Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
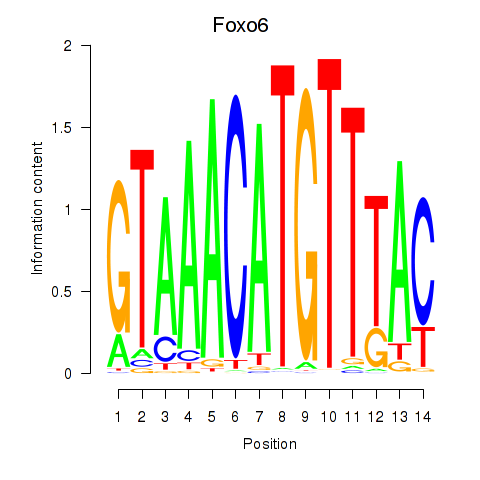
Results for Foxo6
Z-value: 0.71
Transcription factors associated with Foxo6
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Foxo6
|
ENSMUSG00000052135.9 | Foxo6 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Foxo6 | mm39_v1_chr4_-_120144546_120144546 | -0.52 | 3.0e-06 | Click! |
Activity profile of Foxo6 motif
Sorted Z-values of Foxo6 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Foxo6
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr15_-_74983786 | 5.02 |
ENSMUST00000191451.2
ENSMUST00000100542.10 |
Ly6c2
|
lymphocyte antigen 6 complex, locus C2 |
| chr1_-_162726234 | 4.62 |
ENSMUST00000111510.8
ENSMUST00000045902.13 |
Fmo2
|
flavin containing monooxygenase 2 |
| chr6_-_129449739 | 4.49 |
ENSMUST00000112076.9
ENSMUST00000184581.3 |
Clec7a
|
C-type lectin domain family 7, member a |
| chr1_-_162726053 | 3.92 |
ENSMUST00000143123.3
|
Fmo2
|
flavin containing monooxygenase 2 |
| chr15_-_74855264 | 3.67 |
ENSMUST00000023250.11
ENSMUST00000166694.2 |
Ly6i
|
lymphocyte antigen 6 complex, locus I |
| chr7_+_25597045 | 3.29 |
ENSMUST00000072438.13
ENSMUST00000005477.6 |
Cyp2b10
|
cytochrome P450, family 2, subfamily b, polypeptide 10 |
| chr12_-_113223839 | 3.27 |
ENSMUST00000194738.6
ENSMUST00000178282.3 |
Igha
|
immunoglobulin heavy constant alpha |
| chr6_+_122929410 | 3.09 |
ENSMUST00000117173.8
|
Clec4a3
|
C-type lectin domain family 4, member a3 |
| chr7_+_43874854 | 2.85 |
ENSMUST00000206144.2
|
Klk1
|
kallikrein 1 |
| chr16_-_88360037 | 2.79 |
ENSMUST00000049697.5
|
Cldn8
|
claudin 8 |
| chr17_-_31496301 | 2.54 |
ENSMUST00000235144.2
|
Rsph1
|
radial spoke head 1 homolog (Chlamydomonas) |
| chr11_+_87684299 | 2.52 |
ENSMUST00000020779.11
|
Mpo
|
myeloperoxidase |
| chr6_+_67532481 | 2.49 |
ENSMUST00000103302.3
|
Igkv2-137
|
immunoglobulin kappa chain variable 2-137 |
| chr6_+_124470053 | 2.40 |
ENSMUST00000049124.10
|
C1rl
|
complement component 1, r subcomponent-like |
| chrX_-_73325318 | 2.38 |
ENSMUST00000239458.2
ENSMUST00000019232.10 |
Dnase1l1
|
deoxyribonuclease 1-like 1 |
| chr15_+_75140241 | 2.38 |
ENSMUST00000023247.13
|
Ly6f
|
lymphocyte antigen 6 complex, locus F |
| chr17_-_24915037 | 2.36 |
ENSMUST00000234235.2
|
Gfer
|
growth factor, augmenter of liver regeneration |
| chr14_-_49482846 | 2.23 |
ENSMUST00000227113.2
ENSMUST00000130853.2 ENSMUST00000228936.2 ENSMUST00000022398.15 |
ccdc198
|
coiled-coil domain containing 198 |
| chr12_-_115832846 | 2.16 |
ENSMUST00000199373.2
|
Ighv1-78
|
immunoglobulin heavy variable 1-78 |
| chr9_+_7272514 | 2.16 |
ENSMUST00000015394.10
|
Mmp13
|
matrix metallopeptidase 13 |
| chr2_-_84255602 | 2.15 |
ENSMUST00000074262.9
|
Calcrl
|
calcitonin receptor-like |
| chrX_-_73325204 | 2.08 |
ENSMUST00000114189.10
ENSMUST00000119361.4 |
Dnase1l1
|
deoxyribonuclease 1-like 1 |
| chr6_-_130044234 | 2.08 |
ENSMUST00000119096.2
|
Klra4
|
killer cell lectin-like receptor, subfamily A, member 4 |
| chr7_+_28140352 | 2.00 |
ENSMUST00000078845.13
|
Gmfg
|
glia maturation factor, gamma |
| chr12_-_113802603 | 1.95 |
ENSMUST00000103458.3
ENSMUST00000193652.2 |
Ighv5-16
|
immunoglobulin heavy variable 5-16 |
| chr6_+_122929627 | 1.86 |
ENSMUST00000204427.2
|
Clec4a3
|
C-type lectin domain family 4, member a3 |
| chr17_-_24915121 | 1.85 |
ENSMUST00000046839.10
|
Gfer
|
growth factor, augmenter of liver regeneration |
| chr12_-_91351177 | 1.82 |
ENSMUST00000141429.8
|
Cep128
|
centrosomal protein 128 |
| chr7_+_44358144 | 1.73 |
ENSMUST00000085422.4
|
Izumo2
|
IZUMO family member 2 |
| chr3_+_84573499 | 1.71 |
ENSMUST00000107682.2
|
Tmem154
|
transmembrane protein 154 |
| chr19_-_17316906 | 1.68 |
ENSMUST00000169897.2
|
Gcnt1
|
glucosaminyl (N-acetyl) transferase 1, core 2 |
| chr2_-_130928658 | 1.64 |
ENSMUST00000028794.10
ENSMUST00000110227.8 ENSMUST00000110226.2 |
Siglec1
|
sialic acid binding Ig-like lectin 1, sialoadhesin |
| chr4_-_156134922 | 1.63 |
ENSMUST00000184684.8
|
Ttll10
|
tubulin tyrosine ligase-like family, member 10 |
| chrX_-_8059597 | 1.63 |
ENSMUST00000143223.2
ENSMUST00000033509.15 |
Ebp
|
phenylalkylamine Ca2+ antagonist (emopamil) binding protein |
| chr18_+_60345152 | 1.58 |
ENSMUST00000031549.6
|
Gm4951
|
predicted gene 4951 |
| chr7_+_28140450 | 1.53 |
ENSMUST00000135686.2
|
Gmfg
|
glia maturation factor, gamma |
| chr15_+_102012782 | 1.48 |
ENSMUST00000230474.2
|
Tns2
|
tensin 2 |
| chr7_-_122969064 | 1.47 |
ENSMUST00000207010.2
ENSMUST00000167309.8 ENSMUST00000205262.2 ENSMUST00000106442.9 ENSMUST00000098060.5 |
Arhgap17
|
Rho GTPase activating protein 17 |
| chr1_-_180073492 | 1.38 |
ENSMUST00000010753.14
|
Psen2
|
presenilin 2 |
| chr5_+_92534738 | 1.32 |
ENSMUST00000128246.8
|
Art3
|
ADP-ribosyltransferase 3 |
| chrX_+_143253677 | 1.26 |
ENSMUST00000178233.2
|
Trpc5os
|
transient receptor potential cation channel, subfamily C, member 5, opposite strand |
| chr3_-_146302343 | 1.23 |
ENSMUST00000029836.9
|
Dnase2b
|
deoxyribonuclease II beta |
| chr15_+_103362195 | 1.18 |
ENSMUST00000047405.9
|
Nckap1l
|
NCK associated protein 1 like |
| chr6_-_54949587 | 1.14 |
ENSMUST00000060655.15
|
Nod1
|
nucleotide-binding oligomerization domain containing 1 |
| chr3_-_49711765 | 1.14 |
ENSMUST00000035931.13
|
Pcdh18
|
protocadherin 18 |
| chr14_+_96118660 | 1.11 |
ENSMUST00000228913.2
ENSMUST00000045892.3 |
Spertl
|
spermatid associated like |
| chr3_-_49711706 | 1.11 |
ENSMUST00000191794.2
|
Pcdh18
|
protocadherin 18 |
| chr4_+_62204678 | 1.10 |
ENSMUST00000084530.9
|
Slc31a2
|
solute carrier family 31, member 2 |
| chr5_+_10997772 | 1.09 |
ENSMUST00000168945.9
ENSMUST00000170064.3 |
Gm8871
Gm8857
|
predicted pseudogene 8871 predicted gene 8857 |
| chr16_-_38649107 | 1.04 |
ENSMUST00000122078.3
|
Tex55
|
testis expressed 55 |
| chr5_+_11968010 | 1.03 |
ENSMUST00000170301.3
ENSMUST00000168329.7 |
4933402N22Rik
|
RIKEN cDNA 4933402N22 gene |
| chr5_+_11820515 | 1.03 |
ENSMUST00000167566.8
|
Gm8926
|
predicted gene 8926 |
| chr5_+_11306000 | 1.02 |
ENSMUST00000164262.8
|
Gm8890
|
predicted gene 8890 |
| chr17_+_25690538 | 1.02 |
ENSMUST00000234449.2
ENSMUST00000025002.3 ENSMUST00000235033.2 |
Tekt4
|
tektin 4 |
| chr5_+_11177601 | 1.00 |
ENSMUST00000170260.8
|
Gm8879
|
predicted gene 8879 |
| chrX_+_133486391 | 0.97 |
ENSMUST00000113211.8
|
Rpl36a
|
ribosomal protein L36A |
| chr1_-_174078542 | 0.96 |
ENSMUST00000061990.5
|
Olfr419
|
olfactory receptor 419 |
| chr5_+_11466385 | 0.96 |
ENSMUST00000167041.8
|
Gm8897
|
predicted gene 8897 |
| chr5_+_11644880 | 0.96 |
ENSMUST00000088616.12
ENSMUST00000163787.3 |
Gm6460
|
predicted gene 6460 |
| chr1_-_184615415 | 0.94 |
ENSMUST00000048308.6
|
C130074G19Rik
|
RIKEN cDNA C130074G19 gene |
| chr5_+_11391483 | 0.93 |
ENSMUST00000172106.9
|
Speer1
|
spermatogenesis associated glutamate (E)-rich protein 1 |
| chr17_-_33879224 | 0.92 |
ENSMUST00000130946.8
|
Hnrnpm
|
heterogeneous nuclear ribonucleoprotein M |
| chr5_+_11731926 | 0.92 |
ENSMUST00000170057.8
|
Gm8922
|
predicted gene 8922 |
| chr6_+_42015319 | 0.92 |
ENSMUST00000117406.8
ENSMUST00000024059.5 |
Sva
|
seminal vesicle antigen |
| chr5_+_11552652 | 0.92 |
ENSMUST00000164651.8
|
Gm8906
|
predicted gene 8906 |
| chr17_+_25097199 | 0.91 |
ENSMUST00000050714.8
|
Igfals
|
insulin-like growth factor binding protein, acid labile subunit |
| chr5_-_88823049 | 0.89 |
ENSMUST00000133532.8
ENSMUST00000150438.2 |
Grsf1
|
G-rich RNA sequence binding factor 1 |
| chr5_-_10296459 | 0.89 |
ENSMUST00000081735.13
ENSMUST00000088625.7 |
Gm5152
|
predicted gene 5152 |
| chr5_+_11895506 | 0.88 |
ENSMUST00000168154.2
|
Gm6465
|
predicted gene 6465 |
| chr16_-_20144154 | 0.86 |
ENSMUST00000182508.9
|
Cyp2ab1
|
cytochrome P450, family 2, subfamily ab, polypeptide 1 |
| chr4_+_48663504 | 0.85 |
ENSMUST00000030033.5
|
Cavin4
|
caveolae associated 4 |
| chr14_+_53478202 | 0.84 |
ENSMUST00000179583.3
|
Trav12n-3
|
T cell receptor alpha variable 12N-3 |
| chr5_+_11233039 | 0.80 |
ENSMUST00000171093.8
|
Gm5861
|
predicted gene 5861 |
| chr7_+_122969004 | 0.76 |
ENSMUST00000206721.2
|
Lcmt1
|
leucine carboxyl methyltransferase 1 |
| chr5_-_86345705 | 0.76 |
ENSMUST00000094654.3
|
Gnrhr
|
gonadotropin releasing hormone receptor |
| chrX_+_56454507 | 0.76 |
ENSMUST00000114725.3
|
Tm9sf5
|
transmembrane 9 superfamily member 5 |
| chr11_+_49916136 | 0.75 |
ENSMUST00000054684.14
ENSMUST00000238748.2 ENSMUST00000102776.5 |
Rnf130
|
ring finger protein 130 |
| chr7_-_107357104 | 0.74 |
ENSMUST00000208217.2
ENSMUST00000052438.8 |
Cyb5r2
|
cytochrome b5 reductase 2 |
| chr15_+_3300249 | 0.74 |
ENSMUST00000082424.12
ENSMUST00000159158.9 ENSMUST00000159216.10 ENSMUST00000160311.3 |
Selenop
|
selenoprotein P |
| chr1_+_93855759 | 0.74 |
ENSMUST00000177958.2
|
Gal3st2b
|
galactose-3-O-sulfotransferase 2B |
| chr2_+_129435115 | 0.73 |
ENSMUST00000099113.10
ENSMUST00000103202.10 |
Sirpa
|
signal-regulatory protein alpha |
| chr1_-_72739704 | 0.73 |
ENSMUST00000053499.6
ENSMUST00000212710.2 ENSMUST00000212573.2 |
Ankar
|
ankyrin and armadillo repeat containing |
| chr5_+_11090082 | 0.72 |
ENSMUST00000163544.2
|
Gm8871
|
predicted pseudogene 8871 |
| chr2_+_153716958 | 0.71 |
ENSMUST00000028983.3
|
Bpifb2
|
BPI fold containing family B, member 2 |
| chr5_-_86345729 | 0.70 |
ENSMUST00000031172.9
ENSMUST00000113372.2 |
Gnrhr
|
gonadotropin releasing hormone receptor |
| chr5_+_11307124 | 0.70 |
ENSMUST00000177727.2
|
Gm8890
|
predicted gene 8890 |
| chr5_+_110478558 | 0.69 |
ENSMUST00000112481.2
|
Pole
|
polymerase (DNA directed), epsilon |
| chr9_+_106377153 | 0.69 |
ENSMUST00000164965.3
|
Iqcf1
|
IQ motif containing F1 |
| chr7_-_14172434 | 0.68 |
ENSMUST00000210396.2
ENSMUST00000168252.9 |
Sult2a8
|
sulfotransferase family 2A, dehydroepiandrosterone (DHEA)-preferring, member 8 |
| chr6_+_87405968 | 0.63 |
ENSMUST00000032125.7
|
Bmp10
|
bone morphogenetic protein 10 |
| chr1_-_72739691 | 0.62 |
ENSMUST00000211837.2
|
Ankar
|
ankyrin and armadillo repeat containing |
| chrX_+_105964224 | 0.59 |
ENSMUST00000060576.8
|
Lpar4
|
lysophosphatidic acid receptor 4 |
| chr4_+_134238310 | 0.57 |
ENSMUST00000105866.3
|
Aunip
|
aurora kinase A and ninein interacting protein |
| chr3_+_146302832 | 0.57 |
ENSMUST00000029837.14
ENSMUST00000147409.2 ENSMUST00000121133.2 |
Uox
|
urate oxidase |
| chr10_+_100426346 | 0.56 |
ENSMUST00000218464.2
ENSMUST00000188930.7 |
1700017N19Rik
|
RIKEN cDNA 1700017N19 gene |
| chr9_-_51240201 | 0.56 |
ENSMUST00000039959.11
ENSMUST00000238450.3 |
1810046K07Rik
|
RIKEN cDNA 1810046K07 gene |
| chr14_+_31888061 | 0.55 |
ENSMUST00000164341.2
|
Ncoa4
|
nuclear receptor coactivator 4 |
| chr10_+_77650454 | 0.54 |
ENSMUST00000220062.2
ENSMUST00000220393.2 |
Gm49918
|
predicted gene, 49918 |
| chr1_-_180073322 | 0.54 |
ENSMUST00000111104.2
|
Psen2
|
presenilin 2 |
| chr2_+_118428690 | 0.47 |
ENSMUST00000038341.8
|
Bub1b
|
BUB1B, mitotic checkpoint serine/threonine kinase |
| chr3_-_88455556 | 0.47 |
ENSMUST00000131775.2
ENSMUST00000008745.13 |
Rab25
|
RAB25, member RAS oncogene family |
| chr5_-_87288177 | 0.46 |
ENSMUST00000067790.7
|
Ugt2b5
|
UDP glucuronosyltransferase 2 family, polypeptide B5 |
| chr6_-_119925387 | 0.45 |
ENSMUST00000162541.8
|
Wnk1
|
WNK lysine deficient protein kinase 1 |
| chr14_+_65842716 | 0.44 |
ENSMUST00000079469.7
|
Nuggc
|
nuclear GTPase, germinal center associated |
| chr1_+_93789548 | 0.44 |
ENSMUST00000187896.2
|
Gal3st2
|
galactose-3-O-sulfotransferase 2 |
| chr4_+_43851565 | 0.43 |
ENSMUST00000107860.3
|
Olfr155
|
olfactory receptor 155 |
| chr3_-_27950491 | 0.43 |
ENSMUST00000058077.4
|
Tmem212
|
transmembrane protein 212 |
| chr14_+_51648277 | 0.43 |
ENSMUST00000159674.4
ENSMUST00000163019.9 ENSMUST00000233394.2 ENSMUST00000232951.2 ENSMUST00000232797.2 ENSMUST00000233766.2 ENSMUST00000233848.2 ENSMUST00000233280.2 ENSMUST00000233920.2 ENSMUST00000233126.2 ENSMUST00000233651.2 |
Vmn2r88
|
vomeronasal 2, receptor 88 |
| chr6_-_41012435 | 0.42 |
ENSMUST00000031931.6
|
2210010C04Rik
|
RIKEN cDNA 2210010C04 gene |
| chr3_-_51316347 | 0.42 |
ENSMUST00000193279.2
ENSMUST00000038108.12 |
Ndufc1
|
NADH:ubiquinone oxidoreductase subunit C1 |
| chr5_-_65548723 | 0.42 |
ENSMUST00000120094.5
ENSMUST00000127874.6 ENSMUST00000057885.13 |
Rpl9
|
ribosomal protein L9 |
| chr1_-_139786421 | 0.42 |
ENSMUST00000194186.6
ENSMUST00000094489.5 ENSMUST00000239380.2 |
Cfhr2
|
complement factor H-related 2 |
| chr16_+_17149235 | 0.40 |
ENSMUST00000023450.15
ENSMUST00000231884.2 |
Serpind1
|
serine (or cysteine) peptidase inhibitor, clade D, member 1 |
| chr17_-_40553176 | 0.40 |
ENSMUST00000026499.6
|
Crisp3
|
cysteine-rich secretory protein 3 |
| chr8_+_47439916 | 0.37 |
ENSMUST00000039840.15
|
Enpp6
|
ectonucleotide pyrophosphatase/phosphodiesterase 6 |
| chr19_-_39451509 | 0.35 |
ENSMUST00000035488.3
|
Cyp2c38
|
cytochrome P450, family 2, subfamily c, polypeptide 38 |
| chr4_+_156683398 | 0.34 |
ENSMUST00000074107.13
ENSMUST00000096792.8 |
Vmn2r129
|
vomeronasal 2, receptor 129 |
| chr10_+_33913593 | 0.33 |
ENSMUST00000095758.3
|
Trappc3l
|
trafficking protein particle complex 3 like |
| chr7_+_79882609 | 0.32 |
ENSMUST00000062915.9
|
Gdpgp1
|
GDP-D-glucose phosphorylase 1 |
| chr6_-_68784692 | 0.28 |
ENSMUST00000103334.4
|
Igkv4-90
|
immunoglobulin kappa chain variable 4-90 |
| chr17_-_41225241 | 0.28 |
ENSMUST00000166343.3
|
Glyatl3
|
glycine-N-acyltransferase-like 3 |
| chr17_+_32877851 | 0.25 |
ENSMUST00000235086.2
|
Cyp4f40
|
cytochrome P450, family 4, subfamily f, polypeptide 40 |
| chr3_-_129616136 | 0.25 |
ENSMUST00000029648.11
ENSMUST00000171313.8 ENSMUST00000200079.5 |
Rrh
|
retinal pigment epithelium derived rhodopsin homolog |
| chr4_+_80752360 | 0.24 |
ENSMUST00000133655.8
ENSMUST00000006151.13 |
Tyrp1
|
tyrosinase-related protein 1 |
| chr13_-_4329421 | 0.24 |
ENSMUST00000021632.5
|
Akr1c12
|
aldo-keto reductase family 1, member C12 |
| chr14_+_53167246 | 0.23 |
ENSMUST00000177703.3
|
Trav12d-3
|
T cell receptor alpha variable 12D-3 |
| chr6_+_89684522 | 0.22 |
ENSMUST00000228642.2
ENSMUST00000226925.2 ENSMUST00000228485.2 ENSMUST00000227669.2 |
Vmn1r40
|
vomeronasal 1 receptor 40 |
| chrY_-_6681243 | 0.22 |
ENSMUST00000115940.2
|
Gm21719
|
predicted gene, 21719 |
| chr16_-_19226828 | 0.19 |
ENSMUST00000052516.5
ENSMUST00000206410.3 |
Olfr165
|
olfactory receptor 165 |
| chr14_-_70945434 | 0.19 |
ENSMUST00000228346.2
|
Xpo7
|
exportin 7 |
| chr7_+_28508220 | 0.19 |
ENSMUST00000172529.8
|
Hnrnpl
|
heterogeneous nuclear ribonucleoprotein L |
| chr14_-_36657517 | 0.19 |
ENSMUST00000183038.8
|
Ccser2
|
coiled-coil serine rich 2 |
| chr14_-_50476340 | 0.18 |
ENSMUST00000059565.2
|
Olfr731
|
olfactory receptor 731 |
| chr6_+_40546421 | 0.16 |
ENSMUST00000216942.2
|
Olfr460
|
olfactory receptor 460 |
| chr19_-_13807805 | 0.15 |
ENSMUST00000214475.2
ENSMUST00000067670.5 ENSMUST00000219674.2 ENSMUST00000216287.2 ENSMUST00000215760.2 |
Olfr1500
|
olfactory receptor 1500 |
| chr6_+_132832401 | 0.15 |
ENSMUST00000032315.3
|
Tas2r116
|
taste receptor, type 2, member 116 |
| chr14_+_51648458 | 0.15 |
ENSMUST00000022438.12
|
Vmn2r88
|
vomeronasal 2, receptor 88 |
| chr5_+_109478028 | 0.14 |
ENSMUST00000165180.3
ENSMUST00000232912.2 |
Vmn2r16
|
vomeronasal 2, receptor 16 |
| chr9_+_25000718 | 0.14 |
ENSMUST00000086238.3
|
Gm10181
|
predicted gene 10181 |
| chr7_+_84502761 | 0.13 |
ENSMUST00000217039.3
ENSMUST00000211582.2 |
Olfr291
|
olfactory receptor 291 |
| chr2_-_111820618 | 0.13 |
ENSMUST00000216948.2
ENSMUST00000214935.2 ENSMUST00000217452.2 ENSMUST00000215045.2 |
Olfr1309
|
olfactory receptor 1309 |
| chr7_+_142660095 | 0.12 |
ENSMUST00000186488.7
|
Kcnq1
|
potassium voltage-gated channel, subfamily Q, member 1 |
| chr7_-_103710652 | 0.12 |
ENSMUST00000074064.5
|
4930516K23Rik
|
RIKEN cDNA 4930516K23 gene |
| chr3_-_64473251 | 0.12 |
ENSMUST00000176481.9
|
Vmn2r6
|
vomeronasal 2, receptor 6 |
| chr14_+_51689419 | 0.11 |
ENSMUST00000159611.10
ENSMUST00000159734.4 |
Vmn2r89
|
vomeronasal 2, receptor 89 |
| chr4_-_136626073 | 0.11 |
ENSMUST00000046285.6
|
C1qa
|
complement component 1, q subcomponent, alpha polypeptide |
| chr7_+_29883569 | 0.09 |
ENSMUST00000098594.4
|
Cox7a1
|
cytochrome c oxidase subunit 7A1 |
| chr2_-_89583170 | 0.08 |
ENSMUST00000099765.2
|
Olfr1253
|
olfactory receptor 1253 |
| chr8_+_71176713 | 0.07 |
ENSMUST00000034307.14
ENSMUST00000239435.2 ENSMUST00000239487.2 ENSMUST00000110095.3 |
Pde4c
|
phosphodiesterase 4C, cAMP specific |
| chr3_-_97318495 | 0.05 |
ENSMUST00000060912.4
|
Olfr1402
|
olfactory receptor 1402 |
| chr7_-_102566717 | 0.03 |
ENSMUST00000214160.2
ENSMUST00000215773.2 |
Olfr571
|
olfactory receptor 571 |
| chr2_-_85966272 | 0.02 |
ENSMUST00000216566.3
ENSMUST00000214364.2 |
Olfr1039
|
olfactory receptor 1039 |
| chr4_+_114020581 | 0.01 |
ENSMUST00000079915.10
ENSMUST00000164297.8 |
Skint11
|
selection and upkeep of intraepithelial T cells 11 |
| chr17_-_30831576 | 0.01 |
ENSMUST00000235171.2
ENSMUST00000236335.2 ENSMUST00000167624.2 |
Glo1
|
glyoxalase 1 |
| chr16_+_51870001 | 0.01 |
ENSMUST00000227756.2
|
Cblb
|
Casitas B-lineage lymphoma b |
| chrX_+_20984611 | 0.01 |
ENSMUST00000133987.3
|
Ssxa1
|
synovial sarcoma, X member A1 |
| chr11_-_121563894 | 0.00 |
ENSMUST00000067399.14
ENSMUST00000062654.8 |
B3gntl1
|
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase-like 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 8.5 | GO:0072592 | oxygen metabolic process(GO:0072592) |
| 1.2 | 3.5 | GO:0071846 | actin filament debranching(GO:0071846) |
| 0.8 | 2.5 | GO:0002149 | hypochlorous acid metabolic process(GO:0002148) hypochlorous acid biosynthetic process(GO:0002149) |
| 0.5 | 2.2 | GO:0090187 | positive regulation of pancreatic juice secretion(GO:0090187) |
| 0.5 | 2.1 | GO:0034285 | response to sucrose(GO:0009744) response to disaccharide(GO:0034285) |
| 0.5 | 3.3 | GO:0042738 | exogenous drug catabolic process(GO:0042738) |
| 0.3 | 4.2 | GO:0097421 | liver regeneration(GO:0097421) |
| 0.3 | 1.2 | GO:0043376 | regulation of CD8-positive, alpha-beta T cell differentiation(GO:0043376) actin polymerization-dependent cell motility(GO:0070358) |
| 0.3 | 1.6 | GO:0018094 | protein polyglycylation(GO:0018094) |
| 0.2 | 0.7 | GO:0045004 | DNA replication proofreading(GO:0045004) |
| 0.2 | 1.0 | GO:0080154 | regulation of fertilization(GO:0080154) |
| 0.2 | 1.5 | GO:0097210 | response to gonadotropin-releasing hormone(GO:0097210) cellular response to gonadotropin-releasing hormone(GO:0097211) |
| 0.2 | 0.8 | GO:0006481 | C-terminal protein methylation(GO:0006481) |
| 0.1 | 1.1 | GO:0002606 | positive regulation of dendritic cell antigen processing and presentation(GO:0002606) |
| 0.1 | 5.7 | GO:0006308 | DNA catabolic process(GO:0006308) |
| 0.1 | 1.9 | GO:0043589 | skin morphogenesis(GO:0043589) |
| 0.1 | 3.3 | GO:0002385 | mucosal immune response(GO:0002385) |
| 0.1 | 0.7 | GO:0060474 | positive regulation of sperm motility involved in capacitation(GO:0060474) |
| 0.1 | 0.5 | GO:0051754 | meiotic sister chromatid cohesion, centromeric(GO:0051754) |
| 0.1 | 0.7 | GO:0001887 | selenium compound metabolic process(GO:0001887) |
| 0.1 | 1.1 | GO:0035434 | copper ion transmembrane transport(GO:0035434) |
| 0.1 | 0.4 | GO:0023016 | signal transduction by trans-phosphorylation(GO:0023016) |
| 0.1 | 1.7 | GO:0060352 | cell adhesion molecule production(GO:0060352) |
| 0.1 | 0.3 | GO:0042376 | menaquinone metabolic process(GO:0009233) phylloquinone metabolic process(GO:0042374) phylloquinone catabolic process(GO:0042376) quinone catabolic process(GO:1901662) |
| 0.1 | 0.6 | GO:1903242 | regulation of cardiac muscle adaptation(GO:0010612) regulation of cardiac muscle hypertrophy in response to stress(GO:1903242) |
| 0.1 | 0.2 | GO:0006583 | melanin biosynthetic process from tyrosine(GO:0006583) |
| 0.1 | 1.6 | GO:0070234 | positive regulation of T cell apoptotic process(GO:0070234) |
| 0.1 | 2.2 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.1 | 1.5 | GO:0014850 | response to muscle activity(GO:0014850) |
| 0.0 | 6.6 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.0 | 0.1 | GO:0072347 | response to anesthetic(GO:0072347) |
| 0.0 | 2.5 | GO:0035082 | axoneme assembly(GO:0035082) |
| 0.0 | 0.4 | GO:0016446 | somatic hypermutation of immunoglobulin genes(GO:0016446) |
| 0.0 | 0.9 | GO:0055003 | cardiac myofibril assembly(GO:0055003) |
| 0.0 | 1.6 | GO:0035458 | cellular response to interferon-beta(GO:0035458) |
| 0.0 | 1.3 | GO:0006471 | protein ADP-ribosylation(GO:0006471) |
| 0.0 | 0.5 | GO:0006622 | protein targeting to lysosome(GO:0006622) |
| 0.0 | 0.6 | GO:0046415 | urate metabolic process(GO:0046415) |
| 0.0 | 0.5 | GO:0031268 | pseudopodium organization(GO:0031268) |
| 0.0 | 0.2 | GO:0019695 | choline metabolic process(GO:0019695) |
| 0.0 | 0.4 | GO:0051923 | sulfation(GO:0051923) |
| 0.0 | 0.2 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.0 | 0.6 | GO:0051482 | positive regulation of cytosolic calcium ion concentration involved in phospholipase C-activating G-protein coupled signaling pathway(GO:0051482) |
| 0.0 | 3.8 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.0 | 0.2 | GO:1902416 | positive regulation of mRNA binding(GO:1902416) |
| 0.0 | 0.4 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.0 | 0.3 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.5 | GO:0072687 | meiotic spindle(GO:0072687) |
| 0.1 | 2.5 | GO:0042582 | primary lysosome(GO:0005766) azurophil granule(GO:0042582) |
| 0.1 | 0.7 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 0.1 | 2.2 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.1 | 0.9 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.1 | 0.9 | GO:0042567 | insulin-like growth factor ternary complex(GO:0042567) |
| 0.1 | 2.8 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.1 | 1.2 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.1 | 1.9 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.1 | 7.4 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.1 | 12.0 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 0.5 | GO:0044754 | autolysosome(GO:0044754) |
| 0.0 | 1.0 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.0 | 0.5 | GO:0000778 | condensed nuclear chromosome kinetochore(GO:0000778) |
| 0.0 | 0.2 | GO:0045009 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.0 | 0.5 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.0 | 1.7 | GO:0031985 | Golgi cisterna(GO:0031985) |
| 0.0 | 1.4 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.0 | 0.9 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 0.3 | GO:0030008 | TRAPP complex(GO:0030008) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 4.2 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) |
| 0.7 | 2.1 | GO:0004948 | calcitonin receptor activity(GO:0004948) |
| 0.7 | 8.5 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.5 | 1.5 | GO:0004968 | gonadotropin-releasing hormone receptor activity(GO:0004968) |
| 0.3 | 1.7 | GO:0003829 | beta-1,3-galactosyl-O-glycosyl-glycoprotein beta-1,6-N-acetylglucosaminyltransferase activity(GO:0003829) |
| 0.3 | 1.6 | GO:0070735 | protein-glycine ligase activity(GO:0070735) |
| 0.3 | 3.5 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.2 | 1.2 | GO:0004531 | deoxyribonuclease II activity(GO:0004531) |
| 0.2 | 1.6 | GO:0004769 | steroid delta-isomerase activity(GO:0004769) |
| 0.2 | 1.9 | GO:0042500 | aspartic endopeptidase activity, intramembrane cleaving(GO:0042500) |
| 0.2 | 0.8 | GO:0003880 | protein C-terminal carboxyl O-methyltransferase activity(GO:0003880) |
| 0.1 | 3.6 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.1 | 1.1 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.1 | 1.3 | GO:0003956 | NAD(P)+-protein-arginine ADP-ribosyltransferase activity(GO:0003956) |
| 0.1 | 0.6 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 0.1 | 1.1 | GO:0005375 | copper ion transmembrane transporter activity(GO:0005375) |
| 0.1 | 0.7 | GO:0008310 | single-stranded DNA 3'-5' exodeoxyribonuclease activity(GO:0008310) |
| 0.1 | 0.4 | GO:0050694 | galactose 3-O-sulfotransferase activity(GO:0050694) |
| 0.1 | 0.7 | GO:0004128 | cytochrome-b5 reductase activity, acting on NAD(P)H(GO:0004128) |
| 0.1 | 0.9 | GO:1990405 | protein antigen binding(GO:1990405) |
| 0.1 | 2.2 | GO:0001968 | fibronectin binding(GO:0001968) |
| 0.1 | 7.4 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.1 | 4.5 | GO:0004536 | deoxyribonuclease activity(GO:0004536) |
| 0.1 | 0.3 | GO:0004645 | phosphorylase activity(GO:0004645) |
| 0.1 | 0.6 | GO:0031433 | telethonin binding(GO:0031433) |
| 0.0 | 0.6 | GO:0016661 | oxidoreductase activity, acting on other nitrogenous compounds as donors(GO:0016661) |
| 0.0 | 2.5 | GO:0004601 | peroxidase activity(GO:0004601) |
| 0.0 | 0.7 | GO:0008430 | selenium binding(GO:0008430) |
| 0.0 | 0.2 | GO:0016716 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, another compound as one donor, and incorporation of one atom of oxygen(GO:0016716) |
| 0.0 | 0.9 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.0 | 0.4 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) |
| 0.0 | 2.8 | GO:0016811 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amides(GO:0016811) |
| 0.0 | 0.2 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.0 | 0.2 | GO:0008889 | glycerophosphodiester phosphodiesterase activity(GO:0008889) |
| 0.0 | 0.9 | GO:0008395 | steroid hydroxylase activity(GO:0008395) |
| 0.0 | 0.1 | GO:0086089 | voltage-gated potassium channel activity involved in atrial cardiac muscle cell action potential repolarization(GO:0086089) |
| 0.0 | 0.2 | GO:0097157 | pre-mRNA intronic binding(GO:0097157) |
| 0.0 | 0.5 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.0 | 0.2 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.0 | 2.6 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 1.2 | GO:0030295 | protein kinase activator activity(GO:0030295) |
| 0.0 | 0.7 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.0 | 3.7 | GO:0030246 | carbohydrate binding(GO:0030246) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 2.5 | PID IL23 PATHWAY | IL23-mediated signaling events |
| 0.0 | 2.2 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.0 | 0.6 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.0 | 1.9 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.0 | 4.6 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 1.6 | SIG PIP3 SIGNALING IN CARDIAC MYOCTES | Genes related to PIP3 signaling in cardiac myocytes |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 8.5 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.1 | 1.5 | REACTOME HORMONE LIGAND BINDING RECEPTORS | Genes involved in Hormone ligand-binding receptors |
| 0.1 | 1.9 | REACTOME REGULATED PROTEOLYSIS OF P75NTR | Genes involved in Regulated proteolysis of p75NTR |
| 0.1 | 4.2 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.1 | 2.2 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.1 | 2.8 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.0 | 0.9 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.0 | 1.1 | REACTOME ACTIVATED TAK1 MEDIATES P38 MAPK ACTIVATION | Genes involved in activated TAK1 mediates p38 MAPK activation |
| 0.0 | 0.6 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.0 | 1.7 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 0.2 | REACTOME OPSINS | Genes involved in Opsins |
| 0.0 | 0.5 | REACTOME INHIBITION OF THE PROTEOLYTIC ACTIVITY OF APC C REQUIRED FOR THE ONSET OF ANAPHASE BY MITOTIC SPINDLE CHECKPOINT COMPONENTS | Genes involved in Inhibition of the proteolytic activity of APC/C required for the onset of anaphase by mitotic spindle checkpoint components |
| 0.0 | 0.7 | REACTOME ACTIVATION OF THE PRE REPLICATIVE COMPLEX | Genes involved in Activation of the pre-replicative complex |
| 0.0 | 2.1 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 2.3 | REACTOME INFLUENZA VIRAL RNA TRANSCRIPTION AND REPLICATION | Genes involved in Influenza Viral RNA Transcription and Replication |
| 0.0 | 0.1 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.0 | 0.4 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |