Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
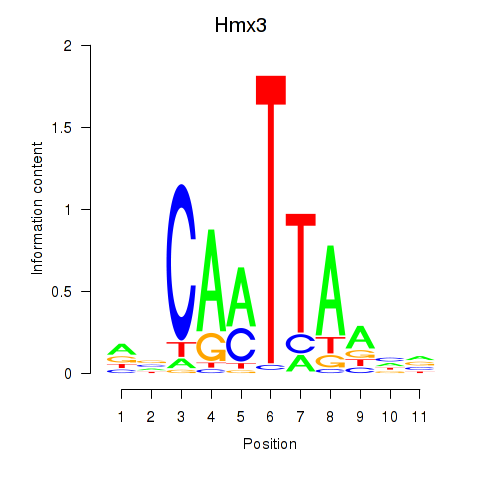
Results for Hmx3
Z-value: 0.68
Transcription factors associated with Hmx3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Hmx3
|
ENSMUSG00000040148.6 | Hmx3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Hmx3 | mm39_v1_chr7_+_131144596_131144596 | 0.20 | 9.0e-02 | Click! |
Activity profile of Hmx3 motif
Sorted Z-values of Hmx3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Hmx3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_-_103114105 | 4.91 |
ENSMUST00000111174.8
|
Ehf
|
ets homologous factor |
| chr16_+_34815177 | 4.73 |
ENSMUST00000231589.2
|
Mylk
|
myosin, light polypeptide kinase |
| chr3_-_49711706 | 4.65 |
ENSMUST00000191794.2
|
Pcdh18
|
protocadherin 18 |
| chr3_-_49711765 | 4.62 |
ENSMUST00000035931.13
|
Pcdh18
|
protocadherin 18 |
| chr1_-_93088562 | 4.61 |
ENSMUST00000143419.2
|
Mab21l4
|
mab-21-like 4 |
| chr5_-_53864874 | 4.40 |
ENSMUST00000031093.5
|
Cckar
|
cholecystokinin A receptor |
| chr11_+_96209093 | 4.26 |
ENSMUST00000049241.9
|
Hoxb4
|
homeobox B4 |
| chr2_+_27055245 | 3.94 |
ENSMUST00000000910.7
|
Dbh
|
dopamine beta hydroxylase |
| chr1_-_45542442 | 3.62 |
ENSMUST00000086430.5
|
Col5a2
|
collagen, type V, alpha 2 |
| chr2_+_74528071 | 3.32 |
ENSMUST00000059272.10
|
Hoxd9
|
homeobox D9 |
| chr3_+_3699205 | 3.29 |
ENSMUST00000108394.3
|
Hnf4g
|
hepatocyte nuclear factor 4, gamma |
| chrX_+_138464065 | 3.06 |
ENSMUST00000113027.8
|
Rnf128
|
ring finger protein 128 |
| chr1_-_149836974 | 2.92 |
ENSMUST00000190507.2
ENSMUST00000070200.15 |
Pla2g4a
|
phospholipase A2, group IVA (cytosolic, calcium-dependent) |
| chr9_-_110453427 | 2.65 |
ENSMUST00000196876.2
ENSMUST00000035069.14 |
Nradd
|
neurotrophin receptor associated death domain |
| chr2_+_57887896 | 2.60 |
ENSMUST00000112616.8
ENSMUST00000166729.2 |
Galnt5
|
polypeptide N-acetylgalactosaminyltransferase 5 |
| chr7_+_78922947 | 2.47 |
ENSMUST00000037315.13
|
Abhd2
|
abhydrolase domain containing 2 |
| chr3_-_86455575 | 2.12 |
ENSMUST00000077524.4
|
Mab21l2
|
mab-21-like 2 |
| chr7_+_83281193 | 2.04 |
ENSMUST00000117410.2
|
Stard5
|
StAR-related lipid transfer (START) domain containing 5 |
| chr7_-_45159739 | 1.98 |
ENSMUST00000211682.2
ENSMUST00000033096.16 ENSMUST00000209436.2 ENSMUST00000211343.2 |
Nucb1
|
nucleobindin 1 |
| chr6_-_87312743 | 1.97 |
ENSMUST00000042025.12
ENSMUST00000205033.2 |
Antxr1
|
anthrax toxin receptor 1 |
| chr7_+_79944198 | 1.89 |
ENSMUST00000163812.9
ENSMUST00000047558.14 ENSMUST00000174199.8 ENSMUST00000173824.8 ENSMUST00000174172.8 |
Prc1
|
protein regulator of cytokinesis 1 |
| chr5_+_110987839 | 1.82 |
ENSMUST00000200172.2
ENSMUST00000066160.3 |
Chek2
|
checkpoint kinase 2 |
| chr4_+_135870808 | 1.82 |
ENSMUST00000008016.3
|
Id3
|
inhibitor of DNA binding 3 |
| chr6_-_87312681 | 1.79 |
ENSMUST00000204805.3
|
Antxr1
|
anthrax toxin receptor 1 |
| chr14_+_51333816 | 1.77 |
ENSMUST00000169895.3
|
Rnase4
|
ribonuclease, RNase A family 4 |
| chr4_-_45532470 | 1.74 |
ENSMUST00000147448.2
|
Shb
|
src homology 2 domain-containing transforming protein B |
| chr2_-_150510116 | 1.68 |
ENSMUST00000028944.4
|
Acss1
|
acyl-CoA synthetase short-chain family member 1 |
| chr12_-_104439589 | 1.63 |
ENSMUST00000021513.6
|
Gsc
|
goosecoid homeobox |
| chr11_+_114559350 | 1.62 |
ENSMUST00000106602.10
ENSMUST00000077915.10 ENSMUST00000106599.8 ENSMUST00000082092.5 |
Rpl38
|
ribosomal protein L38 |
| chr7_-_45159790 | 1.59 |
ENSMUST00000211765.2
|
Nucb1
|
nucleobindin 1 |
| chr17_-_36501112 | 1.49 |
ENSMUST00000025312.13
ENSMUST00000102675.10 |
H2-T3
|
histocompatibility 2, T region locus 3 |
| chr7_+_43339842 | 1.33 |
ENSMUST00000056329.7
|
Klk14
|
kallikrein related-peptidase 14 |
| chr6_-_129599645 | 1.28 |
ENSMUST00000032252.8
|
Klrk1
|
killer cell lectin-like receptor subfamily K, member 1 |
| chr18_-_88912446 | 1.27 |
ENSMUST00000070116.12
ENSMUST00000125362.8 |
Socs6
|
suppressor of cytokine signaling 6 |
| chr4_+_19818718 | 1.21 |
ENSMUST00000035890.8
|
Slc7a13
|
solute carrier family 7, (cationic amino acid transporter, y+ system) member 13 |
| chr6_+_86348286 | 1.20 |
ENSMUST00000089558.7
|
Snrpg
|
small nuclear ribonucleoprotein polypeptide G |
| chr14_-_73286535 | 1.06 |
ENSMUST00000169168.3
|
Cysltr2
|
cysteinyl leukotriene receptor 2 |
| chr14_-_73286504 | 1.04 |
ENSMUST00000044664.12
|
Cysltr2
|
cysteinyl leukotriene receptor 2 |
| chr2_-_45002902 | 1.04 |
ENSMUST00000076836.13
ENSMUST00000176732.8 ENSMUST00000200844.4 |
Zeb2
|
zinc finger E-box binding homeobox 2 |
| chr14_+_53562089 | 1.03 |
ENSMUST00000178100.3
|
Trav7n-6
|
T cell receptor alpha variable 7N-6 |
| chr11_-_99265721 | 1.00 |
ENSMUST00000006963.3
|
Krt28
|
keratin 28 |
| chr12_-_83968507 | 0.90 |
ENSMUST00000222439.2
ENSMUST00000135962.8 ENSMUST00000155112.8 ENSMUST00000136848.8 ENSMUST00000126943.2 ENSMUST00000117217.8 |
Numb
|
NUMB endocytic adaptor protein |
| chr4_+_134124691 | 0.90 |
ENSMUST00000105870.8
|
Pafah2
|
platelet-activating factor acetylhydrolase 2 |
| chr15_-_101801351 | 0.89 |
ENSMUST00000100179.2
|
Krt76
|
keratin 76 |
| chr17_-_24054717 | 0.82 |
ENSMUST00000059906.8
|
Prss33
|
protease, serine 33 |
| chr2_-_152857239 | 0.79 |
ENSMUST00000028972.9
|
Pdrg1
|
p53 and DNA damage regulated 1 |
| chr17_-_57554631 | 0.75 |
ENSMUST00000233568.2
ENSMUST00000005975.8 |
Gpr108
|
G protein-coupled receptor 108 |
| chr18_-_88912415 | 0.68 |
ENSMUST00000145120.3
|
Socs6
|
suppressor of cytokine signaling 6 |
| chr3_-_154034271 | 0.67 |
ENSMUST00000204403.2
|
Lhx8
|
LIM homeobox protein 8 |
| chr5_+_110988095 | 0.65 |
ENSMUST00000198373.2
|
Chek2
|
checkpoint kinase 2 |
| chr15_+_101371353 | 0.64 |
ENSMUST00000088049.5
|
Krt86
|
keratin 86 |
| chr17_-_36008863 | 0.62 |
ENSMUST00000146472.8
|
Ddr1
|
discoidin domain receptor family, member 1 |
| chr19_-_56810593 | 0.61 |
ENSMUST00000118592.8
|
Ccdc186
|
coiled-coil domain containing 186 |
| chr4_+_80752535 | 0.56 |
ENSMUST00000102831.8
|
Tyrp1
|
tyrosinase-related protein 1 |
| chr5_-_87630117 | 0.54 |
ENSMUST00000079811.13
ENSMUST00000144144.3 |
Ugt2a2
|
UDP glucuronosyltransferase 2 family, polypeptide A2 |
| chr7_+_126808016 | 0.54 |
ENSMUST00000206204.2
ENSMUST00000206772.2 |
Mylpf
|
myosin light chain, phosphorylatable, fast skeletal muscle |
| chr13_+_23930717 | 0.52 |
ENSMUST00000099703.5
|
H2bc3
|
H2B clustered histone 3 |
| chr10_-_107321938 | 0.51 |
ENSMUST00000000445.2
|
Myf5
|
myogenic factor 5 |
| chr10_-_80257681 | 0.49 |
ENSMUST00000156244.2
|
Tcf3
|
transcription factor 3 |
| chr1_-_144427302 | 0.43 |
ENSMUST00000184189.3
|
Rgs21
|
regulator of G-protein signalling 21 |
| chr2_-_84545504 | 0.42 |
ENSMUST00000035840.6
|
Zdhhc5
|
zinc finger, DHHC domain containing 5 |
| chr13_+_118851214 | 0.41 |
ENSMUST00000022246.9
|
Fgf10
|
fibroblast growth factor 10 |
| chr10_+_99099084 | 0.39 |
ENSMUST00000020118.5
ENSMUST00000220291.2 |
Dusp6
|
dual specificity phosphatase 6 |
| chr7_-_4909515 | 0.33 |
ENSMUST00000210663.2
|
Gm36210
|
predicted gene, 36210 |
| chr11_+_62770275 | 0.29 |
ENSMUST00000014321.5
|
Tvp23b
|
trans-golgi network vesicle protein 23B |
| chr9_+_106048116 | 0.29 |
ENSMUST00000020490.13
|
Wdr82
|
WD repeat domain containing 82 |
| chr4_+_80752360 | 0.28 |
ENSMUST00000133655.8
ENSMUST00000006151.13 |
Tyrp1
|
tyrosinase-related protein 1 |
| chr19_-_41921676 | 0.27 |
ENSMUST00000075280.12
ENSMUST00000112123.4 |
Exosc1
|
exosome component 1 |
| chr14_+_53698556 | 0.27 |
ENSMUST00000181728.3
|
Trav7-4
|
T cell receptor alpha variable 7-4 |
| chr11_-_99494134 | 0.26 |
ENSMUST00000072306.4
|
Gm11938
|
predicted gene 11938 |
| chr5_+_73164226 | 0.21 |
ENSMUST00000031127.11
ENSMUST00000201304.2 |
Slc10a4
|
solute carrier family 10 (sodium/bile acid cotransporter family), member 4 |
| chr11_+_6366259 | 0.20 |
ENSMUST00000213200.2
|
Ppia
|
peptidylprolyl isomerase A |
| chr7_-_106341163 | 0.19 |
ENSMUST00000050541.5
|
Olfr697
|
olfactory receptor 697 |
| chr3_+_96537484 | 0.17 |
ENSMUST00000200647.2
|
Rbm8a
|
RNA binding motif protein 8a |
| chr8_-_33374825 | 0.16 |
ENSMUST00000238791.2
|
Nrg1
|
neuregulin 1 |
| chr9_-_56151334 | 0.16 |
ENSMUST00000188142.7
|
Peak1
|
pseudopodium-enriched atypical kinase 1 |
| chr3_+_96537235 | 0.15 |
ENSMUST00000048915.11
ENSMUST00000196456.5 ENSMUST00000198027.5 |
Rbm8a
|
RNA binding motif protein 8a |
| chr19_-_12209960 | 0.13 |
ENSMUST00000207710.3
|
Olfr1432
|
olfactory receptor 1432 |
| chr16_-_58940431 | 0.12 |
ENSMUST00000072608.2
|
Olfr194
|
olfactory receptor 194 |
| chr8_-_23727639 | 0.10 |
ENSMUST00000033950.7
|
Gins4
|
GINS complex subunit 4 (Sld5 homolog) |
| chr14_+_52892115 | 0.08 |
ENSMUST00000198019.2
|
Trav7-1
|
T cell receptor alpha variable 7-1 |
| chr3_+_59914164 | 0.07 |
ENSMUST00000169794.2
|
Aadacl2
|
arylacetamide deacetylase like 2 |
| chr14_+_53007210 | 0.06 |
ENSMUST00000178768.4
|
Trav7d-4
|
T cell receptor alpha variable 7D-4 |
| chr4_+_136038243 | 0.05 |
ENSMUST00000131671.8
|
Hnrnpr
|
heterogeneous nuclear ribonucleoprotein R |
| chr19_-_12147438 | 0.03 |
ENSMUST00000207679.3
ENSMUST00000219261.2 |
Olfr1555-ps1
|
olfactory receptor 1555, pseudogene 1 |
| chr2_+_3115250 | 0.03 |
ENSMUST00000072955.12
|
Fam171a1
|
family with sequence similarity 171, member A1 |
| chr7_-_126496534 | 0.03 |
ENSMUST00000120007.8
|
Tmem219
|
transmembrane protein 219 |
| chr7_+_140147799 | 0.02 |
ENSMUST00000210973.3
|
Olfr538
|
olfactory receptor 538 |
| chr6_+_136530970 | 0.00 |
ENSMUST00000189535.2
|
Atf7ip
|
activating transcription factor 7 interacting protein |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 3.9 | GO:0006589 | octopamine biosynthetic process(GO:0006589) octopamine metabolic process(GO:0046333) |
| 0.9 | 4.7 | GO:0060414 | aorta smooth muscle tissue morphogenesis(GO:0060414) |
| 0.9 | 3.6 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 0.7 | 4.4 | GO:0002023 | reduction of food intake in response to dietary excess(GO:0002023) positive regulation of somatostatin secretion(GO:0090274) |
| 0.6 | 3.8 | GO:1903054 | negative regulation of extracellular matrix organization(GO:1903054) |
| 0.6 | 2.5 | GO:0051325 | interphase(GO:0051325) mitotic interphase(GO:0051329) |
| 0.6 | 1.7 | GO:0019427 | acetate biosynthetic process(GO:0019413) acetyl-CoA biosynthetic process from acetate(GO:0019427) propionate biosynthetic process(GO:0019542) |
| 0.5 | 4.3 | GO:0048539 | bone marrow development(GO:0048539) |
| 0.4 | 1.3 | GO:0030887 | positive regulation of myeloid dendritic cell activation(GO:0030887) |
| 0.4 | 2.9 | GO:2001280 | positive regulation of prostaglandin biosynthetic process(GO:0031394) positive regulation of unsaturated fatty acid biosynthetic process(GO:2001280) |
| 0.3 | 0.9 | GO:0030860 | regulation of polarized epithelial cell differentiation(GO:0030860) |
| 0.3 | 0.8 | GO:0006583 | melanin biosynthetic process from tyrosine(GO:0006583) |
| 0.3 | 9.3 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.2 | 1.7 | GO:1900194 | negative regulation of oocyte maturation(GO:1900194) |
| 0.1 | 1.0 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.1 | 1.6 | GO:0014029 | neural crest formation(GO:0014029) |
| 0.1 | 1.6 | GO:0048318 | axial mesoderm development(GO:0048318) |
| 0.1 | 0.4 | GO:0060435 | bronchiole development(GO:0060435) secretion by lung epithelial cell involved in lung growth(GO:0061033) |
| 0.1 | 2.5 | GO:0032570 | response to progesterone(GO:0032570) |
| 0.1 | 1.3 | GO:0007320 | insemination(GO:0007320) |
| 0.1 | 1.9 | GO:0048935 | peripheral nervous system neuron differentiation(GO:0048934) peripheral nervous system neuron development(GO:0048935) |
| 0.1 | 2.0 | GO:0035376 | sterol import(GO:0035376) cholesterol import(GO:0070508) |
| 0.1 | 0.5 | GO:0002326 | B cell lineage commitment(GO:0002326) |
| 0.1 | 3.1 | GO:0042036 | negative regulation of cytokine biosynthetic process(GO:0042036) |
| 0.1 | 0.4 | GO:1903224 | regulation of endodermal cell differentiation(GO:1903224) |
| 0.1 | 0.5 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 0.1 | 2.1 | GO:0010172 | embryonic body morphogenesis(GO:0010172) |
| 0.1 | 1.8 | GO:0030903 | notochord development(GO:0030903) |
| 0.0 | 0.6 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.0 | 0.9 | GO:0048733 | sebaceous gland development(GO:0048733) |
| 0.0 | 1.3 | GO:0002474 | antigen processing and presentation of peptide antigen via MHC class I(GO:0002474) |
| 0.0 | 1.9 | GO:1904893 | negative regulation of JAK-STAT cascade(GO:0046426) negative regulation of STAT cascade(GO:1904893) |
| 0.0 | 2.1 | GO:0007218 | neuropeptide signaling pathway(GO:0007218) |
| 0.0 | 0.2 | GO:0021840 | directional guidance of interneurons involved in migration from the subpallium to the cortex(GO:0021840) chemorepulsion involved in interneuron migration from the subpallium to the cortex(GO:0021842) |
| 0.0 | 0.1 | GO:0000727 | double-strand break repair via break-induced replication(GO:0000727) |
| 0.0 | 0.3 | GO:0080182 | histone H3-K4 trimethylation(GO:0080182) |
| 0.0 | 1.9 | GO:0001578 | microtubule bundle formation(GO:0001578) |
| 0.0 | 0.7 | GO:0021884 | forebrain neuron development(GO:0021884) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 3.9 | GO:0034774 | secretory granule lumen(GO:0034774) cytoplasmic membrane-bounded vesicle lumen(GO:0060205) |
| 1.2 | 3.6 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.6 | 3.6 | GO:0090498 | extrinsic component of Golgi membrane(GO:0090498) |
| 0.5 | 1.6 | GO:0070992 | translation initiation complex(GO:0070992) |
| 0.2 | 3.8 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.2 | 2.5 | GO:0097524 | sperm plasma membrane(GO:0097524) |
| 0.2 | 2.7 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.1 | 0.8 | GO:0045009 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.1 | 1.2 | GO:0005687 | U4 snRNP(GO:0005687) |
| 0.1 | 1.9 | GO:0070938 | contractile ring(GO:0070938) |
| 0.1 | 2.9 | GO:0042588 | zymogen granule(GO:0042588) |
| 0.1 | 0.8 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.1 | 4.7 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.1 | 0.7 | GO:0001674 | female germ cell nucleus(GO:0001674) |
| 0.0 | 0.5 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.0 | 1.5 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 0.3 | GO:0072357 | PTW/PP1 phosphatase complex(GO:0072357) |
| 0.0 | 0.1 | GO:0000811 | GINS complex(GO:0000811) |
| 0.0 | 1.9 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.0 | 4.4 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 0.3 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.0 | 2.5 | GO:0016605 | PML body(GO:0016605) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 3.9 | GO:0004500 | dopamine beta-monooxygenase activity(GO:0004500) |
| 0.5 | 4.7 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.4 | 1.3 | GO:0032394 | MHC class Ib receptor activity(GO:0032394) |
| 0.4 | 2.1 | GO:0004974 | leukotriene receptor activity(GO:0004974) |
| 0.3 | 1.7 | GO:0003987 | acetate-CoA ligase activity(GO:0003987) |
| 0.3 | 2.7 | GO:0005166 | neurotrophin p75 receptor binding(GO:0005166) |
| 0.3 | 2.9 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.2 | 2.0 | GO:0032052 | bile acid binding(GO:0032052) |
| 0.2 | 2.5 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
| 0.1 | 0.8 | GO:0016716 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, another compound as one donor, and incorporation of one atom of oxygen(GO:0016716) |
| 0.1 | 2.6 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.1 | 1.2 | GO:1990446 | U1 snRNP binding(GO:1990446) |
| 0.1 | 0.5 | GO:0070644 | vitamin D response element binding(GO:0070644) |
| 0.1 | 3.6 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 0.9 | GO:0003847 | 1-alkyl-2-acetylglycerophosphocholine esterase activity(GO:0003847) |
| 0.1 | 0.4 | GO:0005111 | type 2 fibroblast growth factor receptor binding(GO:0005111) |
| 0.1 | 0.6 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 0.1 | 0.9 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.1 | 3.6 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 3.8 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 3.3 | GO:0098531 | RNA polymerase II transcription factor activity, ligand-activated sequence-specific DNA binding(GO:0004879) transcription factor activity, direct ligand regulated sequence-specific DNA binding(GO:0098531) |
| 0.0 | 1.3 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 0.0 | 1.6 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.0 | 4.4 | GO:0008528 | G-protein coupled peptide receptor activity(GO:0008528) |
| 0.0 | 1.8 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.0 | 0.2 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.0 | 1.7 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.0 | 1.0 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.0 | 0.2 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.0 | 0.4 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.0 | 1.9 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.0 | 1.9 | GO:0004860 | protein kinase inhibitor activity(GO:0004860) |
| 0.0 | 0.5 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.0 | 1.8 | GO:0004540 | ribonuclease activity(GO:0004540) |
| 0.0 | 0.2 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 0.0 | 1.2 | GO:0015297 | antiporter activity(GO:0015297) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.8 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.1 | 4.7 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.1 | 2.5 | PID ATM PATHWAY | ATM pathway |
| 0.1 | 3.6 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.1 | 5.0 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.0 | 1.7 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.0 | 1.7 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.0 | 3.1 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.0 | 1.9 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.0 | 1.6 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.0 | 0.5 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.5 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.2 | 3.9 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.2 | 2.9 | REACTOME ACYL CHAIN REMODELLING OF PS | Genes involved in Acyl chain remodelling of PS |
| 0.2 | 1.7 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.1 | 2.1 | REACTOME EICOSANOID LIGAND BINDING RECEPTORS | Genes involved in Eicosanoid ligand-binding receptors |
| 0.1 | 3.3 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.1 | 1.2 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.1 | 4.7 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.1 | 3.6 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.1 | 1.9 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.1 | 0.2 | REACTOME BINDING AND ENTRY OF HIV VIRION | Genes involved in Binding and entry of HIV virion |
| 0.0 | 2.0 | REACTOME STEROID HORMONES | Genes involved in Steroid hormones |
| 0.0 | 2.6 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 0.3 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.0 | 4.4 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 0.4 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.0 | 1.3 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.0 | 0.4 | REACTOME REGULATION OF BETA CELL DEVELOPMENT | Genes involved in Regulation of beta-cell development |
| 0.0 | 0.9 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.0 | 1.0 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 1.6 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.0 | 0.3 | REACTOME NONSENSE MEDIATED DECAY ENHANCED BY THE EXON JUNCTION COMPLEX | Genes involved in Nonsense Mediated Decay Enhanced by the Exon Junction Complex |
| 0.0 | 0.2 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |