Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
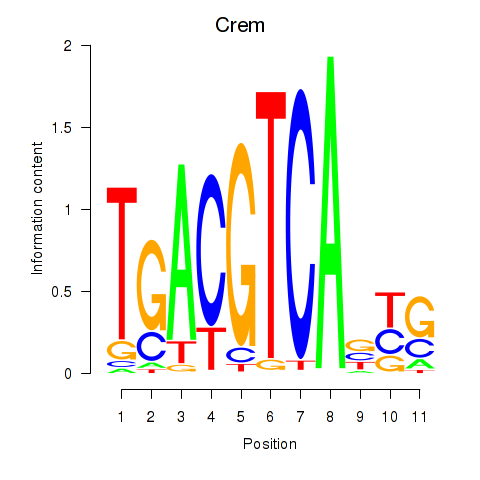
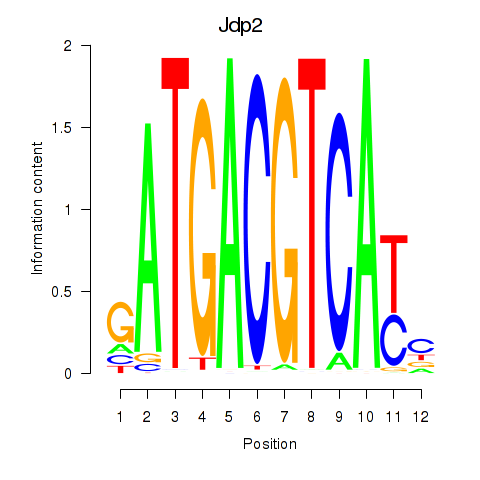
Results for Crem_Jdp2
Z-value: 2.38
Transcription factors associated with Crem_Jdp2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Crem
|
ENSMUSG00000063889.17 | Crem |
|
Jdp2
|
ENSMUSG00000034271.17 | Jdp2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Crem | mm39_v1_chr18_-_3299452_3299524 | -0.71 | 4.5e-12 | Click! |
| Jdp2 | mm39_v1_chr12_+_85646162_85646190 | 0.35 | 2.4e-03 | Click! |
Activity profile of Crem_Jdp2 motif
Sorted Z-values of Crem_Jdp2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Crem_Jdp2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr7_+_97437709 | 41.50 |
ENSMUST00000206984.2
|
Pak1
|
p21 (RAC1) activated kinase 1 |
| chr1_-_79417732 | 41.17 |
ENSMUST00000185234.2
ENSMUST00000049972.6 |
Scg2
|
secretogranin II |
| chr7_-_141649003 | 33.12 |
ENSMUST00000039926.10
|
Dusp8
|
dual specificity phosphatase 8 |
| chr2_+_155118217 | 31.99 |
ENSMUST00000029128.4
|
Map1lc3a
|
microtubule-associated protein 1 light chain 3 alpha |
| chr2_+_132623198 | 25.67 |
ENSMUST00000028826.4
|
Chgb
|
chromogranin B |
| chr9_-_57375269 | 22.69 |
ENSMUST00000215059.2
ENSMUST00000046587.8 ENSMUST00000214256.2 |
Scamp5
|
secretory carrier membrane protein 5 |
| chr11_-_6556053 | 21.55 |
ENSMUST00000045713.4
|
Nacad
|
NAC alpha domain containing |
| chrX_-_20787150 | 21.51 |
ENSMUST00000081893.7
ENSMUST00000115345.8 |
Syn1
|
synapsin I |
| chr7_+_112278534 | 21.51 |
ENSMUST00000106638.10
|
Tead1
|
TEA domain family member 1 |
| chr9_-_96634874 | 21.49 |
ENSMUST00000152594.8
|
Zbtb38
|
zinc finger and BTB domain containing 38 |
| chr14_+_66581818 | 20.58 |
ENSMUST00000118426.8
ENSMUST00000121955.8 ENSMUST00000120229.8 ENSMUST00000134440.2 |
Stmn4
|
stathmin-like 4 |
| chr10_+_29087658 | 20.28 |
ENSMUST00000213489.2
|
9330159F19Rik
|
RIKEN cDNA 9330159F19 gene |
| chr1_-_36748985 | 19.97 |
ENSMUST00000043951.10
|
Actr1b
|
ARP1 actin-related protein 1B, centractin beta |
| chr8_-_24928953 | 19.22 |
ENSMUST00000052622.6
|
Tcim
|
transcriptional and immune response regulator |
| chr14_+_66581745 | 18.82 |
ENSMUST00000152093.8
ENSMUST00000074523.13 |
Stmn4
|
stathmin-like 4 |
| chr4_-_150736554 | 18.67 |
ENSMUST00000117997.2
ENSMUST00000037827.10 |
Slc45a1
|
solute carrier family 45, member 1 |
| chr3_+_82265474 | 18.57 |
ENSMUST00000195471.6
ENSMUST00000195640.2 |
Map9
|
microtubule-associated protein 9 |
| chr2_+_143388062 | 18.08 |
ENSMUST00000028905.10
|
Pcsk2
|
proprotein convertase subtilisin/kexin type 2 |
| chr3_+_82265351 | 17.96 |
ENSMUST00000193559.6
ENSMUST00000192595.6 ENSMUST00000091014.10 |
Map9
|
microtubule-associated protein 9 |
| chr13_-_34261975 | 17.33 |
ENSMUST00000056427.10
|
Tubb2a
|
tubulin, beta 2A class IIA |
| chr18_-_35348049 | 17.23 |
ENSMUST00000091636.5
ENSMUST00000236680.2 |
Lrrtm2
|
leucine rich repeat transmembrane neuronal 2 |
| chr5_-_114582097 | 16.74 |
ENSMUST00000031560.14
|
Mmab
|
methylmalonic aciduria (cobalamin deficiency) cblB type homolog (human) |
| chr10_+_29087602 | 16.47 |
ENSMUST00000092627.6
|
9330159F19Rik
|
RIKEN cDNA 9330159F19 gene |
| chr7_+_112278520 | 16.21 |
ENSMUST00000084705.13
ENSMUST00000239442.2 ENSMUST00000239404.2 ENSMUST00000059768.18 |
Tead1
|
TEA domain family member 1 |
| chr17_-_26727437 | 15.93 |
ENSMUST00000236661.2
ENSMUST00000025025.7 |
Dusp1
|
dual specificity phosphatase 1 |
| chr6_+_54658609 | 15.67 |
ENSMUST00000190641.7
ENSMUST00000187701.2 |
Mturn
|
maturin, neural progenitor differentiation regulator homolog (Xenopus) |
| chr7_+_16716989 | 15.53 |
ENSMUST00000206129.3
|
Gm42372
|
predicted gene, 42372 |
| chr3_-_88455556 | 15.51 |
ENSMUST00000131775.2
ENSMUST00000008745.13 |
Rab25
|
RAB25, member RAS oncogene family |
| chr2_+_136555364 | 15.42 |
ENSMUST00000028727.11
ENSMUST00000110098.4 |
Snap25
|
synaptosomal-associated protein 25 |
| chr11_+_101358990 | 15.19 |
ENSMUST00000001347.7
|
Rnd2
|
Rho family GTPase 2 |
| chr17_-_24908874 | 15.07 |
ENSMUST00000007236.5
|
Syngr3
|
synaptogyrin 3 |
| chr6_-_124441731 | 14.90 |
ENSMUST00000008297.5
|
Clstn3
|
calsyntenin 3 |
| chr19_-_5135510 | 14.68 |
ENSMUST00000140389.8
ENSMUST00000151413.2 ENSMUST00000077066.8 |
Tmem151a
|
transmembrane protein 151A |
| chr9_-_97915227 | 14.66 |
ENSMUST00000035027.13
|
Clstn2
|
calsyntenin 2 |
| chr7_-_126548671 | 14.33 |
ENSMUST00000106339.2
ENSMUST00000052937.12 |
Asphd1
|
aspartate beta-hydroxylase domain containing 1 |
| chr5_-_114582053 | 14.28 |
ENSMUST00000123256.3
ENSMUST00000112245.6 |
Mmab
|
methylmalonic aciduria (cobalamin deficiency) cblB type homolog (human) |
| chr17_+_44263890 | 14.22 |
ENSMUST00000177857.9
ENSMUST00000044792.6 |
Rcan2
|
regulator of calcineurin 2 |
| chr15_+_79975520 | 14.12 |
ENSMUST00000009728.13
ENSMUST00000009727.12 |
Syngr1
|
synaptogyrin 1 |
| chr9_-_52591030 | 13.74 |
ENSMUST00000213937.2
|
AI593442
|
expressed sequence AI593442 |
| chr11_-_90281721 | 13.66 |
ENSMUST00000004051.8
|
Hlf
|
hepatic leukemia factor |
| chr13_-_54836059 | 13.53 |
ENSMUST00000122935.2
ENSMUST00000128257.8 |
Rnf44
|
ring finger protein 44 |
| chrX_-_20928219 | 13.34 |
ENSMUST00000040628.6
ENSMUST00000115333.9 ENSMUST00000115334.8 |
Zfp182
|
zinc finger protein 182 |
| chr2_+_49509288 | 13.24 |
ENSMUST00000028102.14
|
Kif5c
|
kinesin family member 5C |
| chr1_-_92401459 | 12.97 |
ENSMUST00000185251.2
ENSMUST00000027478.7 |
Ndufa10
|
NADH:ubiquinone oxidoreductase subunit A10 |
| chr11_+_69920542 | 12.87 |
ENSMUST00000232266.2
ENSMUST00000132597.5 |
Dlg4
|
discs large MAGUK scaffold protein 4 |
| chr18_+_35347983 | 12.79 |
ENSMUST00000235449.2
ENSMUST00000235269.2 |
Ctnna1
|
catenin (cadherin associated protein), alpha 1 |
| chr7_-_45016138 | 12.77 |
ENSMUST00000211067.2
ENSMUST00000003961.16 |
Ppfia3
|
protein tyrosine phosphatase, receptor type, f polypeptide (PTPRF), interacting protein (liprin), alpha 3 |
| chr5_+_114582327 | 12.73 |
ENSMUST00000137167.8
ENSMUST00000112239.9 ENSMUST00000124260.8 ENSMUST00000125650.6 ENSMUST00000043760.15 |
Mvk
|
mevalonate kinase |
| chr17_+_44264130 | 12.58 |
ENSMUST00000229240.2
|
Rcan2
|
regulator of calcineurin 2 |
| chr13_-_54835996 | 12.53 |
ENSMUST00000150806.8
ENSMUST00000125927.8 |
Rnf44
|
ring finger protein 44 |
| chr13_-_54835878 | 12.31 |
ENSMUST00000125871.8
|
Rnf44
|
ring finger protein 44 |
| chr1_-_93029532 | 12.18 |
ENSMUST00000171796.8
|
Kif1a
|
kinesin family member 1A |
| chr1_-_77491683 | 12.09 |
ENSMUST00000186930.2
ENSMUST00000027451.13 ENSMUST00000188797.7 |
Epha4
|
Eph receptor A4 |
| chr9_+_102595628 | 12.07 |
ENSMUST00000156485.2
ENSMUST00000145937.2 ENSMUST00000134483.2 ENSMUST00000190047.7 |
Amotl2
|
angiomotin-like 2 |
| chr10_-_67748461 | 12.01 |
ENSMUST00000064656.8
|
Zfp365
|
zinc finger protein 365 |
| chr2_-_130484689 | 11.78 |
ENSMUST00000045761.7
|
Lzts3
|
leucine zipper, putative tumor suppressor family member 3 |
| chr12_+_102521225 | 11.58 |
ENSMUST00000021610.7
|
Chga
|
chromogranin A |
| chr16_-_23709564 | 11.50 |
ENSMUST00000004480.5
|
Sst
|
somatostatin |
| chr11_-_78056347 | 11.49 |
ENSMUST00000017530.4
|
Traf4
|
TNF receptor associated factor 4 |
| chr5_+_142946098 | 11.46 |
ENSMUST00000031565.15
ENSMUST00000198017.5 |
Fscn1
|
fascin actin-bundling protein 1 |
| chr3_-_88669551 | 11.33 |
ENSMUST00000183267.2
|
Syt11
|
synaptotagmin XI |
| chr14_+_47710005 | 11.29 |
ENSMUST00000043112.9
ENSMUST00000163324.8 ENSMUST00000228668.2 ENSMUST00000168833.9 |
Fbxo34
|
F-box protein 34 |
| chr8_+_94537460 | 11.28 |
ENSMUST00000034198.15
ENSMUST00000125716.8 |
Gnao1
|
guanine nucleotide binding protein, alpha O |
| chr5_-_124170305 | 11.26 |
ENSMUST00000040967.9
|
Vps37b
|
vacuolar protein sorting 37B |
| chr16_-_4608084 | 11.25 |
ENSMUST00000118703.8
|
Cdip1
|
cell death inducing Trp53 target 1 |
| chr1_-_33946802 | 11.17 |
ENSMUST00000115161.8
ENSMUST00000129464.8 ENSMUST00000062289.11 |
Bend6
|
BEN domain containing 6 |
| chr9_+_20914211 | 11.05 |
ENSMUST00000214124.2
ENSMUST00000216818.2 |
Mrpl4
|
mitochondrial ribosomal protein L4 |
| chr1_-_93029547 | 11.02 |
ENSMUST00000112958.9
ENSMUST00000186861.2 ENSMUST00000171556.8 |
Kif1a
|
kinesin family member 1A |
| chr3_+_144998233 | 10.89 |
ENSMUST00000029848.5
ENSMUST00000139001.2 |
Col24a1
|
collagen, type XXIV, alpha 1 |
| chr2_-_109111064 | 10.87 |
ENSMUST00000147770.2
|
Mettl15
|
methyltransferase like 15 |
| chr9_-_97915036 | 10.78 |
ENSMUST00000162295.2
|
Clstn2
|
calsyntenin 2 |
| chr10_-_102326286 | 10.69 |
ENSMUST00000020040.5
|
Nts
|
neurotensin |
| chr7_-_6733411 | 10.64 |
ENSMUST00000239104.2
ENSMUST00000051209.11 |
Peg3
|
paternally expressed 3 |
| chr6_-_30304512 | 10.63 |
ENSMUST00000094543.3
ENSMUST00000102993.10 |
Ube2h
|
ubiquitin-conjugating enzyme E2H |
| chrX_+_73352694 | 10.56 |
ENSMUST00000130581.2
|
Gdi1
|
guanosine diphosphate (GDP) dissociation inhibitor 1 |
| chr10_+_44144346 | 10.27 |
ENSMUST00000039286.5
|
Atg5
|
autophagy related 5 |
| chr6_-_50543514 | 10.26 |
ENSMUST00000161401.2
|
Cycs
|
cytochrome c, somatic |
| chr5_-_137529251 | 10.18 |
ENSMUST00000132525.8
|
Gnb2
|
guanine nucleotide binding protein (G protein), beta 2 |
| chr17_-_31731190 | 10.08 |
ENSMUST00000237127.2
|
Wdr4
|
WD repeat domain 4 |
| chr6_+_124908341 | 10.05 |
ENSMUST00000203021.3
|
Mlf2
|
myeloid leukemia factor 2 |
| chr12_-_17226889 | 9.96 |
ENSMUST00000170580.3
|
Kcnf1
|
potassium voltage-gated channel, subfamily F, member 1 |
| chr4_-_131565542 | 9.93 |
ENSMUST00000030741.9
ENSMUST00000105987.9 |
Ptpru
|
protein tyrosine phosphatase, receptor type, U |
| chr5_-_137529465 | 9.91 |
ENSMUST00000150063.9
|
Gnb2
|
guanine nucleotide binding protein (G protein), beta 2 |
| chr9_+_66620959 | 9.83 |
ENSMUST00000071889.13
|
Car12
|
carbonic anhydrase 12 |
| chr9_+_66621001 | 9.78 |
ENSMUST00000085420.12
|
Car12
|
carbonic anhydrase 12 |
| chr15_-_33687986 | 9.76 |
ENSMUST00000042021.5
|
Tspyl5
|
testis-specific protein, Y-encoded-like 5 |
| chr7_-_19043955 | 9.76 |
ENSMUST00000207334.2
ENSMUST00000208505.2 ENSMUST00000207716.2 ENSMUST00000208326.2 ENSMUST00000003640.4 |
Fosb
|
FBJ osteosarcoma oncogene B |
| chr14_+_47710085 | 9.75 |
ENSMUST00000095941.9
|
Fbxo34
|
F-box protein 34 |
| chr6_+_129510145 | 9.74 |
ENSMUST00000204487.3
|
Gabarapl1
|
gamma-aminobutyric acid (GABA) A receptor-associated protein-like 1 |
| chr11_+_69214895 | 9.63 |
ENSMUST00000060956.13
|
Trappc1
|
trafficking protein particle complex 1 |
| chr5_-_121329385 | 9.56 |
ENSMUST00000054547.9
ENSMUST00000100770.9 |
Ptpn11
|
protein tyrosine phosphatase, non-receptor type 11 |
| chr11_+_52655461 | 9.55 |
ENSMUST00000036796.8
|
Fstl4
|
follistatin-like 4 |
| chrX_+_65696608 | 9.45 |
ENSMUST00000036043.5
|
Slitrk2
|
SLIT and NTRK-like family, member 2 |
| chr7_+_130294262 | 9.42 |
ENSMUST00000033141.7
|
Tacc2
|
transforming, acidic coiled-coil containing protein 2 |
| chr15_-_83989801 | 9.39 |
ENSMUST00000229826.2
ENSMUST00000082365.6 |
Sult4a1
|
sulfotransferase family 4A, member 1 |
| chr17_-_56783376 | 9.39 |
ENSMUST00000223859.2
|
Ptprs
|
protein tyrosine phosphatase, receptor type, S |
| chr19_-_61215743 | 9.31 |
ENSMUST00000237386.2
|
Csf2ra
|
colony stimulating factor 2 receptor, alpha, low-affinity (granulocyte-macrophage) |
| chr17_-_56783462 | 9.27 |
ENSMUST00000067538.6
|
Ptprs
|
protein tyrosine phosphatase, receptor type, S |
| chr2_-_36026614 | 9.26 |
ENSMUST00000122456.8
|
Rbm18
|
RNA binding motif protein 18 |
| chr1_-_75240551 | 9.11 |
ENSMUST00000186178.7
ENSMUST00000189769.7 ENSMUST00000027404.12 |
Ptprn
|
protein tyrosine phosphatase, receptor type, N |
| chr9_+_20914012 | 9.04 |
ENSMUST00000003386.7
|
Mrpl4
|
mitochondrial ribosomal protein L4 |
| chr6_+_124908439 | 9.02 |
ENSMUST00000032214.14
|
Mlf2
|
myeloid leukemia factor 2 |
| chr5_-_113285852 | 9.00 |
ENSMUST00000212276.2
|
2900026A02Rik
|
RIKEN cDNA 2900026A02 gene |
| chr4_+_21931290 | 8.99 |
ENSMUST00000029908.8
|
Faxc
|
failed axon connections homolog |
| chr8_+_4375212 | 8.90 |
ENSMUST00000127460.8
ENSMUST00000136191.8 |
Ccl25
|
chemokine (C-C motif) ligand 25 |
| chrX_+_72716756 | 8.89 |
ENSMUST00000033752.14
ENSMUST00000114467.9 |
Slc6a8
|
solute carrier family 6 (neurotransmitter transporter, creatine), member 8 |
| chr9_-_20638233 | 8.89 |
ENSMUST00000217198.2
|
Olfm2
|
olfactomedin 2 |
| chr13_-_54759086 | 8.89 |
ENSMUST00000049575.8
|
Cltb
|
clathrin, light polypeptide (Lcb) |
| chr1_-_93029576 | 8.83 |
ENSMUST00000190723.7
|
Kif1a
|
kinesin family member 1A |
| chr10_+_79977291 | 8.74 |
ENSMUST00000105367.8
|
Atp5d
|
ATP synthase, H+ transporting, mitochondrial F1 complex, delta subunit |
| chr11_+_110858842 | 8.69 |
ENSMUST00000180023.8
ENSMUST00000106636.8 |
Kcnj16
|
potassium inwardly-rectifying channel, subfamily J, member 16 |
| chr10_+_121575819 | 8.68 |
ENSMUST00000065600.8
ENSMUST00000136432.2 |
BC048403
|
cDNA sequence BC048403 |
| chr11_+_69214789 | 8.65 |
ENSMUST00000102602.8
|
Trappc1
|
trafficking protein particle complex 1 |
| chr6_+_135042649 | 8.65 |
ENSMUST00000050104.8
|
Gprc5a
|
G protein-coupled receptor, family C, group 5, member A |
| chr11_+_69214883 | 8.64 |
ENSMUST00000102601.10
|
Trappc1
|
trafficking protein particle complex 1 |
| chr10_-_17823736 | 8.64 |
ENSMUST00000037879.8
|
Heca
|
hdc homolog, cell cycle regulator |
| chr6_+_124908389 | 8.55 |
ENSMUST00000180095.4
|
Mlf2
|
myeloid leukemia factor 2 |
| chr7_+_24230063 | 8.51 |
ENSMUST00000049020.9
|
Irgq
|
immunity-related GTPase family, Q |
| chr4_+_42950367 | 8.44 |
ENSMUST00000084662.12
|
Dnajb5
|
DnaJ heat shock protein family (Hsp40) member B5 |
| chr7_+_54485336 | 8.43 |
ENSMUST00000082373.8
|
Luzp2
|
leucine zipper protein 2 |
| chr11_+_69214971 | 8.40 |
ENSMUST00000108662.2
|
Trappc1
|
trafficking protein particle complex 1 |
| chr17_-_31731222 | 8.39 |
ENSMUST00000236665.2
|
Wdr4
|
WD repeat domain 4 |
| chr19_-_34452537 | 8.25 |
ENSMUST00000050562.6
|
Ch25h
|
cholesterol 25-hydroxylase |
| chr4_+_85123654 | 8.19 |
ENSMUST00000030212.15
ENSMUST00000107189.8 ENSMUST00000107184.8 |
Sh3gl2
|
SH3-domain GRB2-like 2 |
| chr11_+_52122836 | 8.14 |
ENSMUST00000037324.12
ENSMUST00000166537.8 |
Skp1
|
S-phase kinase-associated protein 1 |
| chr15_+_12321558 | 8.13 |
ENSMUST00000226517.2
|
Golph3
|
golgi phosphoprotein 3 |
| chr8_+_46338557 | 8.13 |
ENSMUST00000210422.2
|
Pdlim3
|
PDZ and LIM domain 3 |
| chrX_+_169106356 | 8.09 |
ENSMUST00000178693.4
|
Asmt
|
acetylserotonin O-methyltransferase |
| chr4_+_141028539 | 8.09 |
ENSMUST00000006614.3
|
Epha2
|
Eph receptor A2 |
| chr7_+_130294403 | 8.08 |
ENSMUST00000207282.2
|
Tacc2
|
transforming, acidic coiled-coil containing protein 2 |
| chr15_-_37459570 | 8.05 |
ENSMUST00000119730.8
ENSMUST00000120746.8 |
Ncald
|
neurocalcin delta |
| chr1_+_24717968 | 8.04 |
ENSMUST00000095062.10
|
Lmbrd1
|
LMBR1 domain containing 1 |
| chr6_+_125016723 | 8.02 |
ENSMUST00000140131.8
ENSMUST00000032480.14 |
Ing4
|
inhibitor of growth family, member 4 |
| chr13_-_54759145 | 8.01 |
ENSMUST00000091609.11
|
Cltb
|
clathrin, light polypeptide (Lcb) |
| chr8_+_45388466 | 7.99 |
ENSMUST00000191428.7
|
Fat1
|
FAT atypical cadherin 1 |
| chr5_+_25427860 | 7.90 |
ENSMUST00000045737.14
|
Galnt11
|
polypeptide N-acetylgalactosaminyltransferase 11 |
| chr12_+_79177523 | 7.90 |
ENSMUST00000021550.7
|
Arg2
|
arginase type II |
| chr6_-_115569504 | 7.86 |
ENSMUST00000112957.2
|
Mkrn2os
|
makorin, ring finger protein 2, opposite strand |
| chr4_+_85123358 | 7.83 |
ENSMUST00000107188.10
|
Sh3gl2
|
SH3-domain GRB2-like 2 |
| chr6_-_55658242 | 7.82 |
ENSMUST00000044767.10
|
Neurod6
|
neurogenic differentiation 6 |
| chr11_-_3454766 | 7.80 |
ENSMUST00000044507.12
|
Inpp5j
|
inositol polyphosphate 5-phosphatase J |
| chr2_-_36026785 | 7.73 |
ENSMUST00000028251.10
|
Rbm18
|
RNA binding motif protein 18 |
| chr7_-_7301760 | 7.65 |
ENSMUST00000210061.2
|
Clcn4
|
chloride channel, voltage-sensitive 4 |
| chr3_-_19749489 | 7.65 |
ENSMUST00000061294.5
|
Crh
|
corticotropin releasing hormone |
| chr1_+_75456173 | 7.55 |
ENSMUST00000113575.9
ENSMUST00000148980.2 ENSMUST00000050899.7 ENSMUST00000187411.2 |
Tmem198
|
transmembrane protein 198 |
| chr9_+_40180569 | 7.53 |
ENSMUST00000176185.8
|
Scn3b
|
sodium channel, voltage-gated, type III, beta |
| chr17_+_9068805 | 7.50 |
ENSMUST00000115720.8
|
Pde10a
|
phosphodiesterase 10A |
| chr6_+_129510117 | 7.49 |
ENSMUST00000032264.9
|
Gabarapl1
|
gamma-aminobutyric acid (GABA) A receptor-associated protein-like 1 |
| chr16_+_57369595 | 7.44 |
ENSMUST00000159414.2
|
Filip1l
|
filamin A interacting protein 1-like |
| chr11_+_83193495 | 7.42 |
ENSMUST00000176430.8
ENSMUST00000065692.14 ENSMUST00000142680.2 |
Ap2b1
|
adaptor-related protein complex 2, beta 1 subunit |
| chr11_+_113510135 | 7.38 |
ENSMUST00000146390.3
|
Sstr2
|
somatostatin receptor 2 |
| chr2_+_105499233 | 7.36 |
ENSMUST00000111086.11
ENSMUST00000111087.10 |
Pax6
|
paired box 6 |
| chrX_+_142447286 | 7.35 |
ENSMUST00000112868.8
|
Pak3
|
p21 (RAC1) activated kinase 3 |
| chr9_+_64086553 | 7.34 |
ENSMUST00000034965.8
|
Snapc5
|
small nuclear RNA activating complex, polypeptide 5 |
| chr11_+_113510207 | 7.34 |
ENSMUST00000106630.2
|
Sstr2
|
somatostatin receptor 2 |
| chr9_+_40180497 | 7.31 |
ENSMUST00000049941.12
|
Scn3b
|
sodium channel, voltage-gated, type III, beta |
| chr11_+_93935156 | 7.28 |
ENSMUST00000024979.15
|
Spag9
|
sperm associated antigen 9 |
| chr17_-_24388384 | 7.27 |
ENSMUST00000024932.12
|
Atp6v0c
|
ATPase, H+ transporting, lysosomal V0 subunit C |
| chr15_+_12321620 | 7.26 |
ENSMUST00000228671.2
|
Golph3
|
golgi phosphoprotein 3 |
| chr18_-_43925932 | 7.23 |
ENSMUST00000237926.2
ENSMUST00000096570.4 |
Gm94
|
predicted gene 94 |
| chr2_+_112285342 | 7.21 |
ENSMUST00000069747.6
|
Emc7
|
ER membrane protein complex subunit 7 |
| chr18_+_36481706 | 7.17 |
ENSMUST00000235864.2
ENSMUST00000050584.10 |
Cystm1
|
cysteine-rich transmembrane module containing 1 |
| chr9_-_45847344 | 7.16 |
ENSMUST00000034590.4
|
Tagln
|
transgelin |
| chr11_+_102992508 | 7.14 |
ENSMUST00000107040.10
ENSMUST00000140372.8 ENSMUST00000024492.15 ENSMUST00000134884.8 |
Acbd4
|
acyl-Coenzyme A binding domain containing 4 |
| chr11_+_93934940 | 7.08 |
ENSMUST00000132079.8
|
Spag9
|
sperm associated antigen 9 |
| chr15_+_81820954 | 7.05 |
ENSMUST00000038757.8
ENSMUST00000230633.2 |
Csdc2
|
cold shock domain containing C2, RNA binding |
| chr17_-_33979280 | 6.99 |
ENSMUST00000173860.8
|
Rab11b
|
RAB11B, member RAS oncogene family |
| chr5_+_137059127 | 6.96 |
ENSMUST00000041543.9
ENSMUST00000186451.2 |
Vgf
|
VGF nerve growth factor inducible |
| chr16_-_44153498 | 6.94 |
ENSMUST00000047446.13
|
Sidt1
|
SID1 transmembrane family, member 1 |
| chr11_+_52287239 | 6.93 |
ENSMUST00000036952.5
ENSMUST00000109057.8 |
9530068E07Rik
|
RIKEN cDNA 9530068E07 gene |
| chr15_-_77129786 | 6.91 |
ENSMUST00000228558.2
|
Rbfox2
|
RNA binding protein, fox-1 homolog (C. elegans) 2 |
| chrX_-_8042129 | 6.86 |
ENSMUST00000143984.2
|
Tbc1d25
|
TBC1 domain family, member 25 |
| chr8_+_46338498 | 6.82 |
ENSMUST00000034053.7
|
Pdlim3
|
PDZ and LIM domain 3 |
| chr2_+_30127692 | 6.78 |
ENSMUST00000113654.8
ENSMUST00000095078.3 |
Lrrc8a
|
leucine rich repeat containing 8A VRAC subunit A |
| chr11_+_52123016 | 6.77 |
ENSMUST00000109072.2
|
Skp1
|
S-phase kinase-associated protein 1 |
| chr10_-_20600797 | 6.77 |
ENSMUST00000020165.14
|
Pde7b
|
phosphodiesterase 7B |
| chr7_+_6733684 | 6.76 |
ENSMUST00000197117.5
|
Usp29
|
ubiquitin specific peptidase 29 |
| chr3_+_36120128 | 6.76 |
ENSMUST00000011492.15
|
Acad9
|
acyl-Coenzyme A dehydrogenase family, member 9 |
| chrX_-_55643429 | 6.75 |
ENSMUST00000059899.3
|
Mmgt1
|
membrane magnesium transporter 1 |
| chr17_+_44389704 | 6.75 |
ENSMUST00000154166.8
ENSMUST00000024756.5 |
Enpp5
|
ectonucleotide pyrophosphatase/phosphodiesterase 5 |
| chr11_-_86561980 | 6.71 |
ENSMUST00000143991.3
|
Vmp1
|
vacuole membrane protein 1 |
| chr11_-_76134513 | 6.71 |
ENSMUST00000017430.12
|
Glod4
|
glyoxalase domain containing 4 |
| chr1_+_75145275 | 6.60 |
ENSMUST00000162768.8
ENSMUST00000160439.8 ENSMUST00000027394.12 |
Zfand2b
|
zinc finger, AN1 type domain 2B |
| chr19_-_61216834 | 6.57 |
ENSMUST00000076046.7
|
Csf2ra
|
colony stimulating factor 2 receptor, alpha, low-affinity (granulocyte-macrophage) |
| chr11_+_93935066 | 6.56 |
ENSMUST00000103168.10
|
Spag9
|
sperm associated antigen 9 |
| chr15_-_77129706 | 6.56 |
ENSMUST00000228361.2
|
Rbfox2
|
RNA binding protein, fox-1 homolog (C. elegans) 2 |
| chr15_+_12321502 | 6.56 |
ENSMUST00000059680.7
|
Golph3
|
golgi phosphoprotein 3 |
| chr4_+_130001349 | 6.56 |
ENSMUST00000030563.6
|
Pef1
|
penta-EF hand domain containing 1 |
| chr5_+_33176160 | 6.55 |
ENSMUST00000019109.8
|
Ywhah
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, eta polypeptide |
| chr5_+_91287448 | 6.54 |
ENSMUST00000031325.6
|
Areg
|
amphiregulin |
| chr16_-_4607751 | 6.53 |
ENSMUST00000117713.8
|
Cdip1
|
cell death inducing Trp53 target 1 |
| chr17_-_35954573 | 6.46 |
ENSMUST00000095467.4
|
Mucl3
|
mucin like 3 |
| chr18_+_23937019 | 6.43 |
ENSMUST00000025127.5
|
Mapre2
|
microtubule-associated protein, RP/EB family, member 2 |
| chr11_+_93935021 | 6.38 |
ENSMUST00000075695.13
ENSMUST00000092777.11 |
Spag9
|
sperm associated antigen 9 |
| chr16_-_44153288 | 6.31 |
ENSMUST00000136381.8
|
Sidt1
|
SID1 transmembrane family, member 1 |
| chr11_-_70578905 | 6.24 |
ENSMUST00000108544.8
|
Camta2
|
calmodulin binding transcription activator 2 |
| chr5_-_97259302 | 6.23 |
ENSMUST00000196078.5
|
Paqr3
|
progestin and adipoQ receptor family member III |
| chr7_+_6733561 | 6.22 |
ENSMUST00000200535.6
|
Usp29
|
ubiquitin specific peptidase 29 |
| chr11_-_76134436 | 6.19 |
ENSMUST00000164022.8
ENSMUST00000168055.2 ENSMUST00000169701.8 |
Glod4
|
glyoxalase domain containing 4 |
| chr2_-_173118315 | 6.19 |
ENSMUST00000036248.13
|
Pmepa1
|
prostate transmembrane protein, androgen induced 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 9.3 | 37.3 | GO:0061534 | gamma-aminobutyric acid secretion, neurotransmission(GO:0061534) |
| 6.0 | 18.1 | GO:0030070 | insulin processing(GO:0030070) |
| 5.7 | 11.3 | GO:1990927 | calcium ion regulated lysosome exocytosis(GO:1990927) |
| 5.5 | 22.0 | GO:0045053 | protein retention in Golgi apparatus(GO:0045053) |
| 5.2 | 31.0 | GO:0009235 | cobalamin metabolic process(GO:0009235) |
| 5.1 | 15.4 | GO:1990926 | short-term synaptic potentiation(GO:1990926) |
| 4.0 | 64.6 | GO:0043562 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 4.0 | 12.1 | GO:0097156 | fasciculation of motor neuron axon(GO:0097156) |
| 3.7 | 18.7 | GO:0034164 | negative regulation of toll-like receptor 9 signaling pathway(GO:0034164) |
| 3.7 | 36.5 | GO:1902412 | regulation of mitotic cytokinesis(GO:1902412) |
| 3.4 | 27.3 | GO:0090074 | negative regulation of protein homodimerization activity(GO:0090074) |
| 3.3 | 13.2 | GO:0098963 | dendritic transport of ribonucleoprotein complex(GO:0098961) dendritic transport of messenger ribonucleoprotein complex(GO:0098963) anterograde dendritic transport of messenger ribonucleoprotein complex(GO:0098964) |
| 3.2 | 9.6 | GO:2000850 | negative regulation of corticosteroid hormone secretion(GO:2000847) negative regulation of glucocorticoid secretion(GO:2000850) |
| 3.2 | 12.7 | GO:0019287 | isopentenyl diphosphate biosynthetic process, mevalonate pathway(GO:0019287) |
| 3.1 | 9.4 | GO:1904633 | glomerular visceral epithelial cell apoptotic process(GO:1903210) regulation of glomerular visceral epithelial cell apoptotic process(GO:1904633) positive regulation of glomerular visceral epithelial cell apoptotic process(GO:1904635) positive regulation of progesterone biosynthetic process(GO:2000184) |
| 3.0 | 12.0 | GO:0033566 | gamma-tubulin complex localization(GO:0033566) |
| 3.0 | 8.9 | GO:0015881 | creatine transport(GO:0015881) |
| 2.9 | 11.6 | GO:1901079 | positive regulation of relaxation of muscle(GO:1901079) regulation of dense core granule biogenesis(GO:2000705) |
| 2.9 | 17.1 | GO:0060702 | negative regulation of ribonuclease activity(GO:0060701) negative regulation of endoribonuclease activity(GO:0060702) |
| 2.8 | 11.3 | GO:0021917 | pancreatic A cell development(GO:0003322) forebrain-midbrain boundary formation(GO:0021905) somatic motor neuron fate commitment(GO:0021917) regulation of transcription from RNA polymerase II promoter involved in somatic motor neuron fate commitment(GO:0021918) sensory neuron migration(GO:1904937) |
| 2.7 | 41.2 | GO:0048245 | eosinophil chemotaxis(GO:0048245) |
| 2.7 | 8.1 | GO:2000019 | negative regulation of male gonad development(GO:2000019) |
| 2.7 | 8.1 | GO:0048320 | axial mesoderm formation(GO:0048320) |
| 2.5 | 7.6 | GO:0008628 | hormone-mediated apoptotic signaling pathway(GO:0008628) |
| 2.5 | 10.2 | GO:0080120 | CAAX-box protein processing(GO:0071586) CAAX-box protein maturation(GO:0080120) |
| 2.5 | 49.5 | GO:0022027 | interkinetic nuclear migration(GO:0022027) |
| 2.4 | 7.1 | GO:0097401 | synaptic vesicle lumen acidification(GO:0097401) |
| 2.3 | 11.6 | GO:0045054 | constitutive secretory pathway(GO:0045054) |
| 2.3 | 18.5 | GO:0036265 | RNA (guanine-N7)-methylation(GO:0036265) |
| 2.3 | 11.3 | GO:1903774 | positive regulation of viral budding via host ESCRT complex(GO:1903774) |
| 2.2 | 8.9 | GO:1903237 | negative regulation of leukocyte tethering or rolling(GO:1903237) |
| 2.2 | 6.6 | GO:1902527 | positive regulation of protein monoubiquitination(GO:1902527) |
| 2.2 | 6.6 | GO:0006713 | glucocorticoid catabolic process(GO:0006713) |
| 2.1 | 12.8 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 2.0 | 6.1 | GO:0097052 | L-kynurenine metabolic process(GO:0097052) |
| 2.0 | 8.0 | GO:0038016 | insulin receptor internalization(GO:0038016) |
| 2.0 | 19.6 | GO:0015670 | carbon dioxide transport(GO:0015670) |
| 1.9 | 48.4 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 1.9 | 5.7 | GO:0071288 | cellular response to mercury ion(GO:0071288) |
| 1.9 | 37.7 | GO:1902459 | positive regulation of stem cell population maintenance(GO:1902459) |
| 1.8 | 5.5 | GO:0045226 | extracellular polysaccharide biosynthetic process(GO:0045226) extracellular polysaccharide metabolic process(GO:0046379) |
| 1.8 | 3.7 | GO:0051795 | positive regulation of catagen(GO:0051795) |
| 1.8 | 5.5 | GO:0000451 | rRNA 2'-O-methylation(GO:0000451) |
| 1.8 | 10.6 | GO:0010991 | negative regulation of SMAD protein complex assembly(GO:0010991) |
| 1.8 | 5.3 | GO:0021529 | spinal cord oligodendrocyte cell differentiation(GO:0021529) spinal cord oligodendrocyte cell fate specification(GO:0021530) |
| 1.7 | 18.7 | GO:0033227 | dsRNA transport(GO:0033227) |
| 1.6 | 3.3 | GO:0003331 | regulation of extracellular matrix constituent secretion(GO:0003330) positive regulation of extracellular matrix constituent secretion(GO:0003331) |
| 1.6 | 6.5 | GO:0060598 | dichotomous subdivision of terminal units involved in mammary gland duct morphogenesis(GO:0060598) |
| 1.6 | 8.0 | GO:0097309 | cap1 mRNA methylation(GO:0097309) |
| 1.6 | 16.0 | GO:1990416 | cellular response to brain-derived neurotrophic factor stimulus(GO:1990416) |
| 1.6 | 7.9 | GO:0090467 | regulation of amino acid import(GO:0010958) L-arginine import(GO:0043091) arginine import(GO:0090467) |
| 1.5 | 12.3 | GO:0003376 | sphingosine-1-phosphate signaling pathway(GO:0003376) |
| 1.5 | 23.8 | GO:0090091 | positive regulation of extracellular matrix disassembly(GO:0090091) |
| 1.5 | 8.9 | GO:0051365 | cellular response to potassium ion starvation(GO:0051365) |
| 1.4 | 5.5 | GO:0002879 | positive regulation of acute inflammatory response to non-antigenic stimulus(GO:0002879) |
| 1.4 | 10.9 | GO:0070475 | rRNA base methylation(GO:0070475) |
| 1.4 | 29.8 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 1.3 | 4.0 | GO:2000686 | regulation of rubidium ion transmembrane transporter activity(GO:2000686) |
| 1.3 | 8.1 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 1.3 | 9.2 | GO:0015842 | aminergic neurotransmitter loading into synaptic vesicle(GO:0015842) |
| 1.3 | 30.1 | GO:0002091 | negative regulation of receptor internalization(GO:0002091) |
| 1.3 | 1.3 | GO:0061386 | closure of optic fissure(GO:0061386) |
| 1.3 | 3.9 | GO:0048597 | post-embryonic camera-type eye morphogenesis(GO:0048597) |
| 1.3 | 3.9 | GO:2000328 | regulation of T-helper 17 cell lineage commitment(GO:2000328) |
| 1.2 | 14.8 | GO:0060373 | regulation of ventricular cardiac muscle cell membrane depolarization(GO:0060373) |
| 1.2 | 7.4 | GO:0099590 | neurotransmitter receptor internalization(GO:0099590) |
| 1.2 | 3.7 | GO:0021629 | olfactory nerve morphogenesis(GO:0021627) olfactory nerve structural organization(GO:0021629) |
| 1.2 | 3.6 | GO:1903660 | transforming growth factor beta activation(GO:0036363) complement-dependent cytotoxicity(GO:0097278) regulation of complement-dependent cytotoxicity(GO:1903659) negative regulation of complement-dependent cytotoxicity(GO:1903660) |
| 1.2 | 7.1 | GO:0048294 | negative regulation of isotype switching to IgE isotypes(GO:0048294) |
| 1.2 | 4.7 | GO:1902219 | negative regulation of intrinsic apoptotic signaling pathway in response to osmotic stress(GO:1902219) |
| 1.2 | 3.5 | GO:2000503 | positive regulation of natural killer cell chemotaxis(GO:2000503) |
| 1.2 | 7.0 | GO:0006384 | transcription initiation from RNA polymerase III promoter(GO:0006384) |
| 1.2 | 3.5 | GO:0019918 | peptidyl-arginine methylation, to symmetrical-dimethyl arginine(GO:0019918) |
| 1.2 | 5.8 | GO:0098957 | anterograde axonal transport of mitochondrion(GO:0098957) |
| 1.1 | 14.9 | GO:1902474 | positive regulation of protein localization to synapse(GO:1902474) |
| 1.1 | 8.0 | GO:0051581 | negative regulation of neurotransmitter uptake(GO:0051581) negative regulation of serotonin uptake(GO:0051612) |
| 1.1 | 4.6 | GO:0001306 | age-dependent response to oxidative stress(GO:0001306) age-dependent general metabolic decline(GO:0007571) |
| 1.1 | 29.7 | GO:0007614 | short-term memory(GO:0007614) |
| 1.1 | 5.6 | GO:0060279 | positive regulation of ovulation(GO:0060279) |
| 1.1 | 3.4 | GO:2000118 | regulation of sodium-dependent phosphate transport(GO:2000118) |
| 1.1 | 8.9 | GO:0031038 | myosin II filament organization(GO:0031038) regulation of myosin II filament organization(GO:0043519) |
| 1.1 | 5.4 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 1.0 | 13.5 | GO:0010724 | regulation of definitive erythrocyte differentiation(GO:0010724) |
| 1.0 | 15.2 | GO:0048672 | positive regulation of collateral sprouting(GO:0048672) |
| 1.0 | 3.0 | GO:0051030 | snRNA transport(GO:0051030) |
| 1.0 | 20.1 | GO:0031268 | pseudopodium organization(GO:0031268) |
| 1.0 | 5.9 | GO:0070164 | negative regulation of adiponectin secretion(GO:0070164) |
| 1.0 | 7.9 | GO:0018243 | protein O-linked glycosylation via threonine(GO:0018243) |
| 1.0 | 3.9 | GO:0090063 | positive regulation of microtubule nucleation(GO:0090063) |
| 1.0 | 6.7 | GO:0061084 | negative regulation of protein refolding(GO:0061084) |
| 0.9 | 2.8 | GO:1901856 | negative regulation of cellular respiration(GO:1901856) |
| 0.9 | 6.6 | GO:0006616 | SRP-dependent cotranslational protein targeting to membrane, translocation(GO:0006616) |
| 0.9 | 15.9 | GO:0097011 | cellular response to granulocyte macrophage colony-stimulating factor stimulus(GO:0097011) |
| 0.9 | 7.4 | GO:0000042 | protein targeting to Golgi(GO:0000042) |
| 0.9 | 4.6 | GO:1902775 | mitochondrial large ribosomal subunit assembly(GO:1902775) |
| 0.9 | 14.7 | GO:0030432 | peristalsis(GO:0030432) |
| 0.9 | 5.4 | GO:0035865 | response to potassium ion(GO:0035864) cellular response to potassium ion(GO:0035865) |
| 0.9 | 15.2 | GO:0061000 | negative regulation of dendritic spine development(GO:0061000) |
| 0.9 | 2.7 | GO:1900369 | negative regulation of RNA interference(GO:1900369) |
| 0.9 | 7.9 | GO:0051835 | positive regulation of synapse structural plasticity(GO:0051835) |
| 0.9 | 21.5 | GO:0001553 | luteinization(GO:0001553) |
| 0.9 | 4.3 | GO:0008616 | queuosine biosynthetic process(GO:0008616) queuosine metabolic process(GO:0046116) |
| 0.9 | 10.3 | GO:0008635 | mitochondrial electron transport, ubiquinol to cytochrome c(GO:0006122) activation of cysteine-type endopeptidase activity involved in apoptotic process by cytochrome c(GO:0008635) |
| 0.8 | 13.3 | GO:0051152 | positive regulation of smooth muscle cell differentiation(GO:0051152) |
| 0.8 | 2.5 | GO:0015819 | lysine transport(GO:0015819) |
| 0.8 | 8.2 | GO:0035754 | B cell chemotaxis(GO:0035754) |
| 0.8 | 1.6 | GO:2000286 | receptor internalization involved in canonical Wnt signaling pathway(GO:2000286) |
| 0.8 | 16.3 | GO:0034067 | protein localization to Golgi apparatus(GO:0034067) |
| 0.8 | 5.4 | GO:1901029 | negative regulation of mitochondrial outer membrane permeabilization involved in apoptotic signaling pathway(GO:1901029) |
| 0.7 | 11.1 | GO:0048680 | positive regulation of axon regeneration(GO:0048680) |
| 0.7 | 6.5 | GO:0042795 | snRNA transcription from RNA polymerase II promoter(GO:0042795) |
| 0.7 | 3.5 | GO:1903071 | positive regulation of ER-associated ubiquitin-dependent protein catabolic process(GO:1903071) |
| 0.7 | 2.1 | GO:0021764 | amygdala development(GO:0021764) |
| 0.7 | 13.0 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.7 | 11.6 | GO:0043084 | penile erection(GO:0043084) |
| 0.7 | 2.7 | GO:1900864 | mitochondrial tRNA modification(GO:0070900) mitochondrial RNA modification(GO:1900864) |
| 0.7 | 5.4 | GO:1901750 | peptide modification(GO:0031179) leukotriene D4 metabolic process(GO:1901748) leukotriene D4 biosynthetic process(GO:1901750) |
| 0.7 | 10.6 | GO:0090315 | negative regulation of protein targeting to membrane(GO:0090315) |
| 0.6 | 3.2 | GO:0090669 | telomerase RNA stabilization(GO:0090669) |
| 0.6 | 10.3 | GO:0046069 | cGMP catabolic process(GO:0046069) |
| 0.6 | 13.6 | GO:0010763 | positive regulation of fibroblast migration(GO:0010763) |
| 0.6 | 4.1 | GO:0014053 | negative regulation of gamma-aminobutyric acid secretion(GO:0014053) negative regulation of dopamine secretion(GO:0033602) |
| 0.6 | 4.7 | GO:0009644 | response to high light intensity(GO:0009644) |
| 0.6 | 8.7 | GO:0061299 | retina vasculature morphogenesis in camera-type eye(GO:0061299) |
| 0.6 | 5.2 | GO:0086028 | bundle of His cell to Purkinje myocyte signaling(GO:0086028) bundle of His cell action potential(GO:0086043) |
| 0.6 | 8.7 | GO:1904262 | negative regulation of TORC1 signaling(GO:1904262) |
| 0.6 | 5.2 | GO:0033353 | S-adenosylmethionine cycle(GO:0033353) epithelial fluid transport(GO:0042045) |
| 0.6 | 11.5 | GO:0090073 | positive regulation of protein homodimerization activity(GO:0090073) |
| 0.6 | 2.8 | GO:0072368 | regulation of lipid transport by negative regulation of transcription from RNA polymerase II promoter(GO:0072368) |
| 0.6 | 3.9 | GO:0050847 | progesterone receptor signaling pathway(GO:0050847) |
| 0.6 | 2.2 | GO:0002949 | tRNA threonylcarbamoyladenosine modification(GO:0002949) |
| 0.6 | 8.8 | GO:0043983 | histone H4-K12 acetylation(GO:0043983) |
| 0.5 | 2.2 | GO:0022007 | neural plate elongation(GO:0014022) convergent extension involved in neural plate elongation(GO:0022007) |
| 0.5 | 5.9 | GO:0070863 | positive regulation of protein exit from endoplasmic reticulum(GO:0070863) |
| 0.5 | 4.2 | GO:0032790 | ribosome disassembly(GO:0032790) |
| 0.5 | 1.6 | GO:0045014 | carbon catabolite repression of transcription(GO:0045013) negative regulation of transcription by glucose(GO:0045014) |
| 0.5 | 1.6 | GO:0046462 | monoacylglycerol metabolic process(GO:0046462) |
| 0.5 | 7.3 | GO:0007042 | lysosomal lumen acidification(GO:0007042) |
| 0.5 | 3.9 | GO:0046543 | development of secondary female sexual characteristics(GO:0046543) |
| 0.5 | 6.8 | GO:0002329 | pre-B cell differentiation(GO:0002329) |
| 0.5 | 6.8 | GO:0006824 | cobalt ion transport(GO:0006824) |
| 0.5 | 11.5 | GO:0034389 | lipid particle organization(GO:0034389) |
| 0.5 | 5.2 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 0.5 | 4.3 | GO:0006776 | vitamin A metabolic process(GO:0006776) |
| 0.5 | 6.9 | GO:1901096 | regulation of autophagosome maturation(GO:1901096) |
| 0.5 | 5.0 | GO:0070940 | dephosphorylation of RNA polymerase II C-terminal domain(GO:0070940) |
| 0.4 | 3.5 | GO:2000766 | negative regulation of cytoplasmic translation(GO:2000766) |
| 0.4 | 5.5 | GO:1902004 | positive regulation of beta-amyloid formation(GO:1902004) |
| 0.4 | 51.1 | GO:0031110 | regulation of microtubule polymerization or depolymerization(GO:0031110) |
| 0.4 | 3.3 | GO:0019660 | glucose catabolic process to lactate(GO:0019659) glycolytic fermentation(GO:0019660) glucose catabolic process to lactate via pyruvate(GO:0019661) |
| 0.4 | 3.3 | GO:0042985 | negative regulation of amyloid precursor protein biosynthetic process(GO:0042985) |
| 0.4 | 2.8 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 0.4 | 22.7 | GO:0045806 | negative regulation of endocytosis(GO:0045806) |
| 0.4 | 9.8 | GO:0071480 | cellular response to gamma radiation(GO:0071480) |
| 0.4 | 5.8 | GO:0071243 | cellular response to arsenic-containing substance(GO:0071243) |
| 0.4 | 1.9 | GO:1901026 | ripoptosome assembly(GO:0097343) ripoptosome assembly involved in necroptotic process(GO:1901026) |
| 0.4 | 1.9 | GO:0046167 | glycerol-3-phosphate biosynthetic process(GO:0046167) |
| 0.4 | 3.4 | GO:0021960 | anterior commissure morphogenesis(GO:0021960) |
| 0.4 | 1.9 | GO:0097167 | circadian regulation of translation(GO:0097167) negative regulation of glucocorticoid receptor signaling pathway(GO:2000323) |
| 0.4 | 3.0 | GO:0033299 | secretion of lysosomal enzymes(GO:0033299) |
| 0.4 | 4.7 | GO:0022615 | protein to membrane docking(GO:0022615) |
| 0.4 | 43.8 | GO:0006888 | ER to Golgi vesicle-mediated transport(GO:0006888) |
| 0.4 | 5.4 | GO:1902231 | positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage(GO:1902231) |
| 0.4 | 3.2 | GO:0003406 | retinal pigment epithelium development(GO:0003406) |
| 0.4 | 8.7 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.4 | 5.3 | GO:0030049 | muscle filament sliding(GO:0030049) |
| 0.4 | 5.3 | GO:0060501 | positive regulation of epithelial cell proliferation involved in lung morphogenesis(GO:0060501) |
| 0.4 | 16.5 | GO:2000279 | negative regulation of DNA biosynthetic process(GO:2000279) |
| 0.3 | 3.5 | GO:0060149 | negative regulation of posttranscriptional gene silencing(GO:0060149) negative regulation of gene silencing by miRNA(GO:0060965) negative regulation of gene silencing by RNA(GO:0060967) |
| 0.3 | 14.6 | GO:0003299 | muscle hypertrophy in response to stress(GO:0003299) cardiac muscle adaptation(GO:0014887) cardiac muscle hypertrophy in response to stress(GO:0014898) |
| 0.3 | 2.1 | GO:0006450 | regulation of translational fidelity(GO:0006450) |
| 0.3 | 12.1 | GO:0035329 | hippo signaling(GO:0035329) |
| 0.3 | 10.7 | GO:0030575 | nuclear body organization(GO:0030575) |
| 0.3 | 4.1 | GO:0034982 | mitochondrial protein processing(GO:0034982) |
| 0.3 | 5.8 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.3 | 3.6 | GO:0032463 | negative regulation of protein homooligomerization(GO:0032463) |
| 0.3 | 34.9 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.3 | 5.1 | GO:0046007 | negative regulation of activated T cell proliferation(GO:0046007) |
| 0.3 | 20.8 | GO:0042771 | intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator(GO:0042771) |
| 0.3 | 2.8 | GO:0032471 | negative regulation of endoplasmic reticulum calcium ion concentration(GO:0032471) |
| 0.3 | 1.9 | GO:0070895 | transposon integration(GO:0070893) regulation of transposon integration(GO:0070894) negative regulation of transposon integration(GO:0070895) |
| 0.3 | 1.5 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.3 | 1.9 | GO:0071918 | urea transmembrane transport(GO:0071918) |
| 0.3 | 3.2 | GO:0016558 | protein import into peroxisome matrix(GO:0016558) |
| 0.3 | 2.9 | GO:0018231 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.3 | 2.9 | GO:0032287 | peripheral nervous system myelin maintenance(GO:0032287) |
| 0.3 | 2.0 | GO:2000767 | regulation of cytoplasmic translation(GO:2000765) positive regulation of cytoplasmic translation(GO:2000767) |
| 0.3 | 13.3 | GO:0010107 | potassium ion import(GO:0010107) |
| 0.3 | 8.7 | GO:0015985 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.3 | 0.8 | GO:0000349 | generation of catalytic spliceosome for first transesterification step(GO:0000349) |
| 0.3 | 11.1 | GO:0097031 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.3 | 3.9 | GO:0030828 | positive regulation of cGMP biosynthetic process(GO:0030828) |
| 0.3 | 3.3 | GO:1901078 | negative regulation of relaxation of muscle(GO:1901078) negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.3 | 4.8 | GO:0071625 | vocalization behavior(GO:0071625) |
| 0.2 | 11.2 | GO:0045746 | negative regulation of Notch signaling pathway(GO:0045746) |
| 0.2 | 2.0 | GO:1903026 | negative regulation of RNA polymerase II regulatory region sequence-specific DNA binding(GO:1903026) |
| 0.2 | 2.9 | GO:0034625 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.2 | 4.4 | GO:0021542 | dentate gyrus development(GO:0021542) |
| 0.2 | 3.5 | GO:0033617 | mitochondrial respiratory chain complex IV assembly(GO:0033617) |
| 0.2 | 16.9 | GO:0072583 | clathrin-mediated endocytosis(GO:0072583) |
| 0.2 | 3.6 | GO:0048733 | sebaceous gland development(GO:0048733) |
| 0.2 | 9.0 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.2 | 8.1 | GO:0008299 | isoprenoid biosynthetic process(GO:0008299) |
| 0.2 | 13.9 | GO:0071277 | cellular response to calcium ion(GO:0071277) |
| 0.2 | 11.9 | GO:0051926 | negative regulation of calcium ion transport(GO:0051926) |
| 0.2 | 4.1 | GO:0010832 | negative regulation of myotube differentiation(GO:0010832) |
| 0.2 | 2.6 | GO:0072422 | signal transduction involved in cell cycle checkpoint(GO:0072395) signal transduction involved in DNA integrity checkpoint(GO:0072401) signal transduction involved in DNA damage checkpoint(GO:0072422) |
| 0.2 | 2.8 | GO:0006622 | protein targeting to lysosome(GO:0006622) |
| 0.2 | 2.1 | GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0000083) |
| 0.2 | 2.3 | GO:0043278 | response to isoquinoline alkaloid(GO:0014072) response to morphine(GO:0043278) |
| 0.2 | 2.5 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.2 | 4.5 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade(GO:0070886) |
| 0.2 | 0.8 | GO:0044208 | 'de novo' AMP biosynthetic process(GO:0044208) |
| 0.2 | 5.2 | GO:0031643 | positive regulation of myelination(GO:0031643) |
| 0.2 | 14.1 | GO:1902476 | chloride transmembrane transport(GO:1902476) |
| 0.2 | 8.0 | GO:0045197 | establishment or maintenance of epithelial cell apical/basal polarity(GO:0045197) |
| 0.2 | 8.6 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.2 | 8.0 | GO:0031648 | protein destabilization(GO:0031648) |
| 0.2 | 3.5 | GO:0006107 | oxaloacetate metabolic process(GO:0006107) |
| 0.2 | 2.0 | GO:0000022 | mitotic spindle elongation(GO:0000022) mitotic spindle midzone assembly(GO:0051256) |
| 0.2 | 7.8 | GO:0060351 | cartilage development involved in endochondral bone morphogenesis(GO:0060351) |
| 0.2 | 1.8 | GO:0034058 | endosomal vesicle fusion(GO:0034058) |
| 0.2 | 5.5 | GO:0070918 | dsRNA fragmentation(GO:0031050) production of miRNAs involved in gene silencing by miRNA(GO:0035196) production of small RNA involved in gene silencing by RNA(GO:0070918) |
| 0.2 | 2.2 | GO:0043649 | dicarboxylic acid catabolic process(GO:0043649) |
| 0.2 | 3.7 | GO:0060441 | epithelial tube branching involved in lung morphogenesis(GO:0060441) |
| 0.2 | 16.7 | GO:0035914 | skeletal muscle cell differentiation(GO:0035914) |
| 0.2 | 15.1 | GO:0032411 | positive regulation of transporter activity(GO:0032411) |
| 0.2 | 1.4 | GO:0048671 | negative regulation of collateral sprouting(GO:0048671) |
| 0.2 | 0.6 | GO:1901894 | regulation of calcium-transporting ATPase activity(GO:1901894) |
| 0.2 | 0.8 | GO:0046341 | CDP-diacylglycerol metabolic process(GO:0046341) |
| 0.1 | 3.1 | GO:0003094 | glomerular filtration(GO:0003094) |
| 0.1 | 9.9 | GO:0034394 | protein localization to cell surface(GO:0034394) |
| 0.1 | 1.8 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.1 | 5.8 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.1 | 9.6 | GO:0008542 | visual learning(GO:0008542) |
| 0.1 | 4.7 | GO:0010761 | fibroblast migration(GO:0010761) |
| 0.1 | 1.7 | GO:0021520 | spinal cord motor neuron cell fate specification(GO:0021520) |
| 0.1 | 11.6 | GO:0007030 | Golgi organization(GO:0007030) |
| 0.1 | 20.0 | GO:0006275 | regulation of DNA replication(GO:0006275) |
| 0.1 | 3.6 | GO:0034766 | negative regulation of ion transmembrane transport(GO:0034766) |
| 0.1 | 4.2 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.1 | 14.1 | GO:0071805 | potassium ion transmembrane transport(GO:0071805) |
| 0.1 | 1.1 | GO:0030206 | chondroitin sulfate biosynthetic process(GO:0030206) |
| 0.1 | 14.4 | GO:0008645 | hexose transport(GO:0008645) glucose transport(GO:0015758) |
| 0.1 | 1.2 | GO:0021942 | radial glia guided migration of Purkinje cell(GO:0021942) |
| 0.1 | 2.2 | GO:0002053 | positive regulation of mesenchymal cell proliferation(GO:0002053) |
| 0.1 | 2.9 | GO:0031424 | keratinization(GO:0031424) |
| 0.1 | 1.7 | GO:0061436 | establishment of skin barrier(GO:0061436) |
| 0.1 | 3.8 | GO:0016601 | Rac protein signal transduction(GO:0016601) |
| 0.1 | 1.5 | GO:0071470 | myeloid dendritic cell differentiation(GO:0043011) cellular response to osmotic stress(GO:0071470) |
| 0.1 | 0.2 | GO:0033140 | negative regulation of peptidyl-serine phosphorylation of STAT protein(GO:0033140) |
| 0.1 | 2.3 | GO:0060716 | labyrinthine layer blood vessel development(GO:0060716) |
| 0.1 | 1.3 | GO:0034204 | lipid translocation(GO:0034204) |
| 0.1 | 0.9 | GO:0060253 | negative regulation of glial cell proliferation(GO:0060253) |
| 0.1 | 2.3 | GO:0045109 | intermediate filament organization(GO:0045109) |
| 0.1 | 2.4 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.1 | 3.6 | GO:0030521 | androgen receptor signaling pathway(GO:0030521) |
| 0.1 | 0.5 | GO:0051012 | microtubule sliding(GO:0051012) |
| 0.1 | 6.4 | GO:0003073 | regulation of systemic arterial blood pressure(GO:0003073) |
| 0.1 | 3.7 | GO:0051489 | regulation of filopodium assembly(GO:0051489) |
| 0.1 | 6.1 | GO:0043488 | regulation of mRNA stability(GO:0043488) |
| 0.1 | 2.0 | GO:0040020 | regulation of meiotic nuclear division(GO:0040020) |
| 0.1 | 0.1 | GO:1902035 | positive regulation of hematopoietic stem cell proliferation(GO:1902035) |
| 0.1 | 1.1 | GO:0080182 | histone H3-K4 trimethylation(GO:0080182) |
| 0.1 | 3.5 | GO:0060612 | adipose tissue development(GO:0060612) |
| 0.1 | 4.9 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.1 | 4.3 | GO:0090263 | positive regulation of canonical Wnt signaling pathway(GO:0090263) |
| 0.1 | 0.7 | GO:0043923 | positive regulation by host of viral transcription(GO:0043923) |
| 0.1 | 0.3 | GO:0071635 | negative regulation of transforming growth factor beta production(GO:0071635) |
| 0.1 | 4.2 | GO:0070527 | platelet aggregation(GO:0070527) |
| 0.1 | 0.5 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.1 | 10.9 | GO:0002244 | hematopoietic progenitor cell differentiation(GO:0002244) |
| 0.1 | 1.8 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.1 | 0.7 | GO:1904778 | regulation of protein localization to cell cortex(GO:1904776) positive regulation of protein localization to cell cortex(GO:1904778) |
| 0.1 | 2.8 | GO:0046626 | regulation of insulin receptor signaling pathway(GO:0046626) |
| 0.1 | 20.3 | GO:0007015 | actin filament organization(GO:0007015) |
| 0.1 | 5.2 | GO:0000187 | activation of MAPK activity(GO:0000187) |
| 0.1 | 1.5 | GO:2000134 | negative regulation of G1/S transition of mitotic cell cycle(GO:2000134) |
| 0.1 | 5.5 | GO:0008277 | regulation of G-protein coupled receptor protein signaling pathway(GO:0008277) |
| 0.1 | 5.4 | GO:0010811 | positive regulation of cell-substrate adhesion(GO:0010811) |
| 0.1 | 3.3 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 0.5 | GO:0031629 | synaptic vesicle fusion to presynaptic active zone membrane(GO:0031629) vesicle fusion to plasma membrane(GO:0099500) |
| 0.0 | 3.9 | GO:0048706 | embryonic skeletal system development(GO:0048706) |
| 0.0 | 5.8 | GO:0043123 | positive regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043123) |
| 0.0 | 2.4 | GO:0045666 | positive regulation of neuron differentiation(GO:0045666) |
| 0.0 | 1.9 | GO:0006695 | cholesterol biosynthetic process(GO:0006695) |
| 0.0 | 8.3 | GO:0008202 | steroid metabolic process(GO:0008202) |
| 0.0 | 0.7 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.0 | 0.6 | GO:0010818 | T cell chemotaxis(GO:0010818) |
| 0.0 | 1.2 | GO:0032012 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.0 | 10.8 | GO:0006412 | translation(GO:0006412) |
| 0.0 | 1.1 | GO:0045773 | positive regulation of axon extension(GO:0045773) |
| 0.0 | 0.7 | GO:0071577 | zinc II ion transmembrane transport(GO:0071577) |
| 0.0 | 2.0 | GO:0030307 | positive regulation of cell growth(GO:0030307) |
| 0.0 | 7.9 | GO:0051260 | protein homooligomerization(GO:0051260) |
| 0.0 | 1.9 | GO:0000045 | autophagosome assembly(GO:0000045) |
| 0.0 | 1.7 | GO:0006687 | glycosphingolipid metabolic process(GO:0006687) |
| 0.0 | 0.2 | GO:0070166 | enamel mineralization(GO:0070166) |
| 0.0 | 1.1 | GO:0000184 | nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:0000184) |
| 0.0 | 1.4 | GO:0009166 | nucleotide catabolic process(GO:0009166) |
| 0.0 | 3.7 | GO:0016358 | dendrite development(GO:0016358) |
| 0.0 | 21.0 | GO:0055114 | oxidation-reduction process(GO:0055114) |
| 0.0 | 1.2 | GO:0032543 | mitochondrial translation(GO:0032543) |
| 0.0 | 0.5 | GO:0070306 | lens fiber cell differentiation(GO:0070306) |
| 0.0 | 1.1 | GO:0031338 | regulation of vesicle fusion(GO:0031338) |
| 0.0 | 1.2 | GO:0007601 | visual perception(GO:0007601) sensory perception of light stimulus(GO:0050953) |
| 0.0 | 0.7 | GO:0030512 | negative regulation of transforming growth factor beta receptor signaling pathway(GO:0030512) negative regulation of cellular response to transforming growth factor beta stimulus(GO:1903845) |
| 0.0 | 3.1 | GO:0051090 | regulation of sequence-specific DNA binding transcription factor activity(GO:0051090) |
| 0.0 | 27.1 | GO:0007186 | G-protein coupled receptor signaling pathway(GO:0007186) |
| 0.0 | 1.0 | GO:0019221 | cytokine-mediated signaling pathway(GO:0019221) |
| 0.0 | 2.9 | GO:0006869 | lipid transport(GO:0006869) |
| 0.0 | 0.2 | GO:0030033 | microvillus assembly(GO:0030033) |
| 0.0 | 0.3 | GO:0060135 | maternal process involved in female pregnancy(GO:0060135) |
| 0.0 | 1.7 | GO:0006874 | cellular calcium ion homeostasis(GO:0006874) |
| 0.0 | 3.2 | GO:0030334 | regulation of cell migration(GO:0030334) |
| 0.0 | 1.0 | GO:0007596 | blood coagulation(GO:0007596) |
| 0.0 | 2.9 | GO:0042254 | ribosome biogenesis(GO:0042254) |
| 0.0 | 6.4 | GO:0048666 | neuron development(GO:0048666) |
| 0.0 | 0.7 | GO:0007368 | determination of left/right symmetry(GO:0007368) |
| 0.0 | 10.8 | GO:0006952 | defense response(GO:0006952) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 9.4 | 37.7 | GO:0071148 | TEAD-1-YAP complex(GO:0071148) |
| 7.2 | 21.6 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 4.1 | 12.4 | GO:1903754 | cortical microtubule plus-end(GO:1903754) cytoplasmic microtubule plus-end(GO:1904511) |
| 4.1 | 20.3 | GO:0098560 | cytoplasmic side of late endosome membrane(GO:0098560) |
| 3.5 | 20.8 | GO:0043527 | tRNA methyltransferase complex(GO:0043527) |
| 3.4 | 10.3 | GO:0034274 | Atg12-Atg5-Atg16 complex(GO:0034274) |
| 3.3 | 36.5 | GO:0000235 | astral microtubule(GO:0000235) |
| 3.1 | 9.4 | GO:1990844 | subsarcolemmal mitochondrion(GO:1990843) interfibrillar mitochondrion(GO:1990844) |
| 3.1 | 15.4 | GO:0070033 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin I complex(GO:0070032) synaptobrevin 2-SNAP-25-syntaxin-1a-complexin II complex(GO:0070033) |
| 2.3 | 11.5 | GO:0044393 | microspike(GO:0044393) |
| 2.3 | 45.6 | GO:0031045 | dense core granule(GO:0031045) |
| 2.1 | 61.0 | GO:0000421 | autophagosome membrane(GO:0000421) |
| 1.9 | 11.6 | GO:0042583 | chromaffin granule(GO:0042583) |
| 1.9 | 41.5 | GO:0071437 | invadopodium(GO:0071437) |
| 1.6 | 23.8 | GO:0005869 | dynactin complex(GO:0005869) |
| 1.5 | 16.9 | GO:0030130 | clathrin coat of trans-Golgi network vesicle(GO:0030130) |
| 1.5 | 35.3 | GO:0030008 | TRAPP complex(GO:0030008) |
| 1.3 | 5.2 | GO:0046691 | intracellular canaliculus(GO:0046691) |
| 1.1 | 14.4 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 1.1 | 4.3 | GO:0005745 | m-AAA complex(GO:0005745) |
| 1.0 | 8.7 | GO:0045261 | mitochondrial proton-transporting ATP synthase complex, catalytic core F(1)(GO:0000275) proton-transporting ATP synthase complex, catalytic core F(1)(GO:0045261) |
| 0.9 | 7.3 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.9 | 4.5 | GO:0035976 | AP1 complex(GO:0035976) |
| 0.9 | 5.2 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.9 | 12.8 | GO:0005915 | zonula adherens(GO:0005915) |
| 0.9 | 4.3 | GO:0035363 | histone locus body(GO:0035363) |
| 0.8 | 4.1 | GO:0038039 | G-protein coupled receptor heterodimeric complex(GO:0038039) |
| 0.8 | 18.7 | GO:0030285 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.8 | 11.3 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.8 | 37.2 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.8 | 3.2 | GO:1990429 | Pex17p-Pex14p docking complex(GO:1990415) peroxisomal importomer complex(GO:1990429) |
| 0.8 | 4.7 | GO:0070820 | tertiary granule(GO:0070820) |
| 0.8 | 44.3 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.8 | 15.5 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.7 | 2.9 | GO:0097447 | dendritic tree(GO:0097447) |
| 0.7 | 3.6 | GO:0034751 | aryl hydrocarbon receptor complex(GO:0034751) |
| 0.7 | 26.1 | GO:0032839 | dendrite cytoplasm(GO:0032839) |
| 0.7 | 4.0 | GO:0005947 | mitochondrial alpha-ketoglutarate dehydrogenase complex(GO:0005947) |
| 0.6 | 4.5 | GO:0070381 | endosome to plasma membrane transport vesicle(GO:0070381) |
| 0.6 | 5.7 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.6 | 4.8 | GO:0045009 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.6 | 14.8 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.6 | 8.4 | GO:0044232 | organelle membrane contact site(GO:0044232) |
| 0.5 | 3.4 | GO:0098642 | collagen type IV trimer(GO:0005587) network-forming collagen trimer(GO:0098642) collagen network(GO:0098645) basement membrane collagen trimer(GO:0098651) |
| 0.5 | 24.3 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.5 | 2.8 | GO:0031313 | extrinsic component of endosome membrane(GO:0031313) |
| 0.4 | 3.6 | GO:0070847 | core mediator complex(GO:0070847) |
| 0.4 | 31.4 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.4 | 10.3 | GO:0043196 | varicosity(GO:0043196) |
| 0.4 | 2.7 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 0.4 | 4.1 | GO:0060053 | neurofilament cytoskeleton(GO:0060053) |
| 0.4 | 6.8 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 0.4 | 7.1 | GO:0042581 | specific granule(GO:0042581) |
| 0.4 | 7.4 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.4 | 12.0 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.4 | 3.1 | GO:0000322 | storage vacuole(GO:0000322) |
| 0.4 | 6.6 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.4 | 31.3 | GO:0031985 | Golgi cisterna(GO:0031985) |
| 0.4 | 8.9 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.4 | 2.8 | GO:0032937 | SREBP-SCAP-Insig complex(GO:0032937) |
| 0.3 | 21.0 | GO:0005776 | autophagosome(GO:0005776) |
| 0.3 | 5.1 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.3 | 20.6 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.3 | 1.8 | GO:0033263 | CORVET complex(GO:0033263) |
| 0.3 | 7.7 | GO:0045334 | clathrin-coated endocytic vesicle(GO:0045334) |
| 0.3 | 14.7 | GO:0005747 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.3 | 12.8 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.3 | 5.8 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.3 | 3.0 | GO:0031083 | BLOC-1 complex(GO:0031083) |
| 0.3 | 1.0 | GO:0031084 | BLOC-2 complex(GO:0031084) |
| 0.3 | 96.9 | GO:0045211 | postsynaptic membrane(GO:0045211) |
| 0.2 | 12.9 | GO:0031228 | intrinsic component of Golgi membrane(GO:0031228) |
| 0.2 | 33.0 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.2 | 4.2 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.2 | 1.7 | GO:0019907 | cyclin-dependent protein kinase activating kinase holoenzyme complex(GO:0019907) |
| 0.2 | 6.4 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.2 | 2.1 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.2 | 5.0 | GO:0016460 | myosin II complex(GO:0016460) |
| 0.2 | 3.1 | GO:0016461 | unconventional myosin complex(GO:0016461) |
| 0.2 | 1.2 | GO:0043224 | nuclear SCF ubiquitin ligase complex(GO:0043224) |
| 0.2 | 2.0 | GO:0030478 | actin cap(GO:0030478) |
| 0.2 | 2.2 | GO:0000408 | EKC/KEOPS complex(GO:0000408) |
| 0.2 | 22.3 | GO:0043679 | axon terminus(GO:0043679) |
| 0.2 | 3.4 | GO:0000974 | Prp19 complex(GO:0000974) |
| 0.2 | 6.1 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.2 | 2.3 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.2 | 2.2 | GO:1990909 | Wnt signalosome(GO:1990909) |
| 0.2 | 40.9 | GO:0005769 | early endosome(GO:0005769) |
| 0.2 | 20.1 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) |
| 0.2 | 3.6 | GO:0005852 | eukaryotic translation initiation factor 3 complex(GO:0005852) |
| 0.2 | 25.6 | GO:0043204 | perikaryon(GO:0043204) |
| 0.2 | 9.4 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.2 | 18.5 | GO:0036477 | somatodendritic compartment(GO:0036477) |
| 0.2 | 1.9 | GO:0097542 | ciliary tip(GO:0097542) |
| 0.1 | 7.8 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.1 | 13.1 | GO:0030018 | Z disc(GO:0030018) |
| 0.1 | 24.9 | GO:0005923 | bicellular tight junction(GO:0005923) |
| 0.1 | 11.1 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.1 | 2.0 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.1 | 7.4 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.1 | 5.9 | GO:0055037 | recycling endosome(GO:0055037) |
| 0.1 | 5.5 | GO:0014704 | intercalated disc(GO:0014704) |
| 0.1 | 1.1 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.1 | 12.7 | GO:0001725 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.1 | 1.5 | GO:0033256 | I-kappaB/NF-kappaB complex(GO:0033256) |
| 0.1 | 23.5 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.1 | 118.6 | GO:0043005 | neuron projection(GO:0043005) |
| 0.1 | 13.1 | GO:0016605 | PML body(GO:0016605) |
| 0.1 | 3.3 | GO:0030660 | Golgi-associated vesicle membrane(GO:0030660) |
| 0.1 | 11.0 | GO:0045177 | apical part of cell(GO:0045177) |
| 0.1 | 1.3 | GO:0008541 | proteasome regulatory particle, lid subcomplex(GO:0008541) |
| 0.1 | 10.9 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.1 | 3.5 | GO:0031305 | intrinsic component of mitochondrial inner membrane(GO:0031304) integral component of mitochondrial inner membrane(GO:0031305) |
| 0.1 | 1.3 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.1 | 1.8 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 0.1 | 4.6 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.1 | 46.2 | GO:0005768 | endosome(GO:0005768) |
| 0.1 | 21.5 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.1 | 3.5 | GO:0030864 | cortical actin cytoskeleton(GO:0030864) |
| 0.1 | 2.9 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.1 | 0.8 | GO:0008024 | cyclin/CDK positive transcription elongation factor complex(GO:0008024) |
| 0.1 | 9.0 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.1 | 13.7 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.1 | 21.8 | GO:0048471 | perinuclear region of cytoplasm(GO:0048471) |
| 0.1 | 0.6 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.1 | 15.6 | GO:0000323 | lytic vacuole(GO:0000323) lysosome(GO:0005764) |
| 0.1 | 3.5 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.1 | 93.9 | GO:0005739 | mitochondrion(GO:0005739) |
| 0.1 | 0.5 | GO:0070187 | telosome(GO:0070187) |
| 0.1 | 1.1 | GO:0031672 | A band(GO:0031672) |
| 0.1 | 1.6 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.0 | 0.3 | GO:0072357 | PTW/PP1 phosphatase complex(GO:0072357) |
| 0.0 | 256.3 | GO:0016021 | integral component of membrane(GO:0016021) |
| 0.0 | 0.7 | GO:0030863 | cortical cytoskeleton(GO:0030863) |
| 0.0 | 0.3 | GO:0070552 | BRISC complex(GO:0070552) |
| 0.0 | 9.2 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 3.9 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.0 | 4.4 | GO:0000781 | chromosome, telomeric region(GO:0000781) |
| 0.0 | 91.9 | GO:0005576 | extracellular region(GO:0005576) |
| 0.0 | 3.3 | GO:0000123 | histone acetyltransferase complex(GO:0000123) |
| 0.0 | 0.8 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.0 | 0.3 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.0 | 0.7 | GO:0030017 | sarcomere(GO:0030017) |
| 0.0 | 1.6 | GO:0030054 | cell junction(GO:0030054) |
| 0.0 | 11.0 | GO:0005886 | plasma membrane(GO:0005886) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.3 | 15.9 | GO:0008330 | protein tyrosine/threonine phosphatase activity(GO:0008330) |
| 5.3 | 15.9 | GO:0051916 | granulocyte colony-stimulating factor binding(GO:0051916) |
| 4.6 | 18.5 | GO:0008176 | tRNA (guanine-N7-)-methyltransferase activity(GO:0008176) |
| 3.4 | 10.3 | GO:0019776 | Atg8 ligase activity(GO:0019776) |
| 3.3 | 29.7 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 3.1 | 18.7 | GO:0051033 | nucleic acid transmembrane transporter activity(GO:0051032) RNA transmembrane transporter activity(GO:0051033) |
| 3.0 | 8.9 | GO:0005308 | creatine transmembrane transporter activity(GO:0005308) |
| 2.9 | 17.1 | GO:0060698 | endoribonuclease inhibitor activity(GO:0060698) |
| 2.7 | 8.1 | GO:0004421 | hydroxymethylglutaryl-CoA synthase activity(GO:0004421) |
| 2.6 | 10.6 | GO:0005093 | Rab GDP-dissociation inhibitor activity(GO:0005093) |
| 2.6 | 7.8 | GO:0052658 | inositol-1,4,5-trisphosphate 5-phosphatase activity(GO:0052658) |
| 2.5 | 27.7 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 2.5 | 14.7 | GO:0004994 | somatostatin receptor activity(GO:0004994) |
| 2.4 | 9.8 | GO:0008802 | betaine-aldehyde dehydrogenase activity(GO:0008802) |
| 2.2 | 8.9 | GO:0031735 | CCR10 chemokine receptor binding(GO:0031735) |
| 2.2 | 13.2 | GO:0034190 | apolipoprotein receptor binding(GO:0034190) |
| 2.2 | 10.9 | GO:0016434 | rRNA (cytosine) methyltransferase activity(GO:0016434) |
| 2.0 | 6.1 | GO:0047312 | L-phenylalanine:pyruvate aminotransferase activity(GO:0047312) glutamine-phenylpyruvate transaminase activity(GO:0047316) L-glutamine:pyruvate aminotransferase activity(GO:0047945) |
| 1.9 | 18.7 | GO:0035374 | chondroitin sulfate binding(GO:0035374) |
| 1.9 | 14.8 | GO:0086006 | voltage-gated sodium channel activity involved in cardiac muscle cell action potential(GO:0086006) |
| 1.8 | 12.9 | GO:0031812 | G-protein coupled nucleotide receptor binding(GO:0031811) P2Y1 nucleotide receptor binding(GO:0031812) |
| 1.8 | 5.5 | GO:0070039 | rRNA (guanosine-2'-O-)-methyltransferase activity(GO:0070039) |
| 1.8 | 27.0 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 1.8 | 7.1 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 1.8 | 40.3 | GO:0001134 | transcription factor activity, transcription factor recruiting(GO:0001134) |
| 1.7 | 5.2 | GO:0000171 | ribonuclease MRP activity(GO:0000171) |
| 1.7 | 6.9 | GO:0051431 | corticotropin-releasing hormone receptor 2 binding(GO:0051431) |
| 1.7 | 12.1 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 1.7 | 5.0 | GO:0004611 | phosphoenolpyruvate carboxykinase activity(GO:0004611) phosphoenolpyruvate carboxykinase (GTP) activity(GO:0004613) |
| 1.6 | 27.3 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 1.6 | 4.7 | GO:0004947 | bradykinin receptor activity(GO:0004947) |
| 1.5 | 6.0 | GO:0031800 | type 3 metabotropic glutamate receptor binding(GO:0031800) |
| 1.4 | 2.9 | GO:0033549 | MAP kinase phosphatase activity(GO:0033549) |
| 1.4 | 4.3 | GO:0016501 | prostacyclin receptor activity(GO:0016501) |
| 1.4 | 11.3 | GO:0031821 | G-protein coupled serotonin receptor binding(GO:0031821) |
| 1.4 | 5.5 | GO:0050501 | hyaluronan synthase activity(GO:0050501) |
| 1.3 | 8.0 | GO:0004483 | mRNA (nucleoside-2'-O-)-methyltransferase activity(GO:0004483) |
| 1.3 | 11.7 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 1.2 | 7.5 | GO:0004118 | cGMP-stimulated cyclic-nucleotide phosphodiesterase activity(GO:0004118) |
| 1.2 | 4.8 | GO:0034617 | tetrahydrobiopterin binding(GO:0034617) |
| 1.2 | 9.6 | GO:0004726 | non-membrane spanning protein tyrosine phosphatase activity(GO:0004726) |
| 1.2 | 22.3 | GO:0030957 | Tat protein binding(GO:0030957) |
| 1.2 | 3.5 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) |
| 1.1 | 17.2 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 1.1 | 16.9 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 1.1 | 3.4 | GO:0005519 | cytoskeletal regulatory protein binding(GO:0005519) |
| 1.1 | 4.4 | GO:0015222 | serotonin transmembrane transporter activity(GO:0015222) |
| 1.1 | 31.7 | GO:0031489 | myosin V binding(GO:0031489) |
| 1.1 | 19.6 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 1.0 | 41.2 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 1.0 | 9.2 | GO:0032564 | dATP binding(GO:0032564) |
| 1.0 | 12.7 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.9 | 10.9 | GO:0097322 | 7SK snRNA binding(GO:0097322) |
| 0.9 | 2.7 | GO:0003692 | left-handed Z-DNA binding(GO:0003692) |
| 0.9 | 24.8 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.9 | 3.5 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.9 | 3.5 | GO:0035243 | protein-arginine omega-N symmetric methyltransferase activity(GO:0035243) |
| 0.8 | 2.4 | GO:0016426 | tRNA (adenine) methyltransferase activity(GO:0016426) |
| 0.8 | 12.3 | GO:0038036 | sphingosine-1-phosphate receptor activity(GO:0038036) |
| 0.8 | 11.5 | GO:0031996 | thioesterase binding(GO:0031996) |
| 0.8 | 6.8 | GO:0015087 | cobalt ion transmembrane transporter activity(GO:0015087) |
| 0.7 | 5.2 | GO:0086089 | voltage-gated potassium channel activity involved in atrial cardiac muscle cell action potential repolarization(GO:0086089) |
| 0.7 | 5.2 | GO:0004013 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.7 | 2.2 | GO:0061711 | N(6)-L-threonylcarbamoyladenine synthase(GO:0061711) |
| 0.7 | 9.6 | GO:0048403 | brain-derived neurotrophic factor binding(GO:0048403) |
| 0.7 | 8.0 | GO:0031419 | cobalamin binding(GO:0031419) |
| 0.7 | 16.1 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.7 | 2.2 | GO:0051022 | Rho GDP-dissociation inhibitor binding(GO:0051022) |
| 0.7 | 7.9 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.7 | 3.5 | GO:0070012 | oligopeptidase activity(GO:0070012) |
| 0.7 | 14.1 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.7 | 2.8 | GO:0004741 | [pyruvate dehydrogenase (lipoamide)] phosphatase activity(GO:0004741) |
| 0.7 | 4.1 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.7 | 6.8 | GO:0005225 | volume-sensitive anion channel activity(GO:0005225) |
| 0.7 | 22.3 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.7 | 3.9 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.7 | 3.9 | GO:0010853 | cyclase activator activity(GO:0010853) guanylate cyclase activator activity(GO:0030250) |
| 0.7 | 36.5 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.7 | 20.2 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.6 | 6.5 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.6 | 22.0 | GO:0070273 | phosphatidylinositol-4-phosphate binding(GO:0070273) |
| 0.6 | 3.9 | GO:0004720 | protein-lysine 6-oxidase activity(GO:0004720) |
| 0.6 | 10.1 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.6 | 2.5 | GO:0031698 | beta-2 adrenergic receptor binding(GO:0031698) |
| 0.6 | 4.3 | GO:0043184 | vascular endothelial growth factor receptor 2 binding(GO:0043184) |
| 0.6 | 56.4 | GO:0048365 | Rac GTPase binding(GO:0048365) |
| 0.6 | 5.9 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.6 | 4.7 | GO:0035727 | lysophosphatidic acid binding(GO:0035727) |
| 0.6 | 24.2 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.6 | 5.7 | GO:0045340 | mercury ion binding(GO:0045340) |
| 0.6 | 3.9 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.5 | 8.1 | GO:0005003 | ephrin receptor activity(GO:0005003) |
| 0.5 | 6.8 | GO:0035473 | lipase binding(GO:0035473) |
| 0.5 | 2.1 | GO:1990932 | 5.8S rRNA binding(GO:1990932) |
| 0.5 | 18.7 | GO:0005355 | glucose transmembrane transporter activity(GO:0005355) |
| 0.5 | 5.7 | GO:0043008 | ATP-dependent protein binding(GO:0043008) |
| 0.5 | 2.5 | GO:0015189 | L-lysine transmembrane transporter activity(GO:0015189) |
| 0.5 | 5.8 | GO:0031802 | type 5 metabotropic glutamate receptor binding(GO:0031802) |
| 0.5 | 1.4 | GO:0061629 | RNA polymerase II sequence-specific DNA binding transcription factor binding(GO:0061629) |
| 0.5 | 3.7 | GO:0097108 | hedgehog family protein binding(GO:0097108) |
| 0.5 | 8.7 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.4 | 2.7 | GO:0009019 | tRNA (guanine-N1-)-methyltransferase activity(GO:0009019) |
| 0.4 | 5.4 | GO:0036374 | glutathione hydrolase activity(GO:0036374) |
| 0.4 | 12.8 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.4 | 13.0 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.4 | 20.1 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.4 | 4.5 | GO:0002151 | G-quadruplex RNA binding(GO:0002151) G-quadruplex DNA binding(GO:0051880) |
| 0.4 | 39.3 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.4 | 5.1 | GO:0008603 | cAMP-dependent protein kinase regulator activity(GO:0008603) |
| 0.4 | 8.1 | GO:0008171 | O-methyltransferase activity(GO:0008171) |
| 0.4 | 7.3 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.4 | 1.9 | GO:0004370 | glycerol kinase activity(GO:0004370) |
| 0.4 | 1.9 | GO:0004045 | aminoacyl-tRNA hydrolase activity(GO:0004045) |
| 0.4 | 17.4 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.4 | 13.3 | GO:0005242 | inward rectifier potassium channel activity(GO:0005242) |
| 0.4 | 1.1 | GO:0050510 | N-acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase activity(GO:0050510) |
| 0.4 | 1.8 | GO:1904288 | BAT3 complex binding(GO:1904288) |
| 0.4 | 7.6 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 0.4 | 7.9 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.4 | 21.9 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.3 | 14.5 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.3 | 6.7 | GO:0004551 | nucleotide diphosphatase activity(GO:0004551) |
| 0.3 | 31.0 | GO:0016765 | transferase activity, transferring alkyl or aryl (other than methyl) groups(GO:0016765) |
| 0.3 | 3.3 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 0.3 | 3.5 | GO:1990715 | mRNA CDS binding(GO:1990715) |
| 0.3 | 6.8 | GO:0003995 | acyl-CoA dehydrogenase activity(GO:0003995) |
| 0.3 | 5.7 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.3 | 14.3 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.3 | 4.1 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 0.3 | 18.7 | GO:0030276 | clathrin binding(GO:0030276) |
| 0.3 | 1.6 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 0.3 | 5.8 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.3 | 15.1 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.3 | 3.9 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.3 | 0.8 | GO:0004019 | adenylosuccinate synthase activity(GO:0004019) |
| 0.3 | 12.1 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.3 | 0.8 | GO:0004605 | phosphatidate cytidylyltransferase activity(GO:0004605) |
| 0.2 | 9.4 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.2 | 3.2 | GO:0008376 | acetylgalactosaminyltransferase activity(GO:0008376) |
| 0.2 | 13.7 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.2 | 2.9 | GO:0102336 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.2 | 1.9 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.2 | 3.5 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.2 | 1.9 | GO:0032451 | demethylase activity(GO:0032451) |
| 0.2 | 4.6 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.2 | 3.1 | GO:0008097 | 5S rRNA binding(GO:0008097) |
| 0.2 | 3.1 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
| 0.2 | 10.4 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.2 | 0.8 | GO:0034211 | GTP-dependent protein kinase activity(GO:0034211) |
| 0.2 | 75.2 | GO:0015631 | tubulin binding(GO:0015631) |
| 0.2 | 1.5 | GO:0004723 | calcium-dependent protein serine/threonine phosphatase activity(GO:0004723) |
| 0.2 | 1.9 | GO:0015204 | urea transmembrane transporter activity(GO:0015204) |
| 0.2 | 12.0 | GO:0009055 | electron carrier activity(GO:0009055) |
| 0.2 | 9.9 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.2 | 3.9 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.2 | 3.5 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.2 | 36.6 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.2 | 5.5 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.2 | 1.2 | GO:0019788 | NEDD8 transferase activity(GO:0019788) |
| 0.2 | 26.3 | GO:0047485 | protein N-terminus binding(GO:0047485) |
| 0.2 | 3.0 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.2 | 1.1 | GO:1990381 | ubiquitin-specific protease binding(GO:1990381) |
| 0.2 | 14.1 | GO:0005249 | voltage-gated potassium channel activity(GO:0005249) |
| 0.2 | 4.7 | GO:0070888 | E-box binding(GO:0070888) |
| 0.2 | 2.0 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.1 | 11.8 | GO:0043621 | protein self-association(GO:0043621) |
| 0.1 | 2.3 | GO:0048156 | tau protein binding(GO:0048156) |
| 0.1 | 10.4 | GO:0003724 | RNA helicase activity(GO:0003724) ATP-dependent RNA helicase activity(GO:0004004) RNA-dependent ATPase activity(GO:0008186) |
| 0.1 | 3.4 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.1 | 9.4 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.1 | 0.6 | GO:0017150 | tRNA dihydrouridine synthase activity(GO:0017150) |
| 0.1 | 0.6 | GO:0005041 | low-density lipoprotein receptor activity(GO:0005041) |
| 0.1 | 17.5 | GO:0035257 | nuclear hormone receptor binding(GO:0035257) |
| 0.1 | 8.1 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.1 | 6.5 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.1 | 6.2 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.1 | 4.4 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 8.2 | GO:0008395 | steroid hydroxylase activity(GO:0008395) |
| 0.1 | 7.8 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.1 | 14.3 | GO:0051213 | dioxygenase activity(GO:0051213) |
| 0.1 | 38.7 | GO:0061630 | ubiquitin protein ligase activity(GO:0061630) |
| 0.1 | 9.7 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.1 | 1.2 | GO:0009008 | DNA-methyltransferase activity(GO:0009008) |
| 0.1 | 5.7 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.1 | 46.1 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.1 | 14.3 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 10.3 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.1 | 21.6 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.1 | 1.7 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.1 | 3.6 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.1 | 11.5 | GO:0005179 | hormone activity(GO:0005179) |
| 0.1 | 2.3 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) |
| 0.1 | 21.8 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.1 | 1.6 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
| 0.1 | 3.0 | GO:0070001 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.1 | 1.0 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) ADP transmembrane transporter activity(GO:0015217) |
| 0.1 | 2.9 | GO:0019707 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.1 | 0.9 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.1 | 8.5 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.1 | 3.9 | GO:0031491 | nucleosome binding(GO:0031491) |
| 0.1 | 3.3 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.1 | 5.4 | GO:0005518 | collagen binding(GO:0005518) |
| 0.1 | 45.2 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.1 | 2.5 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.1 | 0.5 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.1 | 2.7 | GO:0051539 | 4 iron, 4 sulfur cluster binding(GO:0051539) |
| 0.1 | 13.3 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.1 | 1.2 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.1 | 8.6 | GO:0015171 | amino acid transmembrane transporter activity(GO:0015171) |
| 0.1 | 0.9 | GO:0004861 | cyclin-dependent protein serine/threonine kinase inhibitor activity(GO:0004861) |
| 0.1 | 2.3 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.1 | 4.1 | GO:0008237 | metallopeptidase activity(GO:0008237) |
| 0.1 | 8.0 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.1 | 16.6 | GO:0045296 | cadherin binding(GO:0045296) |
| 0.1 | 0.3 | GO:0035614 | snRNA stem-loop binding(GO:0035614) |
| 0.1 | 4.6 | GO:0004497 | monooxygenase activity(GO:0004497) |
| 0.1 | 2.2 | GO:0016878 | acid-thiol ligase activity(GO:0016878) |
| 0.1 | 2.0 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.0 | 2.5 | GO:0002039 | p53 binding(GO:0002039) |
| 0.0 | 2.4 | GO:0046906 | tetrapyrrole binding(GO:0046906) |
| 0.0 | 25.0 | GO:0008289 | lipid binding(GO:0008289) |
| 0.0 | 4.0 | GO:0004702 | receptor signaling protein serine/threonine kinase activity(GO:0004702) |
| 0.0 | 1.8 | GO:0005160 | transforming growth factor beta receptor binding(GO:0005160) |
| 0.0 | 1.5 | GO:0004714 | transmembrane receptor protein tyrosine kinase activity(GO:0004714) |
| 0.0 | 0.5 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.0 | 1.7 | GO:0030145 | manganese ion binding(GO:0030145) |
| 0.0 | 3.6 | GO:0050839 | cell adhesion molecule binding(GO:0050839) |
| 0.0 | 1.5 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.0 | 2.5 | GO:0001046 | core promoter sequence-specific DNA binding(GO:0001046) |
| 0.0 | 5.0 | GO:0003779 | actin binding(GO:0003779) |
| 0.0 | 3.2 | GO:0008022 | protein C-terminus binding(GO:0008022) |
| 0.0 | 4.5 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.0 | 0.7 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.5 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.0 | 1.3 | GO:0005319 | lipid transporter activity(GO:0005319) |
| 0.0 | 0.3 | GO:0042800 | histone methyltransferase activity (H3-K4 specific)(GO:0042800) |
| 0.0 | 4.6 | GO:0005525 | GTP binding(GO:0005525) |
| 0.0 | 2.6 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.0 | 7.4 | GO:0042803 | protein homodimerization activity(GO:0042803) |
| 0.0 | 0.2 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.0 | 9.5 | GO:0005102 | receptor binding(GO:0005102) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.9 | 99.2 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.9 | 9.9 | ST TYPE I INTERFERON PATHWAY | Type I Interferon (alpha/beta IFN) Pathway |
| 0.8 | 25.7 | PID S1P S1P2 PATHWAY | S1P2 pathway |
| 0.8 | 27.9 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.7 | 26.7 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.6 | 16.9 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.6 | 21.6 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.6 | 20.2 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.6 | 40.1 | PID ARF6 TRAFFICKING PATHWAY | Arf6 trafficking events |
| 0.6 | 28.0 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.6 | 3.9 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.5 | 7.6 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.4 | 15.0 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.4 | 5.9 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.4 | 22.0 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.4 | 13.3 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.3 | 25.1 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.3 | 13.4 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.3 | 8.9 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.2 | 14.3 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.2 | 2.8 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.2 | 5.1 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.2 | 10.3 | PID HIV NEF PATHWAY | HIV-1 Nef: Negative effector of Fas and TNF-alpha |
| 0.2 | 7.1 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.2 | 5.5 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.2 | 9.4 | PID PI3KCI PATHWAY | Class I PI3K signaling events |
| 0.2 | 13.0 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.2 | 5.6 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.2 | 8.8 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.2 | 0.3 | PID VEGFR1 PATHWAY | VEGFR1 specific signals |
| 0.2 | 10.4 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.2 | 2.5 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.1 | 3.7 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.1 | 10.9 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.1 | 18.6 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.1 | 3.5 | PID ER NONGENOMIC PATHWAY | Plasma membrane estrogen receptor signaling |
| 0.1 | 4.4 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.1 | 4.3 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.1 | 3.6 | PID VEGFR1 2 PATHWAY | Signaling events mediated by VEGFR1 and VEGFR2 |
| 0.1 | 2.5 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.1 | 1.9 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.1 | 1.9 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.1 | 4.4 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.1 | 1.5 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.1 | 1.3 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.1 | 1.7 | PID CXCR4 PATHWAY | CXCR4-mediated signaling events |
| 0.1 | 2.0 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.1 | 3.9 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 0.1 | 2.6 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.1 | 1.5 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.0 | 15.3 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.0 | 3.3 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 0.7 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.0 | 0.8 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.0 | 1.0 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.0 | 5.2 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.9 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 1.1 | PID PDGFRB PATHWAY | PDGFR-beta signaling pathway |
| 0.0 | 2.8 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.4 | 41.5 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 2.4 | 41.4 | REACTOME DOPAMINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Dopamine Neurotransmitter Release Cycle |
| 1.4 | 16.9 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 1.2 | 23.4 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.9 | 37.7 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.9 | 18.1 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.8 | 16.1 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.7 | 14.0 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.7 | 9.4 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.7 | 20.8 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.7 | 37.5 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.6 | 11.3 | REACTOME G PROTEIN ACTIVATION | Genes involved in G-protein activation |
| 0.6 | 8.9 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.6 | 9.4 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.5 | 11.3 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.5 | 14.9 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.4 | 8.7 | REACTOME FORMATION OF ATP BY CHEMIOSMOTIC COUPLING | Genes involved in Formation of ATP by chemiosmotic coupling |
| 0.4 | 2.9 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.4 | 21.3 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.4 | 10.1 | REACTOME SPRY REGULATION OF FGF SIGNALING | Genes involved in Spry regulation of FGF signaling |
| 0.4 | 7.0 | REACTOME CTNNB1 PHOSPHORYLATION CASCADE | Genes involved in Beta-catenin phosphorylation cascade |
| 0.4 | 3.9 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.4 | 11.0 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.4 | 12.8 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.4 | 26.7 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.3 | 11.0 | REACTOME DOWNREGULATION OF TGF BETA RECEPTOR SIGNALING | Genes involved in Downregulation of TGF-beta receptor signaling |
| 0.3 | 10.3 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.3 | 5.0 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.3 | 5.5 | REACTOME HYALURONAN METABOLISM | Genes involved in Hyaluronan metabolism |
| 0.3 | 15.9 | REACTOME IL RECEPTOR SHC SIGNALING | Genes involved in Interleukin receptor SHC signaling |
| 0.3 | 8.2 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS | Genes involved in Synthesis of bile acids and bile salts |
| 0.3 | 3.9 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.3 | 53.4 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.3 | 3.6 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.3 | 15.2 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.2 | 6.1 | REACTOME PYRUVATE METABOLISM | Genes involved in Pyruvate metabolism |
| 0.2 | 12.7 | REACTOME CIRCADIAN CLOCK | Genes involved in Circadian Clock |
| 0.2 | 12.2 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.2 | 7.3 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 3 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 3 Promoter |
| 0.2 | 2.5 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.2 | 8.4 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.2 | 4.8 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.2 | 2.2 | REACTOME CD28 DEPENDENT VAV1 PATHWAY | Genes involved in CD28 dependent Vav1 pathway |
| 0.2 | 7.8 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.2 | 7.3 | REACTOME INSULIN RECEPTOR RECYCLING | Genes involved in Insulin receptor recycling |
| 0.2 | 14.3 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.2 | 2.1 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 0.2 | 9.0 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.2 | 3.5 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.2 | 3.8 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.2 | 4.5 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.2 | 2.9 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.1 | 15.0 | REACTOME CLASS A1 RHODOPSIN LIKE RECEPTORS | Genes involved in Class A/1 (Rhodopsin-like receptors) |
| 0.1 | 3.3 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.1 | 1.5 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.1 | 14.0 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.1 | 6.0 | REACTOME MUSCLE CONTRACTION | Genes involved in Muscle contraction |
| 0.1 | 7.9 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.1 | 27.8 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.1 | 11.7 | REACTOME G ALPHA S SIGNALLING EVENTS | Genes involved in G alpha (s) signalling events |
| 0.1 | 2.7 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.1 | 5.5 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.1 | 4.5 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 0.9 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.1 | 3.6 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.1 | 6.3 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.1 | 0.5 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.1 | 8.0 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.1 | 1.6 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.1 | 6.4 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.1 | 15.9 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.1 | 4.6 | REACTOME SIGNALING BY PDGF | Genes involved in Signaling by PDGF |
| 0.1 | 3.3 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.1 | 3.5 | REACTOME REGULATION OF MRNA STABILITY BY PROTEINS THAT BIND AU RICH ELEMENTS | Genes involved in Regulation of mRNA Stability by Proteins that Bind AU-rich Elements |
| 0.1 | 2.5 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.1 | 2.7 | REACTOME MRNA 3 END PROCESSING | Genes involved in mRNA 3'-end processing |
| 0.0 | 1.1 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |
| 0.0 | 1.9 | REACTOME AMINE COMPOUND SLC TRANSPORTERS | Genes involved in Amine compound SLC transporters |
| 0.0 | 2.7 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 0.9 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.0 | 2.5 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.0 | 9.7 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.0 | 0.5 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.0 | 1.6 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.0 | 1.3 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.0 | 2.4 | REACTOME ACTIVATION OF THE MRNA UPON BINDING OF THE CAP BINDING COMPLEX AND EIFS AND SUBSEQUENT BINDING TO 43S | Genes involved in Activation of the mRNA upon binding of the cap-binding complex and eIFs, and subsequent binding to 43S |
| 0.0 | 0.7 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.0 | 0.3 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.0 | 2.7 | REACTOME GLYCEROPHOSPHOLIPID BIOSYNTHESIS | Genes involved in Glycerophospholipid biosynthesis |
| 0.0 | 0.3 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.0 | 1.3 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |