Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
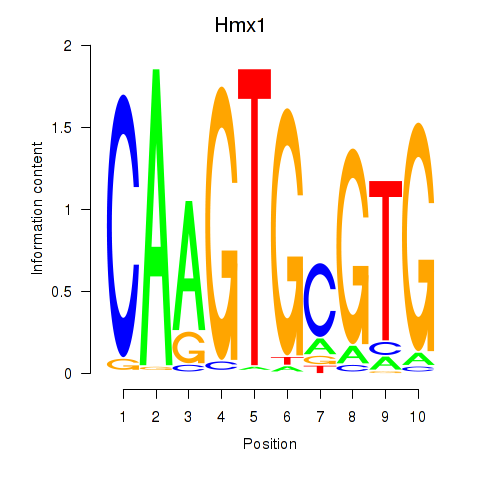
Results for Hmx1
Z-value: 1.22
Transcription factors associated with Hmx1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Hmx1
|
ENSMUSG00000067438.6 | Hmx1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Hmx1 | mm39_v1_chr5_+_35546363_35546461 | -0.41 | 3.3e-04 | Click! |
Activity profile of Hmx1 motif
Sorted Z-values of Hmx1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Hmx1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_135656885 | 8.91 |
ENSMUST00000027677.8
|
Csrp1
|
cysteine and glycine-rich protein 1 |
| chr7_+_18618605 | 7.64 |
ENSMUST00000032573.8
|
Pglyrp1
|
peptidoglycan recognition protein 1 |
| chr13_+_91609169 | 7.63 |
ENSMUST00000004094.15
ENSMUST00000042122.15 |
Ssbp2
|
single-stranded DNA binding protein 2 |
| chr13_+_91609264 | 7.46 |
ENSMUST00000231481.2
|
Ssbp2
|
single-stranded DNA binding protein 2 |
| chr11_+_11636213 | 7.31 |
ENSMUST00000076700.11
ENSMUST00000048122.13 |
Ikzf1
|
IKAROS family zinc finger 1 |
| chr9_-_21223551 | 7.30 |
ENSMUST00000003397.9
ENSMUST00000213250.2 |
Ap1m2
|
adaptor protein complex AP-1, mu 2 subunit |
| chr6_-_67512768 | 6.60 |
ENSMUST00000058178.6
|
Tacstd2
|
tumor-associated calcium signal transducer 2 |
| chrX_-_73290140 | 5.81 |
ENSMUST00000101454.9
ENSMUST00000033699.13 |
Flna
|
filamin, alpha |
| chr9_-_21223631 | 5.68 |
ENSMUST00000115433.11
|
Ap1m2
|
adaptor protein complex AP-1, mu 2 subunit |
| chr12_+_36042899 | 5.65 |
ENSMUST00000020898.12
|
Agr2
|
anterior gradient 2 |
| chr15_-_79718423 | 5.53 |
ENSMUST00000109623.8
ENSMUST00000109625.8 ENSMUST00000023060.13 ENSMUST00000089299.6 |
Cbx6
Npcd
|
chromobox 6 neuronal pentraxin chromo domain |
| chr3_-_92393193 | 5.40 |
ENSMUST00000054599.8
|
Sprr1a
|
small proline-rich protein 1A |
| chr3_-_88455556 | 5.09 |
ENSMUST00000131775.2
ENSMUST00000008745.13 |
Rab25
|
RAB25, member RAS oncogene family |
| chrX_-_73289970 | 5.00 |
ENSMUST00000130007.8
|
Flna
|
filamin, alpha |
| chrX_-_7537580 | 4.98 |
ENSMUST00000033486.6
|
Plp2
|
proteolipid protein 2 |
| chr2_+_164404499 | 4.62 |
ENSMUST00000017867.10
ENSMUST00000109344.9 ENSMUST00000109345.9 |
Wfdc2
|
WAP four-disulfide core domain 2 |
| chr11_+_69856222 | 4.58 |
ENSMUST00000018713.13
|
Cldn7
|
claudin 7 |
| chrX_+_49930311 | 4.49 |
ENSMUST00000114887.9
|
Stk26
|
serine/threonine kinase 26 |
| chr2_-_113588983 | 4.48 |
ENSMUST00000099575.4
|
Grem1
|
gremlin 1, DAN family BMP antagonist |
| chr12_+_51640097 | 4.43 |
ENSMUST00000164782.10
ENSMUST00000085412.7 |
Coch
|
cochlin |
| chr11_-_115258508 | 4.41 |
ENSMUST00000044152.13
ENSMUST00000106542.9 |
Hid1
|
HID1 domain containing |
| chr15_-_79718462 | 4.28 |
ENSMUST00000148358.2
|
Cbx6
|
chromobox 6 |
| chr6_+_41107047 | 4.08 |
ENSMUST00000103271.2
|
Trbv13-3
|
T cell receptor beta, variable 13-3 |
| chr17_+_35413415 | 4.04 |
ENSMUST00000025262.6
ENSMUST00000173600.2 |
Ltb
|
lymphotoxin B |
| chr3_+_108498595 | 3.97 |
ENSMUST00000051145.15
|
Wdr47
|
WD repeat domain 47 |
| chr4_+_134195631 | 3.93 |
ENSMUST00000030636.11
ENSMUST00000127279.8 ENSMUST00000105867.8 |
Stmn1
|
stathmin 1 |
| chr2_+_92745426 | 3.90 |
ENSMUST00000028648.3
|
Syt13
|
synaptotagmin XIII |
| chr17_-_34109513 | 3.88 |
ENSMUST00000173386.2
ENSMUST00000114361.9 ENSMUST00000173492.9 |
Kifc1
|
kinesin family member C1 |
| chr13_+_55517545 | 3.86 |
ENSMUST00000063771.14
|
Rgs14
|
regulator of G-protein signaling 14 |
| chr4_-_152561896 | 3.66 |
ENSMUST00000238738.2
ENSMUST00000162017.3 ENSMUST00000030768.10 |
Kcnab2
|
potassium voltage-gated channel, shaker-related subfamily, beta member 2 |
| chr2_+_165436929 | 3.61 |
ENSMUST00000088132.13
|
Eya2
|
EYA transcriptional coactivator and phosphatase 2 |
| chrX_-_133709733 | 3.53 |
ENSMUST00000035559.11
|
Armcx2
|
armadillo repeat containing, X-linked 2 |
| chr7_-_83384711 | 3.52 |
ENSMUST00000001792.12
|
Il16
|
interleukin 16 |
| chr6_-_127127959 | 3.51 |
ENSMUST00000201637.2
|
Ccnd2
|
cyclin D2 |
| chr14_-_79539063 | 3.49 |
ENSMUST00000022595.8
|
Rgcc
|
regulator of cell cycle |
| chr13_-_32522548 | 3.40 |
ENSMUST00000041859.9
|
Gmds
|
GDP-mannose 4, 6-dehydratase |
| chrX_-_165368675 | 3.36 |
ENSMUST00000000412.3
|
Egfl6
|
EGF-like-domain, multiple 6 |
| chr6_-_127127993 | 3.30 |
ENSMUST00000201066.2
|
Ccnd2
|
cyclin D2 |
| chr11_-_46203047 | 3.27 |
ENSMUST00000129474.2
ENSMUST00000093166.11 ENSMUST00000165599.9 |
Cyfip2
|
cytoplasmic FMR1 interacting protein 2 |
| chr2_+_129040677 | 3.26 |
ENSMUST00000028880.10
|
Slc20a1
|
solute carrier family 20, member 1 |
| chr9_-_110483210 | 3.25 |
ENSMUST00000196488.5
ENSMUST00000133191.8 ENSMUST00000167320.8 |
Nbeal2
|
neurobeachin-like 2 |
| chr7_-_125968653 | 3.14 |
ENSMUST00000205642.2
ENSMUST00000032997.8 ENSMUST00000206793.2 |
Lat
|
linker for activation of T cells |
| chr15_-_78456898 | 3.12 |
ENSMUST00000043214.8
|
Rac2
|
Rac family small GTPase 2 |
| chr6_-_87312743 | 3.05 |
ENSMUST00000042025.12
ENSMUST00000205033.2 |
Antxr1
|
anthrax toxin receptor 1 |
| chr2_+_152950388 | 2.98 |
ENSMUST00000189688.2
ENSMUST00000109799.8 ENSMUST00000003370.14 |
Hck
|
hemopoietic cell kinase |
| chr15_-_98507913 | 2.98 |
ENSMUST00000226500.2
ENSMUST00000227501.2 |
Adcy6
|
adenylate cyclase 6 |
| chr15_+_78312764 | 2.97 |
ENSMUST00000162517.8
ENSMUST00000166142.10 ENSMUST00000089414.11 |
Kctd17
|
potassium channel tetramerisation domain containing 17 |
| chr1_-_160134873 | 2.97 |
ENSMUST00000193185.6
|
Rabgap1l
|
RAB GTPase activating protein 1-like |
| chr1_+_152642291 | 2.95 |
ENSMUST00000077755.11
ENSMUST00000097536.6 |
Arpc5
|
actin related protein 2/3 complex, subunit 5 |
| chr17_+_34811217 | 2.94 |
ENSMUST00000038149.13
|
Pbx2
|
pre B cell leukemia homeobox 2 |
| chr5_-_34445662 | 2.92 |
ENSMUST00000094868.10
|
Zfyve28
|
zinc finger, FYVE domain containing 28 |
| chr11_+_23234644 | 2.90 |
ENSMUST00000150750.3
|
Xpo1
|
exportin 1 |
| chr15_+_54609011 | 2.90 |
ENSMUST00000050027.9
|
Ccn3
|
cellular communication network factor 3 |
| chr15_+_103411689 | 2.89 |
ENSMUST00000226493.2
|
Pde1b
|
phosphodiesterase 1B, Ca2+-calmodulin dependent |
| chr2_+_32766126 | 2.88 |
ENSMUST00000028135.15
|
Niban2
|
niban apoptosis regulator 2 |
| chr15_+_78312851 | 2.86 |
ENSMUST00000159771.8
|
Kctd17
|
potassium channel tetramerisation domain containing 17 |
| chr6_-_127128007 | 2.80 |
ENSMUST00000000188.12
|
Ccnd2
|
cyclin D2 |
| chr11_+_68322945 | 2.74 |
ENSMUST00000021283.8
|
Pik3r5
|
phosphoinositide-3-kinase regulatory subunit 5 |
| chr1_+_135252508 | 2.74 |
ENSMUST00000059352.3
|
Lmod1
|
leiomodin 1 (smooth muscle) |
| chr7_+_4467730 | 2.74 |
ENSMUST00000086372.8
ENSMUST00000169820.8 ENSMUST00000163893.8 ENSMUST00000170635.2 |
Eps8l1
|
EPS8-like 1 |
| chr11_+_44409775 | 2.62 |
ENSMUST00000019333.10
|
Rnf145
|
ring finger protein 145 |
| chr6_-_124710084 | 2.62 |
ENSMUST00000112484.10
|
Ptpn6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr1_+_61017057 | 2.59 |
ENSMUST00000027162.12
ENSMUST00000102827.4 |
Icos
|
inducible T cell co-stimulator |
| chr6_-_87312681 | 2.59 |
ENSMUST00000204805.3
|
Antxr1
|
anthrax toxin receptor 1 |
| chr2_+_162916551 | 2.58 |
ENSMUST00000142729.3
|
Mybl2
|
myeloblastosis oncogene-like 2 |
| chr9_-_42035560 | 2.55 |
ENSMUST00000060989.9
|
Sorl1
|
sortilin-related receptor, LDLR class A repeats-containing |
| chr11_-_115258493 | 2.53 |
ENSMUST00000123428.2
|
Hid1
|
HID1 domain containing |
| chr19_-_14575395 | 2.50 |
ENSMUST00000052011.15
ENSMUST00000167776.3 |
Tle4
|
transducin-like enhancer of split 4 |
| chr8_+_121264161 | 2.48 |
ENSMUST00000118136.2
|
Gse1
|
genetic suppressor element 1, coiled-coil protein |
| chr2_-_73284262 | 2.45 |
ENSMUST00000102679.8
|
Wipf1
|
WAS/WASL interacting protein family, member 1 |
| chr5_-_39801940 | 2.44 |
ENSMUST00000152057.2
ENSMUST00000053116.7 |
Hs3st1
|
heparan sulfate (glucosamine) 3-O-sulfotransferase 1 |
| chr18_-_62313019 | 2.42 |
ENSMUST00000053640.5
|
Adrb2
|
adrenergic receptor, beta 2 |
| chrX_-_151110425 | 2.41 |
ENSMUST00000195280.3
|
Kantr
|
Kdm5c adjacent non-coding transcript |
| chr1_-_181669891 | 2.38 |
ENSMUST00000193028.2
ENSMUST00000191878.6 ENSMUST00000005003.12 |
Lbr
|
lamin B receptor |
| chr5_-_134643805 | 2.34 |
ENSMUST00000202085.2
ENSMUST00000036362.13 ENSMUST00000077636.8 |
Lat2
|
linker for activation of T cells family, member 2 |
| chr5_+_121358254 | 2.30 |
ENSMUST00000042614.13
|
Hectd4
|
HECT domain E3 ubiquitin protein ligase 4 |
| chr9_+_95836797 | 2.26 |
ENSMUST00000034981.14
ENSMUST00000185633.7 |
Xrn1
|
5'-3' exoribonuclease 1 |
| chr12_-_56392646 | 2.26 |
ENSMUST00000021416.9
|
Mbip
|
MAP3K12 binding inhibitory protein 1 |
| chr8_+_121262528 | 2.23 |
ENSMUST00000120493.8
|
Gse1
|
genetic suppressor element 1, coiled-coil protein |
| chr17_+_27136065 | 2.21 |
ENSMUST00000078961.6
|
Kifc5b
|
kinesin family member C5B |
| chr17_+_48769383 | 2.21 |
ENSMUST00000162132.8
|
Unc5cl
|
unc-5 family C-terminal like |
| chr6_-_124710030 | 2.17 |
ENSMUST00000173647.2
|
Ptpn6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr11_+_120612278 | 2.16 |
ENSMUST00000018156.12
|
Rac3
|
Rac family small GTPase 3 |
| chr2_+_130116344 | 2.15 |
ENSMUST00000103198.11
|
Nop56
|
NOP56 ribonucleoprotein |
| chr3_-_89325594 | 2.12 |
ENSMUST00000029679.4
|
Cks1b
|
CDC28 protein kinase 1b |
| chr3_-_107992662 | 2.10 |
ENSMUST00000078912.7
|
Ampd2
|
adenosine monophosphate deaminase 2 |
| chr9_+_95836869 | 2.09 |
ENSMUST00000190665.2
|
Xrn1
|
5'-3' exoribonuclease 1 |
| chr11_+_120612369 | 2.07 |
ENSMUST00000142229.2
|
Rac3
|
Rac family small GTPase 3 |
| chr14_-_56339915 | 2.07 |
ENSMUST00000015583.2
|
Ctsg
|
cathepsin G |
| chr15_+_78294154 | 2.07 |
ENSMUST00000229739.2
|
Mpst
|
mercaptopyruvate sulfurtransferase |
| chr9_+_32607301 | 2.06 |
ENSMUST00000034534.13
ENSMUST00000050797.14 ENSMUST00000184887.2 |
Ets1
|
E26 avian leukemia oncogene 1, 5' domain |
| chr15_-_85462271 | 2.05 |
ENSMUST00000229191.2
|
Wnt7b
|
wingless-type MMTV integration site family, member 7B |
| chr6_+_124973752 | 2.05 |
ENSMUST00000162000.4
|
Pianp
|
PILR alpha associated neural protein |
| chr18_+_38088597 | 2.04 |
ENSMUST00000070709.9
ENSMUST00000177058.8 ENSMUST00000169360.9 ENSMUST00000163591.9 ENSMUST00000091932.12 |
Rell2
|
RELT-like 2 |
| chr9_+_108437485 | 2.02 |
ENSMUST00000081111.14
ENSMUST00000193421.2 |
Impdh2
|
inosine monophosphate dehydrogenase 2 |
| chr6_-_40413056 | 2.01 |
ENSMUST00000039008.10
ENSMUST00000101492.10 |
Dennd11
|
DENN domain containing 11 |
| chr5_-_143513732 | 2.00 |
ENSMUST00000100489.4
ENSMUST00000080537.14 |
Rac1
|
Rac family small GTPase 1 |
| chr3_+_130906434 | 1.99 |
ENSMUST00000098611.4
|
Lef1
|
lymphoid enhancer binding factor 1 |
| chr6_-_91093766 | 1.98 |
ENSMUST00000113509.2
ENSMUST00000032179.14 |
Nup210
|
nucleoporin 210 |
| chr1_+_43769750 | 1.96 |
ENSMUST00000027217.9
|
Ecrg4
|
ECRG4 augurin precursor |
| chr3_+_60408678 | 1.96 |
ENSMUST00000191747.6
ENSMUST00000194069.6 |
Mbnl1
|
muscleblind like splicing factor 1 |
| chr5_-_144160397 | 1.93 |
ENSMUST00000085701.7
|
Tecpr1
|
tectonin beta-propeller repeat containing 1 |
| chr6_+_124973644 | 1.92 |
ENSMUST00000032479.11
|
Pianp
|
PILR alpha associated neural protein |
| chr9_-_31824758 | 1.91 |
ENSMUST00000116615.5
|
Barx2
|
BarH-like homeobox 2 |
| chrX_-_73416869 | 1.87 |
ENSMUST00000073067.11
ENSMUST00000037967.6 |
Slc10a3
|
solute carrier family 10 (sodium/bile acid cotransporter family), member 3 |
| chr16_+_49519561 | 1.86 |
ENSMUST00000046777.11
ENSMUST00000142682.9 |
Ift57
|
intraflagellar transport 57 |
| chr2_+_130116357 | 1.86 |
ENSMUST00000136621.9
ENSMUST00000141872.2 |
Nop56
|
NOP56 ribonucleoprotein |
| chr2_-_27032441 | 1.85 |
ENSMUST00000151224.3
|
Fam163b
|
family with sequence similarity 163, member B |
| chr14_+_56813060 | 1.85 |
ENSMUST00000161553.2
|
Parp4
|
poly (ADP-ribose) polymerase family, member 4 |
| chr11_-_69749549 | 1.83 |
ENSMUST00000001626.10
ENSMUST00000108626.8 |
Tnk1
|
tyrosine kinase, non-receptor, 1 |
| chr19_-_4665509 | 1.83 |
ENSMUST00000053597.3
|
Lrfn4
|
leucine rich repeat and fibronectin type III domain containing 4 |
| chr11_+_62711057 | 1.82 |
ENSMUST00000055006.12
ENSMUST00000072639.10 |
Trim16
|
tripartite motif-containing 16 |
| chr9_-_49479791 | 1.80 |
ENSMUST00000194252.6
|
Ncam1
|
neural cell adhesion molecule 1 |
| chr2_+_3705824 | 1.80 |
ENSMUST00000115054.9
|
Fam107b
|
family with sequence similarity 107, member B |
| chr12_-_46767619 | 1.78 |
ENSMUST00000219886.2
|
Nova1
|
NOVA alternative splicing regulator 1 |
| chr14_-_56472102 | 1.75 |
ENSMUST00000015585.4
|
Gzmc
|
granzyme C |
| chr11_-_102210568 | 1.74 |
ENSMUST00000173870.8
|
Ubtf
|
upstream binding transcription factor, RNA polymerase I |
| chr18_+_68066328 | 1.68 |
ENSMUST00000063775.5
|
Ldlrad4
|
low density lipoprotein receptor class A domain containing 4 |
| chr11_-_61384998 | 1.67 |
ENSMUST00000101085.9
ENSMUST00000079080.13 ENSMUST00000108714.2 |
Mapk7
|
mitogen-activated protein kinase 7 |
| chr4_+_156098292 | 1.67 |
ENSMUST00000030952.6
|
Tnfrsf4
|
tumor necrosis factor receptor superfamily, member 4 |
| chr2_+_25162487 | 1.66 |
ENSMUST00000028341.11
|
Anapc2
|
anaphase promoting complex subunit 2 |
| chr4_-_94444975 | 1.61 |
ENSMUST00000030313.9
|
Caap1
|
caspase activity and apoptosis inhibitor 1 |
| chr11_+_62711295 | 1.59 |
ENSMUST00000108703.2
|
Trim16
|
tripartite motif-containing 16 |
| chr14_-_56499690 | 1.58 |
ENSMUST00000015581.6
|
Gzmb
|
granzyme B |
| chr19_-_4665668 | 1.58 |
ENSMUST00000113822.3
|
Lrfn4
|
leucine rich repeat and fibronectin type III domain containing 4 |
| chrX_+_158038778 | 1.58 |
ENSMUST00000126686.8
ENSMUST00000033671.13 |
Rps6ka3
|
ribosomal protein S6 kinase polypeptide 3 |
| chr3_+_108191398 | 1.58 |
ENSMUST00000135636.6
ENSMUST00000102632.7 |
Sort1
|
sortilin 1 |
| chrX_-_73436293 | 1.57 |
ENSMUST00000114138.8
|
Fam3a
|
family with sequence similarity 3, member A |
| chr18_+_75953244 | 1.55 |
ENSMUST00000058997.15
|
Zbtb7c
|
zinc finger and BTB domain containing 7C |
| chr18_-_24736848 | 1.55 |
ENSMUST00000070726.10
|
Slc39a6
|
solute carrier family 39 (metal ion transporter), member 6 |
| chr13_-_55477535 | 1.54 |
ENSMUST00000021941.8
|
Mxd3
|
Max dimerization protein 3 |
| chr14_-_56812839 | 1.53 |
ENSMUST00000225951.2
|
Cenpj
|
centromere protein J |
| chr2_-_25162347 | 1.49 |
ENSMUST00000028342.7
|
Ssna1
|
SS nuclear autoantigen 1 |
| chr12_-_34956910 | 1.48 |
ENSMUST00000239321.2
|
Hdac9
|
histone deacetylase 9 |
| chr2_+_28082943 | 1.46 |
ENSMUST00000113920.8
|
Olfm1
|
olfactomedin 1 |
| chr17_+_87270707 | 1.44 |
ENSMUST00000139344.2
|
Rhoq
|
ras homolog family member Q |
| chr11_-_119438569 | 1.43 |
ENSMUST00000026670.5
|
Nptx1
|
neuronal pentraxin 1 |
| chr16_-_97412169 | 1.43 |
ENSMUST00000232141.2
ENSMUST00000000395.8 |
Tmprss2
|
transmembrane protease, serine 2 |
| chr17_-_57023788 | 1.43 |
ENSMUST00000067931.7
|
Vmac
|
vimentin-type intermediate filament associated coiled-coil protein |
| chr1_+_165957784 | 1.43 |
ENSMUST00000060833.14
|
Gpa33
|
glycoprotein A33 (transmembrane) |
| chrX_+_158039107 | 1.42 |
ENSMUST00000148570.8
|
Rps6ka3
|
ribosomal protein S6 kinase polypeptide 3 |
| chr15_-_53765869 | 1.41 |
ENSMUST00000078673.14
|
Samd12
|
sterile alpha motif domain containing 12 |
| chrX_+_158038915 | 1.40 |
ENSMUST00000112492.8
|
Rps6ka3
|
ribosomal protein S6 kinase polypeptide 3 |
| chr2_+_28083105 | 1.39 |
ENSMUST00000100244.10
|
Olfm1
|
olfactomedin 1 |
| chr4_+_129407374 | 1.39 |
ENSMUST00000062356.7
|
Marcksl1
|
MARCKS-like 1 |
| chr12_+_24622274 | 1.38 |
ENSMUST00000085553.13
|
Grhl1
|
grainyhead like transcription factor 1 |
| chr10_-_128505096 | 1.36 |
ENSMUST00000238610.2
ENSMUST00000238712.2 |
Ikzf4
|
IKAROS family zinc finger 4 |
| chr10_-_85847697 | 1.35 |
ENSMUST00000105304.2
ENSMUST00000061699.12 |
Bpifc
|
BPI fold containing family C |
| chr13_+_45660905 | 1.35 |
ENSMUST00000000260.13
|
Gmpr
|
guanosine monophosphate reductase |
| chr15_+_32244947 | 1.34 |
ENSMUST00000067458.7
|
Sema5a
|
sema domain, seven thrombospondin repeats (type 1 and type 1-like), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 5A |
| chr4_+_116078874 | 1.34 |
ENSMUST00000106490.3
|
Pik3r3
|
phosphoinositide-3-kinase regulatory subunit 3 |
| chr8_+_84627332 | 1.34 |
ENSMUST00000045393.15
ENSMUST00000132500.8 ENSMUST00000152978.8 |
Adgrl1
|
adhesion G protein-coupled receptor L1 |
| chr5_+_147367237 | 1.33 |
ENSMUST00000176600.8
|
Pan3
|
PAN3 poly(A) specific ribonuclease subunit |
| chr1_-_127605660 | 1.33 |
ENSMUST00000160616.8
|
Tmem163
|
transmembrane protein 163 |
| chr11_-_97520511 | 1.32 |
ENSMUST00000052281.6
|
Epop
|
elongin BC and polycomb repressive complex 2 associated protein |
| chr15_+_18819033 | 1.30 |
ENSMUST00000166873.9
|
Cdh10
|
cadherin 10 |
| chr17_-_37178079 | 1.29 |
ENSMUST00000025329.13
ENSMUST00000174195.8 |
Trim15
|
tripartite motif-containing 15 |
| chr16_+_10652910 | 1.28 |
ENSMUST00000037913.9
|
Rmi2
|
RecQ mediated genome instability 2 |
| chr17_+_8559230 | 1.28 |
ENSMUST00000069742.8
|
Prr18
|
proline rich 18 |
| chr9_+_95836839 | 1.27 |
ENSMUST00000189106.2
|
Xrn1
|
5'-3' exoribonuclease 1 |
| chr9_+_98868418 | 1.25 |
ENSMUST00000035038.8
ENSMUST00000112911.9 |
Faim
|
Fas apoptotic inhibitory molecule |
| chr8_-_126625029 | 1.24 |
ENSMUST00000047239.13
ENSMUST00000131127.3 |
Pcnx2
|
pecanex homolog 2 |
| chr15_+_79025523 | 1.23 |
ENSMUST00000040077.8
|
Polr2f
|
polymerase (RNA) II (DNA directed) polypeptide F |
| chr7_+_48438751 | 1.22 |
ENSMUST00000118927.8
ENSMUST00000125280.8 |
Zdhhc13
|
zinc finger, DHHC domain containing 13 |
| chr2_-_92201342 | 1.21 |
ENSMUST00000176810.8
ENSMUST00000090582.11 ENSMUST00000068586.13 |
Large2
|
LARGE xylosyl- and glucuronyltransferase 2 |
| chr19_-_6285827 | 1.21 |
ENSMUST00000025695.10
|
Ppp2r5b
|
protein phosphatase 2, regulatory subunit B', beta |
| chr14_-_78545472 | 1.20 |
ENSMUST00000022592.8
|
Tnfsf11
|
tumor necrosis factor (ligand) superfamily, member 11 |
| chr18_+_38088179 | 1.19 |
ENSMUST00000176902.8
ENSMUST00000176104.8 |
Rell2
|
RELT-like 2 |
| chr8_+_106024294 | 1.18 |
ENSMUST00000015003.10
|
E2f4
|
E2F transcription factor 4 |
| chr1_+_130793406 | 1.18 |
ENSMUST00000038829.7
|
Fcmr
|
Fc fragment of IgM receptor |
| chr6_+_34453142 | 1.18 |
ENSMUST00000045372.6
ENSMUST00000138668.2 ENSMUST00000139067.2 |
Bpgm
|
2,3-bisphosphoglycerate mutase |
| chr7_-_117842787 | 1.17 |
ENSMUST00000032891.15
|
Smg1
|
SMG1 homolog, phosphatidylinositol 3-kinase-related kinase (C. elegans) |
| chr2_+_118756973 | 1.16 |
ENSMUST00000099546.6
ENSMUST00000110837.2 |
Chst14
|
carbohydrate sulfotransferase 14 |
| chr11_-_94133527 | 1.16 |
ENSMUST00000061469.4
|
Wfikkn2
|
WAP, follistatin/kazal, immunoglobulin, kunitz and netrin domain containing 2 |
| chr1_-_15962881 | 1.15 |
ENSMUST00000040695.5
|
Sbspon
|
somatomedin B and thrombospondin, type 1 domain containing |
| chr1_+_165958036 | 1.14 |
ENSMUST00000166860.2
|
Gpa33
|
glycoprotein A33 (transmembrane) |
| chr17_-_18075450 | 1.14 |
ENSMUST00000003762.8
|
Has1
|
hyaluronan synthase 1 |
| chr5_-_38719025 | 1.13 |
ENSMUST00000005234.13
|
Wdr1
|
WD repeat domain 1 |
| chr11_-_120515799 | 1.13 |
ENSMUST00000106183.3
ENSMUST00000080202.12 |
Sirt7
|
sirtuin 7 |
| chrX_+_9138995 | 1.12 |
ENSMUST00000015486.7
|
Xk
|
X-linked Kx blood group |
| chr14_-_52481907 | 1.10 |
ENSMUST00000226307.2
|
Chd8
|
chromodomain helicase DNA binding protein 8 |
| chr8_+_94879235 | 1.10 |
ENSMUST00000034211.10
ENSMUST00000211930.2 ENSMUST00000211915.2 |
Mt3
|
metallothionein 3 |
| chr5_-_74863514 | 1.09 |
ENSMUST00000117388.8
|
Lnx1
|
ligand of numb-protein X 1 |
| chr15_+_102412157 | 1.09 |
ENSMUST00000096145.5
|
Gm10337
|
predicted gene 10337 |
| chr11_-_102209767 | 1.07 |
ENSMUST00000174302.8
ENSMUST00000178839.8 ENSMUST00000006754.14 |
Ubtf
|
upstream binding transcription factor, RNA polymerase I |
| chr2_-_34716083 | 1.07 |
ENSMUST00000113077.8
ENSMUST00000028220.10 |
Fbxw2
|
F-box and WD-40 domain protein 2 |
| chr4_+_116078830 | 1.06 |
ENSMUST00000030464.14
|
Pik3r3
|
phosphoinositide-3-kinase regulatory subunit 3 |
| chr2_-_156833932 | 1.06 |
ENSMUST00000109558.2
ENSMUST00000069600.13 ENSMUST00000072298.13 |
Ndrg3
|
N-myc downstream regulated gene 3 |
| chrX_-_73436609 | 1.04 |
ENSMUST00000015427.13
|
Fam3a
|
family with sequence similarity 3, member A |
| chr11_-_115024807 | 1.01 |
ENSMUST00000106561.8
ENSMUST00000051264.14 ENSMUST00000106562.3 |
Cd300lf
|
CD300 molecule like family member F |
| chrX_-_73416824 | 1.01 |
ENSMUST00000178691.2
ENSMUST00000114146.8 |
Ubl4a
Slc10a3
|
ubiquitin-like 4A solute carrier family 10 (sodium/bile acid cotransporter family), member 3 |
| chr5_+_102992873 | 0.99 |
ENSMUST00000070000.6
|
Arhgap24
|
Rho GTPase activating protein 24 |
| chr11_-_48836975 | 0.99 |
ENSMUST00000104958.2
|
Psme2b
|
protease (prosome, macropain) activator subunit 2B |
| chr9_-_107474221 | 0.98 |
ENSMUST00000238519.2
|
Lsmem2
|
leucine-rich single-pass membrane protein 2 |
| chr18_+_34353943 | 0.98 |
ENSMUST00000171187.8
|
Apc
|
APC, WNT signaling pathway regulator |
| chr7_-_119319965 | 0.97 |
ENSMUST00000033236.9
|
Thumpd1
|
THUMP domain containing 1 |
| chr4_+_129878890 | 0.95 |
ENSMUST00000106017.8
ENSMUST00000121049.8 |
Adgrb2
|
adhesion G protein-coupled receptor B2 |
| chr7_-_117842892 | 0.95 |
ENSMUST00000179047.3
|
Smg1
|
SMG1 homolog, phosphatidylinositol 3-kinase-related kinase (C. elegans) |
| chr5_+_31107390 | 0.92 |
ENSMUST00000006814.9
|
Abhd1
|
abhydrolase domain containing 1 |
| chr4_+_62537750 | 0.92 |
ENSMUST00000084521.11
ENSMUST00000107424.2 |
Rgs3
|
regulator of G-protein signaling 3 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.6 | 10.8 | GO:1905000 | regulation of membrane repolarization during atrial cardiac muscle cell action potential(GO:1905000) |
| 2.5 | 7.6 | GO:0051714 | regulation of cytolysis in other organism(GO:0051710) positive regulation of cytolysis in other organism(GO:0051714) |
| 2.4 | 7.3 | GO:0045660 | positive regulation of neutrophil differentiation(GO:0045660) |
| 2.2 | 11.1 | GO:0090191 | negative regulation of branching involved in ureteric bud morphogenesis(GO:0090191) |
| 1.9 | 5.7 | GO:1903896 | positive regulation of IRE1-mediated unfolded protein response(GO:1903896) |
| 1.2 | 3.5 | GO:0003330 | regulation of extracellular matrix constituent secretion(GO:0003330) positive regulation of extracellular matrix constituent secretion(GO:0003331) |
| 1.0 | 3.0 | GO:1904117 | response to vasopressin(GO:1904116) cellular response to vasopressin(GO:1904117) |
| 1.0 | 3.9 | GO:0072385 | minus-end-directed organelle transport along microtubule(GO:0072385) |
| 0.9 | 5.6 | GO:1903054 | negative regulation of extracellular matrix organization(GO:1903054) |
| 0.7 | 3.5 | GO:1905034 | regulation of antifungal innate immune response(GO:1905034) negative regulation of antifungal innate immune response(GO:1905035) |
| 0.7 | 3.3 | GO:0008626 | granzyme-mediated apoptotic signaling pathway(GO:0008626) |
| 0.6 | 2.5 | GO:2001137 | protein retention in Golgi apparatus(GO:0045053) positive regulation of endocytic recycling(GO:2001137) |
| 0.6 | 2.4 | GO:0002025 | vasodilation by norepinephrine-epinephrine involved in regulation of systemic arterial blood pressure(GO:0002025) |
| 0.6 | 9.6 | GO:0071481 | cellular response to X-ray(GO:0071481) |
| 0.6 | 4.8 | GO:0033277 | abortive mitotic cell cycle(GO:0033277) |
| 0.6 | 6.2 | GO:0021894 | cerebral cortex GABAergic interneuron development(GO:0021894) |
| 0.6 | 3.9 | GO:0070495 | regulation of thrombin receptor signaling pathway(GO:0070494) negative regulation of thrombin receptor signaling pathway(GO:0070495) |
| 0.6 | 1.7 | GO:0070376 | regulation of ERK5 cascade(GO:0070376) |
| 0.5 | 1.1 | GO:0097212 | lysosomal membrane organization(GO:0097212) |
| 0.5 | 2.1 | GO:0016332 | establishment or maintenance of polarity of embryonic epithelium(GO:0016332) |
| 0.5 | 1.0 | GO:0035666 | TRIF-dependent toll-like receptor signaling pathway(GO:0035666) |
| 0.5 | 2.0 | GO:0061153 | negative regulation of interleukin-13 production(GO:0032696) trachea submucosa development(GO:0061152) trachea gland development(GO:0061153) odontoblast differentiation(GO:0071895) |
| 0.5 | 5.6 | GO:0090503 | histone mRNA catabolic process(GO:0071044) RNA phosphodiester bond hydrolysis, exonucleolytic(GO:0090503) |
| 0.5 | 3.2 | GO:0070889 | platelet alpha granule organization(GO:0070889) |
| 0.5 | 1.8 | GO:0001928 | regulation of exocyst assembly(GO:0001928) |
| 0.4 | 4.0 | GO:0045084 | positive regulation of interleukin-12 biosynthetic process(GO:0045084) |
| 0.4 | 4.5 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 0.4 | 1.2 | GO:0030208 | dermatan sulfate biosynthetic process(GO:0030208) |
| 0.4 | 1.1 | GO:0045226 | extracellular polysaccharide biosynthetic process(GO:0045226) extracellular polysaccharide metabolic process(GO:0046379) |
| 0.4 | 3.4 | GO:0019673 | GDP-mannose metabolic process(GO:0019673) |
| 0.4 | 1.5 | GO:0060830 | ciliary receptor clustering involved in smoothened signaling pathway(GO:0060830) |
| 0.3 | 6.9 | GO:0031268 | pseudopodium organization(GO:0031268) |
| 0.3 | 1.2 | GO:0051466 | positive regulation of corticotropin-releasing hormone secretion(GO:0051466) |
| 0.3 | 1.8 | GO:0061511 | centriole elongation(GO:0061511) |
| 0.3 | 2.9 | GO:0032264 | IMP salvage(GO:0032264) |
| 0.3 | 2.9 | GO:0000056 | ribosomal small subunit export from nucleus(GO:0000056) |
| 0.3 | 2.0 | GO:0006177 | GMP biosynthetic process(GO:0006177) |
| 0.3 | 4.6 | GO:0032463 | negative regulation of protein homooligomerization(GO:0032463) |
| 0.3 | 1.1 | GO:0045897 | regulation of transcription during mitosis(GO:0045896) positive regulation of transcription during mitosis(GO:0045897) |
| 0.3 | 2.8 | GO:0006362 | transcription elongation from RNA polymerase I promoter(GO:0006362) |
| 0.3 | 1.7 | GO:0010991 | negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.3 | 3.9 | GO:0031914 | negative regulation of synaptic plasticity(GO:0031914) |
| 0.3 | 1.3 | GO:0035616 | histone H2B conserved C-terminal lysine deubiquitination(GO:0035616) |
| 0.3 | 1.6 | GO:0051005 | negative regulation of lipoprotein lipase activity(GO:0051005) |
| 0.3 | 3.4 | GO:0048386 | positive regulation of retinoic acid receptor signaling pathway(GO:0048386) |
| 0.3 | 1.3 | GO:1901252 | regulation of intracellular transport of viral material(GO:1901252) |
| 0.3 | 2.1 | GO:0070944 | neutrophil mediated killing of bacterium(GO:0070944) |
| 0.3 | 2.3 | GO:0040032 | post-embryonic body morphogenesis(GO:0040032) |
| 0.3 | 4.8 | GO:0006968 | cellular defense response(GO:0006968) |
| 0.3 | 2.3 | GO:0043545 | molybdopterin cofactor biosynthetic process(GO:0032324) molybdopterin cofactor metabolic process(GO:0043545) prosthetic group metabolic process(GO:0051189) |
| 0.2 | 0.7 | GO:0007352 | zygotic specification of dorsal/ventral axis(GO:0007352) |
| 0.2 | 3.6 | GO:0016576 | histone dephosphorylation(GO:0016576) |
| 0.2 | 2.9 | GO:0023041 | neuronal signal transduction(GO:0023041) |
| 0.2 | 1.3 | GO:0048842 | positive regulation of axon extension involved in axon guidance(GO:0048842) |
| 0.2 | 1.1 | GO:0031133 | regulation of axon diameter(GO:0031133) |
| 0.2 | 2.0 | GO:0060753 | regulation of mast cell chemotaxis(GO:0060753) |
| 0.2 | 2.7 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.2 | 2.1 | GO:0009092 | homoserine metabolic process(GO:0009092) transsulfuration(GO:0019346) hydrogen sulfide biosynthetic process(GO:0070814) |
| 0.2 | 2.9 | GO:0036005 | response to macrophage colony-stimulating factor(GO:0036005) cellular response to macrophage colony-stimulating factor stimulus(GO:0036006) |
| 0.2 | 1.4 | GO:0035865 | response to potassium ion(GO:0035864) cellular response to potassium ion(GO:0035865) axonogenesis involved in innervation(GO:0060385) |
| 0.2 | 1.0 | GO:1902949 | positive regulation of tau-protein kinase activity(GO:1902949) |
| 0.2 | 3.7 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 0.2 | 1.1 | GO:0030043 | actin filament fragmentation(GO:0030043) |
| 0.2 | 2.1 | GO:0030578 | PML body organization(GO:0030578) |
| 0.2 | 0.6 | GO:0072720 | response to dithiothreitol(GO:0072720) |
| 0.2 | 2.7 | GO:1900029 | positive regulation of ruffle assembly(GO:1900029) |
| 0.2 | 6.6 | GO:0045724 | positive regulation of cilium assembly(GO:0045724) |
| 0.2 | 1.6 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 0.2 | 3.8 | GO:0030011 | maintenance of cell polarity(GO:0030011) |
| 0.2 | 1.2 | GO:0030263 | apoptotic chromosome condensation(GO:0030263) |
| 0.1 | 0.9 | GO:0034184 | positive regulation of maintenance of mitotic sister chromatid cohesion(GO:0034184) |
| 0.1 | 5.4 | GO:0031424 | keratinization(GO:0031424) |
| 0.1 | 1.4 | GO:0046598 | positive regulation of viral entry into host cell(GO:0046598) |
| 0.1 | 3.0 | GO:2000251 | positive regulation of actin cytoskeleton reorganization(GO:2000251) |
| 0.1 | 2.1 | GO:0050930 | induction of positive chemotaxis(GO:0050930) |
| 0.1 | 4.5 | GO:0030033 | microvillus assembly(GO:0030033) |
| 0.1 | 1.7 | GO:0031915 | positive regulation of synaptic plasticity(GO:0031915) |
| 0.1 | 3.0 | GO:0035855 | megakaryocyte development(GO:0035855) |
| 0.1 | 0.9 | GO:1904426 | positive regulation of GTP binding(GO:1904426) |
| 0.1 | 1.1 | GO:2000270 | negative regulation of fibroblast apoptotic process(GO:2000270) |
| 0.1 | 0.4 | GO:0010512 | negative regulation of phosphatidylinositol biosynthetic process(GO:0010512) |
| 0.1 | 1.3 | GO:0000290 | deadenylation-dependent decapping of nuclear-transcribed mRNA(GO:0000290) |
| 0.1 | 5.5 | GO:0043551 | regulation of phosphatidylinositol 3-kinase activity(GO:0043551) |
| 0.1 | 1.5 | GO:0034983 | peptidyl-lysine deacetylation(GO:0034983) |
| 0.1 | 1.0 | GO:0071816 | tail-anchored membrane protein insertion into ER membrane(GO:0071816) |
| 0.1 | 6.9 | GO:0046677 | response to antibiotic(GO:0046677) |
| 0.1 | 1.5 | GO:1903025 | regulation of RNA polymerase II regulatory region sequence-specific DNA binding(GO:1903025) |
| 0.1 | 2.6 | GO:0002517 | T cell tolerance induction(GO:0002517) |
| 0.1 | 2.2 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.1 | 8.9 | GO:0070527 | platelet aggregation(GO:0070527) |
| 0.1 | 4.3 | GO:0001562 | response to protozoan(GO:0001562) |
| 0.1 | 1.4 | GO:0002934 | desmosome organization(GO:0002934) |
| 0.1 | 3.3 | GO:0006817 | phosphate ion transport(GO:0006817) |
| 0.1 | 4.0 | GO:0000154 | rRNA modification(GO:0000154) |
| 0.1 | 0.4 | GO:0071374 | cellular response to parathyroid hormone stimulus(GO:0071374) |
| 0.1 | 2.4 | GO:0046827 | positive regulation of protein export from nucleus(GO:0046827) |
| 0.1 | 1.5 | GO:0071578 | zinc II ion transmembrane import(GO:0071578) |
| 0.1 | 3.9 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.1 | 1.9 | GO:0044458 | motile cilium assembly(GO:0044458) |
| 0.1 | 0.4 | GO:0045925 | positive regulation of female receptivity(GO:0045925) |
| 0.1 | 1.7 | GO:0090129 | positive regulation of synapse maturation(GO:0090129) |
| 0.1 | 0.2 | GO:0035544 | negative regulation of SNARE complex assembly(GO:0035544) |
| 0.1 | 1.9 | GO:0001502 | cartilage condensation(GO:0001502) |
| 0.1 | 2.9 | GO:0009954 | proximal/distal pattern formation(GO:0009954) |
| 0.1 | 0.2 | GO:0060010 | Sertoli cell fate commitment(GO:0060010) |
| 0.1 | 0.6 | GO:1902514 | regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1902514) |
| 0.1 | 1.2 | GO:1990001 | inhibition of cysteine-type endopeptidase activity involved in apoptotic process(GO:1990001) |
| 0.1 | 2.1 | GO:0000184 | nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:0000184) |
| 0.1 | 1.4 | GO:0046039 | GTP metabolic process(GO:0046039) |
| 0.0 | 2.4 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.0 | 0.2 | GO:0090306 | spindle assembly involved in meiosis(GO:0090306) |
| 0.0 | 0.4 | GO:0060120 | auditory receptor cell fate commitment(GO:0009912) inner ear receptor cell fate commitment(GO:0060120) |
| 0.0 | 2.3 | GO:0043303 | mast cell activation involved in immune response(GO:0002279) mast cell degranulation(GO:0043303) |
| 0.0 | 1.3 | GO:0006144 | purine nucleobase metabolic process(GO:0006144) |
| 0.0 | 0.3 | GO:0003418 | growth plate cartilage chondrocyte differentiation(GO:0003418) |
| 0.0 | 1.2 | GO:1902042 | negative regulation of extrinsic apoptotic signaling pathway via death domain receptors(GO:1902042) |
| 0.0 | 4.4 | GO:0043154 | negative regulation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0043154) |
| 0.0 | 0.1 | GO:0045348 | positive regulation of transcription from RNA polymerase II promoter involved in unfolded protein response(GO:0006990) positive regulation of MHC class II biosynthetic process(GO:0045348) |
| 0.0 | 2.6 | GO:0090307 | mitotic spindle assembly(GO:0090307) |
| 0.0 | 0.9 | GO:0006336 | DNA replication-independent nucleosome assembly(GO:0006336) |
| 0.0 | 0.4 | GO:0001766 | membrane raft polarization(GO:0001766) membrane raft distribution(GO:0031580) |
| 0.0 | 6.6 | GO:0010466 | negative regulation of peptidase activity(GO:0010466) |
| 0.0 | 0.1 | GO:0036066 | protein O-linked fucosylation(GO:0036066) |
| 0.0 | 0.6 | GO:0016226 | iron-sulfur cluster assembly(GO:0016226) metallo-sulfur cluster assembly(GO:0031163) |
| 0.0 | 0.7 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.0 | 0.5 | GO:0001731 | formation of translation preinitiation complex(GO:0001731) |
| 0.0 | 1.8 | GO:0051972 | regulation of telomerase activity(GO:0051972) |
| 0.0 | 2.0 | GO:0046580 | negative regulation of Ras protein signal transduction(GO:0046580) |
| 0.0 | 0.3 | GO:0051151 | negative regulation of smooth muscle cell differentiation(GO:0051151) |
| 0.0 | 0.7 | GO:0022011 | myelination in peripheral nervous system(GO:0022011) peripheral nervous system axon ensheathment(GO:0032292) |
| 0.0 | 0.5 | GO:1903861 | positive regulation of dendrite extension(GO:1903861) |
| 0.0 | 3.4 | GO:0043123 | positive regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043123) |
| 0.0 | 0.3 | GO:0008090 | retrograde axonal transport(GO:0008090) |
| 0.0 | 4.5 | GO:0045089 | positive regulation of innate immune response(GO:0045089) |
| 0.0 | 1.3 | GO:0031338 | regulation of vesicle fusion(GO:0031338) |
| 0.0 | 0.7 | GO:0051351 | positive regulation of ligase activity(GO:0051351) |
| 0.0 | 1.3 | GO:0033045 | regulation of sister chromatid segregation(GO:0033045) |
| 0.0 | 0.5 | GO:0045747 | positive regulation of Notch signaling pathway(GO:0045747) regulation of clathrin-mediated endocytosis(GO:2000369) |
| 0.0 | 0.2 | GO:0043983 | histone H4-K12 acetylation(GO:0043983) |
| 0.0 | 0.5 | GO:0045879 | negative regulation of smoothened signaling pathway(GO:0045879) |
| 0.0 | 0.1 | GO:1903142 | positive regulation of endothelial cell development(GO:1901552) positive regulation of establishment of endothelial barrier(GO:1903142) |
| 0.0 | 1.0 | GO:0006400 | tRNA modification(GO:0006400) |
| 0.0 | 1.2 | GO:0006094 | gluconeogenesis(GO:0006094) |
| 0.0 | 0.1 | GO:0048711 | positive regulation of astrocyte differentiation(GO:0048711) |
| 0.0 | 0.2 | GO:0043517 | positive regulation of DNA damage response, signal transduction by p53 class mediator(GO:0043517) |
| 0.0 | 0.6 | GO:0001938 | positive regulation of endothelial cell proliferation(GO:0001938) |
| 0.0 | 0.4 | GO:0006884 | cell volume homeostasis(GO:0006884) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.2 | 13.4 | GO:0031523 | Myb complex(GO:0031523) |
| 1.2 | 3.7 | GO:1990031 | pinceau fiber(GO:1990031) |
| 1.2 | 9.6 | GO:0097129 | cyclin D2-CDK4 complex(GO:0097129) |
| 1.2 | 6.9 | GO:0090498 | extrinsic component of Golgi membrane(GO:0090498) |
| 0.5 | 2.7 | GO:0005944 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) |
| 0.4 | 1.3 | GO:1902560 | GMP reductase complex(GO:1902560) |
| 0.4 | 4.0 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.4 | 7.2 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.4 | 4.8 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 0.4 | 1.1 | GO:0042643 | actomyosin, actin portion(GO:0042643) |
| 0.4 | 7.3 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.3 | 1.6 | GO:0044194 | cytolytic granule(GO:0044194) |
| 0.3 | 1.3 | GO:0031251 | PAN complex(GO:0031251) |
| 0.3 | 13.0 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.2 | 5.1 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.2 | 2.9 | GO:0005642 | annulate lamellae(GO:0005642) |
| 0.2 | 2.0 | GO:1990907 | beta-catenin-TCF complex(GO:1990907) |
| 0.2 | 1.8 | GO:0034750 | Scrib-APC-beta-catenin complex(GO:0034750) |
| 0.2 | 1.3 | GO:0070449 | elongin complex(GO:0070449) |
| 0.2 | 0.6 | GO:0097361 | CIA complex(GO:0097361) |
| 0.1 | 0.7 | GO:0032444 | activin responsive factor complex(GO:0032444) SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.1 | 2.9 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.1 | 3.9 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.1 | 3.0 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.1 | 4.6 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.1 | 1.4 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.1 | 0.9 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.1 | 1.2 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.1 | 2.4 | GO:0005652 | integral component of nuclear inner membrane(GO:0005639) nuclear lamina(GO:0005652) intrinsic component of nuclear inner membrane(GO:0031229) |
| 0.1 | 7.4 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.1 | 1.0 | GO:0072379 | BAT3 complex(GO:0071818) ER membrane insertion complex(GO:0072379) |
| 0.1 | 2.3 | GO:0005671 | Ada2/Gcn5/Ada3 transcription activator complex(GO:0005671) |
| 0.1 | 8.2 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.1 | 2.5 | GO:0034362 | low-density lipoprotein particle(GO:0034362) |
| 0.1 | 5.4 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.1 | 1.2 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.1 | 0.9 | GO:0005775 | vacuolar lumen(GO:0005775) |
| 0.1 | 0.9 | GO:0030915 | Smc5-Smc6 complex(GO:0030915) |
| 0.1 | 1.9 | GO:0044292 | dendrite terminus(GO:0044292) |
| 0.1 | 0.5 | GO:0071439 | clathrin complex(GO:0071439) |
| 0.1 | 6.6 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.1 | 6.1 | GO:0005798 | Golgi-associated vesicle(GO:0005798) |
| 0.1 | 2.1 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.1 | 13.6 | GO:0005884 | actin filament(GO:0005884) |
| 0.1 | 3.3 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.1 | 1.9 | GO:0000421 | autophagosome membrane(GO:0000421) |
| 0.1 | 5.3 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.1 | 1.7 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.1 | 0.5 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.0 | 6.9 | GO:0005604 | basement membrane(GO:0005604) |
| 0.0 | 2.5 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.0 | 3.1 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.0 | 1.3 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.0 | 2.9 | GO:0035097 | histone methyltransferase complex(GO:0035097) |
| 0.0 | 1.8 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.0 | 5.1 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 5.6 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 2.2 | GO:0032809 | neuronal cell body membrane(GO:0032809) |
| 0.0 | 0.3 | GO:0031673 | hemidesmosome(GO:0030056) H zone(GO:0031673) |
| 0.0 | 3.0 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 3.9 | GO:0000922 | spindle pole(GO:0000922) |
| 0.0 | 1.2 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.0 | 2.1 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 0.2 | GO:0071203 | WASH complex(GO:0071203) |
| 0.0 | 2.6 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 0.3 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.0 | 2.0 | GO:0031903 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.0 | 7.5 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 0.2 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.0 | 0.1 | GO:0035749 | myelin sheath adaxonal region(GO:0035749) |
| 0.0 | 0.6 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.0 | 7.3 | GO:0005874 | microtubule(GO:0005874) |
| 0.0 | 0.1 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.0 | 0.2 | GO:0042582 | primary lysosome(GO:0005766) azurophil granule(GO:0042582) |
| 0.0 | 8.4 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 1.2 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.0 | 1.3 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.0 | 3.0 | GO:0005769 | early endosome(GO:0005769) |
| 0.0 | 1.1 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.0 | 4.2 | GO:0098793 | presynapse(GO:0098793) |
| 0.0 | 6.4 | GO:0015629 | actin cytoskeleton(GO:0015629) |
| 0.0 | 0.7 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 0.2 | GO:0017101 | aminoacyl-tRNA synthetase multienzyme complex(GO:0017101) |
| 0.0 | 0.5 | GO:0005581 | collagen trimer(GO:0005581) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.8 | 10.8 | GO:0034988 | Fc-gamma receptor I complex binding(GO:0034988) |
| 1.3 | 7.6 | GO:0016019 | N-acetylmuramoyl-L-alanine amidase activity(GO:0008745) peptidoglycan receptor activity(GO:0016019) |
| 1.2 | 4.6 | GO:0019828 | aspartic-type endopeptidase inhibitor activity(GO:0019828) |
| 1.1 | 5.7 | GO:0004534 | 5'-3' exoribonuclease activity(GO:0004534) |
| 1.1 | 4.5 | GO:0030297 | transmembrane receptor protein tyrosine kinase activator activity(GO:0030297) |
| 0.9 | 5.4 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.7 | 2.1 | GO:0016784 | 3-mercaptopyruvate sulfurtransferase activity(GO:0016784) |
| 0.7 | 1.3 | GO:0035373 | chondroitin sulfate proteoglycan binding(GO:0035373) |
| 0.7 | 3.3 | GO:0015319 | sodium:inorganic phosphate symporter activity(GO:0015319) |
| 0.6 | 6.1 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.6 | 2.4 | GO:0004939 | beta-adrenergic receptor activity(GO:0004939) |
| 0.5 | 2.1 | GO:1902379 | chemoattractant activity involved in axon guidance(GO:1902379) |
| 0.5 | 2.8 | GO:0001165 | RNA polymerase I upstream control element sequence-specific DNA binding(GO:0001165) |
| 0.4 | 1.3 | GO:0016657 | GMP reductase activity(GO:0003920) oxidoreductase activity, acting on NAD(P)H, nitrogenous group as acceptor(GO:0016657) |
| 0.4 | 3.0 | GO:0008294 | calcium- and calmodulin-responsive adenylate cyclase activity(GO:0008294) |
| 0.4 | 2.4 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 0.4 | 1.2 | GO:0004619 | bisphosphoglycerate mutase activity(GO:0004082) phosphoglycerate mutase activity(GO:0004619) 2,3-bisphosphoglycerate-dependent phosphoglycerate mutase activity(GO:0046538) |
| 0.4 | 4.8 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.4 | 2.9 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.4 | 2.1 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 0.3 | 2.0 | GO:0003938 | IMP dehydrogenase activity(GO:0003938) |
| 0.3 | 7.3 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.3 | 2.9 | GO:0003876 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.3 | 1.1 | GO:0050501 | hyaluronan synthase activity(GO:0050501) |
| 0.3 | 2.0 | GO:0030284 | estrogen receptor activity(GO:0030284) |
| 0.3 | 1.1 | GO:0034739 | histone deacetylase activity (H4-K16 specific)(GO:0034739) |
| 0.3 | 4.5 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.3 | 2.9 | GO:0004117 | calmodulin-dependent cyclic-nucleotide phosphodiesterase activity(GO:0004117) calcium- and calmodulin-regulated 3',5'-cyclic-GMP phosphodiesterase activity(GO:0048101) |
| 0.3 | 3.4 | GO:0019966 | interleukin-1 binding(GO:0019966) |
| 0.3 | 2.5 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.3 | 1.0 | GO:0005136 | interleukin-4 receptor binding(GO:0005136) |
| 0.3 | 3.5 | GO:0042609 | CD4 receptor binding(GO:0042609) |
| 0.2 | 1.2 | GO:0001537 | N-acetylgalactosamine 4-O-sulfotransferase activity(GO:0001537) |
| 0.2 | 2.9 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.2 | 4.0 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.2 | 2.0 | GO:0051022 | Rho GDP-dissociation inhibitor binding(GO:0051022) |
| 0.2 | 2.4 | GO:0070087 | chromo shadow domain binding(GO:0070087) |
| 0.2 | 5.8 | GO:0097602 | cullin family protein binding(GO:0097602) |
| 0.2 | 2.3 | GO:0016783 | sulfurtransferase activity(GO:0016783) |
| 0.2 | 7.3 | GO:0008187 | poly-pyrimidine tract binding(GO:0008187) |
| 0.2 | 2.3 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) |
| 0.2 | 2.4 | GO:0001069 | regulatory region RNA binding(GO:0001069) |
| 0.2 | 5.1 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.2 | 1.6 | GO:0005030 | neurotrophin receptor activity(GO:0005030) |
| 0.2 | 5.0 | GO:0019956 | chemokine binding(GO:0019956) |
| 0.1 | 0.7 | GO:0030618 | transforming growth factor beta receptor, pathway-specific cytoplasmic mediator activity(GO:0030618) |
| 0.1 | 3.9 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.1 | 5.2 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.1 | 3.7 | GO:0004033 | aldo-keto reductase (NADP) activity(GO:0004033) |
| 0.1 | 3.9 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) |
| 0.1 | 1.2 | GO:0001055 | RNA polymerase I activity(GO:0001054) RNA polymerase II activity(GO:0001055) |
| 0.1 | 1.4 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.1 | 2.7 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.1 | 14.9 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.1 | 10.7 | GO:0005518 | collagen binding(GO:0005518) |
| 0.1 | 0.3 | GO:0098640 | protein binding involved in cell-matrix adhesion(GO:0098634) integrin binding involved in cell-matrix adhesion(GO:0098640) |
| 0.1 | 5.2 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.1 | 0.5 | GO:0033592 | RNA strand annealing activity(GO:0033592) |
| 0.1 | 8.1 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.1 | 1.7 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.1 | 1.9 | GO:0042608 | T cell receptor binding(GO:0042608) |
| 0.1 | 1.4 | GO:0005522 | profilin binding(GO:0005522) |
| 0.1 | 4.4 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.1 | 2.1 | GO:0042162 | telomeric DNA binding(GO:0042162) |
| 0.1 | 1.3 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.1 | 0.4 | GO:0004645 | phosphorylase activity(GO:0004645) |
| 0.1 | 1.3 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.1 | 1.7 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.1 | 1.3 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.1 | 2.3 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.1 | 1.5 | GO:0031078 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.1 | 1.8 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.1 | 0.4 | GO:0033142 | progesterone receptor binding(GO:0033142) |
| 0.1 | 1.2 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.1 | 3.1 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.1 | 1.1 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.1 | 5.0 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.1 | 0.6 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.1 | 1.2 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.1 | 3.1 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.1 | 4.1 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.1 | 1.7 | GO:0005035 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.1 | 4.6 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 2.6 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.0 | 3.4 | GO:0016836 | hydro-lyase activity(GO:0016836) |
| 0.0 | 2.3 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 2.1 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.0 | 0.5 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.0 | 1.5 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.2 | GO:0035014 | phosphatidylinositol 3-kinase regulator activity(GO:0035014) |
| 0.0 | 0.8 | GO:0030275 | LRR domain binding(GO:0030275) |
| 0.0 | 0.9 | GO:0031491 | nucleosome binding(GO:0031491) |
| 0.0 | 1.2 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.0 | 1.5 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.0 | 2.5 | GO:0070491 | repressing transcription factor binding(GO:0070491) |
| 0.0 | 0.9 | GO:0019789 | SUMO transferase activity(GO:0019789) |
| 0.0 | 2.0 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.0 | 0.4 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 0.1 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) |
| 0.0 | 0.8 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.0 | 0.6 | GO:0030331 | estrogen receptor binding(GO:0030331) |
| 0.0 | 0.8 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.0 | 0.3 | GO:0001618 | virus receptor activity(GO:0001618) |
| 0.0 | 0.4 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 0.0 | 2.3 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.0 | 0.8 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) |
| 0.0 | 1.5 | GO:0031490 | chromatin DNA binding(GO:0031490) |
| 0.0 | 0.4 | GO:0005112 | Notch binding(GO:0005112) |
| 0.0 | 4.4 | GO:0061659 | ubiquitin protein ligase activity(GO:0061630) ubiquitin-like protein ligase activity(GO:0061659) |
| 0.0 | 0.1 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.0 | 0.1 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.0 | 1.7 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.0 | 3.4 | GO:0015631 | tubulin binding(GO:0015631) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 12.8 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
| 0.2 | 3.1 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.2 | 5.6 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.2 | 2.4 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.2 | 9.6 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.1 | 7.6 | PID ERBB1 RECEPTOR PROXIMAL PATHWAY | EGF receptor (ErbB1) signaling pathway |
| 0.1 | 9.8 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.1 | 2.9 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.1 | 4.2 | SA B CELL RECEPTOR COMPLEXES | Antigen binding to B cell receptors activates protein tyrosine kinases, such as the Src family, which ultimate activate MAP kinases. |
| 0.1 | 5.7 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.1 | 3.0 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.1 | 0.9 | ST INTERFERON GAMMA PATHWAY | Interferon gamma pathway. |
| 0.1 | 1.2 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.1 | 3.4 | PID CD8 TCR DOWNSTREAM PATHWAY | Downstream signaling in naïve CD8+ T cells |
| 0.1 | 3.1 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.1 | 6.2 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.1 | 4.8 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.1 | 2.7 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.1 | 1.8 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.1 | 4.5 | PID BMP PATHWAY | BMP receptor signaling |
| 0.1 | 1.7 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.1 | 1.5 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.1 | 2.1 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.1 | 2.4 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.1 | 0.7 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.1 | 2.1 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.1 | 0.9 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 1.2 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.0 | 1.4 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.0 | 2.1 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.0 | 1.8 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 2.3 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.0 | 1.1 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.0 | 1.8 | PID FGF PATHWAY | FGF signaling pathway |
| 0.0 | 2.6 | PID E2F PATHWAY | E2F transcription factor network |
| 0.0 | 1.5 | PID HDAC CLASSII PATHWAY | Signaling events mediated by HDAC Class II |
| 0.0 | 3.1 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 1.7 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 0.3 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.0 | 6.3 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.0 | 0.4 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.0 | 0.5 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 0.8 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.0 | 0.5 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 0.1 | PID S1P S1P2 PATHWAY | S1P2 pathway |
| 0.0 | 0.2 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.0 | 2.1 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 13.0 | REACTOME NEF MEDIATED DOWNREGULATION OF MHC CLASS I COMPLEX CELL SURFACE EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.3 | 5.0 | REACTOME THE ROLE OF NEF IN HIV1 REPLICATION AND DISEASE PATHOGENESIS | Genes involved in The role of Nef in HIV-1 replication and disease pathogenesis |
| 0.3 | 5.6 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.2 | 4.8 | REACTOME PD1 SIGNALING | Genes involved in PD-1 signaling |
| 0.2 | 11.6 | REACTOME GPVI MEDIATED ACTIVATION CASCADE | Genes involved in GPVI-mediated activation cascade |
| 0.2 | 2.9 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.2 | 11.4 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.2 | 2.8 | REACTOME RNA POL I PROMOTER OPENING | Genes involved in RNA Polymerase I Promoter Opening |
| 0.2 | 3.0 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.2 | 4.3 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.2 | 1.7 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.2 | 3.0 | REACTOME CTNNB1 PHOSPHORYLATION CASCADE | Genes involved in Beta-catenin phosphorylation cascade |
| 0.2 | 2.4 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.1 | 4.4 | REACTOME ERK MAPK TARGETS | Genes involved in ERK/MAPK targets |
| 0.1 | 3.9 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.1 | 1.7 | REACTOME PHOSPHORYLATION OF THE APC C | Genes involved in Phosphorylation of the APC/C |
| 0.1 | 2.1 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.1 | 2.9 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.1 | 2.6 | REACTOME COSTIMULATION BY THE CD28 FAMILY | Genes involved in Costimulation by the CD28 family |
| 0.1 | 3.7 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.1 | 1.1 | REACTOME HYALURONAN METABOLISM | Genes involved in Hyaluronan metabolism |
| 0.1 | 1.2 | REACTOME VIRAL MESSENGER RNA SYNTHESIS | Genes involved in Viral Messenger RNA Synthesis |
| 0.1 | 1.8 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 2.4 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.1 | 2.4 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.1 | 1.5 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.1 | 3.7 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 2.7 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.1 | 4.0 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.1 | 2.3 | REACTOME PROSTACYCLIN SIGNALLING THROUGH PROSTACYCLIN RECEPTOR | Genes involved in Prostacyclin signalling through prostacyclin receptor |
| 0.0 | 1.3 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 1.8 | REACTOME SIGNAL TRANSDUCTION BY L1 | Genes involved in Signal transduction by L1 |
| 0.0 | 0.4 | REACTOME RECEPTOR LIGAND BINDING INITIATES THE SECOND PROTEOLYTIC CLEAVAGE OF NOTCH RECEPTOR | Genes involved in Receptor-ligand binding initiates the second proteolytic cleavage of Notch receptor |
| 0.0 | 3.0 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.0 | 0.2 | REACTOME RAF MAP KINASE CASCADE | Genes involved in RAF/MAP kinase cascade |
| 0.0 | 1.6 | REACTOME INTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Intrinsic Pathway for Apoptosis |
| 0.0 | 1.2 | REACTOME NEP NS2 INTERACTS WITH THE CELLULAR EXPORT MACHINERY | Genes involved in NEP/NS2 Interacts with the Cellular Export Machinery |
| 0.0 | 1.2 | REACTOME CHONDROITIN SULFATE DERMATAN SULFATE METABOLISM | Genes involved in Chondroitin sulfate/dermatan sulfate metabolism |
| 0.0 | 0.5 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.0 | 2.4 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.7 | REACTOME SMAD2 SMAD3 SMAD4 HETEROTRIMER REGULATES TRANSCRIPTION | Genes involved in SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription |
| 0.0 | 5.2 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.5 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 0.6 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.0 | 2.8 | REACTOME NONSENSE MEDIATED DECAY ENHANCED BY THE EXON JUNCTION COMPLEX | Genes involved in Nonsense Mediated Decay Enhanced by the Exon Junction Complex |
| 0.0 | 1.2 | REACTOME APOPTOTIC CLEAVAGE OF CELLULAR PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |
| 0.0 | 0.4 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.0 | 1.4 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.0 | 0.1 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.0 | 0.5 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |