Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
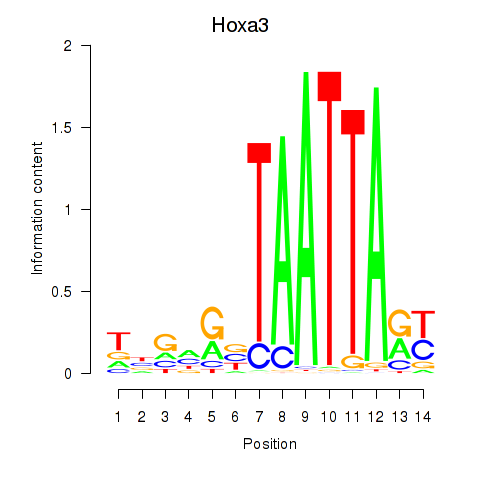
Results for Hoxa3
Z-value: 0.48
Transcription factors associated with Hoxa3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Hoxa3
|
ENSMUSG00000079560.14 | Hoxa3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Hoxa3 | mm39_v1_chr6_-_52160816_52160837 | 0.08 | 5.3e-01 | Click! |
Activity profile of Hoxa3 motif
Sorted Z-values of Hoxa3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Hoxa3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_+_172994841 | 4.27 |
ENSMUST00000029017.6
|
Pck1
|
phosphoenolpyruvate carboxykinase 1, cytosolic |
| chr14_+_79753055 | 3.58 |
ENSMUST00000110835.3
ENSMUST00000227192.2 |
Elf1
|
E74-like factor 1 |
| chr17_-_71160477 | 3.52 |
ENSMUST00000118283.8
|
Tgif1
|
TGFB-induced factor homeobox 1 |
| chr6_-_129449739 | 3.06 |
ENSMUST00000112076.9
ENSMUST00000184581.3 |
Clec7a
|
C-type lectin domain family 7, member a |
| chr6_-_136758716 | 3.04 |
ENSMUST00000078095.11
ENSMUST00000032338.10 |
Gucy2c
|
guanylate cyclase 2c |
| chr5_-_65855511 | 2.80 |
ENSMUST00000201948.4
|
Pds5a
|
PDS5 cohesin associated factor A |
| chr3_-_72875187 | 2.75 |
ENSMUST00000167334.8
|
Sis
|
sucrase isomaltase (alpha-glucosidase) |
| chr14_+_26722319 | 2.08 |
ENSMUST00000035433.10
|
Hesx1
|
homeobox gene expressed in ES cells |
| chr3_-_86455575 | 1.68 |
ENSMUST00000077524.4
|
Mab21l2
|
mab-21-like 2 |
| chr7_-_44753168 | 1.63 |
ENSMUST00000211085.2
ENSMUST00000210642.2 ENSMUST00000003512.9 |
Fcgrt
|
Fc fragment of IgG receptor and transporter |
| chr19_-_7943365 | 1.61 |
ENSMUST00000182102.8
ENSMUST00000075619.5 |
Slc22a27
|
solute carrier family 22, member 27 |
| chr1_-_171854818 | 1.55 |
ENSMUST00000138714.2
ENSMUST00000027837.13 ENSMUST00000111264.8 |
Vangl2
|
VANGL planar cell polarity 2 |
| chr2_-_168608949 | 1.51 |
ENSMUST00000075044.10
|
Sall4
|
spalt like transcription factor 4 |
| chr16_+_35861554 | 1.44 |
ENSMUST00000042203.10
|
Wdr5b
|
WD repeat domain 5B |
| chr10_-_37014859 | 1.42 |
ENSMUST00000092584.6
|
Marcks
|
myristoylated alanine rich protein kinase C substrate |
| chr11_-_99265721 | 1.29 |
ENSMUST00000006963.3
|
Krt28
|
keratin 28 |
| chr5_+_115373895 | 1.21 |
ENSMUST00000081497.13
|
Pop5
|
processing of precursor 5, ribonuclease P/MRP family (S. cerevisiae) |
| chr2_-_27365633 | 1.19 |
ENSMUST00000138693.8
ENSMUST00000113941.9 ENSMUST00000077737.13 |
Brd3
|
bromodomain containing 3 |
| chr12_-_99529767 | 1.13 |
ENSMUST00000176928.3
ENSMUST00000223484.2 |
Foxn3
|
forkhead box N3 |
| chrX_-_74460137 | 1.09 |
ENSMUST00000033542.11
|
Mtcp1
|
mature T cell proliferation 1 |
| chr2_+_36120438 | 1.09 |
ENSMUST00000062069.6
|
Ptgs1
|
prostaglandin-endoperoxide synthase 1 |
| chr9_+_40092216 | 0.99 |
ENSMUST00000218134.2
ENSMUST00000216720.2 ENSMUST00000214763.2 |
Olfr986
|
olfactory receptor 986 |
| chr13_-_103042294 | 0.98 |
ENSMUST00000167462.8
|
Mast4
|
microtubule associated serine/threonine kinase family member 4 |
| chr3_-_92441809 | 0.97 |
ENSMUST00000193521.2
|
2310046K23Rik
|
RIKEN cDNA 2310046K23 gene |
| chr12_+_59060162 | 0.93 |
ENSMUST00000021379.8
|
Gemin2
|
gem nuclear organelle associated protein 2 |
| chr15_-_101801351 | 0.93 |
ENSMUST00000100179.2
|
Krt76
|
keratin 76 |
| chrX_+_16485937 | 0.92 |
ENSMUST00000026013.6
|
Maoa
|
monoamine oxidase A |
| chrX_+_159551171 | 0.90 |
ENSMUST00000112368.3
|
Rs1
|
retinoschisis (X-linked, juvenile) 1 (human) |
| chr11_-_99441687 | 0.89 |
ENSMUST00000092700.5
|
Krtap3-3
|
keratin associated protein 3-3 |
| chr14_+_65612788 | 0.86 |
ENSMUST00000224687.2
|
Zfp395
|
zinc finger protein 395 |
| chrX_+_159551009 | 0.86 |
ENSMUST00000033650.14
|
Rs1
|
retinoschisis (X-linked, juvenile) 1 (human) |
| chr1_-_37955569 | 0.81 |
ENSMUST00000078307.7
|
Lyg2
|
lysozyme G-like 2 |
| chr17_+_38104635 | 0.77 |
ENSMUST00000214770.3
|
Olfr123
|
olfactory receptor 123 |
| chr2_-_168609110 | 0.73 |
ENSMUST00000029061.12
ENSMUST00000103074.2 |
Sall4
|
spalt like transcription factor 4 |
| chr6_-_41752111 | 0.65 |
ENSMUST00000214976.3
|
Olfr459
|
olfactory receptor 459 |
| chr13_-_103042554 | 0.64 |
ENSMUST00000171791.8
|
Mast4
|
microtubule associated serine/threonine kinase family member 4 |
| chr7_-_115637970 | 0.63 |
ENSMUST00000166877.8
|
Sox6
|
SRY (sex determining region Y)-box 6 |
| chr2_-_111843053 | 0.63 |
ENSMUST00000213559.3
|
Olfr1310
|
olfactory receptor 1310 |
| chr15_-_34356567 | 0.63 |
ENSMUST00000179647.2
|
9430069I07Rik
|
RIKEN cDNA 9430069I07 gene |
| chr5_+_90708962 | 0.59 |
ENSMUST00000094615.8
ENSMUST00000200765.2 |
Albfm1
|
albumin superfamily member 1 |
| chr5_+_138185747 | 0.53 |
ENSMUST00000110934.9
|
Cnpy4
|
canopy FGF signaling regulator 4 |
| chr10_-_129000620 | 0.53 |
ENSMUST00000214271.2
|
Olfr771
|
olfactory receptor 771 |
| chr2_+_88505972 | 0.41 |
ENSMUST00000216767.2
ENSMUST00000213893.2 |
Olfr1193
|
olfactory receptor 1193 |
| chr18_-_38336893 | 0.39 |
ENSMUST00000194312.2
|
Pcdh1
|
protocadherin 1 |
| chr14_-_36820304 | 0.34 |
ENSMUST00000022337.11
|
Cdhr1
|
cadherin-related family member 1 |
| chr2_+_109747984 | 0.31 |
ENSMUST00000046548.14
ENSMUST00000111037.3 |
Lgr4
|
leucine-rich repeat-containing G protein-coupled receptor 4 |
| chr17_+_38104420 | 0.31 |
ENSMUST00000216051.3
|
Olfr123
|
olfactory receptor 123 |
| chr7_-_44752508 | 0.28 |
ENSMUST00000209830.2
|
Fcgrt
|
Fc fragment of IgG receptor and transporter |
| chr6_+_63232955 | 0.27 |
ENSMUST00000095852.5
|
Grid2
|
glutamate receptor, ionotropic, delta 2 |
| chr10_+_73657753 | 0.24 |
ENSMUST00000134009.9
ENSMUST00000177420.7 ENSMUST00000125006.9 |
Pcdh15
|
protocadherin 15 |
| chr4_-_14621669 | 0.23 |
ENSMUST00000143105.2
|
Slc26a7
|
solute carrier family 26, member 7 |
| chr2_+_89642395 | 0.23 |
ENSMUST00000214508.2
|
Olfr1255
|
olfactory receptor 1255 |
| chr4_-_43710231 | 0.22 |
ENSMUST00000217544.2
ENSMUST00000107862.3 |
Olfr71
|
olfactory receptor 71 |
| chrX_+_110808231 | 0.19 |
ENSMUST00000207546.2
|
Gm45194
|
predicted gene 45194 |
| chr2_+_110551976 | 0.18 |
ENSMUST00000090332.5
|
Muc15
|
mucin 15 |
| chr13_-_43634695 | 0.17 |
ENSMUST00000144326.4
|
Ranbp9
|
RAN binding protein 9 |
| chr11_-_100653754 | 0.17 |
ENSMUST00000107360.3
ENSMUST00000055083.4 |
Hcrt
|
hypocretin |
| chr2_-_111820618 | 0.17 |
ENSMUST00000216948.2
ENSMUST00000214935.2 ENSMUST00000217452.2 ENSMUST00000215045.2 |
Olfr1309
|
olfactory receptor 1309 |
| chr3_-_58729732 | 0.14 |
ENSMUST00000191233.4
|
Mindy4b-ps
|
MINDY lysine 48 deubiquitinase 4B, pseudogene |
| chr1_-_173018204 | 0.12 |
ENSMUST00000215878.2
ENSMUST00000201132.3 |
Olfr1406
|
olfactory receptor 1406 |
| chr5_+_108842294 | 0.06 |
ENSMUST00000013633.12
|
Fgfrl1
|
fibroblast growth factor receptor-like 1 |
| chr2_+_22959223 | 0.05 |
ENSMUST00000114523.10
|
Acbd5
|
acyl-Coenzyme A binding domain containing 5 |
| chr14_-_52277310 | 0.05 |
ENSMUST00000216907.2
ENSMUST00000214071.2 ENSMUST00000216188.2 |
Olfr221
|
olfactory receptor 221 |
| chr5_-_138185686 | 0.04 |
ENSMUST00000110936.8
|
Taf6
|
TATA-box binding protein associated factor 6 |
| chr3_+_159545309 | 0.04 |
ENSMUST00000068952.10
ENSMUST00000198878.2 |
Wls
|
wntless WNT ligand secretion mediator |
| chr5_-_138185438 | 0.04 |
ENSMUST00000110937.8
ENSMUST00000139276.2 ENSMUST00000048698.14 ENSMUST00000123415.8 |
Taf6
|
TATA-box binding protein associated factor 6 |
| chr11_-_99979052 | 0.04 |
ENSMUST00000107419.2
|
Krt32
|
keratin 32 |
| chr15_+_65682066 | 0.03 |
ENSMUST00000211878.2
|
Efr3a
|
EFR3 homolog A |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 4.3 | GO:0061402 | glycerol biosynthetic process(GO:0006114) positive regulation of transcription from RNA polymerase II promoter in response to acidic pH(GO:0061402) |
| 0.5 | 1.5 | GO:0060448 | dichotomous subdivision of terminal units involved in lung branching(GO:0060448) |
| 0.4 | 1.8 | GO:0036367 | adaptation of rhodopsin mediated signaling(GO:0016062) light adaption(GO:0036367) |
| 0.2 | 2.1 | GO:0030916 | otic vesicle formation(GO:0030916) |
| 0.2 | 1.1 | GO:0019371 | cyclooxygenase pathway(GO:0019371) maintenance of blood-brain barrier(GO:0035633) |
| 0.2 | 3.5 | GO:0048387 | negative regulation of retinoic acid receptor signaling pathway(GO:0048387) |
| 0.1 | 1.2 | GO:0009249 | protein lipoylation(GO:0009249) |
| 0.1 | 0.9 | GO:0042420 | dopamine catabolic process(GO:0042420) |
| 0.1 | 1.6 | GO:0015747 | urate transport(GO:0015747) |
| 0.1 | 0.3 | GO:0061290 | cell-cell signaling involved in kidney development(GO:0060995) Wnt signaling pathway involved in kidney development(GO:0061289) canonical Wnt signaling pathway involved in metanephric kidney development(GO:0061290) cell-cell signaling involved in metanephros development(GO:0072204) |
| 0.1 | 0.8 | GO:0000270 | peptidoglycan metabolic process(GO:0000270) peptidoglycan catabolic process(GO:0009253) |
| 0.1 | 3.6 | GO:0050860 | negative regulation of T cell receptor signaling pathway(GO:0050860) |
| 0.1 | 2.2 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 0.1 | 2.8 | GO:0007064 | mitotic sister chromatid cohesion(GO:0007064) |
| 0.1 | 0.6 | GO:2000741 | positive regulation of mesenchymal stem cell differentiation(GO:2000741) |
| 0.1 | 3.0 | GO:0006182 | cGMP biosynthetic process(GO:0006182) |
| 0.0 | 1.7 | GO:0010172 | embryonic body morphogenesis(GO:0010172) |
| 0.0 | 1.4 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.0 | 1.1 | GO:0097094 | craniofacial suture morphogenesis(GO:0097094) |
| 0.0 | 0.9 | GO:0048733 | sebaceous gland development(GO:0048733) |
| 0.0 | 0.9 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.0 | 0.2 | GO:0051970 | negative regulation of transmission of nerve impulse(GO:0051970) |
| 0.0 | 0.3 | GO:1900454 | positive regulation of long term synaptic depression(GO:1900454) |
| 0.0 | 0.0 | GO:0061357 | positive regulation of Wnt protein secretion(GO:0061357) |
| 0.0 | 0.4 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.5 | GO:0060187 | cell pole(GO:0060187) |
| 0.2 | 1.4 | GO:0042585 | germinal vesicle(GO:0042585) |
| 0.2 | 1.2 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.1 | 3.0 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.1 | 0.9 | GO:0097504 | Gemini of coiled bodies(GO:0097504) |
| 0.0 | 0.9 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 0.3 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.0 | 1.3 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 2.2 | GO:0000792 | heterochromatin(GO:0000792) |
| 0.0 | 2.7 | GO:0005903 | brush border(GO:0005903) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 4.3 | GO:0004611 | phosphoenolpyruvate carboxykinase activity(GO:0004611) phosphoenolpyruvate carboxykinase (GTP) activity(GO:0004613) |
| 0.7 | 2.7 | GO:0004574 | oligo-1,6-glucosidase activity(GO:0004574) |
| 0.4 | 1.2 | GO:0000171 | ribonuclease MRP activity(GO:0000171) |
| 0.3 | 1.9 | GO:0019770 | IgG receptor activity(GO:0019770) |
| 0.2 | 1.8 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 0.2 | 3.0 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.1 | 3.5 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.1 | 0.9 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.1 | 1.6 | GO:0015143 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.0 | 0.8 | GO:0003796 | lysozyme activity(GO:0003796) |
| 0.0 | 1.2 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.0 | 0.1 | GO:0001571 | non-tyrosine kinase fibroblast growth factor receptor activity(GO:0001571) |
| 0.0 | 0.3 | GO:0016500 | protein-hormone receptor activity(GO:0016500) |
| 0.0 | 1.1 | GO:0004601 | peroxidase activity(GO:0004601) |
| 0.0 | 1.4 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.0 | 2.7 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 4.3 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 3.6 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.0 | 3.7 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.0 | 2.2 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 3.1 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 4.3 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.1 | 1.4 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.1 | 3.5 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.1 | 0.9 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.0 | 1.1 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.0 | 0.9 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |