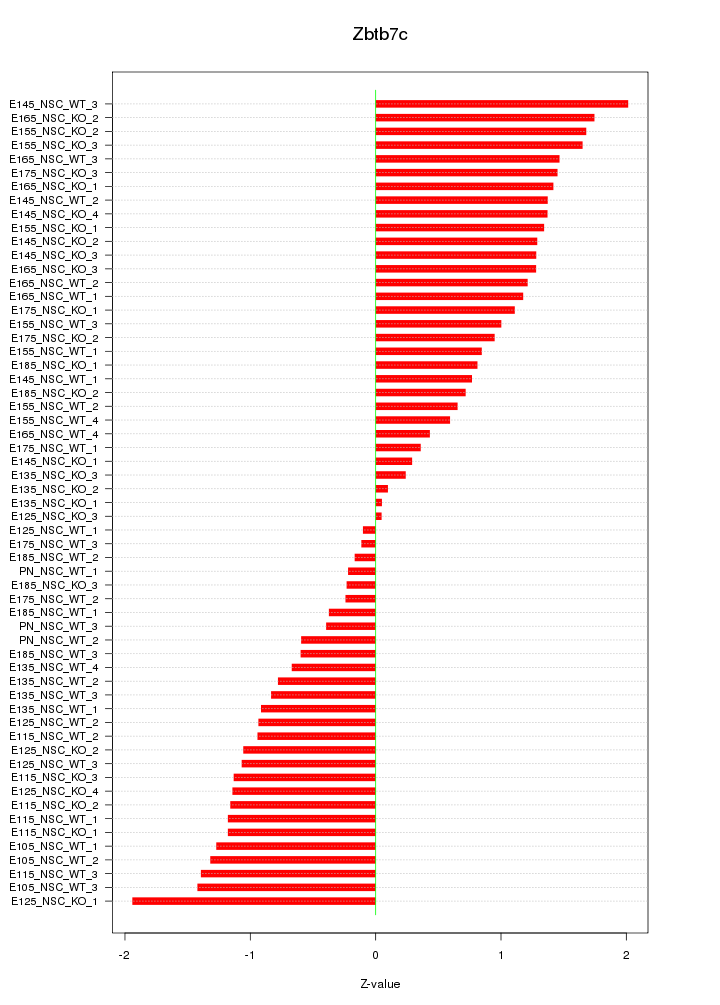
Motif ID: Zbtb7c
Z-value: 1.048

Transcription factors associated with Zbtb7c:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| Zbtb7c | ENSMUSG00000044646.8 | Zbtb7c |
Activity-expression correlation:
| Gene Symbol | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Zbtb7c | mm10_v2_chr18_+_76059458_76059501 | 0.16 | 2.2e-01 | Click! |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.4 | 37.8 | GO:1900454 | positive regulation of long term synaptic depression(GO:1900454) |
| 2.3 | 9.3 | GO:0021812 | neuronal-glial interaction involved in cerebral cortex radial glia guided migration(GO:0021812) |
| 1.1 | 3.4 | GO:0035864 | response to potassium ion(GO:0035864) cellular response to potassium ion(GO:0035865) |
| 0.9 | 3.7 | GO:0010891 | negative regulation of sequestering of triglyceride(GO:0010891) |
| 0.8 | 4.5 | GO:1903056 | regulation of lens fiber cell differentiation(GO:1902746) positive regulation of lens fiber cell differentiation(GO:1902748) regulation of melanosome organization(GO:1903056) |
| 0.7 | 3.7 | GO:1900122 | positive regulation of receptor binding(GO:1900122) |
| 0.7 | 11.4 | GO:0043011 | myeloid dendritic cell differentiation(GO:0043011) |
| 0.6 | 1.8 | GO:1904742 | protein poly-ADP-ribosylation(GO:0070212) regulation of telomeric DNA binding(GO:1904742) |
| 0.6 | 2.9 | GO:0045053 | protein retention in Golgi apparatus(GO:0045053) |
| 0.5 | 3.8 | GO:1903423 | positive regulation of synaptic vesicle recycling(GO:1903423) |
| 0.4 | 2.0 | GO:0030242 | pexophagy(GO:0030242) |
| 0.4 | 3.2 | GO:0090074 | negative regulation of protein homodimerization activity(GO:0090074) |
| 0.4 | 6.8 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.3 | 4.2 | GO:0021869 | forebrain ventricular zone progenitor cell division(GO:0021869) |
| 0.3 | 1.0 | GO:0006990 | positive regulation of transcription from RNA polymerase II promoter involved in unfolded protein response(GO:0006990) |
| 0.3 | 1.5 | GO:0090467 | lysine transport(GO:0015819) L-arginine import(GO:0043091) arginine import(GO:0090467) L-arginine transport(GO:1902023) |
| 0.3 | 3.1 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.3 | 1.9 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.2 | 2.2 | GO:0010793 | regulation of mRNA export from nucleus(GO:0010793) regulation of ribonucleoprotein complex localization(GO:2000197) |
| 0.2 | 2.3 | GO:0006002 | fructose 6-phosphate metabolic process(GO:0006002) |
| 0.2 | 3.1 | GO:0070445 | oligodendrocyte progenitor proliferation(GO:0070444) regulation of oligodendrocyte progenitor proliferation(GO:0070445) |
| 0.2 | 1.8 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.2 | 4.4 | GO:0014898 | muscle hypertrophy in response to stress(GO:0003299) cardiac muscle hypertrophy in response to stress(GO:0014898) |
| 0.2 | 12.9 | GO:0015807 | L-amino acid transport(GO:0015807) |
| 0.2 | 2.1 | GO:2000465 | regulation of glycogen (starch) synthase activity(GO:2000465) |
| 0.2 | 1.0 | GO:1900246 | positive regulation of RIG-I signaling pathway(GO:1900246) |
| 0.2 | 2.2 | GO:0000042 | protein targeting to Golgi(GO:0000042) |
| 0.2 | 0.9 | GO:0051013 | microtubule severing(GO:0051013) |
| 0.1 | 2.0 | GO:0032438 | melanosome organization(GO:0032438) endosome to melanosome transport(GO:0035646) endosome to pigment granule transport(GO:0043485) pigment granule maturation(GO:0048757) |
| 0.1 | 3.3 | GO:0021702 | cerebellar Purkinje cell differentiation(GO:0021702) |
| 0.1 | 1.6 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.1 | 9.1 | GO:0060999 | positive regulation of dendritic spine development(GO:0060999) |
| 0.1 | 7.1 | GO:0035418 | protein localization to synapse(GO:0035418) |
| 0.1 | 15.1 | GO:0030705 | cytoskeleton-dependent intracellular transport(GO:0030705) |
| 0.1 | 14.5 | GO:0071805 | potassium ion transmembrane transport(GO:0071805) |
| 0.1 | 2.3 | GO:0046854 | phosphatidylinositol phosphorylation(GO:0046854) |
| 0.1 | 2.4 | GO:0007189 | adenylate cyclase-activating G-protein coupled receptor signaling pathway(GO:0007189) |
| 0.1 | 0.5 | GO:0042095 | interferon-gamma biosynthetic process(GO:0042095) |
| 0.1 | 1.5 | GO:0010955 | negative regulation of protein processing(GO:0010955) negative regulation of protein maturation(GO:1903318) |
| 0.0 | 0.2 | GO:0045657 | positive regulation of monocyte differentiation(GO:0045657) |
| 0.0 | 0.9 | GO:0046827 | positive regulation of protein export from nucleus(GO:0046827) |
| 0.0 | 1.0 | GO:2000279 | negative regulation of DNA biosynthetic process(GO:2000279) |
| 0.0 | 16.0 | GO:0007409 | axonogenesis(GO:0007409) |
| 0.0 | 0.5 | GO:0045116 | protein neddylation(GO:0045116) |
| 0.0 | 0.5 | GO:0097345 | mitochondrial outer membrane permeabilization(GO:0097345) |
| 0.0 | 1.3 | GO:0006919 | activation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0006919) |
| 0.0 | 4.0 | GO:0000209 | protein polyubiquitination(GO:0000209) |
| 0.0 | 0.4 | GO:0070734 | histone H3-K27 methylation(GO:0070734) |
| 0.0 | 2.2 | GO:0051291 | protein heterooligomerization(GO:0051291) |
| 0.0 | 2.8 | GO:0006457 | protein folding(GO:0006457) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.8 | 37.8 | GO:0045298 | tubulin complex(GO:0045298) |
| 1.6 | 6.5 | GO:0043259 | laminin-1 complex(GO:0005606) laminin-10 complex(GO:0043259) |
| 1.0 | 15.1 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.7 | 3.7 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 0.7 | 6.8 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.6 | 2.6 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.4 | 14.2 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.4 | 12.0 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.3 | 4.7 | GO:0097470 | ribbon synapse(GO:0097470) |
| 0.3 | 3.8 | GO:0043196 | varicosity(GO:0043196) |
| 0.2 | 2.2 | GO:0000347 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.2 | 2.0 | GO:0030008 | TRAPP complex(GO:0030008) |
| 0.2 | 1.8 | GO:0002178 | palmitoyltransferase complex(GO:0002178) |
| 0.2 | 15.0 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) |
| 0.2 | 1.9 | GO:0045095 | keratin filament(GO:0045095) |
| 0.1 | 3.1 | GO:0043034 | costamere(GO:0043034) |
| 0.1 | 2.2 | GO:0030137 | COPI-coated vesicle(GO:0030137) |
| 0.1 | 2.0 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.0 | 2.5 | GO:0001725 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.0 | 0.2 | GO:0008091 | spectrin(GO:0008091) |
| 0.0 | 24.2 | GO:0005730 | nucleolus(GO:0005730) |
| 0.0 | 2.7 | GO:0000784 | nuclear chromosome, telomeric region(GO:0000784) |
| 0.0 | 1.0 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.0 | 12.1 | GO:0005887 | integral component of plasma membrane(GO:0005887) |
| 0.0 | 1.6 | GO:0005901 | caveola(GO:0005901) |
| 0.0 | 1.8 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.0 | 0.1 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.7 | 5.1 | GO:0060072 | large conductance calcium-activated potassium channel activity(GO:0060072) |
| 1.6 | 37.8 | GO:0034185 | apolipoprotein binding(GO:0034185) |
| 1.0 | 15.1 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.8 | 6.5 | GO:0043208 | glycosphingolipid binding(GO:0043208) |
| 0.5 | 2.1 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.5 | 11.4 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.5 | 5.6 | GO:0015271 | outward rectifier potassium channel activity(GO:0015271) |
| 0.4 | 3.1 | GO:0031802 | type 5 metabotropic glutamate receptor binding(GO:0031802) |
| 0.4 | 3.8 | GO:0031749 | D2 dopamine receptor binding(GO:0031749) |
| 0.3 | 3.9 | GO:0070679 | inositol 1,4,5 trisphosphate binding(GO:0070679) |
| 0.3 | 12.9 | GO:0015175 | neutral amino acid transmembrane transporter activity(GO:0015175) |
| 0.3 | 3.2 | GO:0048273 | mitogen-activated protein kinase p38 binding(GO:0048273) |
| 0.2 | 5.4 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.2 | 1.5 | GO:0015189 | L-ornithine transmembrane transporter activity(GO:0000064) arginine transmembrane transporter activity(GO:0015181) L-lysine transmembrane transporter activity(GO:0015189) |
| 0.2 | 2.6 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.2 | 4.2 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.2 | 3.4 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.1 | 1.6 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.1 | 3.7 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.1 | 4.5 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.1 | 2.3 | GO:0008483 | transaminase activity(GO:0008483) |
| 0.1 | 1.8 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.1 | 5.7 | GO:0031593 | polyubiquitin binding(GO:0031593) |
| 0.1 | 0.9 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 0.1 | 1.9 | GO:0008330 | protein tyrosine/threonine phosphatase activity(GO:0008330) |
| 0.1 | 1.8 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) |
| 0.1 | 0.4 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.1 | 1.0 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.1 | 0.5 | GO:0019788 | NEDD8 transferase activity(GO:0019788) |
| 0.1 | 7.8 | GO:0016810 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds(GO:0016810) |
| 0.1 | 1.5 | GO:0005112 | Notch binding(GO:0005112) |
| 0.1 | 6.4 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.1 | 7.5 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.1 | 0.8 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.1 | 2.4 | GO:0035255 | ionotropic glutamate receptor binding(GO:0035255) |
| 0.0 | 2.7 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.0 | 2.8 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.0 | 3.1 | GO:0017048 | Rho GTPase binding(GO:0017048) |
| 0.0 | 0.9 | GO:0042162 | telomeric DNA binding(GO:0042162) |
| 0.0 | 3.0 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.0 | 1.0 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.0 | 2.2 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.0 | 12.4 | GO:0043565 | sequence-specific DNA binding(GO:0043565) |
| 0.0 | 2.4 | GO:0004930 | G-protein coupled receptor activity(GO:0004930) |
| 0.0 | 1.4 | GO:0003729 | mRNA binding(GO:0003729) |