Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
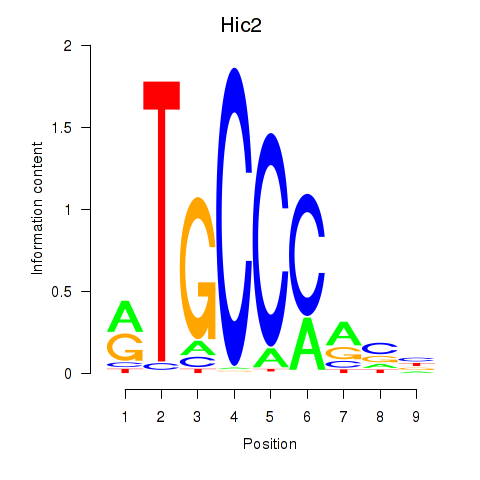
Results for Hic2
Z-value: 2.34
Transcription factors associated with Hic2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Hic2
|
ENSMUSG00000050240.17 | Hic2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Hic2 | mm39_v1_chr16_+_17051423_17051528 | 0.06 | 5.9e-01 | Click! |
Activity profile of Hic2 motif
Sorted Z-values of Hic2 motif
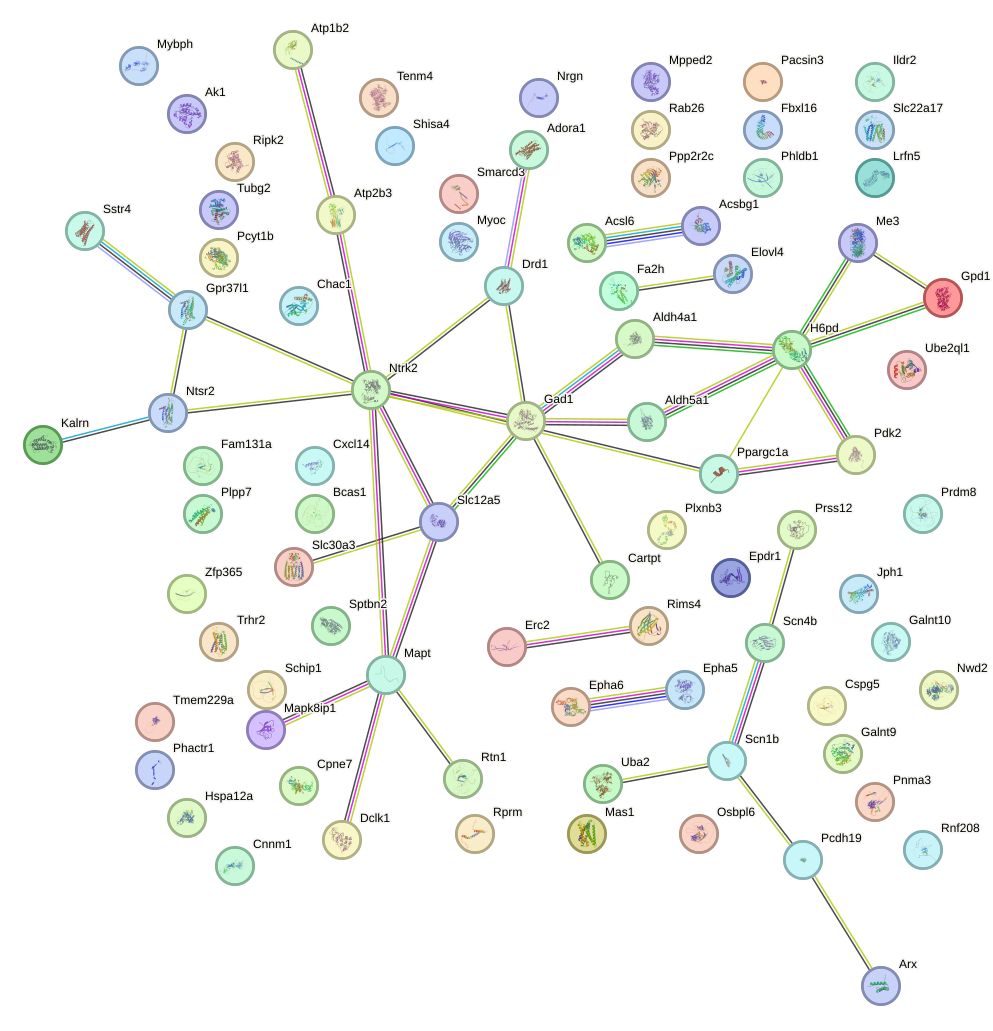
Network of associatons between targets according to the STRING database.
First level regulatory network of Hic2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_+_16703383 | 23.58 |
ENSMUST00000221596.2
ENSMUST00000111064.3 ENSMUST00000220892.2 |
Ntsr2
|
neurotensin receptor 2 |
| chr13_-_19803928 | 19.90 |
ENSMUST00000221014.2
ENSMUST00000002885.8 ENSMUST00000220944.2 |
Epdr1
|
ependymin related protein 1 (zebrafish) |
| chr13_+_58956495 | 16.88 |
ENSMUST00000225950.2
ENSMUST00000225583.2 |
Ntrk2
|
neurotrophic tyrosine kinase, receptor, type 2 |
| chr9_+_50662625 | 16.46 |
ENSMUST00000217475.2
|
Cryab
|
crystallin, alpha B |
| chr12_+_16703709 | 15.71 |
ENSMUST00000221049.2
|
Ntsr2
|
neurotensin receptor 2 |
| chr11_+_104122216 | 14.99 |
ENSMUST00000106992.10
|
Mapt
|
microtubule-associated protein tau |
| chr11_+_78215026 | 13.45 |
ENSMUST00000102478.4
|
Aldoc
|
aldolase C, fructose-bisphosphate |
| chr11_+_104122399 | 12.37 |
ENSMUST00000132977.8
ENSMUST00000132245.8 ENSMUST00000100347.11 |
Mapt
|
microtubule-associated protein tau |
| chr11_+_104122341 | 12.20 |
ENSMUST00000106993.10
|
Mapt
|
microtubule-associated protein tau |
| chr9_+_45049687 | 12.15 |
ENSMUST00000060125.7
|
Scn4b
|
sodium channel, type IV, beta |
| chr1_-_135095344 | 11.76 |
ENSMUST00000027682.9
|
Gpr37l1
|
G protein-coupled receptor 37-like 1 |
| chr2_+_119181703 | 11.68 |
ENSMUST00000028780.4
|
Chac1
|
ChaC, cation transport regulator 1 |
| chr9_-_121324705 | 11.62 |
ENSMUST00000216138.2
|
Cck
|
cholecystokinin |
| chr11_+_104122291 | 11.33 |
ENSMUST00000145227.8
|
Mapt
|
microtubule-associated protein tau |
| chr15_+_91949032 | 10.78 |
ENSMUST00000169825.8
|
Cntn1
|
contactin 1 |
| chr7_+_19144950 | 10.65 |
ENSMUST00000208710.2
ENSMUST00000003643.3 |
Ckm
|
creatine kinase, muscle |
| chr5_-_24829395 | 10.61 |
ENSMUST00000195943.2
|
Smarcd3
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3 |
| chr1_+_134121170 | 10.57 |
ENSMUST00000038445.13
ENSMUST00000191577.2 |
Mybph
|
myosin binding protein H |
| chr1_-_134162231 | 10.35 |
ENSMUST00000169927.2
|
Adora1
|
adenosine A1 receptor |
| chr12_+_61570669 | 10.33 |
ENSMUST00000055815.14
ENSMUST00000119481.2 |
Lrfn5
|
leucine rich repeat and fibronectin type III domain containing 5 |
| chr10_-_67748461 | 10.29 |
ENSMUST00000064656.8
|
Zfp365
|
zinc finger protein 365 |
| chr1_-_134163102 | 9.87 |
ENSMUST00000187631.2
ENSMUST00000038191.8 ENSMUST00000086465.6 |
Adora1
|
adenosine A1 receptor |
| chr5_+_110692162 | 9.84 |
ENSMUST00000040001.14
|
Galnt9
|
polypeptide N-acetylgalactosaminyltransferase 9 |
| chr13_+_58956077 | 9.68 |
ENSMUST00000109838.10
|
Ntrk2
|
neurotrophic tyrosine kinase, receptor, type 2 |
| chrX_+_72546680 | 9.64 |
ENSMUST00000033744.12
ENSMUST00000088429.8 ENSMUST00000114479.2 |
Atp2b3
|
ATPase, Ca++ transporting, plasma membrane 3 |
| chr18_-_23171713 | 9.41 |
ENSMUST00000081423.13
|
Nol4
|
nucleolar protein 4 |
| chr2_+_106525938 | 9.26 |
ENSMUST00000016530.14
|
Mpped2
|
metallophosphoesterase domain containing 2 |
| chr1_-_17168063 | 8.83 |
ENSMUST00000038382.5
|
Jph1
|
junctophilin 1 |
| chr2_+_70393782 | 8.79 |
ENSMUST00000123330.3
|
Gad1
|
glutamate decarboxylase 1 |
| chr13_-_54209669 | 8.76 |
ENSMUST00000021932.6
ENSMUST00000221470.2 |
Drd1
|
dopamine receptor D1 |
| chrX_-_132589727 | 8.75 |
ENSMUST00000149154.8
|
Pcdh19
|
protocadherin 19 |
| chr1_-_135302971 | 8.72 |
ENSMUST00000041240.4
|
Shisa4
|
shisa family member 4 |
| chr9_-_37464200 | 8.67 |
ENSMUST00000065668.12
|
Nrgn
|
neurogranin |
| chrX_+_72800675 | 8.60 |
ENSMUST00000002079.7
|
Plxnb3
|
plexin B3 |
| chr1_+_162466717 | 8.58 |
ENSMUST00000028020.11
|
Myoc
|
myocilin |
| chr13_-_100037149 | 8.40 |
ENSMUST00000022150.8
|
Cartpt
|
CART prepropeptide |
| chr13_-_56444118 | 8.32 |
ENSMUST00000224801.2
|
Cxcl14
|
chemokine (C-X-C motif) ligand 14 |
| chrX_+_72108393 | 8.28 |
ENSMUST00000060418.8
|
Pnma3
|
paraneoplastic antigen MA3 |
| chr16_-_60425608 | 8.14 |
ENSMUST00000068860.13
|
Epha6
|
Eph receptor A6 |
| chr19_+_10366753 | 8.13 |
ENSMUST00000169121.9
ENSMUST00000076968.11 ENSMUST00000235479.2 ENSMUST00000223586.2 ENSMUST00000235784.2 ENSMUST00000224135.3 ENSMUST00000225452.3 ENSMUST00000237366.2 |
Syt7
|
synaptotagmin VII |
| chr7_-_30826376 | 8.08 |
ENSMUST00000098548.8
|
Scn1b
|
sodium channel, voltage-gated, type I, beta |
| chr2_-_170269748 | 8.06 |
ENSMUST00000013667.3
ENSMUST00000109152.9 ENSMUST00000068137.11 |
Bcas1
|
breast carcinoma amplified sequence 1 |
| chr2_+_148237258 | 8.06 |
ENSMUST00000109962.4
|
Sstr4
|
somatostatin receptor 4 |
| chr2_+_32519479 | 8.01 |
ENSMUST00000068271.5
|
Ak1
|
adenylate kinase 1 |
| chr8_+_96404713 | 7.97 |
ENSMUST00000041318.14
|
Ndrg4
|
N-myc downstream regulated gene 4 |
| chr3_+_68491487 | 7.95 |
ENSMUST00000182997.3
|
Schip1
|
schwannomin interacting protein 1 |
| chr9_-_54569128 | 7.91 |
ENSMUST00000034822.12
|
Acsbg1
|
acyl-CoA synthetase bubblegum family member 1 |
| chr9_+_110075133 | 7.89 |
ENSMUST00000199736.2
|
Cspg5
|
chondroitin sulfate proteoglycan 5 |
| chr4_-_91260265 | 7.87 |
ENSMUST00000107110.8
ENSMUST00000008633.15 ENSMUST00000107118.8 |
Elavl2
|
ELAV like RNA binding protein 1 |
| chr6_-_24956296 | 7.63 |
ENSMUST00000127247.4
|
Tmem229a
|
transmembrane protein 229A |
| chr14_+_27598021 | 7.50 |
ENSMUST00000211684.2
ENSMUST00000210924.2 |
Erc2
|
ELKS/RAB6-interacting/CAST family member 2 |
| chr12_-_87037204 | 7.49 |
ENSMUST00000222543.2
|
Zdhhc22
|
zinc finger, DHHC-type containing 22 |
| chr8_-_112120442 | 7.44 |
ENSMUST00000038475.9
|
Fa2h
|
fatty acid 2-hydroxylase |
| chr2_-_92222979 | 7.38 |
ENSMUST00000111279.9
|
Mapk8ip1
|
mitogen-activated protein kinase 8 interacting protein 1 |
| chr9_-_121323906 | 7.37 |
ENSMUST00000215228.2
ENSMUST00000213106.2 |
Cck
|
cholecystokinin |
| chr14_-_55150547 | 7.29 |
ENSMUST00000228495.3
ENSMUST00000228119.3 ENSMUST00000050772.10 ENSMUST00000231305.2 |
Slc22a17
|
solute carrier family 22 (organic cation transporter), member 17 |
| chr11_-_101875575 | 7.21 |
ENSMUST00000176261.2
ENSMUST00000143177.2 ENSMUST00000003612.13 |
Dusp3
|
dual specificity phosphatase 3 (vaccinia virus phosphatase VH1-related) |
| chr13_+_58955675 | 7.19 |
ENSMUST00000224402.2
|
Ntrk2
|
neurotrophic tyrosine kinase, receptor, type 2 |
| chr8_-_123087485 | 7.15 |
ENSMUST00000044123.2
|
Trhr2
|
thyrotropin releasing hormone receptor 2 |
| chr17_-_24752683 | 7.11 |
ENSMUST00000061764.14
|
Rab26
|
RAB26, member RAS oncogene family |
| chr16_+_20513658 | 6.99 |
ENSMUST00000056518.13
|
Fam131a
|
family with sequence similarity 131, member A |
| chr2_+_31985528 | 6.91 |
ENSMUST00000057423.6
ENSMUST00000140762.2 |
Plpp7
|
phospholipid phosphatase 7 (inactive) |
| chr13_+_42862957 | 6.90 |
ENSMUST00000066928.12
ENSMUST00000148891.8 |
Phactr1
|
phosphatase and actin regulator 1 |
| chr5_+_98328723 | 6.85 |
ENSMUST00000112959.4
|
Prdm8
|
PR domain containing 8 |
| chr5_+_37025810 | 6.85 |
ENSMUST00000031003.11
|
Ppp2r2c
|
protein phosphatase 2, regulatory subunit B, gamma |
| chr11_+_60956624 | 6.82 |
ENSMUST00000041944.9
ENSMUST00000108717.3 |
Kcnj12
|
potassium inwardly-rectifying channel, subfamily J, member 12 |
| chrX_+_92718695 | 6.82 |
ENSMUST00000045898.4
|
Pcyt1b
|
phosphate cytidylyltransferase 1, choline, beta isoform |
| chr16_-_34083549 | 6.78 |
ENSMUST00000114949.8
ENSMUST00000114954.8 |
Kalrn
|
kalirin, RhoGEF kinase |
| chr8_+_123844090 | 6.76 |
ENSMUST00000037900.9
|
Cpne7
|
copine VII |
| chr2_+_164802729 | 6.73 |
ENSMUST00000202623.4
|
Slc12a5
|
solute carrier family 12, member 5 |
| chr2_+_164802766 | 6.72 |
ENSMUST00000202223.4
|
Slc12a5
|
solute carrier family 12, member 5 |
| chr7_+_95860863 | 6.71 |
ENSMUST00000107165.8
|
Tenm4
|
teneurin transmembrane protein 4 |
| chr3_+_55149947 | 6.61 |
ENSMUST00000167204.8
ENSMUST00000054237.14 |
Dclk1
|
doublecortin-like kinase 1 |
| chr7_-_30826184 | 6.60 |
ENSMUST00000211945.2
|
Scn1b
|
sodium channel, voltage-gated, type I, beta |
| chr4_-_149391963 | 6.57 |
ENSMUST00000055647.15
ENSMUST00000030806.6 ENSMUST00000238956.2 ENSMUST00000060537.13 |
Kif1b
|
kinesin family member 1B |
| chr13_-_25121568 | 6.57 |
ENSMUST00000037615.7
|
Aldh5a1
|
aldhehyde dehydrogenase family 5, subfamily A1 |
| chr11_+_101046708 | 6.51 |
ENSMUST00000043654.10
|
Tubg2
|
tubulin, gamma 2 |
| chr2_-_163760603 | 6.48 |
ENSMUST00000044734.3
|
Rims4
|
regulating synaptic membrane exocytosis 4 |
| chr5_+_63957805 | 6.48 |
ENSMUST00000162757.4
|
Nwd2
|
NACHT and WD repeat domain containing 2 |
| chr7_+_89281897 | 6.48 |
ENSMUST00000032856.13
|
Me3
|
malic enzyme 3, NADP(+)-dependent, mitochondrial |
| chr5_+_37025926 | 6.45 |
ENSMUST00000201156.2
|
Ppp2r2c
|
protein phosphatase 2, regulatory subunit B, gamma |
| chr11_+_54205722 | 6.43 |
ENSMUST00000072178.11
ENSMUST00000101211.9 ENSMUST00000101213.9 ENSMUST00000064690.10 ENSMUST00000108899.8 |
Acsl6
|
acyl-CoA synthetase long-chain family member 6 |
| chr3_+_123240562 | 6.42 |
ENSMUST00000029603.10
|
Prss12
|
protease, serine 12 neurotrypsin (motopsin) |
| chr9_-_83688294 | 6.37 |
ENSMUST00000034796.14
ENSMUST00000183614.2 |
Elovl4
|
elongation of very long chain fatty acids (FEN1/Elo2, SUR4/Elo3, yeast)-like 4 |
| chr5_-_51725059 | 6.33 |
ENSMUST00000127135.3
|
Ppargc1a
|
peroxisome proliferative activated receptor, gamma, coactivator 1 alpha |
| chr19_+_43428843 | 6.25 |
ENSMUST00000223787.2
ENSMUST00000165311.3 |
Cnnm1
|
cyclin M1 |
| chrX_+_92330102 | 6.23 |
ENSMUST00000046565.13
ENSMUST00000113947.6 |
Arx
|
aristaless related homeobox |
| chr17_+_26036893 | 6.22 |
ENSMUST00000235694.2
|
Fbxl16
|
F-box and leucine-rich repeat protein 16 |
| chr17_-_13070780 | 6.21 |
ENSMUST00000162389.2
ENSMUST00000162119.8 ENSMUST00000159223.8 |
Mas1
|
MAS1 oncogene |
| chr12_-_72283465 | 6.19 |
ENSMUST00000021497.16
ENSMUST00000137990.2 |
Rtn1
|
reticulon 1 |
| chr8_+_46080840 | 6.19 |
ENSMUST00000135336.9
|
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr19_-_58932026 | 6.19 |
ENSMUST00000237297.2
|
Hspa12a
|
heat shock protein 12A |
| chr15_+_99615396 | 6.16 |
ENSMUST00000023760.13
ENSMUST00000162194.2 |
Gpd1
|
glycerol-3-phosphate dehydrogenase 1 (soluble) |
| chr9_-_121324744 | 6.15 |
ENSMUST00000035120.6
|
Cck
|
cholecystokinin |
| chr2_+_76236870 | 6.04 |
ENSMUST00000077972.11
ENSMUST00000111929.8 ENSMUST00000111930.9 |
Osbpl6
|
oxysterol binding protein-like 6 |
| chr11_-_94932158 | 6.01 |
ENSMUST00000038431.8
|
Pdk2
|
pyruvate dehydrogenase kinase, isoenzyme 2 |
| chr1_+_166081755 | 5.94 |
ENSMUST00000194964.6
ENSMUST00000192638.6 ENSMUST00000192426.6 ENSMUST00000195557.6 ENSMUST00000192732.6 ENSMUST00000193860.2 |
Ildr2
|
immunoglobulin-like domain containing receptor 2 |
| chr5_-_84565218 | 5.92 |
ENSMUST00000113401.4
|
Epha5
|
Eph receptor A5 |
| chr11_-_69493567 | 5.91 |
ENSMUST00000138694.2
|
Atp1b2
|
ATPase, Na+/K+ transporting, beta 2 polypeptide |
| chr8_+_46080746 | 5.91 |
ENSMUST00000145458.9
ENSMUST00000134321.8 |
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr15_+_82152994 | 5.88 |
ENSMUST00000238416.3
ENSMUST00000239048.2 |
Septin3
|
septin 3 |
| chr2_+_91090167 | 5.88 |
ENSMUST00000138470.2
|
Pacsin3
|
protein kinase C and casein kinase substrate in neurons 3 |
| chr9_-_44624496 | 5.87 |
ENSMUST00000144251.8
ENSMUST00000156918.8 |
Phldb1
|
pleckstrin homology like domain, family B, member 1 |
| chr14_+_70793340 | 5.85 |
ENSMUST00000163060.2
|
Hr
|
lysine demethylase and nuclear receptor corepressor |
| chr2_-_53975501 | 5.84 |
ENSMUST00000100089.3
|
Rprm
|
reprimo, TP53 dependent G2 arrest mediator candidate |
| chr19_+_4771089 | 5.75 |
ENSMUST00000238976.3
|
Sptbn2
|
spectrin beta, non-erythrocytic 2 |
| chr13_-_69887964 | 5.74 |
ENSMUST00000065118.7
|
Ube2ql1
|
ubiquitin-conjugating enzyme E2Q family-like 1 |
| chr19_+_8641369 | 5.70 |
ENSMUST00000035444.10
ENSMUST00000163785.2 |
Chrm1
|
cholinergic receptor, muscarinic 1, CNS |
| chr5_-_31265562 | 5.70 |
ENSMUST00000201396.2
ENSMUST00000202740.4 |
Slc30a3
|
solute carrier family 30 (zinc transporter), member 3 |
| chr9_-_44646487 | 5.60 |
ENSMUST00000034611.15
|
Phldb1
|
pleckstrin homology like domain, family B, member 1 |
| chr2_+_25132941 | 5.60 |
ENSMUST00000114355.2
ENSMUST00000060818.2 |
Rnf208
|
ring finger protein 208 |
| chr11_-_74413564 | 5.58 |
ENSMUST00000208896.2
|
Rap1gap2
|
RAP1 GTPase activating protein 2 |
| chr5_-_71253107 | 5.58 |
ENSMUST00000197284.5
|
Gabra2
|
gamma-aminobutyric acid (GABA) A receptor, subunit alpha 2 |
| chr4_+_129030710 | 5.56 |
ENSMUST00000102600.4
|
Fndc5
|
fibronectin type III domain containing 5 |
| chr7_+_122270599 | 5.55 |
ENSMUST00000182563.2
|
Cacng3
|
calcium channel, voltage-dependent, gamma subunit 3 |
| chr8_+_31581635 | 5.50 |
ENSMUST00000161713.2
|
Dusp26
|
dual specificity phosphatase 26 (putative) |
| chr10_+_40759468 | 5.47 |
ENSMUST00000019975.14
|
Wasf1
|
WASP family, member 1 |
| chr10_+_57660948 | 5.47 |
ENSMUST00000020024.12
|
Fabp7
|
fatty acid binding protein 7, brain |
| chr16_-_34083315 | 5.41 |
ENSMUST00000114953.8
|
Kalrn
|
kalirin, RhoGEF kinase |
| chr11_-_83959380 | 5.39 |
ENSMUST00000164891.8
|
Dusp14
|
dual specificity phosphatase 14 |
| chr10_+_80192293 | 5.37 |
ENSMUST00000039836.15
ENSMUST00000105351.2 |
Plk5
|
polo like kinase 5 |
| chrX_-_58179754 | 5.37 |
ENSMUST00000033473.12
|
Fgf13
|
fibroblast growth factor 13 |
| chr12_-_72455708 | 5.36 |
ENSMUST00000078505.14
|
Rtn1
|
reticulon 1 |
| chr4_-_25800083 | 5.36 |
ENSMUST00000084770.5
|
Fut9
|
fucosyltransferase 9 |
| chr7_+_122270623 | 5.34 |
ENSMUST00000182095.2
|
Cacng3
|
calcium channel, voltage-dependent, gamma subunit 3 |
| chr4_-_15149051 | 5.34 |
ENSMUST00000041606.14
|
Necab1
|
N-terminal EF-hand calcium binding protein 1 |
| chr9_+_27702243 | 5.24 |
ENSMUST00000115243.9
|
Opcml
|
opioid binding protein/cell adhesion molecule-like |
| chr9_+_45341589 | 5.24 |
ENSMUST00000239471.2
ENSMUST00000034592.11 ENSMUST00000239429.2 |
Dscaml1
|
DS cell adhesion molecule like 1 |
| chr10_+_40759815 | 5.24 |
ENSMUST00000105509.2
|
Wasf1
|
WASP family, member 1 |
| chr11_+_98632696 | 5.23 |
ENSMUST00000103139.11
|
Thra
|
thyroid hormone receptor alpha |
| chr11_-_81859241 | 5.22 |
ENSMUST00000066197.7
|
Asic2
|
acid-sensing (proton-gated) ion channel 2 |
| chr8_+_13485451 | 5.21 |
ENSMUST00000167505.3
ENSMUST00000167071.9 |
Tmem255b
|
transmembrane protein 255B |
| chr11_-_119910986 | 5.20 |
ENSMUST00000134319.2
|
Aatk
|
apoptosis-associated tyrosine kinase |
| chr15_+_83676140 | 5.19 |
ENSMUST00000172115.8
ENSMUST00000172398.2 |
Mpped1
|
metallophosphoesterase domain containing 1 |
| chr4_-_41697040 | 5.16 |
ENSMUST00000102962.10
|
Cntfr
|
ciliary neurotrophic factor receptor |
| chr17_+_44263890 | 5.14 |
ENSMUST00000177857.9
ENSMUST00000044792.6 |
Rcan2
|
regulator of calcineurin 2 |
| chr11_+_77353218 | 5.14 |
ENSMUST00000102493.8
|
Coro6
|
coronin 6 |
| chr17_+_7437500 | 5.14 |
ENSMUST00000024575.8
|
Rps6ka2
|
ribosomal protein S6 kinase, polypeptide 2 |
| chr8_-_95405234 | 5.13 |
ENSMUST00000213043.2
|
Pllp
|
plasma membrane proteolipid |
| chr9_-_57375269 | 5.11 |
ENSMUST00000215059.2
ENSMUST00000046587.8 ENSMUST00000214256.2 |
Scamp5
|
secretory carrier membrane protein 5 |
| chr11_+_102284229 | 5.07 |
ENSMUST00000107105.9
ENSMUST00000107102.8 ENSMUST00000107103.8 ENSMUST00000006750.8 |
Rundc3a
|
RUN domain containing 3A |
| chr7_+_46045862 | 5.07 |
ENSMUST00000025202.8
|
Kcnc1
|
potassium voltage gated channel, Shaw-related subfamily, member 1 |
| chrX_-_72924436 | 5.05 |
ENSMUST00000102871.10
|
L1cam
|
L1 cell adhesion molecule |
| chr18_+_36098090 | 5.04 |
ENSMUST00000176873.8
ENSMUST00000177432.8 ENSMUST00000175734.2 |
Psd2
|
pleckstrin and Sec7 domain containing 2 |
| chr3_+_64884839 | 5.03 |
ENSMUST00000239069.2
|
Kcnab1
|
potassium voltage-gated channel, shaker-related subfamily, beta member 1 |
| chrX_-_58613428 | 5.03 |
ENSMUST00000119833.8
ENSMUST00000131319.8 |
Fgf13
|
fibroblast growth factor 13 |
| chr15_+_78312764 | 5.03 |
ENSMUST00000162517.8
ENSMUST00000166142.10 ENSMUST00000089414.11 |
Kctd17
|
potassium channel tetramerisation domain containing 17 |
| chr3_-_88679881 | 5.03 |
ENSMUST00000090945.5
|
Syt11
|
synaptotagmin XI |
| chr9_+_104446682 | 4.99 |
ENSMUST00000057742.15
|
Cpne4
|
copine IV |
| chr2_+_70393195 | 4.97 |
ENSMUST00000130998.8
|
Gad1
|
glutamate decarboxylase 1 |
| chr14_+_7030760 | 4.92 |
ENSMUST00000055211.6
|
Lrrc3b
|
leucine rich repeat containing 3B |
| chr19_-_50667079 | 4.91 |
ENSMUST00000209413.2
ENSMUST00000072685.13 ENSMUST00000164039.9 |
Sorcs1
|
sortilin-related VPS10 domain containing receptor 1 |
| chr5_+_27022355 | 4.89 |
ENSMUST00000071500.13
|
Dpp6
|
dipeptidylpeptidase 6 |
| chr7_+_29007349 | 4.88 |
ENSMUST00000108230.8
ENSMUST00000065181.12 |
Dpf1
|
D4, zinc and double PHD fingers family 1 |
| chr1_+_74371755 | 4.86 |
ENSMUST00000087225.6
|
Pnkd
|
paroxysmal nonkinesiogenic dyskinesia |
| chr11_+_78389913 | 4.85 |
ENSMUST00000017488.5
|
Vtn
|
vitronectin |
| chr1_+_75522902 | 4.85 |
ENSMUST00000124341.8
|
Slc4a3
|
solute carrier family 4 (anion exchanger), member 3 |
| chr12_+_80565764 | 4.84 |
ENSMUST00000021558.8
|
Galnt16
|
polypeptide N-acetylgalactosaminyltransferase 16 |
| chr9_-_77159152 | 4.83 |
ENSMUST00000183955.8
|
Mlip
|
muscular LMNA-interacting protein |
| chr4_+_43414696 | 4.83 |
ENSMUST00000131668.3
|
Rusc2
|
RUN and SH3 domain containing 2 |
| chr10_-_108846816 | 4.80 |
ENSMUST00000105276.8
ENSMUST00000064054.14 |
Syt1
|
synaptotagmin I |
| chr14_+_9646630 | 4.79 |
ENSMUST00000112658.8
ENSMUST00000112657.9 ENSMUST00000177814.2 ENSMUST00000067491.14 |
Cadps
|
Ca2+-dependent secretion activator |
| chr15_+_87509413 | 4.79 |
ENSMUST00000068088.8
|
Tafa5
|
TAFA chemokine like family member 5 |
| chr11_-_83959175 | 4.77 |
ENSMUST00000100705.11
|
Dusp14
|
dual specificity phosphatase 14 |
| chr5_+_35971767 | 4.76 |
ENSMUST00000114203.8
ENSMUST00000150146.2 |
Ablim2
|
actin-binding LIM protein 2 |
| chr4_-_134431534 | 4.75 |
ENSMUST00000054096.13
ENSMUST00000038628.4 |
Man1c1
|
mannosidase, alpha, class 1C, member 1 |
| chr14_-_34310602 | 4.75 |
ENSMUST00000064098.14
ENSMUST00000090040.12 ENSMUST00000022330.9 ENSMUST00000022327.13 |
Ldb3
|
LIM domain binding 3 |
| chr9_+_104446957 | 4.73 |
ENSMUST00000077190.7
|
Cpne4
|
copine IV |
| chr16_-_34083200 | 4.71 |
ENSMUST00000114947.2
|
Kalrn
|
kalirin, RhoGEF kinase |
| chr6_-_88851027 | 4.69 |
ENSMUST00000038409.12
|
Podxl2
|
podocalyxin-like 2 |
| chr6_+_42263609 | 4.69 |
ENSMUST00000238845.2
ENSMUST00000031894.13 |
Clcn1
|
chloride channel, voltage-sensitive 1 |
| chr11_+_71641505 | 4.69 |
ENSMUST00000021168.14
|
Wscd1
|
WSC domain containing 1 |
| chr14_+_70768257 | 4.69 |
ENSMUST00000047331.8
|
Lgi3
|
leucine-rich repeat LGI family, member 3 |
| chr4_+_65523223 | 4.68 |
ENSMUST00000050850.14
ENSMUST00000107366.2 |
Trim32
|
tripartite motif-containing 32 |
| chrX_+_119199956 | 4.66 |
ENSMUST00000113364.10
ENSMUST00000050239.16 ENSMUST00000113358.10 |
Pcdh11x
|
protocadherin 11 X-linked |
| chr11_+_69231274 | 4.66 |
ENSMUST00000129321.2
|
Rnf227
|
ring finger protein 227 |
| chr3_-_95811993 | 4.66 |
ENSMUST00000147962.3
ENSMUST00000036181.15 |
Car14
|
carbonic anhydrase 14 |
| chr1_-_75240551 | 4.65 |
ENSMUST00000186178.7
ENSMUST00000189769.7 ENSMUST00000027404.12 |
Ptprn
|
protein tyrosine phosphatase, receptor type, N |
| chr8_-_46664321 | 4.64 |
ENSMUST00000034049.5
|
Slc25a4
|
solute carrier family 25 (mitochondrial carrier, adenine nucleotide translocator), member 4 |
| chr3_-_89153135 | 4.63 |
ENSMUST00000041022.15
|
Trim46
|
tripartite motif-containing 46 |
| chr7_+_142052569 | 4.59 |
ENSMUST00000078497.15
ENSMUST00000105953.10 ENSMUST00000179658.8 ENSMUST00000105954.10 ENSMUST00000105952.10 ENSMUST00000105955.8 ENSMUST00000074187.13 ENSMUST00000169299.9 ENSMUST00000105957.10 ENSMUST00000180152.8 ENSMUST00000105950.11 ENSMUST00000105958.10 ENSMUST00000105949.8 |
Tnnt3
|
troponin T3, skeletal, fast |
| chr2_+_67948057 | 4.57 |
ENSMUST00000112346.3
|
B3galt1
|
UDP-Gal:betaGlcNAc beta 1,3-galactosyltransferase, polypeptide 1 |
| chr9_+_121211820 | 4.56 |
ENSMUST00000209995.2
|
Trak1
|
trafficking protein, kinesin binding 1 |
| chr9_-_100916910 | 4.54 |
ENSMUST00000142676.8
ENSMUST00000149322.8 ENSMUST00000189498.2 |
Pccb
|
propionyl Coenzyme A carboxylase, beta polypeptide |
| chr13_+_43019718 | 4.52 |
ENSMUST00000131942.2
|
Phactr1
|
phosphatase and actin regulator 1 |
| chr14_+_70768289 | 4.52 |
ENSMUST00000226548.2
|
Lgi3
|
leucine-rich repeat LGI family, member 3 |
| chr15_+_78312851 | 4.52 |
ENSMUST00000159771.8
|
Kctd17
|
potassium channel tetramerisation domain containing 17 |
| chr7_-_105218472 | 4.51 |
ENSMUST00000187683.7
ENSMUST00000210079.2 ENSMUST00000187051.7 ENSMUST00000189265.7 ENSMUST00000190369.7 |
Apbb1
|
amyloid beta (A4) precursor protein-binding, family B, member 1 |
| chr18_+_40390013 | 4.50 |
ENSMUST00000096572.2
ENSMUST00000236889.2 |
Kctd16
|
potassium channel tetramerisation domain containing 16 |
| chr8_+_84703495 | 4.48 |
ENSMUST00000211558.2
|
Prkaca
|
protein kinase, cAMP dependent, catalytic, alpha |
| chr11_-_119937896 | 4.47 |
ENSMUST00000064307.10
|
Aatk
|
apoptosis-associated tyrosine kinase |
| chr3_-_84178089 | 4.47 |
ENSMUST00000054990.11
|
Trim2
|
tripartite motif-containing 2 |
| chr2_-_85027041 | 4.46 |
ENSMUST00000099930.9
ENSMUST00000111601.2 |
Lrrc55
|
leucine rich repeat containing 55 |
| chr7_-_142453722 | 4.46 |
ENSMUST00000000219.10
|
Th
|
tyrosine hydroxylase |
| chr19_+_48194464 | 4.46 |
ENSMUST00000078880.6
|
Sorcs3
|
sortilin-related VPS10 domain containing receptor 3 |
| chr9_-_54568950 | 4.46 |
ENSMUST00000128624.2
|
Acsbg1
|
acyl-CoA synthetase bubblegum family member 1 |
| chr19_+_53665719 | 4.41 |
ENSMUST00000164202.9
|
Rbm20
|
RNA binding motif protein 20 |
| chr7_+_4925781 | 4.41 |
ENSMUST00000207527.2
ENSMUST00000207687.2 ENSMUST00000208754.2 |
Nat14
|
N-acetyltransferase 14 |
| chr12_+_87073338 | 4.39 |
ENSMUST00000110187.8
ENSMUST00000156162.8 |
Tmem63c
|
transmembrane protein 63c |
| chr15_-_43733389 | 4.38 |
ENSMUST00000067469.6
|
Tmem74
|
transmembrane protein 74 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 6.7 | 20.2 | GO:0032242 | positive regulation of nucleobase-containing compound transport(GO:0032241) regulation of nucleoside transport(GO:0032242) negative regulation of circadian sleep/wake cycle, non-REM sleep(GO:0042323) negative regulation of mucus secretion(GO:0070256) |
| 5.6 | 33.8 | GO:0099551 | synaptic signaling via neuropeptide(GO:0099538) trans-synaptic signaling by neuropeptide(GO:0099540) trans-synaptic signaling by neuropeptide, modulating synaptic transmission(GO:0099551) |
| 5.1 | 50.9 | GO:1900034 | regulation of cellular response to heat(GO:1900034) |
| 4.9 | 14.7 | GO:0021966 | corticospinal neuron axon guidance(GO:0021966) |
| 4.5 | 13.5 | GO:0040040 | thermosensory behavior(GO:0040040) |
| 3.4 | 13.6 | GO:0017055 | negative regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0017055) |
| 3.3 | 13.3 | GO:0003360 | brainstem development(GO:0003360) |
| 3.3 | 13.2 | GO:1990927 | calcium ion regulated lysosome exocytosis(GO:1990927) |
| 2.9 | 8.6 | GO:0010593 | negative regulation of lamellipodium assembly(GO:0010593) |
| 2.8 | 27.6 | GO:0046959 | habituation(GO:0046959) |
| 2.7 | 2.7 | GO:0051542 | elastin biosynthetic process(GO:0051542) |
| 2.6 | 10.3 | GO:0033566 | gamma-tubulin complex localization(GO:0033566) |
| 2.4 | 7.3 | GO:0015891 | iron chelate transport(GO:0015688) siderophore transport(GO:0015891) |
| 2.3 | 9.3 | GO:0010510 | regulation of acetyl-CoA biosynthetic process from pyruvate(GO:0010510) |
| 2.3 | 20.3 | GO:0009448 | gamma-aminobutyric acid metabolic process(GO:0009448) |
| 2.2 | 33.7 | GO:0051901 | positive regulation of mitochondrial depolarization(GO:0051901) |
| 2.1 | 6.4 | GO:0006175 | adenosine salvage(GO:0006169) dATP biosynthetic process(GO:0006175) |
| 2.1 | 6.3 | GO:1903210 | glomerular visceral epithelial cell apoptotic process(GO:1903210) regulation of glomerular visceral epithelial cell apoptotic process(GO:1904633) positive regulation of glomerular visceral epithelial cell apoptotic process(GO:1904635) positive regulation of progesterone biosynthetic process(GO:2000184) |
| 2.1 | 8.3 | GO:2000503 | positive regulation of natural killer cell chemotaxis(GO:2000503) |
| 2.1 | 6.2 | GO:0046168 | glycerol-3-phosphate catabolic process(GO:0046168) |
| 1.8 | 12.9 | GO:0045196 | establishment or maintenance of neuroblast polarity(GO:0045196) establishment of neuroblast polarity(GO:0045200) |
| 1.8 | 10.9 | GO:0098917 | retrograde trans-synaptic signaling(GO:0098917) |
| 1.8 | 3.6 | GO:0071579 | regulation of zinc ion transport(GO:0071579) |
| 1.8 | 8.8 | GO:0070093 | negative regulation of glucagon secretion(GO:0070093) |
| 1.7 | 5.2 | GO:0098885 | modification of postsynaptic actin cytoskeleton(GO:0098885) |
| 1.7 | 6.8 | GO:0031448 | regulation of fast-twitch skeletal muscle fiber contraction(GO:0031446) positive regulation of fast-twitch skeletal muscle fiber contraction(GO:0031448) |
| 1.7 | 6.7 | GO:0060912 | cardiac cell fate specification(GO:0060912) |
| 1.6 | 11.3 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 1.6 | 4.8 | GO:0061461 | lysine import(GO:0034226) L-lysine import(GO:0061461) L-lysine import into cell(GO:1903410) |
| 1.5 | 10.6 | GO:0003219 | cardiac right ventricle formation(GO:0003219) |
| 1.5 | 16.5 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 1.5 | 4.5 | GO:0042414 | epinephrine metabolic process(GO:0042414) |
| 1.5 | 4.4 | GO:2000313 | fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:0060825) regulation of fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:2000313) |
| 1.4 | 4.2 | GO:0033058 | directional locomotion(GO:0033058) |
| 1.4 | 4.2 | GO:0033693 | neurofilament bundle assembly(GO:0033693) |
| 1.4 | 4.2 | GO:0021759 | globus pallidus development(GO:0021759) |
| 1.4 | 9.6 | GO:1990034 | calcium ion export from cell(GO:1990034) |
| 1.4 | 5.4 | GO:0051866 | general adaptation syndrome(GO:0051866) |
| 1.3 | 8.0 | GO:0046103 | ADP biosynthetic process(GO:0006172) inosine biosynthetic process(GO:0046103) |
| 1.3 | 9.0 | GO:0097501 | stress response to metal ion(GO:0097501) |
| 1.3 | 6.4 | GO:0010747 | positive regulation of plasma membrane long-chain fatty acid transport(GO:0010747) |
| 1.3 | 5.1 | GO:0016332 | establishment or maintenance of polarity of embryonic epithelium(GO:0016332) |
| 1.3 | 3.8 | GO:0006434 | seryl-tRNA aminoacylation(GO:0006434) |
| 1.3 | 6.3 | GO:2000277 | positive regulation of oxidative phosphorylation uncoupler activity(GO:2000277) |
| 1.3 | 7.6 | GO:0099525 | presynaptic dense core granule exocytosis(GO:0099525) |
| 1.3 | 6.3 | GO:0098957 | anterograde axonal transport of mitochondrion(GO:0098957) |
| 1.2 | 3.6 | GO:0046166 | alditol catabolic process(GO:0019405) glyceraldehyde-3-phosphate biosynthetic process(GO:0046166) |
| 1.2 | 4.8 | GO:0006436 | tryptophanyl-tRNA aminoacylation(GO:0006436) |
| 1.2 | 5.9 | GO:0036324 | vascular endothelial growth factor receptor-2 signaling pathway(GO:0036324) |
| 1.2 | 5.9 | GO:1903288 | positive regulation of potassium ion import(GO:1903288) |
| 1.2 | 3.5 | GO:0061107 | seminal vesicle development(GO:0061107) |
| 1.2 | 11.6 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 1.2 | 4.6 | GO:2000293 | regulation of defecation(GO:2000292) negative regulation of defecation(GO:2000293) |
| 1.1 | 5.7 | GO:0007197 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) phospholipase C-activating G-protein coupled acetylcholine receptor signaling pathway(GO:0007207) |
| 1.1 | 3.4 | GO:0006500 | N-terminal protein palmitoylation(GO:0006500) |
| 1.1 | 4.5 | GO:0050923 | regulation of negative chemotaxis(GO:0050923) |
| 1.1 | 2.2 | GO:0006106 | fumarate metabolic process(GO:0006106) |
| 1.1 | 3.3 | GO:1901053 | sarcosine metabolic process(GO:1901052) sarcosine catabolic process(GO:1901053) |
| 1.1 | 3.3 | GO:1904020 | regulation of G-protein coupled receptor internalization(GO:1904020) |
| 1.1 | 5.5 | GO:1902310 | positive regulation of peptidyl-serine dephosphorylation(GO:1902310) |
| 1.1 | 3.2 | GO:0051586 | positive regulation of neurotransmitter uptake(GO:0051582) positive regulation of dopamine uptake involved in synaptic transmission(GO:0051586) positive regulation of catecholamine uptake involved in synaptic transmission(GO:0051944) |
| 1.0 | 6.3 | GO:0060160 | negative regulation of dopamine receptor signaling pathway(GO:0060160) |
| 1.0 | 3.1 | GO:0010133 | proline catabolic process to glutamate(GO:0010133) |
| 1.0 | 8.1 | GO:1904714 | regulation of chaperone-mediated autophagy(GO:1904714) |
| 1.0 | 4.0 | GO:0050883 | cerebellar molecular layer development(GO:0021679) vestibular nucleus development(GO:0021750) musculoskeletal movement, spinal reflex action(GO:0050883) |
| 1.0 | 8.1 | GO:0098532 | histone H3-K27 trimethylation(GO:0098532) |
| 1.0 | 3.0 | GO:0042560 | 10-formyltetrahydrofolate catabolic process(GO:0009258) folic acid-containing compound catabolic process(GO:0009397) pteridine-containing compound catabolic process(GO:0042560) |
| 1.0 | 2.0 | GO:0046958 | nonassociative learning(GO:0046958) |
| 1.0 | 13.4 | GO:0006751 | glutathione catabolic process(GO:0006751) |
| 0.9 | 1.8 | GO:2000981 | negative regulation of mechanoreceptor differentiation(GO:0045632) negative regulation of pro-B cell differentiation(GO:2000974) negative regulation of inner ear receptor cell differentiation(GO:2000981) |
| 0.9 | 15.5 | GO:0071688 | striated muscle myosin thick filament assembly(GO:0071688) |
| 0.9 | 2.7 | GO:2000041 | regulation of planar cell polarity pathway involved in axis elongation(GO:2000040) negative regulation of planar cell polarity pathway involved in axis elongation(GO:2000041) |
| 0.9 | 2.7 | GO:0009095 | tyrosine biosynthetic process(GO:0006571) aromatic amino acid family biosynthetic process(GO:0009073) aromatic amino acid family biosynthetic process, prephenate pathway(GO:0009095) |
| 0.9 | 3.6 | GO:1902080 | regulation of calcium ion import into sarcoplasmic reticulum(GO:1902080) negative regulation of calcium ion import into sarcoplasmic reticulum(GO:1902081) |
| 0.9 | 7.9 | GO:0006108 | malate metabolic process(GO:0006108) |
| 0.9 | 3.5 | GO:2000297 | negative regulation of synapse maturation(GO:2000297) |
| 0.9 | 4.3 | GO:0072363 | regulation of glycolytic process by positive regulation of transcription from RNA polymerase II promoter(GO:0072363) |
| 0.9 | 2.6 | GO:0060618 | nipple development(GO:0060618) |
| 0.8 | 13.3 | GO:0047497 | establishment of mitochondrion localization, microtubule-mediated(GO:0034643) mitochondrion transport along microtubule(GO:0047497) |
| 0.8 | 3.3 | GO:0035524 | proline transmembrane transport(GO:0035524) |
| 0.8 | 7.4 | GO:0032286 | central nervous system myelin maintenance(GO:0032286) |
| 0.8 | 2.4 | GO:0030327 | prenylated protein catabolic process(GO:0030327) |
| 0.8 | 4.0 | GO:0036151 | phosphatidylcholine acyl-chain remodeling(GO:0036151) |
| 0.8 | 3.1 | GO:0035582 | sequestering of BMP in extracellular matrix(GO:0035582) |
| 0.8 | 7.7 | GO:0098887 | neurotransmitter receptor transport, endosome to postsynaptic membrane(GO:0098887) |
| 0.8 | 2.3 | GO:1903348 | positive regulation of bicellular tight junction assembly(GO:1903348) |
| 0.8 | 11.5 | GO:1904259 | regulation of basement membrane assembly involved in embryonic body morphogenesis(GO:1904259) positive regulation of basement membrane assembly involved in embryonic body morphogenesis(GO:1904261) basement membrane assembly involved in embryonic body morphogenesis(GO:2001197) |
| 0.8 | 2.3 | GO:0014878 | response to electrical stimulus involved in regulation of muscle adaptation(GO:0014878) |
| 0.8 | 4.5 | GO:0050760 | thymidylate synthase biosynthetic process(GO:0050757) regulation of thymidylate synthase biosynthetic process(GO:0050758) negative regulation of thymidylate synthase biosynthetic process(GO:0050760) |
| 0.8 | 8.3 | GO:0006013 | mannose metabolic process(GO:0006013) |
| 0.7 | 9.6 | GO:0021902 | commitment of neuronal cell to specific neuron type in forebrain(GO:0021902) |
| 0.7 | 13.3 | GO:0070262 | peptidyl-serine dephosphorylation(GO:0070262) |
| 0.7 | 11.7 | GO:0042182 | ketone catabolic process(GO:0042182) |
| 0.7 | 2.9 | GO:0038026 | reelin-mediated signaling pathway(GO:0038026) |
| 0.7 | 2.9 | GO:1903237 | negative regulation of leukocyte tethering or rolling(GO:1903237) |
| 0.7 | 3.6 | GO:0001830 | trophectodermal cell fate commitment(GO:0001830) |
| 0.7 | 4.3 | GO:0060082 | response to carbon monoxide(GO:0034465) eye blink reflex(GO:0060082) |
| 0.7 | 9.2 | GO:0032482 | Rab protein signal transduction(GO:0032482) |
| 0.7 | 13.4 | GO:0021940 | positive regulation of cerebellar granule cell precursor proliferation(GO:0021940) |
| 0.7 | 12.5 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 0.7 | 2.1 | GO:0051385 | response to mineralocorticoid(GO:0051385) |
| 0.7 | 2.0 | GO:1904954 | canonical Wnt signaling pathway involved in midbrain dopaminergic neuron differentiation(GO:1904954) |
| 0.7 | 6.1 | GO:1901029 | negative regulation of mitochondrial outer membrane permeabilization involved in apoptotic signaling pathway(GO:1901029) |
| 0.7 | 2.0 | GO:0033563 | dorsal/ventral axon guidance(GO:0033563) |
| 0.7 | 2.7 | GO:0002125 | maternal aggressive behavior(GO:0002125) |
| 0.7 | 1.3 | GO:0009644 | response to high light intensity(GO:0009644) |
| 0.7 | 2.0 | GO:1903984 | positive regulation of TRAIL-activated apoptotic signaling pathway(GO:1903984) |
| 0.7 | 2.0 | GO:1900477 | negative regulation of G1/S transition of mitotic cell cycle by negative regulation of transcription from RNA polymerase II promoter(GO:1900477) |
| 0.7 | 2.0 | GO:0006216 | cytidine catabolic process(GO:0006216) cytidine deamination(GO:0009972) cytidine metabolic process(GO:0046087) |
| 0.7 | 7.8 | GO:0098870 | neuronal action potential propagation(GO:0019227) action potential propagation(GO:0098870) |
| 0.7 | 13.7 | GO:0045792 | negative regulation of cell size(GO:0045792) |
| 0.6 | 3.2 | GO:0035616 | histone H2B conserved C-terminal lysine deubiquitination(GO:0035616) |
| 0.6 | 5.7 | GO:2000969 | positive regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000969) |
| 0.6 | 1.9 | GO:0032487 | regulation of Rap protein signal transduction(GO:0032487) |
| 0.6 | 2.5 | GO:0036228 | protein targeting to nuclear inner membrane(GO:0036228) |
| 0.6 | 2.5 | GO:0021914 | negative regulation of smoothened signaling pathway involved in ventral spinal cord patterning(GO:0021914) |
| 0.6 | 2.5 | GO:0046951 | ketone body biosynthetic process(GO:0046951) |
| 0.6 | 4.3 | GO:0072092 | ureteric bud invasion(GO:0072092) metanephric renal vesicle formation(GO:0072093) |
| 0.6 | 3.6 | GO:0014826 | midgut development(GO:0007494) vein smooth muscle contraction(GO:0014826) |
| 0.6 | 2.4 | GO:2000049 | positive regulation of cell-cell adhesion mediated by cadherin(GO:2000049) |
| 0.6 | 1.7 | GO:0070681 | glutaminyl-tRNAGln biosynthesis via transamidation(GO:0070681) |
| 0.6 | 5.2 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.6 | 2.3 | GO:0021812 | neuronal-glial interaction involved in cerebral cortex radial glia guided migration(GO:0021812) |
| 0.6 | 4.0 | GO:0001766 | membrane raft polarization(GO:0001766) membrane raft distribution(GO:0031580) |
| 0.6 | 1.2 | GO:0010956 | negative regulation of calcidiol 1-monooxygenase activity(GO:0010956) |
| 0.6 | 3.4 | GO:2000124 | regulation of endocannabinoid signaling pathway(GO:2000124) |
| 0.6 | 8.0 | GO:2001135 | regulation of endocytic recycling(GO:2001135) |
| 0.6 | 3.4 | GO:0021999 | neural plate anterior/posterior regionalization(GO:0021999) |
| 0.6 | 3.4 | GO:0072318 | clathrin coat disassembly(GO:0072318) |
| 0.6 | 5.6 | GO:0007196 | adenylate cyclase-inhibiting G-protein coupled glutamate receptor signaling pathway(GO:0007196) |
| 0.6 | 6.1 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
| 0.6 | 2.8 | GO:1903054 | negative regulation of extracellular matrix organization(GO:1903054) |
| 0.6 | 2.2 | GO:0072053 | renal inner medulla development(GO:0072053) |
| 0.5 | 3.8 | GO:0045163 | clustering of voltage-gated potassium channels(GO:0045163) |
| 0.5 | 2.7 | GO:0070837 | dehydroascorbic acid transport(GO:0070837) |
| 0.5 | 1.6 | GO:0035627 | ceramide transport(GO:0035627) |
| 0.5 | 13.4 | GO:0001553 | luteinization(GO:0001553) |
| 0.5 | 5.9 | GO:0071372 | cellular response to follicle-stimulating hormone stimulus(GO:0071372) |
| 0.5 | 1.1 | GO:0090035 | regulation of chaperone-mediated protein complex assembly(GO:0090034) positive regulation of chaperone-mediated protein complex assembly(GO:0090035) |
| 0.5 | 1.6 | GO:0061075 | cerebral cortex GABAergic interneuron fate commitment(GO:0021893) positive regulation of neural retina development(GO:0061075) positive regulation of retina development in camera-type eye(GO:1902868) positive regulation of amacrine cell differentiation(GO:1902871) |
| 0.5 | 9.5 | GO:1903818 | positive regulation of voltage-gated potassium channel activity(GO:1903818) |
| 0.5 | 5.2 | GO:0015712 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.5 | 3.1 | GO:0050915 | sensory perception of sour taste(GO:0050915) |
| 0.5 | 1.6 | GO:2000170 | positive regulation of peptidyl-cysteine S-nitrosylation(GO:2000170) |
| 0.5 | 14.5 | GO:0099625 | ventricular cardiac muscle cell membrane repolarization(GO:0099625) |
| 0.5 | 1.5 | GO:0032474 | otolith morphogenesis(GO:0032474) |
| 0.5 | 4.6 | GO:0019375 | galactosylceramide biosynthetic process(GO:0006682) galactolipid biosynthetic process(GO:0019375) |
| 0.5 | 3.0 | GO:0008295 | spermidine biosynthetic process(GO:0008295) |
| 0.5 | 3.0 | GO:0014823 | response to activity(GO:0014823) |
| 0.5 | 2.5 | GO:0031944 | negative regulation of glucocorticoid metabolic process(GO:0031944) negative regulation of glucocorticoid biosynthetic process(GO:0031947) response to glycoside(GO:1903416) |
| 0.5 | 7.4 | GO:0036065 | fucosylation(GO:0036065) |
| 0.5 | 4.8 | GO:0097421 | liver regeneration(GO:0097421) |
| 0.5 | 1.9 | GO:0060785 | regulation of apoptosis involved in tissue homeostasis(GO:0060785) |
| 0.5 | 4.7 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.5 | 11.1 | GO:0031987 | locomotion involved in locomotory behavior(GO:0031987) |
| 0.5 | 3.2 | GO:0061743 | motor learning(GO:0061743) |
| 0.5 | 3.2 | GO:0019254 | carnitine metabolic process, CoA-linked(GO:0019254) |
| 0.5 | 7.7 | GO:0014850 | response to muscle activity(GO:0014850) |
| 0.5 | 1.8 | GO:2000043 | cardiogenic plate morphogenesis(GO:0003142) regulation of transcription from RNA polymerase II promoter involved in definitive endodermal cell fate specification(GO:0060807) regulation of cardiac cell fate specification(GO:2000043) |
| 0.5 | 5.4 | GO:0045161 | neuronal ion channel clustering(GO:0045161) |
| 0.4 | 8.5 | GO:0060134 | prepulse inhibition(GO:0060134) |
| 0.4 | 1.8 | GO:0008355 | olfactory learning(GO:0008355) |
| 0.4 | 3.1 | GO:0060295 | regulation of cilium movement involved in cell motility(GO:0060295) regulation of cilium beat frequency involved in ciliary motility(GO:0060296) regulation of cilium-dependent cell motility(GO:1902019) |
| 0.4 | 2.6 | GO:0035022 | positive regulation of Rac protein signal transduction(GO:0035022) |
| 0.4 | 18.6 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.4 | 5.5 | GO:1901621 | negative regulation of smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:1901621) |
| 0.4 | 3.8 | GO:0097283 | keratinocyte apoptotic process(GO:0097283) regulation of keratinocyte apoptotic process(GO:1902172) |
| 0.4 | 4.5 | GO:0002035 | brain renin-angiotensin system(GO:0002035) |
| 0.4 | 10.2 | GO:0030815 | negative regulation of cAMP metabolic process(GO:0030815) |
| 0.4 | 3.6 | GO:0045919 | positive regulation of cytolysis(GO:0045919) |
| 0.4 | 10.9 | GO:2000311 | regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000311) |
| 0.4 | 2.4 | GO:1903275 | positive regulation of sodium ion export(GO:1903275) positive regulation of sodium ion export from cell(GO:1903278) |
| 0.4 | 1.6 | GO:0001579 | medium-chain fatty acid transport(GO:0001579) |
| 0.4 | 0.8 | GO:0060594 | mammary gland specification(GO:0060594) |
| 0.4 | 2.0 | GO:0015888 | thiamine transport(GO:0015888) |
| 0.4 | 2.0 | GO:0050916 | sensory perception of sweet taste(GO:0050916) |
| 0.4 | 8.2 | GO:1900452 | regulation of long term synaptic depression(GO:1900452) |
| 0.4 | 3.1 | GO:0035902 | response to immobilization stress(GO:0035902) |
| 0.4 | 17.4 | GO:0035640 | exploration behavior(GO:0035640) |
| 0.4 | 1.9 | GO:0038128 | ERBB2 signaling pathway(GO:0038128) |
| 0.4 | 4.6 | GO:0019368 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.4 | 0.4 | GO:0001661 | conditioned taste aversion(GO:0001661) |
| 0.4 | 3.4 | GO:0019695 | choline metabolic process(GO:0019695) |
| 0.4 | 9.1 | GO:0007413 | axonal fasciculation(GO:0007413) |
| 0.4 | 5.7 | GO:0021957 | corticospinal tract morphogenesis(GO:0021957) |
| 0.4 | 9.1 | GO:0006744 | ubiquinone biosynthetic process(GO:0006744) |
| 0.4 | 8.7 | GO:1900273 | positive regulation of long-term synaptic potentiation(GO:1900273) |
| 0.4 | 4.0 | GO:1901386 | negative regulation of voltage-gated calcium channel activity(GO:1901386) |
| 0.4 | 2.2 | GO:1902260 | negative regulation of delayed rectifier potassium channel activity(GO:1902260) |
| 0.4 | 1.8 | GO:1904457 | positive regulation of neuronal action potential(GO:1904457) |
| 0.4 | 8.8 | GO:0010832 | negative regulation of myotube differentiation(GO:0010832) |
| 0.4 | 4.9 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.4 | 40.3 | GO:0007200 | phospholipase C-activating G-protein coupled receptor signaling pathway(GO:0007200) |
| 0.3 | 1.0 | GO:0060830 | ciliary receptor clustering involved in smoothened signaling pathway(GO:0060830) |
| 0.3 | 1.7 | GO:0051835 | positive regulation of synapse structural plasticity(GO:0051835) |
| 0.3 | 6.1 | GO:0007183 | SMAD protein complex assembly(GO:0007183) |
| 0.3 | 2.7 | GO:0015868 | purine ribonucleotide transport(GO:0015868) |
| 0.3 | 4.0 | GO:0099601 | regulation of neurotransmitter receptor activity(GO:0099601) |
| 0.3 | 0.7 | GO:1901671 | positive regulation of superoxide dismutase activity(GO:1901671) positive regulation of removal of superoxide radicals(GO:1904833) |
| 0.3 | 1.0 | GO:0098707 | ferrous iron import into cell(GO:0097460) ferrous iron import across plasma membrane(GO:0098707) |
| 0.3 | 12.4 | GO:0000038 | very long-chain fatty acid metabolic process(GO:0000038) |
| 0.3 | 2.9 | GO:0042905 | 9-cis-retinoic acid biosynthetic process(GO:0042904) 9-cis-retinoic acid metabolic process(GO:0042905) |
| 0.3 | 1.3 | GO:0000160 | phosphorelay signal transduction system(GO:0000160) |
| 0.3 | 2.8 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.3 | 2.2 | GO:0031547 | brain-derived neurotrophic factor receptor signaling pathway(GO:0031547) |
| 0.3 | 2.1 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
| 0.3 | 2.7 | GO:0048149 | behavioral response to ethanol(GO:0048149) |
| 0.3 | 2.7 | GO:0060718 | chorionic trophoblast cell differentiation(GO:0060718) |
| 0.3 | 1.2 | GO:0072021 | ascending thin limb development(GO:0072021) thick ascending limb development(GO:0072023) metanephric ascending thin limb development(GO:0072218) metanephric thick ascending limb development(GO:0072233) |
| 0.3 | 2.7 | GO:0048630 | skeletal muscle tissue growth(GO:0048630) |
| 0.3 | 6.6 | GO:0021952 | central nervous system projection neuron axonogenesis(GO:0021952) |
| 0.3 | 4.2 | GO:0060019 | radial glial cell differentiation(GO:0060019) |
| 0.3 | 2.7 | GO:0006102 | isocitrate metabolic process(GO:0006102) |
| 0.3 | 9.9 | GO:0097320 | membrane tubulation(GO:0097320) |
| 0.3 | 0.9 | GO:0030472 | mitotic spindle organization in nucleus(GO:0030472) |
| 0.3 | 2.9 | GO:0042135 | neurotransmitter catabolic process(GO:0042135) |
| 0.3 | 2.0 | GO:0042421 | norepinephrine biosynthetic process(GO:0042421) |
| 0.3 | 3.9 | GO:2000463 | positive regulation of excitatory postsynaptic potential(GO:2000463) |
| 0.3 | 0.8 | GO:0000066 | mitochondrial ornithine transport(GO:0000066) |
| 0.3 | 1.4 | GO:0016081 | synaptic vesicle docking(GO:0016081) positive regulation of trophoblast cell migration(GO:1901165) |
| 0.3 | 2.8 | GO:0034983 | peptidyl-lysine deacetylation(GO:0034983) |
| 0.3 | 1.1 | GO:2000588 | positive regulation of platelet-derived growth factor receptor-beta signaling pathway(GO:2000588) |
| 0.3 | 2.7 | GO:0006105 | succinate metabolic process(GO:0006105) |
| 0.3 | 3.5 | GO:1901898 | negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.3 | 0.5 | GO:0051823 | regulation of synapse structural plasticity(GO:0051823) |
| 0.3 | 0.8 | GO:0090403 | oxidative stress-induced premature senescence(GO:0090403) |
| 0.3 | 1.9 | GO:0021873 | forebrain neuroblast division(GO:0021873) |
| 0.3 | 4.5 | GO:1900119 | positive regulation of execution phase of apoptosis(GO:1900119) |
| 0.3 | 1.1 | GO:1902268 | negative regulation of polyamine transmembrane transport(GO:1902268) |
| 0.3 | 3.4 | GO:0016082 | synaptic vesicle priming(GO:0016082) |
| 0.3 | 2.6 | GO:0032261 | purine nucleotide salvage(GO:0032261) IMP salvage(GO:0032264) |
| 0.3 | 1.3 | GO:0072139 | glomerular parietal epithelial cell differentiation(GO:0072139) positive regulation of nephron tubule epithelial cell differentiation(GO:2000768) |
| 0.3 | 0.8 | GO:0061537 | glycine secretion(GO:0061536) glycine secretion, neurotransmission(GO:0061537) |
| 0.3 | 2.3 | GO:0048672 | positive regulation of collateral sprouting(GO:0048672) |
| 0.3 | 1.5 | GO:0007217 | tachykinin receptor signaling pathway(GO:0007217) |
| 0.3 | 3.1 | GO:1901534 | positive regulation of hematopoietic progenitor cell differentiation(GO:1901534) |
| 0.3 | 1.8 | GO:0060316 | positive regulation of ryanodine-sensitive calcium-release channel activity(GO:0060316) |
| 0.3 | 1.8 | GO:0001880 | Mullerian duct regression(GO:0001880) |
| 0.2 | 2.2 | GO:0003147 | neural crest cell migration involved in heart formation(GO:0003147) anterior neural tube closure(GO:0061713) cellular response to folic acid(GO:0071231) |
| 0.2 | 1.5 | GO:0048069 | eye pigmentation(GO:0048069) |
| 0.2 | 3.1 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.2 | 4.1 | GO:0030828 | positive regulation of cGMP biosynthetic process(GO:0030828) |
| 0.2 | 3.6 | GO:0071625 | vocalization behavior(GO:0071625) |
| 0.2 | 1.2 | GO:0070966 | nuclear-transcribed mRNA catabolic process, no-go decay(GO:0070966) |
| 0.2 | 1.2 | GO:0009946 | proximal/distal axis specification(GO:0009946) |
| 0.2 | 5.4 | GO:0021859 | pyramidal neuron differentiation(GO:0021859) |
| 0.2 | 3.2 | GO:0006044 | N-acetylglucosamine metabolic process(GO:0006044) |
| 0.2 | 4.8 | GO:0010614 | negative regulation of cardiac muscle hypertrophy(GO:0010614) |
| 0.2 | 2.3 | GO:1902412 | regulation of mitotic cytokinesis(GO:1902412) |
| 0.2 | 5.0 | GO:0045956 | positive regulation of calcium ion-dependent exocytosis(GO:0045956) |
| 0.2 | 2.0 | GO:0060732 | positive regulation of inositol phosphate biosynthetic process(GO:0060732) |
| 0.2 | 12.8 | GO:0010765 | positive regulation of sodium ion transport(GO:0010765) |
| 0.2 | 4.6 | GO:0099612 | protein localization to axon(GO:0099612) |
| 0.2 | 3.5 | GO:0045199 | maintenance of epithelial cell apical/basal polarity(GO:0045199) |
| 0.2 | 4.1 | GO:0006054 | N-acetylneuraminate metabolic process(GO:0006054) |
| 0.2 | 1.5 | GO:0071638 | negative regulation of monocyte chemotactic protein-1 production(GO:0071638) |
| 0.2 | 5.8 | GO:0016079 | synaptic vesicle exocytosis(GO:0016079) |
| 0.2 | 0.6 | GO:0090063 | positive regulation of microtubule nucleation(GO:0090063) |
| 0.2 | 1.5 | GO:0016255 | attachment of GPI anchor to protein(GO:0016255) |
| 0.2 | 1.2 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 0.2 | 7.6 | GO:0045724 | positive regulation of cilium assembly(GO:0045724) |
| 0.2 | 2.7 | GO:0071318 | cellular response to ATP(GO:0071318) |
| 0.2 | 1.2 | GO:1901509 | regulation of endothelial tube morphogenesis(GO:1901509) |
| 0.2 | 1.0 | GO:0017182 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 0.2 | 2.6 | GO:2000465 | regulation of glycogen (starch) synthase activity(GO:2000465) |
| 0.2 | 2.6 | GO:0010804 | negative regulation of tumor necrosis factor-mediated signaling pathway(GO:0010804) |
| 0.2 | 2.0 | GO:0090394 | negative regulation of excitatory postsynaptic potential(GO:0090394) |
| 0.2 | 9.4 | GO:0045214 | sarcomere organization(GO:0045214) |
| 0.2 | 9.0 | GO:0035335 | peptidyl-tyrosine dephosphorylation(GO:0035335) |
| 0.2 | 4.1 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 0.2 | 5.4 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.2 | 3.7 | GO:0007288 | sperm axoneme assembly(GO:0007288) |
| 0.2 | 6.7 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.2 | 1.5 | GO:0097428 | protein maturation by iron-sulfur cluster transfer(GO:0097428) |
| 0.2 | 2.7 | GO:0070389 | chaperone cofactor-dependent protein refolding(GO:0070389) |
| 0.2 | 2.8 | GO:0015747 | urate transport(GO:0015747) |
| 0.2 | 2.8 | GO:0009128 | purine nucleoside monophosphate catabolic process(GO:0009128) |
| 0.2 | 7.4 | GO:0043001 | Golgi to plasma membrane protein transport(GO:0043001) |
| 0.2 | 0.6 | GO:1904690 | regulation of cap-independent translational initiation(GO:1903677) positive regulation of cap-independent translational initiation(GO:1903679) regulation of cytoplasmic translational initiation(GO:1904688) positive regulation of cytoplasmic translational initiation(GO:1904690) |
| 0.2 | 4.9 | GO:1901381 | positive regulation of potassium ion transmembrane transport(GO:1901381) |
| 0.2 | 1.6 | GO:0001778 | plasma membrane repair(GO:0001778) |
| 0.2 | 6.5 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.2 | 0.5 | GO:0033540 | fatty acid beta-oxidation using acyl-CoA oxidase(GO:0033540) |
| 0.2 | 3.0 | GO:0045725 | positive regulation of glycogen biosynthetic process(GO:0045725) |
| 0.2 | 1.6 | GO:0034372 | very-low-density lipoprotein particle remodeling(GO:0034372) |
| 0.2 | 6.7 | GO:0010107 | potassium ion import(GO:0010107) |
| 0.2 | 0.7 | GO:1900739 | regulation of protein insertion into mitochondrial membrane involved in apoptotic signaling pathway(GO:1900739) positive regulation of protein insertion into mitochondrial membrane involved in apoptotic signaling pathway(GO:1900740) |
| 0.2 | 1.7 | GO:0002934 | desmosome organization(GO:0002934) |
| 0.2 | 0.7 | GO:0044805 | late nucleophagy(GO:0044805) |
| 0.2 | 7.1 | GO:0003298 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.2 | 2.0 | GO:1903861 | positive regulation of dendrite extension(GO:1903861) |
| 0.2 | 2.2 | GO:0021955 | central nervous system neuron axonogenesis(GO:0021955) |
| 0.2 | 1.2 | GO:0009405 | pathogenesis(GO:0009405) |
| 0.2 | 3.5 | GO:0046069 | cGMP catabolic process(GO:0046069) |
| 0.2 | 1.8 | GO:0034058 | endosomal vesicle fusion(GO:0034058) |
| 0.2 | 5.5 | GO:0002347 | response to tumor cell(GO:0002347) |
| 0.2 | 1.3 | GO:0097039 | protein linear polyubiquitination(GO:0097039) |
| 0.2 | 6.5 | GO:0008333 | endosome to lysosome transport(GO:0008333) |
| 0.2 | 6.0 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.2 | 3.7 | GO:0035025 | positive regulation of Rho protein signal transduction(GO:0035025) |
| 0.2 | 17.9 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.1 | 1.5 | GO:0007216 | G-protein coupled glutamate receptor signaling pathway(GO:0007216) |
| 0.1 | 3.2 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.1 | 1.2 | GO:0008611 | ether lipid biosynthetic process(GO:0008611) glycerol ether biosynthetic process(GO:0046504) ether biosynthetic process(GO:1901503) |
| 0.1 | 0.9 | GO:0007468 | regulation of rhodopsin gene expression(GO:0007468) |
| 0.1 | 1.5 | GO:1900121 | negative regulation of receptor binding(GO:1900121) |
| 0.1 | 1.3 | GO:0043931 | ossification involved in bone maturation(GO:0043931) |
| 0.1 | 0.9 | GO:0090315 | negative regulation of protein targeting to membrane(GO:0090315) |
| 0.1 | 3.0 | GO:0001573 | ganglioside metabolic process(GO:0001573) |
| 0.1 | 1.4 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.1 | 1.1 | GO:0036265 | RNA (guanine-N7)-methylation(GO:0036265) |
| 0.1 | 1.0 | GO:0034214 | protein hexamerization(GO:0034214) |
| 0.1 | 0.4 | GO:0018900 | dichloromethane metabolic process(GO:0018900) |
| 0.1 | 3.6 | GO:0007020 | microtubule nucleation(GO:0007020) |
| 0.1 | 2.5 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.1 | 1.9 | GO:0006388 | tRNA splicing, via endonucleolytic cleavage and ligation(GO:0006388) |
| 0.1 | 0.4 | GO:1904180 | negative regulation of mitochondrial depolarization(GO:0051902) cellular response to progesterone stimulus(GO:0071393) negative regulation of membrane depolarization(GO:1904180) |
| 0.1 | 0.8 | GO:0030917 | midbrain-hindbrain boundary morphogenesis(GO:0021555) midbrain-hindbrain boundary development(GO:0030917) |
| 0.1 | 0.7 | GO:1903378 | positive regulation of oxidative stress-induced neuron intrinsic apoptotic signaling pathway(GO:1903378) |
| 0.1 | 3.7 | GO:0046325 | negative regulation of glucose import(GO:0046325) |
| 0.1 | 4.6 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.1 | 9.4 | GO:0060395 | SMAD protein signal transduction(GO:0060395) |
| 0.1 | 0.8 | GO:0045188 | regulation of circadian sleep/wake cycle, non-REM sleep(GO:0045188) |
| 0.1 | 1.2 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.1 | 2.3 | GO:2001223 | negative regulation of neuron migration(GO:2001223) |
| 0.1 | 0.4 | GO:0008292 | acetylcholine biosynthetic process(GO:0008292) acetate ester biosynthetic process(GO:1900620) |
| 0.1 | 0.4 | GO:1903336 | negative regulation of vacuolar transport(GO:1903336) negative regulation of early endosome to late endosome transport(GO:2000642) |
| 0.1 | 3.3 | GO:0015813 | L-glutamate transport(GO:0015813) |
| 0.1 | 0.9 | GO:0035469 | determination of pancreatic left/right asymmetry(GO:0035469) |
| 0.1 | 0.7 | GO:0006537 | glutamate biosynthetic process(GO:0006537) glutamine catabolic process(GO:0006543) |
| 0.1 | 1.2 | GO:2000574 | regulation of microtubule motor activity(GO:2000574) positive regulation of microtubule motor activity(GO:2000576) |
| 0.1 | 0.4 | GO:0019413 | acetate biosynthetic process(GO:0019413) acetyl-CoA biosynthetic process from acetate(GO:0019427) propionate metabolic process(GO:0019541) propionate biosynthetic process(GO:0019542) |
| 0.1 | 1.8 | GO:0000478 | endonucleolytic cleavage involved in rRNA processing(GO:0000478) |
| 0.1 | 0.9 | GO:0038042 | dimeric G-protein coupled receptor signaling pathway(GO:0038042) calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.1 | 4.0 | GO:0045746 | negative regulation of Notch signaling pathway(GO:0045746) |
| 0.1 | 0.6 | GO:0090370 | negative regulation of cholesterol efflux(GO:0090370) |
| 0.1 | 1.7 | GO:0034497 | protein localization to pre-autophagosomal structure(GO:0034497) |
| 0.1 | 1.0 | GO:0001842 | neural fold formation(GO:0001842) |
| 0.1 | 1.6 | GO:0043567 | regulation of insulin-like growth factor receptor signaling pathway(GO:0043567) |
| 0.1 | 2.9 | GO:0032094 | response to food(GO:0032094) |
| 0.1 | 1.5 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.1 | 0.8 | GO:0006983 | ER overload response(GO:0006983) |
| 0.1 | 1.0 | GO:0006122 | mitochondrial electron transport, ubiquinol to cytochrome c(GO:0006122) |
| 0.1 | 0.3 | GO:1902220 | suppression by virus of host apoptotic process(GO:0019050) modulation by virus of host apoptotic process(GO:0039526) positive regulation of intrinsic apoptotic signaling pathway in response to osmotic stress(GO:1902220) |
| 0.1 | 0.3 | GO:0006562 | proline catabolic process(GO:0006562) |
| 0.1 | 0.8 | GO:0046628 | positive regulation of insulin receptor signaling pathway(GO:0046628) |
| 0.1 | 0.3 | GO:0009080 | alanine catabolic process(GO:0006524) pyruvate family amino acid catabolic process(GO:0009080) glycine biosynthetic process, by transamination of glyoxylate(GO:0019265) L-alanine catabolic process(GO:0042853) oxalic acid secretion(GO:0046724) |
| 0.1 | 1.9 | GO:1901385 | regulation of voltage-gated calcium channel activity(GO:1901385) |
| 0.1 | 0.2 | GO:0007191 | adenylate cyclase-activating dopamine receptor signaling pathway(GO:0007191) |
| 0.1 | 2.0 | GO:0043501 | skeletal muscle adaptation(GO:0043501) |
| 0.1 | 0.7 | GO:2001106 | regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001106) |
| 0.1 | 2.4 | GO:0016180 | snRNA processing(GO:0016180) |
| 0.1 | 1.0 | GO:0006662 | glycerol ether metabolic process(GO:0006662) |
| 0.1 | 0.6 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 0.1 | 1.5 | GO:0070050 | neuron cellular homeostasis(GO:0070050) |
| 0.1 | 0.9 | GO:0000301 | retrograde transport, vesicle recycling within Golgi(GO:0000301) |
| 0.1 | 0.4 | GO:0045955 | negative regulation of calcium ion-dependent exocytosis(GO:0045955) |
| 0.1 | 0.2 | GO:0072708 | response to sorbitol(GO:0072708) |
| 0.1 | 1.0 | GO:0045187 | regulation of circadian sleep/wake cycle, sleep(GO:0045187) |
| 0.1 | 0.6 | GO:0061462 | protein localization to lysosome(GO:0061462) |
| 0.1 | 3.2 | GO:0090662 | energy coupled proton transmembrane transport, against electrochemical gradient(GO:0015988) ATP hydrolysis coupled proton transport(GO:0015991) ATP hydrolysis coupled transmembrane transport(GO:0090662) |
| 0.1 | 0.5 | GO:0035063 | nuclear speck organization(GO:0035063) |
| 0.1 | 3.0 | GO:0046839 | phospholipid dephosphorylation(GO:0046839) |
| 0.1 | 0.6 | GO:2000601 | positive regulation of Arp2/3 complex-mediated actin nucleation(GO:2000601) |
| 0.1 | 2.0 | GO:0046579 | positive regulation of Ras protein signal transduction(GO:0046579) |
| 0.1 | 1.2 | GO:0042754 | negative regulation of circadian rhythm(GO:0042754) |
| 0.1 | 1.9 | GO:0015985 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.1 | 6.3 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.1 | 0.5 | GO:0043589 | skin morphogenesis(GO:0043589) |
| 0.1 | 2.6 | GO:0019835 | cytolysis(GO:0019835) |
| 0.1 | 2.4 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.1 | 4.3 | GO:0042398 | cellular modified amino acid biosynthetic process(GO:0042398) |
| 0.1 | 7.2 | GO:0035308 | negative regulation of protein dephosphorylation(GO:0035308) |
| 0.1 | 3.0 | GO:0007416 | synapse assembly(GO:0007416) |
| 0.1 | 3.6 | GO:0010257 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.1 | 3.5 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.1 | 0.2 | GO:0072205 | metanephric collecting duct development(GO:0072205) |
| 0.1 | 1.4 | GO:0003254 | regulation of membrane depolarization(GO:0003254) |
| 0.1 | 0.9 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.1 | 4.2 | GO:0048821 | erythrocyte development(GO:0048821) |
| 0.1 | 1.1 | GO:1904869 | protein localization to nuclear body(GO:1903405) protein localization to Cajal body(GO:1904867) regulation of protein localization to Cajal body(GO:1904869) positive regulation of protein localization to Cajal body(GO:1904871) protein localization to nucleoplasm(GO:1990173) |
| 0.1 | 1.7 | GO:0034389 | lipid particle organization(GO:0034389) |
| 0.1 | 0.9 | GO:0046473 | phosphatidic acid metabolic process(GO:0046473) |
| 0.1 | 0.7 | GO:0061158 | 3'-UTR-mediated mRNA destabilization(GO:0061158) |
| 0.1 | 5.7 | GO:0051865 | protein autoubiquitination(GO:0051865) |
| 0.1 | 0.1 | GO:0007509 | mesoderm migration involved in gastrulation(GO:0007509) |
| 0.1 | 3.7 | GO:0001541 | ovarian follicle development(GO:0001541) |
| 0.1 | 1.0 | GO:0006828 | manganese ion transport(GO:0006828) |
| 0.1 | 0.7 | GO:0090042 | tubulin deacetylation(GO:0090042) regulation of tubulin deacetylation(GO:0090043) |
| 0.1 | 3.4 | GO:0042733 | embryonic digit morphogenesis(GO:0042733) |
| 0.1 | 1.4 | GO:0007193 | adenylate cyclase-inhibiting G-protein coupled receptor signaling pathway(GO:0007193) |
| 0.1 | 2.2 | GO:0000266 | mitochondrial fission(GO:0000266) |
| 0.1 | 1.3 | GO:0030502 | negative regulation of bone mineralization(GO:0030502) |
| 0.1 | 1.5 | GO:0071501 | response to sterol depletion(GO:0006991) SREBP signaling pathway(GO:0032933) cellular response to sterol depletion(GO:0071501) |
| 0.1 | 0.1 | GO:0060849 | regulation of transcription involved in lymphatic endothelial cell fate commitment(GO:0060849) |
| 0.1 | 0.2 | GO:0009750 | response to fructose(GO:0009750) |
| 0.1 | 0.2 | GO:0006427 | histidyl-tRNA aminoacylation(GO:0006427) |
| 0.1 | 1.1 | GO:1903504 | regulation of mitotic cell cycle spindle assembly checkpoint(GO:0090266) regulation of mitotic spindle checkpoint(GO:1903504) |
| 0.1 | 0.5 | GO:0032532 | regulation of microvillus length(GO:0032532) |
| 0.1 | 0.9 | GO:0032754 | positive regulation of interleukin-5 production(GO:0032754) |
| 0.1 | 0.4 | GO:2000786 | positive regulation of autophagosome assembly(GO:2000786) |
| 0.1 | 1.4 | GO:0032456 | endocytic recycling(GO:0032456) |
| 0.1 | 1.3 | GO:0007041 | lysosomal transport(GO:0007041) |
| 0.1 | 0.3 | GO:0009992 | cellular water homeostasis(GO:0009992) |
| 0.1 | 1.1 | GO:0039703 | viral RNA genome replication(GO:0039694) RNA replication(GO:0039703) |
| 0.1 | 0.3 | GO:0008090 | retrograde axonal transport(GO:0008090) |
| 0.0 | 1.0 | GO:0009081 | branched-chain amino acid metabolic process(GO:0009081) |
| 0.0 | 0.5 | GO:2001224 | positive regulation of neuron migration(GO:2001224) |
| 0.0 | 0.2 | GO:0070125 | mitochondrial translational elongation(GO:0070125) |
| 0.0 | 1.2 | GO:2000785 | regulation of autophagosome assembly(GO:2000785) |
| 0.0 | 1.5 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.0 | 1.2 | GO:0061001 | regulation of dendritic spine morphogenesis(GO:0061001) |
| 0.0 | 0.7 | GO:0006309 | apoptotic DNA fragmentation(GO:0006309) |
| 0.0 | 2.3 | GO:0007029 | endoplasmic reticulum organization(GO:0007029) |
| 0.0 | 0.5 | GO:0023019 | signal transduction involved in regulation of gene expression(GO:0023019) |
| 0.0 | 3.0 | GO:0009062 | fatty acid catabolic process(GO:0009062) |
| 0.0 | 0.1 | GO:1990022 | RNA polymerase II complex import to nucleus(GO:0044376) RNA polymerase III complex localization to nucleus(GO:1990022) |
| 0.0 | 0.4 | GO:1901409 | positive regulation of phosphorylation of RNA polymerase II C-terminal domain(GO:1901409) |
| 0.0 | 1.8 | GO:0045880 | positive regulation of smoothened signaling pathway(GO:0045880) |
| 0.0 | 0.2 | GO:0043102 | amino acid salvage(GO:0043102) L-methionine salvage(GO:0071267) |
| 0.0 | 0.7 | GO:0050650 | chondroitin sulfate proteoglycan biosynthetic process(GO:0050650) |
| 0.0 | 0.9 | GO:0071577 | zinc II ion transmembrane transport(GO:0071577) |
| 0.0 | 1.2 | GO:0009072 | aromatic amino acid family metabolic process(GO:0009072) |
| 0.0 | 5.1 | GO:0019236 | response to pheromone(GO:0019236) |
| 0.0 | 0.4 | GO:0006024 | glycosaminoglycan biosynthetic process(GO:0006024) |
| 0.0 | 0.4 | GO:0051026 | chiasma assembly(GO:0051026) |
| 0.0 | 0.2 | GO:0035721 | intraciliary retrograde transport(GO:0035721) |
| 0.0 | 0.4 | GO:0032776 | DNA methylation on cytosine(GO:0032776) |
| 0.0 | 0.4 | GO:0006610 | ribosomal protein import into nucleus(GO:0006610) |
| 0.0 | 0.3 | GO:0042048 | olfactory behavior(GO:0042048) |
| 0.0 | 0.2 | GO:0007403 | glial cell fate determination(GO:0007403) |
| 0.0 | 0.4 | GO:0045040 | protein import into mitochondrial outer membrane(GO:0045040) |
| 0.0 | 0.2 | GO:0050847 | progesterone receptor signaling pathway(GO:0050847) |
| 0.0 | 0.6 | GO:0017144 | drug metabolic process(GO:0017144) |
| 0.0 | 0.0 | GO:0003221 | right ventricular cardiac muscle tissue morphogenesis(GO:0003221) |
| 0.0 | 0.5 | GO:0016973 | poly(A)+ mRNA export from nucleus(GO:0016973) |
| 0.0 | 1.1 | GO:0046677 | response to antibiotic(GO:0046677) |
| 0.0 | 1.2 | GO:0071277 | cellular response to calcium ion(GO:0071277) |
| 0.0 | 0.1 | GO:0006004 | fucose metabolic process(GO:0006004) |
| 0.0 | 0.3 | GO:0016226 | iron-sulfur cluster assembly(GO:0016226) metallo-sulfur cluster assembly(GO:0031163) |
| 0.0 | 0.6 | GO:0006458 | 'de novo' protein folding(GO:0006458) |
| 0.0 | 0.2 | GO:0006536 | glutamate metabolic process(GO:0006536) |
| 0.0 | 2.8 | GO:0034284 | response to hexose(GO:0009746) response to monosaccharide(GO:0034284) |
| 0.0 | 0.1 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 0.0 | 0.2 | GO:0030948 | negative regulation of vascular endothelial growth factor receptor signaling pathway(GO:0030948) |
| 0.0 | 1.2 | GO:0032543 | mitochondrial translation(GO:0032543) |
| 0.0 | 0.3 | GO:0046339 | diacylglycerol metabolic process(GO:0046339) |
| 0.0 | 0.1 | GO:0031022 | nuclear migration along microfilament(GO:0031022) |
| 0.0 | 21.2 | GO:0007608 | sensory perception of smell(GO:0007608) |
| 0.0 | 0.8 | GO:0045600 | positive regulation of fat cell differentiation(GO:0045600) |
| 0.0 | 0.5 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.0 | 0.2 | GO:0043248 | proteasome assembly(GO:0043248) |
| 0.0 | 0.1 | GO:0000492 | box C/D snoRNP assembly(GO:0000492) |
| 0.0 | 0.7 | GO:0006637 | acyl-CoA metabolic process(GO:0006637) thioester metabolic process(GO:0035383) |
| 0.0 | 1.0 | GO:0031532 | actin cytoskeleton reorganization(GO:0031532) |
| 0.0 | 1.0 | GO:0006885 | regulation of pH(GO:0006885) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.7 | 50.9 | GO:0045298 | tubulin complex(GO:0045298) |
| 3.8 | 11.3 | GO:0044302 | dentate gyrus mossy fiber(GO:0044302) |
| 2.7 | 13.3 | GO:0097059 | CNTFR-CLCF1 complex(GO:0097059) |
| 2.2 | 8.8 | GO:0014701 | junctional sarcoplasmic reticulum membrane(GO:0014701) |
| 2.1 | 6.3 | GO:1990843 | subsarcolemmal mitochondrion(GO:1990843) interfibrillar mitochondrion(GO:1990844) |
| 2.0 | 6.0 | GO:0005967 | mitochondrial pyruvate dehydrogenase complex(GO:0005967) |
| 1.6 | 4.7 | GO:0005863 | striated muscle myosin thick filament(GO:0005863) |
| 1.5 | 25.1 | GO:0043203 | axon hillock(GO:0043203) |
| 1.5 | 5.9 | GO:1990032 | parallel fiber(GO:1990032) |
| 1.4 | 96.5 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 1.2 | 17.4 | GO:0044327 | dendritic spine head(GO:0044327) |
| 1.2 | 6.2 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 1.2 | 4.6 | GO:1990769 | proximal neuron projection(GO:1990769) |
| 1.2 | 3.5 | GO:0034677 | integrin alpha7-beta1 complex(GO:0034677) |
| 1.0 | 26.8 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 1.0 | 8.1 | GO:0098574 | cytoplasmic side of lysosomal membrane(GO:0098574) |
| 0.9 | 8.4 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.9 | 9.9 | GO:0031931 | TORC1 complex(GO:0031931) |
| 0.9 | 20.8 | GO:0098563 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.8 | 14.9 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 0.8 | 18.8 | GO:0032982 | myosin filament(GO:0032982) |
| 0.8 | 4.6 | GO:0042719 | mitochondrial intermembrane space protein transporter complex(GO:0042719) |
| 0.8 | 2.3 | GO:0005607 | laminin-2 complex(GO:0005607) |
| 0.8 | 11.3 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.7 | 3.0 | GO:0034686 | integrin alphav-beta8 complex(GO:0034686) |
| 0.7 | 3.7 | GO:1990130 | Iml1 complex(GO:1990130) Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.7 | 8.5 | GO:0032279 | asymmetric synapse(GO:0032279) |
| 0.7 | 4.3 | GO:0008091 | spectrin(GO:0008091) |
| 0.7 | 4.8 | GO:0048237 | rough endoplasmic reticulum lumen(GO:0048237) |
| 0.7 | 20.3 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.7 | 2.6 | GO:0032437 | cuticular plate(GO:0032437) |
| 0.6 | 7.8 | GO:0035748 | myelin sheath abaxonal region(GO:0035748) |
| 0.6 | 7.7 | GO:0098845 | postsynaptic endosome(GO:0098845) |
| 0.6 | 11.5 | GO:0045180 | basal cortex(GO:0045180) |
| 0.6 | 1.8 | GO:0044316 | cone cell pedicle(GO:0044316) |
| 0.6 | 6.4 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.6 | 1.7 | GO:0030956 | glutamyl-tRNA(Gln) amidotransferase complex(GO:0030956) |
| 0.6 | 6.3 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.6 | 1.7 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 0.6 | 4.5 | GO:1990812 | growth cone filopodium(GO:1990812) |
| 0.6 | 4.5 | GO:0045009 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.5 | 3.2 | GO:0070449 | elongin complex(GO:0070449) |
| 0.5 | 3.1 | GO:0005947 | mitochondrial alpha-ketoglutarate dehydrogenase complex(GO:0005947) |
| 0.5 | 15.5 | GO:0071565 | nBAF complex(GO:0071565) |
| 0.5 | 56.6 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.5 | 6.1 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.5 | 6.7 | GO:0001520 | outer dense fiber(GO:0001520) |
| 0.5 | 13.1 | GO:0090545 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.5 | 32.8 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.5 | 1.4 | GO:0033596 | TSC1-TSC2 complex(GO:0033596) |
| 0.5 | 14.9 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.4 | 5.7 | GO:0031315 | extrinsic component of mitochondrial outer membrane(GO:0031315) |
| 0.4 | 16.0 | GO:0030673 | axolemma(GO:0030673) |
| 0.4 | 1.7 | GO:0070722 | Tle3-Aes complex(GO:0070722) |
| 0.4 | 4.2 | GO:0005883 | neurofilament(GO:0005883) |
| 0.4 | 9.2 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 0.4 | 14.2 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.4 | 2.4 | GO:0070876 | SOSS complex(GO:0070876) |
| 0.4 | 4.6 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.4 | 2.3 | GO:0000235 | astral microtubule(GO:0000235) aster(GO:0005818) |
| 0.4 | 5.6 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.4 | 2.9 | GO:0044292 | dendrite terminus(GO:0044292) |
| 0.4 | 4.6 | GO:0005861 | troponin complex(GO:0005861) |
| 0.3 | 2.1 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.3 | 0.9 | GO:0097361 | CIA complex(GO:0097361) |
| 0.3 | 1.8 | GO:0030314 | junctional membrane complex(GO:0030314) |
| 0.3 | 1.4 | GO:1990745 | EARP complex(GO:1990745) |
| 0.3 | 9.6 | GO:0005790 | smooth endoplasmic reticulum(GO:0005790) |
| 0.3 | 7.4 | GO:0005922 | connexon complex(GO:0005922) |
| 0.3 | 3.2 | GO:0005859 | muscle myosin complex(GO:0005859) |
| 0.3 | 3.9 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.3 | 2.1 | GO:0017071 | intracellular cyclic nucleotide activated cation channel complex(GO:0017071) |
| 0.2 | 1.4 | GO:0043527 | tRNA methyltransferase complex(GO:0043527) |
| 0.2 | 2.5 | GO:0044613 | nuclear pore central transport channel(GO:0044613) |
| 0.2 | 1.8 | GO:0033263 | CORVET complex(GO:0033263) |
| 0.2 | 44.1 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.2 | 14.3 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.2 | 6.3 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.2 | 2.3 | GO:0000137 | Golgi cis cisterna(GO:0000137) |
| 0.2 | 1.5 | GO:0042765 | GPI-anchor transamidase complex(GO:0042765) |
| 0.2 | 3.7 | GO:0043196 | varicosity(GO:0043196) |
| 0.2 | 1.4 | GO:0044300 | cerebellar mossy fiber(GO:0044300) |
| 0.2 | 2.4 | GO:0016600 | flotillin complex(GO:0016600) |
| 0.2 | 3.3 | GO:0045263 | proton-transporting ATP synthase complex, coupling factor F(o)(GO:0045263) |
| 0.2 | 15.6 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) |
| 0.2 | 1.3 | GO:0071914 | prominosome(GO:0071914) |
| 0.2 | 4.6 | GO:0032589 | neuron projection membrane(GO:0032589) |
| 0.2 | 18.9 | GO:0030315 | T-tubule(GO:0030315) |
| 0.2 | 7.6 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.2 | 4.5 | GO:0043218 | compact myelin(GO:0043218) |
| 0.2 | 1.7 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.2 | 0.7 | GO:1990452 | Parkin-FBXW7-Cul1 ubiquitin ligase complex(GO:1990452) |
| 0.2 | 2.2 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 0.2 | 3.5 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.2 | 2.7 | GO:0001939 | female pronucleus(GO:0001939) |
| 0.2 | 6.5 | GO:0030660 | Golgi-associated vesicle membrane(GO:0030660) |
| 0.2 | 45.8 | GO:0014069 | postsynaptic density(GO:0014069) |
| 0.2 | 0.9 | GO:1903439 | calcitonin family receptor complex(GO:1903439) amylin receptor complex(GO:1903440) |
| 0.1 | 10.2 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.1 | 2.1 | GO:0031371 | ubiquitin conjugating enzyme complex(GO:0031371) |
| 0.1 | 1.2 | GO:0061617 | MICOS complex(GO:0061617) |
| 0.1 | 0.6 | GO:0072534 | perineuronal net(GO:0072534) |
| 0.1 | 3.9 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.1 | 12.3 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.1 | 7.0 | GO:0030964 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.1 | 0.9 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.1 | 6.9 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.1 | 1.0 | GO:0005750 | mitochondrial respiratory chain complex III(GO:0005750) |
| 0.1 | 12.0 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.1 | 7.9 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.1 | 1.0 | GO:1990712 | HFE-transferrin receptor complex(GO:1990712) |
| 0.1 | 0.4 | GO:0042272 | nuclear RNA export factor complex(GO:0042272) |
| 0.1 | 1.1 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.1 | 3.0 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.1 | 8.0 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.1 | 23.3 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.1 | 1.5 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 0.1 | 3.8 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.1 | 1.4 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 0.1 | 2.1 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.1 | 2.6 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.1 | 1.7 | GO:0034045 | pre-autophagosomal structure membrane(GO:0034045) |
| 0.1 | 9.2 | GO:0030018 | Z disc(GO:0030018) |
| 0.1 | 17.8 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.1 | 3.7 | GO:0035869 | ciliary transition zone(GO:0035869) |
| 0.1 | 0.6 | GO:0036396 | MIS complex(GO:0036396) |
| 0.1 | 1.0 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.1 | 0.2 | GO:0031310 | integral component of vacuolar membrane(GO:0031166) intrinsic component of vacuolar membrane(GO:0031310) |
| 0.1 | 24.3 | GO:0000139 | Golgi membrane(GO:0000139) |
| 0.1 | 4.1 | GO:0005770 | late endosome(GO:0005770) |
| 0.1 | 2.2 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.1 | 0.9 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.1 | 37.3 | GO:0045202 | synapse(GO:0045202) |
| 0.1 | 3.3 | GO:0016459 | myosin complex(GO:0016459) |
| 0.1 | 10.5 | GO:0043292 | contractile fiber(GO:0043292) |
| 0.1 | 0.4 | GO:0035032 | phosphatidylinositol 3-kinase complex, class III(GO:0035032) |
| 0.1 | 4.0 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.1 | 1.2 | GO:0043205 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.1 | 0.3 | GO:0097550 | transcriptional preinitiation complex(GO:0097550) |
| 0.1 | 2.0 | GO:0005771 | multivesicular body(GO:0005771) |
| 0.1 | 0.2 | GO:0045257 | mitochondrial respiratory chain complex II, succinate dehydrogenase complex (ubiquinone)(GO:0005749) succinate dehydrogenase complex (ubiquinone)(GO:0045257) respiratory chain complex II(GO:0045273) succinate dehydrogenase complex(GO:0045281) fumarate reductase complex(GO:0045283) |
| 0.1 | 11.6 | GO:0031301 | integral component of organelle membrane(GO:0031301) |
| 0.1 | 3.7 | GO:0005901 | caveola(GO:0005901) |
| 0.1 | 19.8 | GO:0005764 | lytic vacuole(GO:0000323) lysosome(GO:0005764) |
| 0.1 | 0.7 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.0 | 1.7 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.0 | 0.2 | GO:0097255 | R2TP complex(GO:0097255) |
| 0.0 | 1.0 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.0 | 6.9 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 1.1 | GO:0071782 | endoplasmic reticulum tubular network(GO:0071782) endoplasmic reticulum subcompartment(GO:0098827) |
| 0.0 | 1.1 | GO:0005838 | proteasome regulatory particle(GO:0005838) |
| 0.0 | 2.7 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.0 | 0.7 | GO:0000421 | autophagosome membrane(GO:0000421) |
| 0.0 | 0.6 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.0 | 2.3 | GO:0055037 | recycling endosome(GO:0055037) |
| 0.0 | 0.2 | GO:0044614 | nuclear pore cytoplasmic filaments(GO:0044614) |
| 0.0 | 10.4 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.0 | 0.7 | GO:0000124 | SAGA complex(GO:0000124) |
| 0.0 | 14.9 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 3.4 | GO:0035097 | histone methyltransferase complex(GO:0035097) |
| 0.0 | 69.0 | GO:0005739 | mitochondrion(GO:0005739) |
| 0.0 | 0.9 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.0 | 0.4 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.0 | 0.4 | GO:0005797 | Golgi medial cisterna(GO:0005797) |
| 0.0 | 0.2 | GO:0034708 | methyltransferase complex(GO:0034708) |
| 0.0 | 2.9 | GO:0031012 | extracellular matrix(GO:0031012) |
| 0.0 | 0.0 | GO:0044317 | rod spherule(GO:0044317) |
| 0.0 | 0.1 | GO:0019773 | proteasome core complex, alpha-subunit complex(GO:0019773) |
| 0.0 | 2.6 | GO:0005635 | nuclear envelope(GO:0005635) |
| 0.0 | 0.4 | GO:0001741 | XY body(GO:0001741) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 13.1 | 39.3 | GO:0016492 | G-protein coupled neurotensin receptor activity(GO:0016492) |
| 10.2 | 50.9 | GO:0099609 | microtubule lateral binding(GO:0099609) |
| 5.6 | 33.8 | GO:0060175 | brain-derived neurotrophic factor-activated receptor activity(GO:0060175) |
| 4.9 | 14.7 | GO:0086062 | voltage-gated sodium channel activity involved in Purkinje myocyte action potential(GO:0086062) |
| 3.4 | 13.6 | GO:0002153 | steroid receptor RNA activator RNA binding(GO:0002153) |
| 2.8 | 8.3 | GO:0004658 | propionyl-CoA carboxylase activity(GO:0004658) |
| 2.7 | 10.8 | GO:0001588 | dopamine neurotransmitter receptor activity, coupled via Gs(GO:0001588) |
| 2.6 | 7.9 | GO:0004470 | malic enzyme activity(GO:0004470) malate dehydrogenase (decarboxylating) (NAD+) activity(GO:0004471) malate dehydrogenase (decarboxylating) (NADP+) activity(GO:0004473) |
| 2.4 | 12.1 | GO:0086006 | voltage-gated sodium channel activity involved in cardiac muscle cell action potential(GO:0086006) |
| 2.4 | 7.1 | GO:0019002 | GMP binding(GO:0019002) |
| 2.3 | 9.3 | GO:0004105 | choline-phosphate cytidylyltransferase activity(GO:0004105) |
| 2.3 | 9.3 | GO:0004740 | pyruvate dehydrogenase (acetyl-transferring) kinase activity(GO:0004740) |
| 2.3 | 6.9 | GO:0043404 | corticotrophin-releasing factor receptor activity(GO:0015056) corticotropin-releasing hormone receptor activity(GO:0043404) |
| 2.2 | 13.1 | GO:0003839 | gamma-glutamylcyclotransferase activity(GO:0003839) |
| 2.1 | 6.4 | GO:0004001 | adenosine kinase activity(GO:0004001) |
| 2.1 | 4.2 | GO:0004517 | nitric-oxide synthase activity(GO:0004517) |
| 2.1 | 6.2 | GO:0016901 | glycerol-3-phosphate dehydrogenase activity(GO:0004368) oxidoreductase activity, acting on the CH-OH group of donors, quinone or similar compound as acceptor(GO:0016901) |
| 2.0 | 8.1 | GO:0061628 | H3K27me3 modified histone binding(GO:0061628) |
| 2.0 | 18.1 | GO:0032795 | heterotrimeric G-protein binding(GO:0032795) |
| 2.0 | 13.8 | GO:0004351 | glutamate decarboxylase activity(GO:0004351) |
| 1.8 | 12.3 | GO:0032027 | myosin light chain binding(GO:0032027) |
| 1.7 | 6.7 | GO:0046997 | oxidoreductase activity, acting on the CH-NH group of donors, flavin as acceptor(GO:0046997) |
| 1.6 | 4.8 | GO:0030348 | syntaxin-3 binding(GO:0030348) |
| 1.6 | 9.5 | GO:0046920 | alpha-(1->3)-fucosyltransferase activity(GO:0046920) |
| 1.6 | 1.6 | GO:0008948 | oxaloacetate decarboxylase activity(GO:0008948) |
| 1.5 | 10.6 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 1.5 | 13.3 | GO:0004897 | ciliary neurotrophic factor receptor activity(GO:0004897) |
| 1.4 | 5.6 | GO:0030156 | benzodiazepine receptor binding(GO:0030156) |
| 1.4 | 4.1 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 1.3 | 8.1 | GO:0004994 | somatostatin receptor activity(GO:0004994) |
| 1.3 | 5.3 | GO:0000010 | trans-hexaprenyltranstransferase activity(GO:0000010) trans-octaprenyltranstransferase activity(GO:0050347) |
| 1.3 | 5.2 | GO:0004346 | glucose-6-phosphatase activity(GO:0004346) sugar-terminal-phosphatase activity(GO:0050309) |
| 1.3 | 5.1 | GO:0004711 | ribosomal protein S6 kinase activity(GO:0004711) |
| 1.3 | 5.1 | GO:1902379 | chemoattractant activity involved in axon guidance(GO:1902379) |
| 1.3 | 3.8 | GO:0004828 | serine-tRNA ligase activity(GO:0004828) |
| 1.2 | 13.1 | GO:0022820 | potassium:chloride symporter activity(GO:0015379) potassium ion symporter activity(GO:0022820) |
| 1.2 | 4.8 | GO:0004830 | tryptophan-tRNA ligase activity(GO:0004830) |
| 1.1 | 11.4 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 1.1 | 4.6 | GO:0047275 | glucosaminylgalactosylglucosylceramide beta-galactosyltransferase activity(GO:0047275) |
| 1.1 | 5.7 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 1.1 | 9.1 | GO:0005042 | netrin receptor activity(GO:0005042) |
| 1.1 | 2.3 | GO:0001639 | PLC activating G-protein coupled glutamate receptor activity(GO:0001639) G-protein coupled receptor activity involved in regulation of postsynaptic membrane potential(GO:0099530) |
| 1.1 | 3.4 | GO:0010428 | methyl-CpNpG binding(GO:0010428) |
| 1.1 | 20.5 | GO:0033549 | MAP kinase phosphatase activity(GO:0033549) |
| 1.1 | 3.2 | GO:0051538 | 3 iron, 4 sulfur cluster binding(GO:0051538) |
| 1.0 | 6.3 | GO:0031750 | D3 dopamine receptor binding(GO:0031750) |
| 1.0 | 3.1 | GO:0016015 | morphogen activity(GO:0016015) |
| 1.0 | 35.7 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 1.0 | 3.0 | GO:0004774 | succinate-CoA ligase activity(GO:0004774) |
| 1.0 | 3.0 | GO:0016155 | formyltetrahydrofolate dehydrogenase activity(GO:0016155) |
| 1.0 | 2.9 | GO:0034189 | very-low-density lipoprotein particle binding(GO:0034189) |
| 1.0 | 8.6 | GO:0051022 | Rho GDP-dissociation inhibitor binding(GO:0051022) |
| 0.9 | 18.8 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.9 | 2.8 | GO:0001566 | phorbol ester receptor activity(GO:0001565) non-kinase phorbol ester receptor activity(GO:0001566) |
| 0.9 | 5.5 | GO:0004647 | phosphoserine phosphatase activity(GO:0004647) |
| 0.9 | 4.5 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.9 | 6.9 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.8 | 6.8 | GO:0004416 | hydroxyacylglutathione hydrolase activity(GO:0004416) |
| 0.8 | 3.4 | GO:0034617 | tetrahydrobiopterin binding(GO:0034617) |
| 0.8 | 5.1 | GO:0030250 | cyclase activator activity(GO:0010853) guanylate cyclase activator activity(GO:0030250) |
| 0.8 | 3.3 | GO:0031687 | A2A adenosine receptor binding(GO:0031687) |
| 0.8 | 3.2 | GO:0031800 | type 3 metabotropic glutamate receptor binding(GO:0031800) |
| 0.8 | 3.2 | GO:0004092 | carnitine O-acetyltransferase activity(GO:0004092) |
| 0.8 | 5.5 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.8 | 2.3 | GO:0003858 | 3-hydroxybutyrate dehydrogenase activity(GO:0003858) |
| 0.8 | 2.3 | GO:0004968 | gonadotropin-releasing hormone receptor activity(GO:0004968) |
| 0.8 | 4.6 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 0.7 | 2.2 | GO:0080130 | L-phenylalanine:2-oxoglutarate aminotransferase activity(GO:0080130) |
| 0.7 | 16.3 | GO:0005003 | ephrin receptor activity(GO:0005003) |
| 0.7 | 5.9 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 0.7 | 21.8 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.7 | 8.5 | GO:0032051 | clathrin light chain binding(GO:0032051) |
| 0.7 | 5.6 | GO:0001595 | angiotensin receptor activity(GO:0001595) |
| 0.7 | 8.2 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.7 | 10.1 | GO:0031432 | titin binding(GO:0031432) |
| 0.7 | 4.0 | GO:0034186 | apolipoprotein A-I binding(GO:0034186) |
| 0.7 | 9.9 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 0.7 | 4.6 | GO:0030172 | troponin C binding(GO:0030172) calcium-dependent ATPase activity(GO:0030899) troponin I binding(GO:0031013) |
| 0.6 | 5.1 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.6 | 7.0 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) ADP transmembrane transporter activity(GO:0015217) |
| 0.6 | 1.8 | GO:0019150 | D-ribulokinase activity(GO:0019150) |
| 0.6 | 1.8 | GO:0004967 | glucagon receptor activity(GO:0004967) |
| 0.6 | 2.3 | GO:0016833 | oxo-acid-lyase activity(GO:0016833) |
| 0.6 | 1.7 | GO:0050567 | glutaminyl-tRNA synthase (glutamine-hydrolyzing) activity(GO:0050567) |
| 0.6 | 2.9 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.6 | 3.5 | GO:0004118 | cGMP-stimulated cyclic-nucleotide phosphodiesterase activity(GO:0004118) |
| 0.6 | 8.0 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 0.6 | 12.5 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.6 | 5.6 | GO:0022851 | GABA-gated chloride ion channel activity(GO:0022851) |
| 0.5 | 5.4 | GO:0016861 | intramolecular oxidoreductase activity, interconverting aldoses and ketoses(GO:0016861) |
| 0.5 | 2.7 | GO:0055056 | dehydroascorbic acid transporter activity(GO:0033300) D-glucose transmembrane transporter activity(GO:0055056) |
| 0.5 | 4.3 | GO:0060072 | large conductance calcium-activated potassium channel activity(GO:0060072) |
| 0.5 | 1.6 | GO:0031711 | bradykinin receptor binding(GO:0031711) |
| 0.5 | 4.7 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.5 | 9.3 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 0.5 | 3.5 | GO:0030160 | GKAP/Homer scaffold activity(GO:0030160) |
| 0.5 | 5.0 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.5 | 8.0 | GO:0042301 | phosphate ion binding(GO:0042301) |
| 0.5 | 2.0 | GO:0004566 | beta-glucuronidase activity(GO:0004566) |
| 0.5 | 2.0 | GO:0015403 | thiamine uptake transmembrane transporter activity(GO:0015403) |
| 0.5 | 3.4 | GO:0001642 | group III metabotropic glutamate receptor activity(GO:0001642) |
| 0.5 | 3.4 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.5 | 14.8 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.5 | 1.9 | GO:0072541 | peroxynitrite reductase activity(GO:0072541) |
| 0.5 | 8.5 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.5 | 3.2 | GO:0001517 | N-acetylglucosamine 6-O-sulfotransferase activity(GO:0001517) |
| 0.5 | 5.9 | GO:0019855 | calcium channel inhibitor activity(GO:0019855) |
| 0.4 | 2.7 | GO:0004505 | phenylalanine 4-monooxygenase activity(GO:0004505) |
| 0.4 | 16.1 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.4 | 16.9 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.4 | 3.0 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.4 | 1.7 | GO:0036478 | tyrosine 3-monooxygenase activator activity(GO:0036470) L-dopa decarboxylase activator activity(GO:0036478) |
| 0.4 | 2.0 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.4 | 1.2 | GO:0003868 | 4-hydroxyphenylpyruvate dioxygenase activity(GO:0003868) |
| 0.4 | 2.0 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.4 | 5.9 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.4 | 9.6 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.4 | 4.6 | GO:0102338 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.4 | 4.9 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) |
| 0.4 | 10.4 | GO:0070300 | phosphatidic acid binding(GO:0070300) |
| 0.4 | 6.2 | GO:0001206 | transcriptional repressor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001206) |
| 0.4 | 1.4 | GO:0008176 | tRNA (guanine-N7-)-methyltransferase activity(GO:0008176) |
| 0.3 | 2.1 | GO:0046624 | sphingolipid transporter activity(GO:0046624) |
| 0.3 | 2.1 | GO:0005222 | intracellular cAMP activated cation channel activity(GO:0005222) |
| 0.3 | 2.4 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.3 | 7.5 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.3 | 13.5 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.3 | 2.4 | GO:0097109 | neuroligin family protein binding(GO:0097109) |
| 0.3 | 1.3 | GO:0008260 | 3-oxoacid CoA-transferase activity(GO:0008260) |
| 0.3 | 3.3 | GO:0015193 | L-proline transmembrane transporter activity(GO:0015193) |
| 0.3 | 2.0 | GO:0004614 | phosphoglucomutase activity(GO:0004614) |
| 0.3 | 8.4 | GO:0005227 | calcium activated cation channel activity(GO:0005227) |
| 0.3 | 0.9 | GO:0005292 | high-affinity basic amino acid transmembrane transporter activity(GO:0005287) high-affinity arginine transmembrane transporter activity(GO:0005289) high-affinity lysine transmembrane transporter activity(GO:0005292) |
| 0.3 | 0.9 | GO:0004948 | calcitonin receptor activity(GO:0004948) |
| 0.3 | 1.2 | GO:0004103 | choline kinase activity(GO:0004103) |
| 0.3 | 3.1 | GO:0001665 | alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase activity(GO:0001665) |
| 0.3 | 2.2 | GO:0048403 | brain-derived neurotrophic factor binding(GO:0048403) |
| 0.3 | 8.6 | GO:0097602 | cullin family protein binding(GO:0097602) |
| 0.3 | 4.2 | GO:0016595 | glutamate binding(GO:0016595) |
| 0.3 | 19.1 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.3 | 2.1 | GO:0016421 | CoA carboxylase activity(GO:0016421) |
| 0.3 | 2.1 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.3 | 3.0 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.3 | 5.4 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.3 | 18.8 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.3 | 7.1 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.3 | 6.3 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.3 | 4.0 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.3 | 1.1 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
| 0.3 | 2.9 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.3 | 1.1 | GO:0030911 | TPR domain binding(GO:0030911) |
| 0.3 | 1.6 | GO:0019958 | C-X-C chemokine binding(GO:0019958) |
| 0.3 | 2.6 | GO:0003876 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.3 | 2.1 | GO:0031821 | G-protein coupled serotonin receptor binding(GO:0031821) |
| 0.3 | 4.7 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.3 | 5.2 | GO:0005248 | voltage-gated sodium channel activity(GO:0005248) voltage-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1905030) |
| 0.3 | 1.0 | GO:0008113 | peptide-methionine (S)-S-oxide reductase activity(GO:0008113) |
| 0.3 | 26.8 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
| 0.3 | 3.3 | GO:0005314 | high-affinity glutamate transmembrane transporter activity(GO:0005314) |
| 0.2 | 7.2 | GO:0010485 | H4 histone acetyltransferase activity(GO:0010485) |
| 0.2 | 10.4 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.2 | 21.1 | GO:0030276 | clathrin binding(GO:0030276) |
| 0.2 | 2.4 | GO:0016670 | oxidoreductase activity, acting on a sulfur group of donors, oxygen as acceptor(GO:0016670) |
| 0.2 | 3.1 | GO:0004439 | phosphatidylinositol-4,5-bisphosphate 5-phosphatase activity(GO:0004439) |
| 0.2 | 9.5 | GO:0008009 | chemokine activity(GO:0008009) |
| 0.2 | 4.5 | GO:0004559 | alpha-mannosidase activity(GO:0004559) |
| 0.2 | 0.7 | GO:0050816 | phosphothreonine binding(GO:0050816) |
| 0.2 | 26.3 | GO:0005249 | voltage-gated potassium channel activity(GO:0005249) |
| 0.2 | 0.9 | GO:0032564 | dATP binding(GO:0032564) |
| 0.2 | 2.0 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.2 | 1.5 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.2 | 2.4 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.2 | 1.7 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.2 | 0.9 | GO:0005093 | Rab GDP-dissociation inhibitor activity(GO:0005093) |
| 0.2 | 1.1 | GO:0071987 | WD40-repeat domain binding(GO:0071987) |
| 0.2 | 0.8 | GO:0000064 | L-ornithine transmembrane transporter activity(GO:0000064) |
| 0.2 | 1.0 | GO:0018479 | benzaldehyde dehydrogenase (NAD+) activity(GO:0018479) |
| 0.2 | 1.5 | GO:0003923 | GPI-anchor transamidase activity(GO:0003923) |
| 0.2 | 1.4 | GO:0005087 | Ran guanyl-nucleotide exchange factor activity(GO:0005087) |
| 0.2 | 10.1 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.2 | 3.7 | GO:0005243 | gap junction channel activity(GO:0005243) |
| 0.2 | 1.2 | GO:0004792 | thiosulfate sulfurtransferase activity(GO:0004792) |
| 0.2 | 4.7 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.2 | 4.4 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.2 | 2.2 | GO:0051870 | methotrexate binding(GO:0051870) |
| 0.2 | 1.8 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.2 | 11.7 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.2 | 1.0 | GO:0004998 | transferrin receptor activity(GO:0004998) |
| 0.2 | 4.1 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.2 | 5.3 | GO:0030742 | GTP-dependent protein binding(GO:0030742) |
| 0.2 | 4.7 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.2 | 0.6 | GO:0051185 | coenzyme transporter activity(GO:0051185) |
| 0.2 | 1.0 | GO:0016499 | orexin receptor activity(GO:0016499) |
| 0.2 | 1.7 | GO:0034711 | inhibin binding(GO:0034711) |
| 0.2 | 3.4 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.2 | 1.9 | GO:0008889 | glycerophosphodiester phosphodiesterase activity(GO:0008889) |
| 0.2 | 0.7 | GO:0019212 | phosphatase inhibitor activity(GO:0019212) |
| 0.2 | 11.2 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.2 | 1.6 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.2 | 0.9 | GO:0002046 | opsin binding(GO:0002046) |
| 0.2 | 0.8 | GO:0005143 | interleukin-12 receptor binding(GO:0005143) |
| 0.2 | 5.4 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.2 | 2.6 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
| 0.2 | 1.3 | GO:0071253 | connexin binding(GO:0071253) |
| 0.2 | 0.5 | GO:0031752 | D5 dopamine receptor binding(GO:0031752) |
| 0.2 | 2.8 | GO:0005452 | inorganic anion exchanger activity(GO:0005452) |
| 0.2 | 0.6 | GO:0097001 | ceramide binding(GO:0097001) |
| 0.1 | 2.1 | GO:0051378 | serotonin binding(GO:0051378) |
| 0.1 | 3.3 | GO:0042923 | neuropeptide binding(GO:0042923) |
| 0.1 | 1.5 | GO:0030550 | acetylcholine receptor inhibitor activity(GO:0030550) |
| 0.1 | 0.4 | GO:0008499 | UDP-galactose:beta-N-acetylglucosamine beta-1,3-galactosyltransferase activity(GO:0008499) |
| 0.1 | 2.8 | GO:0070403 | NAD+ binding(GO:0070403) |
| 0.1 | 4.3 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.1 | 0.7 | GO:0015018 | galactosylgalactosylxylosylprotein 3-beta-glucuronosyltransferase activity(GO:0015018) |
| 0.1 | 0.4 | GO:0016824 | hydrolase activity, acting on acid halide bonds(GO:0016824) hydrolase activity, acting on acid halide bonds, in C-halide compounds(GO:0019120) alkylhalidase activity(GO:0047651) |
| 0.1 | 3.6 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.1 | 0.6 | GO:1904047 | mRNA (2'-O-methyladenosine-N6-)-methyltransferase activity(GO:0016422) S-adenosyl-L-methionine binding(GO:1904047) |
| 0.1 | 2.5 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.1 | 3.5 | GO:0000030 | mannosyltransferase activity(GO:0000030) |
| 0.1 | 0.7 | GO:0004531 | deoxyribonuclease II activity(GO:0004531) |
| 0.1 | 0.8 | GO:0008273 | calcium, potassium:sodium antiporter activity(GO:0008273) |
| 0.1 | 0.4 | GO:0016309 | 1-phosphatidylinositol-5-phosphate 4-kinase activity(GO:0016309) |
| 0.1 | 1.2 | GO:0000268 | peroxisome targeting sequence binding(GO:0000268) |
| 0.1 | 0.9 | GO:0047184 | 1-acylglycerophosphocholine O-acyltransferase activity(GO:0047184) |
| 0.1 | 1.2 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 0.1 | 6.7 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.1 | 1.0 | GO:0008121 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 0.1 | 1.8 | GO:0003708 | retinoic acid receptor activity(GO:0003708) |
| 0.1 | 1.2 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.1 | 20.2 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 0.7 | GO:0004359 | glutaminase activity(GO:0004359) |
| 0.1 | 2.2 | GO:0019841 | retinol binding(GO:0019841) |
| 0.1 | 3.0 | GO:0004383 | guanylate cyclase activity(GO:0004383) |
| 0.1 | 2.5 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.1 | 0.8 | GO:0018685 | alkane 1-monooxygenase activity(GO:0018685) leukotriene-B4 20-monooxygenase activity(GO:0050051) |
| 0.1 | 1.2 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.1 | 5.5 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.1 | 3.9 | GO:0071949 | FAD binding(GO:0071949) |
| 0.1 | 3.3 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.1 | 1.9 | GO:0005337 | nucleoside transmembrane transporter activity(GO:0005337) |
| 0.1 | 0.6 | GO:0016634 | oxidoreductase activity, acting on the CH-CH group of donors, oxygen as acceptor(GO:0016634) |
| 0.1 | 1.1 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.1 | 6.6 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.1 | 3.1 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.1 | 1.5 | GO:0008430 | selenium binding(GO:0008430) |
| 0.1 | 1.0 | GO:0005283 | sodium:amino acid symporter activity(GO:0005283) |
| 0.1 | 4.8 | GO:0031369 | translation initiation factor binding(GO:0031369) |
| 0.1 | 4.3 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.1 | 0.6 | GO:0019808 | polyamine binding(GO:0019808) |
| 0.1 | 0.8 | GO:0005519 | cytoskeletal regulatory protein binding(GO:0005519) |
| 0.1 | 5.0 | GO:0016620 | oxidoreductase activity, acting on the aldehyde or oxo group of donors, NAD or NADP as acceptor(GO:0016620) |
| 0.1 | 4.1 | GO:0019213 | deacetylase activity(GO:0019213) |
| 0.1 | 0.4 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.1 | 0.8 | GO:0016892 | endoribonuclease activity, producing 3'-phosphomonoesters(GO:0016892) |
| 0.1 | 0.3 | GO:0050501 | hyaluronan synthase activity(GO:0050501) |
| 0.1 | 1.5 | GO:0042166 | acetylcholine binding(GO:0042166) |
| 0.1 | 3.2 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.1 | 1.0 | GO:0003956 | NAD(P)+-protein-arginine ADP-ribosyltransferase activity(GO:0003956) |
| 0.1 | 0.2 | GO:0031871 | proteinase activated receptor binding(GO:0031871) |
| 0.1 | 1.4 | GO:0045499 | chemorepellent activity(GO:0045499) |
| 0.1 | 0.4 | GO:0003987 | acetate-CoA ligase activity(GO:0003987) |
| 0.1 | 0.7 | GO:0034511 | U3 snoRNA binding(GO:0034511) |
| 0.1 | 1.2 | GO:0044548 | S100 protein binding(GO:0044548) |
| 0.1 | 0.2 | GO:0070698 | type I activin receptor binding(GO:0070698) |
| 0.1 | 2.4 | GO:0035255 | ionotropic glutamate receptor binding(GO:0035255) |
| 0.1 | 0.3 | GO:0008147 | structural constituent of bone(GO:0008147) |
| 0.1 | 4.2 | GO:1990939 | ATP-dependent microtubule motor activity(GO:1990939) |
| 0.1 | 0.3 | GO:0047696 | beta-adrenergic receptor kinase activity(GO:0047696) |
| 0.1 | 0.4 | GO:0033265 | choline binding(GO:0033265) |
| 0.1 | 1.5 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.1 | 0.4 | GO:0019788 | NEDD8 transferase activity(GO:0019788) |
| 0.1 | 0.2 | GO:0004821 | histidine-tRNA ligase activity(GO:0004821) |
| 0.1 | 4.2 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.1 | 0.6 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.1 | 0.9 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.1 | 0.4 | GO:0008174 | mRNA methyltransferase activity(GO:0008174) |
| 0.1 | 2.8 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 1.4 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.1 | 0.9 | GO:0004993 | G-protein coupled serotonin receptor activity(GO:0004993) |
| 0.1 | 7.1 | GO:0002020 | protease binding(GO:0002020) |
| 0.1 | 1.7 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
| 0.1 | 1.9 | GO:0030552 | cAMP binding(GO:0030552) |
| 0.1 | 1.5 | GO:0016676 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.0 | 0.3 | GO:0008453 | alanine-glyoxylate transaminase activity(GO:0008453) |
| 0.0 | 0.3 | GO:0030294 | receptor signaling protein tyrosine kinase inhibitor activity(GO:0030294) |
| 0.0 | 0.3 | GO:0000298 | endopolyphosphatase activity(GO:0000298) diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) bis(5'-adenosyl)-hexaphosphatase activity(GO:0034431) bis(5'-adenosyl)-pentaphosphatase activity(GO:0034432) |
| 0.0 | 1.3 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.0 | 5.1 | GO:0005550 | pheromone binding(GO:0005550) |
| 0.0 | 0.6 | GO:0004983 | neuropeptide Y receptor activity(GO:0004983) |
| 0.0 | 1.4 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.0 | 1.7 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.0 | 0.5 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.0 | 0.3 | GO:0004046 | aminoacylase activity(GO:0004046) |
| 0.0 | 2.6 | GO:0016820 | hydrolase activity, acting on acid anhydrides, catalyzing transmembrane movement of substances(GO:0016820) |
| 0.0 | 0.3 | GO:0047179 | platelet-activating factor acetyltransferase activity(GO:0047179) |
| 0.0 | 0.2 | GO:0032296 | ribonuclease III activity(GO:0004525) double-stranded RNA-specific ribonuclease activity(GO:0032296) |
| 0.0 | 2.2 | GO:0070888 | E-box binding(GO:0070888) |
| 0.0 | 0.5 | GO:0016500 | protein-hormone receptor activity(GO:0016500) |
| 0.0 | 0.3 | GO:0008656 | cysteine-type endopeptidase activator activity involved in apoptotic process(GO:0008656) |
| 0.0 | 0.3 | GO:0000990 | transcription factor activity, core RNA polymerase binding(GO:0000990) |
| 0.0 | 3.1 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.0 | 0.3 | GO:0004366 | glycerol-3-phosphate O-acyltransferase activity(GO:0004366) |
| 0.0 | 3.0 | GO:0003774 | motor activity(GO:0003774) |
| 0.0 | 0.1 | GO:0004560 | alpha-L-fucosidase activity(GO:0004560) fucosidase activity(GO:0015928) |
| 0.0 | 0.4 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.0 | 0.2 | GO:0044390 | ubiquitin-like protein conjugating enzyme binding(GO:0044390) |
| 0.0 | 3.1 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 9.5 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.0 | 0.9 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.0 | 3.0 | GO:0019209 | kinase activator activity(GO:0019209) |
| 0.0 | 2.3 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 0.8 | GO:0005112 | Notch binding(GO:0005112) |
| 0.0 | 0.2 | GO:0008172 | S-methyltransferase activity(GO:0008172) |
| 0.0 | 0.9 | GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.0 | 0.3 | GO:0070095 | fructose-6-phosphate binding(GO:0070095) |
| 0.0 | 0.6 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.0 | 11.0 | GO:0004930 | G-protein coupled receptor activity(GO:0004930) |
| 0.0 | 0.5 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.0 | 0.2 | GO:0005166 | neurotrophin p75 receptor binding(GO:0005166) |
| 0.0 | 0.2 | GO:0019864 | IgG binding(GO:0019864) |
| 0.0 | 2.4 | GO:0001191 | transcriptional repressor activity, RNA polymerase II transcription factor binding(GO:0001191) |
| 0.0 | 0.3 | GO:0036374 | glutathione hydrolase activity(GO:0036374) |
| 0.0 | 0.8 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.0 | 0.1 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.0 | 19.6 | GO:0004984 | olfactory receptor activity(GO:0004984) |
| 0.0 | 0.9 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 0.8 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 1.5 | GO:0015078 | hydrogen ion transmembrane transporter activity(GO:0015078) |
| 0.0 | 0.4 | GO:0016709 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen(GO:0016709) |
| 0.0 | 7.8 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 0.0 | 0.3 | GO:0070742 | C2H2 zinc finger domain binding(GO:0070742) |
| 0.0 | 0.4 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.0 | 0.3 | GO:0031435 | mitogen-activated protein kinase kinase kinase binding(GO:0031435) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.0 | 50.9 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.6 | 20.2 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.3 | 4.2 | ST INTEGRIN SIGNALING PATHWAY | Integrin Signaling Pathway |
| 0.3 | 20.1 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.3 | 16.9 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.3 | 35.4 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.3 | 20.5 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.2 | 11.7 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.2 | 14.0 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.2 | 2.3 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.2 | 1.6 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.2 | 2.6 | PID S1P S1P4 PATHWAY | S1P4 pathway |
| 0.2 | 4.7 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
| 0.2 | 5.4 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.2 | 10.2 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.2 | 11.1 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.2 | 3.7 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.1 | 3.1 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.1 | 3.5 | PID EPHB FWD PATHWAY | EPHB forward signaling |
| 0.1 | 5.7 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.1 | 4.8 | SIG CD40PATHWAYMAP | Genes related to CD40 signaling |
| 0.1 | 1.7 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.1 | 4.8 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.1 | 2.9 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.1 | 2.8 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.1 | 2.2 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.1 | 4.8 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.1 | 2.6 | PID IFNG PATHWAY | IFN-gamma pathway |
| 0.1 | 22.3 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.1 | 3.2 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.1 | 1.8 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.1 | 2.3 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.1 | 3.5 | PID ARF6 TRAFFICKING PATHWAY | Arf6 trafficking events |
| 0.1 | 6.5 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.1 | 3.5 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.1 | 0.5 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.1 | 2.0 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.1 | 3.1 | PID BMP PATHWAY | BMP receptor signaling |
| 0.1 | 1.0 | PID PI3K PLC TRK PATHWAY | Trk receptor signaling mediated by PI3K and PLC-gamma |
| 0.1 | 2.3 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.1 | 3.2 | PID ERBB1 DOWNSTREAM PATHWAY | ErbB1 downstream signaling |
| 0.1 | 0.4 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.0 | 1.9 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.0 | 0.8 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.0 | 0.4 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.0 | 2.3 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 1.0 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.0 | 10.5 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.5 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.0 | 0.7 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.0 | 0.5 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.0 | 0.5 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.0 | 1.0 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.0 | 7.1 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.0 | 0.5 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.0 | 0.6 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.0 | 0.3 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 1.2 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 52.0 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.9 | 25.4 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.7 | 9.3 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.7 | 9.1 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.6 | 16.2 | REACTOME NUCLEOTIDE LIKE PURINERGIC RECEPTORS | Genes involved in Nucleotide-like (purinergic) receptors |
| 0.6 | 30.3 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.6 | 21.4 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.5 | 1.6 | REACTOME SHC1 EVENTS IN EGFR SIGNALING | Genes involved in SHC1 events in EGFR signaling |
| 0.5 | 9.5 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.5 | 20.8 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.5 | 25.1 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.4 | 4.9 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.4 | 21.1 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.4 | 9.0 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.3 | 11.0 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.3 | 16.9 | REACTOME TRAFFICKING OF AMPA RECEPTORS | Genes involved in Trafficking of AMPA receptors |
| 0.3 | 65.6 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.3 | 1.3 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.3 | 7.5 | REACTOME CREB PHOSPHORYLATION THROUGH THE ACTIVATION OF CAMKII | Genes involved in CREB phosphorylation through the activation of CaMKII |
| 0.3 | 7.4 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.3 | 6.7 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.3 | 18.0 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.3 | 5.0 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.3 | 3.5 | REACTOME GLUTAMATE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Glutamate Neurotransmitter Release Cycle |
| 0.3 | 7.7 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.3 | 7.6 | REACTOME AKT PHOSPHORYLATES TARGETS IN THE CYTOSOL | Genes involved in AKT phosphorylates targets in the cytosol |
| 0.3 | 7.4 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.3 | 9.2 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.3 | 2.7 | REACTOME INTERACTIONS OF VPR WITH HOST CELLULAR PROTEINS | Genes involved in Interactions of Vpr with host cellular proteins |
| 0.3 | 4.7 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.3 | 7.8 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.3 | 11.1 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.2 | 4.1 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.2 | 2.3 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.2 | 5.6 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.2 | 6.9 | REACTOME ACTIVATION OF NMDA RECEPTOR UPON GLUTAMATE BINDING AND POSTSYNAPTIC EVENTS | Genes involved in Activation of NMDA receptor upon glutamate binding and postsynaptic events |
| 0.2 | 4.5 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.2 | 4.5 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.2 | 0.6 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 0.2 | 6.7 | REACTOME INSULIN RECEPTOR RECYCLING | Genes involved in Insulin receptor recycling |
| 0.2 | 0.8 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.2 | 7.0 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.2 | 11.4 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.2 | 0.4 | REACTOME ACETYLCHOLINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Acetylcholine Neurotransmitter Release Cycle |
| 0.2 | 2.9 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.2 | 5.8 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.2 | 3.0 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.2 | 1.0 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.2 | 4.7 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.2 | 22.8 | REACTOME NGF SIGNALLING VIA TRKA FROM THE PLASMA MEMBRANE | Genes involved in NGF signalling via TRKA from the plasma membrane |
| 0.2 | 0.5 | REACTOME THROMBIN SIGNALLING THROUGH PROTEINASE ACTIVATED RECEPTORS PARS | Genes involved in Thrombin signalling through proteinase activated receptors (PARs) |
| 0.2 | 5.3 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.2 | 2.2 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.2 | 6.2 | REACTOME TRIGLYCERIDE BIOSYNTHESIS | Genes involved in Triglyceride Biosynthesis |
| 0.2 | 5.5 | REACTOME DOWNREGULATION OF TGF BETA RECEPTOR SIGNALING | Genes involved in Downregulation of TGF-beta receptor signaling |
| 0.2 | 3.4 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.2 | 3.5 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.2 | 2.4 | REACTOME BOTULINUM NEUROTOXICITY | Genes involved in Botulinum neurotoxicity |
| 0.1 | 8.5 | REACTOME NUCLEAR SIGNALING BY ERBB4 | Genes involved in Nuclear signaling by ERBB4 |
| 0.1 | 3.5 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.1 | 1.4 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.1 | 4.5 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.1 | 3.8 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.1 | 1.5 | REACTOME POST TRANSLATIONAL MODIFICATION SYNTHESIS OF GPI ANCHORED PROTEINS | Genes involved in Post-translational modification: synthesis of GPI-anchored proteins |
| 0.1 | 3.1 | REACTOME ACYL CHAIN REMODELLING OF PE | Genes involved in Acyl chain remodelling of PE |
| 0.1 | 3.7 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.1 | 3.9 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.1 | 10.2 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.1 | 1.8 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.1 | 1.4 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.1 | 3.2 | REACTOME NITRIC OXIDE STIMULATES GUANYLATE CYCLASE | Genes involved in Nitric oxide stimulates guanylate cyclase |
| 0.1 | 10.1 | REACTOME G ALPHA Q SIGNALLING EVENTS | Genes involved in G alpha (q) signalling events |
| 0.1 | 4.7 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.1 | 3.7 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.1 | 1.5 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.1 | 4.0 | REACTOME POTASSIUM CHANNELS | Genes involved in Potassium Channels |
| 0.1 | 5.9 | REACTOME TRANSMISSION ACROSS CHEMICAL SYNAPSES | Genes involved in Transmission across Chemical Synapses |
| 0.1 | 6.8 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.1 | 1.5 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.1 | 3.3 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.1 | 4.7 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.1 | 1.7 | REACTOME SIGNALING BY FGFR3 MUTANTS | Genes involved in Signaling by FGFR3 mutants |
| 0.1 | 1.6 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.1 | 1.1 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.1 | 7.6 | REACTOME FATTY ACID TRIACYLGLYCEROL AND KETONE BODY METABOLISM | Genes involved in Fatty acid, triacylglycerol, and ketone body metabolism |
| 0.1 | 2.3 | REACTOME L1CAM INTERACTIONS | Genes involved in L1CAM interactions |
| 0.1 | 1.0 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.1 | 1.3 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.1 | 0.5 | REACTOME NFKB IS ACTIVATED AND SIGNALS SURVIVAL | Genes involved in NF-kB is activated and signals survival |
| 0.1 | 1.0 | REACTOME TRNA AMINOACYLATION | Genes involved in tRNA Aminoacylation |
| 0.1 | 2.8 | REACTOME TRANSPORT TO THE GOLGI AND SUBSEQUENT MODIFICATION | Genes involved in Transport to the Golgi and subsequent modification |
| 0.0 | 6.9 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.0 | 0.9 | REACTOME P75 NTR RECEPTOR MEDIATED SIGNALLING | Genes involved in p75 NTR receptor-mediated signalling |
| 0.0 | 0.8 | REACTOME KERATAN SULFATE BIOSYNTHESIS | Genes involved in Keratan sulfate biosynthesis |
| 0.0 | 0.3 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.0 | 1.5 | REACTOME SIGNALING BY EGFR IN CANCER | Genes involved in Signaling by EGFR in Cancer |
| 0.0 | 3.5 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.7 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.0 | 1.1 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.0 | 0.3 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.0 | 1.0 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 1.3 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.0 | 0.3 | REACTOME DOWNSTREAM SIGNALING EVENTS OF B CELL RECEPTOR BCR | Genes involved in Downstream Signaling Events Of B Cell Receptor (BCR) |
| 0.0 | 1.0 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF PIPS AT THE GOLGI MEMBRANE | Genes involved in Synthesis of PIPs at the Golgi membrane |
| 0.0 | 0.4 | REACTOME DOWNSTREAM SIGNAL TRANSDUCTION | Genes involved in Downstream signal transduction |
| 0.0 | 1.0 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.0 | 4.7 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.0 | 0.5 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.0 | 4.8 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.0 | 0.5 | REACTOME A TETRASACCHARIDE LINKER SEQUENCE IS REQUIRED FOR GAG SYNTHESIS | Genes involved in A tetrasaccharide linker sequence is required for GAG synthesis |
| 0.0 | 0.9 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.0 | 0.4 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 0.9 | REACTOME MUSCLE CONTRACTION | Genes involved in Muscle contraction |