Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
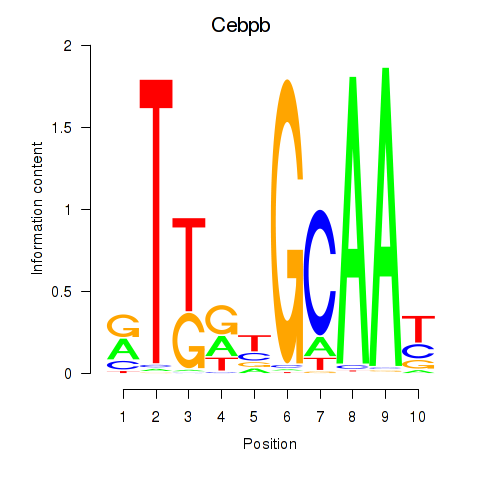
Results for Cebpb
Z-value: 2.97
Transcription factors associated with Cebpb
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Cebpb
|
ENSMUSG00000056501.4 | Cebpb |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Cebpb | mm39_v1_chr2_+_167530835_167530894 | 0.64 | 1.3e-09 | Click! |
Activity profile of Cebpb motif
Sorted Z-values of Cebpb motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Cebpb
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr10_+_93324624 | 42.67 |
ENSMUST00000129421.8
|
Hal
|
histidine ammonia lyase |
| chr16_+_22877000 | 39.25 |
ENSMUST00000039492.14
ENSMUST00000023589.15 ENSMUST00000089902.8 |
Kng1
|
kininogen 1 |
| chr10_+_23727325 | 37.02 |
ENSMUST00000020190.8
|
Vnn3
|
vanin 3 |
| chr3_+_82915031 | 33.09 |
ENSMUST00000048486.13
ENSMUST00000194175.2 |
Fgg
|
fibrinogen gamma chain |
| chr16_+_22710785 | 30.25 |
ENSMUST00000023583.7
ENSMUST00000232098.2 |
Ahsg
|
alpha-2-HS-glycoprotein |
| chr11_+_69983459 | 30.03 |
ENSMUST00000102572.8
|
Asgr2
|
asialoglycoprotein receptor 2 |
| chr14_+_40827108 | 28.79 |
ENSMUST00000224514.2
|
Mat1a
|
methionine adenosyltransferase I, alpha |
| chr3_+_82933383 | 27.99 |
ENSMUST00000029630.15
ENSMUST00000166581.4 |
Fga
|
fibrinogen alpha chain |
| chr11_+_69983479 | 26.54 |
ENSMUST00000143772.8
|
Asgr2
|
asialoglycoprotein receptor 2 |
| chr14_+_40827317 | 25.83 |
ENSMUST00000047286.7
|
Mat1a
|
methionine adenosyltransferase I, alpha |
| chr4_+_63262775 | 25.73 |
ENSMUST00000030044.3
|
Orm1
|
orosomucoid 1 |
| chr14_+_51328534 | 24.69 |
ENSMUST00000022428.13
ENSMUST00000171688.9 |
Rnase4
Ang
|
ribonuclease, RNase A family 4 angiogenin, ribonuclease, RNase A family, 5 |
| chr9_-_71070506 | 24.16 |
ENSMUST00000074465.9
|
Aqp9
|
aquaporin 9 |
| chr14_+_40826970 | 23.58 |
ENSMUST00000225720.2
|
Mat1a
|
methionine adenosyltransferase I, alpha |
| chr5_-_124003553 | 23.24 |
ENSMUST00000057145.7
|
Hcar2
|
hydroxycarboxylic acid receptor 2 |
| chr14_+_75479727 | 23.17 |
ENSMUST00000022576.10
|
Cpb2
|
carboxypeptidase B2 (plasma) |
| chr6_-_124519240 | 22.71 |
ENSMUST00000159463.8
ENSMUST00000162844.2 ENSMUST00000160505.8 ENSMUST00000162443.8 |
C1s1
|
complement component 1, s subcomponent 1 |
| chr12_-_81014849 | 22.37 |
ENSMUST00000095572.5
|
Slc10a1
|
solute carrier family 10 (sodium/bile acid cotransporter family), member 1 |
| chr1_+_67162176 | 22.34 |
ENSMUST00000027144.8
|
Cps1
|
carbamoyl-phosphate synthetase 1 |
| chrX_+_59044796 | 21.72 |
ENSMUST00000033477.5
|
F9
|
coagulation factor IX |
| chr11_+_69983531 | 21.24 |
ENSMUST00000124721.2
|
Asgr2
|
asialoglycoprotein receptor 2 |
| chr19_-_38113249 | 20.81 |
ENSMUST00000112335.4
|
Rbp4
|
retinol binding protein 4, plasma |
| chr17_-_56428968 | 19.82 |
ENSMUST00000041357.9
|
Lrg1
|
leucine-rich alpha-2-glycoprotein 1 |
| chr19_-_38113056 | 19.76 |
ENSMUST00000236283.2
|
Rbp4
|
retinol binding protein 4, plasma |
| chr9_+_107957621 | 19.76 |
ENSMUST00000035211.14
|
Mst1
|
macrophage stimulating 1 (hepatocyte growth factor-like) |
| chr6_-_3968365 | 19.01 |
ENSMUST00000031674.11
|
Tfpi2
|
tissue factor pathway inhibitor 2 |
| chr12_-_81014755 | 18.94 |
ENSMUST00000218342.2
|
Slc10a1
|
solute carrier family 10 (sodium/bile acid cotransporter family), member 1 |
| chr19_-_38113696 | 18.85 |
ENSMUST00000025951.14
ENSMUST00000237287.2 |
Rbp4
|
retinol binding protein 4, plasma |
| chr9_+_107957640 | 18.68 |
ENSMUST00000162886.2
|
Mst1
|
macrophage stimulating 1 (hepatocyte growth factor-like) |
| chr12_+_104304631 | 18.65 |
ENSMUST00000043058.5
ENSMUST00000101078.12 |
Serpina3k
Serpina3m
|
serine (or cysteine) peptidase inhibitor, clade A, member 3K serine (or cysteine) peptidase inhibitor, clade A, member 3M |
| chr11_-_75313412 | 18.58 |
ENSMUST00000138661.8
ENSMUST00000000769.14 |
Serpinf1
|
serine (or cysteine) peptidase inhibitor, clade F, member 1 |
| chr9_-_103107460 | 18.25 |
ENSMUST00000165296.8
ENSMUST00000112645.8 |
Trf
|
transferrin |
| chr6_+_37507108 | 17.83 |
ENSMUST00000040987.11
|
Akr1d1
|
aldo-keto reductase family 1, member D1 |
| chr9_-_103107495 | 17.66 |
ENSMUST00000035158.16
|
Trf
|
transferrin |
| chr11_-_75313350 | 17.33 |
ENSMUST00000125982.2
ENSMUST00000137103.8 |
Serpinf1
|
serine (or cysteine) peptidase inhibitor, clade F, member 1 |
| chr5_-_104125270 | 16.05 |
ENSMUST00000112803.3
|
Hsd17b13
|
hydroxysteroid (17-beta) dehydrogenase 13 |
| chr5_-_104125192 | 15.99 |
ENSMUST00000120320.8
|
Hsd17b13
|
hydroxysteroid (17-beta) dehydrogenase 13 |
| chr12_+_8027640 | 15.94 |
ENSMUST00000171271.8
ENSMUST00000037811.13 |
Apob
|
apolipoprotein B |
| chr12_+_8027767 | 15.87 |
ENSMUST00000037520.14
|
Apob
|
apolipoprotein B |
| chr7_+_27147403 | 15.20 |
ENSMUST00000037399.16
ENSMUST00000108358.8 |
Blvrb
|
biliverdin reductase B (flavin reductase (NADPH)) |
| chr2_+_24235300 | 15.12 |
ENSMUST00000114485.9
ENSMUST00000114482.3 |
Il1rn
|
interleukin 1 receptor antagonist |
| chr16_+_37400500 | 15.08 |
ENSMUST00000160847.2
|
Hgd
|
homogentisate 1, 2-dioxygenase |
| chr1_-_162726234 | 14.88 |
ENSMUST00000111510.8
ENSMUST00000045902.13 |
Fmo2
|
flavin containing monooxygenase 2 |
| chr16_+_37400590 | 14.64 |
ENSMUST00000159787.8
|
Hgd
|
homogentisate 1, 2-dioxygenase |
| chr3_-_89820451 | 14.56 |
ENSMUST00000029559.7
ENSMUST00000197679.5 |
Il6ra
|
interleukin 6 receptor, alpha |
| chr6_+_17463748 | 14.55 |
ENSMUST00000115443.8
|
Met
|
met proto-oncogene |
| chr9_-_109678685 | 14.45 |
ENSMUST00000112022.5
|
Camp
|
cathelicidin antimicrobial peptide |
| chr4_+_63280675 | 14.29 |
ENSMUST00000075341.4
|
Orm2
|
orosomucoid 2 |
| chr7_+_30463175 | 14.23 |
ENSMUST00000165887.8
ENSMUST00000085691.11 ENSMUST00000054427.13 ENSMUST00000085688.11 |
Dmkn
|
dermokine |
| chr3_+_20011201 | 14.22 |
ENSMUST00000091309.12
ENSMUST00000108329.8 ENSMUST00000003714.13 |
Cp
|
ceruloplasmin |
| chr10_+_62860291 | 14.16 |
ENSMUST00000020262.5
|
Pbld2
|
phenazine biosynthesis-like protein domain containing 2 |
| chr11_-_99383938 | 13.99 |
ENSMUST00000006969.8
|
Krt23
|
keratin 23 |
| chr16_+_90535212 | 13.95 |
ENSMUST00000038197.3
|
Mrap
|
melanocortin 2 receptor accessory protein |
| chr5_-_104125226 | 13.90 |
ENSMUST00000048118.15
|
Hsd17b13
|
hydroxysteroid (17-beta) dehydrogenase 13 |
| chr7_-_45109262 | 13.78 |
ENSMUST00000094434.13
|
Ftl1
|
ferritin light polypeptide 1 |
| chr8_+_75820240 | 13.77 |
ENSMUST00000005548.8
|
Hmox1
|
heme oxygenase 1 |
| chr8_+_46924074 | 13.76 |
ENSMUST00000034046.13
ENSMUST00000211644.2 |
Acsl1
|
acyl-CoA synthetase long-chain family member 1 |
| chr19_+_55240357 | 13.71 |
ENSMUST00000225551.2
|
Acsl5
|
acyl-CoA synthetase long-chain family member 5 |
| chr3_+_20011251 | 13.70 |
ENSMUST00000108328.8
|
Cp
|
ceruloplasmin |
| chr7_+_46401214 | 13.69 |
ENSMUST00000210769.2
ENSMUST00000210272.2 ENSMUST00000075982.4 |
Saa2
|
serum amyloid A 2 |
| chr7_-_45109075 | 13.49 |
ENSMUST00000210864.2
|
Ftl1
|
ferritin light polypeptide 1 |
| chr4_-_130068902 | 13.41 |
ENSMUST00000105998.8
|
Tinagl1
|
tubulointerstitial nephritis antigen-like 1 |
| chr5_+_8010445 | 13.37 |
ENSMUST00000115421.3
|
Steap4
|
STEAP family member 4 |
| chr2_-_164198427 | 13.22 |
ENSMUST00000109367.10
|
Slpi
|
secretory leukocyte peptidase inhibitor |
| chr8_-_34419826 | 12.85 |
ENSMUST00000033995.14
ENSMUST00000033994.15 ENSMUST00000191473.7 ENSMUST00000053251.12 |
Rbpms
|
RNA binding protein gene with multiple splicing |
| chr16_-_22847808 | 12.81 |
ENSMUST00000115349.9
|
Kng2
|
kininogen 2 |
| chr17_+_37253802 | 12.54 |
ENSMUST00000040498.12
|
Rnf39
|
ring finger protein 39 |
| chr14_-_55950939 | 12.34 |
ENSMUST00000168729.8
ENSMUST00000228123.2 ENSMUST00000178034.9 |
Tgm1
|
transglutaminase 1, K polypeptide |
| chr10_+_62860094 | 12.28 |
ENSMUST00000124784.8
|
Pbld2
|
phenazine biosynthesis-like protein domain containing 2 |
| chr7_+_27147475 | 12.19 |
ENSMUST00000133750.8
|
Blvrb
|
biliverdin reductase B (flavin reductase (NADPH)) |
| chr16_-_22847760 | 12.13 |
ENSMUST00000039338.13
|
Kng2
|
kininogen 2 |
| chr14_-_55950545 | 11.89 |
ENSMUST00000002389.9
ENSMUST00000226907.2 |
Tgm1
|
transglutaminase 1, K polypeptide |
| chr17_+_57665691 | 11.88 |
ENSMUST00000086763.13
ENSMUST00000233376.2 ENSMUST00000233840.2 ENSMUST00000232808.2 ENSMUST00000004850.8 |
Adgre1
|
adhesion G protein-coupled receptor E1 |
| chr7_+_13467422 | 11.66 |
ENSMUST00000086148.8
|
Sult2a2
|
sulfotransferase family 2A, dehydroepiandrosterone (DHEA)-preferring, member 2 |
| chr1_-_162726053 | 11.62 |
ENSMUST00000143123.3
|
Fmo2
|
flavin containing monooxygenase 2 |
| chr9_-_48516447 | 11.61 |
ENSMUST00000034808.12
ENSMUST00000119426.2 |
Nnmt
|
nicotinamide N-methyltransferase |
| chr3_+_20011405 | 11.28 |
ENSMUST00000108325.9
|
Cp
|
ceruloplasmin |
| chr17_-_74257164 | 11.00 |
ENSMUST00000024866.6
|
Xdh
|
xanthine dehydrogenase |
| chr3_+_106393348 | 10.91 |
ENSMUST00000183271.2
|
Dennd2d
|
DENN/MADD domain containing 2D |
| chr7_-_28078671 | 10.89 |
ENSMUST00000209061.2
|
Zfp36
|
zinc finger protein 36 |
| chr15_-_101848689 | 10.76 |
ENSMUST00000023799.8
|
Krt79
|
keratin 79 |
| chr16_-_22847829 | 10.69 |
ENSMUST00000100046.9
|
Kng2
|
kininogen 2 |
| chr2_-_32277773 | 10.64 |
ENSMUST00000050785.14
|
Lcn2
|
lipocalin 2 |
| chr9_+_21746785 | 10.39 |
ENSMUST00000058777.8
|
Angptl8
|
angiopoietin-like 8 |
| chr6_+_72281587 | 10.13 |
ENSMUST00000183018.8
ENSMUST00000182014.10 |
Sftpb
|
surfactant associated protein B |
| chr2_+_158148413 | 9.88 |
ENSMUST00000109491.8
ENSMUST00000016168.9 |
Lbp
|
lipopolysaccharide binding protein |
| chr9_+_110248815 | 9.38 |
ENSMUST00000035061.9
|
Ngp
|
neutrophilic granule protein |
| chr6_+_17463819 | 9.26 |
ENSMUST00000140070.8
|
Met
|
met proto-oncogene |
| chr7_-_44181477 | 9.17 |
ENSMUST00000098483.9
ENSMUST00000035323.6 |
Spib
|
Spi-B transcription factor (Spi-1/PU.1 related) |
| chr9_+_110867807 | 9.16 |
ENSMUST00000197575.2
|
Ltf
|
lactotransferrin |
| chr13_+_25127127 | 9.15 |
ENSMUST00000021773.13
|
Gpld1
|
glycosylphosphatidylinositol specific phospholipase D1 |
| chr15_+_25940859 | 9.00 |
ENSMUST00000226750.2
|
Retreg1
|
reticulophagy regulator 1 |
| chr2_-_110144869 | 8.99 |
ENSMUST00000133608.2
|
Bbox1
|
butyrobetaine (gamma), 2-oxoglutarate dioxygenase 1 (gamma-butyrobetaine hydroxylase) |
| chr4_-_107928567 | 8.92 |
ENSMUST00000106701.2
|
Scp2
|
sterol carrier protein 2, liver |
| chr7_-_46365108 | 8.90 |
ENSMUST00000006956.9
ENSMUST00000210913.2 |
Saa3
|
serum amyloid A 3 |
| chr3_+_84573499 | 8.86 |
ENSMUST00000107682.2
|
Tmem154
|
transmembrane protein 154 |
| chr16_+_45044678 | 8.83 |
ENSMUST00000102802.10
ENSMUST00000063654.6 |
Btla
|
B and T lymphocyte associated |
| chr1_+_74414354 | 8.80 |
ENSMUST00000187516.7
ENSMUST00000027368.6 |
Slc11a1
|
solute carrier family 11 (proton-coupled divalent metal ion transporters), member 1 |
| chr17_+_37253916 | 8.77 |
ENSMUST00000173072.2
|
Rnf39
|
ring finger protein 39 |
| chr11_-_99280177 | 8.76 |
ENSMUST00000103131.5
|
Krt10
|
keratin 10 |
| chr14_+_51181956 | 8.65 |
ENSMUST00000178092.2
ENSMUST00000227052.2 |
Pnp
Gm49342
|
purine-nucleoside phosphorylase predicted gene, 49342 |
| chr14_-_68893253 | 8.41 |
ENSMUST00000225767.3
ENSMUST00000111072.8 ENSMUST00000022642.6 ENSMUST00000224039.2 |
Adam28
|
a disintegrin and metallopeptidase domain 28 |
| chr2_-_152185901 | 8.38 |
ENSMUST00000040312.7
|
Trib3
|
tribbles pseudokinase 3 |
| chr16_-_22848153 | 8.27 |
ENSMUST00000232459.2
|
Kng2
|
kininogen 2 |
| chr15_-_103161237 | 8.18 |
ENSMUST00000154510.8
|
Nfe2
|
nuclear factor, erythroid derived 2 |
| chr5_+_91222470 | 8.18 |
ENSMUST00000031324.6
|
Ereg
|
epiregulin |
| chr10_-_66932685 | 7.96 |
ENSMUST00000020023.9
|
Reep3
|
receptor accessory protein 3 |
| chr6_+_34686543 | 7.81 |
ENSMUST00000031775.13
|
Cald1
|
caldesmon 1 |
| chrX_-_166047275 | 7.72 |
ENSMUST00000112170.2
|
Tlr8
|
toll-like receptor 8 |
| chrX_-_137985960 | 7.70 |
ENSMUST00000033626.15
ENSMUST00000060824.4 |
Serpina7
|
serine (or cysteine) peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 7 |
| chr18_+_20643325 | 7.67 |
ENSMUST00000070892.8
ENSMUST00000234945.2 |
Dsg3
|
desmoglein 3 |
| chr16_+_57369595 | 7.62 |
ENSMUST00000159414.2
|
Filip1l
|
filamin A interacting protein 1-like |
| chr16_-_93726399 | 7.56 |
ENSMUST00000177648.8
ENSMUST00000142083.2 |
Cldn14
|
claudin 14 |
| chr1_+_164624200 | 7.41 |
ENSMUST00000027861.6
|
Dpt
|
dermatopontin |
| chr19_+_33985684 | 7.39 |
ENSMUST00000225505.2
|
Lipk
|
lipase, family member K |
| chr1_-_131204651 | 7.16 |
ENSMUST00000161764.8
|
Ikbke
|
inhibitor of kappaB kinase epsilon |
| chr19_-_34724689 | 7.05 |
ENSMUST00000009522.4
|
Slc16a12
|
solute carrier family 16 (monocarboxylic acid transporters), member 12 |
| chr19_+_33985701 | 7.02 |
ENSMUST00000054260.8
|
Lipk
|
lipase, family member K |
| chr3_-_59102517 | 6.96 |
ENSMUST00000200095.2
|
Gpr87
|
G protein-coupled receptor 87 |
| chr19_-_11618192 | 6.66 |
ENSMUST00000112984.4
|
Ms4a3
|
membrane-spanning 4-domains, subfamily A, member 3 |
| chr16_+_57173456 | 6.66 |
ENSMUST00000159816.8
|
Filip1l
|
filamin A interacting protein 1-like |
| chr6_-_87786736 | 6.60 |
ENSMUST00000032134.9
|
Rab43
|
RAB43, member RAS oncogene family |
| chr1_-_131204422 | 6.53 |
ENSMUST00000159195.2
|
Ikbke
|
inhibitor of kappaB kinase epsilon |
| chr6_+_115398996 | 6.51 |
ENSMUST00000000450.5
|
Pparg
|
peroxisome proliferator activated receptor gamma |
| chr10_-_127358300 | 6.51 |
ENSMUST00000026470.6
|
Shmt2
|
serine hydroxymethyltransferase 2 (mitochondrial) |
| chr15_+_25940912 | 6.51 |
ENSMUST00000226438.2
|
Retreg1
|
reticulophagy regulator 1 |
| chr19_-_11618165 | 6.37 |
ENSMUST00000186023.7
|
Ms4a3
|
membrane-spanning 4-domains, subfamily A, member 3 |
| chr7_+_13357892 | 6.28 |
ENSMUST00000108525.4
|
Sult2a5
|
sulfotransferase family 2A, dehydroepiandrosterone (DHEA)-preferring, member 5 |
| chr15_-_101759212 | 6.27 |
ENSMUST00000023790.5
|
Krt1
|
keratin 1 |
| chr2_-_32278245 | 6.26 |
ENSMUST00000192241.2
|
Lcn2
|
lipocalin 2 |
| chr19_-_34855278 | 6.18 |
ENSMUST00000112460.3
|
Pank1
|
pantothenate kinase 1 |
| chr9_+_7558449 | 6.07 |
ENSMUST00000018765.4
|
Mmp8
|
matrix metallopeptidase 8 |
| chr13_-_53135064 | 6.01 |
ENSMUST00000071065.8
|
Nfil3
|
nuclear factor, interleukin 3, regulated |
| chr10_+_97318223 | 5.92 |
ENSMUST00000163448.4
|
Dcn
|
decorin |
| chr3_-_121608809 | 5.86 |
ENSMUST00000197383.5
|
Abcd3
|
ATP-binding cassette, sub-family D (ALD), member 3 |
| chr10_-_127358231 | 5.82 |
ENSMUST00000219239.2
|
Shmt2
|
serine hydroxymethyltransferase 2 (mitochondrial) |
| chr7_-_126873219 | 5.77 |
ENSMUST00000082428.6
|
Sephs2
|
selenophosphate synthetase 2 |
| chr19_-_34855242 | 5.63 |
ENSMUST00000238065.2
|
Pank1
|
pantothenate kinase 1 |
| chr7_-_46392403 | 5.63 |
ENSMUST00000128088.4
|
Saa1
|
serum amyloid A 1 |
| chr9_-_71678814 | 5.52 |
ENSMUST00000122065.2
ENSMUST00000121322.8 ENSMUST00000072899.9 |
Cgnl1
|
cingulin-like 1 |
| chr1_-_106980033 | 5.51 |
ENSMUST00000112717.3
|
Serpinb3a
|
serine (or cysteine) peptidase inhibitor, clade B (ovalbumin), member 3A |
| chr6_+_141194886 | 5.51 |
ENSMUST00000043259.10
|
Pde3a
|
phosphodiesterase 3A, cGMP inhibited |
| chr17_+_29333116 | 5.44 |
ENSMUST00000233717.2
ENSMUST00000141239.2 |
Rab44
|
RAB44, member RAS oncogene family |
| chr9_+_123902143 | 5.42 |
ENSMUST00000168841.3
ENSMUST00000055918.7 |
Ccr2
|
chemokine (C-C motif) receptor 2 |
| chr7_+_78563150 | 5.41 |
ENSMUST00000118867.8
|
Isg20
|
interferon-stimulated protein |
| chr19_+_6451667 | 5.39 |
ENSMUST00000113471.3
ENSMUST00000113469.3 |
Rasgrp2
|
RAS, guanyl releasing protein 2 |
| chr3_-_92528480 | 5.37 |
ENSMUST00000170676.3
|
Lce6a
|
late cornified envelope 6A |
| chr11_-_115968745 | 5.33 |
ENSMUST00000156545.2
ENSMUST00000075036.9 ENSMUST00000106451.8 |
Unc13d
|
unc-13 homolog D |
| chr3_-_121608859 | 5.29 |
ENSMUST00000029770.8
|
Abcd3
|
ATP-binding cassette, sub-family D (ALD), member 3 |
| chr11_+_115353290 | 5.22 |
ENSMUST00000106532.4
ENSMUST00000092445.12 ENSMUST00000153466.2 |
Slc16a5
|
solute carrier family 16 (monocarboxylic acid transporters), member 5 |
| chr6_-_11907392 | 5.20 |
ENSMUST00000204084.3
ENSMUST00000031637.8 ENSMUST00000204978.3 ENSMUST00000204714.2 |
Ndufa4
|
Ndufa4, mitochondrial complex associated |
| chr13_+_51254852 | 5.11 |
ENSMUST00000095797.6
|
Spin1
|
spindlin 1 |
| chr18_-_43925932 | 5.10 |
ENSMUST00000237926.2
ENSMUST00000096570.4 |
Gm94
|
predicted gene 94 |
| chr6_-_123266820 | 5.09 |
ENSMUST00000032239.11
ENSMUST00000177367.8 |
Clec4e
|
C-type lectin domain family 4, member e |
| chr4_+_114914607 | 5.03 |
ENSMUST00000136946.8
|
Tal1
|
T cell acute lymphocytic leukemia 1 |
| chr19_+_34268053 | 4.97 |
ENSMUST00000025691.13
|
Fas
|
Fas (TNF receptor superfamily member 6) |
| chr18_+_4994600 | 4.97 |
ENSMUST00000140448.8
|
Svil
|
supervillin |
| chr6_+_40605758 | 4.91 |
ENSMUST00000202636.4
ENSMUST00000201148.4 ENSMUST00000071535.10 |
Mgam
|
maltase-glucoamylase |
| chr13_-_61084358 | 4.84 |
ENSMUST00000225859.2
ENSMUST00000225167.2 ENSMUST00000021880.10 |
Gm49391
Ctla2a
|
predicted gene, 49391 cytotoxic T lymphocyte-associated protein 2 alpha |
| chr10_+_102348076 | 4.70 |
ENSMUST00000219445.2
|
Rassf9
|
Ras association (RalGDS/AF-6) domain family (N-terminal) member 9 |
| chrX_-_137985979 | 4.67 |
ENSMUST00000152457.2
|
Serpina7
|
serine (or cysteine) peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 7 |
| chr11_+_97697128 | 4.67 |
ENSMUST00000138919.2
|
Lasp1
|
LIM and SH3 protein 1 |
| chr13_+_31809774 | 4.65 |
ENSMUST00000042054.3
|
Foxf2
|
forkhead box F2 |
| chr15_+_25940781 | 4.64 |
ENSMUST00000227275.2
|
Retreg1
|
reticulophagy regulator 1 |
| chr8_+_95113066 | 4.64 |
ENSMUST00000161576.8
ENSMUST00000034220.8 |
Herpud1
|
homocysteine-inducible, endoplasmic reticulum stress-inducible, ubiquitin-like domain member 1 |
| chr17_+_35863025 | 4.61 |
ENSMUST00000044804.8
|
Cdsn
|
corneodesmosin |
| chr15_+_59520199 | 4.56 |
ENSMUST00000067543.8
|
Trib1
|
tribbles pseudokinase 1 |
| chr17_+_29077385 | 4.47 |
ENSMUST00000056866.8
|
Pnpla1
|
patatin-like phospholipase domain containing 1 |
| chr1_+_40123858 | 4.45 |
ENSMUST00000027243.13
|
Il1r2
|
interleukin 1 receptor, type II |
| chr4_+_155646807 | 4.43 |
ENSMUST00000030939.14
|
Nadk
|
NAD kinase |
| chr16_+_42727926 | 4.40 |
ENSMUST00000151244.8
ENSMUST00000114694.9 |
Zbtb20
|
zinc finger and BTB domain containing 20 |
| chr19_+_34268071 | 4.39 |
ENSMUST00000112472.4
ENSMUST00000235232.2 |
Fas
|
Fas (TNF receptor superfamily member 6) |
| chr11_-_33153587 | 4.34 |
ENSMUST00000037746.8
|
Tlx3
|
T cell leukemia, homeobox 3 |
| chr18_-_52662917 | 4.34 |
ENSMUST00000171470.8
|
Lox
|
lysyl oxidase |
| chr14_+_66208253 | 4.30 |
ENSMUST00000138191.8
|
Clu
|
clusterin |
| chr11_-_115968576 | 4.29 |
ENSMUST00000106450.8
|
Unc13d
|
unc-13 homolog D |
| chr18_+_44237577 | 4.27 |
ENSMUST00000239465.2
|
Spink12
|
serine peptidase inhibitor, Kazal type 12 |
| chr13_+_16186410 | 4.25 |
ENSMUST00000042603.14
|
Inhba
|
inhibin beta-A |
| chr1_-_107088844 | 4.24 |
ENSMUST00000086694.11
|
Serpinb3b
|
serine (or cysteine) peptidase inhibitor, clade B (ovalbumin), member 3B |
| chr7_+_30475819 | 4.20 |
ENSMUST00000041703.10
|
Dmkn
|
dermokine |
| chr2_+_155593030 | 4.17 |
ENSMUST00000029140.12
ENSMUST00000132608.2 |
Procr
|
protein C receptor, endothelial |
| chr3_+_60436163 | 4.12 |
ENSMUST00000194201.6
ENSMUST00000193517.6 |
Mbnl1
|
muscleblind like splicing factor 1 |
| chr15_+_25940931 | 4.10 |
ENSMUST00000110438.3
|
Retreg1
|
reticulophagy regulator 1 |
| chr19_+_12438125 | 4.08 |
ENSMUST00000081035.9
|
Mpeg1
|
macrophage expressed gene 1 |
| chrX_-_166047289 | 4.04 |
ENSMUST00000133722.2
|
Tlr8
|
toll-like receptor 8 |
| chr15_+_59520493 | 3.99 |
ENSMUST00000118228.2
|
Trib1
|
tribbles pseudokinase 1 |
| chr2_-_118378020 | 3.97 |
ENSMUST00000110859.3
|
Bmf
|
BCL2 modifying factor |
| chr11_-_83540175 | 3.87 |
ENSMUST00000001008.6
|
Ccl3
|
chemokine (C-C motif) ligand 3 |
| chr10_+_79722081 | 3.74 |
ENSMUST00000046091.7
|
Elane
|
elastase, neutrophil expressed |
| chr2_+_117942357 | 3.73 |
ENSMUST00000039559.9
|
Thbs1
|
thrombospondin 1 |
| chr11_-_115968373 | 3.67 |
ENSMUST00000174822.8
|
Unc13d
|
unc-13 homolog D |
| chr12_-_101784727 | 3.67 |
ENSMUST00000222587.2
|
Fbln5
|
fibulin 5 |
| chr11_-_100139728 | 3.65 |
ENSMUST00000007280.9
|
Krt16
|
keratin 16 |
| chr6_-_129428746 | 3.64 |
ENSMUST00000204012.2
ENSMUST00000037481.10 |
Clec1a
|
C-type lectin domain family 1, member a |
| chr17_-_29226965 | 3.58 |
ENSMUST00000009138.13
ENSMUST00000119274.3 |
Stk38
|
serine/threonine kinase 38 |
| chr6_-_129428869 | 3.55 |
ENSMUST00000203162.3
|
Clec1a
|
C-type lectin domain family 1, member a |
| chr9_-_55190902 | 3.50 |
ENSMUST00000164721.8
|
Nrg4
|
neuregulin 4 |
| chr17_-_57501170 | 3.48 |
ENSMUST00000005976.8
|
Tnfsf14
|
tumor necrosis factor (ligand) superfamily, member 14 |
| chr18_-_52662728 | 3.45 |
ENSMUST00000025409.9
|
Lox
|
lysyl oxidase |
| chr13_-_40887444 | 3.43 |
ENSMUST00000224665.2
|
Tfap2a
|
transcription factor AP-2, alpha |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 26.1 | 78.2 | GO:0009087 | methionine catabolic process(GO:0009087) |
| 11.9 | 59.4 | GO:0048807 | female genitalia morphogenesis(GO:0048807) |
| 8.5 | 42.7 | GO:0006548 | histidine catabolic process(GO:0006548) |
| 7.7 | 23.2 | GO:0003331 | regulation of extracellular matrix constituent secretion(GO:0003330) positive regulation of extracellular matrix constituent secretion(GO:0003331) |
| 7.0 | 28.0 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 6.0 | 24.2 | GO:0015855 | nucleobase transport(GO:0015851) pyrimidine nucleobase transport(GO:0015855) |
| 5.9 | 17.8 | GO:0030573 | bile acid catabolic process(GO:0030573) |
| 5.6 | 16.9 | GO:0015688 | iron chelate transport(GO:0015688) siderophore transport(GO:0015891) |
| 5.1 | 41.2 | GO:0042167 | porphyrin-containing compound catabolic process(GO:0006787) tetrapyrrole catabolic process(GO:0033015) heme catabolic process(GO:0042167) pigment catabolic process(GO:0046149) |
| 4.9 | 14.6 | GO:0010536 | positive regulation of activation of Janus kinase activity(GO:0010536) |
| 4.6 | 18.4 | GO:1903575 | cornified envelope assembly(GO:1903575) |
| 4.5 | 35.9 | GO:0070447 | positive regulation of oligodendrocyte progenitor proliferation(GO:0070447) |
| 4.4 | 26.5 | GO:0072592 | oxygen metabolic process(GO:0072592) |
| 4.3 | 38.4 | GO:0060763 | mammary duct terminal end bud growth(GO:0060763) |
| 4.1 | 12.4 | GO:0033189 | response to vitamin A(GO:0033189) |
| 4.1 | 24.7 | GO:0006651 | diacylglycerol biosynthetic process(GO:0006651) positive regulation of phospholipase A2 activity(GO:0032430) |
| 4.1 | 12.3 | GO:0019264 | glycine biosynthetic process from serine(GO:0019264) |
| 3.9 | 19.6 | GO:2000660 | negative regulation of interleukin-1-mediated signaling pathway(GO:2000660) |
| 3.7 | 3.7 | GO:0045079 | negative regulation of chemokine biosynthetic process(GO:0045079) |
| 3.6 | 10.9 | GO:1904582 | positive regulation of intracellular mRNA localization(GO:1904582) |
| 3.4 | 23.8 | GO:0070494 | regulation of thrombin receptor signaling pathway(GO:0070494) negative regulation of thrombin receptor signaling pathway(GO:0070495) |
| 3.3 | 29.7 | GO:0006572 | tyrosine catabolic process(GO:0006572) |
| 3.3 | 9.9 | GO:0015920 | lipopolysaccharide transport(GO:0015920) |
| 3.2 | 25.9 | GO:0070163 | adiponectin secretion(GO:0070162) regulation of adiponectin secretion(GO:0070163) |
| 3.1 | 9.4 | GO:0031104 | dendrite regeneration(GO:0031104) |
| 3.1 | 21.5 | GO:0006207 | 'de novo' pyrimidine nucleobase biosynthetic process(GO:0006207) pyrimidine nucleobase biosynthetic process(GO:0019856) |
| 3.1 | 9.2 | GO:0010040 | response to iron(II) ion(GO:0010040) |
| 3.0 | 35.9 | GO:0060770 | negative regulation of epithelial cell proliferation involved in prostate gland development(GO:0060770) |
| 2.9 | 8.8 | GO:0070839 | divalent metal ion export(GO:0070839) |
| 2.9 | 8.7 | GO:0046122 | purine deoxyribonucleoside metabolic process(GO:0046122) |
| 2.9 | 8.6 | GO:0045645 | regulation of eosinophil differentiation(GO:0045643) positive regulation of eosinophil differentiation(GO:0045645) |
| 2.8 | 11.1 | GO:0015910 | peroxisomal long-chain fatty acid import(GO:0015910) |
| 2.8 | 33.1 | GO:0072378 | blood coagulation, fibrin clot formation(GO:0072378) |
| 2.7 | 13.7 | GO:0010747 | positive regulation of plasma membrane long-chain fatty acid transport(GO:0010747) |
| 2.7 | 8.2 | GO:0002436 | immune complex clearance by monocytes and macrophages(GO:0002436) regulation of immune complex clearance by monocytes and macrophages(GO:0090264) positive regulation of immune complex clearance by monocytes and macrophages(GO:0090265) negative regulation of eosinophil activation(GO:1902567) positive regulation of CD8-positive, alpha-beta T cell extravasation(GO:2000451) |
| 2.7 | 27.3 | GO:0097577 | intracellular sequestering of iron ion(GO:0006880) sequestering of iron ion(GO:0097577) |
| 2.7 | 24.2 | GO:0061709 | reticulophagy(GO:0061709) |
| 2.4 | 31.8 | GO:0010886 | positive regulation of cholesterol storage(GO:0010886) |
| 2.4 | 24.2 | GO:0010838 | positive regulation of keratinocyte proliferation(GO:0010838) |
| 2.4 | 7.1 | GO:0015881 | creatine transport(GO:0015881) |
| 2.3 | 9.4 | GO:1901491 | negative regulation of lymphangiogenesis(GO:1901491) |
| 2.3 | 9.2 | GO:0051673 | biofilm formation(GO:0042710) single-species biofilm formation(GO:0044010) single-species biofilm formation in or on host organism(GO:0044407) membrane disruption in other organism(GO:0051673) regulation of single-species biofilm formation(GO:1900190) negative regulation of single-species biofilm formation(GO:1900191) regulation of single-species biofilm formation in or on host organism(GO:1900228) negative regulation of single-species biofilm formation in or on host organism(GO:1900229) |
| 2.2 | 9.0 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 2.2 | 8.9 | GO:0032382 | positive regulation of intracellular lipid transport(GO:0032379) positive regulation of intracellular sterol transport(GO:0032382) positive regulation of intracellular cholesterol transport(GO:0032385) lipid hydroperoxide transport(GO:1901373) |
| 2.2 | 29.0 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 2.2 | 6.6 | GO:0035526 | retrograde transport, plasma membrane to Golgi(GO:0035526) |
| 2.2 | 11.0 | GO:0009115 | xanthine catabolic process(GO:0009115) xanthine metabolic process(GO:0046110) |
| 1.9 | 11.3 | GO:1902998 | regulation of neuronal signal transduction(GO:1902847) positive regulation of neurofibrillary tangle assembly(GO:1902998) |
| 1.8 | 98.5 | GO:0006953 | acute-phase response(GO:0006953) |
| 1.8 | 10.7 | GO:0045356 | positive regulation of interferon-alpha biosynthetic process(GO:0045356) |
| 1.7 | 10.4 | GO:0051005 | negative regulation of lipoprotein lipase activity(GO:0051005) |
| 1.7 | 5.0 | GO:0060217 | hemangioblast cell differentiation(GO:0060217) |
| 1.7 | 13.3 | GO:0002432 | granuloma formation(GO:0002432) |
| 1.6 | 8.2 | GO:0048165 | ovarian cumulus expansion(GO:0001550) fused antrum stage(GO:0048165) |
| 1.6 | 6.5 | GO:2000230 | negative regulation of pancreatic stellate cell proliferation(GO:2000230) |
| 1.6 | 8.0 | GO:0007084 | mitotic nuclear envelope reassembly(GO:0007084) |
| 1.5 | 53.8 | GO:0006825 | copper ion transport(GO:0006825) |
| 1.4 | 14.3 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 1.4 | 78.1 | GO:0050819 | negative regulation of coagulation(GO:0050819) |
| 1.3 | 41.3 | GO:0015721 | bile acid and bile salt transport(GO:0015721) |
| 1.3 | 4.0 | GO:1904092 | regulation of autophagic cell death(GO:1904092) negative regulation of autophagic cell death(GO:1904093) |
| 1.3 | 7.8 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 1.3 | 13.8 | GO:0044539 | long-chain fatty acid import(GO:0044539) |
| 1.2 | 14.5 | GO:0002227 | innate immune response in mucosa(GO:0002227) |
| 1.2 | 3.5 | GO:0008588 | release of cytoplasmic sequestered NF-kappaB(GO:0008588) |
| 1.1 | 4.3 | GO:0060279 | positive regulation of ovulation(GO:0060279) |
| 0.9 | 37.0 | GO:0006767 | water-soluble vitamin metabolic process(GO:0006767) |
| 0.9 | 5.5 | GO:0060282 | positive regulation of oocyte development(GO:0060282) |
| 0.9 | 2.7 | GO:1904211 | membrane protein proteolysis involved in retrograde protein transport, ER to cytosol(GO:1904211) |
| 0.8 | 5.9 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.8 | 5.8 | GO:0016259 | selenocysteine metabolic process(GO:0016259) |
| 0.8 | 3.2 | GO:0042851 | L-alanine metabolic process(GO:0042851) |
| 0.8 | 7.0 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 0.8 | 4.6 | GO:1903071 | positive regulation of ER-associated ubiquitin-dependent protein catabolic process(GO:1903071) |
| 0.7 | 11.8 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.7 | 2.8 | GO:0006049 | UDP-N-acetylglucosamine catabolic process(GO:0006049) |
| 0.7 | 5.5 | GO:0042270 | protection from natural killer cell mediated cytotoxicity(GO:0042270) |
| 0.7 | 3.4 | GO:0003409 | optic cup structural organization(GO:0003409) |
| 0.7 | 2.0 | GO:0071726 | response to diacyl bacterial lipopeptide(GO:0071724) cellular response to diacyl bacterial lipopeptide(GO:0071726) |
| 0.7 | 13.2 | GO:0019731 | antibacterial humoral response(GO:0019731) |
| 0.6 | 2.6 | GO:1903984 | positive regulation of TRAIL-activated apoptotic signaling pathway(GO:1903984) |
| 0.6 | 5.1 | GO:0038094 | Fc-gamma receptor signaling pathway(GO:0038094) |
| 0.6 | 13.7 | GO:0010884 | positive regulation of lipid storage(GO:0010884) |
| 0.5 | 8.2 | GO:0034242 | negative regulation of syncytium formation by plasma membrane fusion(GO:0034242) |
| 0.5 | 3.2 | GO:0046878 | positive regulation of saliva secretion(GO:0046878) |
| 0.5 | 8.8 | GO:0046642 | negative regulation of alpha-beta T cell proliferation(GO:0046642) |
| 0.5 | 35.6 | GO:0050873 | brown fat cell differentiation(GO:0050873) |
| 0.5 | 12.9 | GO:0060391 | positive regulation of SMAD protein import into nucleus(GO:0060391) |
| 0.5 | 10.1 | GO:0045717 | negative regulation of fatty acid biosynthetic process(GO:0045717) |
| 0.5 | 4.4 | GO:0006741 | NADP biosynthetic process(GO:0006741) |
| 0.5 | 1.5 | GO:0009397 | 10-formyltetrahydrofolate catabolic process(GO:0009258) folic acid-containing compound catabolic process(GO:0009397) pteridine-containing compound catabolic process(GO:0042560) |
| 0.5 | 72.6 | GO:0055088 | lipid homeostasis(GO:0055088) |
| 0.5 | 8.8 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.4 | 44.0 | GO:0046889 | positive regulation of lipid biosynthetic process(GO:0046889) |
| 0.4 | 1.3 | GO:0060672 | epithelial cell differentiation involved in embryonic placenta development(GO:0060671) epithelial cell morphogenesis involved in placental branching(GO:0060672) |
| 0.4 | 2.7 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.4 | 5.6 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.4 | 1.1 | GO:0000448 | cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000448) |
| 0.4 | 3.2 | GO:2000298 | regulation of Rho-dependent protein serine/threonine kinase activity(GO:2000298) |
| 0.3 | 4.2 | GO:2000675 | negative regulation of type B pancreatic cell apoptotic process(GO:2000675) |
| 0.3 | 3.6 | GO:0051546 | keratinocyte migration(GO:0051546) |
| 0.3 | 0.9 | GO:0006419 | alanyl-tRNA aminoacylation(GO:0006419) |
| 0.3 | 4.6 | GO:0043589 | skin morphogenesis(GO:0043589) |
| 0.3 | 2.4 | GO:0002326 | B cell lineage commitment(GO:0002326) |
| 0.3 | 3.9 | GO:0043615 | astrocyte cell migration(GO:0043615) |
| 0.2 | 1.7 | GO:0007527 | adult somatic muscle development(GO:0007527) |
| 0.2 | 5.6 | GO:0030574 | collagen catabolic process(GO:0030574) |
| 0.2 | 3.4 | GO:0044154 | histone H3-K14 acetylation(GO:0044154) |
| 0.2 | 9.2 | GO:0030225 | macrophage differentiation(GO:0030225) |
| 0.2 | 2.1 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) |
| 0.2 | 4.6 | GO:0042249 | establishment of planar polarity of embryonic epithelium(GO:0042249) |
| 0.2 | 4.3 | GO:0002087 | regulation of respiratory gaseous exchange by neurological system process(GO:0002087) |
| 0.2 | 7.4 | GO:0030199 | collagen fibril organization(GO:0030199) |
| 0.2 | 5.0 | GO:0045589 | regulation of regulatory T cell differentiation(GO:0045589) |
| 0.2 | 3.4 | GO:0090026 | positive regulation of monocyte chemotaxis(GO:0090026) |
| 0.2 | 3.4 | GO:0006337 | nucleosome disassembly(GO:0006337) |
| 0.2 | 34.6 | GO:0007596 | blood coagulation(GO:0007596) coagulation(GO:0050817) |
| 0.2 | 5.1 | GO:0009303 | rRNA transcription(GO:0009303) |
| 0.2 | 6.0 | GO:0071353 | cellular response to interleukin-4(GO:0071353) |
| 0.1 | 1.1 | GO:0042226 | interleukin-6 biosynthetic process(GO:0042226) |
| 0.1 | 2.5 | GO:0007250 | activation of NF-kappaB-inducing kinase activity(GO:0007250) |
| 0.1 | 5.7 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.1 | 0.8 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) |
| 0.1 | 3.2 | GO:0040034 | regulation of development, heterochronic(GO:0040034) |
| 0.1 | 1.1 | GO:2000254 | regulation of male germ cell proliferation(GO:2000254) |
| 0.1 | 0.3 | GO:0034374 | low-density lipoprotein particle remodeling(GO:0034374) |
| 0.1 | 7.8 | GO:0006940 | regulation of smooth muscle contraction(GO:0006940) |
| 0.1 | 4.4 | GO:0032728 | positive regulation of interferon-beta production(GO:0032728) |
| 0.1 | 0.7 | GO:0043305 | negative regulation of mast cell degranulation(GO:0043305) |
| 0.1 | 23.4 | GO:0010466 | negative regulation of peptidase activity(GO:0010466) |
| 0.1 | 0.2 | GO:0006711 | estrogen catabolic process(GO:0006711) |
| 0.1 | 2.7 | GO:0006879 | cellular iron ion homeostasis(GO:0006879) |
| 0.1 | 1.2 | GO:0050832 | defense response to fungus(GO:0050832) |
| 0.1 | 4.0 | GO:0043407 | negative regulation of MAP kinase activity(GO:0043407) |
| 0.1 | 0.6 | GO:0009048 | dosage compensation(GO:0007549) dosage compensation by inactivation of X chromosome(GO:0009048) |
| 0.1 | 0.5 | GO:0046477 | glycosylceramide catabolic process(GO:0046477) |
| 0.1 | 14.0 | GO:0016042 | lipid catabolic process(GO:0016042) |
| 0.0 | 0.3 | GO:0006933 | negative regulation of cell adhesion involved in substrate-bound cell migration(GO:0006933) |
| 0.0 | 10.0 | GO:0008202 | steroid metabolic process(GO:0008202) |
| 0.0 | 2.0 | GO:0007585 | respiratory gaseous exchange(GO:0007585) |
| 0.0 | 0.6 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.0 | 0.5 | GO:0046085 | adenosine metabolic process(GO:0046085) |
| 0.0 | 0.6 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 0.0 | 5.4 | GO:0043547 | positive regulation of GTPase activity(GO:0043547) |
| 0.0 | 2.3 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.0 | 0.2 | GO:0034501 | protein localization to kinetochore(GO:0034501) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 10.6 | 31.8 | GO:0034359 | mature chylomicron(GO:0034359) |
| 8.2 | 24.7 | GO:0032311 | angiogenin-PRI complex(GO:0032311) |
| 7.2 | 64.8 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 3.6 | 14.6 | GO:0005896 | interleukin-6 receptor complex(GO:0005896) |
| 3.3 | 35.9 | GO:0097433 | dense body(GO:0097433) |
| 2.1 | 35.9 | GO:0043203 | axon hillock(GO:0043203) |
| 1.9 | 29.9 | GO:0044754 | autolysosome(GO:0044754) |
| 1.8 | 8.8 | GO:0030312 | external encapsulating structure(GO:0030312) |
| 1.7 | 5.0 | GO:0033193 | Lsd1/2 complex(GO:0033193) |
| 1.6 | 6.5 | GO:0071953 | elastic fiber(GO:0071953) |
| 1.5 | 7.7 | GO:0005914 | spot adherens junction(GO:0005914) |
| 1.5 | 9.2 | GO:0044279 | other organism cell membrane(GO:0044218) other organism membrane(GO:0044279) |
| 1.4 | 4.3 | GO:0043512 | inhibin A complex(GO:0043512) |
| 1.3 | 13.3 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 1.2 | 12.3 | GO:0070552 | BRISC complex(GO:0070552) |
| 1.2 | 48.7 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 1.2 | 4.6 | GO:1990037 | Lewy body core(GO:1990037) |
| 1.0 | 9.4 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) |
| 1.0 | 10.9 | GO:0070578 | RISC-loading complex(GO:0070578) |
| 1.0 | 7.8 | GO:0030478 | actin cap(GO:0030478) |
| 0.9 | 10.1 | GO:0097208 | alveolar lamellar body(GO:0097208) |
| 0.8 | 204.1 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.7 | 8.9 | GO:0031315 | extrinsic component of mitochondrial outer membrane(GO:0031315) |
| 0.7 | 14.5 | GO:0042581 | specific granule(GO:0042581) |
| 0.5 | 74.3 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.5 | 51.9 | GO:0005811 | lipid particle(GO:0005811) |
| 0.5 | 12.9 | GO:0005685 | U1 snRNP(GO:0005685) |
| 0.5 | 19.5 | GO:0045095 | keratin filament(GO:0045095) |
| 0.4 | 1.8 | GO:0070531 | BRCA1-A complex(GO:0070531) |
| 0.4 | 11.1 | GO:0005782 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.4 | 22.7 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.3 | 2.0 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
| 0.3 | 3.4 | GO:0008024 | cyclin/CDK positive transcription elongation factor complex(GO:0008024) |
| 0.3 | 5.2 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 0.3 | 5.9 | GO:0098644 | complex of collagen trimers(GO:0098644) |
| 0.3 | 4.7 | GO:0012510 | trans-Golgi network transport vesicle membrane(GO:0012510) |
| 0.2 | 17.3 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.2 | 2.7 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.2 | 17.6 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.2 | 406.3 | GO:0005615 | extracellular space(GO:0005615) |
| 0.2 | 11.8 | GO:0030118 | clathrin coat(GO:0030118) |
| 0.2 | 2.4 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.2 | 13.8 | GO:0031903 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.2 | 0.5 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 0.2 | 5.0 | GO:0043034 | costamere(GO:0043034) |
| 0.1 | 7.8 | GO:0015030 | Cajal body(GO:0015030) |
| 0.1 | 13.8 | GO:0005901 | caveola(GO:0005901) |
| 0.1 | 41.1 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.1 | 6.6 | GO:0045335 | phagocytic vesicle(GO:0045335) |
| 0.1 | 14.8 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.1 | 11.5 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.1 | 1.7 | GO:0070938 | contractile ring(GO:0070938) |
| 0.1 | 4.2 | GO:0030864 | cortical actin cytoskeleton(GO:0030864) |
| 0.1 | 7.6 | GO:0016605 | PML body(GO:0016605) |
| 0.1 | 7.0 | GO:0016459 | myosin complex(GO:0016459) |
| 0.1 | 15.7 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.1 | 7.6 | GO:0005923 | bicellular tight junction(GO:0005923) |
| 0.1 | 8.6 | GO:0032993 | protein-DNA complex(GO:0032993) |
| 0.0 | 6.5 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.0 | 2.1 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 10.9 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 1.1 | GO:0045111 | intermediate filament cytoskeleton(GO:0045111) |
| 0.0 | 33.2 | GO:0005783 | endoplasmic reticulum(GO:0005783) |
| 0.0 | 3.0 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.0 | 36.1 | GO:0005576 | extracellular region(GO:0005576) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 19.8 | 59.4 | GO:0034632 | retinol transporter activity(GO:0034632) |
| 19.6 | 78.2 | GO:0004478 | methionine adenosyltransferase activity(GO:0004478) |
| 9.1 | 27.4 | GO:0042602 | riboflavin reductase (NADPH) activity(GO:0042602) |
| 7.2 | 21.5 | GO:0004087 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 6.1 | 42.7 | GO:0016841 | ammonia-lyase activity(GO:0016841) |
| 6.0 | 23.8 | GO:0005008 | hepatocyte growth factor-activated receptor activity(GO:0005008) |
| 5.2 | 41.3 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 5.1 | 35.9 | GO:0015091 | ferric iron transmembrane transporter activity(GO:0015091) trivalent inorganic cation transmembrane transporter activity(GO:0072510) |
| 5.0 | 30.2 | GO:0030294 | receptor signaling protein tyrosine kinase inhibitor activity(GO:0030294) |
| 4.9 | 14.6 | GO:0004915 | interleukin-6 receptor activity(GO:0004915) interleukin-6 binding(GO:0019981) |
| 4.8 | 24.2 | GO:0005345 | purine nucleobase transmembrane transporter activity(GO:0005345) pyrimidine nucleobase transmembrane transporter activity(GO:0005350) nucleobase transmembrane transporter activity(GO:0015205) glycerol channel activity(GO:0015254) |
| 4.6 | 13.9 | GO:0070996 | type 1 melanocortin receptor binding(GO:0070996) |
| 4.6 | 13.8 | GO:0004392 | heme oxygenase (decyclizing) activity(GO:0004392) |
| 4.1 | 12.3 | GO:0004372 | glycine hydroxymethyltransferase activity(GO:0004372) |
| 3.9 | 11.6 | GO:0035717 | chemokine (C-C motif) ligand 7 binding(GO:0035717) |
| 3.8 | 15.1 | GO:0005152 | interleukin-1 receptor antagonist activity(GO:0005152) |
| 3.6 | 17.8 | GO:0033765 | steroid dehydrogenase activity, acting on the CH-CH group of donors(GO:0033765) |
| 3.4 | 40.4 | GO:0004322 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 2.9 | 8.7 | GO:0004731 | purine-nucleoside phosphorylase activity(GO:0004731) |
| 2.8 | 13.8 | GO:0043758 | acetate-CoA ligase (ADP-forming) activity(GO:0043758) |
| 2.6 | 12.9 | GO:0035614 | snRNA stem-loop binding(GO:0035614) |
| 2.4 | 31.8 | GO:0035473 | lipase binding(GO:0035473) |
| 2.4 | 7.1 | GO:0005308 | creatine transmembrane transporter activity(GO:0005308) |
| 2.3 | 9.4 | GO:0043120 | tumor necrosis factor binding(GO:0043120) |
| 2.3 | 13.7 | GO:0008384 | IkappaB kinase activity(GO:0008384) |
| 2.2 | 8.9 | GO:0033814 | propanoyl-CoA C-acyltransferase activity(GO:0033814) propionyl-CoA C2-trimethyltridecanoyltransferase activity(GO:0050632) phosphatidylethanolamine transporter activity(GO:1904121) |
| 2.2 | 26.5 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 2.2 | 24.2 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 2.2 | 11.0 | GO:0004854 | xanthine dehydrogenase activity(GO:0004854) oxidoreductase activity, acting on CH or CH2 groups, NAD or NADP as acceptor(GO:0016726) molybdenum ion binding(GO:0030151) molybdopterin cofactor binding(GO:0043546) |
| 2.0 | 11.8 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 1.9 | 5.8 | GO:0016781 | selenide, water dikinase activity(GO:0004756) phosphotransferase activity, paired acceptors(GO:0016781) |
| 1.7 | 11.9 | GO:0071723 | lipopeptide binding(GO:0071723) |
| 1.7 | 13.4 | GO:0008823 | cupric reductase activity(GO:0008823) ferric-chelate reductase (NADPH) activity(GO:0052851) |
| 1.6 | 7.8 | GO:0008859 | exoribonuclease II activity(GO:0008859) |
| 1.3 | 7.8 | GO:0004720 | protein-lysine 6-oxidase activity(GO:0004720) |
| 1.2 | 11.1 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 1.1 | 9.2 | GO:0004630 | phospholipase D activity(GO:0004630) |
| 1.1 | 4.5 | GO:0004910 | interleukin-1, Type II, blocking receptor activity(GO:0004910) |
| 1.1 | 8.9 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 1.0 | 6.3 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 1.0 | 30.2 | GO:0016502 | purinergic nucleotide receptor activity(GO:0001614) nucleotide receptor activity(GO:0016502) |
| 1.0 | 10.9 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 1.0 | 111.7 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.9 | 3.7 | GO:0070052 | collagen V binding(GO:0070052) |
| 0.9 | 2.8 | GO:0051916 | granulocyte colony-stimulating factor binding(GO:0051916) |
| 0.9 | 13.7 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.8 | 29.7 | GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.8 | 16.9 | GO:0055106 | ubiquitin-protein transferase regulator activity(GO:0055106) |
| 0.8 | 4.9 | GO:0032450 | maltose alpha-glucosidase activity(GO:0032450) |
| 0.7 | 2.8 | GO:0003827 | alpha-1,3-mannosylglycoprotein 2-beta-N-acetylglucosaminyltransferase activity(GO:0003827) |
| 0.7 | 3.4 | GO:0034211 | GTP-dependent protein kinase activity(GO:0034211) |
| 0.6 | 101.3 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.6 | 3.8 | GO:0050656 | 3'-phosphoadenosine 5'-phosphosulfate binding(GO:0050656) |
| 0.6 | 3.2 | GO:0047635 | L-alanine:2-oxoglutarate aminotransferase activity(GO:0004021) alanine-oxo-acid transaminase activity(GO:0047635) |
| 0.6 | 18.9 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.6 | 6.5 | GO:0050543 | icosatetraenoic acid binding(GO:0050543) arachidonic acid binding(GO:0050544) |
| 0.5 | 4.3 | GO:0070699 | type II activin receptor binding(GO:0070699) |
| 0.5 | 8.8 | GO:0015299 | solute:proton antiporter activity(GO:0015299) |
| 0.5 | 5.6 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.5 | 1.5 | GO:0016155 | formyltetrahydrofolate dehydrogenase activity(GO:0016155) |
| 0.5 | 19.3 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 0.4 | 5.7 | GO:0001069 | regulatory region RNA binding(GO:0001069) |
| 0.4 | 11.3 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.4 | 24.7 | GO:0005507 | copper ion binding(GO:0005507) |
| 0.4 | 82.2 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.3 | 35.4 | GO:0016811 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amides(GO:0016811) |
| 0.3 | 13.4 | GO:0043236 | laminin binding(GO:0043236) |
| 0.3 | 0.9 | GO:0004813 | alanine-tRNA ligase activity(GO:0004813) |
| 0.3 | 19.8 | GO:0005160 | transforming growth factor beta receptor binding(GO:0005160) |
| 0.3 | 2.4 | GO:0070644 | vitamin D response element binding(GO:0070644) |
| 0.3 | 1.5 | GO:0004174 | electron-transferring-flavoprotein dehydrogenase activity(GO:0004174) oxidoreductase activity, acting on the CH-NH group of donors, quinone or similar compound as acceptor(GO:0016649) |
| 0.3 | 2.5 | GO:0008061 | chitin binding(GO:0008061) |
| 0.2 | 44.2 | GO:0005506 | iron ion binding(GO:0005506) |
| 0.2 | 4.5 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.2 | 70.2 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.2 | 0.4 | GO:0004616 | phosphogluconate dehydrogenase (decarboxylating) activity(GO:0004616) |
| 0.2 | 10.9 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.2 | 5.2 | GO:0016676 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.2 | 3.4 | GO:0003688 | DNA replication origin binding(GO:0003688) |
| 0.2 | 6.1 | GO:0031435 | mitogen-activated protein kinase kinase kinase binding(GO:0031435) |
| 0.2 | 3.3 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.2 | 38.3 | GO:0030674 | protein binding, bridging(GO:0030674) |
| 0.2 | 5.7 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.2 | 0.5 | GO:0052692 | raffinose alpha-galactosidase activity(GO:0052692) |
| 0.2 | 1.1 | GO:0051731 | polynucleotide 5'-hydroxyl-kinase activity(GO:0051731) |
| 0.2 | 4.4 | GO:0003951 | NAD+ kinase activity(GO:0003951) |
| 0.1 | 8.2 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.1 | 26.4 | GO:0016853 | isomerase activity(GO:0016853) |
| 0.1 | 2.6 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.1 | 5.5 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.1 | 1.8 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.1 | 2.7 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.1 | 3.2 | GO:0031005 | filamin binding(GO:0031005) |
| 0.1 | 3.9 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.1 | 11.8 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.1 | 5.2 | GO:0008028 | monocarboxylic acid transmembrane transporter activity(GO:0008028) |
| 0.1 | 0.3 | GO:0043532 | angiostatin binding(GO:0043532) |
| 0.1 | 4.0 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.1 | 0.6 | GO:0004966 | galanin receptor activity(GO:0004966) |
| 0.1 | 2.5 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.1 | 1.3 | GO:0044213 | intronic transcription regulatory region sequence-specific DNA binding(GO:0001161) intronic transcription regulatory region DNA binding(GO:0044213) |
| 0.1 | 10.4 | GO:0005179 | hormone activity(GO:0005179) |
| 0.1 | 5.1 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.1 | 5.0 | GO:0070888 | E-box binding(GO:0070888) |
| 0.1 | 5.1 | GO:0016810 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds(GO:0016810) |
| 0.1 | 7.2 | GO:0008170 | N-methyltransferase activity(GO:0008170) |
| 0.1 | 0.2 | GO:0050294 | steroid sulfotransferase activity(GO:0050294) |
| 0.1 | 6.3 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 8.1 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 11.2 | GO:0001664 | G-protein coupled receptor binding(GO:0001664) |
| 0.0 | 9.2 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 26.1 | GO:0016491 | oxidoreductase activity(GO:0016491) |
| 0.0 | 9.8 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.0 | 5.5 | GO:0003774 | motor activity(GO:0003774) |
| 0.0 | 0.5 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.0 | 0.2 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.0 | 3.0 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.0 | 2.6 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.0 | 1.6 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 8.4 | GO:0005198 | structural molecule activity(GO:0005198) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.9 | 100.3 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 1.6 | 14.5 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 1.2 | 70.2 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.6 | 9.4 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.5 | 50.1 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.4 | 18.2 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.4 | 11.7 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.4 | 105.0 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.4 | 23.8 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.4 | 23.2 | PID BMP PATHWAY | BMP receptor signaling |
| 0.3 | 6.6 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.3 | 9.2 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.2 | 19.8 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.2 | 52.8 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.2 | 0.7 | ST IL 13 PATHWAY | Interleukin 13 (IL-13) Pathway |
| 0.2 | 5.9 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.2 | 11.7 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.2 | 6.5 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
| 0.1 | 4.3 | PID ALK1 PATHWAY | ALK1 signaling events |
| 0.1 | 5.4 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.1 | 23.1 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.1 | 2.4 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.1 | 3.4 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.0 | 3.4 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 1.1 | PID IL23 PATHWAY | IL23-mediated signaling events |
| 0.0 | 2.6 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.0 | 1.5 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.8 | 41.3 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 3.6 | 61.1 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 3.0 | 61.0 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 2.3 | 78.2 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 1.7 | 24.2 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 1.7 | 41.2 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 1.4 | 26.7 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 7ALPHA HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 7alpha-hydroxycholesterol |
| 1.2 | 4.9 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 1.1 | 48.0 | REACTOME METAL ION SLC TRANSPORTERS | Genes involved in Metal ion SLC transporters |
| 1.1 | 31.8 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 1.0 | 19.7 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.9 | 8.2 | REACTOME BETA DEFENSINS | Genes involved in Beta defensins |
| 0.8 | 11.8 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION IN TLR7 8 OR 9 SIGNALING | Genes involved in TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling |
| 0.8 | 27.5 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.8 | 14.6 | REACTOME ACTIVATION OF IRF3 IRF7 MEDIATED BY TBK1 IKK EPSILON | Genes involved in Activation of IRF3/IRF7 mediated by TBK1/IKK epsilon |
| 0.7 | 11.8 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.6 | 8.4 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.5 | 9.4 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.5 | 5.6 | REACTOME OPSINS | Genes involved in Opsins |
| 0.5 | 14.8 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.5 | 10.9 | REACTOME DESTABILIZATION OF MRNA BY TRISTETRAPROLIN TTP | Genes involved in Destabilization of mRNA by Tristetraprolin (TTP) |
| 0.5 | 11.1 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.4 | 22.3 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.4 | 5.1 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.4 | 5.9 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.4 | 24.8 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.4 | 90.8 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.4 | 16.9 | REACTOME SEMA4D IN SEMAPHORIN SIGNALING | Genes involved in Sema4D in semaphorin signaling |
| 0.3 | 7.7 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.3 | 9.8 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.3 | 11.7 | REACTOME GRB2 EVENTS IN ERBB2 SIGNALING | Genes involved in GRB2 events in ERBB2 signaling |
| 0.3 | 1.8 | REACTOME DESTABILIZATION OF MRNA BY KSRP | Genes involved in Destabilization of mRNA by KSRP |
| 0.2 | 7.6 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.2 | 4.0 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.2 | 7.7 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.2 | 5.4 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.2 | 7.2 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.1 | 12.0 | REACTOME PHASE II CONJUGATION | Genes involved in Phase II conjugation |
| 0.1 | 15.0 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.1 | 7.8 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.1 | 8.8 | REACTOME COSTIMULATION BY THE CD28 FAMILY | Genes involved in Costimulation by the CD28 family |
| 0.1 | 22.4 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.1 | 1.8 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.1 | 0.7 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.1 | 2.4 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.1 | 9.0 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.1 | 3.3 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.1 | 8.7 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.1 | 2.8 | REACTOME TRANSPORT TO THE GOLGI AND SUBSEQUENT MODIFICATION | Genes involved in Transport to the Golgi and subsequent modification |
| 0.0 | 5.5 | REACTOME G ALPHA S SIGNALLING EVENTS | Genes involved in G alpha (s) signalling events |
| 0.0 | 2.1 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.0 | 0.5 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.0 | 0.3 | REACTOME ACYL CHAIN REMODELLING OF PS | Genes involved in Acyl chain remodelling of PS |
| 0.0 | 0.3 | REACTOME FORMATION OF ATP BY CHEMIOSMOTIC COUPLING | Genes involved in Formation of ATP by chemiosmotic coupling |
| 0.0 | 3.1 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.0 | 0.5 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.0 | 0.9 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |