.0 | 1.9 | REACTOME_FORMATION_OF_THE_TERNARY_COMPLEX_AND_SUBSEQUENTLY_THE_43S_COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.0 | 1.1 | REACTOME_SEMA4D_INDUCED_CELL_MIGRATION_AND_GROWTH_CONE_COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.0 | 6.0 | REACTOME_ANTIGEN_PROCESSING_UBIQUITINATION_PROTEASOME_DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.0 | 0.4 | REACTOME_ASSOCIATION_OF_TRIC_CCT_WITH_TARGET_PROTEINS_DURING_BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 0.3 | REACTOME_DESTABILIZATION_OF_MRNA_BY_TRISTETRAPROLIN_TTP | Genes involved in Destabilization of mRNA by Tristetraprolin (TTP) |
| 0.0 | 1.2 | REACTOME_NONSENSE_MEDIATED_DECAY_ENHANCED_BY_THE_EXON_JUNCTION_COMPLEX | Genes involved in Nonsense Mediated Decay Enhanced by the Exon Junction Complex |
| 0.0 | 0.9 | REACTOME_TRANSCRIPTIONAL_REGULATION_OF_WHITE_ADIPOCYTE_DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |