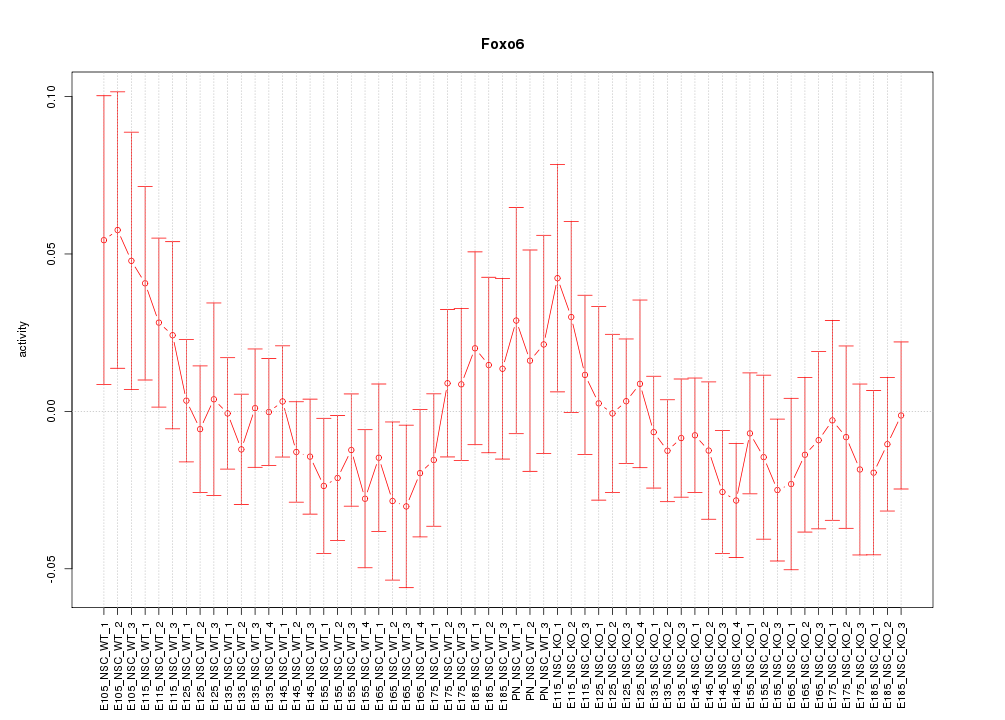
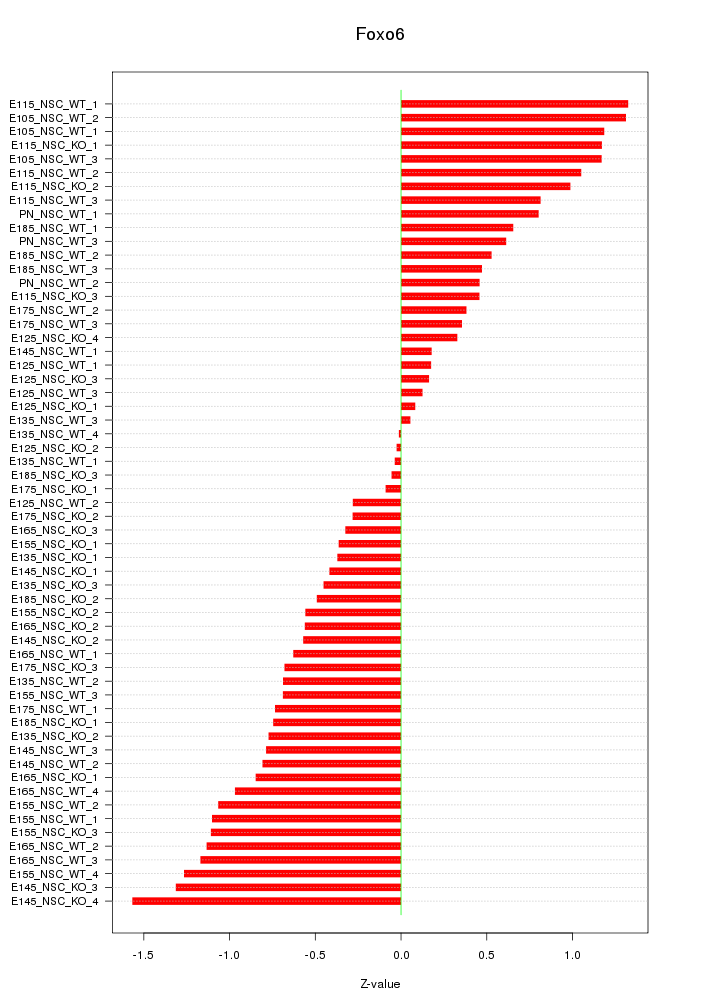
Motif ID: Foxo6
Z-value: 0.757

Transcription factors associated with Foxo6:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| Foxo6 | ENSMUSG00000052135.8 | Foxo6 |
Activity-expression correlation:
| Gene Symbol | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Foxo6 | mm10_v2_chr4_-_120287349_120287349 | -0.87 | 2.5e-19 | Click! |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 4.7 | GO:0060129 | thyroid-stimulating hormone-secreting cell differentiation(GO:0060129) |
| 1.5 | 8.7 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) |
| 1.0 | 3.1 | GO:0007521 | muscle cell fate determination(GO:0007521) mammary placode formation(GO:0060596) |
| 1.0 | 2.9 | GO:0035696 | monocyte extravasation(GO:0035696) regulation of monocyte extravasation(GO:2000437) |
| 0.9 | 3.8 | GO:0034285 | response to sucrose(GO:0009744) response to disaccharide(GO:0034285) |
| 0.9 | 6.0 | GO:0001887 | selenium compound metabolic process(GO:0001887) |
| 0.8 | 4.8 | GO:0006287 | base-excision repair, gap-filling(GO:0006287) |
| 0.6 | 5.3 | GO:0045719 | negative regulation of glycogen biosynthetic process(GO:0045719) negative regulation of glycogen metabolic process(GO:0070874) |
| 0.4 | 1.5 | GO:1903966 | monounsaturated fatty acid metabolic process(GO:1903964) monounsaturated fatty acid biosynthetic process(GO:1903966) |
| 0.3 | 2.8 | GO:1900262 | regulation of DNA-directed DNA polymerase activity(GO:1900262) positive regulation of DNA-directed DNA polymerase activity(GO:1900264) |
| 0.3 | 2.2 | GO:1902416 | positive regulation of mRNA binding(GO:1902416) positive regulation of RNA binding(GO:1905216) |
| 0.3 | 2.5 | GO:0050957 | equilibrioception(GO:0050957) |
| 0.3 | 3.1 | GO:0036289 | peptidyl-serine autophosphorylation(GO:0036289) |
| 0.3 | 1.1 | GO:0072592 | oxygen metabolic process(GO:0072592) |
| 0.2 | 1.0 | GO:0002949 | tRNA threonylcarbamoyladenosine modification(GO:0002949) |
| 0.2 | 0.9 | GO:0007039 | protein catabolic process in the vacuole(GO:0007039) |
| 0.2 | 1.2 | GO:1903887 | motile primary cilium assembly(GO:1903887) |
| 0.2 | 1.4 | GO:0048752 | semicircular canal morphogenesis(GO:0048752) |
| 0.2 | 1.9 | GO:1903204 | liver regeneration(GO:0097421) negative regulation of oxidative stress-induced neuron death(GO:1903204) |
| 0.2 | 1.2 | GO:0034316 | negative regulation of Arp2/3 complex-mediated actin nucleation(GO:0034316) |
| 0.2 | 0.5 | GO:0009106 | lipoate metabolic process(GO:0009106) |
| 0.2 | 4.3 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.1 | 5.9 | GO:0035082 | axoneme assembly(GO:0035082) |
| 0.1 | 0.8 | GO:0019695 | choline metabolic process(GO:0019695) |
| 0.1 | 0.8 | GO:0016255 | attachment of GPI anchor to protein(GO:0016255) |
| 0.1 | 0.7 | GO:0060294 | cilium movement involved in cell motility(GO:0060294) |
| 0.1 | 1.1 | GO:0043248 | proteasome assembly(GO:0043248) |
| 0.1 | 2.6 | GO:0030865 | cortical cytoskeleton organization(GO:0030865) |
| 0.1 | 1.2 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.1 | 3.5 | GO:2000134 | negative regulation of G1/S transition of mitotic cell cycle(GO:2000134) |
| 0.1 | 0.2 | GO:0035434 | copper ion transmembrane transport(GO:0035434) |
| 0.1 | 8.7 | GO:0007051 | spindle organization(GO:0007051) |
| 0.1 | 0.4 | GO:2000124 | regulation of endocannabinoid signaling pathway(GO:2000124) |
| 0.1 | 0.2 | GO:0006624 | vacuolar protein processing(GO:0006624) |
| 0.1 | 0.7 | GO:0003215 | cardiac right ventricle morphogenesis(GO:0003215) |
| 0.0 | 3.0 | GO:0002181 | cytoplasmic translation(GO:0002181) |
| 0.0 | 0.4 | GO:2000059 | negative regulation of protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:2000059) |
| 0.0 | 1.6 | GO:1902653 | cholesterol biosynthetic process(GO:0006695) secondary alcohol biosynthetic process(GO:1902653) |
| 0.0 | 2.6 | GO:0017156 | calcium ion regulated exocytosis(GO:0017156) |
| 0.0 | 1.2 | GO:0000380 | alternative mRNA splicing, via spliceosome(GO:0000380) |
| 0.0 | 1.7 | GO:0032436 | positive regulation of proteasomal ubiquitin-dependent protein catabolic process(GO:0032436) |
| 0.0 | 0.4 | GO:0016180 | snRNA processing(GO:0016180) |
| 0.0 | 0.7 | GO:0030521 | androgen receptor signaling pathway(GO:0030521) |
| 0.0 | 4.3 | GO:0042060 | wound healing(GO:0042060) |
| 0.0 | 3.1 | GO:0019221 | cytokine-mediated signaling pathway(GO:0019221) |
| 0.0 | 0.8 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 4.8 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 0.7 | 5.9 | GO:0001520 | outer dense fiber(GO:0001520) |
| 0.5 | 2.8 | GO:0005663 | DNA replication factor C complex(GO:0005663) |
| 0.3 | 1.0 | GO:0000408 | EKC/KEOPS complex(GO:0000408) |
| 0.3 | 1.0 | GO:0071008 | U2-type post-mRNA release spliceosomal complex(GO:0071008) |
| 0.3 | 3.2 | GO:1990023 | mitotic spindle midzone(GO:1990023) |
| 0.2 | 3.1 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.1 | 0.8 | GO:0042765 | GPI-anchor transamidase complex(GO:0042765) |
| 0.1 | 1.1 | GO:0008541 | proteasome regulatory particle, lid subcomplex(GO:0008541) |
| 0.1 | 2.2 | GO:0045120 | pronucleus(GO:0045120) |
| 0.1 | 8.0 | GO:0000922 | spindle pole(GO:0000922) |
| 0.1 | 5.9 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.1 | 1.2 | GO:0036038 | MKS complex(GO:0036038) |
| 0.1 | 0.7 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.0 | 0.7 | GO:0097228 | sperm midpiece(GO:0097225) sperm principal piece(GO:0097228) |
| 0.0 | 5.4 | GO:0005923 | bicellular tight junction(GO:0005923) |
| 0.0 | 0.4 | GO:0032039 | integrator complex(GO:0032039) |
| 0.0 | 0.4 | GO:0043196 | varicosity(GO:0043196) |
| 0.0 | 0.9 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.0 | 4.2 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.0 | 0.7 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 0.8 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.0 | 1.5 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 0.0 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.0 | 3.5 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.0 | 4.5 | GO:0005667 | transcription factor complex(GO:0005667) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 4.7 | GO:0005105 | type 1 fibroblast growth factor receptor binding(GO:0005105) |
| 1.1 | 4.3 | GO:0032422 | purine-rich negative regulatory element binding(GO:0032422) |
| 0.9 | 3.8 | GO:0097642 | calcitonin family receptor activity(GO:0097642) |
| 0.8 | 3.1 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 0.5 | 1.6 | GO:0004769 | steroid delta-isomerase activity(GO:0004769) |
| 0.5 | 3.0 | GO:0035727 | lysophosphatidic acid binding(GO:0035727) lysophosphatidic acid receptor activity(GO:0070915) |
| 0.5 | 1.9 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) |
| 0.5 | 1.4 | GO:0004492 | methylmalonyl-CoA decarboxylase activity(GO:0004492) |
| 0.4 | 6.0 | GO:0008430 | selenium binding(GO:0008430) |
| 0.4 | 3.5 | GO:0008420 | CTD phosphatase activity(GO:0008420) |
| 0.4 | 1.5 | GO:0032896 | palmitoyl-CoA 9-desaturase activity(GO:0032896) |
| 0.3 | 1.0 | GO:0061711 | N(6)-L-threonylcarbamoyladenine synthase(GO:0061711) |
| 0.3 | 2.8 | GO:0003689 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.2 | 5.3 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.2 | 2.2 | GO:1990715 | mRNA CDS binding(GO:1990715) |
| 0.2 | 1.1 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.2 | 4.3 | GO:0045309 | protein phosphorylated amino acid binding(GO:0045309) |
| 0.2 | 2.9 | GO:0005161 | platelet-derived growth factor receptor binding(GO:0005161) |
| 0.2 | 0.8 | GO:0034235 | GPI anchor binding(GO:0034235) |
| 0.1 | 1.2 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.1 | 2.6 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.1 | 0.8 | GO:0008889 | glycerophosphodiester phosphodiesterase activity(GO:0008889) |
| 0.1 | 0.9 | GO:1990459 | transferrin receptor binding(GO:1990459) |
| 0.1 | 13.2 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.1 | 0.5 | GO:0016783 | sulfurtransferase activity(GO:0016783) |
| 0.0 | 1.7 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.0 | 0.4 | GO:0042500 | aspartic endopeptidase activity, intramembrane cleaving(GO:0042500) |
| 0.0 | 0.4 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
| 0.0 | 3.0 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.0 | 0.7 | GO:0046966 | thyroid hormone receptor binding(GO:0046966) |
| 0.0 | 0.2 | GO:0005375 | copper ion transmembrane transporter activity(GO:0005375) |
| 0.0 | 3.0 | GO:0004721 | phosphoprotein phosphatase activity(GO:0004721) |
| 0.0 | 1.0 | GO:0008186 | ATP-dependent RNA helicase activity(GO:0004004) RNA-dependent ATPase activity(GO:0008186) |