Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
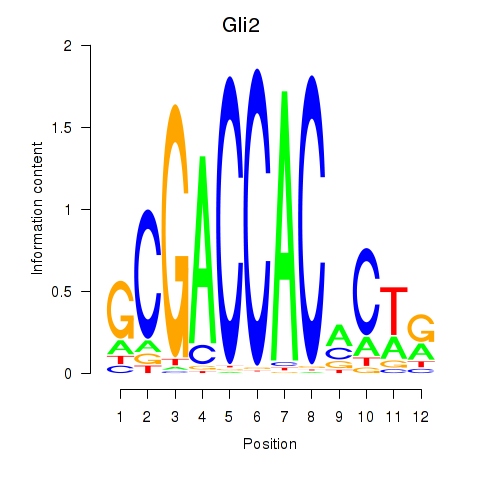
Results for Gli2
Z-value: 0.63
Transcription factors associated with Gli2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Gli2
|
ENSMUSG00000048402.15 | Gli2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Gli2 | mm39_v1_chr1_-_118981341_118981368 | 0.33 | 4.2e-03 | Click! |
Activity profile of Gli2 motif
Sorted Z-values of Gli2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Gli2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr7_+_28240262 | 6.58 |
ENSMUST00000119180.4
|
Sycn
|
syncollin |
| chr16_+_4825216 | 5.63 |
ENSMUST00000185147.8
|
Smim22
|
small integral membrane protein 22 |
| chr11_+_115802828 | 4.33 |
ENSMUST00000132961.2
|
Smim6
|
small integral membrane protein 6 |
| chr5_-_53370761 | 3.82 |
ENSMUST00000031090.8
|
Sel1l3
|
sel-1 suppressor of lin-12-like 3 (C. elegans) |
| chr7_-_144292288 | 3.35 |
ENSMUST00000238848.2
ENSMUST00000118556.9 ENSMUST00000033393.15 |
Ano1
|
anoctamin 1, calcium activated chloride channel |
| chr18_+_35347983 | 2.88 |
ENSMUST00000235449.2
ENSMUST00000235269.2 |
Ctnna1
|
catenin (cadherin associated protein), alpha 1 |
| chr11_+_96820220 | 2.71 |
ENSMUST00000062172.6
|
Prr15l
|
proline rich 15-like |
| chr11_+_96820091 | 2.65 |
ENSMUST00000054311.6
ENSMUST00000107636.4 |
Prr15l
|
proline rich 15-like |
| chr7_-_144292257 | 2.62 |
ENSMUST00000121758.8
|
Ano1
|
anoctamin 1, calcium activated chloride channel |
| chr5_+_117457126 | 2.59 |
ENSMUST00000111967.8
|
Vsig10
|
V-set and immunoglobulin domain containing 10 |
| chr6_+_113448388 | 2.44 |
ENSMUST00000058300.14
|
Il17rc
|
interleukin 17 receptor C |
| chr5_+_117457356 | 2.40 |
ENSMUST00000086464.8
|
Vsig10
|
V-set and immunoglobulin domain containing 10 |
| chr5_+_144127102 | 2.27 |
ENSMUST00000060747.8
|
Bhlha15
|
basic helix-loop-helix family, member a15 |
| chr5_+_73563418 | 2.16 |
ENSMUST00000031040.13
ENSMUST00000065543.8 |
Cwh43
|
cell wall biogenesis 43 C-terminal homolog |
| chr11_-_103235475 | 2.14 |
ENSMUST00000041385.14
|
Arhgap27
|
Rho GTPase activating protein 27 |
| chr9_+_44966464 | 2.05 |
ENSMUST00000114664.8
|
Mpzl3
|
myelin protein zero-like 3 |
| chr8_-_27664651 | 1.98 |
ENSMUST00000054212.7
ENSMUST00000033878.14 ENSMUST00000209377.2 |
Rab11fip1
|
RAB11 family interacting protein 1 (class I) |
| chr16_+_19579651 | 1.83 |
ENSMUST00000119468.2
|
B3gnt5
|
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 5 |
| chr1_-_51955054 | 1.81 |
ENSMUST00000018561.14
ENSMUST00000114537.9 |
Myo1b
|
myosin IB |
| chr11_-_70924288 | 1.78 |
ENSMUST00000238695.2
|
6330403K07Rik
|
RIKEN cDNA 6330403K07 gene |
| chr7_+_101034604 | 1.69 |
ENSMUST00000130016.8
ENSMUST00000134143.2 |
Arap1
|
ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 1 |
| chr5_-_139805661 | 1.67 |
ENSMUST00000147328.2
|
Tmem184a
|
transmembrane protein 184a |
| chr8_+_127790772 | 1.54 |
ENSMUST00000079777.12
ENSMUST00000160272.8 ENSMUST00000162907.8 ENSMUST00000162536.8 ENSMUST00000026921.13 ENSMUST00000162665.8 ENSMUST00000162602.8 ENSMUST00000160581.8 ENSMUST00000161355.8 ENSMUST00000162531.8 ENSMUST00000160766.8 ENSMUST00000159537.8 |
Pard3
|
par-3 family cell polarity regulator |
| chr6_-_72765935 | 1.51 |
ENSMUST00000114053.9
|
Tcf7l1
|
transcription factor 7 like 1 (T cell specific, HMG box) |
| chr1_-_51955126 | 1.51 |
ENSMUST00000046390.14
|
Myo1b
|
myosin IB |
| chr7_-_65020955 | 1.47 |
ENSMUST00000102592.10
|
Tjp1
|
tight junction protein 1 |
| chrX_+_99773523 | 1.43 |
ENSMUST00000019503.14
|
Gdpd2
|
glycerophosphodiester phosphodiesterase domain containing 2 |
| chrX_+_99773784 | 1.43 |
ENSMUST00000113744.2
|
Gdpd2
|
glycerophosphodiester phosphodiesterase domain containing 2 |
| chr6_-_72766224 | 1.42 |
ENSMUST00000069536.12
|
Tcf7l1
|
transcription factor 7 like 1 (T cell specific, HMG box) |
| chr6_-_137626207 | 1.38 |
ENSMUST00000134630.6
ENSMUST00000058210.13 ENSMUST00000111878.8 |
Eps8
|
epidermal growth factor receptor pathway substrate 8 |
| chr12_+_58258558 | 1.30 |
ENSMUST00000110671.3
ENSMUST00000044299.3 |
Sstr1
|
somatostatin receptor 1 |
| chr7_-_29204812 | 1.27 |
ENSMUST00000183096.8
ENSMUST00000085809.11 |
Sipa1l3
|
signal-induced proliferation-associated 1 like 3 |
| chr8_+_77628916 | 1.22 |
ENSMUST00000109912.8
ENSMUST00000128862.2 ENSMUST00000109911.8 |
Nr3c2
|
nuclear receptor subfamily 3, group C, member 2 |
| chr5_-_18565353 | 1.15 |
ENSMUST00000074694.7
|
Gnai1
|
guanine nucleotide binding protein (G protein), alpha inhibiting 1 |
| chr7_-_132724889 | 1.14 |
ENSMUST00000166439.8
|
Ctbp2
|
C-terminal binding protein 2 |
| chr7_-_132725075 | 1.06 |
ENSMUST00000163601.8
ENSMUST00000033269.15 ENSMUST00000124096.8 |
Ctbp2
Fgfr2
|
C-terminal binding protein 2 fibroblast growth factor receptor 2 |
| chr11_+_72686990 | 1.04 |
ENSMUST00000069395.7
ENSMUST00000172220.8 |
Zzef1
|
zinc finger, ZZ-type with EF hand domain 1 |
| chr15_+_95688763 | 1.03 |
ENSMUST00000227791.2
|
Ano6
|
anoctamin 6 |
| chrX_+_10351360 | 0.96 |
ENSMUST00000076354.13
ENSMUST00000115526.2 |
Tspan7
|
tetraspanin 7 |
| chr4_+_120896582 | 0.94 |
ENSMUST00000030372.6
|
Col9a2
|
collagen, type IX, alpha 2 |
| chr2_+_164327988 | 0.94 |
ENSMUST00000109350.9
|
Dbndd2
|
dysbindin (dystrobrevin binding protein 1) domain containing 2 |
| chr11_+_72687080 | 0.93 |
ENSMUST00000207107.2
|
Zzef1
|
zinc finger, ZZ-type with EF hand domain 1 |
| chrX_-_73067514 | 0.91 |
ENSMUST00000033769.15
ENSMUST00000114352.8 ENSMUST00000068286.12 ENSMUST00000114360.10 ENSMUST00000114354.10 |
Irak1
|
interleukin-1 receptor-associated kinase 1 |
| chr4_+_43401232 | 0.90 |
ENSMUST00000125399.2
|
Rusc2
|
RUN and SH3 domain containing 2 |
| chr7_-_132725137 | 0.86 |
ENSMUST00000170459.8
|
Ctbp2
|
C-terminal binding protein 2 |
| chr9_+_114230144 | 0.86 |
ENSMUST00000063042.11
ENSMUST00000111820.4 |
Glb1
Tmppe
|
galactosidase, beta 1 transmembrane protein with metallophosphoesterase domain |
| chr6_+_137731526 | 0.83 |
ENSMUST00000203216.3
ENSMUST00000087675.9 ENSMUST00000203693.3 |
Dera
|
deoxyribose-phosphate aldolase (putative) |
| chr2_+_117942357 | 0.81 |
ENSMUST00000039559.9
|
Thbs1
|
thrombospondin 1 |
| chr8_+_127790626 | 0.80 |
ENSMUST00000162309.8
|
Pard3
|
par-3 family cell polarity regulator |
| chr5_+_145051090 | 0.78 |
ENSMUST00000196111.5
ENSMUST00000141602.2 |
Arpc1b
|
actin related protein 2/3 complex, subunit 1B |
| chr2_+_164328375 | 0.76 |
ENSMUST00000069385.15
ENSMUST00000143690.8 |
Dbndd2
|
dysbindin (dystrobrevin binding protein 1) domain containing 2 |
| chr4_-_46536088 | 0.76 |
ENSMUST00000102924.3
ENSMUST00000046897.13 |
Trim14
|
tripartite motif-containing 14 |
| chrX_-_166165743 | 0.76 |
ENSMUST00000026839.5
|
Prps2
|
phosphoribosyl pyrophosphate synthetase 2 |
| chr8_+_10056654 | 0.75 |
ENSMUST00000033892.9
|
Tnfsf13b
|
tumor necrosis factor (ligand) superfamily, member 13b |
| chr11_+_70453666 | 0.74 |
ENSMUST00000072237.13
ENSMUST00000072873.14 |
Mink1
|
misshapen-like kinase 1 (zebrafish) |
| chr7_-_132724344 | 0.73 |
ENSMUST00000167218.8
|
Ctbp2
|
C-terminal binding protein 2 |
| chr1_-_172418058 | 0.71 |
ENSMUST00000065679.8
|
Slamf8
|
SLAM family member 8 |
| chr4_-_43499608 | 0.70 |
ENSMUST00000136005.3
ENSMUST00000054538.13 |
Arhgef39
|
Rho guanine nucleotide exchange factor (GEF) 39 |
| chr7_-_65020655 | 0.70 |
ENSMUST00000032729.8
|
Tjp1
|
tight junction protein 1 |
| chr5_+_145051025 | 0.69 |
ENSMUST00000085679.13
|
Arpc1b
|
actin related protein 2/3 complex, subunit 1B |
| chr3_+_97536120 | 0.66 |
ENSMUST00000107050.8
ENSMUST00000029729.15 ENSMUST00000107049.2 |
Fmo5
|
flavin containing monooxygenase 5 |
| chr17_+_45874800 | 0.66 |
ENSMUST00000224905.2
ENSMUST00000226086.2 ENSMUST00000041353.7 |
Slc35b2
|
solute carrier family 35, member B2 |
| chr6_+_137731599 | 0.65 |
ENSMUST00000204356.2
|
Dera
|
deoxyribose-phosphate aldolase (putative) |
| chr17_+_24851647 | 0.64 |
ENSMUST00000047611.4
|
Nthl1
|
nth (endonuclease III)-like 1 (E.coli) |
| chr11_+_70453724 | 0.64 |
ENSMUST00000102559.11
|
Mink1
|
misshapen-like kinase 1 (zebrafish) |
| chr3_-_135313982 | 0.62 |
ENSMUST00000132668.8
|
Nfkb1
|
nuclear factor of kappa light polypeptide gene enhancer in B cells 1, p105 |
| chr15_+_95688712 | 0.62 |
ENSMUST00000071874.8
|
Ano6
|
anoctamin 6 |
| chr7_-_97970388 | 0.61 |
ENSMUST00000120520.8
|
Acer3
|
alkaline ceramidase 3 |
| chr9_+_100956145 | 0.61 |
ENSMUST00000189616.2
|
Msl2
|
MSL complex subunit 2 |
| chr13_-_54759086 | 0.61 |
ENSMUST00000049575.8
|
Cltb
|
clathrin, light polypeptide (Lcb) |
| chrX_-_9529189 | 0.56 |
ENSMUST00000033519.3
|
Dynlt3
|
dynein light chain Tctex-type 3 |
| chr7_-_132725041 | 0.55 |
ENSMUST00000171022.8
|
Ctbp2
|
C-terminal binding protein 2 |
| chr13_+_13612136 | 0.54 |
ENSMUST00000005532.9
|
Nid1
|
nidogen 1 |
| chr8_+_10056631 | 0.53 |
ENSMUST00000207792.3
|
Tnfsf13b
|
tumor necrosis factor (ligand) superfamily, member 13b |
| chr13_-_54759145 | 0.51 |
ENSMUST00000091609.11
|
Cltb
|
clathrin, light polypeptide (Lcb) |
| chr7_+_40547608 | 0.48 |
ENSMUST00000044705.12
|
Vstm2b
|
V-set and transmembrane domain containing 2B |
| chr9_+_100525807 | 0.47 |
ENSMUST00000133388.2
|
Stag1
|
stromal antigen 1 |
| chr2_+_74566740 | 0.46 |
ENSMUST00000111982.8
|
Hoxd3
|
homeobox D3 |
| chr6_-_38614155 | 0.46 |
ENSMUST00000096030.7
ENSMUST00000201345.2 |
Klrg2
|
killer cell lectin-like receptor subfamily G, member 2 |
| chr1_-_93406091 | 0.44 |
ENSMUST00000188165.2
|
Hdlbp
|
high density lipoprotein (HDL) binding protein |
| chr17_-_24851549 | 0.43 |
ENSMUST00000227607.2
ENSMUST00000227509.2 ENSMUST00000227745.2 |
Tsc2
|
TSC complex subunit 2 |
| chr18_+_61688329 | 0.42 |
ENSMUST00000165123.8
|
Csnk1a1
|
casein kinase 1, alpha 1 |
| chr11_+_100510043 | 0.42 |
ENSMUST00000107376.8
|
Nkiras2
|
NFKB inhibitor interacting Ras-like protein 2 |
| chr2_-_37593287 | 0.40 |
ENSMUST00000072186.12
|
Strbp
|
spermatid perinuclear RNA binding protein |
| chr14_+_51162425 | 0.39 |
ENSMUST00000049411.12
ENSMUST00000136753.8 ENSMUST00000154288.3 |
Apex1
|
apurinic/apyrimidinic endonuclease 1 |
| chr1_-_72576089 | 0.38 |
ENSMUST00000047786.6
|
Marchf4
|
membrane associated ring-CH-type finger 4 |
| chr4_-_150039486 | 0.38 |
ENSMUST00000038562.9
|
Spsb1
|
splA/ryanodine receptor domain and SOCS box containing 1 |
| chr15_-_56557920 | 0.38 |
ENSMUST00000050544.8
|
Has2
|
hyaluronan synthase 2 |
| chr8_+_85786684 | 0.37 |
ENSMUST00000095220.4
|
Fbxw9
|
F-box and WD-40 domain protein 9 |
| chr17_-_56916771 | 0.37 |
ENSMUST00000052832.6
|
Micos13
|
mitochondrial contact site and cristae organizing system subunit 13 |
| chr9_+_62249730 | 0.37 |
ENSMUST00000156461.2
|
Anp32a
|
acidic (leucine-rich) nuclear phosphoprotein 32 family, member A |
| chr4_-_82939330 | 0.37 |
ENSMUST00000071708.12
|
Frem1
|
Fras1 related extracellular matrix protein 1 |
| chrX_+_160500051 | 0.35 |
ENSMUST00000112338.2
|
Rai2
|
retinoic acid induced 2 |
| chr9_+_13573756 | 0.34 |
ENSMUST00000177755.3
|
Maml2
|
mastermind like transcriptional coactivator 2 |
| chr11_+_29476385 | 0.34 |
ENSMUST00000133452.8
|
Mtif2
|
mitochondrial translational initiation factor 2 |
| chr4_+_41760454 | 0.34 |
ENSMUST00000108040.8
|
Il11ra1
|
interleukin 11 receptor, alpha chain 1 |
| chr8_+_91635192 | 0.33 |
ENSMUST00000211403.2
|
Chd9
|
chromodomain helicase DNA binding protein 9 |
| chr5_+_147206769 | 0.33 |
ENSMUST00000085591.7
|
Pdx1
|
pancreatic and duodenal homeobox 1 |
| chr2_+_130116357 | 0.33 |
ENSMUST00000136621.9
ENSMUST00000141872.2 |
Nop56
|
NOP56 ribonucleoprotein |
| chr5_-_115438971 | 0.32 |
ENSMUST00000112090.2
|
Dynll1
|
dynein light chain LC8-type 1 |
| chr12_-_32000169 | 0.32 |
ENSMUST00000176520.8
|
Hbp1
|
high mobility group box transcription factor 1 |
| chr17_-_84495364 | 0.31 |
ENSMUST00000060366.7
|
Zfp36l2
|
zinc finger protein 36, C3H type-like 2 |
| chr11_+_70453806 | 0.31 |
ENSMUST00000079244.12
ENSMUST00000102558.11 |
Mink1
|
misshapen-like kinase 1 (zebrafish) |
| chr11_+_117198958 | 0.31 |
ENSMUST00000153668.8
|
Septin9
|
septin 9 |
| chr1_-_93406515 | 0.31 |
ENSMUST00000170883.8
ENSMUST00000189025.7 |
Hdlbp
|
high density lipoprotein (HDL) binding protein |
| chr11_+_101066867 | 0.30 |
ENSMUST00000103109.4
|
Cntnap1
|
contactin associated protein-like 1 |
| chr16_+_8647959 | 0.26 |
ENSMUST00000023150.7
|
1810013L24Rik
|
RIKEN cDNA 1810013L24 gene |
| chr18_+_61688378 | 0.25 |
ENSMUST00000165721.8
ENSMUST00000115246.9 ENSMUST00000166990.8 ENSMUST00000163205.8 ENSMUST00000170862.8 |
Csnk1a1
|
casein kinase 1, alpha 1 |
| chr2_+_120439858 | 0.24 |
ENSMUST00000124187.8
|
Haus2
|
HAUS augmin-like complex, subunit 2 |
| chr2_+_85600147 | 0.24 |
ENSMUST00000065626.3
|
Olfr1013
|
olfactory receptor 1013 |
| chr17_-_25716189 | 0.23 |
ENSMUST00000165183.10
ENSMUST00000051864.5 |
Sstr5
|
somatostatin receptor 5 |
| chr9_+_107454114 | 0.23 |
ENSMUST00000112387.9
ENSMUST00000123005.8 ENSMUST00000010195.14 ENSMUST00000144392.2 |
Hyal1
|
hyaluronoglucosaminidase 1 |
| chr14_-_65187287 | 0.23 |
ENSMUST00000067843.10
ENSMUST00000176489.8 ENSMUST00000175905.8 ENSMUST00000022544.14 ENSMUST00000175744.8 ENSMUST00000176128.8 |
Hmbox1
|
homeobox containing 1 |
| chrX_+_160500623 | 0.23 |
ENSMUST00000061514.8
|
Rai2
|
retinoic acid induced 2 |
| chr11_-_121279062 | 0.22 |
ENSMUST00000106107.3
|
Rab40b
|
Rab40B, member RAS oncogene family |
| chr5_-_92231517 | 0.22 |
ENSMUST00000202258.4
ENSMUST00000113127.7 |
G3bp2
|
GTPase activating protein (SH3 domain) binding protein 2 |
| chr10_-_93425553 | 0.21 |
ENSMUST00000020203.7
|
Snrpf
|
small nuclear ribonucleoprotein polypeptide F |
| chr14_-_55354392 | 0.21 |
ENSMUST00000022819.13
|
Jph4
|
junctophilin 4 |
| chr6_+_134897473 | 0.21 |
ENSMUST00000204807.2
|
Cdkn1b
|
cyclin-dependent kinase inhibitor 1B |
| chr6_+_134897364 | 0.20 |
ENSMUST00000067327.11
ENSMUST00000003115.9 |
Cdkn1b
|
cyclin-dependent kinase inhibitor 1B |
| chr6_+_135175031 | 0.20 |
ENSMUST00000130612.2
|
Fam234b
|
family with sequence similarity 234, member B |
| chr12_-_32000209 | 0.20 |
ENSMUST00000176084.2
ENSMUST00000176103.8 ENSMUST00000167458.9 |
Hbp1
|
high mobility group box transcription factor 1 |
| chr2_+_84867783 | 0.20 |
ENSMUST00000168266.8
ENSMUST00000130729.3 |
Ssrp1
|
structure specific recognition protein 1 |
| chr9_+_100525501 | 0.19 |
ENSMUST00000146312.8
ENSMUST00000129269.8 |
Stag1
|
stromal antigen 1 |
| chr14_+_51162635 | 0.18 |
ENSMUST00000128395.2
|
Apex1
|
apurinic/apyrimidinic endonuclease 1 |
| chr17_-_54605479 | 0.17 |
ENSMUST00000233758.2
|
Slc5a7
|
solute carrier family 5 (choline transporter), member 7 |
| chr1_-_175453117 | 0.17 |
ENSMUST00000027810.14
|
Fh1
|
fumarate hydratase 1 |
| chr14_+_65187485 | 0.16 |
ENSMUST00000043914.8
ENSMUST00000239450.2 |
Ints9
|
integrator complex subunit 9 |
| chr9_+_108924457 | 0.16 |
ENSMUST00000072093.13
|
Plxnb1
|
plexin B1 |
| chr11_+_115921129 | 0.15 |
ENSMUST00000021116.12
ENSMUST00000106452.2 |
Unk
|
unkempt family zinc finger |
| chr2_+_130116344 | 0.14 |
ENSMUST00000103198.11
|
Nop56
|
NOP56 ribonucleoprotein |
| chrX_-_140508177 | 0.13 |
ENSMUST00000067841.8
|
Irs4
|
insulin receptor substrate 4 |
| chr11_+_29476423 | 0.13 |
ENSMUST00000136351.8
ENSMUST00000020749.13 ENSMUST00000144321.8 ENSMUST00000093239.11 |
Mtif2
|
mitochondrial translational initiation factor 2 |
| chr9_+_100525637 | 0.12 |
ENSMUST00000041418.13
|
Stag1
|
stromal antigen 1 |
| chr2_+_69691982 | 0.12 |
ENSMUST00000112260.3
|
Ssb
|
Sjogren syndrome antigen B |
| chrX_-_95000496 | 0.11 |
ENSMUST00000079987.13
ENSMUST00000113864.3 |
Las1l
|
LAS1-like (S. cerevisiae) |
| chr10_-_18890281 | 0.09 |
ENSMUST00000146388.2
|
Tnfaip3
|
tumor necrosis factor, alpha-induced protein 3 |
| chr11_-_100510423 | 0.09 |
ENSMUST00000137688.9
ENSMUST00000239389.2 ENSMUST00000155152.9 ENSMUST00000154972.9 ENSMUST00000239479.2 ENSMUST00000132886.3 |
Dnajc7
|
DnaJ heat shock protein family (Hsp40) member C7 |
| chrX_+_7439839 | 0.09 |
ENSMUST00000144719.9
ENSMUST00000234896.2 |
Flicr
Foxp3
|
Foxp3 regulating long intergenic noncoding RNA forkhead box P3 |
| chr17_-_24428351 | 0.08 |
ENSMUST00000024931.6
|
Ntn3
|
netrin 3 |
| chr9_-_107648144 | 0.08 |
ENSMUST00000183248.3
ENSMUST00000182022.8 ENSMUST00000035199.13 ENSMUST00000182659.8 |
Rbm5
|
RNA binding motif protein 5 |
| chr2_-_121211410 | 0.08 |
ENSMUST00000038389.15
|
Strc
|
stereocilin |
| chr1_+_55127110 | 0.08 |
ENSMUST00000075242.7
|
Hspe1
|
heat shock protein 1 (chaperonin 10) |
| chr14_-_51162083 | 0.07 |
ENSMUST00000160375.8
ENSMUST00000162177.8 |
Osgep
|
O-sialoglycoprotein endopeptidase |
| chr4_-_119151717 | 0.07 |
ENSMUST00000079644.13
|
Ybx1
|
Y box protein 1 |
| chr16_+_3648742 | 0.07 |
ENSMUST00000214238.2
ENSMUST00000214590.2 |
Olfr15
|
olfactory receptor 15 |
| chr2_-_157408239 | 0.07 |
ENSMUST00000109528.9
ENSMUST00000088494.3 |
Blcap
|
bladder cancer associated protein |
| chr9_-_108455899 | 0.06 |
ENSMUST00000068700.7
|
Wdr6
|
WD repeat domain 6 |
| chr19_+_6952319 | 0.05 |
ENSMUST00000070850.8
|
Ppp1r14b
|
protein phosphatase 1, regulatory inhibitor subunit 14B |
| chr7_-_30349548 | 0.05 |
ENSMUST00000108141.2
ENSMUST00000042726.14 |
Rbm42
|
RNA binding motif protein 42 |
| chr12_+_38830081 | 0.04 |
ENSMUST00000095767.11
|
Etv1
|
ets variant 1 |
| chr14_-_51162346 | 0.03 |
ENSMUST00000159292.8
|
Osgep
|
O-sialoglycoprotein endopeptidase |
| chr2_+_158636727 | 0.02 |
ENSMUST00000029186.14
ENSMUST00000109478.9 ENSMUST00000156893.2 |
Dhx35
|
DEAH (Asp-Glu-Ala-His) box polypeptide 35 |
| chr17_+_28910393 | 0.02 |
ENSMUST00000124886.9
ENSMUST00000114758.9 |
Mapk14
|
mitogen-activated protein kinase 14 |
| chr10_+_83379805 | 0.01 |
ENSMUST00000038388.7
|
Washc4
|
WASH complex subunit 4 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 2.9 | GO:0046021 | regulation of transcription from RNA polymerase II promoter, mitotic(GO:0046021) positive regulation of transcription from RNA polymerase II promoter during mitosis(GO:0046022) |
| 0.5 | 1.6 | GO:0097045 | activation of blood coagulation via clotting cascade(GO:0002543) phosphatidylserine exposure on blood platelet(GO:0097045) |
| 0.5 | 6.0 | GO:0015705 | iodide transport(GO:0015705) |
| 0.5 | 2.9 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.4 | 1.1 | GO:0035602 | fibroblast growth factor receptor signaling pathway involved in negative regulation of apoptotic process in bone marrow(GO:0035602) fibroblast growth factor receptor signaling pathway involved in hemopoiesis(GO:0035603) fibroblast growth factor receptor signaling pathway involved in positive regulation of cell proliferation in bone marrow(GO:0035604) coronal suture morphogenesis(GO:0060365) |
| 0.3 | 1.4 | GO:0070358 | actin polymerization-dependent cell motility(GO:0070358) |
| 0.3 | 2.3 | GO:0003383 | apical constriction(GO:0003383) |
| 0.3 | 2.0 | GO:0070164 | negative regulation of adiponectin secretion(GO:0070164) |
| 0.3 | 1.2 | GO:0031959 | mineralocorticoid receptor signaling pathway(GO:0031959) |
| 0.3 | 1.7 | GO:0018992 | germ-line sex determination(GO:0018992) |
| 0.3 | 3.3 | GO:0048386 | positive regulation of retinoic acid receptor signaling pathway(GO:0048386) |
| 0.3 | 0.8 | GO:0010752 | negative regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002581) regulation of cGMP-mediated signaling(GO:0010752) |
| 0.2 | 1.5 | GO:0046121 | deoxyribonucleoside catabolic process(GO:0046121) |
| 0.2 | 0.7 | GO:1902623 | negative regulation of neutrophil migration(GO:1902623) |
| 0.2 | 0.6 | GO:0006296 | base-excision repair, AP site formation(GO:0006285) nucleotide-excision repair, DNA incision, 5'-to lesion(GO:0006296) |
| 0.2 | 0.6 | GO:0045083 | negative regulation of interleukin-12 biosynthetic process(GO:0045083) response to diterpene(GO:1904629) cellular response to diterpene(GO:1904630) response to glucoside(GO:1904631) cellular response to glucoside(GO:1904632) |
| 0.2 | 1.3 | GO:0031296 | positive regulation of germinal center formation(GO:0002636) B cell costimulation(GO:0031296) |
| 0.2 | 2.7 | GO:0090527 | actin filament reorganization(GO:0090527) |
| 0.1 | 0.9 | GO:0019388 | galactose catabolic process(GO:0019388) |
| 0.1 | 0.8 | GO:0006167 | AMP biosynthetic process(GO:0006167) |
| 0.1 | 0.4 | GO:0046379 | extracellular polysaccharide biosynthetic process(GO:0045226) extracellular polysaccharide metabolic process(GO:0046379) |
| 0.1 | 2.4 | GO:1900017 | positive regulation of cytokine production involved in inflammatory response(GO:1900017) |
| 0.1 | 2.2 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.1 | 2.3 | GO:0006851 | mitochondrial calcium ion transport(GO:0006851) |
| 0.1 | 0.2 | GO:1900106 | hyaluranon cable assembly(GO:0036118) regulation of hyaluranon cable assembly(GO:1900104) positive regulation of hyaluranon cable assembly(GO:1900106) |
| 0.1 | 0.9 | GO:0090370 | negative regulation of cholesterol efflux(GO:0090370) |
| 0.1 | 0.4 | GO:0044861 | protein transport into plasma membrane raft(GO:0044861) |
| 0.1 | 1.7 | GO:0045060 | negative thymic T cell selection(GO:0045060) |
| 0.1 | 0.5 | GO:0021615 | glossopharyngeal nerve morphogenesis(GO:0021615) |
| 0.1 | 0.7 | GO:0051503 | adenine nucleotide transport(GO:0051503) |
| 0.1 | 1.2 | GO:1904322 | response to forskolin(GO:1904321) cellular response to forskolin(GO:1904322) |
| 0.1 | 0.3 | GO:0034287 | detection of carbohydrate stimulus(GO:0009730) detection of hexose stimulus(GO:0009732) detection of monosaccharide stimulus(GO:0034287) detection of glucose(GO:0051594) |
| 0.1 | 0.2 | GO:1900620 | acetylcholine biosynthetic process(GO:0008292) acetate ester biosynthetic process(GO:1900620) |
| 0.1 | 0.2 | GO:0006106 | fumarate metabolic process(GO:0006106) |
| 0.0 | 1.3 | GO:0071392 | cellular response to estradiol stimulus(GO:0071392) |
| 0.0 | 0.6 | GO:0043247 | telomere maintenance in response to DNA damage(GO:0043247) |
| 0.0 | 0.4 | GO:1904706 | negative regulation of vascular smooth muscle cell proliferation(GO:1904706) |
| 0.0 | 0.2 | GO:1900220 | semaphorin-plexin signaling pathway involved in bone trabecula morphogenesis(GO:1900220) |
| 0.0 | 1.7 | GO:0001921 | positive regulation of receptor recycling(GO:0001921) |
| 0.0 | 0.6 | GO:0046512 | sphingosine biosynthetic process(GO:0046512) |
| 0.0 | 2.0 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.0 | 0.7 | GO:1904424 | regulation of GTP binding(GO:1904424) |
| 0.0 | 0.5 | GO:0032836 | glomerular basement membrane development(GO:0032836) |
| 0.0 | 0.3 | GO:1904628 | response to phorbol 13-acetate 12-myristate(GO:1904627) cellular response to phorbol 13-acetate 12-myristate(GO:1904628) |
| 0.0 | 0.1 | GO:0002631 | regulation of granuloma formation(GO:0002631) negative regulation of granuloma formation(GO:0002632) regulation of toll-like receptor 5 signaling pathway(GO:0034147) negative regulation of toll-like receptor 5 signaling pathway(GO:0034148) negative regulation of nucleotide-binding oligomerization domain containing 1 signaling pathway(GO:0070429) |
| 0.0 | 0.1 | GO:0002851 | peripheral T cell tolerance induction(GO:0002458) peripheral tolerance induction(GO:0002465) positive regulation of tolerance induction dependent upon immune response(GO:0002654) regulation of peripheral tolerance induction(GO:0002658) positive regulation of peripheral tolerance induction(GO:0002660) regulation of peripheral T cell tolerance induction(GO:0002849) positive regulation of peripheral T cell tolerance induction(GO:0002851) |
| 0.0 | 1.3 | GO:0090162 | establishment of epithelial cell polarity(GO:0090162) |
| 0.0 | 0.8 | GO:0032897 | negative regulation of viral transcription(GO:0032897) |
| 0.0 | 3.3 | GO:0006892 | post-Golgi vesicle-mediated transport(GO:0006892) |
| 0.0 | 0.3 | GO:0002175 | protein localization to paranode region of axon(GO:0002175) |
| 0.0 | 0.3 | GO:0007221 | positive regulation of transcription of Notch receptor target(GO:0007221) |
| 0.0 | 0.1 | GO:0002949 | tRNA threonylcarbamoyladenosine modification(GO:0002949) |
| 0.0 | 0.3 | GO:0035721 | intraciliary retrograde transport(GO:0035721) |
| 0.0 | 0.6 | GO:0043984 | histone H4-K16 acetylation(GO:0043984) |
| 0.0 | 1.8 | GO:0030148 | sphingolipid biosynthetic process(GO:0030148) |
| 0.0 | 0.8 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.0 | 0.4 | GO:0097094 | craniofacial suture morphogenesis(GO:0097094) |
| 0.0 | 0.1 | GO:0070934 | CRD-mediated mRNA stabilization(GO:0070934) |
| 0.0 | 0.3 | GO:0046697 | decidualization(GO:0046697) |
| 0.0 | 0.2 | GO:0035563 | positive regulation of chromatin binding(GO:0035563) |
| 0.0 | 0.2 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.0 | 1.1 | GO:0072583 | clathrin-mediated endocytosis(GO:0072583) |
| 0.0 | 0.5 | GO:0000154 | rRNA modification(GO:0000154) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.5 | GO:0036284 | tubulobulbar complex(GO:0036284) |
| 0.3 | 2.3 | GO:0033269 | internode region of axon(GO:0033269) |
| 0.2 | 0.9 | GO:0005594 | collagen type IX trimer(GO:0005594) |
| 0.2 | 2.9 | GO:0005915 | zonula adherens(GO:0005915) |
| 0.2 | 0.8 | GO:0002189 | ribose phosphate diphosphokinase complex(GO:0002189) |
| 0.2 | 4.4 | GO:0097470 | ribbon synapse(GO:0097470) |
| 0.1 | 1.3 | GO:0061689 | tricellular tight junction(GO:0061689) |
| 0.1 | 0.4 | GO:0033596 | TSC1-TSC2 complex(GO:0033596) |
| 0.1 | 1.1 | GO:0030130 | clathrin coat of trans-Golgi network vesicle(GO:0030130) |
| 0.1 | 0.8 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.1 | 0.6 | GO:0072487 | MSL complex(GO:0072487) |
| 0.1 | 3.3 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.1 | 8.5 | GO:0030667 | secretory granule membrane(GO:0030667) |
| 0.1 | 2.2 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.1 | 4.8 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.1 | 1.4 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.1 | 0.5 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.0 | 0.6 | GO:0033256 | I-kappaB/NF-kappaB complex(GO:0033256) |
| 0.0 | 0.7 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.0 | 0.2 | GO:0005683 | U7 snRNP(GO:0005683) |
| 0.0 | 0.2 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.0 | 1.3 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.0 | 0.7 | GO:0005868 | cytoplasmic dynein complex(GO:0005868) |
| 0.0 | 0.5 | GO:0005605 | basal lamina(GO:0005605) |
| 0.0 | 0.2 | GO:0030314 | junctional membrane complex(GO:0030314) |
| 0.0 | 1.2 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 0.8 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.0 | 2.8 | GO:0005884 | actin filament(GO:0005884) |
| 0.0 | 0.1 | GO:0070937 | CRD-mediated mRNA stability complex(GO:0070937) histone pre-mRNA 3'end processing complex(GO:0071204) |
| 0.0 | 0.2 | GO:0005697 | telomerase holoenzyme complex(GO:0005697) |
| 0.0 | 0.1 | GO:0000408 | EKC/KEOPS complex(GO:0000408) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 6.0 | GO:0015111 | iodide transmembrane transporter activity(GO:0015111) |
| 0.6 | 1.8 | GO:0047256 | beta-galactosyl-N-acetylglucosaminylgalactosylglucosyl-ceramide beta-1,3-acetylglucosaminyltransferase activity(GO:0008457) lactosylceramide 1,3-N-acetyl-beta-D-glucosaminyltransferase activity(GO:0047256) |
| 0.4 | 4.4 | GO:0016618 | hydroxypyruvate reductase activity(GO:0016618) glyoxylate reductase (NADP) activity(GO:0030267) |
| 0.4 | 1.2 | GO:0017082 | mineralocorticoid receptor activity(GO:0017082) |
| 0.3 | 2.9 | GO:0008889 | glycerophosphodiester phosphodiesterase activity(GO:0008889) |
| 0.3 | 2.4 | GO:0030368 | interleukin-17 receptor activity(GO:0030368) |
| 0.3 | 1.5 | GO:0004994 | somatostatin receptor activity(GO:0004994) |
| 0.2 | 0.7 | GO:0046964 | 3'-phosphoadenosine 5'-phosphosulfate transmembrane transporter activity(GO:0046964) |
| 0.2 | 0.8 | GO:0070052 | collagen V binding(GO:0070052) |
| 0.2 | 0.6 | GO:0016890 | site-specific endodeoxyribonuclease activity, specific for altered base(GO:0016890) |
| 0.2 | 0.8 | GO:0004749 | ribose phosphate diphosphokinase activity(GO:0004749) |
| 0.2 | 2.2 | GO:0071253 | connexin binding(GO:0071253) |
| 0.2 | 0.6 | GO:0000702 | oxidized base lesion DNA N-glycosylase activity(GO:0000702) |
| 0.2 | 0.6 | GO:0050501 | hyaluronan synthase activity(GO:0050501) |
| 0.1 | 1.2 | GO:0031821 | G-protein coupled serotonin receptor binding(GO:0031821) |
| 0.1 | 1.7 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.1 | 7.3 | GO:0005547 | phosphatidylinositol-3,4,5-trisphosphate binding(GO:0005547) |
| 0.1 | 0.9 | GO:0016936 | galactoside binding(GO:0016936) |
| 0.1 | 2.9 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 0.1 | 1.6 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.1 | 0.3 | GO:0004921 | interleukin-11 receptor activity(GO:0004921) interleukin-11 binding(GO:0019970) |
| 0.1 | 0.6 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.1 | 1.1 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.1 | 1.5 | GO:0016832 | aldehyde-lyase activity(GO:0016832) |
| 0.1 | 0.5 | GO:0043237 | laminin-1 binding(GO:0043237) |
| 0.1 | 0.7 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.0 | 0.1 | GO:0061711 | N(6)-L-threonylcarbamoyladenine synthase(GO:0061711) |
| 0.0 | 0.9 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.0 | 0.5 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.0 | 1.3 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 0.2 | GO:0015220 | choline transmembrane transporter activity(GO:0015220) |
| 0.0 | 0.2 | GO:0003720 | telomerase activity(GO:0003720) RNA-directed DNA polymerase activity(GO:0003964) |
| 0.0 | 2.9 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.0 | 0.3 | GO:0030235 | nitric-oxide synthase regulator activity(GO:0030235) |
| 0.0 | 0.4 | GO:0004861 | cyclin-dependent protein serine/threonine kinase inhibitor activity(GO:0004861) |
| 0.0 | 1.4 | GO:0048365 | Rac GTPase binding(GO:0048365) |
| 0.0 | 2.9 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.0 | 0.1 | GO:1990715 | mRNA CDS binding(GO:1990715) |
| 0.0 | 0.2 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.0 | 0.1 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 5.1 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.1 | 7.4 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 1.1 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.0 | 1.2 | PID S1P META PATHWAY | Sphingosine 1-phosphate (S1P) pathway |
| 0.0 | 1.5 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.0 | 2.5 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.0 | 0.4 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.0 | 2.9 | PID ERBB1 DOWNSTREAM PATHWAY | ErbB1 downstream signaling |
| 0.0 | 0.8 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.0 | 0.9 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.0 | 0.6 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 0.6 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.0 | 0.4 | SIG PIP3 SIGNALING IN CARDIAC MYOCTES | Genes related to PIP3 signaling in cardiac myocytes |
| 0.0 | 1.1 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.2 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 0.1 | 2.2 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.1 | 2.9 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.1 | 2.3 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.1 | 0.9 | REACTOME NFKB IS ACTIVATED AND SIGNALS SURVIVAL | Genes involved in NF-kB is activated and signals survival |
| 0.1 | 0.9 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.1 | 1.1 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 0.1 | 1.2 | REACTOME ADENYLATE CYCLASE INHIBITORY PATHWAY | Genes involved in Adenylate cyclase inhibitory pathway |
| 0.0 | 1.1 | REACTOME FGFR2C LIGAND BINDING AND ACTIVATION | Genes involved in FGFR2c ligand binding and activation |
| 0.0 | 0.6 | REACTOME HYALURONAN METABOLISM | Genes involved in Hyaluronan metabolism |
| 0.0 | 0.4 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.0 | 1.8 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 0.5 | REACTOME CTNNB1 PHOSPHORYLATION CASCADE | Genes involved in Beta-catenin phosphorylation cascade |
| 0.0 | 0.3 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.0 | 0.2 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.0 | 0.4 | REACTOME AKT PHOSPHORYLATES TARGETS IN THE CYTOSOL | Genes involved in AKT phosphorylates targets in the cytosol |
| 0.0 | 0.9 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.0 | 0.3 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.0 | 0.8 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.0 | 0.3 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.0 | 0.6 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.7 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.0 | 1.2 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.1 | REACTOME ACETYLCHOLINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Acetylcholine Neurotransmitter Release Cycle |