Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
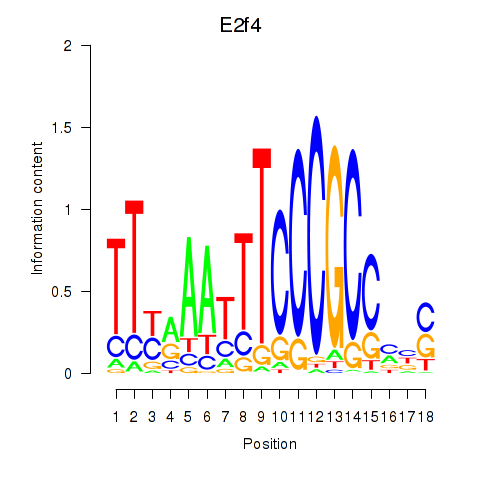
Results for E2f4
Z-value: 2.36
Transcription factors associated with E2f4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
E2f4
|
ENSMUSG00000014859.10 | E2f4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| E2f4 | mm39_v1_chr8_+_106024294_106024367 | 0.88 | 3.2e-24 | Click! |
Activity profile of E2f4 motif
Sorted Z-values of E2f4 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of E2f4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_+_70703409 | 25.59 |
ENSMUST00000103410.3
|
Igkc
|
immunoglobulin kappa constant |
| chr11_+_68936457 | 22.76 |
ENSMUST00000108666.8
ENSMUST00000021277.6 |
Aurkb
|
aurora kinase B |
| chr8_-_54091980 | 21.86 |
ENSMUST00000047768.11
|
Neil3
|
nei like 3 (E. coli) |
| chr6_+_124807176 | 21.79 |
ENSMUST00000131847.8
ENSMUST00000151674.3 |
Cdca3
|
cell division cycle associated 3 |
| chr9_+_72345801 | 21.50 |
ENSMUST00000184604.8
ENSMUST00000034746.10 |
Mns1
|
meiosis-specific nuclear structural protein 1 |
| chr1_-_169358912 | 21.42 |
ENSMUST00000192248.2
ENSMUST00000028000.13 |
Nuf2
|
NUF2, NDC80 kinetochore complex component |
| chr1_-_189420270 | 21.39 |
ENSMUST00000171929.8
ENSMUST00000165962.2 |
Cenpf
|
centromere protein F |
| chr9_+_72345267 | 21.29 |
ENSMUST00000183809.8
|
Mns1
|
meiosis-specific nuclear structural protein 1 |
| chr4_+_134238310 | 20.26 |
ENSMUST00000105866.3
|
Aunip
|
aurora kinase A and ninein interacting protein |
| chr1_-_169359015 | 19.93 |
ENSMUST00000111368.8
|
Nuf2
|
NUF2, NDC80 kinetochore complex component |
| chr15_+_79776597 | 19.83 |
ENSMUST00000175714.8
ENSMUST00000109620.10 ENSMUST00000165537.8 ENSMUST00000175752.8 ENSMUST00000176325.8 |
Apobec3
|
apolipoprotein B mRNA editing enzyme, catalytic polypeptide 3 |
| chr18_-_34884555 | 19.35 |
ENSMUST00000060710.9
|
Cdc25c
|
cell division cycle 25C |
| chr19_+_6135013 | 18.84 |
ENSMUST00000025704.3
|
Cdca5
|
cell division cycle associated 5 |
| chr14_-_87378641 | 16.61 |
ENSMUST00000168889.3
ENSMUST00000022599.14 |
Diaph3
|
diaphanous related formin 3 |
| chr10_-_87982732 | 16.42 |
ENSMUST00000164121.8
ENSMUST00000164803.2 ENSMUST00000168163.8 ENSMUST00000048518.16 |
Parpbp
|
PARP1 binding protein |
| chr4_-_133694607 | 16.15 |
ENSMUST00000105893.8
|
Hmgn2
|
high mobility group nucleosomal binding domain 2 |
| chr7_-_37806912 | 15.41 |
ENSMUST00000108023.10
|
Ccne1
|
cyclin E1 |
| chr11_-_101442663 | 15.20 |
ENSMUST00000017290.11
|
Brca1
|
breast cancer 1, early onset |
| chr4_-_133694543 | 15.13 |
ENSMUST00000123234.8
|
Hmgn2
|
high mobility group nucleosomal binding domain 2 |
| chr7_-_92523396 | 14.95 |
ENSMUST00000209074.2
ENSMUST00000208356.2 ENSMUST00000032877.11 |
Ddias
|
DNA damage-induced apoptosis suppressor |
| chr6_+_113508636 | 14.87 |
ENSMUST00000036340.12
ENSMUST00000204827.3 |
Fancd2
|
Fanconi anemia, complementation group D2 |
| chr4_-_115980813 | 14.27 |
ENSMUST00000102704.4
ENSMUST00000102705.10 |
Rad54l
|
RAD54 like (S. cerevisiae) |
| chr4_-_132072988 | 13.95 |
ENSMUST00000030726.13
|
Rcc1
|
regulator of chromosome condensation 1 |
| chr18_+_56840813 | 13.93 |
ENSMUST00000025486.9
|
Lmnb1
|
lamin B1 |
| chr4_-_132073048 | 13.62 |
ENSMUST00000084250.11
|
Rcc1
|
regulator of chromosome condensation 1 |
| chr1_+_52669849 | 13.35 |
ENSMUST00000165859.8
|
Nemp2
|
nuclear envelope integral membrane protein 2 |
| chr12_+_24758240 | 12.94 |
ENSMUST00000020980.12
|
Rrm2
|
ribonucleotide reductase M2 |
| chr13_-_74085880 | 12.51 |
ENSMUST00000022053.11
|
Trip13
|
thyroid hormone receptor interactor 13 |
| chr16_-_90524214 | 12.33 |
ENSMUST00000099554.5
|
Mis18a
|
MIS18 kinetochore protein A |
| chr2_+_24257576 | 12.05 |
ENSMUST00000140547.2
ENSMUST00000102942.8 |
Psd4
|
pleckstrin and Sec7 domain containing 4 |
| chr17_-_25946370 | 11.74 |
ENSMUST00000170070.3
ENSMUST00000048054.14 |
Chtf18
|
CTF18, chromosome transmission fidelity factor 18 |
| chr18_-_60981981 | 11.33 |
ENSMUST00000177172.8
ENSMUST00000175934.8 ENSMUST00000176630.8 |
Tcof1
|
treacle ribosome biogenesis factor 1 |
| chr4_+_11558905 | 11.13 |
ENSMUST00000095145.12
ENSMUST00000108306.9 ENSMUST00000070755.13 |
Rad54b
|
RAD54 homolog B (S. cerevisiae) |
| chr12_+_117807607 | 10.50 |
ENSMUST00000176735.8
ENSMUST00000177339.2 |
Cdca7l
|
cell division cycle associated 7 like |
| chr16_-_30129591 | 10.37 |
ENSMUST00000061190.8
|
Gp5
|
glycoprotein 5 (platelet) |
| chr4_-_133695264 | 10.36 |
ENSMUST00000102553.11
|
Hmgn2
|
high mobility group nucleosomal binding domain 2 |
| chr9_+_44245981 | 10.28 |
ENSMUST00000052686.4
|
H2ax
|
H2A.X variant histone |
| chr4_-_133695204 | 9.92 |
ENSMUST00000100472.10
ENSMUST00000136327.2 |
Hmgn2
|
high mobility group nucleosomal binding domain 2 |
| chr9_+_107828136 | 9.79 |
ENSMUST00000049348.9
ENSMUST00000194271.2 |
Traip
|
TRAF-interacting protein |
| chr1_-_33708876 | 9.12 |
ENSMUST00000027312.11
|
Prim2
|
DNA primase, p58 subunit |
| chr8_+_108271643 | 8.86 |
ENSMUST00000212543.2
|
Wwp2
|
WW domain containing E3 ubiquitin protein ligase 2 |
| chr6_-_56681657 | 8.68 |
ENSMUST00000176595.3
ENSMUST00000170382.5 ENSMUST00000203958.2 |
Lsm5
|
LSM5 homolog, U6 small nuclear RNA and mRNA degradation associated |
| chr2_+_162896602 | 8.54 |
ENSMUST00000018005.10
|
Mybl2
|
myeloblastosis oncogene-like 2 |
| chrX_+_141464393 | 8.34 |
ENSMUST00000112889.8
ENSMUST00000101198.9 ENSMUST00000112891.8 ENSMUST00000087333.9 |
Tmem164
|
transmembrane protein 164 |
| chrX_-_165992145 | 8.22 |
ENSMUST00000112176.8
|
Tmsb4x
|
thymosin, beta 4, X chromosome |
| chr5_+_31046196 | 8.18 |
ENSMUST00000069705.11
ENSMUST00000201917.4 ENSMUST00000201168.4 ENSMUST00000202060.4 ENSMUST00000201225.4 ENSMUST00000114700.9 |
Agbl5
|
ATP/GTP binding protein-like 5 |
| chr15_-_76524097 | 8.08 |
ENSMUST00000168185.8
|
Tonsl
|
tonsoku-like, DNA repair protein |
| chrX_+_163763588 | 8.00 |
ENSMUST00000167446.8
ENSMUST00000057150.8 |
Fancb
|
Fanconi anemia, complementation group B |
| chr2_+_112114906 | 7.80 |
ENSMUST00000053666.8
|
Slc12a6
|
solute carrier family 12, member 6 |
| chr7_+_126380655 | 7.68 |
ENSMUST00000172352.8
ENSMUST00000094037.5 |
Tbx6
|
T-box 6 |
| chr8_+_18645144 | 7.25 |
ENSMUST00000039412.15
|
Mcph1
|
microcephaly, primary autosomal recessive 1 |
| chr4_-_59783780 | 7.00 |
ENSMUST00000107526.8
ENSMUST00000095063.11 |
Inip
|
INTS3 and NABP interacting protein |
| chr5_-_33809872 | 6.87 |
ENSMUST00000057551.14
|
Slbp
|
stem-loop binding protein |
| chrX_+_8137372 | 6.63 |
ENSMUST00000127103.8
ENSMUST00000115591.8 |
Slc38a5
|
solute carrier family 38, member 5 |
| chr12_+_105976463 | 6.56 |
ENSMUST00000072040.7
|
Vrk1
|
vaccinia related kinase 1 |
| chr5_+_21577640 | 6.38 |
ENSMUST00000035799.6
|
Fgl2
|
fibrinogen-like protein 2 |
| chr17_+_35219941 | 6.26 |
ENSMUST00000087315.14
|
Vars
|
valyl-tRNA synthetase |
| chr5_-_33809640 | 6.21 |
ENSMUST00000151081.2
ENSMUST00000139518.8 ENSMUST00000101354.10 |
Slbp
|
stem-loop binding protein |
| chr2_-_3513783 | 6.10 |
ENSMUST00000124331.8
ENSMUST00000140494.2 ENSMUST00000027961.12 |
Gm45902
Hspa14
|
predicted gene 45902 heat shock protein 14 |
| chr17_+_35201074 | 6.02 |
ENSMUST00000007266.14
|
Lsm2
|
LSM2 homolog, U6 small nuclear RNA and mRNA degradation associated |
| chr12_+_105976533 | 5.98 |
ENSMUST00000085026.12
ENSMUST00000021539.16 |
Vrk1
|
vaccinia related kinase 1 |
| chr9_-_82857528 | 5.92 |
ENSMUST00000034787.12
|
Phip
|
pleckstrin homology domain interacting protein |
| chr5_-_138170644 | 5.78 |
ENSMUST00000000505.16
|
Mcm7
|
minichromosome maintenance complex component 7 |
| chr2_+_180961507 | 5.77 |
ENSMUST00000098971.11
ENSMUST00000054622.15 ENSMUST00000108814.8 ENSMUST00000048608.16 ENSMUST00000108815.8 |
Rtel1
|
regulator of telomere elongation helicase 1 |
| chr17_-_35265702 | 5.76 |
ENSMUST00000097338.11
|
Msh5
|
mutS homolog 5 |
| chr6_-_69394425 | 5.74 |
ENSMUST00000199160.2
|
Igkv4-61
|
immunoglobulin kappa chain variable 4-61 |
| chr17_+_35219998 | 5.68 |
ENSMUST00000173584.8
|
Vars
|
valyl-tRNA synthetase |
| chrX_+_8137620 | 5.60 |
ENSMUST00000033512.11
|
Slc38a5
|
solute carrier family 38, member 5 |
| chr15_+_57775595 | 5.48 |
ENSMUST00000022992.13
|
Tbc1d31
|
TBC1 domain family, member 31 |
| chrX_+_8137881 | 5.33 |
ENSMUST00000115590.2
|
Slc38a5
|
solute carrier family 38, member 5 |
| chr10_+_87982854 | 5.24 |
ENSMUST00000052355.15
|
Nup37
|
nucleoporin 37 |
| chr11_+_16901370 | 5.20 |
ENSMUST00000109635.2
ENSMUST00000061327.2 |
Fbxo48
|
F-box protein 48 |
| chr5_+_123214332 | 5.18 |
ENSMUST00000067505.15
ENSMUST00000111619.10 ENSMUST00000160344.2 |
Tmem120b
|
transmembrane protein 120B |
| chr11_+_74510413 | 5.13 |
ENSMUST00000100866.3
|
Ccdc92b
|
coiled-coil domain containing 92B |
| chr17_-_84495364 | 5.10 |
ENSMUST00000060366.7
|
Zfp36l2
|
zinc finger protein 36, C3H type-like 2 |
| chr19_-_5416626 | 5.05 |
ENSMUST00000237167.2
|
Banf1
|
BAF nuclear assembly factor 1 |
| chr10_+_87982916 | 5.04 |
ENSMUST00000169309.3
|
Nup37
|
nucleoporin 37 |
| chr5_-_24652775 | 4.99 |
ENSMUST00000123167.2
ENSMUST00000030799.15 |
Tmub1
|
transmembrane and ubiquitin-like domain containing 1 |
| chr17_+_35201130 | 4.95 |
ENSMUST00000173004.2
|
Lsm2
|
LSM2 homolog, U6 small nuclear RNA and mRNA degradation associated |
| chr7_+_58307930 | 4.94 |
ENSMUST00000168747.3
|
Atp10a
|
ATPase, class V, type 10A |
| chr9_-_32255604 | 4.93 |
ENSMUST00000034533.7
|
Kcnj5
|
potassium inwardly-rectifying channel, subfamily J, member 5 |
| chr5_-_33809683 | 4.82 |
ENSMUST00000075670.13
|
Slbp
|
stem-loop binding protein |
| chr1_+_9868332 | 4.74 |
ENSMUST00000166384.8
ENSMUST00000168907.8 |
Sgk3
|
serum/glucocorticoid regulated kinase 3 |
| chr4_-_3835595 | 4.72 |
ENSMUST00000138502.2
|
Rps20
|
ribosomal protein S20 |
| chr5_-_92496730 | 4.60 |
ENSMUST00000038816.13
ENSMUST00000118006.3 |
Cxcl10
|
chemokine (C-X-C motif) ligand 10 |
| chr12_+_99850756 | 4.59 |
ENSMUST00000153627.8
|
Tdp1
|
tyrosyl-DNA phosphodiesterase 1 |
| chr9_+_108820846 | 4.53 |
ENSMUST00000198140.5
ENSMUST00000051873.15 |
Pfkfb4
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 4 |
| chr13_-_23929490 | 4.49 |
ENSMUST00000091752.5
|
H3c3
|
H3 clustered histone 3 |
| chr9_+_57615816 | 4.48 |
ENSMUST00000043990.14
ENSMUST00000142807.2 |
Edc3
|
enhancer of mRNA decapping 3 |
| chrX_+_104123341 | 4.42 |
ENSMUST00000033577.11
|
Pbdc1
|
polysaccharide biosynthesis domain containing 1 |
| chr5_-_138170077 | 4.42 |
ENSMUST00000155902.8
ENSMUST00000148879.8 |
Mcm7
|
minichromosome maintenance complex component 7 |
| chr15_-_78280099 | 4.41 |
ENSMUST00000229878.2
ENSMUST00000165170.8 ENSMUST00000074380.14 |
Tex33
|
testis expressed 33 |
| chr8_+_124750133 | 4.36 |
ENSMUST00000034457.9
|
Urb2
|
URB2 ribosome biogenesis 2 homolog (S. cerevisiae) |
| chr2_+_129435448 | 4.31 |
ENSMUST00000049262.14
ENSMUST00000163034.8 ENSMUST00000160276.2 |
Sirpa
|
signal-regulatory protein alpha |
| chr17_+_31515163 | 4.26 |
ENSMUST00000235972.2
ENSMUST00000165149.3 ENSMUST00000236251.2 |
Slc37a1
|
solute carrier family 37 (glycerol-3-phosphate transporter), member 1 |
| chr19_+_6451667 | 4.26 |
ENSMUST00000113471.3
ENSMUST00000113469.3 |
Rasgrp2
|
RAS, guanyl releasing protein 2 |
| chr14_+_54441879 | 4.23 |
ENSMUST00000103726.2
|
Traj15
|
T cell receptor alpha joining 15 |
| chr13_+_23719579 | 4.22 |
ENSMUST00000080859.8
|
H3c8
|
H3 clustered histone 8 |
| chr9_-_83323080 | 4.19 |
ENSMUST00000034791.15
ENSMUST00000188548.7 ENSMUST00000034793.15 |
Lca5
|
Leber congenital amaurosis 5 (human) |
| chr18_+_65933586 | 4.14 |
ENSMUST00000025394.14
ENSMUST00000236847.2 ENSMUST00000153193.3 |
Sec11c
|
SEC11 homolog C, signal peptidase complex subunit |
| chr5_+_31046670 | 3.88 |
ENSMUST00000200850.2
|
Agbl5
|
ATP/GTP binding protein-like 5 |
| chr9_-_32255556 | 3.73 |
ENSMUST00000214223.2
|
Kcnj5
|
potassium inwardly-rectifying channel, subfamily J, member 5 |
| chr13_+_74085916 | 3.72 |
ENSMUST00000222399.2
ENSMUST00000099384.4 |
Brd9
|
bromodomain containing 9 |
| chr2_-_26393826 | 3.69 |
ENSMUST00000028288.5
|
Notch1
|
notch 1 |
| chr7_+_24920840 | 3.55 |
ENSMUST00000055604.6
|
Zfp526
|
zinc finger protein 526 |
| chr10_+_36850532 | 3.46 |
ENSMUST00000019911.14
ENSMUST00000105510.2 |
Hdac2
|
histone deacetylase 2 |
| chr3_-_10273628 | 3.31 |
ENSMUST00000029041.6
|
Fabp4
|
fatty acid binding protein 4, adipocyte |
| chr12_-_4283926 | 3.28 |
ENSMUST00000111169.10
ENSMUST00000020981.12 |
Cenpo
|
centromere protein O |
| chr17_+_35220252 | 3.27 |
ENSMUST00000174260.8
|
Vars
|
valyl-tRNA synthetase |
| chr5_-_113910571 | 3.19 |
ENSMUST00000019118.8
|
Sart3
|
squamous cell carcinoma antigen recognized by T cells 3 |
| chr2_+_30176395 | 3.18 |
ENSMUST00000064447.12
|
Nup188
|
nucleoporin 188 |
| chr4_-_132191181 | 3.11 |
ENSMUST00000102567.4
|
Med18
|
mediator complex subunit 18 |
| chr4_-_141553306 | 3.04 |
ENSMUST00000102481.4
|
Cela2a
|
chymotrypsin-like elastase family, member 2A |
| chr11_-_106890307 | 2.99 |
ENSMUST00000018506.13
|
Kpna2
|
karyopherin (importin) alpha 2 |
| chr2_+_71617266 | 2.98 |
ENSMUST00000112101.8
ENSMUST00000028522.10 |
Itga6
|
integrin alpha 6 |
| chr1_-_179631190 | 2.88 |
ENSMUST00000027768.14
|
Ahctf1
|
AT hook containing transcription factor 1 |
| chr9_-_32255533 | 2.75 |
ENSMUST00000216033.2
|
Kcnj5
|
potassium inwardly-rectifying channel, subfamily J, member 5 |
| chr7_+_97020810 | 2.68 |
ENSMUST00000098300.6
|
Alg8
|
asparagine-linked glycosylation 8 (alpha-1,3-glucosyltransferase) |
| chr2_+_129435115 | 2.43 |
ENSMUST00000099113.10
ENSMUST00000103202.10 |
Sirpa
|
signal-regulatory protein alpha |
| chrX_-_36373910 | 2.36 |
ENSMUST00000076265.13
|
Upf3b
|
UPF3 regulator of nonsense transcripts homolog B (yeast) |
| chr11_+_87295860 | 2.31 |
ENSMUST00000060835.12
|
Tex14
|
testis expressed gene 14 |
| chr11_+_6496875 | 2.28 |
ENSMUST00000000388.15
ENSMUST00000159007.8 |
Ccm2
|
cerebral cavernous malformation 2 |
| chr7_+_24584076 | 2.23 |
ENSMUST00000153451.9
ENSMUST00000108429.8 |
Rps19
|
ribosomal protein S19 |
| chrX_+_161684563 | 2.19 |
ENSMUST00000112303.8
ENSMUST00000033727.14 |
Ctps2
|
cytidine 5'-triphosphate synthase 2 |
| chr19_+_5416769 | 2.18 |
ENSMUST00000025759.9
|
Eif1ad
|
eukaryotic translation initiation factor 1A domain containing |
| chr2_+_30176418 | 2.08 |
ENSMUST00000138666.8
ENSMUST00000113634.3 |
Nup188
|
nucleoporin 188 |
| chr11_+_108811168 | 2.03 |
ENSMUST00000052915.14
|
Axin2
|
axin 2 |
| chr5_-_25910788 | 2.01 |
ENSMUST00000030773.12
|
Xrcc2
|
X-ray repair complementing defective repair in Chinese hamster cells 2 |
| chr3_-_6685492 | 2.01 |
ENSMUST00000091364.4
|
1700008P02Rik
|
RIKEN cDNA 1700008P02 gene |
| chr14_+_30741082 | 1.95 |
ENSMUST00000112098.11
ENSMUST00000112095.8 ENSMUST00000112106.8 ENSMUST00000146325.8 |
Pbrm1
|
polybromo 1 |
| chr4_+_44004437 | 1.92 |
ENSMUST00000107846.10
|
Clta
|
clathrin, light polypeptide (Lca) |
| chr7_+_97229065 | 1.92 |
ENSMUST00000107153.3
|
Rsf1
|
remodeling and spacing factor 1 |
| chr14_+_76725876 | 1.91 |
ENSMUST00000101618.9
|
Tsc22d1
|
TSC22 domain family, member 1 |
| chr8_-_106223502 | 1.89 |
ENSMUST00000212303.2
|
Zdhhc1
|
zinc finger, DHHC domain containing 1 |
| chr4_-_58553553 | 1.89 |
ENSMUST00000107575.9
ENSMUST00000107574.8 ENSMUST00000147354.8 |
Lpar1
|
lysophosphatidic acid receptor 1 |
| chr9_-_123507847 | 1.81 |
ENSMUST00000170591.2
ENSMUST00000171647.9 |
Slc6a20a
|
solute carrier family 6 (neurotransmitter transporter), member 20A |
| chr2_+_101508747 | 1.74 |
ENSMUST00000004949.8
|
Traf6
|
TNF receptor-associated factor 6 |
| chr8_+_18645543 | 1.73 |
ENSMUST00000146819.8
|
Mcph1
|
microcephaly, primary autosomal recessive 1 |
| chr2_+_71617402 | 1.65 |
ENSMUST00000238991.2
|
Itga6
|
integrin alpha 6 |
| chr4_-_43562397 | 1.64 |
ENSMUST00000030187.14
|
Tln1
|
talin 1 |
| chr12_-_118162652 | 1.58 |
ENSMUST00000084806.7
|
Dnah11
|
dynein, axonemal, heavy chain 11 |
| chr17_+_35200823 | 1.56 |
ENSMUST00000114011.11
|
Lsm2
|
LSM2 homolog, U6 small nuclear RNA and mRNA degradation associated |
| chr10_-_62859626 | 1.53 |
ENSMUST00000119814.3
|
Hnrnph3
|
heterogeneous nuclear ribonucleoprotein H3 |
| chr13_+_74086241 | 1.51 |
ENSMUST00000222749.2
|
Brd9
|
bromodomain containing 9 |
| chr4_-_58553311 | 1.50 |
ENSMUST00000107571.8
ENSMUST00000055018.11 |
Lpar1
|
lysophosphatidic acid receptor 1 |
| chr11_-_106890195 | 1.43 |
ENSMUST00000106768.2
ENSMUST00000144834.8 |
Kpna2
|
karyopherin (importin) alpha 2 |
| chrX_+_161684221 | 1.36 |
ENSMUST00000101095.9
|
Ctps2
|
cytidine 5'-triphosphate synthase 2 |
| chr10_+_83558729 | 1.33 |
ENSMUST00000150459.3
|
1500009L16Rik
|
RIKEN cDNA 1500009L16 gene |
| chr2_+_180961599 | 1.32 |
ENSMUST00000153112.2
|
Rtel1
|
regulator of telomere elongation helicase 1 |
| chr1_-_60082246 | 1.32 |
ENSMUST00000027172.13
ENSMUST00000191251.7 |
Ica1l
|
islet cell autoantigen 1-like |
| chr15_-_98729333 | 1.28 |
ENSMUST00000168846.3
|
Prkag1
|
protein kinase, AMP-activated, gamma 1 non-catalytic subunit |
| chr3_-_116456292 | 1.27 |
ENSMUST00000029570.9
|
Mfsd14a
|
major facilitator superfamily domain containing 14A |
| chr11_-_34048538 | 1.26 |
ENSMUST00000223852.2
|
4930469K13Rik
|
RIKEN cDNA 4930469K13 gene |
| chr14_-_33700719 | 1.24 |
ENSMUST00000166737.2
|
Zfp488
|
zinc finger protein 488 |
| chr17_+_88748139 | 1.20 |
ENSMUST00000112238.9
ENSMUST00000155640.2 |
Foxn2
|
forkhead box N2 |
| chr1_-_74640504 | 1.19 |
ENSMUST00000136078.2
ENSMUST00000132081.2 ENSMUST00000113721.8 ENSMUST00000027357.12 |
Rnf25
|
ring finger protein 25 |
| chr17_-_57338468 | 1.18 |
ENSMUST00000007814.10
ENSMUST00000233480.2 |
Khsrp
|
KH-type splicing regulatory protein |
| chr10_-_62859661 | 1.18 |
ENSMUST00000020263.14
ENSMUST00000118898.8 |
Hnrnph3
|
heterogeneous nuclear ribonucleoprotein H3 |
| chr15_+_102314578 | 1.09 |
ENSMUST00000170884.8
ENSMUST00000163709.8 |
Sp1
|
trans-acting transcription factor 1 |
| chrX_+_104123367 | 1.01 |
ENSMUST00000119477.2
|
Pbdc1
|
polysaccharide biosynthesis domain containing 1 |
| chr5_+_30437579 | 0.95 |
ENSMUST00000145167.9
|
Selenoi
|
selenoprotein I |
| chr3_-_142587419 | 0.93 |
ENSMUST00000174422.8
ENSMUST00000173830.8 |
Pkn2
|
protein kinase N2 |
| chrY_-_90850446 | 0.87 |
ENSMUST00000179623.2
|
Gm21748
|
predicted gene, 21748 |
| chr14_-_54754810 | 0.86 |
ENSMUST00000023873.12
|
Prmt5
|
protein arginine N-methyltransferase 5 |
| chr2_-_145776934 | 0.75 |
ENSMUST00000001818.5
|
Crnkl1
|
crooked neck pre-mRNA splicing factor 1 |
| chr4_-_44704006 | 0.66 |
ENSMUST00000146335.8
|
Pax5
|
paired box 5 |
| chr1_+_85822250 | 0.64 |
ENSMUST00000185569.2
|
Itm2c
|
integral membrane protein 2C |
| chrX_-_77627486 | 0.62 |
ENSMUST00000114025.8
ENSMUST00000134602.8 ENSMUST00000114024.9 |
Prrg1
|
proline rich Gla (G-carboxyglutamic acid) 1 |
| chr7_-_16019379 | 0.53 |
ENSMUST00000174270.8
|
Ccdc9
|
coiled-coil domain containing 9 |
| chr3_+_89680867 | 0.53 |
ENSMUST00000038356.13
|
Ube2q1
|
ubiquitin-conjugating enzyme E2Q family member 1 |
| chrX_+_12454031 | 0.51 |
ENSMUST00000033313.3
|
Atp6ap2
|
ATPase, H+ transporting, lysosomal accessory protein 2 |
| chr17_-_87332780 | 0.46 |
ENSMUST00000024957.7
|
Pigf
|
phosphatidylinositol glycan anchor biosynthesis, class F |
| chr15_+_12824901 | 0.45 |
ENSMUST00000169061.8
|
Drosha
|
drosha, ribonuclease type III |
| chr15_-_39807359 | 0.40 |
ENSMUST00000110305.3
|
Lrp12
|
low density lipoprotein-related protein 12 |
| chrX_+_161684735 | 0.38 |
ENSMUST00000112302.8
ENSMUST00000112301.8 |
Ctps2
|
cytidine 5'-triphosphate synthase 2 |
| chr14_+_55815580 | 0.35 |
ENSMUST00000174484.8
|
Psme1
|
proteasome (prosome, macropain) activator subunit 1 (PA28 alpha) |
| chr7_-_16019935 | 0.32 |
ENSMUST00000145519.3
|
Ccdc9
|
coiled-coil domain containing 9 |
| chr10_+_17598961 | 0.31 |
ENSMUST00000038107.9
ENSMUST00000219558.2 ENSMUST00000218370.2 |
Cited2
|
Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 |
| chr1_-_9818601 | 0.30 |
ENSMUST00000057438.7
|
Vcpip1
|
valosin containing protein (p97)/p47 complex interacting protein 1 |
| chr8_-_70426910 | 0.26 |
ENSMUST00000116463.4
|
Gatad2a
|
GATA zinc finger domain containing 2A |
| chr2_-_125993887 | 0.25 |
ENSMUST00000110448.3
ENSMUST00000110446.9 |
Fam227b
|
family with sequence similarity 227, member B |
| chr11_+_77928736 | 0.21 |
ENSMUST00000072289.12
ENSMUST00000100784.9 ENSMUST00000073660.7 |
Flot2
|
flotillin 2 |
| chrX_+_72386220 | 0.21 |
ENSMUST00000114499.8
ENSMUST00000033731.4 |
Zfp275
|
zinc finger protein 275 |
| chr7_-_126859790 | 0.18 |
ENSMUST00000035276.5
|
Dctpp1
|
dCTP pyrophosphatase 1 |
| chr9_-_103243039 | 0.11 |
ENSMUST00000035484.11
|
Cdv3
|
carnitine deficiency-associated gene expressed in ventricle 3 |
| chr4_-_58553171 | 0.10 |
ENSMUST00000145361.2
|
Lpar1
|
lysophosphatidic acid receptor 1 |
| chr19_+_45991907 | 0.07 |
ENSMUST00000099393.4
|
Hps6
|
HPS6, biogenesis of lysosomal organelles complex 2 subunit 3 |
| chr2_+_129434834 | 0.06 |
ENSMUST00000103203.8
|
Sirpa
|
signal-regulatory protein alpha |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.6 | 22.8 | GO:0043988 | histone H3-S28 phosphorylation(GO:0043988) |
| 3.8 | 18.8 | GO:0071922 | establishment of sister chromatid cohesion(GO:0034085) cohesin loading(GO:0071921) regulation of cohesin loading(GO:0071922) |
| 3.5 | 13.9 | GO:1904008 | regulation of cyclin-dependent protein serine/threonine kinase activity involved in G2/M transition of mitotic cell cycle(GO:0031660) positive regulation of cyclin-dependent protein serine/threonine kinase activity involved in G2/M transition of mitotic cell cycle(GO:0031662) response to monosodium glutamate(GO:1904008) cellular response to monosodium glutamate(GO:1904009) |
| 3.1 | 12.5 | GO:0072355 | histone H3-T3 phosphorylation(GO:0072355) |
| 3.0 | 15.2 | GO:2000620 | positive regulation of histone H4-K16 acetylation(GO:2000620) |
| 2.7 | 8.0 | GO:1905168 | positive regulation of double-strand break repair via homologous recombination(GO:1905168) |
| 2.5 | 15.2 | GO:0006438 | valyl-tRNA aminoacylation(GO:0006438) |
| 2.3 | 9.1 | GO:0006269 | DNA replication, synthesis of RNA primer(GO:0006269) |
| 2.2 | 46.8 | GO:0070986 | left/right axis specification(GO:0070986) |
| 2.1 | 12.3 | GO:0061641 | CENP-A containing nucleosome assembly(GO:0034080) CENP-A containing chromatin organization(GO:0061641) |
| 1.9 | 7.8 | GO:0071477 | hypotonic salinity response(GO:0042539) cellular hypotonic salinity response(GO:0071477) |
| 1.8 | 16.4 | GO:2000042 | negative regulation of double-strand break repair via homologous recombination(GO:2000042) |
| 1.8 | 51.6 | GO:0040034 | regulation of development, heterochronic(GO:0040034) |
| 1.8 | 7.1 | GO:1904430 | mitotic telomere maintenance via semi-conservative replication(GO:1902990) negative regulation of t-circle formation(GO:1904430) |
| 1.6 | 17.9 | GO:0006398 | mRNA 3'-end processing by stem-loop binding and cleavage(GO:0006398) |
| 1.5 | 14.9 | GO:2000348 | regulation of CD40 signaling pathway(GO:2000348) |
| 1.4 | 12.5 | GO:0007144 | female meiosis I(GO:0007144) |
| 1.3 | 11.7 | GO:1900262 | regulation of DNA-directed DNA polymerase activity(GO:1900262) positive regulation of DNA-directed DNA polymerase activity(GO:1900264) |
| 1.3 | 9.0 | GO:0060623 | regulation of chromosome condensation(GO:0060623) |
| 1.2 | 15.7 | GO:0000244 | spliceosomal tri-snRNP complex assembly(GO:0000244) |
| 1.2 | 12.1 | GO:0035608 | protein deglutamylation(GO:0035608) |
| 1.1 | 4.6 | GO:1901740 | negative regulation of myoblast fusion(GO:1901740) |
| 1.1 | 11.4 | GO:1990573 | potassium ion import across plasma membrane(GO:1990573) |
| 1.1 | 11.3 | GO:0014029 | neural crest formation(GO:0014029) |
| 1.1 | 19.4 | GO:0015816 | glycine transport(GO:0015816) |
| 1.1 | 14.9 | GO:2000270 | negative regulation of fibroblast apoptotic process(GO:2000270) |
| 1.0 | 5.0 | GO:0015074 | DNA integration(GO:0015074) |
| 1.0 | 40.5 | GO:0051290 | protein heterotetramerization(GO:0051290) |
| 1.0 | 7.7 | GO:0014043 | negative regulation of neuron maturation(GO:0014043) |
| 0.9 | 43.7 | GO:0008608 | attachment of spindle microtubules to kinetochore(GO:0008608) |
| 0.9 | 19.8 | GO:0010529 | regulation of transposition(GO:0010528) negative regulation of transposition(GO:0010529) |
| 0.9 | 25.4 | GO:0008340 | determination of adult lifespan(GO:0008340) |
| 0.9 | 3.5 | GO:1904565 | response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904565) cellular response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904566) |
| 0.9 | 10.2 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.8 | 5.3 | GO:0090521 | glomerular visceral epithelial cell migration(GO:0090521) |
| 0.7 | 9.8 | GO:0010804 | negative regulation of tumor necrosis factor-mediated signaling pathway(GO:0010804) |
| 0.7 | 3.5 | GO:0006344 | maintenance of chromatin silencing(GO:0006344) |
| 0.6 | 2.2 | GO:0060265 | positive regulation of respiratory burst involved in inflammatory response(GO:0060265) |
| 0.5 | 8.2 | GO:0051152 | positive regulation of smooth muscle cell differentiation(GO:0051152) |
| 0.5 | 19.3 | GO:1902751 | positive regulation of cell cycle G2/M phase transition(GO:1902751) |
| 0.5 | 5.1 | GO:1904628 | response to phorbol 13-acetate 12-myristate(GO:1904627) cellular response to phorbol 13-acetate 12-myristate(GO:1904628) |
| 0.5 | 4.5 | GO:0031086 | nuclear-transcribed mRNA catabolic process, deadenylation-independent decay(GO:0031086) deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
| 0.5 | 1.9 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.5 | 15.4 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.5 | 21.4 | GO:0051310 | metaphase plate congression(GO:0051310) |
| 0.5 | 4.5 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.4 | 21.9 | GO:0006284 | base-excision repair(GO:0006284) |
| 0.4 | 4.6 | GO:0000012 | single strand break repair(GO:0000012) |
| 0.4 | 1.9 | GO:0003349 | epicardium-derived cardiac endothelial cell differentiation(GO:0003349) |
| 0.4 | 4.6 | GO:0035878 | nail development(GO:0035878) |
| 0.4 | 10.4 | GO:0010544 | negative regulation of platelet activation(GO:0010544) |
| 0.4 | 3.7 | GO:0015712 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.4 | 1.1 | GO:1904828 | regulation of hydrogen sulfide biosynthetic process(GO:1904826) positive regulation of hydrogen sulfide biosynthetic process(GO:1904828) |
| 0.3 | 2.3 | GO:0060837 | blood vessel endothelial cell differentiation(GO:0060837) |
| 0.3 | 12.0 | GO:0032012 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.3 | 5.9 | GO:0043568 | positive regulation of insulin-like growth factor receptor signaling pathway(GO:0043568) |
| 0.3 | 0.9 | GO:0019918 | peptidyl-arginine methylation, to symmetrical-dimethyl arginine(GO:0019918) |
| 0.3 | 3.1 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.3 | 3.9 | GO:0051026 | chiasma assembly(GO:0051026) |
| 0.3 | 3.9 | GO:0006241 | CTP biosynthetic process(GO:0006241) CTP metabolic process(GO:0046036) |
| 0.2 | 4.1 | GO:0006465 | signal peptide processing(GO:0006465) |
| 0.2 | 32.4 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.2 | 6.2 | GO:0031297 | replication fork processing(GO:0031297) |
| 0.2 | 2.9 | GO:0051292 | nuclear pore complex assembly(GO:0051292) |
| 0.2 | 1.7 | GO:0045084 | positive regulation of interleukin-12 biosynthetic process(GO:0045084) |
| 0.2 | 3.3 | GO:0034508 | centromere complex assembly(GO:0034508) |
| 0.2 | 3.3 | GO:0071285 | cellular response to lithium ion(GO:0071285) |
| 0.2 | 4.4 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.2 | 28.8 | GO:0007051 | spindle organization(GO:0007051) |
| 0.2 | 8.9 | GO:0070534 | protein K63-linked ubiquitination(GO:0070534) |
| 0.1 | 1.6 | GO:0007016 | cytoskeletal anchoring at plasma membrane(GO:0007016) |
| 0.1 | 1.6 | GO:0003356 | regulation of cilium beat frequency(GO:0003356) |
| 0.1 | 2.7 | GO:0006490 | oligosaccharide-lipid intermediate biosynthetic process(GO:0006490) |
| 0.1 | 6.4 | GO:0019835 | cytolysis(GO:0019835) |
| 0.1 | 3.0 | GO:0061436 | establishment of skin barrier(GO:0061436) |
| 0.1 | 0.5 | GO:0070459 | prolactin secretion(GO:0070459) |
| 0.1 | 1.0 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 0.1 | 1.2 | GO:0061158 | 3'-UTR-mediated mRNA destabilization(GO:0061158) |
| 0.1 | 4.7 | GO:1901099 | negative regulation of signal transduction in absence of ligand(GO:1901099) negative regulation of extrinsic apoptotic signaling pathway in absence of ligand(GO:2001240) |
| 0.1 | 11.0 | GO:0000956 | nuclear-transcribed mRNA catabolic process(GO:0000956) |
| 0.1 | 0.6 | GO:0042985 | negative regulation of amyloid precursor protein biosynthetic process(GO:0042985) |
| 0.1 | 4.2 | GO:0042073 | intraciliary transport(GO:0042073) |
| 0.1 | 0.5 | GO:0048069 | positive regulation of transforming growth factor beta1 production(GO:0032914) eye pigmentation(GO:0048069) |
| 0.1 | 0.2 | GO:1903903 | regulation of establishment of T cell polarity(GO:1903903) |
| 0.1 | 4.9 | GO:0015914 | phospholipid transport(GO:0015914) |
| 0.1 | 1.2 | GO:0048714 | positive regulation of oligodendrocyte differentiation(GO:0048714) |
| 0.1 | 0.2 | GO:0042262 | DNA protection(GO:0042262) |
| 0.1 | 1.9 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.1 | 1.9 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.1 | 0.3 | GO:0090168 | Golgi reassembly(GO:0090168) |
| 0.0 | 4.3 | GO:0071277 | cellular response to calcium ion(GO:0071277) |
| 0.0 | 0.2 | GO:2000053 | regulation of Wnt signaling pathway involved in dorsal/ventral axis specification(GO:2000053) negative regulation of Wnt signaling pathway involved in dorsal/ventral axis specification(GO:2000054) |
| 0.0 | 6.2 | GO:0010212 | response to ionizing radiation(GO:0010212) |
| 0.0 | 0.7 | GO:0021670 | lateral ventricle development(GO:0021670) |
| 0.0 | 23.7 | GO:0051301 | cell division(GO:0051301) |
| 0.0 | 0.3 | GO:0021995 | anterior neuropore closure(GO:0021506) neuropore closure(GO:0021995) |
| 0.0 | 5.7 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.0 | 0.1 | GO:0030422 | production of siRNA involved in RNA interference(GO:0030422) |
| 0.0 | 0.4 | GO:0019884 | antigen processing and presentation of exogenous antigen(GO:0019884) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 6.9 | 41.3 | GO:0031262 | Ndc80 complex(GO:0031262) |
| 3.8 | 22.8 | GO:0032133 | chromosome passenger complex(GO:0032133) |
| 2.4 | 21.2 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 2.2 | 17.9 | GO:0071204 | histone pre-mRNA 3'end processing complex(GO:0071204) |
| 2.2 | 12.9 | GO:0005971 | ribonucleoside-diphosphate reductase complex(GO:0005971) |
| 2.0 | 13.9 | GO:0005638 | lamin filament(GO:0005638) |
| 1.6 | 18.8 | GO:0008278 | cohesin complex(GO:0008278) |
| 1.6 | 6.2 | GO:0035101 | FACT complex(GO:0035101) |
| 1.5 | 21.4 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
| 1.4 | 8.5 | GO:0031523 | Myb complex(GO:0031523) |
| 1.4 | 15.2 | GO:0070531 | BRCA1-A complex(GO:0070531) |
| 1.3 | 9.1 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 1.2 | 11.7 | GO:0031390 | Ctf18 RFC-like complex(GO:0031390) |
| 1.2 | 7.0 | GO:0070876 | SOSS complex(GO:0070876) |
| 1.1 | 3.2 | GO:0071001 | U4/U6 snRNP(GO:0071001) |
| 1.0 | 13.2 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.8 | 10.2 | GO:0042555 | MCM complex(GO:0042555) |
| 0.8 | 5.3 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
| 0.6 | 1.9 | GO:0098843 | postsynaptic endocytic zone(GO:0098843) |
| 0.6 | 12.5 | GO:0001673 | male germ cell nucleus(GO:0001673) |
| 0.5 | 3.7 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.4 | 8.0 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.4 | 4.1 | GO:0005787 | signal peptidase complex(GO:0005787) |
| 0.4 | 48.5 | GO:0005930 | axoneme(GO:0005930) |
| 0.3 | 3.5 | GO:0090571 | RNA polymerase II transcription repressor complex(GO:0090571) |
| 0.2 | 15.4 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.2 | 25.6 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.2 | 12.1 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.2 | 51.4 | GO:0000793 | condensed chromosome(GO:0000793) |
| 0.2 | 1.3 | GO:0070847 | core mediator complex(GO:0070847) |
| 0.1 | 4.6 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.1 | 20.3 | GO:0000922 | spindle pole(GO:0000922) |
| 0.1 | 19.9 | GO:0000775 | chromosome, centromeric region(GO:0000775) |
| 0.1 | 5.2 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.1 | 12.5 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.1 | 1.7 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.1 | 1.9 | GO:0031010 | ISWI-type complex(GO:0031010) |
| 0.1 | 5.2 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.1 | 12.0 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.1 | 11.4 | GO:0030315 | T-tubule(GO:0030315) |
| 0.1 | 4.4 | GO:0016235 | aggresome(GO:0016235) |
| 0.1 | 15.9 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.1 | 0.7 | GO:0071014 | post-mRNA release spliceosomal complex(GO:0071014) |
| 0.1 | 0.9 | GO:0034709 | methylosome(GO:0034709) |
| 0.1 | 4.4 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.1 | 2.4 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.1 | 7.1 | GO:0000781 | chromosome, telomeric region(GO:0000781) |
| 0.1 | 0.4 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.0 | 1.3 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 6.5 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.0 | 4.5 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 0.1 | GO:1903095 | microprocessor complex(GO:0070877) ribonuclease III complex(GO:1903095) |
| 0.0 | 1.6 | GO:0001726 | ruffle(GO:0001726) |
| 0.0 | 7.8 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 13.2 | GO:0000785 | chromatin(GO:0000785) |
| 0.0 | 8.2 | GO:0005635 | nuclear envelope(GO:0005635) |
| 0.0 | 0.3 | GO:0090568 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) nuclear transcriptional repressor complex(GO:0090568) |
| 0.0 | 1.2 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.0 | 0.7 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 4.3 | GO:0005840 | ribosome(GO:0005840) |
| 0.0 | 0.1 | GO:0031084 | BLOC-2 complex(GO:0031084) |
| 0.0 | 124.6 | GO:0005634 | nucleus(GO:0005634) |
| 0.0 | 13.9 | GO:0009986 | cell surface(GO:0009986) |
| 0.0 | 1.3 | GO:0001669 | acrosomal vesicle(GO:0001669) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.5 | 17.9 | GO:0071207 | histone pre-mRNA stem-loop binding(GO:0071207) |
| 4.2 | 12.5 | GO:0035175 | histone kinase activity (H3-S10 specific)(GO:0035175) |
| 3.9 | 27.6 | GO:0005087 | Ran guanyl-nucleotide exchange factor activity(GO:0005087) |
| 3.8 | 11.3 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 3.3 | 19.8 | GO:0047844 | deoxycytidine deaminase activity(GO:0047844) |
| 3.1 | 21.9 | GO:0000405 | bubble DNA binding(GO:0000405) |
| 3.0 | 9.1 | GO:0003896 | DNA primase activity(GO:0003896) |
| 2.5 | 15.2 | GO:0004832 | valine-tRNA ligase activity(GO:0004832) |
| 2.2 | 12.9 | GO:0004748 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 1.9 | 7.8 | GO:0035827 | rubidium ion transmembrane transporter activity(GO:0035827) |
| 1.8 | 14.3 | GO:0036310 | annealing helicase activity(GO:0036310) |
| 1.6 | 11.4 | GO:0086089 | voltage-gated potassium channel activity involved in atrial cardiac muscle cell action potential repolarization(GO:0086089) |
| 1.6 | 11.1 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 1.3 | 17.6 | GO:0015187 | glycine transmembrane transporter activity(GO:0015187) |
| 1.2 | 22.8 | GO:0035174 | histone serine kinase activity(GO:0035174) |
| 1.2 | 3.5 | GO:0035851 | Krueppel-associated box domain binding(GO:0035851) |
| 1.1 | 11.7 | GO:0003689 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 1.0 | 3.9 | GO:0003883 | CTP synthase activity(GO:0003883) |
| 0.9 | 2.7 | GO:0000033 | alpha-1,3-mannosyltransferase activity(GO:0000033) |
| 0.8 | 4.6 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 0.7 | 8.5 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.7 | 22.0 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.5 | 21.4 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.5 | 3.7 | GO:0015119 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 0.5 | 4.5 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) |
| 0.5 | 3.5 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 0.5 | 13.9 | GO:0043274 | phospholipase binding(GO:0043274) |
| 0.5 | 4.6 | GO:0038132 | neuregulin binding(GO:0038132) |
| 0.4 | 12.5 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.4 | 12.1 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.4 | 10.4 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.3 | 15.4 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.3 | 19.3 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.3 | 3.9 | GO:0030983 | mismatched DNA binding(GO:0030983) |
| 0.3 | 1.8 | GO:0015193 | L-proline transmembrane transporter activity(GO:0015193) |
| 0.3 | 10.2 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.3 | 12.0 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.3 | 15.2 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.2 | 1.0 | GO:0004307 | ethanolaminephosphotransferase activity(GO:0004307) |
| 0.2 | 3.2 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.2 | 25.6 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.2 | 4.7 | GO:0017081 | chloride channel regulator activity(GO:0017081) |
| 0.2 | 3.2 | GO:0017070 | U6 snRNA binding(GO:0017070) |
| 0.2 | 0.9 | GO:0035243 | protein-arginine omega-N symmetric methyltransferase activity(GO:0035243) |
| 0.2 | 5.3 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.2 | 8.2 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.2 | 1.3 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.2 | 4.6 | GO:0016891 | endoribonuclease activity, producing 5'-phosphomonoesters(GO:0016891) |
| 0.1 | 4.4 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.1 | 4.9 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.1 | 1.9 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.1 | 12.9 | GO:0001190 | transcriptional activator activity, RNA polymerase II transcription factor binding(GO:0001190) transcriptional repressor activity, RNA polymerase II activating transcription factor binding(GO:0098811) |
| 0.1 | 1.7 | GO:0031996 | thioesterase binding(GO:0031996) |
| 0.1 | 7.7 | GO:0001191 | transcriptional repressor activity, RNA polymerase II transcription factor binding(GO:0001191) |
| 0.1 | 1.6 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.1 | 16.6 | GO:0017048 | Rho GTPase binding(GO:0017048) |
| 0.1 | 8.4 | GO:0005518 | collagen binding(GO:0005518) |
| 0.1 | 2.2 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.1 | 6.8 | GO:0045309 | protein phosphorylated amino acid binding(GO:0045309) |
| 0.1 | 1.9 | GO:0019707 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.1 | 3.3 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.1 | 1.6 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) |
| 0.0 | 0.7 | GO:0004697 | protein kinase C activity(GO:0004697) |
| 0.0 | 8.1 | GO:0042393 | histone binding(GO:0042393) |
| 0.0 | 3.8 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.0 | 0.2 | GO:0032552 | deoxyribonucleotide binding(GO:0032552) |
| 0.0 | 0.2 | GO:0070411 | I-SMAD binding(GO:0070411) |
| 0.0 | 1.3 | GO:0001104 | RNA polymerase II transcription cofactor activity(GO:0001104) |
| 0.0 | 1.2 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 0.0 | 5.6 | GO:0005057 | receptor signaling protein activity(GO:0005057) |
| 0.0 | 0.4 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 0.0 | 1.1 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.0 | 5.0 | GO:0047485 | protein N-terminus binding(GO:0047485) |
| 0.0 | 7.2 | GO:0008236 | serine-type peptidase activity(GO:0008236) |
| 0.0 | 4.7 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 10.2 | GO:0003682 | chromatin binding(GO:0003682) |
| 0.0 | 0.5 | GO:0016780 | phosphotransferase activity, for other substituted phosphate groups(GO:0016780) |
| 0.0 | 0.6 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 32.4 | GO:0042802 | identical protein binding(GO:0042802) |
| 0.0 | 0.5 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 2.4 | GO:0003729 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 49.4 | PID ATM PATHWAY | ATM pathway |
| 1.1 | 60.6 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.3 | 8.0 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.3 | 4.6 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.3 | 22.3 | PID E2F PATHWAY | E2F transcription factor network |
| 0.2 | 10.2 | PID ATR PATHWAY | ATR signaling pathway |
| 0.2 | 27.6 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.2 | 11.4 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.2 | 6.2 | ST GAQ PATHWAY | G alpha q Pathway |
| 0.2 | 4.4 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.2 | 13.9 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.1 | 4.6 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 0.1 | 4.3 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.1 | 9.9 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.1 | 4.0 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.1 | 1.6 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.1 | 1.9 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.1 | 3.7 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.1 | 3.5 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.1 | 3.4 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.0 | 6.1 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.7 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.0 | 1.3 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.0 | 0.2 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.0 | 0.5 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.7 | 14.9 | REACTOME REGULATION OF THE FANCONI ANEMIA PATHWAY | Genes involved in Regulation of the Fanconi anemia pathway |
| 1.5 | 23.2 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 1.5 | 19.3 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 1.4 | 17.9 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 1.2 | 28.4 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 1.2 | 25.7 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.9 | 9.1 | REACTOME INHIBITION OF REPLICATION INITIATION OF DAMAGED DNA BY RB1 E2F1 | Genes involved in Inhibition of replication initiation of damaged DNA by RB1/E2F1 |
| 0.8 | 17.5 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.7 | 77.3 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.7 | 10.2 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.7 | 5.0 | REACTOME APOBEC3G MEDIATED RESISTANCE TO HIV1 INFECTION | Genes involved in APOBEC3G mediated resistance to HIV-1 infection |
| 0.6 | 10.4 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.5 | 15.2 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.5 | 8.5 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.3 | 32.8 | REACTOME LATE PHASE OF HIV LIFE CYCLE | Genes involved in Late Phase of HIV Life Cycle |
| 0.3 | 11.8 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.3 | 17.6 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.2 | 6.8 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 0.2 | 11.4 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.2 | 4.1 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.2 | 4.6 | REACTOME DOUBLE STRAND BREAK REPAIR | Genes involved in Double-Strand Break Repair |
| 0.2 | 1.7 | REACTOME NFKB IS ACTIVATED AND SIGNALS SURVIVAL | Genes involved in NF-kB is activated and signals survival |
| 0.1 | 3.9 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.1 | 3.9 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.1 | 5.9 | REACTOME INTEGRIN ALPHAIIB BETA3 SIGNALING | Genes involved in Integrin alphaIIb beta3 signaling |
| 0.1 | 3.3 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.1 | 6.9 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.1 | 1.9 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.1 | 4.5 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 1.3 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.1 | 4.9 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.1 | 4.6 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.1 | 7.8 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.1 | 8.2 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.1 | 2.4 | REACTOME TRANSPORT OF MATURE TRANSCRIPT TO CYTOPLASM | Genes involved in Transport of Mature Transcript to Cytoplasm |
| 0.1 | 1.2 | REACTOME DESTABILIZATION OF MRNA BY KSRP | Genes involved in Destabilization of mRNA by KSRP |
| 0.0 | 0.9 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.0 | 1.1 | REACTOME SMAD2 SMAD3 SMAD4 HETEROTRIMER REGULATES TRANSCRIPTION | Genes involved in SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription |
| 0.0 | 3.5 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.0 | 3.5 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.0 | 1.1 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.0 | 0.5 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.0 | 1.3 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 0.1 | REACTOME MICRORNA MIRNA BIOGENESIS | Genes involved in MicroRNA (miRNA) Biogenesis |