Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
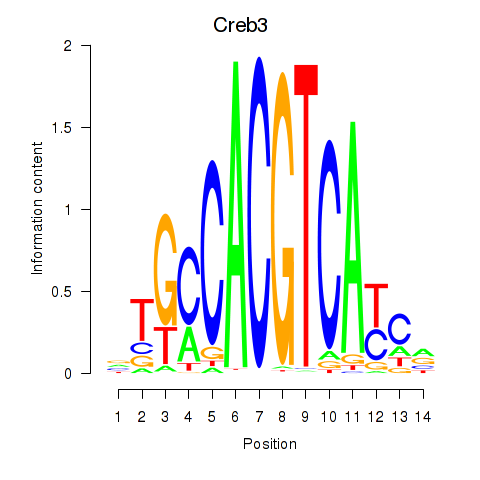
Results for Creb3
Z-value: 1.17
Transcription factors associated with Creb3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Creb3
|
ENSMUSG00000028466.16 | Creb3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Creb3 | mm39_v1_chr4_+_43562706_43562947 | 0.33 | 4.5e-03 | Click! |
Activity profile of Creb3 motif
Sorted Z-values of Creb3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Creb3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr8_-_106863423 | 10.70 |
ENSMUST00000146940.2
|
Esrp2
|
epithelial splicing regulatory protein 2 |
| chr8_-_106863521 | 10.41 |
ENSMUST00000115979.9
|
Esrp2
|
epithelial splicing regulatory protein 2 |
| chr15_+_79400597 | 9.49 |
ENSMUST00000010974.9
|
Kdelr3
|
KDEL (Lys-Asp-Glu-Leu) endoplasmic reticulum protein retention receptor 3 |
| chrX_+_72830668 | 9.31 |
ENSMUST00000002090.3
|
Ssr4
|
signal sequence receptor, delta |
| chrX_+_72830607 | 8.90 |
ENSMUST00000166518.8
|
Ssr4
|
signal sequence receptor, delta |
| chr7_-_44741138 | 7.63 |
ENSMUST00000210527.2
|
Rcn3
|
reticulocalbin 3, EF-hand calcium binding domain |
| chr3_+_123061094 | 7.32 |
ENSMUST00000047923.12
ENSMUST00000200333.2 |
Sec24d
|
Sec24 related gene family, member D (S. cerevisiae) |
| chr2_-_143853122 | 7.32 |
ENSMUST00000016072.12
ENSMUST00000037875.6 |
Rrbp1
|
ribosome binding protein 1 |
| chr1_+_58152295 | 6.88 |
ENSMUST00000040999.14
ENSMUST00000162011.3 |
Aox3
|
aldehyde oxidase 3 |
| chr18_+_35347983 | 6.66 |
ENSMUST00000235449.2
ENSMUST00000235269.2 |
Ctnna1
|
catenin (cadherin associated protein), alpha 1 |
| chr1_+_157334347 | 6.50 |
ENSMUST00000027881.15
|
Sec16b
|
SEC16 homolog B (S. cerevisiae) |
| chr1_+_157334298 | 6.35 |
ENSMUST00000086130.9
|
Sec16b
|
SEC16 homolog B (S. cerevisiae) |
| chr2_+_172994841 | 6.29 |
ENSMUST00000029017.6
|
Pck1
|
phosphoenolpyruvate carboxykinase 1, cytosolic |
| chr2_-_179976458 | 6.07 |
ENSMUST00000015771.3
|
Gata5
|
GATA binding protein 5 |
| chr6_-_71121347 | 6.05 |
ENSMUST00000160918.8
|
Thnsl2
|
threonine synthase-like 2 (bacterial) |
| chr6_-_71121324 | 6.04 |
ENSMUST00000074241.9
|
Thnsl2
|
threonine synthase-like 2 (bacterial) |
| chr18_-_64794338 | 5.87 |
ENSMUST00000025482.10
|
Atp8b1
|
ATPase, class I, type 8B, member 1 |
| chr9_+_103940575 | 5.66 |
ENSMUST00000120854.8
|
Acad11
|
acyl-Coenzyme A dehydrogenase family, member 11 |
| chr17_-_57535003 | 5.21 |
ENSMUST00000177046.2
ENSMUST00000024988.15 |
C3
|
complement component 3 |
| chr15_+_99488527 | 4.48 |
ENSMUST00000169082.3
ENSMUST00000088200.13 |
Aqp5
|
aquaporin 5 |
| chr9_-_20888054 | 4.45 |
ENSMUST00000054197.7
|
S1pr2
|
sphingosine-1-phosphate receptor 2 |
| chr11_-_60702081 | 4.13 |
ENSMUST00000018744.15
|
Shmt1
|
serine hydroxymethyltransferase 1 (soluble) |
| chr8_+_91635192 | 4.13 |
ENSMUST00000211403.2
|
Chd9
|
chromodomain helicase DNA binding protein 9 |
| chr9_-_36678868 | 4.13 |
ENSMUST00000217599.2
ENSMUST00000120381.9 |
Stt3a
|
STT3, subunit of the oligosaccharyltransferase complex, homolog A (S. cerevisiae) |
| chr13_-_4573312 | 4.06 |
ENSMUST00000221564.2
ENSMUST00000078239.5 ENSMUST00000080361.13 |
Akr1c20
|
aldo-keto reductase family 1, member C20 |
| chr13_-_38712387 | 4.05 |
ENSMUST00000035988.16
|
Txndc5
|
thioredoxin domain containing 5 |
| chr15_+_99489018 | 4.05 |
ENSMUST00000229728.2
ENSMUST00000231163.2 |
Aqp5
|
aquaporin 5 |
| chr17_+_35658131 | 3.82 |
ENSMUST00000071951.14
ENSMUST00000116598.10 ENSMUST00000078205.14 ENSMUST00000076256.8 |
H2-Q7
|
histocompatibility 2, Q region locus 7 |
| chr4_+_47474652 | 3.75 |
ENSMUST00000065678.6
|
Sec61b
|
Sec61 beta subunit |
| chr7_-_118091135 | 3.70 |
ENSMUST00000178344.3
|
Itpripl2
|
inositol 1,4,5-triphosphate receptor interacting protein-like 2 |
| chr5_+_52898910 | 3.70 |
ENSMUST00000031081.11
ENSMUST00000031082.8 |
Pi4k2b
|
phosphatidylinositol 4-kinase type 2 beta |
| chr4_+_47474715 | 3.69 |
ENSMUST00000137461.8
ENSMUST00000125622.2 |
Sec61b
|
Sec61 beta subunit |
| chr17_-_28569721 | 3.56 |
ENSMUST00000156862.3
|
Tead3
|
TEA domain family member 3 |
| chr4_-_104967032 | 3.56 |
ENSMUST00000030243.8
|
Prkaa2
|
protein kinase, AMP-activated, alpha 2 catalytic subunit |
| chr10_+_128158413 | 3.55 |
ENSMUST00000219836.2
|
Cnpy2
|
canopy FGF signaling regulator 2 |
| chr15_-_99717956 | 3.54 |
ENSMUST00000109024.9
|
Lima1
|
LIM domain and actin binding 1 |
| chr10_-_128236317 | 3.45 |
ENSMUST00000167859.2
ENSMUST00000218858.2 |
Slc39a5
|
solute carrier family 39 (metal ion transporter), member 5 |
| chr1_-_93373364 | 3.33 |
ENSMUST00000190321.7
ENSMUST00000042498.14 |
Hdlbp
|
high density lipoprotein (HDL) binding protein |
| chr9_-_20887967 | 3.33 |
ENSMUST00000214218.2
|
S1pr2
|
sphingosine-1-phosphate receptor 2 |
| chr1_-_93373145 | 3.31 |
ENSMUST00000186787.7
|
Hdlbp
|
high density lipoprotein (HDL) binding protein |
| chr17_-_35954573 | 3.26 |
ENSMUST00000095467.4
|
Mucl3
|
mucin like 3 |
| chr7_+_128125339 | 3.20 |
ENSMUST00000033136.9
|
Bag3
|
BCL2-associated athanogene 3 |
| chr10_+_41395870 | 3.09 |
ENSMUST00000189300.2
|
Cd164
|
CD164 antigen |
| chr10_+_128158328 | 3.07 |
ENSMUST00000219037.2
ENSMUST00000026446.4 |
Cnpy2
|
canopy FGF signaling regulator 2 |
| chr14_-_55880980 | 3.06 |
ENSMUST00000132338.8
|
Tm9sf1
|
transmembrane 9 superfamily member 1 |
| chr15_+_6416079 | 3.03 |
ENSMUST00000080880.12
|
Dab2
|
disabled 2, mitogen-responsive phosphoprotein |
| chr7_+_45522551 | 3.03 |
ENSMUST00000211234.2
|
Kdelr1
|
KDEL (Lys-Asp-Glu-Leu) endoplasmic reticulum protein retention receptor 1 |
| chr18_-_61344452 | 3.00 |
ENSMUST00000148829.2
|
Slc26a2
|
solute carrier family 26 (sulfate transporter), member 2 |
| chr7_-_44741609 | 3.00 |
ENSMUST00000210734.2
|
Rcn3
|
reticulocalbin 3, EF-hand calcium binding domain |
| chr10_-_17823736 | 2.99 |
ENSMUST00000037879.8
|
Heca
|
hdc homolog, cell cycle regulator |
| chr4_-_118401185 | 2.98 |
ENSMUST00000128098.8
|
Tmem125
|
transmembrane protein 125 |
| chr5_+_28276353 | 2.96 |
ENSMUST00000059155.11
|
Insig1
|
insulin induced gene 1 |
| chr3_+_129007599 | 2.93 |
ENSMUST00000042587.12
|
Pitx2
|
paired-like homeodomain transcription factor 2 |
| chr15_+_6416229 | 2.84 |
ENSMUST00000110664.9
ENSMUST00000110663.9 ENSMUST00000161812.8 ENSMUST00000160134.8 |
Dab2
|
disabled 2, mitogen-responsive phosphoprotein |
| chr16_-_56706494 | 2.75 |
ENSMUST00000023435.6
|
Tmem45a
|
transmembrane protein 45a |
| chr7_-_44741622 | 2.75 |
ENSMUST00000210469.2
ENSMUST00000211352.2 ENSMUST00000019683.11 |
Rcn3
|
reticulocalbin 3, EF-hand calcium binding domain |
| chr11_+_85777185 | 2.69 |
ENSMUST00000108047.8
|
Tbx4
|
T-box 4 |
| chr10_-_128236366 | 2.67 |
ENSMUST00000219131.2
|
Slc39a5
|
solute carrier family 39 (metal ion transporter), member 5 |
| chr4_-_108263873 | 2.66 |
ENSMUST00000184609.2
|
Gpx7
|
glutathione peroxidase 7 |
| chr11_-_96807192 | 2.65 |
ENSMUST00000144731.8
ENSMUST00000127048.8 |
Cdk5rap3
|
CDK5 regulatory subunit associated protein 3 |
| chr11_-_113641980 | 2.63 |
ENSMUST00000153453.2
|
Cdc42ep4
|
CDC42 effector protein (Rho GTPase binding) 4 |
| chr11_-_51647204 | 2.59 |
ENSMUST00000109092.8
ENSMUST00000064297.5 |
Sec24a
|
Sec24 related gene family, member A (S. cerevisiae) |
| chrX_-_72830487 | 2.56 |
ENSMUST00000052761.9
|
Idh3g
|
isocitrate dehydrogenase 3 (NAD+), gamma |
| chr14_-_55881177 | 2.53 |
ENSMUST00000138085.2
|
Tm9sf1
|
transmembrane 9 superfamily member 1 |
| chr3_-_154798982 | 2.51 |
ENSMUST00000066568.6
|
Fpgt
|
fucose-1-phosphate guanylyltransferase |
| chr11_-_113642635 | 2.49 |
ENSMUST00000053536.5
|
Cdc42ep4
|
CDC42 effector protein (Rho GTPase binding) 4 |
| chr11_-_51647290 | 2.48 |
ENSMUST00000109097.9
|
Sec24a
|
Sec24 related gene family, member A (S. cerevisiae) |
| chr1_-_170002444 | 2.44 |
ENSMUST00000111351.10
ENSMUST00000027981.8 |
Uap1
|
UDP-N-acetylglucosamine pyrophosphorylase 1 |
| chr16_-_13720915 | 2.44 |
ENSMUST00000115803.9
|
Pdxdc1
|
pyridoxal-dependent decarboxylase domain containing 1 |
| chr6_+_88061464 | 2.43 |
ENSMUST00000032143.8
|
Rpn1
|
ribophorin I |
| chr14_+_26359390 | 2.42 |
ENSMUST00000112318.10
|
Arf4
|
ADP-ribosylation factor 4 |
| chr14_-_55880708 | 2.36 |
ENSMUST00000120041.8
ENSMUST00000121937.8 ENSMUST00000133707.2 ENSMUST00000002391.15 ENSMUST00000121791.8 |
Tm9sf1
|
transmembrane 9 superfamily member 1 |
| chr13_-_38178059 | 2.35 |
ENSMUST00000225319.2
ENSMUST00000225246.2 ENSMUST00000021864.8 |
Ssr1
|
signal sequence receptor, alpha |
| chr2_+_80145805 | 2.33 |
ENSMUST00000028392.8
|
Dnajc10
|
DnaJ heat shock protein family (Hsp40) member C10 |
| chr9_+_108216233 | 2.25 |
ENSMUST00000082429.8
|
Gpx1
|
glutathione peroxidase 1 |
| chr16_-_13720863 | 2.24 |
ENSMUST00000115804.9
|
Pdxdc1
|
pyridoxal-dependent decarboxylase domain containing 1 |
| chr15_+_34238174 | 2.23 |
ENSMUST00000022867.5
ENSMUST00000226627.2 |
Laptm4b
|
lysosomal-associated protein transmembrane 4B |
| chr17_+_45874800 | 2.22 |
ENSMUST00000224905.2
ENSMUST00000226086.2 ENSMUST00000041353.7 |
Slc35b2
|
solute carrier family 35, member B2 |
| chr7_-_113853894 | 2.21 |
ENSMUST00000033012.9
|
Copb1
|
coatomer protein complex, subunit beta 1 |
| chr6_-_118456198 | 2.21 |
ENSMUST00000161170.2
|
Zfp9
|
zinc finger protein 9 |
| chr3_+_89153704 | 2.21 |
ENSMUST00000168900.3
|
Krtcap2
|
keratinocyte associated protein 2 |
| chrX_-_135641869 | 2.20 |
ENSMUST00000166930.8
ENSMUST00000113095.8 |
Morf4l2
|
mortality factor 4 like 2 |
| chrX_-_73416869 | 2.19 |
ENSMUST00000073067.11
ENSMUST00000037967.6 |
Slc10a3
|
solute carrier family 10 (sodium/bile acid cotransporter family), member 3 |
| chr16_+_36695479 | 2.18 |
ENSMUST00000023534.7
ENSMUST00000114812.9 ENSMUST00000134616.8 |
Golgb1
|
golgi autoantigen, golgin subfamily b, macrogolgin 1 |
| chr11_-_109886569 | 2.17 |
ENSMUST00000106669.3
|
Abca8b
|
ATP-binding cassette, sub-family A (ABC1), member 8b |
| chr10_+_41395410 | 2.16 |
ENSMUST00000019962.15
|
Cd164
|
CD164 antigen |
| chr12_+_51424343 | 2.15 |
ENSMUST00000219434.2
ENSMUST00000021335.7 |
Scfd1
|
Sec1 family domain containing 1 |
| chr11_-_96807273 | 2.14 |
ENSMUST00000103152.11
|
Cdk5rap3
|
CDK5 regulatory subunit associated protein 3 |
| chr7_-_79765042 | 2.14 |
ENSMUST00000206714.2
ENSMUST00000107384.10 |
Idh2
|
isocitrate dehydrogenase 2 (NADP+), mitochondrial |
| chr13_+_76246853 | 2.13 |
ENSMUST00000091466.4
ENSMUST00000224386.2 |
Ttc37
|
tetratricopeptide repeat domain 37 |
| chr1_+_185095232 | 2.11 |
ENSMUST00000046514.13
|
Eprs
|
glutamyl-prolyl-tRNA synthetase |
| chrX_-_135642025 | 2.06 |
ENSMUST00000155207.8
ENSMUST00000080411.13 ENSMUST00000169418.8 |
Morf4l2
|
mortality factor 4 like 2 |
| chr17_-_26727437 | 2.06 |
ENSMUST00000236661.2
ENSMUST00000025025.7 |
Dusp1
|
dual specificity phosphatase 1 |
| chr18_-_61344644 | 2.06 |
ENSMUST00000146409.8
|
Slc26a2
|
solute carrier family 26 (sulfate transporter), member 2 |
| chr5_-_31102829 | 2.05 |
ENSMUST00000031051.8
|
Cgref1
|
cell growth regulator with EF hand domain 1 |
| chr16_-_94171556 | 2.00 |
ENSMUST00000113906.9
|
Pigp
|
phosphatidylinositol glycan anchor biosynthesis, class P |
| chr9_+_44290832 | 2.00 |
ENSMUST00000161318.8
ENSMUST00000217019.2 ENSMUST00000160902.8 |
Hyou1
|
hypoxia up-regulated 1 |
| chr9_+_108216466 | 1.98 |
ENSMUST00000193987.2
|
Gpx1
|
glutathione peroxidase 1 |
| chr9_+_108216433 | 1.98 |
ENSMUST00000191997.2
|
Gpx1
|
glutathione peroxidase 1 |
| chr4_+_46489248 | 1.96 |
ENSMUST00000030018.5
|
Nans
|
N-acetylneuraminic acid synthase (sialic acid synthase) |
| chr9_-_107482462 | 1.96 |
ENSMUST00000194433.6
|
Sema3b
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3B |
| chr3_+_129007917 | 1.95 |
ENSMUST00000174623.3
|
Pitx2
|
paired-like homeodomain transcription factor 2 |
| chr9_+_44290787 | 1.95 |
ENSMUST00000066601.13
|
Hyou1
|
hypoxia up-regulated 1 |
| chr12_+_69230931 | 1.94 |
ENSMUST00000060579.10
|
Mgat2
|
mannoside acetylglucosaminyltransferase 2 |
| chr2_-_104324035 | 1.92 |
ENSMUST00000111124.8
|
Hipk3
|
homeodomain interacting protein kinase 3 |
| chr16_-_13720949 | 1.92 |
ENSMUST00000023361.12
ENSMUST00000115802.2 |
Pdxdc1
|
pyridoxal-dependent decarboxylase domain containing 1 |
| chr11_-_113642135 | 1.87 |
ENSMUST00000106616.2
|
Cdc42ep4
|
CDC42 effector protein (Rho GTPase binding) 4 |
| chr7_+_43424028 | 1.84 |
ENSMUST00000080211.12
|
Klk11
|
kallikrein related-peptidase 11 |
| chr11_-_109886601 | 1.83 |
ENSMUST00000020948.15
|
Abca8b
|
ATP-binding cassette, sub-family A (ABC1), member 8b |
| chr6_-_108162513 | 1.83 |
ENSMUST00000167338.8
ENSMUST00000172188.2 ENSMUST00000032191.16 |
Sumf1
|
sulfatase modifying factor 1 |
| chr4_+_130001349 | 1.82 |
ENSMUST00000030563.6
|
Pef1
|
penta-EF hand domain containing 1 |
| chr7_+_101859542 | 1.78 |
ENSMUST00000140631.2
ENSMUST00000120879.8 ENSMUST00000146996.8 |
Pgap2
|
post-GPI attachment to proteins 2 |
| chrX_+_73356597 | 1.76 |
ENSMUST00000114160.2
|
Fam50a
|
family with sequence similarity 50, member A |
| chr11_-_96807233 | 1.75 |
ENSMUST00000130774.2
|
Cdk5rap3
|
CDK5 regulatory subunit associated protein 3 |
| chr16_-_94171533 | 1.72 |
ENSMUST00000113910.8
|
Pigp
|
phosphatidylinositol glycan anchor biosynthesis, class P |
| chr6_+_29348068 | 1.70 |
ENSMUST00000173216.8
ENSMUST00000173694.5 ENSMUST00000172974.8 ENSMUST00000031779.17 ENSMUST00000090481.14 |
Calu
|
calumenin |
| chr18_+_31742565 | 1.69 |
ENSMUST00000164667.2
|
B930094E09Rik
|
RIKEN cDNA B930094E09 gene |
| chr2_+_32665781 | 1.67 |
ENSMUST00000066352.6
|
Ptrh1
|
peptidyl-tRNA hydrolase 1 homolog |
| chr14_+_26359191 | 1.67 |
ENSMUST00000022429.9
|
Arf4
|
ADP-ribosylation factor 4 |
| chr18_-_84703738 | 1.65 |
ENSMUST00000025546.17
|
Cndp2
|
CNDP dipeptidase 2 (metallopeptidase M20 family) |
| chr11_+_72686990 | 1.60 |
ENSMUST00000069395.7
ENSMUST00000172220.8 |
Zzef1
|
zinc finger, ZZ-type with EF hand domain 1 |
| chr3_+_89366632 | 1.59 |
ENSMUST00000107410.8
|
Pmvk
|
phosphomevalonate kinase |
| chr8_+_126721878 | 1.59 |
ENSMUST00000046765.10
|
Kcnk1
|
potassium channel, subfamily K, member 1 |
| chr4_+_129941696 | 1.57 |
ENSMUST00000142293.8
|
Col16a1
|
collagen, type XVI, alpha 1 |
| chr8_+_46428551 | 1.56 |
ENSMUST00000034051.7
ENSMUST00000150943.2 |
Ufsp2
|
UFM1-specific peptidase 2 |
| chr9_-_103940247 | 1.54 |
ENSMUST00000035166.12
|
Uba5
|
ubiquitin-like modifier activating enzyme 5 |
| chr2_+_155118217 | 1.50 |
ENSMUST00000029128.4
|
Map1lc3a
|
microtubule-associated protein 1 light chain 3 alpha |
| chr17_-_56490887 | 1.49 |
ENSMUST00000019723.8
|
Mydgf
|
myeloid derived growth factor |
| chr10_-_115087309 | 1.46 |
ENSMUST00000020339.10
|
Tbc1d15
|
TBC1 domain family, member 15 |
| chr4_+_115594951 | 1.45 |
ENSMUST00000106522.9
|
Efcab14
|
EF-hand calcium binding domain 14 |
| chr13_-_103911092 | 1.45 |
ENSMUST00000074616.7
|
Srek1
|
splicing regulatory glutamine/lysine-rich protein 1 |
| chr1_-_183126875 | 1.45 |
ENSMUST00000195233.3
|
Mia3
|
melanoma inhibitory activity 3 |
| chr1_-_170002526 | 1.44 |
ENSMUST00000111350.10
|
Uap1
|
UDP-N-acetylglucosamine pyrophosphorylase 1 |
| chr8_+_126722113 | 1.41 |
ENSMUST00000212831.2
|
Kcnk1
|
potassium channel, subfamily K, member 1 |
| chr11_+_53241561 | 1.41 |
ENSMUST00000060945.12
|
Aff4
|
AF4/FMR2 family, member 4 |
| chr18_+_57487908 | 1.40 |
ENSMUST00000238069.2
|
Prrc1
|
proline-rich coiled-coil 1 |
| chr14_-_55881257 | 1.38 |
ENSMUST00000122358.8
|
Tm9sf1
|
transmembrane 9 superfamily member 1 |
| chr17_-_84773544 | 1.38 |
ENSMUST00000047524.10
|
Thada
|
thyroid adenoma associated |
| chr7_-_44983080 | 1.37 |
ENSMUST00000211743.2
ENSMUST00000042194.10 |
Trpm4
|
transient receptor potential cation channel, subfamily M, member 4 |
| chr19_-_60779077 | 1.37 |
ENSMUST00000025955.8
|
Eif3a
|
eukaryotic translation initiation factor 3, subunit A |
| chr10_-_128759817 | 1.36 |
ENSMUST00000131271.2
|
Bloc1s1
|
biogenesis of lysosomal organelles complex-1, subunit 1 |
| chr3_+_99048397 | 1.34 |
ENSMUST00000198044.2
|
Wars2
|
tryptophanyl tRNA synthetase 2 (mitochondrial) |
| chr10_+_126877798 | 1.34 |
ENSMUST00000006915.14
ENSMUST00000120542.8 |
Mettl1
|
methyltransferase like 1 |
| chr11_+_94544593 | 1.33 |
ENSMUST00000025278.8
|
Mrpl27
|
mitochondrial ribosomal protein L27 |
| chr5_+_129970882 | 1.30 |
ENSMUST00000201855.2
ENSMUST00000073945.6 |
Vkorc1l1
|
vitamin K epoxide reductase complex, subunit 1-like 1 |
| chr7_-_34914675 | 1.30 |
ENSMUST00000118444.3
ENSMUST00000122409.8 |
Lrp3
|
low density lipoprotein receptor-related protein 3 |
| chr8_+_26210064 | 1.29 |
ENSMUST00000068916.16
ENSMUST00000139836.8 |
Plpp5
|
phospholipid phosphatase 5 |
| chr4_-_150994260 | 1.29 |
ENSMUST00000134751.8
ENSMUST00000030805.14 |
Park7
|
Parkinson disease (autosomal recessive, early onset) 7 |
| chr1_+_171910073 | 1.27 |
ENSMUST00000135192.8
|
Copa
|
coatomer protein complex subunit alpha |
| chr1_+_130947580 | 1.27 |
ENSMUST00000016673.6
|
Il10
|
interleukin 10 |
| chr7_-_48531344 | 1.26 |
ENSMUST00000119223.2
|
E2f8
|
E2F transcription factor 8 |
| chr12_-_69230760 | 1.25 |
ENSMUST00000110620.2
ENSMUST00000110619.2 ENSMUST00000054544.7 |
Rpl36al
|
ribosomal protein L36A-like |
| chr1_+_75145275 | 1.25 |
ENSMUST00000162768.8
ENSMUST00000160439.8 ENSMUST00000027394.12 |
Zfand2b
|
zinc finger, AN1 type domain 2B |
| chr7_-_46445085 | 1.25 |
ENSMUST00000123725.2
|
Hps5
|
HPS5, biogenesis of lysosomal organelles complex 2 subunit 2 |
| chr7_-_45116197 | 1.24 |
ENSMUST00000211195.2
ENSMUST00000210019.2 |
Bax
|
BCL2-associated X protein |
| chr10_+_121200984 | 1.24 |
ENSMUST00000040344.7
|
Gns
|
glucosamine (N-acetyl)-6-sulfatase |
| chr16_-_94171340 | 1.22 |
ENSMUST00000138514.2
|
Pigp
|
phosphatidylinositol glycan anchor biosynthesis, class P |
| chr3_+_99048379 | 1.22 |
ENSMUST00000004343.7
|
Wars2
|
tryptophanyl tRNA synthetase 2 (mitochondrial) |
| chr9_-_103243039 | 1.20 |
ENSMUST00000035484.11
|
Cdv3
|
carnitine deficiency-associated gene expressed in ventricle 3 |
| chr2_+_155224105 | 1.20 |
ENSMUST00000134218.2
|
Trp53inp2
|
transformation related protein 53 inducible nuclear protein 2 |
| chr4_+_115594918 | 1.17 |
ENSMUST00000106525.9
|
Efcab14
|
EF-hand calcium binding domain 14 |
| chr19_-_53026965 | 1.17 |
ENSMUST00000183274.8
ENSMUST00000182097.2 |
Xpnpep1
|
X-prolyl aminopeptidase (aminopeptidase P) 1, soluble |
| chr5_+_137015873 | 1.12 |
ENSMUST00000004968.11
|
Plod3
|
procollagen-lysine, 2-oxoglutarate 5-dioxygenase 3 |
| chr19_+_11943265 | 1.11 |
ENSMUST00000025590.11
|
Osbp
|
oxysterol binding protein |
| chr15_+_84052028 | 1.11 |
ENSMUST00000045289.6
|
Pnpla3
|
patatin-like phospholipase domain containing 3 |
| chr19_+_8718837 | 1.09 |
ENSMUST00000177373.8
ENSMUST00000010254.16 |
Stx5a
|
syntaxin 5A |
| chr16_-_10952465 | 1.07 |
ENSMUST00000118362.8
ENSMUST00000118679.2 ENSMUST00000038424.14 |
Txndc11
|
thioredoxin domain containing 11 |
| chr13_-_76246689 | 1.07 |
ENSMUST00000239063.2
ENSMUST00000120573.3 |
Arsk
|
arylsulfatase K |
| chr2_-_168584020 | 1.07 |
ENSMUST00000109177.8
|
Atp9a
|
ATPase, class II, type 9A |
| chr9_-_89586960 | 1.05 |
ENSMUST00000058488.9
|
Tmed3
|
transmembrane p24 trafficking protein 3 |
| chr19_+_8718777 | 1.04 |
ENSMUST00000176381.8
|
Stx5a
|
syntaxin 5A |
| chr1_+_171910343 | 1.03 |
ENSMUST00000027833.12
|
Copa
|
coatomer protein complex subunit alpha |
| chr18_-_31742946 | 1.03 |
ENSMUST00000060396.7
|
Slc25a46
|
solute carrier family 25, member 46 |
| chr11_-_118306620 | 1.03 |
ENSMUST00000164927.2
|
Cant1
|
calcium activated nucleotidase 1 |
| chr7_-_28931873 | 1.02 |
ENSMUST00000085818.6
|
Kcnk6
|
potassium inwardly-rectifying channel, subfamily K, member 6 |
| chr1_+_192855776 | 1.01 |
ENSMUST00000161235.3
ENSMUST00000160077.2 ENSMUST00000178744.2 ENSMUST00000192189.2 ENSMUST00000110831.4 ENSMUST00000191613.2 |
A130010J15Rik
|
RIKEN cDNA A130010J15 gene |
| chr3_-_108469468 | 1.00 |
ENSMUST00000106622.3
|
Tmem167b
|
transmembrane protein 167B |
| chr11_-_72686853 | 0.98 |
ENSMUST00000156294.8
|
Cyb5d2
|
cytochrome b5 domain containing 2 |
| chr8_+_109441276 | 0.97 |
ENSMUST00000043896.10
|
Zfhx3
|
zinc finger homeobox 3 |
| chr9_-_64928927 | 0.96 |
ENSMUST00000036615.7
|
Hacd3
|
3-hydroxyacyl-CoA dehydratase 3 |
| chr8_-_105981732 | 0.96 |
ENSMUST00000093217.9
ENSMUST00000161745.3 ENSMUST00000136822.3 |
B3gnt9
|
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 9 |
| chr3_-_108469740 | 0.95 |
ENSMUST00000090546.6
|
Tmem167b
|
transmembrane protein 167B |
| chr12_+_71356632 | 0.94 |
ENSMUST00000061273.12
ENSMUST00000150639.2 |
Dact1
|
dishevelled-binding antagonist of beta-catenin 1 |
| chr1_+_87254729 | 0.93 |
ENSMUST00000172794.8
ENSMUST00000164992.9 ENSMUST00000173173.8 |
Gigyf2
|
GRB10 interacting GYF protein 2 |
| chr2_-_105229653 | 0.91 |
ENSMUST00000006128.7
|
Rcn1
|
reticulocalbin 1 |
| chr7_-_121666486 | 0.91 |
ENSMUST00000033159.4
|
Ears2
|
glutamyl-tRNA synthetase 2, mitochondrial |
| chrX_-_162332673 | 0.88 |
ENSMUST00000033730.3
|
Grpr
|
gastrin releasing peptide receptor |
| chr10_+_19232281 | 0.88 |
ENSMUST00000053225.7
|
Olig3
|
oligodendrocyte transcription factor 3 |
| chr11_+_72687080 | 0.87 |
ENSMUST00000207107.2
|
Zzef1
|
zinc finger, ZZ-type with EF hand domain 1 |
| chr1_-_183150867 | 0.87 |
ENSMUST00000194543.4
|
Mia3
|
melanoma inhibitory activity 3 |
| chr14_-_59835285 | 0.86 |
ENSMUST00000022555.11
ENSMUST00000225839.2 ENSMUST00000056997.15 ENSMUST00000171683.3 ENSMUST00000167100.9 |
Cdadc1
|
cytidine and dCMP deaminase domain containing 1 |
| chr13_-_54836059 | 0.86 |
ENSMUST00000122935.2
ENSMUST00000128257.8 |
Rnf44
|
ring finger protein 44 |
| chr6_-_119825020 | 0.84 |
ENSMUST00000184864.8
|
Erc1
|
ELKS/RAB6-interacting/CAST family member 1 |
| chr5_-_148989821 | 0.84 |
ENSMUST00000139443.8
ENSMUST00000085546.13 |
Hmgb1
|
high mobility group box 1 |
| chr16_-_5021843 | 0.83 |
ENSMUST00000147567.2
ENSMUST00000023911.11 |
Nagpa
|
N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase |
| chr9_-_88364593 | 0.83 |
ENSMUST00000173801.8
ENSMUST00000069221.12 ENSMUST00000172508.2 |
Syncrip
|
synaptotagmin binding, cytoplasmic RNA interacting protein |
| chr10_-_128758757 | 0.83 |
ENSMUST00000135161.2
|
Rdh5
|
retinol dehydrogenase 5 |
| chr3_-_96812610 | 0.80 |
ENSMUST00000029738.14
|
Gpr89
|
G protein-coupled receptor 89 |
| chr19_+_4042752 | 0.76 |
ENSMUST00000041871.9
|
Tbx10
|
T-box 10 |
| chr10_+_81012465 | 0.75 |
ENSMUST00000047864.11
|
Eef2
|
eukaryotic translation elongation factor 2 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.4 | 12.1 | GO:0006566 | threonine metabolic process(GO:0006566) |
| 1.9 | 7.4 | GO:0031204 | posttranslational protein targeting to membrane, translocation(GO:0031204) |
| 1.8 | 12.5 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 1.6 | 6.3 | GO:0006114 | glycerol biosynthetic process(GO:0006114) positive regulation of transcription from RNA polymerase II promoter in response to acidic pH(GO:0061402) |
| 1.6 | 6.2 | GO:0009609 | response to symbiont(GO:0009608) response to symbiotic bacterium(GO:0009609) response to selenium ion(GO:0010269) |
| 1.5 | 14.7 | GO:0048208 | vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) |
| 1.4 | 4.1 | GO:0019264 | glycine biosynthetic process from serine(GO:0019264) |
| 1.4 | 6.9 | GO:0009115 | xanthine catabolic process(GO:0009115) |
| 1.4 | 4.1 | GO:0043686 | co-translational protein modification(GO:0043686) |
| 1.3 | 5.2 | GO:0001970 | positive regulation of activation of membrane attack complex(GO:0001970) |
| 1.2 | 6.1 | GO:0034224 | cellular response to zinc ion starvation(GO:0034224) |
| 1.1 | 6.7 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 1.0 | 18.3 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 1.0 | 8.1 | GO:0071569 | protein ufmylation(GO:0071569) |
| 1.0 | 5.9 | GO:0035026 | leading edge cell differentiation(GO:0035026) |
| 1.0 | 4.9 | GO:0021763 | subthalamic nucleus development(GO:0021763) superior vena cava morphogenesis(GO:0060578) |
| 1.0 | 7.8 | GO:0003376 | sphingosine-1-phosphate signaling pathway(GO:0003376) |
| 0.9 | 6.6 | GO:0045716 | positive regulation of low-density lipoprotein particle receptor biosynthetic process(GO:0045716) |
| 0.8 | 8.5 | GO:0015670 | carbon dioxide transport(GO:0015670) |
| 0.7 | 5.9 | GO:0021650 | vestibulocochlear nerve formation(GO:0021650) |
| 0.7 | 2.1 | GO:1904464 | regulation of matrix metallopeptidase secretion(GO:1904464) matrix metallopeptidase secretion(GO:1990773) |
| 0.7 | 2.0 | GO:0032976 | release of matrix enzymes from mitochondria(GO:0032976) B cell receptor apoptotic signaling pathway(GO:1990117) |
| 0.6 | 3.9 | GO:0006048 | UDP-N-acetylglucosamine biosynthetic process(GO:0006048) |
| 0.6 | 2.6 | GO:0006436 | tryptophanyl-tRNA aminoacylation(GO:0006436) |
| 0.6 | 1.3 | GO:0002874 | regulation of chronic inflammatory response to antigenic stimulus(GO:0002874) |
| 0.6 | 4.0 | GO:0060075 | regulation of resting membrane potential(GO:0060075) |
| 0.5 | 1.4 | GO:1903116 | positive regulation of actin filament-based movement(GO:1903116) |
| 0.4 | 7.0 | GO:0031274 | positive regulation of pseudopodium assembly(GO:0031274) |
| 0.4 | 21.1 | GO:0060445 | branching involved in salivary gland morphogenesis(GO:0060445) |
| 0.4 | 1.3 | GO:0032877 | positive regulation of DNA endoreduplication(GO:0032877) |
| 0.4 | 1.6 | GO:0019287 | isopentenyl diphosphate biosynthetic process, mevalonate pathway(GO:0019287) |
| 0.4 | 2.3 | GO:0034975 | protein folding in endoplasmic reticulum(GO:0034975) |
| 0.4 | 4.1 | GO:0031584 | activation of phospholipase D activity(GO:0031584) |
| 0.4 | 4.0 | GO:1903297 | regulation of hypoxia-induced intrinsic apoptotic signaling pathway(GO:1903297) negative regulation of hypoxia-induced intrinsic apoptotic signaling pathway(GO:1903298) |
| 0.3 | 1.0 | GO:0090149 | mitochondrial membrane fission(GO:0090149) |
| 0.3 | 1.4 | GO:0001732 | formation of cytoplasmic translation initiation complex(GO:0001732) |
| 0.3 | 1.3 | GO:0017187 | peptidyl-glutamic acid carboxylation(GO:0017187) |
| 0.3 | 1.3 | GO:1903184 | enzyme active site formation via L-cysteine sulfinic acid(GO:0018323) primary alcohol biosynthetic process(GO:0034309) cellular response to glyoxal(GO:0036471) neuron intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:0036482) glycolate biosynthetic process(GO:0046295) positive regulation of mitochondrial electron transport, NADH to ubiquinone(GO:1902958) negative regulation of TRAIL-activated apoptotic signaling pathway(GO:1903122) regulation of pyrroline-5-carboxylate reductase activity(GO:1903167) positive regulation of pyrroline-5-carboxylate reductase activity(GO:1903168) regulation of tyrosine 3-monooxygenase activity(GO:1903176) positive regulation of tyrosine 3-monooxygenase activity(GO:1903178) L-dopa metabolic process(GO:1903184) L-dopa biosynthetic process(GO:1903185) glyoxal metabolic process(GO:1903189) regulation of L-dopa biosynthetic process(GO:1903195) positive regulation of L-dopa biosynthetic process(GO:1903197) regulation of L-dopa decarboxylase activity(GO:1903198) positive regulation of L-dopa decarboxylase activity(GO:1903200) regulation of hydrogen peroxide-induced neuron intrinsic apoptotic signaling pathway(GO:1903383) negative regulation of hydrogen peroxide-induced neuron intrinsic apoptotic signaling pathway(GO:1903384) positive regulation of cellular amino acid biosynthetic process(GO:2000284) |
| 0.3 | 2.6 | GO:0006102 | isocitrate metabolic process(GO:0006102) |
| 0.3 | 3.6 | GO:0055059 | asymmetric neuroblast division(GO:0055059) |
| 0.3 | 0.8 | GO:0002270 | plasmacytoid dendritic cell activation(GO:0002270) |
| 0.3 | 6.1 | GO:0060575 | intestinal epithelial cell differentiation(GO:0060575) |
| 0.2 | 2.7 | GO:0090166 | Golgi disassembly(GO:0090166) |
| 0.2 | 5.1 | GO:0008272 | sulfate transport(GO:0008272) |
| 0.2 | 5.7 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.2 | 1.1 | GO:0046947 | hydroxylysine metabolic process(GO:0046946) hydroxylysine biosynthetic process(GO:0046947) |
| 0.2 | 0.9 | GO:0038016 | insulin receptor internalization(GO:0038016) |
| 0.2 | 2.2 | GO:0051503 | adenine nucleotide transport(GO:0051503) |
| 0.2 | 2.7 | GO:0060628 | regulation of ER to Golgi vesicle-mediated transport(GO:0060628) |
| 0.2 | 1.0 | GO:1904059 | regulation of locomotor rhythm(GO:1904059) |
| 0.2 | 1.0 | GO:0046726 | positive regulation by virus of viral protein levels in host cell(GO:0046726) |
| 0.2 | 0.9 | GO:0048619 | embryonic hindgut morphogenesis(GO:0048619) |
| 0.2 | 3.6 | GO:0055089 | fatty acid homeostasis(GO:0055089) |
| 0.2 | 0.9 | GO:0036343 | psychomotor behavior(GO:0036343) |
| 0.2 | 0.9 | GO:0097476 | spinal cord motor neuron migration(GO:0097476) |
| 0.2 | 0.8 | GO:0070934 | CRD-mediated mRNA stabilization(GO:0070934) |
| 0.2 | 0.6 | GO:1903070 | negative regulation of ER-associated ubiquitin-dependent protein catabolic process(GO:1903070) |
| 0.2 | 4.3 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.1 | 5.6 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.1 | 0.9 | GO:0070127 | tRNA aminoacylation for mitochondrial protein translation(GO:0070127) |
| 0.1 | 6.7 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.1 | 0.7 | GO:2000767 | positive regulation of cytoplasmic translation(GO:2000767) |
| 0.1 | 1.2 | GO:0010815 | bradykinin catabolic process(GO:0010815) |
| 0.1 | 0.4 | GO:0022007 | neural plate elongation(GO:0014022) convergent extension involved in neural plate elongation(GO:0022007) |
| 0.1 | 0.7 | GO:0000103 | sulfate assimilation(GO:0000103) |
| 0.1 | 1.9 | GO:0043508 | negative regulation of JUN kinase activity(GO:0043508) |
| 0.1 | 0.5 | GO:1902775 | mitochondrial large ribosomal subunit assembly(GO:1902775) |
| 0.1 | 1.4 | GO:0032471 | negative regulation of endoplasmic reticulum calcium ion concentration(GO:0032471) |
| 0.1 | 1.5 | GO:0006995 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.1 | 0.8 | GO:0033299 | secretion of lysosomal enzymes(GO:0033299) |
| 0.1 | 1.6 | GO:0006613 | cotranslational protein targeting to membrane(GO:0006613) |
| 0.1 | 3.2 | GO:0046827 | positive regulation of protein export from nucleus(GO:0046827) |
| 0.1 | 0.4 | GO:0035519 | protein K29-linked ubiquitination(GO:0035519) |
| 0.1 | 4.1 | GO:0043277 | apoptotic cell clearance(GO:0043277) |
| 0.1 | 2.2 | GO:0018195 | peptidyl-arginine modification(GO:0018195) |
| 0.1 | 0.6 | GO:0032055 | negative regulation of translation in response to stress(GO:0032055) |
| 0.1 | 1.0 | GO:0009191 | ribonucleoside diphosphate catabolic process(GO:0009191) |
| 0.1 | 5.9 | GO:0071346 | cellular response to interferon-gamma(GO:0071346) |
| 0.1 | 1.8 | GO:0007252 | I-kappaB phosphorylation(GO:0007252) |
| 0.1 | 0.1 | GO:0007571 | age-dependent response to oxidative stress(GO:0001306) age-dependent general metabolic decline(GO:0007571) |
| 0.1 | 1.1 | GO:0044126 | regulation of growth of symbiont in host(GO:0044126) |
| 0.1 | 3.7 | GO:0007032 | endosome organization(GO:0007032) |
| 0.1 | 1.1 | GO:0006654 | phosphatidic acid biosynthetic process(GO:0006654) |
| 0.1 | 0.8 | GO:0034982 | mitochondrial protein processing(GO:0034982) |
| 0.1 | 0.5 | GO:0006450 | regulation of translational fidelity(GO:0006450) |
| 0.1 | 1.4 | GO:0099517 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.1 | 0.3 | GO:0021937 | cerebellar Purkinje cell-granule cell precursor cell signaling involved in regulation of granule cell precursor cell proliferation(GO:0021937) |
| 0.1 | 0.5 | GO:0071816 | tail-anchored membrane protein insertion into ER membrane(GO:0071816) |
| 0.1 | 1.7 | GO:0033622 | integrin activation(GO:0033622) |
| 0.0 | 0.0 | GO:0006419 | alanyl-tRNA aminoacylation(GO:0006419) |
| 0.0 | 4.2 | GO:0031529 | ruffle organization(GO:0031529) |
| 0.0 | 0.4 | GO:1904668 | positive regulation of ubiquitin protein ligase activity(GO:1904668) |
| 0.0 | 0.9 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.0 | 2.1 | GO:0001706 | endoderm formation(GO:0001706) |
| 0.0 | 6.6 | GO:0008203 | cholesterol metabolic process(GO:0008203) |
| 0.0 | 1.6 | GO:0033146 | regulation of intracellular estrogen receptor signaling pathway(GO:0033146) |
| 0.0 | 2.0 | GO:0050919 | negative chemotaxis(GO:0050919) |
| 0.0 | 0.4 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.0 | 0.2 | GO:0010587 | miRNA catabolic process(GO:0010587) |
| 0.0 | 0.7 | GO:0044819 | mitotic G1 DNA damage checkpoint(GO:0031571) mitotic G1/S transition checkpoint(GO:0044819) |
| 0.0 | 0.3 | GO:1903026 | negative regulation of RNA polymerase II regulatory region sequence-specific DNA binding(GO:1903026) |
| 0.0 | 4.1 | GO:0006694 | steroid biosynthetic process(GO:0006694) |
| 0.0 | 0.2 | GO:0060005 | vestibular reflex(GO:0060005) |
| 0.0 | 1.3 | GO:0046839 | phospholipid dephosphorylation(GO:0046839) |
| 0.0 | 0.4 | GO:0006620 | posttranslational protein targeting to membrane(GO:0006620) |
| 0.0 | 0.6 | GO:0070734 | histone H3-K27 methylation(GO:0070734) |
| 0.0 | 3.0 | GO:0032526 | response to retinoic acid(GO:0032526) |
| 0.0 | 2.1 | GO:0031338 | regulation of vesicle fusion(GO:0031338) |
| 0.0 | 0.3 | GO:0006465 | signal peptide processing(GO:0006465) |
| 0.0 | 1.5 | GO:0001938 | positive regulation of endothelial cell proliferation(GO:0001938) |
| 0.0 | 1.4 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.0 | 0.1 | GO:0000973 | posttranscriptional tethering of RNA polymerase II gene DNA at nuclear periphery(GO:0000973) |
| 0.0 | 0.2 | GO:0060770 | negative regulation of epithelial cell proliferation involved in prostate gland development(GO:0060770) |
| 0.0 | 0.8 | GO:0051452 | intracellular pH reduction(GO:0051452) |
| 0.0 | 2.7 | GO:0035113 | embryonic limb morphogenesis(GO:0030326) embryonic appendage morphogenesis(GO:0035113) |
| 0.0 | 1.2 | GO:0000045 | autophagosome assembly(GO:0000045) |
| 0.0 | 0.1 | GO:0036215 | response to stem cell factor(GO:0036215) cellular response to stem cell factor stimulus(GO:0036216) Kit signaling pathway(GO:0038109) |
| 0.0 | 0.1 | GO:0046855 | inositol phosphate dephosphorylation(GO:0046855) |
| 0.0 | 0.6 | GO:0060325 | face morphogenesis(GO:0060325) |
| 0.0 | 0.5 | GO:0097031 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.7 | 25.6 | GO:0005784 | Sec61 translocon complex(GO:0005784) translocon complex(GO:0071256) |
| 1.2 | 5.9 | GO:0070022 | transforming growth factor beta receptor homodimeric complex(GO:0070022) |
| 0.8 | 14.8 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.7 | 8.8 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.7 | 5.8 | GO:0097413 | Lewy body(GO:0097413) |
| 0.7 | 2.1 | GO:0055087 | Ski complex(GO:0055087) |
| 0.7 | 2.0 | GO:0097144 | BAX complex(GO:0097144) |
| 0.6 | 3.0 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.5 | 2.9 | GO:0097452 | GAIT complex(GO:0097452) |
| 0.5 | 1.4 | GO:0043614 | multi-eIF complex(GO:0043614) |
| 0.4 | 6.7 | GO:0005915 | zonula adherens(GO:0005915) |
| 0.4 | 6.3 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 0.4 | 5.4 | GO:0030126 | COPI vesicle coat(GO:0030126) |
| 0.4 | 3.0 | GO:0032937 | SREBP-SCAP-Insig complex(GO:0032937) |
| 0.3 | 1.2 | GO:0031084 | BLOC-2 complex(GO:0031084) |
| 0.3 | 4.9 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
| 0.3 | 3.8 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 0.3 | 1.6 | GO:0043527 | tRNA methyltransferase complex(GO:0043527) |
| 0.2 | 6.6 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.1 | 2.2 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.1 | 0.5 | GO:0005745 | m-AAA complex(GO:0005745) |
| 0.1 | 4.3 | GO:1902562 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.1 | 1.8 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.1 | 3.6 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.1 | 2.3 | GO:0070971 | endoplasmic reticulum exit site(GO:0070971) |
| 0.1 | 1.5 | GO:0044754 | autolysosome(GO:0044754) |
| 0.1 | 8.5 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.1 | 5.2 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.1 | 0.5 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.1 | 6.0 | GO:0032420 | stereocilium(GO:0032420) |
| 0.1 | 0.8 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.1 | 4.0 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.1 | 10.7 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.1 | 1.4 | GO:0031083 | BLOC-1 complex(GO:0031083) |
| 0.1 | 0.3 | GO:0044307 | dendritic branch(GO:0044307) |
| 0.1 | 0.8 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.1 | 3.1 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.1 | 0.5 | GO:0071818 | BAT3 complex(GO:0071818) ER membrane insertion complex(GO:0072379) |
| 0.1 | 0.2 | GO:0032783 | ELL-EAF complex(GO:0032783) |
| 0.1 | 8.3 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.0 | 0.7 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.0 | 0.7 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.0 | 0.3 | GO:0005787 | signal peptidase complex(GO:0005787) |
| 0.0 | 1.4 | GO:0034706 | sodium channel complex(GO:0034706) |
| 0.0 | 8.0 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.0 | 4.1 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.0 | 1.4 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 0.9 | GO:0045334 | clathrin-coated endocytic vesicle(GO:0045334) |
| 0.0 | 1.5 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.0 | 1.3 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 5.2 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.0 | 4.5 | GO:0098791 | trans-Golgi network(GO:0005802) Golgi subcompartment(GO:0098791) |
| 0.0 | 0.2 | GO:0042555 | MCM complex(GO:0042555) |
| 0.0 | 1.1 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 0.1 | GO:0044614 | nuclear pore cytoplasmic filaments(GO:0044614) |
| 0.0 | 1.0 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 10.8 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 4.6 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 1.7 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 1.0 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.0 | 3.2 | GO:0030018 | Z disc(GO:0030018) |
| 0.0 | 26.7 | GO:0005783 | endoplasmic reticulum(GO:0005783) |
| 0.0 | 2.6 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.0 | 0.4 | GO:0016235 | aggresome(GO:0016235) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.1 | 16.2 | GO:0070905 | serine binding(GO:0070905) |
| 2.5 | 12.5 | GO:0046923 | ER retention sequence binding(GO:0046923) |
| 2.1 | 6.3 | GO:0004613 | phosphoenolpyruvate carboxykinase activity(GO:0004611) phosphoenolpyruvate carboxykinase (GTP) activity(GO:0004613) |
| 1.9 | 5.7 | GO:0017099 | very-long-chain-acyl-CoA dehydrogenase activity(GO:0017099) |
| 1.4 | 6.9 | GO:0004854 | aldehyde oxidase activity(GO:0004031) xanthine dehydrogenase activity(GO:0004854) oxidoreductase activity, acting on the aldehyde or oxo group of donors, oxygen as acceptor(GO:0016623) oxidoreductase activity, acting on CH or CH2 groups, NAD or NADP as acceptor(GO:0016726) molybdenum ion binding(GO:0030151) |
| 1.3 | 3.9 | GO:0003977 | UDP-N-acetylglucosamine diphosphorylase activity(GO:0003977) |
| 1.2 | 7.4 | GO:0048408 | epidermal growth factor binding(GO:0048408) |
| 0.9 | 8.8 | GO:0004576 | oligosaccharyl transferase activity(GO:0004576) dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.8 | 2.3 | GO:0042954 | lipoprotein transporter activity(GO:0042954) |
| 0.7 | 2.2 | GO:0046964 | 3'-phosphoadenosine 5'-phosphosulfate transmembrane transporter activity(GO:0046964) |
| 0.7 | 3.7 | GO:0004430 | 1-phosphatidylinositol 4-kinase activity(GO:0004430) |
| 0.7 | 2.1 | GO:0008330 | protein tyrosine/threonine phosphatase activity(GO:0008330) |
| 0.7 | 2.7 | GO:0004096 | catalase activity(GO:0004096) |
| 0.6 | 2.6 | GO:0004830 | tryptophan-tRNA ligase activity(GO:0004830) |
| 0.6 | 5.9 | GO:1901612 | cardiolipin binding(GO:1901612) |
| 0.5 | 2.1 | GO:0004450 | isocitrate dehydrogenase (NADP+) activity(GO:0004450) |
| 0.5 | 2.6 | GO:0004448 | isocitrate dehydrogenase activity(GO:0004448) isocitrate dehydrogenase (NAD+) activity(GO:0004449) |
| 0.5 | 7.8 | GO:0038036 | sphingosine-1-phosphate receptor activity(GO:0038036) |
| 0.5 | 8.5 | GO:0015250 | water channel activity(GO:0015250) |
| 0.4 | 1.3 | GO:0016900 | oxidoreductase activity, acting on the CH-OH group of donors, disulfide as acceptor(GO:0016900) vitamin-K-epoxide reductase (warfarin-sensitive) activity(GO:0047057) |
| 0.4 | 2.5 | GO:0070568 | guanylyltransferase activity(GO:0070568) |
| 0.4 | 4.9 | GO:0017176 | phosphatidylinositol N-acetylglucosaminyltransferase activity(GO:0017176) |
| 0.3 | 2.0 | GO:0051434 | BH3 domain binding(GO:0051434) |
| 0.3 | 1.3 | GO:0008176 | tRNA (guanine-N7-)-methyltransferase activity(GO:0008176) |
| 0.3 | 1.7 | GO:0004045 | aminoacyl-tRNA hydrolase activity(GO:0004045) |
| 0.3 | 2.7 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.3 | 1.3 | GO:0036478 | tyrosine 3-monooxygenase activator activity(GO:0036470) L-dopa decarboxylase activator activity(GO:0036478) |
| 0.3 | 5.9 | GO:0035615 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
| 0.3 | 0.9 | GO:0004946 | bombesin receptor activity(GO:0004946) |
| 0.3 | 7.0 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.3 | 3.2 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.3 | 5.1 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.3 | 0.8 | GO:0000402 | open form four-way junction DNA binding(GO:0000401) crossed form four-way junction DNA binding(GO:0000402) |
| 0.3 | 6.5 | GO:0097371 | MDM2/MDM4 family protein binding(GO:0097371) |
| 0.2 | 3.8 | GO:0030881 | beta-2-microglobulin binding(GO:0030881) |
| 0.2 | 1.1 | GO:0008475 | procollagen-lysine 5-dioxygenase activity(GO:0008475) |
| 0.2 | 5.1 | GO:0016864 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.2 | 6.7 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 0.2 | 1.0 | GO:0004382 | guanosine-diphosphatase activity(GO:0004382) |
| 0.2 | 1.0 | GO:0102345 | 3-hydroxy-behenoyl-CoA dehydratase activity(GO:0102344) 3-hydroxy-lignoceroyl-CoA dehydratase activity(GO:0102345) |
| 0.2 | 4.0 | GO:0022841 | potassium ion leak channel activity(GO:0022841) |
| 0.2 | 6.2 | GO:0004602 | glutathione peroxidase activity(GO:0004602) |
| 0.2 | 1.2 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.2 | 3.6 | GO:0035174 | histone serine kinase activity(GO:0035174) |
| 0.2 | 2.3 | GO:0016671 | oxidoreductase activity, acting on a sulfur group of donors, disulfide as acceptor(GO:0016671) |
| 0.2 | 6.3 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.2 | 4.1 | GO:0004033 | aldo-keto reductase (NADP) activity(GO:0004033) |
| 0.1 | 0.6 | GO:0004694 | eukaryotic translation initiation factor 2alpha kinase activity(GO:0004694) |
| 0.1 | 1.1 | GO:0035727 | lysophosphatidic acid binding(GO:0035727) |
| 0.1 | 1.5 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 0.1 | 0.6 | GO:1904288 | BAT3 complex binding(GO:1904288) |
| 0.1 | 6.6 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.1 | 0.5 | GO:1990932 | 5.8S rRNA binding(GO:1990932) |
| 0.1 | 0.7 | GO:0050294 | alcohol sulfotransferase activity(GO:0004027) steroid sulfotransferase activity(GO:0050294) |
| 0.1 | 2.0 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.1 | 1.1 | GO:0008484 | sulfuric ester hydrolase activity(GO:0008484) |
| 0.1 | 0.6 | GO:0005047 | signal recognition particle binding(GO:0005047) |
| 0.1 | 0.3 | GO:0016426 | tRNA (adenine) methyltransferase activity(GO:0016426) |
| 0.1 | 0.9 | GO:0031419 | cobalamin binding(GO:0031419) |
| 0.1 | 3.5 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 1.6 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.1 | 5.0 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) |
| 0.1 | 0.6 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.1 | 3.1 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.1 | 3.1 | GO:0004812 | aminoacyl-tRNA ligase activity(GO:0004812) ligase activity, forming carbon-oxygen bonds(GO:0016875) ligase activity, forming aminoacyl-tRNA and related compounds(GO:0016876) |
| 0.1 | 20.2 | GO:0003729 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.1 | 1.8 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.1 | 0.8 | GO:0015929 | hexosaminidase activity(GO:0015929) |
| 0.1 | 0.2 | GO:0004176 | ATP-dependent peptidase activity(GO:0004176) |
| 0.1 | 1.3 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.1 | 6.8 | GO:0000149 | SNARE binding(GO:0000149) |
| 0.0 | 0.8 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
| 0.0 | 1.7 | GO:0070273 | phosphatidylinositol-4-phosphate binding(GO:0070273) |
| 0.0 | 2.4 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 9.6 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.0 | 1.2 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.0 | 0.6 | GO:0008641 | small protein activating enzyme activity(GO:0008641) |
| 0.0 | 0.6 | GO:0008536 | Ran GTPase binding(GO:0008536) |
| 0.0 | 1.1 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.0 | 0.8 | GO:0008308 | voltage-gated anion channel activity(GO:0008308) |
| 0.0 | 0.3 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.0 | 0.4 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.0 | 1.6 | GO:0016790 | thiolester hydrolase activity(GO:0016790) |
| 0.0 | 1.4 | GO:0005227 | calcium activated cation channel activity(GO:0005227) |
| 0.0 | 4.0 | GO:0042626 | ATPase activity, coupled to transmembrane movement of substances(GO:0042626) ATPase activity, coupled to movement of substances(GO:0043492) |
| 0.0 | 0.5 | GO:0005527 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.0 | 9.6 | GO:0045296 | cadherin binding(GO:0045296) |
| 0.0 | 1.1 | GO:0016776 | phosphotransferase activity, phosphate group as acceptor(GO:0016776) |
| 0.0 | 5.6 | GO:0004866 | endopeptidase inhibitor activity(GO:0004866) |
| 0.0 | 0.5 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.0 | 0.2 | GO:0008381 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.0 | 1.2 | GO:0008375 | acetylglucosaminyltransferase activity(GO:0008375) |
| 0.0 | 2.2 | GO:0008565 | protein transporter activity(GO:0008565) |
| 0.0 | 0.5 | GO:0004177 | aminopeptidase activity(GO:0004177) |
| 0.0 | 11.2 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.0 | 2.3 | GO:0004386 | helicase activity(GO:0004386) |
| 0.0 | 1.7 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 1.7 | GO:0020037 | heme binding(GO:0020037) |
| 0.0 | 1.3 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 0.6 | GO:0031491 | nucleosome binding(GO:0031491) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 5.2 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.2 | 18.7 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.2 | 6.3 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.2 | 7.8 | PID S1P META PATHWAY | Sphingosine 1-phosphate (S1P) pathway |
| 0.1 | 4.9 | ST GAQ PATHWAY | G alpha q Pathway |
| 0.1 | 2.0 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.1 | 6.3 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 1.8 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.0 | 3.6 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.0 | 8.3 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.0 | 1.1 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.0 | 4.1 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 0.9 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.0 | 1.3 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.0 | 0.9 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.0 | 0.8 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 1.7 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 4.1 | PID ERBB1 DOWNSTREAM PATHWAY | ErbB1 downstream signaling |
| 0.0 | 0.8 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.0 | 1.3 | PID AP1 PATHWAY | AP-1 transcription factor network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 8.5 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.5 | 6.2 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.4 | 4.5 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.4 | 6.3 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.4 | 5.2 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.4 | 7.3 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.3 | 2.9 | REACTOME THE ACTIVATION OF ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.3 | 16.6 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.3 | 5.9 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 0.3 | 3.6 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.3 | 4.0 | REACTOME TANDEM PORE DOMAIN POTASSIUM CHANNELS | Genes involved in Tandem pore domain potassium channels |
| 0.2 | 6.2 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.2 | 6.7 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
| 0.2 | 4.7 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.2 | 0.7 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.2 | 6.7 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.2 | 3.7 | REACTOME SYNTHESIS OF PIPS AT THE EARLY ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the early endosome membrane |
| 0.2 | 26.9 | REACTOME SRP DEPENDENT COTRANSLATIONAL PROTEIN TARGETING TO MEMBRANE | Genes involved in SRP-dependent cotranslational protein targeting to membrane |
| 0.2 | 4.9 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.2 | 3.5 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.1 | 6.9 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.1 | 3.6 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.1 | 2.2 | REACTOME TRNA AMINOACYLATION | Genes involved in tRNA Aminoacylation |
| 0.1 | 1.4 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.1 | 0.8 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.1 | 1.9 | REACTOME N GLYCAN ANTENNAE ELONGATION IN THE MEDIAL TRANS GOLGI | Genes involved in N-glycan antennae elongation in the medial/trans-Golgi |
| 0.1 | 0.7 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.1 | 1.6 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 1.6 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.0 | 2.2 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.0 | 2.8 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 6.1 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.0 | 2.0 | REACTOME INTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Intrinsic Pathway for Apoptosis |
| 0.0 | 2.2 | REACTOME ASPARAGINE N LINKED GLYCOSYLATION | Genes involved in Asparagine N-linked glycosylation |
| 0.0 | 2.5 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.0 | 0.7 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.0 | 8.7 | REACTOME CLASS A1 RHODOPSIN LIKE RECEPTORS | Genes involved in Class A/1 (Rhodopsin-like receptors) |
| 0.0 | 1.1 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.6 | REACTOME TRANSPORT OF RIBONUCLEOPROTEINS INTO THE HOST NUCLEUS | Genes involved in Transport of Ribonucleoproteins into the Host Nucleus |
| 0.0 | 1.0 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 1.7 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |