Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
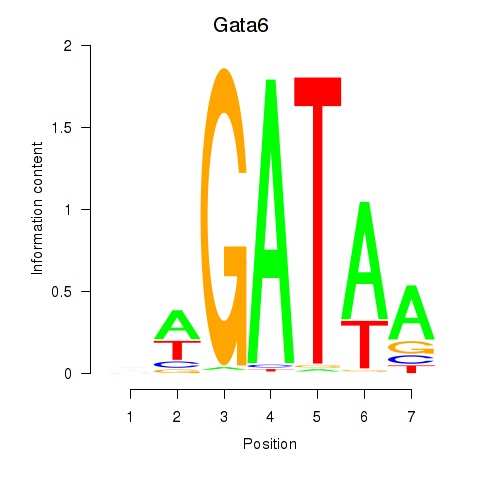
Results for Gata6
Z-value: 0.56
Transcription factors associated with Gata6
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Gata6
|
ENSMUSG00000005836.11 | Gata6 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Gata6 | mm39_v1_chr18_+_11052458_11052479 | 0.32 | 7.0e-03 | Click! |
Activity profile of Gata6 motif
Sorted Z-values of Gata6 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Gata6
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_+_58658838 | 5.95 |
ENSMUST00000238108.2
|
Pnlip
|
pancreatic lipase |
| chr17_+_44445659 | 5.92 |
ENSMUST00000239215.2
|
Clic5
|
chloride intracellular channel 5 |
| chr19_+_58658779 | 5.90 |
ENSMUST00000057270.9
|
Pnlip
|
pancreatic lipase |
| chr10_+_53213763 | 4.33 |
ENSMUST00000219491.2
ENSMUST00000163319.9 ENSMUST00000220197.2 ENSMUST00000046221.8 ENSMUST00000218468.2 ENSMUST00000219921.2 |
Pln
|
phospholamban |
| chr6_+_30639217 | 3.76 |
ENSMUST00000031806.10
|
Cpa1
|
carboxypeptidase A1, pancreatic |
| chr10_+_116881246 | 3.44 |
ENSMUST00000073834.5
|
Lrrc10
|
leucine rich repeat containing 10 |
| chr4_-_137157824 | 2.95 |
ENSMUST00000102522.5
|
Cela3b
|
chymotrypsin-like elastase family, member 3B |
| chr2_-_17465410 | 2.94 |
ENSMUST00000145492.2
|
Nebl
|
nebulette |
| chr7_+_141276575 | 2.64 |
ENSMUST00000185406.8
|
Muc2
|
mucin 2 |
| chr8_-_86091970 | 2.52 |
ENSMUST00000121972.8
|
Mylk3
|
myosin light chain kinase 3 |
| chr18_+_11052458 | 2.38 |
ENSMUST00000047762.10
|
Gata6
|
GATA binding protein 6 |
| chr8_-_86091946 | 2.34 |
ENSMUST00000034133.14
|
Mylk3
|
myosin light chain kinase 3 |
| chr2_+_90948481 | 2.33 |
ENSMUST00000137942.8
ENSMUST00000111430.10 ENSMUST00000169776.2 |
Mybpc3
|
myosin binding protein C, cardiac |
| chr7_-_135130374 | 2.28 |
ENSMUST00000053716.8
|
Clrn3
|
clarin 3 |
| chr11_-_46280281 | 2.15 |
ENSMUST00000101306.4
|
Itk
|
IL2 inducible T cell kinase |
| chr2_+_110427643 | 2.08 |
ENSMUST00000045972.13
ENSMUST00000111026.3 |
Slc5a12
|
solute carrier family 5 (sodium/glucose cotransporter), member 12 |
| chr6_-_136852792 | 2.05 |
ENSMUST00000032342.3
|
Mgp
|
matrix Gla protein |
| chr14_-_70761507 | 1.95 |
ENSMUST00000022692.5
|
Sftpc
|
surfactant associated protein C |
| chr10_+_101517348 | 1.75 |
ENSMUST00000179929.8
ENSMUST00000219195.2 ENSMUST00000127504.9 |
Mgat4c
|
MGAT4 family, member C |
| chr3_-_75177378 | 1.73 |
ENSMUST00000039047.5
|
Serpini2
|
serine (or cysteine) peptidase inhibitor, clade I, member 2 |
| chr1_+_12762501 | 1.46 |
ENSMUST00000177608.8
ENSMUST00000180062.8 |
Sulf1
|
sulfatase 1 |
| chr8_+_46081213 | 1.37 |
ENSMUST00000130850.8
|
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr11_-_46280298 | 1.34 |
ENSMUST00000109237.9
|
Itk
|
IL2 inducible T cell kinase |
| chr8_+_57774010 | 1.29 |
ENSMUST00000040104.5
|
Hand2
|
heart and neural crest derivatives expressed 2 |
| chr11_-_46280336 | 1.19 |
ENSMUST00000020664.13
|
Itk
|
IL2 inducible T cell kinase |
| chr4_-_137137088 | 1.18 |
ENSMUST00000024200.7
|
Cela3a
|
chymotrypsin-like elastase family, member 3A |
| chr9_-_95697441 | 1.09 |
ENSMUST00000119760.2
|
Pls1
|
plastin 1 (I-isoform) |
| chr1_+_45350698 | 1.04 |
ENSMUST00000087883.13
|
Col3a1
|
collagen, type III, alpha 1 |
| chr9_+_48896765 | 1.01 |
ENSMUST00000047349.8
|
Usp28
|
ubiquitin specific peptidase 28 |
| chr5_+_90920353 | 0.99 |
ENSMUST00000202625.2
|
Pf4
|
platelet factor 4 |
| chr6_+_58810674 | 0.97 |
ENSMUST00000041401.11
|
Herc3
|
hect domain and RLD 3 |
| chr3_+_79793237 | 0.92 |
ENSMUST00000029567.9
|
Gask1b
|
golgi associated kinase 1B |
| chr3_+_108272205 | 0.76 |
ENSMUST00000090563.7
|
Mybphl
|
myosin binding protein H-like |
| chr16_+_35861554 | 0.70 |
ENSMUST00000042203.10
|
Wdr5b
|
WD repeat domain 5B |
| chr17_+_34823790 | 0.67 |
ENSMUST00000173242.8
|
Agpat1
|
1-acylglycerol-3-phosphate O-acyltransferase 1 (lysophosphatidic acid acyltransferase, alpha) |
| chr10_+_115653152 | 0.66 |
ENSMUST00000080630.11
ENSMUST00000179196.3 ENSMUST00000035563.15 |
Tspan8
|
tetraspanin 8 |
| chr13_+_19398273 | 0.66 |
ENSMUST00000103558.3
|
Trgc1
|
T cell receptor gamma, constant 1 |
| chr10_-_62178453 | 0.62 |
ENSMUST00000143179.2
ENSMUST00000130422.8 |
Hk1
|
hexokinase 1 |
| chr3_-_30194559 | 0.60 |
ENSMUST00000108271.10
|
Mecom
|
MDS1 and EVI1 complex locus |
| chr9_+_22011488 | 0.53 |
ENSMUST00000213607.2
|
Cnn1
|
calponin 1 |
| chr11_+_87471867 | 0.48 |
ENSMUST00000018544.12
ENSMUST00000063156.11 ENSMUST00000107960.8 |
Septin4
|
septin 4 |
| chr3_+_90201388 | 0.46 |
ENSMUST00000199607.5
|
Gatad2b
|
GATA zinc finger domain containing 2B |
| chr10_+_101517556 | 0.46 |
ENSMUST00000156751.8
|
Mgat4c
|
MGAT4 family, member C |
| chr1_-_144052997 | 0.43 |
ENSMUST00000111941.2
ENSMUST00000052375.8 |
Rgs13
|
regulator of G-protein signaling 13 |
| chr17_+_88933957 | 0.42 |
ENSMUST00000163588.8
ENSMUST00000064035.13 |
Ston1
|
stonin 1 |
| chr4_+_150233362 | 0.42 |
ENSMUST00000059893.8
|
Slc2a7
|
solute carrier family 2 (facilitated glucose transporter), member 7 |
| chr11_-_70111796 | 0.41 |
ENSMUST00000060010.3
ENSMUST00000190533.2 |
Slc16a13
|
solute carrier family 16 (monocarboxylic acid transporters), member 13 |
| chr2_-_59778560 | 0.40 |
ENSMUST00000153136.2
|
Baz2b
|
bromodomain adjacent to zinc finger domain, 2B |
| chr16_-_95260104 | 0.37 |
ENSMUST00000176345.10
ENSMUST00000121809.11 ENSMUST00000233664.2 ENSMUST00000122199.10 |
Erg
|
ETS transcription factor |
| chr2_-_77000936 | 0.35 |
ENSMUST00000164114.9
ENSMUST00000049544.14 |
Ccdc141
|
coiled-coil domain containing 141 |
| chr12_+_95658987 | 0.33 |
ENSMUST00000057324.4
|
Flrt2
|
fibronectin leucine rich transmembrane protein 2 |
| chrX_-_111608339 | 0.33 |
ENSMUST00000039887.4
|
Pof1b
|
premature ovarian failure 1B |
| chr2_-_7086018 | 0.33 |
ENSMUST00000114923.3
ENSMUST00000182706.8 |
Celf2
|
CUGBP, Elav-like family member 2 |
| chr13_-_103042554 | 0.33 |
ENSMUST00000171791.8
|
Mast4
|
microtubule associated serine/threonine kinase family member 4 |
| chr5_+_76988444 | 0.31 |
ENSMUST00000120639.9
ENSMUST00000163347.8 ENSMUST00000121851.2 |
Cracd
|
capping protein inhibiting regulator of actin |
| chr16_+_91444730 | 0.29 |
ENSMUST00000119368.8
ENSMUST00000114037.9 ENSMUST00000114036.9 ENSMUST00000122302.8 |
Son
|
Son DNA binding protein |
| chr2_-_7400690 | 0.28 |
ENSMUST00000182404.8
|
Celf2
|
CUGBP, Elav-like family member 2 |
| chr11_-_53598078 | 0.24 |
ENSMUST00000124352.2
ENSMUST00000020649.14 |
Rad50
|
RAD50 double strand break repair protein |
| chr9_+_62249730 | 0.22 |
ENSMUST00000156461.2
|
Anp32a
|
acidic (leucine-rich) nuclear phosphoprotein 32 family, member A |
| chr11_+_97917520 | 0.22 |
ENSMUST00000092425.11
|
Rpl19
|
ribosomal protein L19 |
| chr1_-_144651157 | 0.19 |
ENSMUST00000027603.4
|
Rgs18
|
regulator of G-protein signaling 18 |
| chr1_-_131161312 | 0.18 |
ENSMUST00000212202.2
|
Rassf5
|
Ras association (RalGDS/AF-6) domain family member 5 |
| chr2_+_143757193 | 0.17 |
ENSMUST00000103172.4
|
Dstn
|
destrin |
| chr2_-_7086066 | 0.16 |
ENSMUST00000183209.8
|
Celf2
|
CUGBP, Elav-like family member 2 |
| chr8_+_46080840 | 0.15 |
ENSMUST00000135336.9
|
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr14_+_54423447 | 0.15 |
ENSMUST00000103709.2
|
Traj32
|
T cell receptor alpha joining 32 |
| chr6_+_58808733 | 0.13 |
ENSMUST00000126292.8
ENSMUST00000031823.12 |
Herc3
|
hect domain and RLD 3 |
| chr13_-_103042294 | 0.12 |
ENSMUST00000167462.8
|
Mast4
|
microtubule associated serine/threonine kinase family member 4 |
| chr2_+_80447389 | 0.10 |
ENSMUST00000028384.5
|
Dusp19
|
dual specificity phosphatase 19 |
| chr15_-_38518458 | 0.10 |
ENSMUST00000127848.2
|
Azin1
|
antizyme inhibitor 1 |
| chr13_+_24986003 | 0.10 |
ENSMUST00000155575.2
|
BC005537
|
cDNA sequence BC005537 |
| chr11_+_97917746 | 0.10 |
ENSMUST00000017548.7
|
Rpl19
|
ribosomal protein L19 |
| chr18_-_39622295 | 0.09 |
ENSMUST00000131885.2
|
Nr3c1
|
nuclear receptor subfamily 3, group C, member 1 |
| chr9_+_64939695 | 0.09 |
ENSMUST00000034960.14
|
Dpp8
|
dipeptidylpeptidase 8 |
| chr11_-_115968373 | 0.08 |
ENSMUST00000174822.8
|
Unc13d
|
unc-13 homolog D |
| chr8_-_84771610 | 0.07 |
ENSMUST00000061923.5
|
Rln3
|
relaxin 3 |
| chr19_-_42190589 | 0.07 |
ENSMUST00000018966.8
|
Sfrp5
|
secreted frizzled-related sequence protein 5 |
| chr9_+_66257747 | 0.07 |
ENSMUST00000042824.13
|
Herc1
|
HECT and RLD domain containing E3 ubiquitin protein ligase family member 1 |
| chr2_+_72306503 | 0.06 |
ENSMUST00000102691.11
ENSMUST00000157019.2 |
Cdca7
|
cell division cycle associated 7 |
| chr10_+_52293617 | 0.06 |
ENSMUST00000023830.16
|
Nus1
|
NUS1 dehydrodolichyl diphosphate synthase subunit |
| chr11_-_115967873 | 0.05 |
ENSMUST00000153408.8
|
Unc13d
|
unc-13 homolog D |
| chr7_+_100435548 | 0.05 |
ENSMUST00000216021.2
|
Fam168a
|
family with sequence similarity 168, member A |
| chr2_+_109522781 | 0.05 |
ENSMUST00000111050.10
|
Bdnf
|
brain derived neurotrophic factor |
| chr2_+_125701054 | 0.05 |
ENSMUST00000028636.13
ENSMUST00000125084.8 |
Galk2
|
galactokinase 2 |
| chr15_-_73114855 | 0.05 |
ENSMUST00000227686.2
|
Ptk2
|
PTK2 protein tyrosine kinase 2 |
| chr5_-_73349191 | 0.03 |
ENSMUST00000176910.3
|
Fryl
|
FRY like transcription coactivator |
| chr3_-_59170245 | 0.00 |
ENSMUST00000050360.14
ENSMUST00000199609.2 |
P2ry12
|
purinergic receptor P2Y, G-protein coupled 12 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 4.9 | GO:0002528 | regulation of vascular permeability involved in acute inflammatory response(GO:0002528) |
| 0.5 | 2.4 | GO:0007493 | endodermal cell fate determination(GO:0007493) |
| 0.5 | 9.0 | GO:0061365 | positive regulation of triglyceride lipase activity(GO:0061365) |
| 0.4 | 4.3 | GO:0086023 | adrenergic receptor signaling pathway involved in heart process(GO:0086023) |
| 0.3 | 5.9 | GO:0002024 | diet induced thermogenesis(GO:0002024) |
| 0.3 | 1.1 | GO:1902896 | terminal web assembly(GO:1902896) |
| 0.2 | 1.5 | GO:0014846 | esophagus smooth muscle contraction(GO:0014846) |
| 0.2 | 4.7 | GO:0001865 | NK T cell differentiation(GO:0001865) |
| 0.2 | 1.0 | GO:0060414 | aorta smooth muscle tissue morphogenesis(GO:0060414) |
| 0.2 | 1.0 | GO:0045347 | negative regulation of MHC class II biosynthetic process(GO:0045347) |
| 0.2 | 1.3 | GO:0003219 | cardiac right ventricle formation(GO:0003219) |
| 0.2 | 3.1 | GO:0071688 | striated muscle myosin thick filament assembly(GO:0071688) |
| 0.2 | 2.6 | GO:0030277 | maintenance of gastrointestinal epithelium(GO:0030277) |
| 0.1 | 0.6 | GO:0006735 | NADH regeneration(GO:0006735) canonical glycolysis(GO:0061621) glucose catabolic process to pyruvate(GO:0061718) |
| 0.1 | 0.4 | GO:0003199 | endocardial cushion to mesenchymal transition involved in heart valve formation(GO:0003199) |
| 0.1 | 0.2 | GO:0031860 | telomeric 3' overhang formation(GO:0031860) |
| 0.1 | 0.1 | GO:0032383 | regulation of intracellular lipid transport(GO:0032377) regulation of intracellular sterol transport(GO:0032380) regulation of intracellular cholesterol transport(GO:0032383) |
| 0.0 | 0.5 | GO:1904706 | negative regulation of vascular smooth muscle cell proliferation(GO:1904706) |
| 0.0 | 0.7 | GO:0006654 | phosphatidic acid biosynthetic process(GO:0006654) |
| 0.0 | 0.6 | GO:0090336 | positive regulation of brown fat cell differentiation(GO:0090336) |
| 0.0 | 0.3 | GO:0061343 | cell adhesion involved in heart morphogenesis(GO:0061343) |
| 0.0 | 0.2 | GO:0030043 | actin filament fragmentation(GO:0030043) |
| 0.0 | 0.1 | GO:1902269 | positive regulation of polyamine transmembrane transport(GO:1902269) |
| 0.0 | 0.1 | GO:2000040 | regulation of planar cell polarity pathway involved in axis elongation(GO:2000040) negative regulation of planar cell polarity pathway involved in axis elongation(GO:2000041) |
| 0.0 | 3.6 | GO:0008203 | cholesterol metabolic process(GO:0008203) |
| 0.0 | 2.8 | GO:0055013 | cardiac muscle cell development(GO:0055013) |
| 0.0 | 0.1 | GO:0061193 | taste bud development(GO:0061193) |
| 0.0 | 0.1 | GO:0002432 | granuloma formation(GO:0002432) |
| 0.0 | 0.8 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.0 | 0.1 | GO:1905053 | regulation of base-excision repair(GO:1905051) positive regulation of base-excision repair(GO:1905053) |
| 0.0 | 1.0 | GO:0042771 | intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator(GO:0042771) |
| 0.0 | 0.0 | GO:0007172 | signal complex assembly(GO:0007172) |
| 0.0 | 0.7 | GO:0030195 | negative regulation of blood coagulation(GO:0030195) negative regulation of hemostasis(GO:1900047) |
| 0.0 | 1.4 | GO:0030500 | regulation of bone mineralization(GO:0030500) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 2.3 | GO:0005863 | striated muscle myosin thick filament(GO:0005863) |
| 0.2 | 2.0 | GO:0097208 | alveolar lamellar body(GO:0097208) |
| 0.1 | 4.3 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.1 | 1.1 | GO:1990357 | terminal web(GO:1990357) |
| 0.1 | 1.0 | GO:0098643 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 0.1 | 5.9 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.1 | 1.0 | GO:0020005 | symbiont-containing vacuole membrane(GO:0020005) |
| 0.0 | 0.2 | GO:0030870 | Mre11 complex(GO:0030870) |
| 0.0 | 0.8 | GO:0005859 | muscle myosin complex(GO:0005859) |
| 0.0 | 4.7 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.0 | 0.6 | GO:0097228 | sperm principal piece(GO:0097228) |
| 0.0 | 0.1 | GO:0033093 | Weibel-Palade body(GO:0033093) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 4.3 | GO:0042030 | ATPase inhibitor activity(GO:0042030) |
| 0.6 | 11.8 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.5 | 4.9 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.2 | 2.1 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.2 | 1.5 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.2 | 2.2 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.2 | 1.0 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 0.1 | 3.1 | GO:0097493 | structural molecule activity conferring elasticity(GO:0097493) |
| 0.1 | 3.7 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.1 | 0.3 | GO:1990932 | 5.8S rRNA binding(GO:1990932) |
| 0.1 | 0.6 | GO:0008865 | fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.1 | 3.4 | GO:0051393 | alpha-actinin binding(GO:0051393) |
| 0.1 | 1.0 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.1 | 4.7 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 1.3 | GO:0003680 | AT DNA binding(GO:0003680) |
| 0.0 | 5.9 | GO:0005254 | chloride channel activity(GO:0005254) |
| 0.0 | 2.4 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.0 | 1.5 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 0.1 | GO:0008330 | protein tyrosine/threonine phosphatase activity(GO:0008330) |
| 0.0 | 0.3 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.0 | 0.7 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.0 | 0.1 | GO:0042978 | ornithine decarboxylase activator activity(GO:0042978) |
| 0.0 | 0.2 | GO:0000014 | single-stranded DNA endodeoxyribonuclease activity(GO:0000014) G-quadruplex DNA binding(GO:0051880) |
| 0.0 | 4.1 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 0.1 | GO:0005169 | neurotrophin TRKB receptor binding(GO:0005169) |
| 0.0 | 0.0 | GO:0033858 | N-acetylgalactosamine kinase activity(GO:0033858) |
| 0.0 | 0.1 | GO:0004883 | glucocorticoid receptor activity(GO:0004883) transcription factor activity, ligand-activated RNA polymerase II transcription factor binding(GO:0038049) glucocorticoid-activated RNA polymerase II transcription factor binding transcription factor activity(GO:0038051) |
| 0.0 | 0.5 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.0 | 0.8 | GO:0036002 | pre-mRNA binding(GO:0036002) |
| 0.0 | 1.7 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 4.3 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.1 | 4.7 | ST T CELL SIGNAL TRANSDUCTION | T Cell Signal Transduction |
| 0.0 | 2.1 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.0 | 2.4 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.0 | 7.2 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 4.6 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 1.0 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 1.0 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 11.5 | REACTOME METABOLISM OF STEROID HORMONES AND VITAMINS A AND D | Genes involved in Metabolism of steroid hormones and vitamins A and D |
| 0.1 | 4.7 | REACTOME GENERATION OF SECOND MESSENGER MOLECULES | Genes involved in Generation of second messenger molecules |
| 0.1 | 2.6 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.1 | 2.2 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.1 | 1.0 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.0 | 0.4 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.0 | 1.0 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.0 | 0.7 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.0 | 0.2 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.0 | 0.6 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.0 | 0.1 | REACTOME REGULATION OF ORNITHINE DECARBOXYLASE ODC | Genes involved in Regulation of ornithine decarboxylase (ODC) |
| 0.0 | 1.7 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |