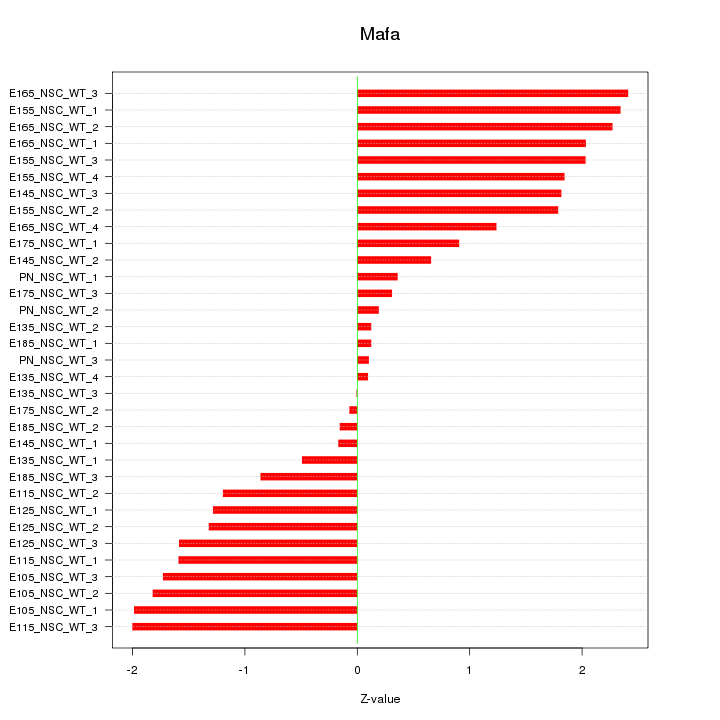
Motif ID: Mafa
Z-value: 1.380
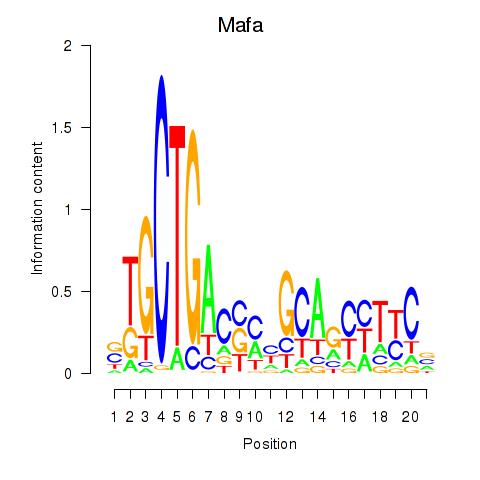
Transcription factors associated with Mafa:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| Mafa | ENSMUSG00000047591.4 | Mafa |
Activity-expression correlation:
| Gene Symbol | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Mafa | mm10_v2_chr15_-_75747922_75747922 | 0.15 | 4.0e-01 | Click! |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.5 | 13.5 | GO:0050925 | negative regulation of negative chemotaxis(GO:0050925) |
| 1.2 | 7.1 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 0.8 | 2.4 | GO:0009174 | UMP biosynthetic process(GO:0006222) pyrimidine ribonucleoside monophosphate metabolic process(GO:0009173) pyrimidine ribonucleoside monophosphate biosynthetic process(GO:0009174) UMP metabolic process(GO:0046049) |
| 0.8 | 7.7 | GO:0033603 | positive regulation of dopamine secretion(GO:0033603) |
| 0.8 | 7.6 | GO:0033631 | cell-cell adhesion mediated by integrin(GO:0033631) |
| 0.7 | 3.7 | GO:0034436 | glycoprotein transport(GO:0034436) |
| 0.7 | 4.9 | GO:0036072 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 0.6 | 9.2 | GO:0060363 | cranial suture morphogenesis(GO:0060363) |
| 0.5 | 7.1 | GO:1901898 | negative regulation of relaxation of muscle(GO:1901078) negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.5 | 4.3 | GO:0035507 | regulation of myosin-light-chain-phosphatase activity(GO:0035507) |
| 0.5 | 2.9 | GO:0000160 | phosphorelay signal transduction system(GO:0000160) |
| 0.5 | 5.0 | GO:0046543 | development of secondary female sexual characteristics(GO:0046543) |
| 0.4 | 1.5 | GO:0046709 | IDP metabolic process(GO:0046707) IDP catabolic process(GO:0046709) |
| 0.3 | 5.7 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.3 | 3.7 | GO:0072010 | renal filtration cell differentiation(GO:0061318) glomerular epithelium development(GO:0072010) glomerular visceral epithelial cell differentiation(GO:0072112) glomerular epithelial cell differentiation(GO:0072311) |
| 0.3 | 19.2 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.3 | 1.3 | GO:0033136 | serine phosphorylation of STAT3 protein(GO:0033136) serine phosphorylation of STAT protein(GO:0042501) |
| 0.2 | 8.0 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.2 | 0.7 | GO:0043201 | response to leucine(GO:0043201) cellular response to leucine(GO:0071233) |
| 0.2 | 2.3 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.2 | 2.1 | GO:1904262 | negative regulation of TORC1 signaling(GO:1904262) |
| 0.1 | 2.8 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.1 | 1.6 | GO:0021940 | positive regulation of cerebellar granule cell precursor proliferation(GO:0021940) |
| 0.1 | 1.3 | GO:0070445 | radial glial cell differentiation(GO:0060019) oligodendrocyte progenitor proliferation(GO:0070444) regulation of oligodendrocyte progenitor proliferation(GO:0070445) |
| 0.1 | 1.6 | GO:1900454 | positive regulation of long term synaptic depression(GO:1900454) |
| 0.1 | 0.4 | GO:2001012 | mesenchymal cell differentiation involved in kidney development(GO:0072161) mesenchymal cell differentiation involved in renal system development(GO:2001012) |
| 0.1 | 7.6 | GO:0048663 | neuron fate commitment(GO:0048663) |
| 0.1 | 6.5 | GO:0072384 | organelle transport along microtubule(GO:0072384) |
| 0.1 | 0.2 | GO:0060821 | inactivation of X chromosome by DNA methylation(GO:0060821) |
| 0.1 | 2.8 | GO:0051926 | negative regulation of calcium ion transport(GO:0051926) |
| 0.1 | 2.1 | GO:0071480 | cellular response to gamma radiation(GO:0071480) |
| 0.0 | 1.5 | GO:1900047 | negative regulation of blood coagulation(GO:0030195) negative regulation of hemostasis(GO:1900047) |
| 0.0 | 0.9 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.0 | 0.2 | GO:0060406 | positive regulation of penile erection(GO:0060406) |
| 0.0 | 1.3 | GO:0045744 | negative regulation of G-protein coupled receptor protein signaling pathway(GO:0045744) |
| 0.0 | 0.6 | GO:0015991 | energy coupled proton transmembrane transport, against electrochemical gradient(GO:0015988) ATP hydrolysis coupled proton transport(GO:0015991) ATP hydrolysis coupled transmembrane transport(GO:0090662) |
| 0.0 | 2.6 | GO:0000187 | activation of MAPK activity(GO:0000187) |
| 0.0 | 1.4 | GO:0051453 | regulation of intracellular pH(GO:0051453) |
| 0.0 | 1.7 | GO:0034446 | substrate adhesion-dependent cell spreading(GO:0034446) |
| 0.0 | 1.6 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.0 | 0.4 | GO:0035458 | cellular response to interferon-beta(GO:0035458) |
| 0.0 | 0.5 | GO:0001919 | regulation of receptor recycling(GO:0001919) |
| 0.0 | 1.4 | GO:0046488 | phosphatidylinositol metabolic process(GO:0046488) |
| 0.0 | 4.7 | GO:0007186 | G-protein coupled receptor signaling pathway(GO:0007186) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 9.2 | GO:0032937 | SREBP-SCAP-Insig complex(GO:0032937) |
| 0.9 | 7.7 | GO:0005883 | neurofilament(GO:0005883) |
| 0.5 | 6.5 | GO:0071598 | neuronal ribonucleoprotein granule(GO:0071598) |
| 0.4 | 1.3 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 0.4 | 13.5 | GO:0030673 | axolemma(GO:0030673) |
| 0.3 | 7.1 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.3 | 3.7 | GO:0034361 | very-low-density lipoprotein particle(GO:0034361) triglyceride-rich lipoprotein particle(GO:0034385) |
| 0.1 | 2.8 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.1 | 5.7 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.1 | 7.1 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.1 | 0.9 | GO:0060053 | neurofilament cytoskeleton(GO:0060053) |
| 0.1 | 3.7 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.1 | 19.2 | GO:0045211 | postsynaptic membrane(GO:0045211) |
| 0.1 | 2.8 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.1 | 0.6 | GO:0033179 | proton-transporting V-type ATPase, V0 domain(GO:0033179) |
| 0.0 | 0.2 | GO:0001740 | Barr body(GO:0001740) |
| 0.0 | 7.6 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 1.6 | GO:0043235 | receptor complex(GO:0043235) |
| 0.0 | 2.1 | GO:0043679 | axon terminus(GO:0043679) |
| 0.0 | 10.8 | GO:0005887 | integral component of plasma membrane(GO:0005887) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 7.6 | GO:0033691 | sialic acid binding(GO:0033691) |
| 1.5 | 13.5 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 1.2 | 3.7 | GO:0034189 | very-low-density lipoprotein particle binding(GO:0034189) glycoprotein transporter activity(GO:0034437) |
| 0.6 | 2.3 | GO:0000832 | inositol hexakisphosphate kinase activity(GO:0000828) inositol hexakisphosphate 5-kinase activity(GO:0000832) inositol hexakisphosphate 1-kinase activity(GO:0052723) inositol hexakisphosphate 3-kinase activity(GO:0052724) |
| 0.5 | 2.9 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.5 | 2.4 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.5 | 2.8 | GO:0051429 | corticotropin-releasing hormone receptor binding(GO:0051429) corticotropin-releasing hormone receptor 1 binding(GO:0051430) |
| 0.4 | 1.3 | GO:0030549 | acetylcholine receptor activator activity(GO:0030549) |
| 0.2 | 7.1 | GO:0034930 | N-acetylglucosamine 6-O-sulfotransferase activity(GO:0001517) heparan sulfate 2-O-sulfotransferase activity(GO:0004394) HNK-1 sulfotransferase activity(GO:0016232) heparan sulfate 6-O-sulfotransferase activity(GO:0017095) trans-9R,10R-dihydrodiolphenanthrene sulfotransferase activity(GO:0018721) 1-phenanthrol sulfotransferase activity(GO:0018722) 3-phenanthrol sulfotransferase activity(GO:0018723) 4-phenanthrol sulfotransferase activity(GO:0018724) trans-3,4-dihydrodiolphenanthrene sulfotransferase activity(GO:0018725) 9-phenanthrol sulfotransferase activity(GO:0018726) 2-phenanthrol sulfotransferase activity(GO:0018727) phenanthrol sulfotransferase activity(GO:0019111) 1-hydroxypyrene sulfotransferase activity(GO:0034930) proteoglycan sulfotransferase activity(GO:0050698) cholesterol sulfotransferase activity(GO:0051922) hydroxyjasmonate sulfotransferase activity(GO:0080131) |
| 0.2 | 1.9 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
| 0.2 | 7.1 | GO:0030552 | cAMP binding(GO:0030552) |
| 0.1 | 1.5 | GO:0050072 | m7G(5')pppN diphosphatase activity(GO:0050072) |
| 0.1 | 0.7 | GO:0070728 | leucine binding(GO:0070728) |
| 0.1 | 9.4 | GO:0015297 | antiporter activity(GO:0015297) |
| 0.1 | 3.8 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.1 | 6.0 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.1 | 0.4 | GO:0008853 | exodeoxyribonuclease III activity(GO:0008853) |
| 0.1 | 0.1 | GO:0004065 | arylsulfatase activity(GO:0004065) |
| 0.1 | 0.9 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.1 | 1.5 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.1 | 0.4 | GO:0004887 | thyroid hormone receptor activity(GO:0004887) |
| 0.1 | 4.3 | GO:0002039 | p53 binding(GO:0002039) |
| 0.1 | 0.2 | GO:0031686 | A1 adenosine receptor binding(GO:0031686) |
| 0.0 | 0.6 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 1.6 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.0 | 2.5 | GO:0004714 | transmembrane receptor protein tyrosine kinase activity(GO:0004714) |
| 0.0 | 5.5 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.0 | 2.6 | GO:0004713 | protein tyrosine kinase activity(GO:0004713) |
| 0.0 | 1.6 | GO:0008528 | G-protein coupled peptide receptor activity(GO:0008528) |
| 0.0 | 3.7 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.0 | 0.3 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.0 | 4.6 | GO:0005516 | calmodulin binding(GO:0005516) |
| 0.0 | 3.1 | GO:0000287 | magnesium ion binding(GO:0000287) |
| 0.0 | 2.1 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.0 | 7.2 | GO:0008134 | transcription factor binding(GO:0008134) |
| 0.0 | 6.7 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.0 | 12.2 | GO:0008270 | zinc ion binding(GO:0008270) |
| 0.0 | 0.1 | GO:0005540 | hyaluronic acid binding(GO:0005540) |