Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
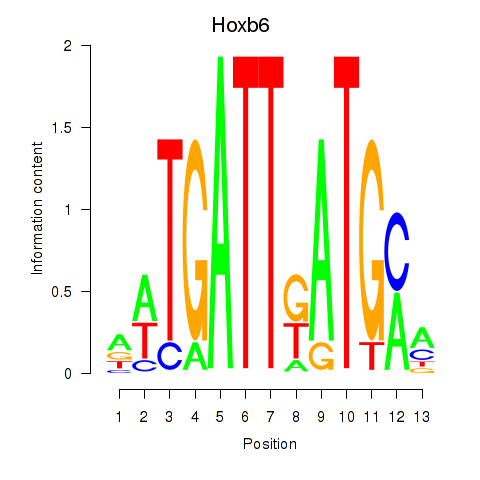
Results for Hoxb6
Z-value: 0.84
Transcription factors associated with Hoxb6
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Hoxb6
|
ENSMUSG00000000690.6 | Hoxb6 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Hoxb6 | mm39_v1_chr11_+_96189963_96189997 | 0.06 | 6.0e-01 | Click! |
Activity profile of Hoxb6 motif
Sorted Z-values of Hoxb6 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Hoxb6
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr16_-_19079594 | 11.02 |
ENSMUST00000103752.3
ENSMUST00000197518.2 |
Iglv2
|
immunoglobulin lambda variable 2 |
| chr11_-_99134885 | 7.72 |
ENSMUST00000103132.10
ENSMUST00000038214.7 |
Krt222
|
keratin 222 |
| chr1_-_138102972 | 6.56 |
ENSMUST00000195533.6
ENSMUST00000183301.8 |
Ptprc
|
protein tyrosine phosphatase, receptor type, C |
| chr1_-_138103021 | 6.25 |
ENSMUST00000182755.8
ENSMUST00000193650.2 ENSMUST00000182283.8 |
Ptprc
|
protein tyrosine phosphatase, receptor type, C |
| chr16_-_18904240 | 5.55 |
ENSMUST00000103746.3
|
Iglv1
|
immunoglobulin lambda variable 1 |
| chrX_+_106132055 | 5.42 |
ENSMUST00000150494.2
|
P2ry10
|
purinergic receptor P2Y, G-protein coupled 10 |
| chr4_+_15881256 | 4.87 |
ENSMUST00000029876.2
|
Calb1
|
calbindin 1 |
| chr2_+_164611812 | 4.02 |
ENSMUST00000088248.13
ENSMUST00000001439.7 |
Ube2c
|
ubiquitin-conjugating enzyme E2C |
| chr3_+_64884839 | 3.66 |
ENSMUST00000239069.2
|
Kcnab1
|
potassium voltage-gated channel, shaker-related subfamily, beta member 1 |
| chr17_+_45866618 | 3.54 |
ENSMUST00000024742.9
ENSMUST00000233929.2 |
Nfkbie
|
nuclear factor of kappa light polypeptide gene enhancer in B cells inhibitor, epsilon |
| chrX_+_56257374 | 3.42 |
ENSMUST00000033466.2
|
Cd40lg
|
CD40 ligand |
| chr8_+_31601837 | 3.00 |
ENSMUST00000046941.8
ENSMUST00000217278.2 |
Rnf122
|
ring finger protein 122 |
| chr8_-_106198112 | 2.57 |
ENSMUST00000014990.13
|
Tppp3
|
tubulin polymerization-promoting protein family member 3 |
| chr6_-_69245427 | 2.47 |
ENSMUST00000103348.3
|
Igkv4-70
|
immunoglobulin kappa chain variable 4-70 |
| chr10_-_117628565 | 2.45 |
ENSMUST00000167943.8
ENSMUST00000064848.7 |
Nup107
|
nucleoporin 107 |
| chr11_-_69686756 | 2.41 |
ENSMUST00000045971.9
|
Chrnb1
|
cholinergic receptor, nicotinic, beta polypeptide 1 (muscle) |
| chr3_+_94320548 | 2.19 |
ENSMUST00000166032.8
ENSMUST00000200486.5 ENSMUST00000196386.5 ENSMUST00000045245.10 ENSMUST00000197901.5 ENSMUST00000198041.2 |
Tdrkh
Gm42463
|
tudor and KH domain containing protein predicted gene 42463 |
| chr8_-_78337226 | 2.12 |
ENSMUST00000034030.15
|
Tmem184c
|
transmembrane protein 184C |
| chr9_-_56151334 | 1.92 |
ENSMUST00000188142.7
|
Peak1
|
pseudopodium-enriched atypical kinase 1 |
| chr3_-_79053182 | 1.80 |
ENSMUST00000118340.7
|
Rapgef2
|
Rap guanine nucleotide exchange factor (GEF) 2 |
| chr3_+_66892979 | 1.74 |
ENSMUST00000162362.8
ENSMUST00000065074.14 ENSMUST00000065047.13 |
Rsrc1
|
arginine/serine-rich coiled-coil 1 |
| chr3_+_68479578 | 1.74 |
ENSMUST00000170788.9
|
Schip1
|
schwannomin interacting protein 1 |
| chr6_+_142359099 | 1.69 |
ENSMUST00000126521.9
ENSMUST00000211094.2 |
Spx
|
spexin hormone |
| chr2_+_29236815 | 1.67 |
ENSMUST00000028139.11
ENSMUST00000113830.11 |
Med27
|
mediator complex subunit 27 |
| chr9_-_35481689 | 1.64 |
ENSMUST00000115110.5
|
Hyls1
|
HYLS1, centriolar and ciliogenesis associated |
| chr8_-_78337297 | 1.60 |
ENSMUST00000141202.2
ENSMUST00000152168.8 |
Tmem184c
|
transmembrane protein 184C |
| chr18_+_37864045 | 1.59 |
ENSMUST00000192535.2
|
Pcdhgb5
|
protocadherin gamma subfamily B, 5 |
| chr6_+_142244145 | 1.57 |
ENSMUST00000041993.3
|
Iapp
|
islet amyloid polypeptide |
| chr16_+_44167484 | 1.52 |
ENSMUST00000050897.7
|
Spice1
|
spindle and centriole associated protein 1 |
| chr6_+_86348286 | 1.51 |
ENSMUST00000089558.7
|
Snrpg
|
small nuclear ribonucleoprotein polypeptide G |
| chr2_-_88157559 | 1.50 |
ENSMUST00000214207.2
|
Olfr1175
|
olfactory receptor 1175 |
| chr5_-_123804745 | 1.49 |
ENSMUST00000149410.2
|
Clip1
|
CAP-GLY domain containing linker protein 1 |
| chr10_-_18103221 | 1.45 |
ENSMUST00000174592.8
|
Ccdc28a
|
coiled-coil domain containing 28A |
| chr12_-_119202527 | 1.35 |
ENSMUST00000026360.9
|
Itgb8
|
integrin beta 8 |
| chr3_-_100876960 | 1.32 |
ENSMUST00000076941.12
|
Ttf2
|
transcription termination factor, RNA polymerase II |
| chr7_+_112806672 | 1.30 |
ENSMUST00000047321.9
ENSMUST00000210074.2 ENSMUST00000210238.2 |
Arntl
|
aryl hydrocarbon receptor nuclear translocator-like |
| chr9_-_70048766 | 1.27 |
ENSMUST00000034749.16
|
Fam81a
|
family with sequence similarity 81, member A |
| chr15_+_34306812 | 1.26 |
ENSMUST00000226766.2
ENSMUST00000163455.9 ENSMUST00000022947.7 ENSMUST00000228570.2 ENSMUST00000227759.2 |
Matn2
|
matrilin 2 |
| chr9_+_74959259 | 1.21 |
ENSMUST00000170310.2
ENSMUST00000166549.2 |
Arpp19
|
cAMP-regulated phosphoprotein 19 |
| chr1_-_36312482 | 1.20 |
ENSMUST00000056946.8
|
Neurl3
|
neuralized E3 ubiquitin protein ligase 3 |
| chr1_-_179572765 | 1.18 |
ENSMUST00000211943.3
ENSMUST00000131716.4 ENSMUST00000221136.2 |
Kif28
|
kinesin family member 28 |
| chr11_-_72106418 | 1.17 |
ENSMUST00000021157.9
|
Med31
|
mediator complex subunit 31 |
| chr19_+_26725589 | 1.14 |
ENSMUST00000207812.2
ENSMUST00000175791.9 ENSMUST00000207118.2 ENSMUST00000209085.2 ENSMUST00000112637.10 ENSMUST00000207054.2 ENSMUST00000208589.2 ENSMUST00000176475.9 ENSMUST00000176698.9 ENSMUST00000207832.2 ENSMUST00000177252.9 ENSMUST00000208712.2 ENSMUST00000208186.2 ENSMUST00000208806.2 ENSMUST00000208027.2 |
Smarca2
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chr2_+_85809620 | 1.06 |
ENSMUST00000056849.3
|
Olfr1030
|
olfactory receptor 1030 |
| chr10_-_63926044 | 1.03 |
ENSMUST00000105439.2
|
Lrrtm3
|
leucine rich repeat transmembrane neuronal 3 |
| chr2_-_125624754 | 1.00 |
ENSMUST00000053699.13
|
Secisbp2l
|
SECIS binding protein 2-like |
| chr4_+_33062999 | 0.97 |
ENSMUST00000108162.8
ENSMUST00000024035.9 |
Gabrr2
|
gamma-aminobutyric acid (GABA) C receptor, subunit rho 2 |
| chr2_+_83642910 | 0.92 |
ENSMUST00000051454.4
|
Fam171b
|
family with sequence similarity 171, member B |
| chr11_-_65636651 | 0.88 |
ENSMUST00000138093.2
|
Map2k4
|
mitogen-activated protein kinase kinase 4 |
| chr11_-_96868483 | 0.84 |
ENSMUST00000107624.8
|
Sp2
|
Sp2 transcription factor |
| chrX_-_8118541 | 0.84 |
ENSMUST00000115594.8
ENSMUST00000115595.8 ENSMUST00000033513.10 |
Ftsj1
|
FtsJ RNA methyltransferase homolog 1 (E. coli) |
| chr5_+_88731386 | 0.84 |
ENSMUST00000031229.11
|
Rufy3
|
RUN and FYVE domain containing 3 |
| chr1_-_175319842 | 0.76 |
ENSMUST00000195324.6
ENSMUST00000192227.6 ENSMUST00000194555.6 |
Rgs7
|
regulator of G protein signaling 7 |
| chr15_-_43733389 | 0.72 |
ENSMUST00000067469.6
|
Tmem74
|
transmembrane protein 74 |
| chr5_+_88731366 | 0.72 |
ENSMUST00000199312.5
|
Rufy3
|
RUN and FYVE domain containing 3 |
| chrX_-_100266032 | 0.71 |
ENSMUST00000120389.8
ENSMUST00000156473.8 ENSMUST00000077876.4 |
Snx12
|
sorting nexin 12 |
| chr11_-_74615496 | 0.70 |
ENSMUST00000021091.15
|
Pafah1b1
|
platelet-activating factor acetylhydrolase, isoform 1b, subunit 1 |
| chr14_-_57353340 | 0.65 |
ENSMUST00000159455.2
|
Gm4491
|
predicted gene 4491 |
| chr6_-_3494587 | 0.62 |
ENSMUST00000049985.15
|
Hepacam2
|
HEPACAM family member 2 |
| chr6_-_69377328 | 0.52 |
ENSMUST00000198345.2
|
Igkv4-62
|
immunoglobulin kappa variable 4-62 |
| chr12_-_79211820 | 0.44 |
ENSMUST00000162569.8
|
Vti1b
|
vesicle transport through interaction with t-SNAREs 1B |
| chrX_+_37689503 | 0.42 |
ENSMUST00000000365.3
|
Mcts1
|
malignant T cell amplified sequence 1 |
| chr5_-_52723607 | 0.35 |
ENSMUST00000199942.5
|
Lgi2
|
leucine-rich repeat LGI family, member 2 |
| chr6_-_133032779 | 0.31 |
ENSMUST00000095391.3
|
Tas2r140
|
taste receptor, type 2, member 140 |
| chr7_-_115459082 | 0.28 |
ENSMUST00000206123.2
|
Sox6
|
SRY (sex determining region Y)-box 6 |
| chr11_+_49410475 | 0.25 |
ENSMUST00000204706.3
|
Olfr1383
|
olfactory receptor 1383 |
| chr16_+_58967409 | 0.25 |
ENSMUST00000216957.3
|
Olfr195
|
olfactory receptor 195 |
| chr2_-_166902307 | 0.25 |
ENSMUST00000155281.8
|
Znfx1
|
zinc finger, NFX1-type containing 1 |
| chr5_-_52723700 | 0.20 |
ENSMUST00000039750.7
|
Lgi2
|
leucine-rich repeat LGI family, member 2 |
| chr3_-_57202546 | 0.20 |
ENSMUST00000196506.2
|
Tm4sf1
|
transmembrane 4 superfamily member 1 |
| chr5_+_31684331 | 0.20 |
ENSMUST00000114533.9
ENSMUST00000202214.4 ENSMUST00000201858.4 ENSMUST00000202950.4 |
Slc4a1ap
|
solute carrier family 4 (anion exchanger), member 1, adaptor protein |
| chr10_-_58511476 | 0.19 |
ENSMUST00000003312.5
|
Edar
|
ectodysplasin-A receptor |
| chr5_-_31684036 | 0.17 |
ENSMUST00000202421.2
ENSMUST00000201769.4 ENSMUST00000065388.11 |
Supt7l
|
SPT7-like, STAGA complex gamma subunit |
| chr2_-_86640362 | 0.14 |
ENSMUST00000216117.2
|
Olfr141
|
olfactory receptor 141 |
| chr10_+_78816884 | 0.13 |
ENSMUST00000058991.5
ENSMUST00000203973.2 |
Olfr1352
|
olfactory receptor 1352 |
| chr2_+_111329683 | 0.12 |
ENSMUST00000219064.3
|
Olfr1291-ps1
|
olfactory receptor 1291, pseudogene 1 |
| chr10_-_38998272 | 0.11 |
ENSMUST00000136546.8
|
Fam229b
|
family with sequence similarity 229, member B |
| chr19_+_46385448 | 0.09 |
ENSMUST00000118440.3
|
Sufu
|
SUFU negative regulator of hedgehog signaling |
| chr17_-_37481536 | 0.09 |
ENSMUST00000174673.3
|
Olfr753-ps1
|
olfactory receptor 753, pseudogene 1 |
| chr19_-_6902698 | 0.02 |
ENSMUST00000237380.2
|
Catsperz
|
cation channel sperm associated auxiliary subunit zeta |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.2 | 12.8 | GO:2000471 | regulation of hematopoietic stem cell migration(GO:2000471) positive regulation of hematopoietic stem cell migration(GO:2000473) |
| 1.2 | 4.9 | GO:0035502 | metanephric part of ureteric bud development(GO:0035502) |
| 1.1 | 3.4 | GO:2001200 | positive regulation of dendritic cell differentiation(GO:2001200) |
| 0.9 | 3.5 | GO:0032329 | serine transport(GO:0032329) |
| 0.6 | 1.8 | GO:2000670 | positive regulation of dendritic cell apoptotic process(GO:2000670) |
| 0.5 | 4.0 | GO:0031536 | positive regulation of exit from mitosis(GO:0031536) |
| 0.5 | 2.5 | GO:0000973 | posttranscriptional tethering of RNA polymerase II gene DNA at nuclear periphery(GO:0000973) |
| 0.4 | 1.3 | GO:0031104 | dendrite regeneration(GO:0031104) |
| 0.3 | 1.5 | GO:0044861 | protein transport into plasma membrane raft(GO:0044861) |
| 0.3 | 1.3 | GO:0090403 | oxidative stress-induced premature senescence(GO:0090403) |
| 0.2 | 0.7 | GO:2000642 | negative regulation of early endosome to late endosome transport(GO:2000642) |
| 0.2 | 1.7 | GO:1904306 | positive regulation of gastro-intestinal system smooth muscle contraction(GO:1904306) |
| 0.2 | 1.6 | GO:0097646 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.2 | 2.4 | GO:0035095 | behavioral response to nicotine(GO:0035095) |
| 0.1 | 5.4 | GO:0051482 | positive regulation of cytosolic calcium ion concentration involved in phospholipase C-activating G-protein coupled signaling pathway(GO:0051482) |
| 0.1 | 1.1 | GO:0035887 | aortic smooth muscle cell differentiation(GO:0035887) |
| 0.1 | 3.7 | GO:1902260 | negative regulation of delayed rectifier potassium channel activity(GO:1902260) |
| 0.1 | 0.7 | GO:0051661 | maintenance of centrosome location(GO:0051661) |
| 0.1 | 1.0 | GO:0001514 | selenocysteine incorporation(GO:0001514) translational readthrough(GO:0006451) |
| 0.1 | 2.2 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 0.1 | 0.9 | GO:2000672 | negative regulation of motor neuron apoptotic process(GO:2000672) |
| 0.1 | 0.4 | GO:0002188 | translation reinitiation(GO:0002188) |
| 0.1 | 1.7 | GO:0001553 | luteinization(GO:0001553) |
| 0.1 | 1.5 | GO:0046599 | regulation of centriole replication(GO:0046599) |
| 0.1 | 1.4 | GO:0001573 | ganglioside metabolic process(GO:0001573) |
| 0.1 | 1.0 | GO:1902004 | positive regulation of beta-amyloid formation(GO:1902004) |
| 0.1 | 1.3 | GO:0006353 | DNA-templated transcription, termination(GO:0006353) |
| 0.0 | 1.6 | GO:2000114 | regulation of establishment of cell polarity(GO:2000114) |
| 0.0 | 1.2 | GO:0045722 | positive regulation of gluconeogenesis(GO:0045722) |
| 0.0 | 0.4 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.0 | 8.4 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.0 | 1.7 | GO:0046677 | response to antibiotic(GO:0046677) |
| 0.0 | 0.3 | GO:2000741 | positive regulation of mesenchymal stem cell differentiation(GO:2000741) |
| 0.0 | 0.8 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.0 | 0.2 | GO:0060662 | tube lumen cavitation(GO:0060605) salivary gland cavitation(GO:0060662) |
| 0.0 | 1.2 | GO:0048147 | negative regulation of fibroblast proliferation(GO:0048147) |
| 0.0 | 1.0 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.0 | 2.6 | GO:0046785 | microtubule polymerization(GO:0046785) |
| 0.0 | 0.1 | GO:0021776 | smoothened signaling pathway involved in ventral spinal cord interneuron specification(GO:0021775) smoothened signaling pathway involved in spinal cord motor neuron cell fate specification(GO:0021776) |
| 0.0 | 1.2 | GO:0072384 | organelle transport along microtubule(GO:0072384) |
| 0.0 | 0.2 | GO:0051457 | maintenance of protein location in nucleus(GO:0051457) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 2.2 | GO:0071547 | piP-body(GO:0071547) |
| 0.3 | 1.4 | GO:0034686 | integrin alphav-beta8 complex(GO:0034686) |
| 0.2 | 3.7 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.2 | 2.6 | GO:0097427 | microtubule bundle(GO:0097427) |
| 0.2 | 2.5 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.1 | 1.5 | GO:0044354 | pinosome(GO:0044352) macropinosome(GO:0044354) |
| 0.1 | 1.2 | GO:0070847 | core mediator complex(GO:0070847) |
| 0.1 | 1.5 | GO:0005687 | U4 snRNP(GO:0005687) |
| 0.1 | 4.0 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.1 | 2.4 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.1 | 0.7 | GO:0000235 | astral microtubule(GO:0000235) |
| 0.1 | 1.7 | GO:0031045 | dense core granule(GO:0031045) |
| 0.1 | 1.3 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.1 | 1.6 | GO:0071437 | invadopodium(GO:0071437) |
| 0.1 | 7.7 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 1.1 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.0 | 1.7 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 1.0 | GO:1902711 | GABA-A receptor complex(GO:1902711) |
| 0.0 | 0.8 | GO:0044292 | dendrite terminus(GO:0044292) |
| 0.0 | 4.9 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 16.2 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.0 | 0.9 | GO:0032839 | dendrite cytoplasm(GO:0032839) |
| 0.0 | 0.4 | GO:0012507 | ER to Golgi transport vesicle membrane(GO:0012507) |
| 0.0 | 3.5 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.0 | 1.2 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 1.5 | GO:0005814 | centriole(GO:0005814) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 3.4 | GO:0005174 | CD40 receptor binding(GO:0005174) |
| 0.6 | 4.9 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.4 | 12.8 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.3 | 5.4 | GO:0045028 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.1 | 0.9 | GO:0008545 | JUN kinase kinase activity(GO:0008545) |
| 0.1 | 3.7 | GO:0070402 | NADPH binding(GO:0070402) |
| 0.1 | 1.8 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) beta-1 adrenergic receptor binding(GO:0031697) |
| 0.1 | 1.4 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.1 | 2.4 | GO:0042166 | acetylcholine binding(GO:0042166) |
| 0.1 | 1.5 | GO:1990446 | U1 snRNP binding(GO:1990446) |
| 0.1 | 1.3 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.1 | 0.8 | GO:0016427 | tRNA (cytosine) methyltransferase activity(GO:0016427) |
| 0.1 | 1.0 | GO:0035368 | selenocysteine insertion sequence binding(GO:0035368) |
| 0.1 | 2.5 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.1 | 4.0 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.1 | 1.5 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.0 | 11.0 | GO:0003823 | antigen binding(GO:0003823) |
| 0.0 | 1.7 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.0 | 0.4 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) |
| 0.0 | 1.0 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.0 | 0.8 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 0.7 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.0 | 1.2 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.0 | 1.9 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 1.1 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) |
| 0.0 | 1.2 | GO:0001104 | RNA polymerase II transcription cofactor activity(GO:0001104) |
| 0.0 | 3.9 | GO:0061630 | ubiquitin protein ligase activity(GO:0061630) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 12.8 | SIG PIP3 SIGNALING IN B LYMPHOCYTES | Genes related to PIP3 signaling in B lymphocytes |
| 0.1 | 3.5 | ST GAQ PATHWAY | G alpha q Pathway |
| 0.1 | 3.4 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.1 | 1.4 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.1 | 1.3 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.1 | 0.9 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.0 | 1.3 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 1.1 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 12.8 | REACTOME PHOSPHORYLATION OF CD3 AND TCR ZETA CHAINS | Genes involved in Phosphorylation of CD3 and TCR zeta chains |
| 0.3 | 5.4 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.2 | 4.0 | REACTOME PHOSPHORYLATION OF THE APC C | Genes involved in Phosphorylation of the APC/C |
| 0.1 | 1.5 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.1 | 2.5 | REACTOME TRANSPORT OF RIBONUCLEOPROTEINS INTO THE HOST NUCLEUS | Genes involved in Transport of Ribonucleoproteins into the Host Nucleus |
| 0.1 | 3.4 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.1 | 3.7 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 1.6 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 3.5 | REACTOME ACTIVATION OF NF KAPPAB IN B CELLS | Genes involved in Activation of NF-kappaB in B Cells |
| 0.0 | 0.9 | REACTOME JNK C JUN KINASES PHOSPHORYLATION AND ACTIVATION MEDIATED BY ACTIVATED HUMAN TAK1 | Genes involved in JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 |
| 0.0 | 1.3 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.0 | 2.8 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 1.0 | REACTOME LIGAND GATED ION CHANNEL TRANSPORT | Genes involved in Ligand-gated ion channel transport |
| 0.0 | 2.2 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.0 | 1.4 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |