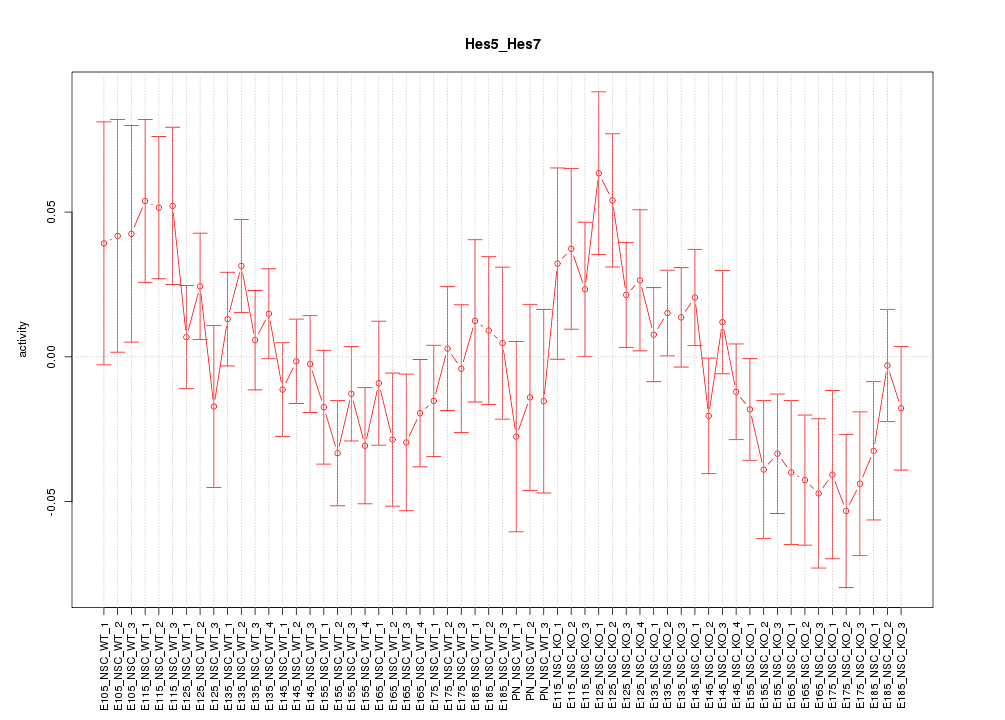
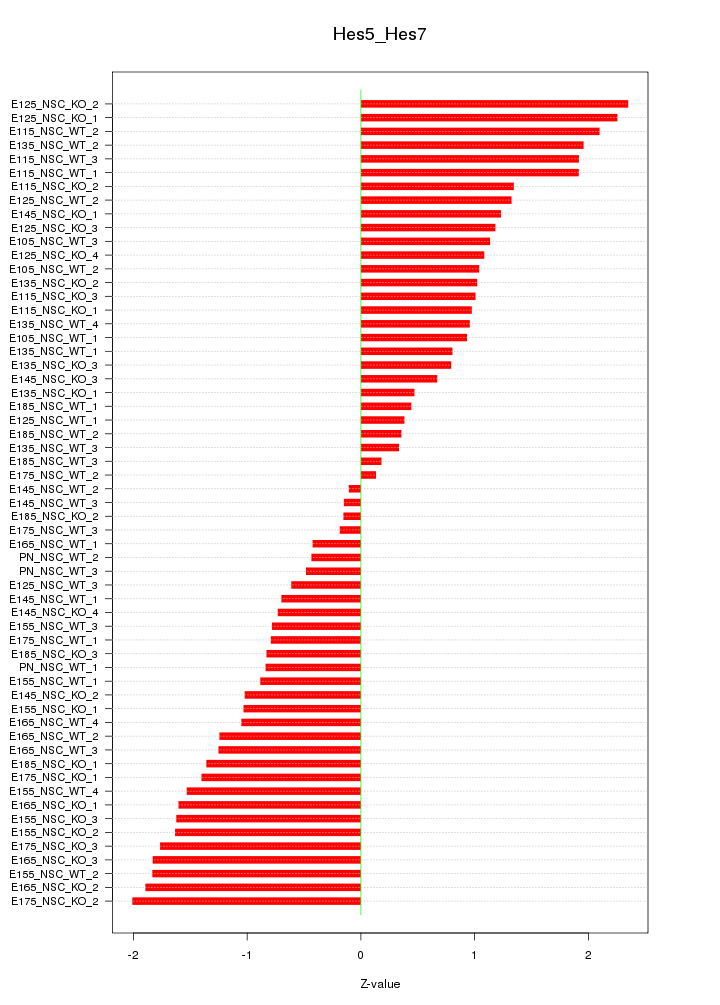
Motif ID: Hes5_Hes7
Z-value: 1.214
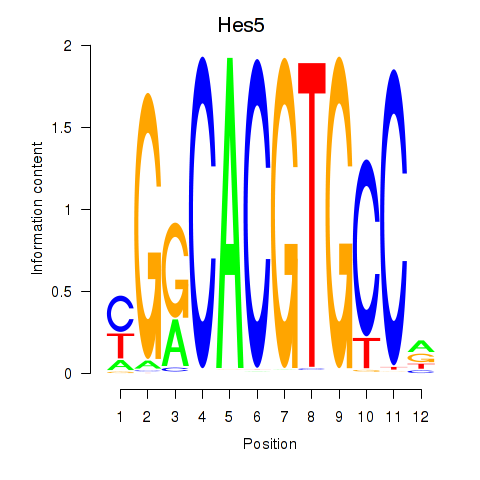
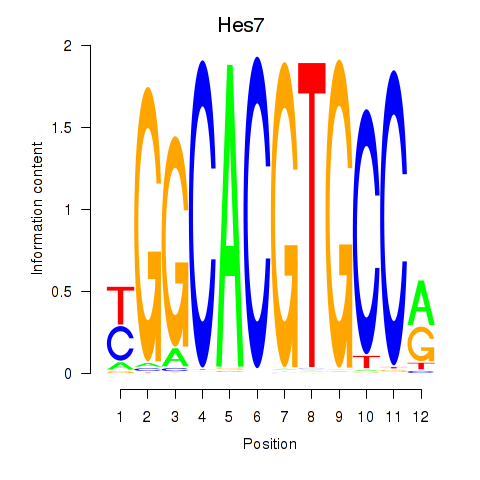
Transcription factors associated with Hes5_Hes7:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| Hes5 | ENSMUSG00000048001.7 | Hes5 |
| Hes7 | ENSMUSG00000023781.2 | Hes7 |
Activity-expression correlation:
| Gene Symbol | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Hes7 | mm10_v2_chr11_+_69120404_69120404 | -0.38 | 2.7e-03 | Click! |
| Hes5 | mm10_v2_chr4_+_154960915_154960930 | 0.30 | 2.3e-02 | Click! |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 10.1 | 91.1 | GO:0045719 | negative regulation of glycogen biosynthetic process(GO:0045719) negative regulation of glycogen metabolic process(GO:0070874) |
| 4.7 | 14.2 | GO:0021759 | globus pallidus development(GO:0021759) |
| 3.7 | 11.1 | GO:0021529 | spinal cord oligodendrocyte cell differentiation(GO:0021529) spinal cord oligodendrocyte cell fate specification(GO:0021530) musculoskeletal movement, spinal reflex action(GO:0050883) olfactory pit development(GO:0060166) |
| 1.6 | 21.9 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 1.4 | 4.3 | GO:0010814 | substance P catabolic process(GO:0010814) calcitonin catabolic process(GO:0010816) endothelin maturation(GO:0034959) |
| 1.4 | 9.8 | GO:0043951 | negative regulation of cAMP-mediated signaling(GO:0043951) |
| 1.4 | 4.2 | GO:0015772 | disaccharide transport(GO:0015766) sucrose transport(GO:0015770) oligosaccharide transport(GO:0015772) |
| 1.0 | 6.0 | GO:0046602 | regulation of mitotic centrosome separation(GO:0046602) |
| 0.9 | 7.5 | GO:0038031 | non-canonical Wnt signaling pathway via JNK cascade(GO:0038031) |
| 0.8 | 2.3 | GO:0010636 | positive regulation of mitochondrial fusion(GO:0010636) |
| 0.7 | 3.4 | GO:0060066 | oviduct development(GO:0060066) |
| 0.6 | 4.5 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.6 | 2.3 | GO:0015744 | succinate transport(GO:0015744) |
| 0.5 | 2.2 | GO:1903416 | response to glycoside(GO:1903416) |
| 0.5 | 1.6 | GO:0009174 | UMP biosynthetic process(GO:0006222) pyrimidine ribonucleoside monophosphate metabolic process(GO:0009173) pyrimidine ribonucleoside monophosphate biosynthetic process(GO:0009174) UMP metabolic process(GO:0046049) |
| 0.5 | 5.7 | GO:0000244 | spliceosomal tri-snRNP complex assembly(GO:0000244) |
| 0.4 | 4.5 | GO:1990403 | embryonic brain development(GO:1990403) |
| 0.4 | 1.1 | GO:1903334 | positive regulation of protein folding(GO:1903334) |
| 0.3 | 12.2 | GO:0045746 | negative regulation of Notch signaling pathway(GO:0045746) |
| 0.3 | 3.4 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.2 | 1.2 | GO:0006344 | maintenance of chromatin silencing(GO:0006344) |
| 0.2 | 2.9 | GO:0033690 | positive regulation of osteoblast proliferation(GO:0033690) |
| 0.2 | 3.9 | GO:0043567 | regulation of insulin-like growth factor receptor signaling pathway(GO:0043567) |
| 0.2 | 1.1 | GO:0010991 | negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.2 | 1.8 | GO:0007000 | nucleolus organization(GO:0007000) |
| 0.2 | 4.1 | GO:0008340 | determination of adult lifespan(GO:0008340) |
| 0.2 | 0.5 | GO:0046368 | GDP-L-fucose metabolic process(GO:0046368) |
| 0.2 | 1.2 | GO:0061484 | hematopoietic stem cell homeostasis(GO:0061484) |
| 0.2 | 5.7 | GO:0030513 | positive regulation of BMP signaling pathway(GO:0030513) |
| 0.2 | 3.9 | GO:0035115 | negative regulation of chondrocyte differentiation(GO:0032331) embryonic forelimb morphogenesis(GO:0035115) |
| 0.1 | 8.9 | GO:0030512 | negative regulation of transforming growth factor beta receptor signaling pathway(GO:0030512) negative regulation of cellular response to transforming growth factor beta stimulus(GO:1903845) |
| 0.1 | 1.1 | GO:0007016 | cytoskeletal anchoring at plasma membrane(GO:0007016) |
| 0.1 | 0.8 | GO:0009098 | branched-chain amino acid biosynthetic process(GO:0009082) leucine biosynthetic process(GO:0009098) valine biosynthetic process(GO:0009099) |
| 0.1 | 2.4 | GO:0035873 | lactate transport(GO:0015727) lactate transmembrane transport(GO:0035873) plasma membrane lactate transport(GO:0035879) |
| 0.1 | 1.4 | GO:0000059 | protein import into nucleus, docking(GO:0000059) |
| 0.1 | 1.3 | GO:0015816 | glycine transport(GO:0015816) |
| 0.1 | 0.5 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 0.1 | 1.0 | GO:0042095 | interferon-gamma biosynthetic process(GO:0042095) |
| 0.1 | 0.3 | GO:0070944 | neutrophil mediated killing of bacterium(GO:0070944) |
| 0.1 | 10.9 | GO:0030177 | positive regulation of Wnt signaling pathway(GO:0030177) |
| 0.1 | 0.3 | GO:0035526 | retrograde transport, plasma membrane to Golgi(GO:0035526) |
| 0.1 | 0.5 | GO:1903755 | regulation of SUMO transferase activity(GO:1903182) positive regulation of SUMO transferase activity(GO:1903755) |
| 0.1 | 0.8 | GO:0032366 | intracellular sterol transport(GO:0032366) |
| 0.1 | 1.9 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.1 | 1.0 | GO:0043248 | proteasome assembly(GO:0043248) |
| 0.0 | 0.7 | GO:2000353 | positive regulation of endothelial cell apoptotic process(GO:2000353) |
| 0.0 | 1.6 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.0 | 1.4 | GO:0048488 | synaptic vesicle endocytosis(GO:0048488) |
| 0.0 | 0.3 | GO:0060371 | regulation of atrial cardiac muscle cell membrane depolarization(GO:0060371) |
| 0.0 | 0.4 | GO:0019368 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.0 | 0.4 | GO:0090073 | positive regulation of protein homodimerization activity(GO:0090073) |
| 0.0 | 1.4 | GO:0014003 | oligodendrocyte development(GO:0014003) |
| 0.0 | 0.1 | GO:0019478 | D-amino acid catabolic process(GO:0019478) |
| 0.0 | 0.0 | GO:0050847 | progesterone receptor signaling pathway(GO:0050847) |
| 0.0 | 0.2 | GO:0090286 | cytoskeletal anchoring at nuclear membrane(GO:0090286) |
| 0.0 | 5.1 | GO:1901657 | glycosyl compound metabolic process(GO:1901657) |
| 0.0 | 0.1 | GO:0034143 | regulation of toll-like receptor 4 signaling pathway(GO:0034143) |
| 0.0 | 0.6 | GO:0002181 | cytoplasmic translation(GO:0002181) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 4.3 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 0.4 | 3.4 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.3 | 2.2 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.3 | 1.4 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
| 0.3 | 3.4 | GO:0043219 | catenin complex(GO:0016342) lateral loop(GO:0043219) |
| 0.2 | 5.7 | GO:0046540 | U4/U6 x U5 tri-snRNP complex(GO:0046540) |
| 0.2 | 1.8 | GO:0070545 | PeBoW complex(GO:0070545) |
| 0.2 | 1.4 | GO:0044233 | ER-mitochondrion membrane contact site(GO:0044233) |
| 0.2 | 0.8 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 0.1 | 2.5 | GO:0005652 | nuclear lamina(GO:0005652) |
| 0.1 | 1.6 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
| 0.1 | 0.6 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.1 | 11.1 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.1 | 6.3 | GO:0005657 | replication fork(GO:0005657) |
| 0.1 | 1.1 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 0.1 | 0.6 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.1 | 0.3 | GO:0034706 | voltage-gated sodium channel complex(GO:0001518) sodium channel complex(GO:0034706) |
| 0.1 | 12.3 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.0 | 4.5 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 0.9 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.0 | 13.4 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 3.9 | GO:0016324 | apical plasma membrane(GO:0016324) |
| 0.0 | 78.4 | GO:0005829 | cytosol(GO:0005829) |
| 0.0 | 5.1 | GO:0005903 | brush border(GO:0005903) |
| 0.0 | 2.9 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 0.5 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.0 | 0.5 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 0.5 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.0 | 0.3 | GO:0036038 | MKS complex(GO:0036038) |
| 0.0 | 3.9 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 1.2 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.0 | 2.4 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.0 | 1.7 | GO:0005840 | ribosome(GO:0005840) |
| 0.0 | 0.8 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.0 | 1.3 | GO:0001726 | ruffle(GO:0001726) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.4 | 21.9 | GO:0018121 | imidazoleglycerol-phosphate synthase activity(GO:0000107) NAD(P)-cysteine ADP-ribosyltransferase activity(GO:0018071) NAD(P)-asparagine ADP-ribosyltransferase activity(GO:0018121) NAD(P)-serine ADP-ribosyltransferase activity(GO:0018127) 7-cyano-7-deazaguanine tRNA-ribosyltransferase activity(GO:0043867) purine deoxyribosyltransferase activity(GO:0044102) |
| 4.1 | 91.1 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 2.0 | 6.0 | GO:0035402 | histone kinase activity (H3-T11 specific)(GO:0035402) |
| 1.4 | 4.2 | GO:0008515 | sucrose:proton symporter activity(GO:0008506) sucrose transmembrane transporter activity(GO:0008515) disaccharide transmembrane transporter activity(GO:0015154) oligosaccharide transmembrane transporter activity(GO:0015157) |
| 1.2 | 14.2 | GO:0001161 | intronic transcription regulatory region sequence-specific DNA binding(GO:0001161) |
| 0.6 | 2.3 | GO:0015141 | succinate transmembrane transporter activity(GO:0015141) |
| 0.6 | 4.5 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.5 | 3.9 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.5 | 2.3 | GO:0008808 | cardiolipin synthase activity(GO:0008808) phosphatidyltransferase activity(GO:0030572) |
| 0.3 | 1.7 | GO:0004614 | phosphoglucomutase activity(GO:0004614) |
| 0.3 | 7.5 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.3 | 3.4 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.3 | 2.2 | GO:0103116 | alpha-D-galactofuranose transporter activity(GO:0103116) steroid hormone binding(GO:1990239) |
| 0.3 | 2.9 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.3 | 11.7 | GO:0043425 | bHLH transcription factor binding(GO:0043425) |
| 0.2 | 9.8 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) ubiquitin-like protein conjugating enzyme activity(GO:0061650) |
| 0.2 | 5.1 | GO:0008422 | beta-glucosidase activity(GO:0008422) |
| 0.2 | 1.1 | GO:0033192 | calcium-dependent protein serine/threonine phosphatase activity(GO:0004723) calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.2 | 2.4 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.1 | 1.3 | GO:0015187 | glycine transmembrane transporter activity(GO:0015187) |
| 0.1 | 0.8 | GO:0052654 | branched-chain-amino-acid transaminase activity(GO:0004084) L-leucine transaminase activity(GO:0052654) L-valine transaminase activity(GO:0052655) L-isoleucine transaminase activity(GO:0052656) |
| 0.1 | 3.9 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.1 | 1.1 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.1 | 0.6 | GO:0033592 | RNA strand annealing activity(GO:0033592) |
| 0.1 | 0.3 | GO:0035851 | histone deacetylase activity (H4-K16 specific)(GO:0034739) Krueppel-associated box domain binding(GO:0035851) |
| 0.1 | 1.4 | GO:0031078 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.1 | 10.0 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.1 | 0.8 | GO:0031996 | thioesterase binding(GO:0031996) |
| 0.1 | 1.4 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.1 | 1.4 | GO:0034185 | apolipoprotein binding(GO:0034185) |
| 0.1 | 4.9 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 1.1 | GO:0016864 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.0 | 0.3 | GO:0086006 | voltage-gated sodium channel activity involved in cardiac muscle cell action potential(GO:0086006) |
| 0.0 | 2.7 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.0 | 1.9 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.0 | 0.4 | GO:0102338 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.0 | 1.6 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.0 | 0.9 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.0 | 0.6 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.0 | 0.0 | GO:0001851 | complement component C3b binding(GO:0001851) proteinase activated receptor binding(GO:0031871) |
| 0.0 | 3.9 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.0 | 3.4 | GO:0008022 | protein C-terminus binding(GO:0008022) |