Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
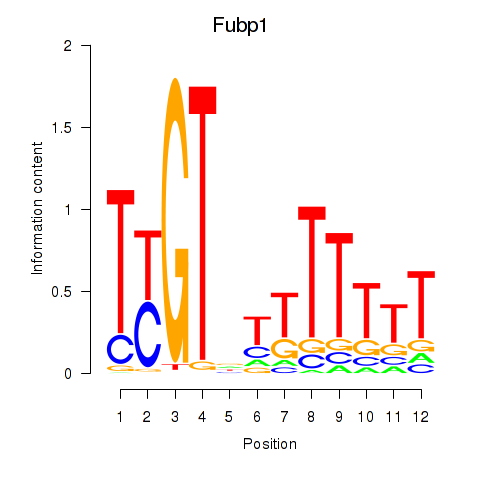
Results for Fubp1
Z-value: 0.82
Transcription factors associated with Fubp1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Fubp1
|
ENSMUSG00000028034.16 | Fubp1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Fubp1 | mm39_v1_chr3_+_151916059_151916102 | -0.05 | 7.1e-01 | Click! |
Activity profile of Fubp1 motif
Sorted Z-values of Fubp1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Fubp1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr7_-_110462446 | 6.12 |
ENSMUST00000033050.5
|
Lyve1
|
lymphatic vessel endothelial hyaluronan receptor 1 |
| chr11_+_44508137 | 5.91 |
ENSMUST00000109268.2
ENSMUST00000101326.10 ENSMUST00000081265.12 |
Ebf1
|
early B cell factor 1 |
| chr6_+_17307639 | 5.65 |
ENSMUST00000115453.2
|
Cav1
|
caveolin 1, caveolae protein |
| chr2_-_60503998 | 5.28 |
ENSMUST00000059888.15
ENSMUST00000154764.2 |
Itgb6
|
integrin beta 6 |
| chr17_-_46343291 | 5.26 |
ENSMUST00000071648.12
ENSMUST00000142351.9 |
Vegfa
|
vascular endothelial growth factor A |
| chr11_-_106605772 | 5.16 |
ENSMUST00000124958.3
|
Pecam1
|
platelet/endothelial cell adhesion molecule 1 |
| chr11_-_106606076 | 4.95 |
ENSMUST00000080853.11
ENSMUST00000183610.8 ENSMUST00000103069.10 ENSMUST00000106796.9 |
Pecam1
|
platelet/endothelial cell adhesion molecule 1 |
| chr17_+_34423054 | 4.29 |
ENSMUST00000138491.2
|
Tap2
|
transporter 2, ATP-binding cassette, sub-family B (MDR/TAP) |
| chr10_+_97315465 | 4.17 |
ENSMUST00000105287.11
|
Dcn
|
decorin |
| chr3_+_137046828 | 4.12 |
ENSMUST00000122064.8
ENSMUST00000119475.6 |
Emcn
|
endomucin |
| chr3_+_137046935 | 3.99 |
ENSMUST00000197511.2
|
Emcn
|
endomucin |
| chr5_-_69699965 | 3.78 |
ENSMUST00000031045.10
|
Yipf7
|
Yip1 domain family, member 7 |
| chr16_-_94171340 | 3.72 |
ENSMUST00000138514.2
|
Pigp
|
phosphatidylinositol glycan anchor biosynthesis, class P |
| chr3_-_151960948 | 3.60 |
ENSMUST00000199423.5
ENSMUST00000198460.5 |
Nexn
|
nexilin |
| chr3_-_151960992 | 3.35 |
ENSMUST00000198750.5
|
Nexn
|
nexilin |
| chr5_-_69699932 | 3.26 |
ENSMUST00000202423.2
|
Yipf7
|
Yip1 domain family, member 7 |
| chr4_+_94627755 | 3.22 |
ENSMUST00000071168.6
|
Tek
|
TEK receptor tyrosine kinase |
| chr15_-_5093222 | 3.00 |
ENSMUST00000110689.5
|
C7
|
complement component 7 |
| chr7_+_89053562 | 2.89 |
ENSMUST00000058755.5
|
Fzd4
|
frizzled class receptor 4 |
| chr16_-_94171556 | 2.88 |
ENSMUST00000113906.9
|
Pigp
|
phosphatidylinositol glycan anchor biosynthesis, class P |
| chr10_+_102348076 | 2.84 |
ENSMUST00000219445.2
|
Rassf9
|
Ras association (RalGDS/AF-6) domain family (N-terminal) member 9 |
| chr11_+_67090878 | 2.81 |
ENSMUST00000124516.8
ENSMUST00000018637.15 ENSMUST00000129018.8 |
Myh1
|
myosin, heavy polypeptide 1, skeletal muscle, adult |
| chrX_+_149377416 | 2.60 |
ENSMUST00000112713.3
|
Pfkfb1
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 |
| chr1_-_97904958 | 2.54 |
ENSMUST00000161567.8
|
Pam
|
peptidylglycine alpha-amidating monooxygenase |
| chr1_-_138103021 | 2.54 |
ENSMUST00000182755.8
ENSMUST00000193650.2 ENSMUST00000182283.8 |
Ptprc
|
protein tyrosine phosphatase, receptor type, C |
| chr6_+_29361408 | 2.48 |
ENSMUST00000156163.2
|
Calu
|
calumenin |
| chr16_-_94171533 | 2.43 |
ENSMUST00000113910.8
|
Pigp
|
phosphatidylinositol glycan anchor biosynthesis, class P |
| chr15_+_102862862 | 2.42 |
ENSMUST00000001701.4
|
Hoxc11
|
homeobox C11 |
| chr7_+_142014546 | 2.39 |
ENSMUST00000105968.8
ENSMUST00000018963.11 ENSMUST00000105967.8 |
Lsp1
|
lymphocyte specific 1 |
| chr4_-_94538329 | 2.32 |
ENSMUST00000107101.2
|
Lrrc19
|
leucine rich repeat containing 19 |
| chr4_-_55532453 | 2.29 |
ENSMUST00000132746.2
ENSMUST00000107619.3 |
Klf4
|
Kruppel-like factor 4 (gut) |
| chr6_+_48624295 | 2.27 |
ENSMUST00000078223.6
ENSMUST00000203509.2 |
Gimap8
|
GTPase, IMAP family member 8 |
| chr11_-_68277799 | 2.23 |
ENSMUST00000135141.2
|
Ntn1
|
netrin 1 |
| chr2_-_28806639 | 2.10 |
ENSMUST00000113847.3
|
Barhl1
|
BarH like homeobox 1 |
| chr15_+_37233280 | 2.09 |
ENSMUST00000161405.8
ENSMUST00000022895.15 ENSMUST00000161532.2 |
Grhl2
|
grainyhead like transcription factor 2 |
| chr13_+_112604037 | 2.05 |
ENSMUST00000183868.8
|
Il6st
|
interleukin 6 signal transducer |
| chr4_-_94538370 | 2.05 |
ENSMUST00000053419.9
|
Lrrc19
|
leucine rich repeat containing 19 |
| chr1_-_138102972 | 2.04 |
ENSMUST00000195533.6
ENSMUST00000183301.8 |
Ptprc
|
protein tyrosine phosphatase, receptor type, C |
| chr3_-_157630690 | 2.02 |
ENSMUST00000118539.2
|
Cth
|
cystathionase (cystathionine gamma-lyase) |
| chr4_-_49549489 | 1.97 |
ENSMUST00000029987.10
|
Aldob
|
aldolase B, fructose-bisphosphate |
| chr3_-_108934916 | 1.94 |
ENSMUST00000171143.2
|
Fam102b
|
family with sequence similarity 102, member B |
| chr14_+_51366512 | 1.93 |
ENSMUST00000095923.4
|
Rnase6
|
ribonuclease, RNase A family, 6 |
| chr1_-_134883645 | 1.87 |
ENSMUST00000045665.13
ENSMUST00000086444.6 ENSMUST00000112163.2 |
Ppp1r12b
|
protein phosphatase 1, regulatory subunit 12B |
| chr6_-_122833109 | 1.86 |
ENSMUST00000042081.9
|
C3ar1
|
complement component 3a receptor 1 |
| chr4_+_144619696 | 1.83 |
ENSMUST00000142808.8
|
Dhrs3
|
dehydrogenase/reductase (SDR family) member 3 |
| chr4_-_147726953 | 1.82 |
ENSMUST00000133006.2
ENSMUST00000037565.14 ENSMUST00000105720.8 |
Zfp979
|
zinc finger protein 979 |
| chr7_-_70010341 | 1.79 |
ENSMUST00000032768.15
|
Nr2f2
|
nuclear receptor subfamily 2, group F, member 2 |
| chr10_+_94412116 | 1.77 |
ENSMUST00000117929.2
|
Tmcc3
|
transmembrane and coiled coil domains 3 |
| chr1_+_156193607 | 1.62 |
ENSMUST00000102782.4
|
Gm2000
|
predicted gene 2000 |
| chr4_+_144619647 | 1.58 |
ENSMUST00000154208.8
|
Dhrs3
|
dehydrogenase/reductase (SDR family) member 3 |
| chr3_+_92192724 | 1.56 |
ENSMUST00000193337.2
|
Sprr2a3
|
small proline-rich protein 2A3 |
| chrX_-_47602395 | 1.50 |
ENSMUST00000114945.9
ENSMUST00000037349.8 |
Aifm1
|
apoptosis-inducing factor, mitochondrion-associated 1 |
| chrX_+_132809166 | 1.47 |
ENSMUST00000033606.15
|
Srpx2
|
sushi-repeat-containing protein, X-linked 2 |
| chr4_-_120682859 | 1.42 |
ENSMUST00000134979.8
ENSMUST00000136236.8 ENSMUST00000043429.12 ENSMUST00000145658.2 |
Nfyc
|
nuclear transcription factor-Y gamma |
| chr5_+_110987839 | 1.41 |
ENSMUST00000200172.2
ENSMUST00000066160.3 |
Chek2
|
checkpoint kinase 2 |
| chr19_-_11301919 | 1.39 |
ENSMUST00000159269.2
|
Ms4a7
|
membrane-spanning 4-domains, subfamily A, member 7 |
| chr9_-_48747232 | 1.36 |
ENSMUST00000093852.5
|
Zbtb16
|
zinc finger and BTB domain containing 16 |
| chr2_+_103800459 | 1.34 |
ENSMUST00000111143.8
ENSMUST00000138815.2 |
Lmo2
|
LIM domain only 2 |
| chr11_-_107238956 | 1.33 |
ENSMUST00000134763.2
|
Pitpnc1
|
phosphatidylinositol transfer protein, cytoplasmic 1 |
| chr11_-_17161504 | 1.27 |
ENSMUST00000020317.8
|
Pno1
|
partner of NOB1 homolog |
| chr9_-_117081518 | 1.25 |
ENSMUST00000111773.10
ENSMUST00000068962.14 ENSMUST00000044901.14 |
Rbms3
|
RNA binding motif, single stranded interacting protein |
| chr13_-_3943433 | 1.24 |
ENSMUST00000222504.2
|
Net1
|
neuroepithelial cell transforming gene 1 |
| chr13_+_36301331 | 1.23 |
ENSMUST00000021857.13
|
Fars2
|
phenylalanine-tRNA synthetase 2 (mitochondrial) |
| chr2_+_103800553 | 1.22 |
ENSMUST00000111140.3
ENSMUST00000111139.3 |
Lmo2
|
LIM domain only 2 |
| chr1_-_134883577 | 1.18 |
ENSMUST00000168381.8
|
Ppp1r12b
|
protein phosphatase 1, regulatory subunit 12B |
| chr10_+_94350687 | 1.18 |
ENSMUST00000065060.12
|
Tmcc3
|
transmembrane and coiled coil domains 3 |
| chr7_-_110681402 | 1.14 |
ENSMUST00000159305.2
|
Eif4g2
|
eukaryotic translation initiation factor 4, gamma 2 |
| chr5_+_110988095 | 1.13 |
ENSMUST00000198373.2
|
Chek2
|
checkpoint kinase 2 |
| chr17_+_35268942 | 1.13 |
ENSMUST00000007257.10
|
Clic1
|
chloride intracellular channel 1 |
| chr9_-_117979147 | 1.10 |
ENSMUST00000215799.2
ENSMUST00000044220.11 |
Cmc1
|
COX assembly mitochondrial protein 1 |
| chr12_-_114487525 | 1.10 |
ENSMUST00000103495.3
|
Ighv10-3
|
immunoglobulin heavy variable V10-3 |
| chr5_+_67125759 | 1.09 |
ENSMUST00000238993.2
ENSMUST00000038188.14 |
Limch1
|
LIM and calponin homology domains 1 |
| chr11_+_67167950 | 1.07 |
ENSMUST00000019625.12
|
Myh8
|
myosin, heavy polypeptide 8, skeletal muscle, perinatal |
| chr14_-_62718581 | 1.06 |
ENSMUST00000186010.8
|
Gm4131
|
predicted gene 4131 |
| chr12_+_21366386 | 1.04 |
ENSMUST00000076813.8
ENSMUST00000221693.2 ENSMUST00000223345.2 ENSMUST00000222344.2 |
Iah1
|
isoamyl acetate-hydrolyzing esterase 1 homolog |
| chr10_-_12689345 | 1.03 |
ENSMUST00000217899.2
|
Utrn
|
utrophin |
| chr11_+_83742961 | 0.97 |
ENSMUST00000146786.8
|
Hnf1b
|
HNF1 homeobox B |
| chrX_-_161612373 | 0.97 |
ENSMUST00000041370.11
ENSMUST00000112316.9 ENSMUST00000112315.2 |
Txlng
|
taxilin gamma |
| chr7_+_122723365 | 0.97 |
ENSMUST00000205514.2
ENSMUST00000094053.7 |
Tnrc6a
|
trinucleotide repeat containing 6a |
| chr12_+_35975619 | 0.90 |
ENSMUST00000042101.5
ENSMUST00000154042.2 |
Agr3
|
anterior gradient 3 |
| chr18_-_84104574 | 0.90 |
ENSMUST00000175783.3
|
Tshz1
|
teashirt zinc finger family member 1 |
| chr19_+_41471395 | 0.85 |
ENSMUST00000237208.2
ENSMUST00000238398.2 |
Lcor
|
ligand dependent nuclear receptor corepressor |
| chr1_+_21419819 | 0.85 |
ENSMUST00000088407.4
|
Khdc1a
|
KH domain containing 1A |
| chr6_-_114898739 | 0.84 |
ENSMUST00000032459.14
|
Vgll4
|
vestigial like family member 4 |
| chr2_+_3115250 | 0.81 |
ENSMUST00000072955.12
|
Fam171a1
|
family with sequence similarity 171, member A1 |
| chr10_+_53472853 | 0.80 |
ENSMUST00000219271.2
|
Asf1a
|
anti-silencing function 1A histone chaperone |
| chr11_-_78074377 | 0.80 |
ENSMUST00000102483.5
|
Rpl23a
|
ribosomal protein L23A |
| chr1_+_157334298 | 0.76 |
ENSMUST00000086130.9
|
Sec16b
|
SEC16 homolog B (S. cerevisiae) |
| chr5_-_44956981 | 0.75 |
ENSMUST00000070748.10
|
Ldb2
|
LIM domain binding 2 |
| chr11_-_40586029 | 0.75 |
ENSMUST00000101347.10
|
Mat2b
|
methionine adenosyltransferase II, beta |
| chr1_+_157334347 | 0.74 |
ENSMUST00000027881.15
|
Sec16b
|
SEC16 homolog B (S. cerevisiae) |
| chr5_-_73789764 | 0.74 |
ENSMUST00000087177.4
|
Lrrc66
|
leucine rich repeat containing 66 |
| chr19_+_43770619 | 0.73 |
ENSMUST00000026208.6
|
Abcc2
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 2 |
| chr5_-_44956932 | 0.72 |
ENSMUST00000199261.2
ENSMUST00000199534.5 |
Ldb2
|
LIM domain binding 2 |
| chr4_-_130169006 | 0.72 |
ENSMUST00000122374.8
|
Serinc2
|
serine incorporator 2 |
| chr18_-_84104507 | 0.71 |
ENSMUST00000060303.10
|
Tshz1
|
teashirt zinc finger family member 1 |
| chr19_+_44551280 | 0.70 |
ENSMUST00000040455.5
|
Hif1an
|
hypoxia-inducible factor 1, alpha subunit inhibitor |
| chr9_-_117080869 | 0.70 |
ENSMUST00000172564.3
|
Rbms3
|
RNA binding motif, single stranded interacting protein |
| chr1_+_36550948 | 0.69 |
ENSMUST00000001166.14
ENSMUST00000097776.4 |
Cnnm3
|
cyclin M3 |
| chrX_-_50106844 | 0.68 |
ENSMUST00000053593.8
|
Rap2c
|
RAP2C, member of RAS oncogene family |
| chr7_+_101619897 | 0.65 |
ENSMUST00000211272.2
|
Numa1
|
nuclear mitotic apparatus protein 1 |
| chr11_-_43317017 | 0.65 |
ENSMUST00000101340.11
ENSMUST00000118368.8 ENSMUST00000020685.10 ENSMUST00000020687.15 |
Pttg1
|
pituitary tumor-transforming gene 1 |
| chr19_-_56536443 | 0.65 |
ENSMUST00000182059.2
|
Dclre1a
|
DNA cross-link repair 1A |
| chr5_-_110987604 | 0.64 |
ENSMUST00000056937.12
|
Hscb
|
HscB iron-sulfur cluster co-chaperone |
| chr4_-_60377932 | 0.63 |
ENSMUST00000107506.9
ENSMUST00000122381.8 ENSMUST00000118759.8 ENSMUST00000132829.3 |
Mup9
|
major urinary protein 9 |
| chr4_-_60222580 | 0.63 |
ENSMUST00000095058.5
ENSMUST00000163931.8 |
Mup8
|
major urinary protein 8 |
| chr13_-_19521337 | 0.63 |
ENSMUST00000103563.3
|
Trgv2
|
T cell receptor gamma variable 2 |
| chr5_-_124492734 | 0.60 |
ENSMUST00000031341.11
|
Cdk2ap1
|
CDK2 (cyclin-dependent kinase 2)-associated protein 1 |
| chr13_-_46881388 | 0.59 |
ENSMUST00000021803.10
|
Nup153
|
nucleoporin 153 |
| chr2_+_103799873 | 0.59 |
ENSMUST00000123437.8
|
Lmo2
|
LIM domain only 2 |
| chr4_+_62398262 | 0.58 |
ENSMUST00000030088.12
ENSMUST00000107449.4 |
Bspry
|
B-box and SPRY domain containing |
| chr7_+_98089623 | 0.57 |
ENSMUST00000206435.2
|
Gucy2d
|
guanylate cyclase 2d |
| chr2_+_85420854 | 0.57 |
ENSMUST00000052307.5
|
Olfr998
|
olfactory receptor 998 |
| chr10_+_122284404 | 0.56 |
ENSMUST00000020323.7
|
Avpr1a
|
arginine vasopressin receptor 1A |
| chr11_-_68864666 | 0.56 |
ENSMUST00000038644.5
|
Rangrf
|
RAN guanine nucleotide release factor |
| chr17_+_8591228 | 0.51 |
ENSMUST00000163578.8
|
T2
|
brachyury 2 |
| chr10_-_116417333 | 0.50 |
ENSMUST00000218744.2
ENSMUST00000105267.8 ENSMUST00000105265.8 ENSMUST00000167706.8 ENSMUST00000168036.8 ENSMUST00000169921.8 ENSMUST00000020374.6 |
Cnot2
|
CCR4-NOT transcription complex, subunit 2 |
| chr5_-_44957016 | 0.50 |
ENSMUST00000199256.5
|
Ldb2
|
LIM domain binding 2 |
| chrX_+_162694397 | 0.49 |
ENSMUST00000140845.2
|
Ap1s2
|
adaptor-related protein complex 1, sigma 2 subunit |
| chr14_+_51366306 | 0.45 |
ENSMUST00000226210.2
|
Rnase6
|
ribonuclease, RNase A family, 6 |
| chr18_-_43610829 | 0.45 |
ENSMUST00000057110.11
|
Eif3j2
|
eukaryotic translation initiation factor 3, subunit J2 |
| chr13_-_103901010 | 0.43 |
ENSMUST00000210489.2
|
Srek1
|
splicing regulatory glutamine/lysine-rich protein 1 |
| chr10_-_53506038 | 0.42 |
ENSMUST00000218549.3
|
Mcm9
|
minichromosome maintenance 9 homologous recombination repair factor |
| chr9_-_48391838 | 0.42 |
ENSMUST00000216470.2
ENSMUST00000217037.2 ENSMUST00000034524.5 ENSMUST00000213895.2 |
Rexo2
|
RNA exonuclease 2 |
| chr6_+_134897473 | 0.42 |
ENSMUST00000204807.2
|
Cdkn1b
|
cyclin-dependent kinase inhibitor 1B |
| chr13_+_83672965 | 0.40 |
ENSMUST00000199432.5
ENSMUST00000198069.5 ENSMUST00000197681.5 ENSMUST00000197722.5 ENSMUST00000197938.5 |
Mef2c
|
myocyte enhancer factor 2C |
| chr1_+_60948149 | 0.40 |
ENSMUST00000027164.9
|
Ctla4
|
cytotoxic T-lymphocyte-associated protein 4 |
| chr7_+_5037117 | 0.38 |
ENSMUST00000076791.4
|
4632433K11Rik
|
RIKEN cDNA 4632433K11 gene |
| chr17_+_26471870 | 0.37 |
ENSMUST00000025023.15
|
Luc7l
|
Luc7-like |
| chr9_-_103242096 | 0.36 |
ENSMUST00000116517.9
|
Cdv3
|
carnitine deficiency-associated gene expressed in ventricle 3 |
| chr5_+_88523967 | 0.35 |
ENSMUST00000073363.2
|
Amtn
|
amelotin |
| chr13_-_21964747 | 0.34 |
ENSMUST00000080511.3
|
H1f5
|
H1.5 linker histone, cluster member |
| chr4_+_32238950 | 0.33 |
ENSMUST00000037416.13
|
Bach2
|
BTB and CNC homology, basic leucine zipper transcription factor 2 |
| chrX_-_58612709 | 0.33 |
ENSMUST00000124402.2
|
Fgf13
|
fibroblast growth factor 13 |
| chr16_-_43484494 | 0.33 |
ENSMUST00000096065.6
|
Tigit
|
T cell immunoreceptor with Ig and ITIM domains |
| chr2_-_164231015 | 0.32 |
ENSMUST00000167427.2
|
Slpi
|
secretory leukocyte peptidase inhibitor |
| chr6_+_134897364 | 0.31 |
ENSMUST00000067327.11
ENSMUST00000003115.9 |
Cdkn1b
|
cyclin-dependent kinase inhibitor 1B |
| chr4_+_109200225 | 0.31 |
ENSMUST00000030281.12
|
Eps15
|
epidermal growth factor receptor pathway substrate 15 |
| chr2_+_89760155 | 0.31 |
ENSMUST00000111516.2
|
Olfr1258
|
olfactory receptor 1258 |
| chr3_+_85946145 | 0.31 |
ENSMUST00000238331.2
|
Sh3d19
|
SH3 domain protein D19 |
| chr2_-_4656870 | 0.30 |
ENSMUST00000192470.2
ENSMUST00000035721.14 ENSMUST00000152362.7 |
Prpf18
|
pre-mRNA processing factor 18 |
| chrX_-_105528503 | 0.30 |
ENSMUST00000138724.8
ENSMUST00000149331.2 |
Fndc3c1
|
fibronectin type III domain containing 3C1 |
| chr2_-_103609703 | 0.28 |
ENSMUST00000143188.2
|
Caprin1
|
cell cycle associated protein 1 |
| chr16_-_45975440 | 0.28 |
ENSMUST00000059524.7
|
Gm4737
|
predicted gene 4737 |
| chr1_-_132958299 | 0.27 |
ENSMUST00000190807.2
|
Mdm4
|
transformed mouse 3T3 cell double minute 4 |
| chr5_+_4073343 | 0.27 |
ENSMUST00000238634.2
|
Akap9
|
A kinase (PRKA) anchor protein (yotiao) 9 |
| chr8_+_84724130 | 0.26 |
ENSMUST00000095228.5
|
Samd1
|
sterile alpha motif domain containing 1 |
| chr4_-_101750986 | 0.26 |
ENSMUST00000051043.4
|
C130073F10Rik
|
RIKEN cDNA C130073F10 gene |
| chr9_-_103079312 | 0.25 |
ENSMUST00000035157.10
|
Srprb
|
signal recognition particle receptor, B subunit |
| chr4_+_147445744 | 0.21 |
ENSMUST00000133078.8
ENSMUST00000154154.2 |
Zfp978
|
zinc finger protein 978 |
| chr1_-_86038394 | 0.21 |
ENSMUST00000155077.2
|
Htr2b
|
5-hydroxytryptamine (serotonin) receptor 2B |
| chr1_-_36982747 | 0.20 |
ENSMUST00000185964.3
|
Tmem131
|
transmembrane protein 131 |
| chr7_-_106491314 | 0.20 |
ENSMUST00000088687.3
|
Olfr707
|
olfactory receptor 707 |
| chr2_-_86528739 | 0.19 |
ENSMUST00000214141.2
|
Olfr1087
|
olfactory receptor 1087 |
| chr4_-_61259997 | 0.19 |
ENSMUST00000071005.9
ENSMUST00000075206.12 |
Mup14
|
major urinary protein 14 |
| chr12_+_87921198 | 0.18 |
ENSMUST00000110145.12
ENSMUST00000181843.2 ENSMUST00000180706.8 ENSMUST00000181394.8 ENSMUST00000181326.8 ENSMUST00000181300.2 |
Gm2042
|
predicted gene 2042 |
| chr5_+_147366953 | 0.18 |
ENSMUST00000031651.15
|
Pan3
|
PAN3 poly(A) specific ribonuclease subunit |
| chr13_+_119599287 | 0.18 |
ENSMUST00000026519.10
ENSMUST00000225186.2 ENSMUST00000224081.2 ENSMUST00000224312.2 |
4833420G17Rik
|
RIKEN cDNA 4833420G17 gene |
| chr2_-_93988229 | 0.18 |
ENSMUST00000028619.5
|
Hsd17b12
|
hydroxysteroid (17-beta) dehydrogenase 12 |
| chr19_+_8595369 | 0.17 |
ENSMUST00000010250.4
|
Slc22a6
|
solute carrier family 22 (organic anion transporter), member 6 |
| chr19_-_56536646 | 0.15 |
ENSMUST00000182276.2
|
Dclre1a
|
DNA cross-link repair 1A |
| chr6_+_134807170 | 0.13 |
ENSMUST00000111937.2
|
Crebl2
|
cAMP responsive element binding protein-like 2 |
| chr4_-_118792037 | 0.11 |
ENSMUST00000081960.5
|
Olfr1328
|
olfactory receptor 1328 |
| chr6_+_30401864 | 0.11 |
ENSMUST00000068240.13
ENSMUST00000068259.10 ENSMUST00000132581.8 |
Klhdc10
|
kelch domain containing 10 |
| chr4_-_118687635 | 0.10 |
ENSMUST00000076019.4
|
Olfr1333
|
olfactory receptor 1333 |
| chr7_-_104296512 | 0.10 |
ENSMUST00000210641.2
ENSMUST00000089296.3 |
Olfr658
|
olfactory receptor 658 |
| chr1_+_173964312 | 0.09 |
ENSMUST00000053941.4
|
Olfr424
|
olfactory receptor 424 |
| chr3_+_138233004 | 0.08 |
ENSMUST00000196990.5
ENSMUST00000200239.5 ENSMUST00000200100.2 |
Eif4e
|
eukaryotic translation initiation factor 4E |
| chr7_-_103393740 | 0.07 |
ENSMUST00000216300.2
|
Olfr629
|
olfactory receptor 629 |
| chr5_-_87634665 | 0.07 |
ENSMUST00000201519.2
|
Gm43638
|
predicted gene 43638 |
| chr2_-_34261121 | 0.06 |
ENSMUST00000127353.3
ENSMUST00000141653.3 |
Pbx3
|
pre B cell leukemia homeobox 3 |
| chr10_-_130116088 | 0.06 |
ENSMUST00000061571.5
|
Neurod4
|
neurogenic differentiation 4 |
| chr16_+_8288627 | 0.05 |
ENSMUST00000046470.16
ENSMUST00000150790.2 ENSMUST00000142899.2 |
Mettl22
|
methyltransferase like 22 |
| chr6_+_83142902 | 0.04 |
ENSMUST00000077407.12
ENSMUST00000113913.8 ENSMUST00000130212.8 |
Dctn1
|
dynactin 1 |
| chr5_-_21150470 | 0.03 |
ENSMUST00000197089.2
|
Rsbn1l
|
round spermatid basic protein 1-like |
| chr19_-_4384029 | 0.02 |
ENSMUST00000176653.2
|
Kdm2a
|
lysine (K)-specific demethylase 2A |
| chr9_-_39465349 | 0.01 |
ENSMUST00000215505.2
ENSMUST00000217227.2 |
Olfr958
|
olfactory receptor 958 |
| chr3_+_89366425 | 0.01 |
ENSMUST00000029564.12
|
Pmvk
|
phosphomevalonate kinase |
| chr9_+_88721217 | 0.01 |
ENSMUST00000163255.9
ENSMUST00000186363.2 |
Trim43c
|
tripartite motif-containing 43C |
| chr2_-_89502276 | 0.01 |
ENSMUST00000214639.2
ENSMUST00000214304.2 |
Olfr1251
|
olfactory receptor 1251 |
| chr12_-_32258469 | 0.00 |
ENSMUST00000085469.6
|
Pik3cg
|
phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma |
| chr12_+_17316546 | 0.00 |
ENSMUST00000057288.7
ENSMUST00000239402.2 |
Pdia6
|
protein disulfide isomerase associated 6 |
| chr4_-_132649798 | 0.00 |
ENSMUST00000097856.10
ENSMUST00000030696.11 |
Fam76a
|
family with sequence similarity 76, member A |
| chr3_-_100591906 | 0.00 |
ENSMUST00000130066.2
|
Man1a2
|
mannosidase, alpha, class 1A, member 2 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.4 | 10.1 | GO:0090673 | endothelial cell-matrix adhesion(GO:0090673) |
| 1.8 | 5.3 | GO:0038189 | neuropilin signaling pathway(GO:0038189) VEGF-activated neuropilin signaling pathway(GO:0038190) |
| 1.4 | 4.3 | GO:0046967 | antigen processing and presentation of endogenous peptide antigen via MHC class Ib(GO:0002476) antigen processing and presentation of exogenous peptide antigen via MHC class Ib(GO:0002477) antigen processing and presentation of exogenous protein antigen via MHC class Ib, TAP-dependent(GO:0002481) cytosol to ER transport(GO:0046967) |
| 1.1 | 4.6 | GO:2000473 | regulation of hematopoietic stem cell migration(GO:2000471) positive regulation of hematopoietic stem cell migration(GO:2000473) |
| 1.0 | 2.9 | GO:0061300 | cerebellum vasculature development(GO:0061300) |
| 0.9 | 5.6 | GO:1903609 | negative regulation of peptidyl-tyrosine autophosphorylation(GO:1900085) negative regulation of inward rectifier potassium channel activity(GO:1903609) |
| 0.7 | 2.1 | GO:0060672 | epithelial cell differentiation involved in embryonic placenta development(GO:0060671) epithelial cell morphogenesis involved in placental branching(GO:0060672) |
| 0.7 | 2.1 | GO:0070104 | negative regulation of interleukin-6-mediated signaling pathway(GO:0070104) |
| 0.6 | 3.2 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.6 | 2.5 | GO:0051329 | interphase(GO:0051325) mitotic interphase(GO:0051329) |
| 0.6 | 5.3 | GO:0038044 | transforming growth factor-beta secretion(GO:0038044) |
| 0.6 | 2.3 | GO:0045415 | negative regulation of interleukin-8 biosynthetic process(GO:0045415) |
| 0.5 | 4.2 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.5 | 2.0 | GO:0019343 | cysteine biosynthetic process via cystathionine(GO:0019343) |
| 0.4 | 1.8 | GO:0060849 | regulation of transcription involved in lymphatic endothelial cell fate commitment(GO:0060849) |
| 0.3 | 7.0 | GO:0048739 | cardiac muscle fiber development(GO:0048739) |
| 0.3 | 2.0 | GO:0006116 | NADH oxidation(GO:0006116) |
| 0.3 | 2.5 | GO:0031179 | peptide modification(GO:0031179) |
| 0.3 | 6.1 | GO:0006027 | glycosaminoglycan catabolic process(GO:0006027) |
| 0.3 | 1.9 | GO:0002430 | complement receptor mediated signaling pathway(GO:0002430) tolerance induction dependent upon immune response(GO:0002461) |
| 0.3 | 2.6 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.2 | 1.2 | GO:0006432 | phenylalanyl-tRNA aminoacylation(GO:0006432) |
| 0.2 | 1.0 | GO:0061228 | regulation of pronephros size(GO:0035565) mesonephros morphogenesis(GO:0061206) mesonephric nephron development(GO:0061215) mesonephric nephron morphogenesis(GO:0061228) mesenchymal stem cell maintenance involved in mesonephric nephron morphogenesis(GO:0061235) regulation of mesenchymal cell apoptotic process involved in mesonephric nephron morphogenesis(GO:0061295) negative regulation of mesenchymal cell apoptotic process involved in mesonephric nephron morphogenesis(GO:0061296) mesenchymal cell apoptotic process involved in mesonephric nephron morphogenesis(GO:1901146) |
| 0.2 | 0.7 | GO:0042264 | peptidyl-aspartic acid hydroxylation(GO:0042264) |
| 0.2 | 1.6 | GO:0060023 | soft palate development(GO:0060023) |
| 0.2 | 2.2 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
| 0.2 | 0.6 | GO:0032240 | negative regulation of nucleobase-containing compound transport(GO:0032240) negative regulation of RNA export from nucleus(GO:0046832) |
| 0.2 | 3.1 | GO:0097067 | cellular response to thyroid hormone stimulus(GO:0097067) |
| 0.2 | 0.7 | GO:0016999 | antibiotic metabolic process(GO:0016999) |
| 0.2 | 2.4 | GO:0060272 | embryonic skeletal joint morphogenesis(GO:0060272) |
| 0.2 | 9.0 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.2 | 0.7 | GO:1902365 | regulation of spindle elongation(GO:0032887) regulation of mitotic spindle elongation(GO:0032888) anastral spindle assembly(GO:0055048) protein localization to spindle pole body(GO:0071988) regulation of protein localization to spindle pole body(GO:1902363) positive regulation of protein localization to spindle pole body(GO:1902365) positive regulation of mitotic spindle elongation(GO:1902846) |
| 0.2 | 1.4 | GO:0048133 | germ-line stem cell division(GO:0042078) male germ-line stem cell asymmetric division(GO:0048133) germline stem cell asymmetric division(GO:0098728) |
| 0.1 | 1.5 | GO:0048208 | vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) |
| 0.1 | 3.4 | GO:0048387 | negative regulation of retinoic acid receptor signaling pathway(GO:0048387) |
| 0.1 | 1.0 | GO:0007527 | adult somatic muscle development(GO:0007527) |
| 0.1 | 0.6 | GO:0002125 | maternal aggressive behavior(GO:0002125) |
| 0.1 | 3.0 | GO:0006883 | cellular sodium ion homeostasis(GO:0006883) |
| 0.1 | 0.7 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 0.1 | 0.3 | GO:0070175 | positive regulation of enamel mineralization(GO:0070175) |
| 0.1 | 1.5 | GO:1902510 | regulation of apoptotic DNA fragmentation(GO:1902510) |
| 0.1 | 1.5 | GO:0090050 | positive regulation of cell migration involved in sprouting angiogenesis(GO:0090050) |
| 0.1 | 0.7 | GO:1904222 | regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904217) positive regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904219) positive regulation of serine C-palmitoyltransferase activity(GO:1904222) |
| 0.1 | 0.8 | GO:0031848 | protection from non-homologous end joining at telomere(GO:0031848) |
| 0.1 | 0.5 | GO:0014028 | notochord formation(GO:0014028) |
| 0.1 | 0.4 | GO:0045590 | negative regulation of regulatory T cell differentiation(GO:0045590) |
| 0.1 | 0.6 | GO:0097428 | protein maturation by iron-sulfur cluster transfer(GO:0097428) |
| 0.1 | 2.7 | GO:0019731 | antibacterial humoral response(GO:0019731) |
| 0.1 | 0.7 | GO:0010606 | positive regulation of cytoplasmic mRNA processing body assembly(GO:0010606) |
| 0.1 | 2.0 | GO:0010669 | epithelial structure maintenance(GO:0010669) |
| 0.1 | 0.2 | GO:0000350 | generation of catalytic spliceosome for second transesterification step(GO:0000350) |
| 0.1 | 0.7 | GO:1904706 | negative regulation of vascular smooth muscle cell proliferation(GO:1904706) |
| 0.1 | 1.1 | GO:0030049 | muscle filament sliding(GO:0030049) |
| 0.1 | 1.2 | GO:0051451 | myoblast migration(GO:0051451) |
| 0.1 | 0.3 | GO:0019065 | receptor-mediated endocytosis of virus by host cell(GO:0019065) endocytosis involved in viral entry into host cell(GO:0075509) |
| 0.1 | 0.7 | GO:0034723 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.1 | 1.0 | GO:0035278 | miRNA mediated inhibition of translation(GO:0035278) negative regulation of translation, ncRNA-mediated(GO:0040033) regulation of translation, ncRNA-mediated(GO:0045974) |
| 0.1 | 0.2 | GO:0007208 | phospholipase C-activating serotonin receptor signaling pathway(GO:0007208) |
| 0.1 | 0.6 | GO:0086028 | bundle of His cell to Purkinje myocyte signaling(GO:0086028) bundle of His cell action potential(GO:0086043) |
| 0.0 | 3.1 | GO:0046426 | negative regulation of JAK-STAT cascade(GO:0046426) negative regulation of STAT cascade(GO:1904893) |
| 0.0 | 0.7 | GO:0032486 | Rap protein signal transduction(GO:0032486) |
| 0.0 | 2.3 | GO:0070232 | regulation of T cell apoptotic process(GO:0070232) |
| 0.0 | 0.4 | GO:0003138 | primary heart field specification(GO:0003138) sinoatrial valve development(GO:0003172) sinoatrial valve morphogenesis(GO:0003185) |
| 0.0 | 1.6 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.0 | 0.8 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.0 | 0.7 | GO:0007064 | mitotic sister chromatid cohesion(GO:0007064) |
| 0.0 | 3.0 | GO:0045727 | positive regulation of translation(GO:0045727) |
| 0.0 | 0.0 | GO:0090063 | positive regulation of microtubule nucleation(GO:0090063) |
| 0.0 | 0.3 | GO:0003283 | atrial septum development(GO:0003283) |
| 0.0 | 0.3 | GO:0032695 | negative regulation of interleukin-12 production(GO:0032695) |
| 0.0 | 0.3 | GO:0051044 | positive regulation of membrane protein ectodomain proteolysis(GO:0051044) |
| 0.0 | 0.6 | GO:0006182 | cGMP biosynthetic process(GO:0006182) |
| 0.0 | 0.2 | GO:0015747 | urate transport(GO:0015747) |
| 0.0 | 1.1 | GO:0051881 | regulation of mitochondrial membrane potential(GO:0051881) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 10.1 | GO:0030485 | smooth muscle contractile fiber(GO:0030485) |
| 1.1 | 5.3 | GO:0034685 | integrin alphav-beta6 complex(GO:0034685) |
| 0.9 | 2.6 | GO:0043540 | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase complex(GO:0043540) |
| 0.7 | 2.1 | GO:0005900 | oncostatin-M receptor complex(GO:0005900) |
| 0.5 | 4.3 | GO:0042825 | TAP complex(GO:0042825) |
| 0.5 | 9.0 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
| 0.4 | 2.0 | GO:0030868 | smooth endoplasmic reticulum membrane(GO:0030868) smooth endoplasmic reticulum part(GO:0097425) |
| 0.3 | 3.0 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.3 | 5.6 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 0.3 | 1.4 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.2 | 0.7 | GO:0048269 | methionine adenosyltransferase complex(GO:0048269) |
| 0.2 | 4.2 | GO:0098644 | complex of collagen trimers(GO:0098644) |
| 0.2 | 2.8 | GO:0012510 | trans-Golgi network transport vesicle membrane(GO:0012510) |
| 0.1 | 3.9 | GO:0032982 | myosin filament(GO:0032982) |
| 0.1 | 0.7 | GO:0061673 | mitotic spindle astral microtubule(GO:0061673) |
| 0.1 | 1.2 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.1 | 1.0 | GO:0035068 | micro-ribonucleoprotein complex(GO:0035068) |
| 0.1 | 0.6 | GO:0044615 | nuclear pore nuclear basket(GO:0044615) |
| 0.1 | 0.3 | GO:0044307 | dendritic branch(GO:0044307) |
| 0.0 | 0.8 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.0 | 0.5 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.0 | 6.6 | GO:0005604 | basement membrane(GO:0005604) |
| 0.0 | 3.5 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.0 | 1.0 | GO:0070938 | contractile ring(GO:0070938) |
| 0.0 | 2.3 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.0 | 2.5 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 0.7 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.0 | 0.2 | GO:0031251 | PAN complex(GO:0031251) |
| 0.0 | 8.0 | GO:0030018 | Z disc(GO:0030018) |
| 0.0 | 0.3 | GO:0005785 | signal recognition particle receptor complex(GO:0005785) |
| 0.0 | 1.6 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.0 | 3.7 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 0.4 | GO:0042555 | MCM complex(GO:0042555) |
| 0.0 | 0.6 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.0 | 0.3 | GO:0005852 | eukaryotic translation initiation factor 3 complex(GO:0005852) |
| 0.0 | 1.5 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.0 | 0.7 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.0 | 1.1 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.0 | 0.4 | GO:0098636 | protein complex involved in cell adhesion(GO:0098636) |
| 0.0 | 0.3 | GO:0005682 | U5 snRNP(GO:0005682) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.8 | 5.3 | GO:0043183 | vascular endothelial growth factor receptor 1 binding(GO:0043183) |
| 1.4 | 4.3 | GO:0015433 | peptide antigen-transporting ATPase activity(GO:0015433) tapasin binding(GO:0046980) |
| 0.9 | 5.6 | GO:0070320 | inward rectifier potassium channel inhibitor activity(GO:0070320) |
| 0.7 | 2.1 | GO:0019981 | interleukin-6 receptor activity(GO:0004915) interleukin-6 binding(GO:0019981) |
| 0.6 | 9.0 | GO:0017176 | phosphatidylinositol N-acetylglucosaminyltransferase activity(GO:0017176) |
| 0.6 | 2.5 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 0.5 | 3.4 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.5 | 2.3 | GO:0001010 | transcription factor activity, sequence-specific DNA binding transcription factor recruiting(GO:0001010) |
| 0.3 | 1.5 | GO:0004174 | electron-transferring-flavoprotein dehydrogenase activity(GO:0004174) oxidoreductase activity, acting on the CH-NH group of donors, quinone or similar compound as acceptor(GO:0016649) |
| 0.3 | 2.6 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) |
| 0.2 | 5.9 | GO:0070742 | C2H2 zinc finger domain binding(GO:0070742) |
| 0.2 | 6.1 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.2 | 1.9 | GO:0004875 | complement receptor activity(GO:0004875) |
| 0.2 | 1.2 | GO:0004826 | phenylalanine-tRNA ligase activity(GO:0004826) |
| 0.2 | 2.0 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.1 | 4.6 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.1 | 2.0 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.1 | 2.1 | GO:0001161 | intronic transcription regulatory region sequence-specific DNA binding(GO:0001161) |
| 0.1 | 2.0 | GO:0016846 | carbon-sulfur lyase activity(GO:0016846) |
| 0.1 | 2.9 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.1 | 0.8 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.1 | 0.6 | GO:0031893 | vasopressin receptor binding(GO:0031893) |
| 0.1 | 1.3 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
| 0.1 | 0.3 | GO:0000386 | second spliceosomal transesterification activity(GO:0000386) |
| 0.1 | 7.0 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.1 | 0.7 | GO:0071532 | ankyrin repeat binding(GO:0071532) |
| 0.1 | 1.7 | GO:0001206 | transcriptional repressor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001206) |
| 0.1 | 0.8 | GO:0035312 | 5'-3' exodeoxyribonuclease activity(GO:0035312) |
| 0.1 | 0.5 | GO:0001226 | RNA polymerase II transcription corepressor binding(GO:0001226) |
| 0.1 | 0.6 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.1 | 1.8 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.1 | 0.2 | GO:0016509 | long-chain-3-hydroxyacyl-CoA dehydrogenase activity(GO:0016509) |
| 0.1 | 8.0 | GO:0004860 | protein kinase inhibitor activity(GO:0004860) |
| 0.1 | 0.9 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.1 | 0.7 | GO:0015194 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.1 | 1.2 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.0 | 3.1 | GO:0043425 | bHLH transcription factor binding(GO:0043425) |
| 0.0 | 0.3 | GO:0005047 | signal recognition particle binding(GO:0005047) |
| 0.0 | 1.0 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.0 | 0.6 | GO:0043495 | protein anchor(GO:0043495) |
| 0.0 | 3.2 | GO:0004714 | transmembrane receptor protein tyrosine kinase activity(GO:0004714) |
| 0.0 | 0.7 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.0 | 9.2 | GO:0019903 | protein phosphatase binding(GO:0019903) |
| 0.0 | 1.1 | GO:0000146 | microfilament motor activity(GO:0000146) |
| 0.0 | 5.3 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.8 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 0.0 | 0.1 | GO:0031370 | eukaryotic initiation factor 4G binding(GO:0031370) |
| 0.0 | 0.6 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.0 | 3.0 | GO:0004540 | ribonuclease activity(GO:0004540) |
| 0.0 | 0.6 | GO:0004383 | guanylate cyclase activity(GO:0004383) |
| 0.0 | 1.5 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.4 | GO:0035198 | miRNA binding(GO:0035198) |
| 0.0 | 2.9 | GO:0003774 | motor activity(GO:0003774) |
| 0.0 | 0.2 | GO:0051378 | serotonin binding(GO:0051378) |
| 0.0 | 0.6 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.0 | 1.5 | GO:0000980 | RNA polymerase II distal enhancer sequence-specific DNA binding(GO:0000980) |
| 0.0 | 0.2 | GO:0031404 | chloride ion binding(GO:0031404) |
| 0.0 | 0.6 | GO:0008536 | Ran GTPase binding(GO:0008536) |
| 0.0 | 1.2 | GO:0032947 | protein complex scaffold(GO:0032947) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 5.3 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.2 | 5.3 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.2 | 10.1 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.2 | 5.6 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.1 | 5.1 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.1 | 2.9 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.1 | 4.6 | SIG PIP3 SIGNALING IN B LYMPHOCYTES | Genes related to PIP3 signaling in B lymphocytes |
| 0.1 | 0.7 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.1 | 2.5 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.1 | 2.4 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.1 | 5.3 | PID SHP2 PATHWAY | SHP2 signaling |
| 0.1 | 2.2 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 8.1 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 1.8 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 2.4 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 2.3 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.0 | 0.7 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.0 | 0.6 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.0 | 1.4 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 1.5 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.0 | 0.6 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.0 | 1.8 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 1.2 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.0 | 0.9 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.0 | 0.4 | PID MAPK TRK PATHWAY | Trk receptor signaling mediated by the MAPK pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 5.3 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.5 | 6.1 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.4 | 4.6 | REACTOME PHOSPHORYLATION OF CD3 AND TCR ZETA CHAINS | Genes involved in Phosphorylation of CD3 and TCR zeta chains |
| 0.4 | 4.2 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.3 | 9.0 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.3 | 9.0 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.2 | 4.3 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.2 | 2.2 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.2 | 2.0 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.2 | 4.5 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.1 | 2.1 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.1 | 3.2 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.1 | 2.3 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.1 | 3.0 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.1 | 4.6 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.1 | 2.8 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.1 | 6.4 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.1 | 1.2 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.0 | 5.3 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 0.7 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.0 | 0.5 | REACTOME NEF MEDIATED DOWNREGULATION OF MHC CLASS I COMPLEX CELL SURFACE EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.0 | 1.0 | REACTOME REGULATORY RNA PATHWAYS | Genes involved in Regulatory RNA pathways |
| 0.0 | 0.7 | REACTOME AKT PHOSPHORYLATES TARGETS IN THE CYTOSOL | Genes involved in AKT phosphorylates targets in the cytosol |
| 0.0 | 0.7 | REACTOME REGULATION OF HYPOXIA INDUCIBLE FACTOR HIF BY OXYGEN | Genes involved in Regulation of Hypoxia-inducible Factor (HIF) by Oxygen |
| 0.0 | 0.4 | REACTOME CTLA4 INHIBITORY SIGNALING | Genes involved in CTLA4 inhibitory signaling |
| 0.0 | 2.9 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 2.1 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.0 | 0.6 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 0.2 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.0 | 1.1 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 0.7 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.0 | 1.2 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.0 | 2.4 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 0.6 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.0 | 0.3 | REACTOME RAS ACTIVATION UOPN CA2 INFUX THROUGH NMDA RECEPTOR | Genes involved in Ras activation uopn Ca2+ infux through NMDA receptor |
| 0.0 | 0.2 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 0.0 | 0.4 | REACTOME REGULATION OF BETA CELL DEVELOPMENT | Genes involved in Regulation of beta-cell development |