Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
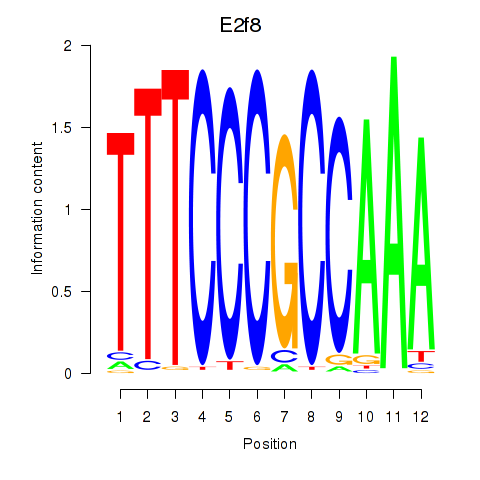
Results for E2f8
Z-value: 1.48
Transcription factors associated with E2f8
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
E2f8
|
ENSMUSG00000046179.18 | E2f8 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| E2f8 | mm39_v1_chr7_-_48531344_48531367 | 0.85 | 5.3e-21 | Click! |
Activity profile of E2f8 motif
Sorted Z-values of E2f8 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of E2f8
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr10_-_21036792 | 21.89 |
ENSMUST00000188495.8
|
Myb
|
myeloblastosis oncogene |
| chr2_+_162896602 | 21.23 |
ENSMUST00000018005.10
|
Mybl2
|
myeloblastosis oncogene-like 2 |
| chr10_-_21036824 | 19.97 |
ENSMUST00000020158.9
|
Myb
|
myeloblastosis oncogene |
| chr17_+_56610321 | 16.40 |
ENSMUST00000001258.15
|
Uhrf1
|
ubiquitin-like, containing PHD and RING finger domains, 1 |
| chr8_+_75836187 | 16.36 |
ENSMUST00000164309.3
ENSMUST00000212426.2 ENSMUST00000212811.2 |
Mcm5
|
minichromosome maintenance complex component 5 |
| chr17_+_56610396 | 15.16 |
ENSMUST00000113038.8
|
Uhrf1
|
ubiquitin-like, containing PHD and RING finger domains, 1 |
| chr3_-_90603013 | 13.93 |
ENSMUST00000069960.12
ENSMUST00000117167.2 |
S100a9
|
S100 calcium binding protein A9 (calgranulin B) |
| chr15_+_61857390 | 12.07 |
ENSMUST00000159327.2
ENSMUST00000167731.8 |
Myc
|
myelocytomatosis oncogene |
| chr15_+_61857226 | 11.93 |
ENSMUST00000161976.8
ENSMUST00000022971.8 |
Myc
|
myelocytomatosis oncogene |
| chr8_+_57964956 | 11.62 |
ENSMUST00000210871.2
|
Hmgb2
|
high mobility group box 2 |
| chr19_+_38919353 | 11.47 |
ENSMUST00000025965.12
|
Hells
|
helicase, lymphoid specific |
| chr12_+_24758724 | 10.81 |
ENSMUST00000153058.8
|
Rrm2
|
ribonucleotide reductase M2 |
| chr1_-_128287347 | 10.77 |
ENSMUST00000190495.2
ENSMUST00000027601.11 |
Mcm6
|
minichromosome maintenance complex component 6 |
| chr6_-_88875646 | 10.36 |
ENSMUST00000058011.8
|
Mcm2
|
minichromosome maintenance complex component 2 |
| chr5_+_123887759 | 10.12 |
ENSMUST00000031366.12
|
Kntc1
|
kinetochore associated 1 |
| chr19_+_38919477 | 10.05 |
ENSMUST00000145051.2
|
Hells
|
helicase, lymphoid specific |
| chr12_+_24758968 | 9.95 |
ENSMUST00000154588.2
|
Rrm2
|
ribonucleotide reductase M2 |
| chr10_+_127851031 | 9.29 |
ENSMUST00000178041.8
ENSMUST00000026461.8 |
Prim1
|
DNA primase, p49 subunit |
| chr7_-_44198157 | 9.28 |
ENSMUST00000145956.2
ENSMUST00000049343.15 |
Pold1
|
polymerase (DNA directed), delta 1, catalytic subunit |
| chr9_-_36637670 | 9.20 |
ENSMUST00000172702.9
ENSMUST00000172742.2 |
Chek1
|
checkpoint kinase 1 |
| chr9_-_36637923 | 8.98 |
ENSMUST00000034625.12
|
Chek1
|
checkpoint kinase 1 |
| chr16_-_18630365 | 8.41 |
ENSMUST00000096990.10
|
Cdc45
|
cell division cycle 45 |
| chr7_-_48530777 | 7.78 |
ENSMUST00000058745.15
|
E2f8
|
E2F transcription factor 8 |
| chr7_-_48531344 | 7.63 |
ENSMUST00000119223.2
|
E2f8
|
E2F transcription factor 8 |
| chr5_-_33809640 | 7.57 |
ENSMUST00000151081.2
ENSMUST00000139518.8 ENSMUST00000101354.10 |
Slbp
|
stem-loop binding protein |
| chr5_-_33809872 | 6.87 |
ENSMUST00000057551.14
|
Slbp
|
stem-loop binding protein |
| chr17_+_56347424 | 6.82 |
ENSMUST00000002914.10
|
Chaf1a
|
chromatin assembly factor 1, subunit A (p150) |
| chr2_-_34803988 | 6.55 |
ENSMUST00000028232.7
ENSMUST00000202907.2 |
Phf19
|
PHD finger protein 19 |
| chr17_+_88282472 | 6.00 |
ENSMUST00000005503.5
|
Msh6
|
mutS homolog 6 |
| chr5_-_33809683 | 5.92 |
ENSMUST00000075670.13
|
Slbp
|
stem-loop binding protein |
| chr16_-_15455141 | 5.86 |
ENSMUST00000023353.4
|
Mcm4
|
minichromosome maintenance complex component 4 |
| chr6_-_125262974 | 5.82 |
ENSMUST00000088246.6
|
Tuba3a
|
tubulin, alpha 3A |
| chr10_+_42736345 | 5.73 |
ENSMUST00000063063.14
|
Scml4
|
Scm polycomb group protein like 4 |
| chr10_-_128361731 | 5.58 |
ENSMUST00000026427.8
|
Esyt1
|
extended synaptotagmin-like protein 1 |
| chr16_+_10652910 | 5.18 |
ENSMUST00000037913.9
|
Rmi2
|
RecQ mediated genome instability 2 |
| chr2_-_157046386 | 5.12 |
ENSMUST00000029170.8
|
Rbl1
|
RB transcriptional corepressor like 1 |
| chr5_-_65492907 | 4.92 |
ENSMUST00000203581.3
|
Rfc1
|
replication factor C (activator 1) 1 |
| chr1_-_93729650 | 4.80 |
ENSMUST00000027503.14
|
Dtymk
|
deoxythymidylate kinase |
| chr6_+_35154319 | 4.72 |
ENSMUST00000201374.4
ENSMUST00000043815.16 |
Nup205
|
nucleoporin 205 |
| chr17_+_34596098 | 4.56 |
ENSMUST00000080254.7
|
Btnl1
|
butyrophilin-like 1 |
| chr5_-_65492940 | 4.45 |
ENSMUST00000203471.3
ENSMUST00000172732.8 ENSMUST00000204965.3 |
Rfc1
|
replication factor C (activator 1) 1 |
| chr7_+_101714943 | 4.35 |
ENSMUST00000094130.4
ENSMUST00000084843.10 |
Xndc1
Xntrpc
|
Xrcc1 N-terminal domain containing 1 Xndc1-transient receptor potential cation channel, subfamily C, member 2 readthrough |
| chr2_-_154411765 | 4.16 |
ENSMUST00000103145.11
|
E2f1
|
E2F transcription factor 1 |
| chr13_+_19807274 | 3.84 |
ENSMUST00000222464.2
ENSMUST00000002883.7 |
Sfrp4
|
secreted frizzled-related protein 4 |
| chr6_+_35154545 | 3.82 |
ENSMUST00000170234.2
|
Nup205
|
nucleoporin 205 |
| chr15_+_78810919 | 3.78 |
ENSMUST00000089377.6
|
Lgals1
|
lectin, galactose binding, soluble 1 |
| chr2_-_154411640 | 3.47 |
ENSMUST00000000894.6
|
E2f1
|
E2F transcription factor 1 |
| chr13_+_44884740 | 3.44 |
ENSMUST00000173246.8
|
Jarid2
|
jumonji, AT rich interactive domain 2 |
| chr16_-_18066591 | 3.42 |
ENSMUST00000115645.10
|
Ranbp1
|
RAN binding protein 1 |
| chr1_+_131838294 | 3.33 |
ENSMUST00000062264.8
|
Nucks1
|
nuclear casein kinase and cyclin-dependent kinase substrate 1 |
| chr11_+_79883885 | 3.25 |
ENSMUST00000163272.2
ENSMUST00000017692.15 |
Suz12
|
SUZ12 polycomb repressive complex 2 subunit |
| chr19_+_4806544 | 3.15 |
ENSMUST00000182821.8
ENSMUST00000036744.8 |
Rbm4b
|
RNA binding motif protein 4B |
| chr8_-_107792264 | 2.96 |
ENSMUST00000034393.7
|
Tmed6
|
transmembrane p24 trafficking protein 6 |
| chr9_-_35028100 | 2.86 |
ENSMUST00000034537.8
|
St3gal4
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 4 |
| chr15_-_97991114 | 2.86 |
ENSMUST00000180657.2
|
Senp1
|
SUMO1/sentrin specific peptidase 1 |
| chr7_+_101714692 | 2.74 |
ENSMUST00000106950.8
ENSMUST00000146450.8 |
Xndc1
|
Xrcc1 N-terminal domain containing 1 |
| chr4_+_132495636 | 2.47 |
ENSMUST00000102561.11
|
Rpa2
|
replication protein A2 |
| chr6_+_117894242 | 2.42 |
ENSMUST00000180020.8
ENSMUST00000177570.2 |
Hnrnpf
|
heterogeneous nuclear ribonucleoprotein F |
| chr1_+_131838220 | 2.19 |
ENSMUST00000189946.7
|
Nucks1
|
nuclear casein kinase and cyclin-dependent kinase substrate 1 |
| chr19_+_4806859 | 2.13 |
ENSMUST00000237921.2
|
Rbm4b
|
RNA binding motif protein 4B |
| chr4_-_126861918 | 2.02 |
ENSMUST00000106108.9
|
Zmym4
|
zinc finger, MYM-type 4 |
| chr2_-_45002902 | 2.00 |
ENSMUST00000076836.13
ENSMUST00000176732.8 ENSMUST00000200844.4 |
Zeb2
|
zinc finger E-box binding homeobox 2 |
| chr2_-_155787197 | 1.92 |
ENSMUST00000040162.3
|
Gdf5
|
growth differentiation factor 5 |
| chr15_+_55420795 | 1.88 |
ENSMUST00000022998.14
|
Mtbp
|
Mdm2, transformed 3T3 cell double minute p53 binding protein |
| chr6_+_117893942 | 1.78 |
ENSMUST00000179478.8
|
Hnrnpf
|
heterogeneous nuclear ribonucleoprotein F |
| chr17_+_34129221 | 1.74 |
ENSMUST00000174541.9
|
Daxx
|
Fas death domain-associated protein |
| chr5_+_137786031 | 1.73 |
ENSMUST00000035852.14
|
Zcwpw1
|
zinc finger, CW type with PWWP domain 1 |
| chr17_-_35454729 | 1.72 |
ENSMUST00000048994.7
|
Nfkbil1
|
nuclear factor of kappa light polypeptide gene enhancer in B cells inhibitor like 1 |
| chr5_+_75312939 | 1.70 |
ENSMUST00000202681.4
ENSMUST00000000476.15 |
Pdgfra
|
platelet derived growth factor receptor, alpha polypeptide |
| chrX_-_41000746 | 1.70 |
ENSMUST00000047037.15
|
Thoc2
|
THO complex 2 |
| chr11_+_87938519 | 1.59 |
ENSMUST00000079866.11
|
Srsf1
|
serine and arginine-rich splicing factor 1 |
| chr2_-_45003270 | 1.53 |
ENSMUST00000202935.4
ENSMUST00000068415.11 ENSMUST00000127520.8 |
Zeb2
|
zinc finger E-box binding homeobox 2 |
| chr2_-_91067212 | 1.43 |
ENSMUST00000111352.8
|
Ddb2
|
damage specific DNA binding protein 2 |
| chr9_+_21914296 | 1.42 |
ENSMUST00000003493.9
|
Prkcsh
|
protein kinase C substrate 80K-H |
| chr13_+_44884757 | 1.41 |
ENSMUST00000174086.2
|
Jarid2
|
jumonji, AT rich interactive domain 2 |
| chr10_+_42736539 | 1.39 |
ENSMUST00000157071.8
|
Scml4
|
Scm polycomb group protein like 4 |
| chr17_+_34128382 | 1.39 |
ENSMUST00000173028.8
ENSMUST00000079421.15 |
Daxx
|
Fas death domain-associated protein |
| chr9_+_21914334 | 1.39 |
ENSMUST00000115331.10
|
Prkcsh
|
protein kinase C substrate 80K-H |
| chr7_-_128342158 | 1.38 |
ENSMUST00000119081.2
ENSMUST00000057557.14 |
Mcmbp
|
minichromosome maintenance complex binding protein |
| chr4_+_63133639 | 1.32 |
ENSMUST00000036300.13
|
Col27a1
|
collagen, type XXVII, alpha 1 |
| chr9_+_21914083 | 1.26 |
ENSMUST00000216344.2
|
Prkcsh
|
protein kinase C substrate 80K-H |
| chr6_+_125016723 | 1.21 |
ENSMUST00000140131.8
ENSMUST00000032480.14 |
Ing4
|
inhibitor of growth family, member 4 |
| chr10_+_19232281 | 1.17 |
ENSMUST00000053225.7
|
Olig3
|
oligodendrocyte transcription factor 3 |
| chr6_+_116241146 | 1.17 |
ENSMUST00000112900.9
ENSMUST00000036503.14 ENSMUST00000223495.2 |
Zfand4
|
zinc finger, AN1-type domain 4 |
| chr17_+_34128455 | 1.13 |
ENSMUST00000173626.8
ENSMUST00000170075.9 |
Daxx
|
Fas death domain-associated protein |
| chr17_+_29020744 | 1.08 |
ENSMUST00000136233.2
|
Brpf3
|
bromodomain and PHD finger containing, 3 |
| chr2_-_91067312 | 1.00 |
ENSMUST00000028696.5
|
Ddb2
|
damage specific DNA binding protein 2 |
| chr16_-_45352346 | 0.94 |
ENSMUST00000232600.2
|
Gm17783
|
predicted gene, 17783 |
| chr9_+_100525501 | 0.90 |
ENSMUST00000146312.8
ENSMUST00000129269.8 |
Stag1
|
stromal antigen 1 |
| chr6_+_38528738 | 0.89 |
ENSMUST00000161227.8
|
Luc7l2
|
LUC7-like 2 (S. cerevisiae) |
| chr11_+_26337194 | 0.83 |
ENSMUST00000136830.2
ENSMUST00000109509.8 |
Fancl
|
Fanconi anemia, complementation group L |
| chr13_+_95012107 | 0.79 |
ENSMUST00000022195.13
|
Otp
|
orthopedia homeobox |
| chr5_-_137785903 | 0.67 |
ENSMUST00000196022.2
|
Mepce
|
methylphosphate capping enzyme |
| chr15_+_101071948 | 0.39 |
ENSMUST00000000544.12
|
Acvr1b
|
activin A receptor, type 1B |
| chrX_-_167970196 | 0.23 |
ENSMUST00000066112.12
ENSMUST00000112118.8 ENSMUST00000112120.8 ENSMUST00000112119.8 |
Amelx
|
amelogenin, X-linked |
| chr9_+_21914513 | 0.22 |
ENSMUST00000215795.2
|
Prkcsh
|
protein kinase C substrate 80K-H |
| chr12_-_55045887 | 0.12 |
ENSMUST00000173529.2
|
Baz1a
|
bromodomain adjacent to zinc finger domain 1A |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 6.1 | 18.2 | GO:0010767 | regulation of transcription from RNA polymerase II promoter in response to UV-induced DNA damage(GO:0010767) |
| 6.0 | 24.0 | GO:0090096 | lactic acid secretion(GO:0046722) regulation of metanephric cap mesenchymal cell proliferation(GO:0090095) positive regulation of metanephric cap mesenchymal cell proliferation(GO:0090096) |
| 5.1 | 15.4 | GO:0032877 | positive regulation of DNA endoreduplication(GO:0032877) |
| 4.6 | 13.9 | GO:0035606 | peptidyl-cysteine S-trans-nitrosylation(GO:0035606) |
| 3.1 | 46.7 | GO:0051574 | positive regulation of histone H3-K9 methylation(GO:0051574) |
| 3.1 | 9.3 | GO:0045004 | DNA replication proofreading(GO:0045004) |
| 2.9 | 53.1 | GO:0010216 | maintenance of DNA methylation(GO:0010216) |
| 2.8 | 8.4 | GO:0036388 | pre-replicative complex assembly involved in nuclear cell cycle DNA replication(GO:0006267) pre-replicative complex assembly(GO:0036388) DNA replication preinitiation complex assembly(GO:0071163) pre-replicative complex assembly involved in cell cycle DNA replication(GO:1902299) |
| 2.5 | 7.6 | GO:0071930 | negative regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0071930) |
| 2.3 | 9.3 | GO:0006269 | DNA replication, synthesis of RNA primer(GO:0006269) |
| 2.2 | 27.0 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 1.9 | 20.4 | GO:0006398 | mRNA 3'-end processing by stem-loop binding and cleavage(GO:0006398) |
| 1.6 | 4.8 | GO:0006233 | dTDP biosynthetic process(GO:0006233) dTDP metabolic process(GO:0046072) |
| 1.4 | 5.5 | GO:0060382 | release from viral latency(GO:0019046) regulation of DNA strand elongation(GO:0060382) |
| 1.3 | 3.8 | GO:2000118 | regulation of sodium-dependent phosphate transport(GO:2000118) |
| 1.3 | 3.8 | GO:0034117 | erythrocyte aggregation(GO:0034117) regulation of erythrocyte aggregation(GO:0034118) |
| 1.2 | 11.6 | GO:0045654 | positive regulation of megakaryocyte differentiation(GO:0045654) |
| 1.0 | 20.8 | GO:0009263 | deoxyribonucleotide biosynthetic process(GO:0009263) |
| 1.0 | 6.0 | GO:0051096 | regulation of helicase activity(GO:0051095) positive regulation of helicase activity(GO:0051096) |
| 0.9 | 3.4 | GO:0046604 | positive regulation of mitotic centrosome separation(GO:0046604) |
| 0.9 | 4.3 | GO:0072738 | response to diamide(GO:0072737) cellular response to diamide(GO:0072738) |
| 0.8 | 4.6 | GO:0045062 | extrathymic T cell selection(GO:0045062) |
| 0.7 | 6.6 | GO:0061087 | positive regulation of histone H3-K27 methylation(GO:0061087) |
| 0.6 | 6.8 | GO:0006335 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.5 | 16.4 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.5 | 3.5 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.5 | 8.5 | GO:0051292 | nuclear pore complex assembly(GO:0051292) |
| 0.5 | 3.2 | GO:0098532 | histone H3-K27 trimethylation(GO:0098532) |
| 0.4 | 1.7 | GO:0072276 | metanephric glomerulus morphogenesis(GO:0072275) metanephric glomerulus vasculature morphogenesis(GO:0072276) metanephric glomerular capillary formation(GO:0072277) |
| 0.3 | 1.7 | GO:0007089 | traversing start control point of mitotic cell cycle(GO:0007089) |
| 0.3 | 2.4 | GO:0006290 | pyrimidine dimer repair(GO:0006290) |
| 0.3 | 1.9 | GO:0060591 | chondroblast differentiation(GO:0060591) |
| 0.3 | 21.2 | GO:0090307 | mitotic spindle assembly(GO:0090307) |
| 0.2 | 1.2 | GO:0097476 | spinal cord motor neuron migration(GO:0097476) |
| 0.2 | 2.9 | GO:0010724 | regulation of definitive erythrocyte differentiation(GO:0010724) |
| 0.2 | 1.7 | GO:0010793 | regulation of mRNA export from nucleus(GO:0010793) |
| 0.2 | 1.3 | GO:0003431 | growth plate cartilage chondrocyte development(GO:0003431) |
| 0.2 | 5.3 | GO:0043153 | entrainment of circadian clock by photoperiod(GO:0043153) |
| 0.2 | 2.5 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.1 | 4.3 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.1 | 1.7 | GO:0031665 | negative regulation of lipopolysaccharide-mediated signaling pathway(GO:0031665) |
| 0.1 | 5.2 | GO:0033045 | regulation of sister chromatid segregation(GO:0033045) |
| 0.1 | 5.1 | GO:0043550 | regulation of lipid kinase activity(GO:0043550) |
| 0.1 | 0.4 | GO:1901165 | positive regulation of trophoblast cell migration(GO:1901165) |
| 0.1 | 1.2 | GO:0043983 | histone H4-K12 acetylation(GO:0043983) |
| 0.1 | 0.7 | GO:0040031 | snRNA modification(GO:0040031) |
| 0.1 | 9.3 | GO:0007093 | mitotic cell cycle checkpoint(GO:0007093) |
| 0.1 | 0.8 | GO:0021979 | hypothalamus cell differentiation(GO:0021979) |
| 0.0 | 2.9 | GO:0018279 | protein N-linked glycosylation via asparagine(GO:0018279) |
| 0.0 | 7.5 | GO:0006260 | DNA replication(GO:0006260) |
| 0.0 | 1.1 | GO:0043966 | histone H3 acetylation(GO:0043966) |
| 0.0 | 5.6 | GO:0006869 | lipid transport(GO:0006869) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.5 | 21.2 | GO:0031523 | Myb complex(GO:0031523) |
| 3.5 | 20.8 | GO:0005971 | ribonucleoside-diphosphate reductase complex(GO:0005971) |
| 3.4 | 10.1 | GO:1990423 | RZZ complex(GO:1990423) |
| 2.8 | 8.4 | GO:0036387 | nuclear pre-replicative complex(GO:0005656) pre-replicative complex(GO:0036387) |
| 2.8 | 44.7 | GO:0042555 | MCM complex(GO:0042555) |
| 2.5 | 20.4 | GO:0071204 | histone pre-mRNA 3'end processing complex(GO:0071204) |
| 1.9 | 9.3 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 1.7 | 6.8 | GO:0033186 | CAF-1 complex(GO:0033186) |
| 1.6 | 9.4 | GO:0005663 | DNA replication factor C complex(GO:0005663) |
| 1.5 | 7.6 | GO:0035189 | Rb-E2F complex(GO:0035189) |
| 1.3 | 9.3 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 1.2 | 8.5 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
| 0.9 | 4.3 | GO:0017177 | glucosidase II complex(GO:0017177) |
| 0.8 | 52.2 | GO:0005657 | replication fork(GO:0005657) |
| 0.7 | 6.0 | GO:0032300 | mismatch repair complex(GO:0032300) |
| 0.6 | 21.5 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.5 | 14.7 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.5 | 24.0 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.3 | 2.4 | GO:0031465 | Cul4B-RING E3 ubiquitin ligase complex(GO:0031465) |
| 0.3 | 5.6 | GO:0044232 | organelle membrane contact site(GO:0044232) |
| 0.2 | 1.7 | GO:0000347 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.2 | 3.4 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.1 | 0.4 | GO:0048179 | activin receptor complex(GO:0048179) |
| 0.1 | 1.1 | GO:0070776 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.1 | 4.3 | GO:0001741 | XY body(GO:0001741) |
| 0.1 | 1.3 | GO:0005583 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 0.0 | 0.8 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.0 | 0.9 | GO:0071004 | U2-type prespliceosome(GO:0071004) |
| 0.0 | 16.3 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 5.8 | GO:0036064 | ciliary basal body(GO:0036064) |
| 0.0 | 0.1 | GO:0008623 | CHRAC(GO:0008623) |
| 0.0 | 12.8 | GO:0000790 | nuclear chromatin(GO:0000790) |
| 0.0 | 3.3 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.0 | 2.2 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.0 | 1.2 | GO:0000123 | histone acetyltransferase complex(GO:0000123) |
| 0.0 | 12.0 | GO:0031012 | extracellular matrix(GO:0031012) |
| 0.0 | 1.7 | GO:0005902 | microvillus(GO:0005902) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 8.4 | 41.9 | GO:0071987 | WD40-repeat domain binding(GO:0071987) |
| 5.1 | 20.4 | GO:0071207 | histone pre-mRNA stem-loop binding(GO:0071207) |
| 4.5 | 18.2 | GO:0035402 | histone kinase activity (H3-T11 specific)(GO:0035402) |
| 4.5 | 31.6 | GO:0044729 | hemi-methylated DNA-binding(GO:0044729) |
| 3.5 | 20.8 | GO:0061731 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 3.1 | 9.3 | GO:0003896 | DNA primase activity(GO:0003896) |
| 2.6 | 25.5 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 2.4 | 21.2 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 2.0 | 6.0 | GO:0032142 | single guanine insertion binding(GO:0032142) |
| 2.0 | 35.1 | GO:0003688 | DNA replication origin binding(GO:0003688) |
| 1.6 | 4.8 | GO:0004798 | thymidylate kinase activity(GO:0004798) |
| 0.9 | 3.8 | GO:0048030 | disaccharide binding(GO:0048030) |
| 0.9 | 9.4 | GO:0003689 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.7 | 2.9 | GO:0070139 | ubiquitin-like protein-specific endopeptidase activity(GO:0070137) SUMO-specific endopeptidase activity(GO:0070139) |
| 0.6 | 6.8 | GO:0070087 | chromo shadow domain binding(GO:0070087) |
| 0.6 | 1.7 | GO:0005017 | platelet-derived growth factor-activated receptor activity(GO:0005017) |
| 0.5 | 9.3 | GO:0008296 | 3'-5'-exodeoxyribonuclease activity(GO:0008296) |
| 0.4 | 2.9 | GO:0047288 | monosialoganglioside sialyltransferase activity(GO:0047288) |
| 0.4 | 16.6 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.3 | 3.2 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.3 | 24.0 | GO:0070888 | E-box binding(GO:0070888) |
| 0.3 | 8.5 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.2 | 1.9 | GO:0036122 | BMP binding(GO:0036122) |
| 0.2 | 2.5 | GO:0098505 | G-rich strand telomeric DNA binding(GO:0098505) |
| 0.2 | 3.8 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.1 | 5.1 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.1 | 21.5 | GO:0004386 | helicase activity(GO:0004386) |
| 0.1 | 4.3 | GO:0008432 | JUN kinase binding(GO:0008432) |
| 0.1 | 3.4 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.1 | 4.9 | GO:0032452 | histone demethylase activity(GO:0032452) |
| 0.1 | 4.2 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.1 | 7.8 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.1 | 3.5 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.1 | 23.0 | GO:0001047 | core promoter binding(GO:0001047) |
| 0.1 | 0.2 | GO:0030021 | extracellular matrix structural constituent conferring compression resistance(GO:0030021) structural constituent of tooth enamel(GO:0030345) |
| 0.1 | 0.4 | GO:0016361 | activin receptor activity, type I(GO:0016361) |
| 0.1 | 2.4 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.0 | 4.2 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.0 | 1.2 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.0 | 1.3 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 4.3 | GO:0051219 | phosphoprotein binding(GO:0051219) |
| 0.0 | 10.2 | GO:0008270 | zinc ion binding(GO:0008270) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.7 | 16.8 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 1.0 | 73.5 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.5 | 50.5 | PID E2F PATHWAY | E2F transcription factor network |
| 0.4 | 13.9 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.3 | 21.5 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.3 | 12.8 | PID ATR PATHWAY | ATR signaling pathway |
| 0.2 | 9.4 | PID FAS PATHWAY | FAS (CD95) signaling pathway |
| 0.2 | 4.3 | PID HIV NEF PATHWAY | HIV-1 Nef: Negative effector of Fas and TNF-alpha |
| 0.1 | 3.8 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.1 | 6.5 | ST T CELL SIGNAL TRANSDUCTION | T Cell Signal Transduction |
| 0.1 | 1.7 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.1 | 5.9 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 2.9 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 0.8 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.0 | 3.8 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 1.3 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 2.4 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.2 | 51.8 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 1.6 | 20.4 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 1.4 | 16.8 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 1.3 | 28.4 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 1.2 | 18.6 | REACTOME POL SWITCHING | Genes involved in Polymerase switching |
| 1.0 | 19.9 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.9 | 11.6 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.6 | 24.0 | REACTOME SMAD2 SMAD3 SMAD4 HETEROTRIMER REGULATES TRANSCRIPTION | Genes involved in SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription |
| 0.5 | 2.5 | REACTOME REMOVAL OF THE FLAP INTERMEDIATE FROM THE C STRAND | Genes involved in Removal of the Flap Intermediate from the C-strand |
| 0.3 | 4.3 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 0.2 | 8.5 | REACTOME TRANSPORT OF RIBONUCLEOPROTEINS INTO THE HOST NUCLEUS | Genes involved in Transport of Ribonucleoproteins into the Host Nucleus |
| 0.2 | 35.9 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.2 | 4.8 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.1 | 2.4 | REACTOME FORMATION OF INCISION COMPLEX IN GG NER | Genes involved in Formation of incision complex in GG-NER |
| 0.1 | 6.6 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.0 | 3.4 | REACTOME LATE PHASE OF HIV LIFE CYCLE | Genes involved in Late Phase of HIV Life Cycle |
| 0.0 | 0.8 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.0 | 3.3 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.0 | 1.3 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.4 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.0 | 1.7 | REACTOME DOWNSTREAM SIGNAL TRANSDUCTION | Genes involved in Downstream signal transduction |