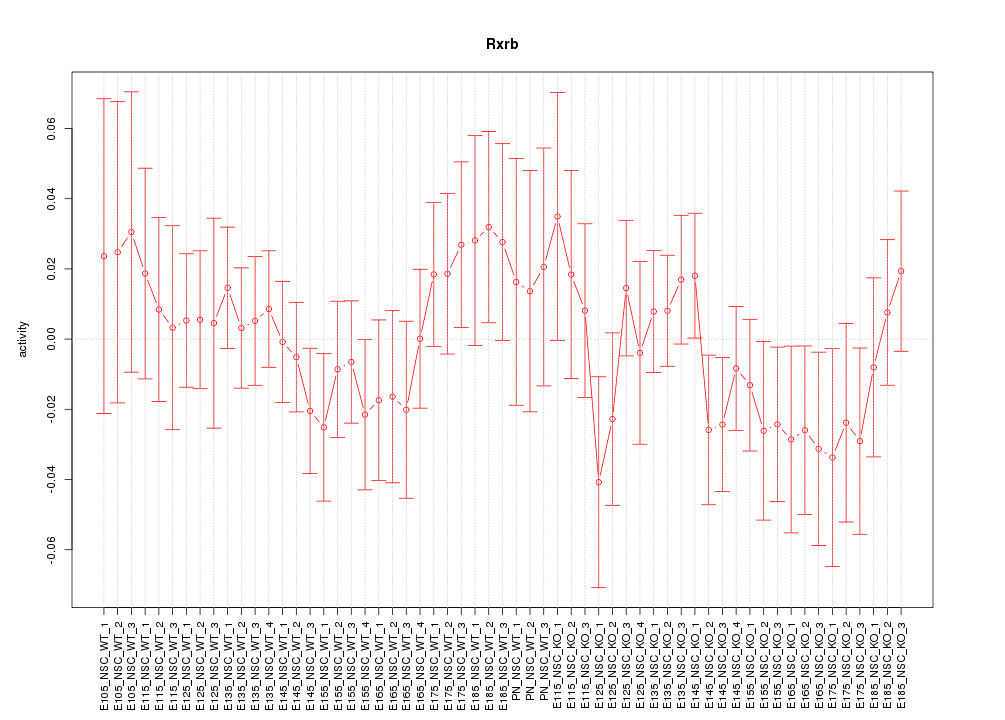
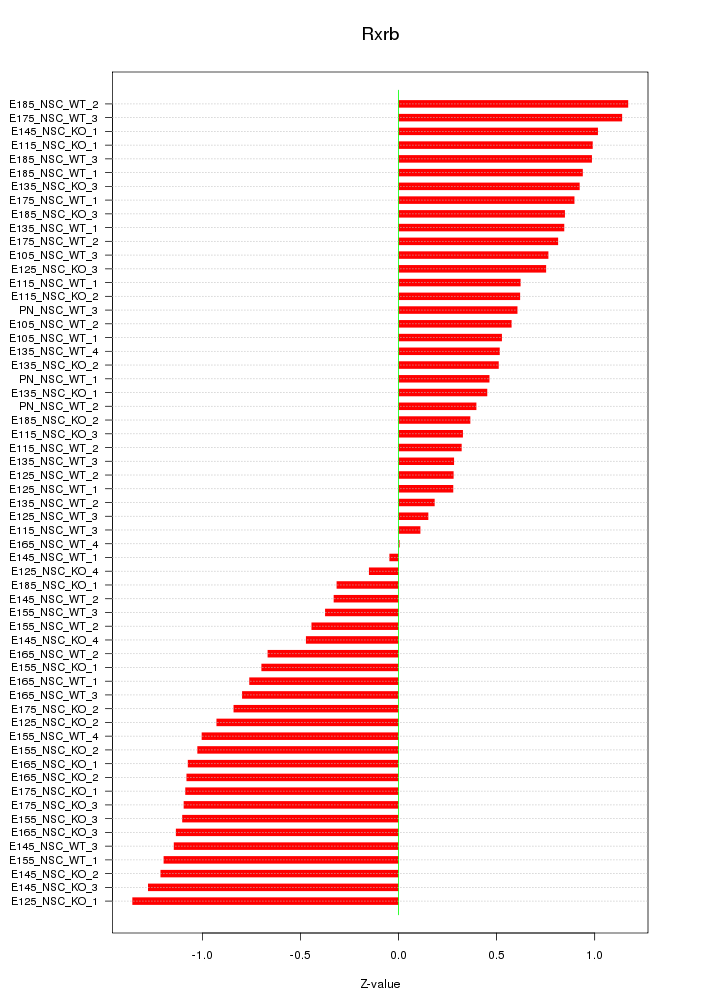
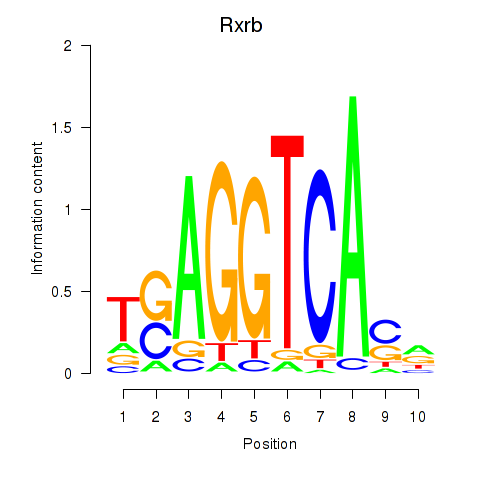
Motif ID: Rxrb
Z-value: 0.786
Transcription factors associated with Rxrb:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| Rxrb | ENSMUSG00000039656.10 | Rxrb |
Activity-expression correlation:
| Gene Symbol | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Rxrb | mm10_v2_chr17_+_34031787_34031821 | -0.62 | 1.4e-07 | Click! |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 3.4 | GO:0060854 | patterning of lymph vessels(GO:0060854) |
| 0.9 | 4.7 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 0.9 | 3.7 | GO:0021847 | ventricular zone neuroblast division(GO:0021847) |
| 0.9 | 5.5 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) |
| 0.9 | 3.6 | GO:1903463 | mitotic cell cycle phase(GO:0098763) regulation of mitotic cell cycle DNA replication(GO:1903463) |
| 0.8 | 4.2 | GO:0014719 | skeletal muscle satellite cell activation(GO:0014719) |
| 0.8 | 2.4 | GO:1903795 | regulation of inorganic anion transmembrane transport(GO:1903795) |
| 0.8 | 4.0 | GO:0035426 | extracellular matrix-cell signaling(GO:0035426) |
| 0.7 | 2.1 | GO:0015910 | peroxisomal long-chain fatty acid import(GO:0015910) |
| 0.6 | 1.9 | GO:0050929 | corticospinal neuron axon guidance through spinal cord(GO:0021972) positive regulation of negative chemotaxis(GO:0050924) induction of negative chemotaxis(GO:0050929) negative regulation of mononuclear cell migration(GO:0071676) negative regulation of retinal ganglion cell axon guidance(GO:0090260) |
| 0.6 | 3.4 | GO:1903753 | cortical microtubule organization(GO:0043622) establishment of centrosome localization(GO:0051660) hard palate development(GO:0060022) negative regulation of p38MAPK cascade(GO:1903753) |
| 0.5 | 3.8 | GO:2000042 | negative regulation of double-strand break repair via homologous recombination(GO:2000042) |
| 0.4 | 3.0 | GO:2000348 | regulation of CD40 signaling pathway(GO:2000348) |
| 0.4 | 3.0 | GO:0010571 | positive regulation of nuclear cell cycle DNA replication(GO:0010571) |
| 0.4 | 1.1 | GO:0002678 | positive regulation of chronic inflammatory response(GO:0002678) |
| 0.3 | 4.9 | GO:0010388 | cullin deneddylation(GO:0010388) |
| 0.3 | 1.6 | GO:0051890 | regulation of cardioblast differentiation(GO:0051890) |
| 0.3 | 2.5 | GO:0061084 | negative regulation of protein refolding(GO:0061084) |
| 0.3 | 2.5 | GO:0006855 | drug transmembrane transport(GO:0006855) |
| 0.3 | 2.3 | GO:0061436 | establishment of skin barrier(GO:0061436) |
| 0.2 | 6.7 | GO:0071398 | cellular response to fatty acid(GO:0071398) |
| 0.2 | 0.7 | GO:1900060 | negative regulation of ceramide biosynthetic process(GO:1900060) |
| 0.2 | 1.1 | GO:0089711 | L-glutamate transmembrane transport(GO:0089711) |
| 0.2 | 1.7 | GO:0042487 | regulation of odontogenesis of dentin-containing tooth(GO:0042487) |
| 0.2 | 1.9 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.1 | 0.8 | GO:0051790 | short-chain fatty acid biosynthetic process(GO:0051790) |
| 0.1 | 3.0 | GO:0070208 | protein heterotrimerization(GO:0070208) |
| 0.1 | 0.9 | GO:0034393 | positive regulation of smooth muscle cell apoptotic process(GO:0034393) |
| 0.1 | 2.0 | GO:0046628 | positive regulation of insulin receptor signaling pathway(GO:0046628) |
| 0.1 | 0.4 | GO:0097494 | regulation of vesicle size(GO:0097494) |
| 0.1 | 1.0 | GO:0051574 | positive regulation of histone H3-K9 methylation(GO:0051574) |
| 0.1 | 3.3 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.1 | 0.6 | GO:2001269 | positive regulation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:2001269) |
| 0.1 | 1.6 | GO:0090136 | epithelial cell-cell adhesion(GO:0090136) |
| 0.1 | 2.1 | GO:0051894 | positive regulation of focal adhesion assembly(GO:0051894) |
| 0.1 | 1.9 | GO:0045742 | positive regulation of epidermal growth factor receptor signaling pathway(GO:0045742) |
| 0.1 | 3.7 | GO:0008631 | intrinsic apoptotic signaling pathway in response to oxidative stress(GO:0008631) |
| 0.1 | 0.3 | GO:0034115 | negative regulation of heterotypic cell-cell adhesion(GO:0034115) cell-cell adhesion involved in gastrulation(GO:0070586) regulation of cell-cell adhesion involved in gastrulation(GO:0070587) |
| 0.1 | 1.1 | GO:2000010 | positive regulation of protein localization to cell surface(GO:2000010) |
| 0.1 | 1.3 | GO:2000114 | regulation of establishment of cell polarity(GO:2000114) |
| 0.1 | 2.1 | GO:0006760 | folic acid-containing compound metabolic process(GO:0006760) |
| 0.1 | 0.3 | GO:0015793 | glycerol transport(GO:0015793) |
| 0.1 | 1.2 | GO:0010225 | response to UV-C(GO:0010225) |
| 0.1 | 0.6 | GO:0015868 | purine ribonucleotide transport(GO:0015868) |
| 0.1 | 0.4 | GO:0006451 | selenocysteine incorporation(GO:0001514) translational readthrough(GO:0006451) |
| 0.1 | 0.7 | GO:0007190 | activation of adenylate cyclase activity(GO:0007190) |
| 0.0 | 0.8 | GO:0043248 | proteasome assembly(GO:0043248) |
| 0.0 | 1.5 | GO:0048488 | synaptic vesicle endocytosis(GO:0048488) |
| 0.0 | 1.5 | GO:0070830 | bicellular tight junction assembly(GO:0070830) |
| 0.0 | 0.5 | GO:0051764 | actin crosslink formation(GO:0051764) positive regulation of actin cytoskeleton reorganization(GO:2000251) |
| 0.0 | 0.7 | GO:0048671 | negative regulation of collateral sprouting(GO:0048671) |
| 0.0 | 3.9 | GO:0051028 | mRNA transport(GO:0051028) |
| 0.0 | 0.7 | GO:0015985 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.0 | 0.4 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.0 | 1.7 | GO:0043149 | contractile actin filament bundle assembly(GO:0030038) stress fiber assembly(GO:0043149) |
| 0.0 | 0.5 | GO:1901522 | positive regulation of transcription from RNA polymerase II promoter involved in cellular response to chemical stimulus(GO:1901522) |
| 0.0 | 0.5 | GO:1902230 | negative regulation of intrinsic apoptotic signaling pathway in response to DNA damage(GO:1902230) |
| 0.0 | 0.8 | GO:0010765 | positive regulation of sodium ion transport(GO:0010765) |
| 0.0 | 0.7 | GO:0061077 | chaperone-mediated protein folding(GO:0061077) |
| 0.0 | 1.9 | GO:0042787 | protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0042787) |
| 0.0 | 0.1 | GO:0046548 | retinal rod cell development(GO:0046548) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 2.5 | GO:0031088 | platelet dense granule membrane(GO:0031088) |
| 0.7 | 3.4 | GO:0097025 | MPP7-DLG1-LIN7 complex(GO:0097025) |
| 0.6 | 1.7 | GO:0071953 | elastic fiber(GO:0071953) |
| 0.4 | 3.9 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.2 | 3.1 | GO:0031011 | Ino80 complex(GO:0031011) |
| 0.2 | 1.5 | GO:0044233 | ER-mitochondrion membrane contact site(GO:0044233) |
| 0.2 | 4.2 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.2 | 0.7 | GO:0035339 | SPOTS complex(GO:0035339) |
| 0.2 | 4.4 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.2 | 2.9 | GO:0002102 | podosome(GO:0002102) |
| 0.1 | 3.7 | GO:0042470 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.1 | 1.6 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.1 | 2.1 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.1 | 2.1 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) |
| 0.1 | 5.1 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.1 | 2.4 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.1 | 0.5 | GO:0071439 | clathrin complex(GO:0071439) |
| 0.0 | 0.6 | GO:0005922 | connexon complex(GO:0005922) |
| 0.0 | 0.7 | GO:0045263 | proton-transporting ATP synthase complex, coupling factor F(o)(GO:0045263) |
| 0.0 | 6.1 | GO:0000776 | kinetochore(GO:0000776) |
| 0.0 | 1.9 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 3.5 | GO:0005604 | basement membrane(GO:0005604) |
| 0.0 | 0.6 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.0 | 0.9 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.0 | 0.8 | GO:0005771 | multivesicular body(GO:0005771) |
| 0.0 | 2.5 | GO:0016234 | inclusion body(GO:0016234) |
| 0.0 | 1.1 | GO:0030673 | axolemma(GO:0030673) |
| 0.0 | 3.0 | GO:0000793 | condensed chromosome(GO:0000793) |
| 0.0 | 1.5 | GO:0016459 | myosin complex(GO:0016459) |
| 0.0 | 2.0 | GO:0043234 | protein complex(GO:0043234) |
| 0.0 | 3.6 | GO:0000781 | chromosome, telomeric region(GO:0000781) |
| 0.0 | 3.7 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 1.1 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 0.6 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.0 | 1.1 | GO:0014704 | intercalated disc(GO:0014704) |
| 0.0 | 4.0 | GO:0005938 | cell cortex(GO:0005938) |
| 0.0 | 0.5 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 0.4 | GO:0045335 | phagocytic vesicle(GO:0045335) |
| 0.0 | 5.6 | GO:0000785 | chromatin(GO:0000785) |
| 0.0 | 2.6 | GO:0043235 | receptor complex(GO:0043235) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 5.9 | GO:0043237 | laminin-1 binding(GO:0043237) |
| 0.7 | 3.4 | GO:0097016 | L27 domain binding(GO:0097016) |
| 0.5 | 2.0 | GO:0000982 | transcription factor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0000982) transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.5 | 4.6 | GO:0015245 | fatty acid transporter activity(GO:0015245) |
| 0.4 | 3.4 | GO:0001224 | RNA polymerase II transcription cofactor binding(GO:0001224) |
| 0.3 | 4.4 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.3 | 3.7 | GO:0016863 | intramolecular oxidoreductase activity, transposing C=C bonds(GO:0016863) |
| 0.3 | 1.1 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 0.2 | 0.8 | GO:0004315 | 3-oxoacyl-[acyl-carrier-protein] synthase activity(GO:0004315) |
| 0.2 | 1.9 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 0.2 | 3.0 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.2 | 3.1 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.2 | 6.7 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.2 | 1.9 | GO:0030296 | protein tyrosine kinase activator activity(GO:0030296) |
| 0.1 | 2.1 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.1 | 1.1 | GO:0103116 | alpha-D-galactofuranose transporter activity(GO:0103116) |
| 0.1 | 1.6 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.1 | 3.7 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.1 | 3.5 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.1 | 1.5 | GO:0034185 | apolipoprotein binding(GO:0034185) |
| 0.1 | 0.4 | GO:0035368 | selenocysteine insertion sequence binding(GO:0035368) |
| 0.1 | 2.0 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.1 | 0.7 | GO:0016500 | protein-hormone receptor activity(GO:0016500) |
| 0.1 | 0.7 | GO:0015078 | hydrogen ion transmembrane transporter activity(GO:0015078) |
| 0.1 | 1.7 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 1.2 | GO:0000400 | four-way junction DNA binding(GO:0000400) |
| 0.1 | 0.6 | GO:0005243 | gap junction channel activity(GO:0005243) |
| 0.0 | 0.3 | GO:0015265 | urea channel activity(GO:0015265) |
| 0.0 | 0.5 | GO:0004886 | 9-cis retinoic acid receptor activity(GO:0004886) |
| 0.0 | 1.7 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.2 | GO:0004301 | epoxide hydrolase activity(GO:0004301) |
| 0.0 | 0.4 | GO:0070181 | small ribosomal subunit rRNA binding(GO:0070181) |
| 0.0 | 2.1 | GO:0051018 | protein kinase A binding(GO:0051018) |
| 0.0 | 0.7 | GO:0030553 | cGMP binding(GO:0030553) |
| 0.0 | 6.3 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.0 | 2.6 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.0 | 0.7 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.0 | 2.1 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.5 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 2.8 | GO:0008017 | microtubule binding(GO:0008017) |
| 0.0 | 0.8 | GO:0032947 | protein complex scaffold(GO:0032947) |
| 0.0 | 4.9 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |