Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
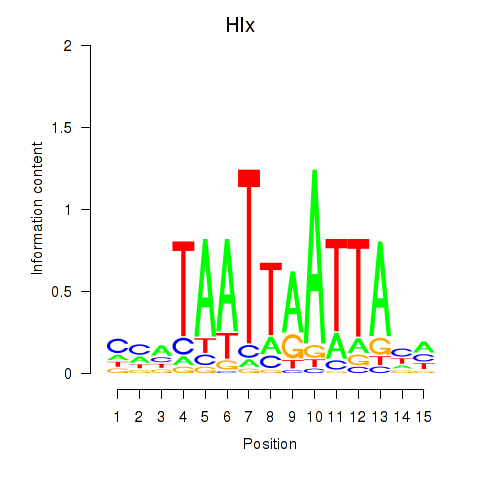
Results for Hlx
Z-value: 0.52
Transcription factors associated with Hlx
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Hlx
|
ENSMUSG00000039377.8 | Hlx |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Hlx | mm39_v1_chr1_-_184464810_184464824 | -0.28 | 1.9e-02 | Click! |
Activity profile of Hlx motif
Sorted Z-values of Hlx motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Hlx
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_+_7555053 | 7.01 |
ENSMUST00000090679.9
ENSMUST00000184986.2 |
Tac1
|
tachykinin 1 |
| chr15_-_8740218 | 5.26 |
ENSMUST00000005493.14
|
Slc1a3
|
solute carrier family 1 (glial high affinity glutamate transporter), member 3 |
| chr11_-_99134885 | 5.12 |
ENSMUST00000103132.10
ENSMUST00000038214.7 |
Krt222
|
keratin 222 |
| chr12_+_37930305 | 4.76 |
ENSMUST00000220990.2
|
Dgkb
|
diacylglycerol kinase, beta |
| chr15_-_8739893 | 4.55 |
ENSMUST00000157065.2
|
Slc1a3
|
solute carrier family 1 (glial high affinity glutamate transporter), member 3 |
| chr3_-_144738526 | 3.66 |
ENSMUST00000029919.7
|
Clca1
|
chloride channel accessory 1 |
| chr3_-_59127571 | 3.53 |
ENSMUST00000199675.2
ENSMUST00000170388.6 |
P2ry12
|
purinergic receptor P2Y, G-protein coupled 12 |
| chrX_-_142716200 | 2.82 |
ENSMUST00000112851.8
ENSMUST00000112856.3 ENSMUST00000033642.10 |
Dcx
|
doublecortin |
| chr12_+_37930169 | 2.69 |
ENSMUST00000221176.2
|
Dgkb
|
diacylglycerol kinase, beta |
| chr10_+_101994841 | 2.17 |
ENSMUST00000020039.13
|
Mgat4c
|
MGAT4 family, member C |
| chr1_-_53011187 | 2.13 |
ENSMUST00000186266.2
|
1700019D03Rik
|
RIKEN cDNA 1700019D03 gene |
| chr3_-_33136153 | 1.76 |
ENSMUST00000108225.10
|
Pex5l
|
peroxisomal biogenesis factor 5-like |
| chr1_+_179936757 | 1.73 |
ENSMUST00000143176.8
ENSMUST00000135056.8 |
Cdc42bpa
|
CDC42 binding protein kinase alpha |
| chr10_+_101994719 | 1.70 |
ENSMUST00000138522.8
ENSMUST00000163753.8 ENSMUST00000138016.8 |
Mgat4c
|
MGAT4 family, member C |
| chr3_+_32490525 | 0.55 |
ENSMUST00000108242.2
|
Pik3ca
|
phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha |
| chrX_-_142716085 | 0.55 |
ENSMUST00000087313.10
|
Dcx
|
doublecortin |
| chr1_-_74639723 | 0.44 |
ENSMUST00000127938.8
ENSMUST00000154874.8 |
Rnf25
|
ring finger protein 25 |
| chr2_-_88577230 | 0.11 |
ENSMUST00000099814.2
|
Olfr1198
|
olfactory receptor 1198 |
| chrX_+_132751729 | 0.03 |
ENSMUST00000033602.9
|
Tnmd
|
tenomodulin |
| chr1_+_106866678 | 0.02 |
ENSMUST00000112724.3
|
Serpinb12
|
serine (or cysteine) peptidase inhibitor, clade B (ovalbumin), member 12 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.0 | 9.8 | GO:0043490 | malate-aspartate shuttle(GO:0043490) |
| 1.8 | 7.0 | GO:2000852 | regulation of corticosterone secretion(GO:2000852) |
| 0.9 | 3.5 | GO:1904124 | microglial cell migration(GO:1904124) regulation of microglial cell migration(GO:1904139) |
| 0.2 | 1.8 | GO:0016560 | protein import into peroxisome matrix, docking(GO:0016560) |
| 0.2 | 3.4 | GO:0048672 | positive regulation of collateral sprouting(GO:0048672) |
| 0.2 | 7.5 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.2 | 0.5 | GO:0044028 | DNA hypomethylation(GO:0044028) hypomethylation of CpG island(GO:0044029) |
| 0.1 | 1.7 | GO:0040023 | nuclear migration(GO:0007097) establishment of nucleus localization(GO:0040023) |
| 0.0 | 3.7 | GO:1902476 | chloride transmembrane transport(GO:1902476) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.8 | GO:0017071 | intracellular cyclic nucleotide activated cation channel complex(GO:0017071) |
| 0.2 | 0.5 | GO:0005943 | phosphatidylinositol 3-kinase complex, class IA(GO:0005943) |
| 0.2 | 3.7 | GO:0042589 | zymogen granule membrane(GO:0042589) |
| 0.0 | 3.5 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.0 | 5.1 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 9.8 | GO:0043197 | dendritic spine(GO:0043197) |
| 0.0 | 10.4 | GO:0030424 | axon(GO:0030424) |
| 0.0 | 1.7 | GO:0042641 | actomyosin(GO:0042641) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 9.8 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 0.4 | 7.5 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.4 | 3.9 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.3 | 3.5 | GO:0001609 | G-protein coupled adenosine receptor activity(GO:0001609) |
| 0.3 | 1.8 | GO:0005052 | peroxisome matrix targeting signal-1 binding(GO:0005052) |
| 0.2 | 7.0 | GO:0071855 | neuropeptide receptor binding(GO:0071855) |
| 0.1 | 3.7 | GO:0005229 | intracellular calcium activated chloride channel activity(GO:0005229) |
| 0.1 | 2.1 | GO:0034237 | protein kinase A regulatory subunit binding(GO:0034237) |
| 0.0 | 0.5 | GO:0052813 | phosphatidylinositol-4,5-bisphosphate 3-kinase activity(GO:0046934) phosphatidylinositol bisphosphate kinase activity(GO:0052813) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.4 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 2.2 | PID CDC42 PATHWAY | CDC42 signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 3.5 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.2 | 7.5 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.1 | 3.9 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.1 | 9.8 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.0 | 7.0 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 3.4 | REACTOME L1CAM INTERACTIONS | Genes involved in L1CAM interactions |
| 0.0 | 0.5 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |