Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
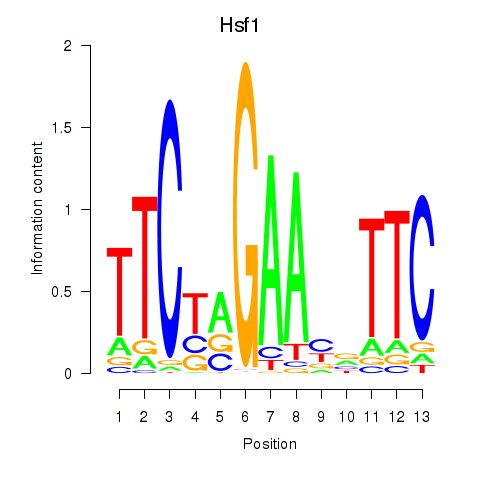
Results for Hsf1
Z-value: 0.93
Transcription factors associated with Hsf1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Hsf1
|
ENSMUSG00000022556.12 | Hsf1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Hsf1 | mm39_v1_chr15_+_76361604_76361774 | 0.42 | 2.2e-04 | Click! |
Activity profile of Hsf1 motif
Sorted Z-values of Hsf1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Hsf1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_+_157401998 | 7.36 |
ENSMUST00000153739.9
ENSMUST00000173595.2 ENSMUST00000109526.2 ENSMUST00000173839.2 ENSMUST00000173041.8 ENSMUST00000173793.8 ENSMUST00000172487.2 ENSMUST00000088484.6 |
Nnat
|
neuronatin |
| chr12_-_72283465 | 6.23 |
ENSMUST00000021497.16
ENSMUST00000137990.2 |
Rtn1
|
reticulon 1 |
| chr17_-_45883421 | 5.14 |
ENSMUST00000130406.2
|
Hsp90ab1
|
heat shock protein 90 alpha (cytosolic), class B member 1 |
| chr1_+_66360865 | 4.58 |
ENSMUST00000114013.8
|
Map2
|
microtubule-associated protein 2 |
| chr5_-_149559636 | 4.58 |
ENSMUST00000201452.4
|
Hsph1
|
heat shock 105kDa/110kDa protein 1 |
| chr10_-_80679859 | 4.57 |
ENSMUST00000053986.9
|
Lingo3
|
leucine rich repeat and Ig domain containing 3 |
| chrX_-_149440388 | 4.51 |
ENSMUST00000151403.9
ENSMUST00000087253.11 ENSMUST00000112709.8 ENSMUST00000163969.8 ENSMUST00000087258.10 |
Tro
|
trophinin |
| chr9_+_50664207 | 4.46 |
ENSMUST00000034562.9
|
Cryab
|
crystallin, alpha B |
| chrX_-_60229164 | 4.43 |
ENSMUST00000166381.3
|
Cdr1
|
cerebellar degeneration related antigen 1 |
| chr9_+_50664288 | 4.36 |
ENSMUST00000214962.2
ENSMUST00000216755.2 |
Cryab
|
crystallin, alpha B |
| chr18_+_67266054 | 4.26 |
ENSMUST00000236771.2
ENSMUST00000237304.2 ENSMUST00000076605.9 |
Gnal
|
guanine nucleotide binding protein, alpha stimulating, olfactory type |
| chr9_-_58448224 | 3.88 |
ENSMUST00000039788.11
|
Cd276
|
CD276 antigen |
| chr4_+_43414696 | 3.72 |
ENSMUST00000131668.3
|
Rusc2
|
RUN and SH3 domain containing 2 |
| chr9_+_18203558 | 3.53 |
ENSMUST00000213605.2
|
Chordc1
|
cysteine and histidine-rich domain (CHORD)-containing, zinc-binding protein 1 |
| chr5_+_66833434 | 3.52 |
ENSMUST00000031131.11
|
Uchl1
|
ubiquitin carboxy-terminal hydrolase L1 |
| chr15_-_58828321 | 3.50 |
ENSMUST00000228067.2
|
Mtss1
|
MTSS I-BAR domain containing 1 |
| chr18_+_43897354 | 3.32 |
ENSMUST00000187157.7
ENSMUST00000043803.13 ENSMUST00000189750.2 |
Scgb3a2
|
secretoglobin, family 3A, member 2 |
| chr5_-_139470169 | 3.27 |
ENSMUST00000150992.2
ENSMUST00000110851.8 ENSMUST00000079996.13 |
Zfand2a
|
zinc finger, AN1-type domain 2A |
| chr16_-_20245071 | 3.25 |
ENSMUST00000115547.9
ENSMUST00000096199.5 |
Abcc5
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 5 |
| chr9_+_18203640 | 3.19 |
ENSMUST00000217031.2
|
Chordc1
|
cysteine and histidine-rich domain (CHORD)-containing, zinc-binding protein 1 |
| chr7_-_110462446 | 3.10 |
ENSMUST00000033050.5
|
Lyve1
|
lymphatic vessel endothelial hyaluronan receptor 1 |
| chr5_-_129864202 | 3.08 |
ENSMUST00000136507.4
|
Psph
|
phosphoserine phosphatase |
| chr9_-_110818679 | 3.03 |
ENSMUST00000084922.6
ENSMUST00000199891.2 |
Rtp3
|
receptor transporter protein 3 |
| chr5_-_149559737 | 3.02 |
ENSMUST00000200805.4
|
Hsph1
|
heat shock 105kDa/110kDa protein 1 |
| chr1_-_55127312 | 2.81 |
ENSMUST00000127861.8
ENSMUST00000144077.3 |
Hspd1
|
heat shock protein 1 (chaperonin) |
| chr4_-_136620376 | 2.78 |
ENSMUST00000046332.6
|
C1qc
|
complement component 1, q subcomponent, C chain |
| chr19_-_7017295 | 2.75 |
ENSMUST00000025918.9
|
Stip1
|
stress-induced phosphoprotein 1 |
| chr16_-_20245138 | 2.68 |
ENSMUST00000079158.13
|
Abcc5
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 5 |
| chrX_-_149440362 | 2.59 |
ENSMUST00000148604.2
|
Tro
|
trophinin |
| chr5_-_149559792 | 2.58 |
ENSMUST00000202361.4
ENSMUST00000202089.4 ENSMUST00000200825.2 ENSMUST00000201559.4 |
Hsph1
|
heat shock 105kDa/110kDa protein 1 |
| chr14_+_66205932 | 2.51 |
ENSMUST00000022616.14
|
Clu
|
clusterin |
| chr7_-_89590404 | 2.50 |
ENSMUST00000153470.9
|
Hikeshi
|
heat shock protein nuclear import factor |
| chr4_+_42949814 | 2.48 |
ENSMUST00000037872.10
ENSMUST00000098112.9 |
Dnajb5
|
DnaJ heat shock protein family (Hsp40) member B5 |
| chr5_-_123887434 | 2.42 |
ENSMUST00000182955.8
ENSMUST00000182489.8 ENSMUST00000183147.9 ENSMUST00000050827.14 ENSMUST00000057795.12 ENSMUST00000111515.8 ENSMUST00000182309.8 |
Rsrc2
|
arginine/serine-rich coiled-coil 2 |
| chr18_-_10706701 | 2.40 |
ENSMUST00000002549.9
ENSMUST00000117726.9 ENSMUST00000117828.9 |
Abhd3
|
abhydrolase domain containing 3 |
| chr8_-_45835234 | 2.36 |
ENSMUST00000093526.13
|
Fam149a
|
family with sequence similarity 149, member A |
| chr14_-_63430937 | 2.34 |
ENSMUST00000225157.2
ENSMUST00000226002.2 ENSMUST00000038229.5 |
Neil2
|
nei like 2 (E. coli) |
| chr12_-_110662256 | 2.31 |
ENSMUST00000149189.2
|
Hsp90aa1
|
heat shock protein 90, alpha (cytosolic), class A member 1 |
| chr4_+_155686311 | 2.27 |
ENSMUST00000118607.2
|
Slc35e2
|
solute carrier family 35, member E2 |
| chr6_+_51447317 | 2.26 |
ENSMUST00000094623.10
|
Cbx3
|
chromobox 3 |
| chr7_-_19043955 | 2.26 |
ENSMUST00000207334.2
ENSMUST00000208505.2 ENSMUST00000207716.2 ENSMUST00000208326.2 ENSMUST00000003640.4 |
Fosb
|
FBJ osteosarcoma oncogene B |
| chr11_+_6510167 | 2.25 |
ENSMUST00000109722.9
|
Ccm2
|
cerebral cavernous malformation 2 |
| chr7_-_89590230 | 2.18 |
ENSMUST00000075010.12
|
Hikeshi
|
heat shock protein nuclear import factor |
| chr7_+_140425460 | 2.17 |
ENSMUST00000035300.7
|
Scgb1c1
|
secretoglobin, family 1C, member 1 |
| chr5_+_129864044 | 2.17 |
ENSMUST00000201414.5
|
Cct6a
|
chaperonin containing Tcp1, subunit 6a (zeta) |
| chr2_+_109523908 | 2.16 |
ENSMUST00000111045.9
|
Bdnf
|
brain derived neurotrophic factor |
| chr3_+_117368483 | 2.10 |
ENSMUST00000039564.11
ENSMUST00000238937.2 |
Plppr5
|
phospholipid phosphatase related 5 |
| chr9_+_18203421 | 2.06 |
ENSMUST00000001825.9
|
Chordc1
|
cysteine and histidine-rich domain (CHORD)-containing, zinc-binding protein 1 |
| chr18_+_37637317 | 2.06 |
ENSMUST00000052179.8
|
Pcdhb20
|
protocadherin beta 20 |
| chr11_-_116226175 | 2.03 |
ENSMUST00000036215.8
|
Foxj1
|
forkhead box J1 |
| chr10_+_59159118 | 2.02 |
ENSMUST00000009789.15
ENSMUST00000092512.11 ENSMUST00000105466.3 |
P4ha1
|
procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), alpha 1 polypeptide |
| chr17_-_50401305 | 2.02 |
ENSMUST00000113195.8
|
Rftn1
|
raftlin lipid raft linker 1 |
| chr7_-_89590334 | 2.00 |
ENSMUST00000207309.2
ENSMUST00000130609.3 |
Hikeshi
|
heat shock protein nuclear import factor |
| chr8_+_88999031 | 1.97 |
ENSMUST00000169037.9
|
Adcy7
|
adenylate cyclase 7 |
| chr16_+_22926504 | 1.96 |
ENSMUST00000187168.7
ENSMUST00000232287.2 ENSMUST00000077605.12 |
Eif4a2
|
eukaryotic translation initiation factor 4A2 |
| chr8_+_36956345 | 1.91 |
ENSMUST00000171777.2
|
Trmt9b
|
tRNA methyltransferase 9B |
| chr3_+_79498663 | 1.89 |
ENSMUST00000029382.13
|
Ppid
|
peptidylprolyl isomerase D (cyclophilin D) |
| chr4_+_155685830 | 1.86 |
ENSMUST00000105608.9
|
Slc35e2
|
solute carrier family 35, member E2 |
| chr1_+_179788675 | 1.85 |
ENSMUST00000076687.12
ENSMUST00000097450.10 ENSMUST00000212756.2 |
Cdc42bpa
|
CDC42 binding protein kinase alpha |
| chr5_+_30486375 | 1.82 |
ENSMUST00000101448.5
|
Drc1
|
dynein regulatory complex subunit 1 |
| chr15_+_81284333 | 1.77 |
ENSMUST00000163754.9
ENSMUST00000041609.11 |
Xpnpep3
|
X-prolyl aminopeptidase 3, mitochondrial |
| chr5_-_149559667 | 1.75 |
ENSMUST00000074846.14
|
Hsph1
|
heat shock 105kDa/110kDa protein 1 |
| chr7_-_46365108 | 1.75 |
ENSMUST00000006956.9
ENSMUST00000210913.2 |
Saa3
|
serum amyloid A 3 |
| chr9_-_53617508 | 1.66 |
ENSMUST00000068449.4
|
Rab39
|
RAB39, member RAS oncogene family |
| chr16_+_22926162 | 1.65 |
ENSMUST00000023599.13
ENSMUST00000168891.8 |
Eif4a2
|
eukaryotic translation initiation factor 4A2 |
| chr4_+_152262583 | 1.64 |
ENSMUST00000075363.10
|
Acot7
|
acyl-CoA thioesterase 7 |
| chr1_+_57813759 | 1.60 |
ENSMUST00000167971.8
ENSMUST00000170139.8 ENSMUST00000171699.8 ENSMUST00000164302.8 |
Spats2l
|
spermatogenesis associated, serine-rich 2-like |
| chr9_+_92339422 | 1.58 |
ENSMUST00000034941.9
|
Plscr4
|
phospholipid scramblase 4 |
| chr1_+_66361252 | 1.58 |
ENSMUST00000123647.8
|
Map2
|
microtubule-associated protein 2 |
| chr16_-_55755208 | 1.55 |
ENSMUST00000121129.8
ENSMUST00000023270.14 |
Cep97
|
centrosomal protein 97 |
| chr9_+_40712562 | 1.55 |
ENSMUST00000117557.8
|
Hspa8
|
heat shock protein 8 |
| chr6_+_58808733 | 1.53 |
ENSMUST00000126292.8
ENSMUST00000031823.12 |
Herc3
|
hect domain and RLD 3 |
| chr7_-_99002430 | 1.51 |
ENSMUST00000094154.6
|
Serpinh1
|
serine (or cysteine) peptidase inhibitor, clade H, member 1 |
| chrX_-_73067514 | 1.50 |
ENSMUST00000033769.15
ENSMUST00000114352.8 ENSMUST00000068286.12 ENSMUST00000114360.10 ENSMUST00000114354.10 |
Irak1
|
interleukin-1 receptor-associated kinase 1 |
| chr19_-_29790352 | 1.49 |
ENSMUST00000099525.5
|
Ranbp6
|
RAN binding protein 6 |
| chrX_-_76968668 | 1.43 |
ENSMUST00000063127.4
|
5430402E10Rik
|
RIKEN cDNA 5430402E10 gene |
| chr10_-_61219229 | 1.43 |
ENSMUST00000020289.10
|
Pald1
|
phosphatase domain containing, paladin 1 |
| chr4_+_17853452 | 1.36 |
ENSMUST00000029881.10
|
Mmp16
|
matrix metallopeptidase 16 |
| chr14_+_34097474 | 1.36 |
ENSMUST00000227130.2
|
Mmrn2
|
multimerin 2 |
| chr9_+_22322802 | 1.33 |
ENSMUST00000058868.9
|
9530077C05Rik
|
RIKEN cDNA 9530077C05 gene |
| chr4_+_40722461 | 1.31 |
ENSMUST00000030118.10
|
Dnaja1
|
DnaJ heat shock protein family (Hsp40) member A1 |
| chr15_-_81283795 | 1.29 |
ENSMUST00000023039.15
|
St13
|
suppression of tumorigenicity 13 |
| chr11_-_5049082 | 1.23 |
ENSMUST00000063232.7
|
Ewsr1
|
Ewing sarcoma breakpoint region 1 |
| chr8_-_49009043 | 1.21 |
ENSMUST00000110343.3
|
Tenm3
|
teneurin transmembrane protein 3 |
| chr5_+_29940935 | 1.17 |
ENSMUST00000114839.8
ENSMUST00000198694.5 ENSMUST00000012734.10 ENSMUST00000196528.5 |
Dnajb6
|
DnaJ heat shock protein family (Hsp40) member B6 |
| chr11_-_23448030 | 1.13 |
ENSMUST00000020529.13
|
Ahsa2
|
AHA1, activator of heat shock protein ATPase 2 |
| chr11_-_23447866 | 1.10 |
ENSMUST00000128559.2
ENSMUST00000147157.8 ENSMUST00000109539.8 |
Ahsa2
|
AHA1, activator of heat shock protein ATPase 2 |
| chr11_+_82802079 | 1.10 |
ENSMUST00000018989.14
ENSMUST00000164945.3 |
Unc45b
|
unc-45 myosin chaperone B |
| chr3_-_106454898 | 1.07 |
ENSMUST00000121231.8
ENSMUST00000141525.2 |
Cept1
|
choline/ethanolaminephosphotransferase 1 |
| chr12_-_110662677 | 1.04 |
ENSMUST00000124156.8
|
Hsp90aa1
|
heat shock protein 90, alpha (cytosolic), class A member 1 |
| chr15_+_28203872 | 1.04 |
ENSMUST00000067048.8
|
Dnah5
|
dynein, axonemal, heavy chain 5 |
| chrX_-_73067351 | 1.03 |
ENSMUST00000114353.10
ENSMUST00000101458.9 |
Irak1
|
interleukin-1 receptor-associated kinase 1 |
| chr3_+_117368876 | 1.01 |
ENSMUST00000106473.5
|
Plppr5
|
phospholipid phosphatase related 5 |
| chr7_-_99002204 | 1.00 |
ENSMUST00000208292.2
ENSMUST00000207989.2 ENSMUST00000208749.2 ENSMUST00000169437.9 ENSMUST00000208119.2 ENSMUST00000207849.2 |
Serpinh1
|
serine (or cysteine) peptidase inhibitor, clade H, member 1 |
| chr11_+_70591299 | 0.98 |
ENSMUST00000152618.9
ENSMUST00000102554.8 ENSMUST00000094499.11 ENSMUST00000072187.12 ENSMUST00000137119.3 |
Kif1c
|
kinesin family member 1C |
| chr15_-_31601652 | 0.98 |
ENSMUST00000161266.2
|
Cct5
|
chaperonin containing Tcp1, subunit 5 (epsilon) |
| chr9_+_22322875 | 0.98 |
ENSMUST00000214436.2
|
9530077C05Rik
|
RIKEN cDNA 9530077C05 gene |
| chr3_-_89309944 | 0.97 |
ENSMUST00000057431.6
|
Lenep
|
lens epithelial protein |
| chr17_+_15261896 | 0.95 |
ENSMUST00000226599.2
ENSMUST00000228518.2 ENSMUST00000226213.2 |
Ermard
|
ER membrane associated RNA degradation |
| chr7_-_19043525 | 0.94 |
ENSMUST00000208446.2
|
Fosb
|
FBJ osteosarcoma oncogene B |
| chr4_+_40722911 | 0.90 |
ENSMUST00000164233.8
ENSMUST00000137246.8 ENSMUST00000125442.8 |
Dnaja1
|
DnaJ heat shock protein family (Hsp40) member A1 |
| chr18_-_36899245 | 0.84 |
ENSMUST00000061522.8
|
Dnd1
|
DND microRNA-mediated repression inhibitor 1 |
| chr11_-_48836975 | 0.84 |
ENSMUST00000104958.2
|
Psme2b
|
protease (prosome, macropain) activator subunit 2B |
| chr16_-_87292592 | 0.82 |
ENSMUST00000176750.2
ENSMUST00000175977.8 |
Cct8
|
chaperonin containing Tcp1, subunit 8 (theta) |
| chr16_-_87292711 | 0.75 |
ENSMUST00000176041.8
ENSMUST00000026704.14 |
Cct8
|
chaperonin containing Tcp1, subunit 8 (theta) |
| chr5_-_86616849 | 0.75 |
ENSMUST00000101073.3
|
Tmprss11a
|
transmembrane protease, serine 11a |
| chr4_-_133273243 | 0.74 |
ENSMUST00000030665.7
|
Nudc
|
nudC nuclear distribution protein |
| chr11_+_83599841 | 0.71 |
ENSMUST00000001009.14
|
Wfdc18
|
WAP four-disulfide core domain 18 |
| chr15_+_31602252 | 0.71 |
ENSMUST00000042702.7
ENSMUST00000161061.3 |
Atpsckmt
|
ATP synthase C subunit lysine N-methyltransferase |
| chr12_+_75355082 | 0.69 |
ENSMUST00000118602.8
ENSMUST00000118966.8 ENSMUST00000055390.6 |
Rhoj
|
ras homolog family member J |
| chr16_-_55755160 | 0.67 |
ENSMUST00000122280.8
ENSMUST00000121703.3 |
Cep97
|
centrosomal protein 97 |
| chr1_-_55127183 | 0.66 |
ENSMUST00000027123.15
|
Hspd1
|
heat shock protein 1 (chaperonin) |
| chr5_-_36853281 | 0.62 |
ENSMUST00000031091.13
ENSMUST00000140653.2 |
D5Ertd579e
|
DNA segment, Chr 5, ERATO Doi 579, expressed |
| chr7_+_121666388 | 0.61 |
ENSMUST00000033158.6
|
Ubfd1
|
ubiquitin family domain containing 1 |
| chr2_+_29014106 | 0.58 |
ENSMUST00000129544.8
|
Setx
|
senataxin |
| chr6_-_136918885 | 0.57 |
ENSMUST00000111891.4
|
Arhgdib
|
Rho, GDP dissociation inhibitor (GDI) beta |
| chr6_-_72416531 | 0.54 |
ENSMUST00000205335.2
ENSMUST00000206692.2 ENSMUST00000059472.10 |
Mat2a
|
methionine adenosyltransferase II, alpha |
| chr8_-_70411054 | 0.53 |
ENSMUST00000211960.2
|
Gatad2a
|
GATA zinc finger domain containing 2A |
| chr11_+_49500090 | 0.53 |
ENSMUST00000020617.3
|
Flt4
|
FMS-like tyrosine kinase 4 |
| chr15_-_100301124 | 0.50 |
ENSMUST00000124324.2
|
Slc11a2
|
solute carrier family 11 (proton-coupled divalent metal ion transporters), member 2 |
| chr17_-_47321430 | 0.46 |
ENSMUST00000113337.10
ENSMUST00000113335.4 |
Ubr2
|
ubiquitin protein ligase E3 component n-recognin 2 |
| chr5_+_135916764 | 0.46 |
ENSMUST00000005077.7
|
Hspb1
|
heat shock protein 1 |
| chr6_-_128803182 | 0.45 |
ENSMUST00000204756.3
ENSMUST00000204394.3 ENSMUST00000204423.3 ENSMUST00000204677.2 ENSMUST00000205130.3 ENSMUST00000174544.2 ENSMUST00000172887.8 ENSMUST00000032472.11 |
Gm44511
Klrb1b
|
predicted gene 44511 killer cell lectin-like receptor subfamily B member 1B |
| chr8_-_110419867 | 0.45 |
ENSMUST00000034164.6
|
Ist1
|
increased sodium tolerance 1 homolog (yeast) |
| chr1_-_160040286 | 0.44 |
ENSMUST00000195654.2
ENSMUST00000014370.11 |
Cacybp
|
calcyclin binding protein |
| chr19_+_13594739 | 0.43 |
ENSMUST00000217061.3
ENSMUST00000209005.4 ENSMUST00000208347.3 |
Olfr1487
|
olfactory receptor 1487 |
| chr9_+_119939414 | 0.38 |
ENSMUST00000035106.12
|
Slc25a38
|
solute carrier family 25, member 38 |
| chr7_+_110628158 | 0.38 |
ENSMUST00000005749.6
|
Ctr9
|
CTR9 homolog, Paf1/RNA polymerase II complex component |
| chr5_+_135916847 | 0.37 |
ENSMUST00000111155.2
|
Hspb1
|
heat shock protein 1 |
| chr9_-_103357564 | 0.35 |
ENSMUST00000124310.5
|
Bfsp2
|
beaded filament structural protein 2, phakinin |
| chr15_-_81284244 | 0.34 |
ENSMUST00000172107.8
ENSMUST00000169204.2 ENSMUST00000163382.2 |
St13
|
suppression of tumorigenicity 13 |
| chr7_-_19621833 | 0.29 |
ENSMUST00000052605.8
|
Ceacam19
|
carcinoembryonic antigen-related cell adhesion molecule 19 |
| chr15_-_31601932 | 0.29 |
ENSMUST00000022842.16
|
Cct5
|
chaperonin containing Tcp1, subunit 5 (epsilon) |
| chr12_+_24881582 | 0.25 |
ENSMUST00000221952.2
ENSMUST00000078902.8 ENSMUST00000110942.11 |
Mboat2
|
membrane bound O-acyltransferase domain containing 2 |
| chr12_-_104718159 | 0.25 |
ENSMUST00000041987.7
|
Dicer1
|
dicer 1, ribonuclease type III |
| chr1_+_57814001 | 0.25 |
ENSMUST00000167085.8
|
Spats2l
|
spermatogenesis associated, serine-rich 2-like |
| chr11_-_106469938 | 0.23 |
ENSMUST00000103070.3
|
Tex2
|
testis expressed gene 2 |
| chr8_-_49008881 | 0.23 |
ENSMUST00000110345.8
|
Tenm3
|
teneurin transmembrane protein 3 |
| chr17_+_15261470 | 0.23 |
ENSMUST00000097393.11
|
Ermard
|
ER membrane associated RNA degradation |
| chr2_+_72128239 | 0.19 |
ENSMUST00000144111.2
|
Map3k20
|
mitogen-activated protein kinase kinase kinase 20 |
| chr6_-_51446850 | 0.19 |
ENSMUST00000069949.13
|
Hnrnpa2b1
|
heterogeneous nuclear ribonucleoprotein A2/B1 |
| chr7_+_35285657 | 0.19 |
ENSMUST00000040844.16
ENSMUST00000188906.7 ENSMUST00000186245.7 ENSMUST00000190503.7 |
Ankrd27
|
ankyrin repeat domain 27 (VPS9 domain) |
| chr17_+_15230438 | 0.18 |
ENSMUST00000097395.5
|
Gm3435
|
predicted gene 3435 |
| chr6_-_51446752 | 0.17 |
ENSMUST00000204188.3
ENSMUST00000203220.3 ENSMUST00000114459.8 ENSMUST00000090002.10 |
Hnrnpa2b1
|
heterogeneous nuclear ribonucleoprotein A2/B1 |
| chr9_-_80347482 | 0.15 |
ENSMUST00000085289.12
ENSMUST00000185068.2 ENSMUST00000113250.10 |
Impg1
|
interphotoreceptor matrix proteoglycan 1 |
| chrX_+_76908364 | 0.15 |
ENSMUST00000114039.3
|
Gm14744
|
predicted gene 14744 |
| chr9_-_105372235 | 0.11 |
ENSMUST00000176190.8
ENSMUST00000163879.9 ENSMUST00000112558.10 ENSMUST00000176363.9 |
Atp2c1
|
ATPase, Ca++-sequestering |
| chr11_-_5049223 | 0.10 |
ENSMUST00000079949.13
|
Ewsr1
|
Ewing sarcoma breakpoint region 1 |
| chr7_+_92210348 | 0.09 |
ENSMUST00000032842.13
ENSMUST00000085017.5 |
Ccdc90b
|
coiled-coil domain containing 90B |
| chr3_+_93052089 | 0.08 |
ENSMUST00000107300.7
ENSMUST00000195515.2 |
Crnn
|
cornulin |
| chr13_+_44883270 | 0.07 |
ENSMUST00000172830.8
|
Jarid2
|
jumonji, AT rich interactive domain 2 |
| chr10_-_86541349 | 0.05 |
ENSMUST00000020238.14
|
Hsp90b1
|
heat shock protein 90, beta (Grp94), member 1 |
| chr17_+_15230597 | 0.05 |
ENSMUST00000232446.2
|
Gm3435
|
predicted gene 3435 |
| chr11_-_5049036 | 0.03 |
ENSMUST00000102930.10
ENSMUST00000093365.12 ENSMUST00000073308.11 |
Ewsr1
|
Ewing sarcoma breakpoint region 1 |
| chr17_+_48606948 | 0.02 |
ENSMUST00000233092.2
|
Treml2
|
triggering receptor expressed on myeloid cells-like 2 |
| chrX_-_72759748 | 0.02 |
ENSMUST00000002091.6
|
Bcap31
|
B cell receptor associated protein 31 |
| chr7_-_121666486 | 0.02 |
ENSMUST00000033159.4
|
Ears2
|
glutamyl-tRNA synthetase 2, mitochondrial |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.9 | 8.8 | GO:2000299 | negative regulation of Rho-dependent protein serine/threonine kinase activity(GO:2000299) |
| 1.7 | 5.1 | GO:0036363 | transforming growth factor beta activation(GO:0036363) |
| 1.2 | 11.9 | GO:1903753 | negative regulation of p38MAPK cascade(GO:1903753) |
| 1.2 | 3.5 | GO:0045041 | protein import into mitochondrial intermembrane space(GO:0045041) |
| 0.9 | 5.7 | GO:0007412 | axon target recognition(GO:0007412) |
| 0.8 | 8.8 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 0.7 | 2.0 | GO:0002514 | B cell tolerance induction(GO:0002514) regulation of B cell tolerance induction(GO:0002661) positive regulation of B cell tolerance induction(GO:0002663) |
| 0.6 | 3.5 | GO:0034334 | adherens junction maintenance(GO:0034334) |
| 0.6 | 3.9 | GO:0045077 | negative regulation of interferon-gamma biosynthetic process(GO:0045077) |
| 0.5 | 4.3 | GO:0009405 | pathogenesis(GO:0009405) |
| 0.5 | 1.5 | GO:0097212 | protein targeting to vacuole involved in autophagy(GO:0071211) lysosomal membrane organization(GO:0097212) positive regulation of protein folding(GO:1903334) |
| 0.4 | 3.0 | GO:0045585 | regulation of cytotoxic T cell differentiation(GO:0045583) positive regulation of cytotoxic T cell differentiation(GO:0045585) |
| 0.4 | 2.5 | GO:1902998 | regulation of neuronal signal transduction(GO:1902847) positive regulation of neurofibrillary tangle assembly(GO:1902998) |
| 0.4 | 1.6 | GO:1900533 | medium-chain fatty-acyl-CoA catabolic process(GO:0036114) long-chain fatty-acyl-CoA catabolic process(GO:0036116) palmitic acid metabolic process(GO:1900533) palmitic acid biosynthetic process(GO:1900535) |
| 0.4 | 3.1 | GO:0006564 | L-serine biosynthetic process(GO:0006564) |
| 0.4 | 1.1 | GO:0060715 | syncytiotrophoblast cell differentiation involved in labyrinthine layer development(GO:0060715) |
| 0.3 | 2.5 | GO:0090370 | negative regulation of cholesterol efflux(GO:0090370) |
| 0.3 | 2.8 | GO:0045650 | negative regulation of macrophage differentiation(GO:0045650) |
| 0.3 | 4.5 | GO:1904871 | protein localization to nuclear body(GO:1903405) protein localization to Cajal body(GO:1904867) regulation of protein localization to Cajal body(GO:1904869) positive regulation of protein localization to Cajal body(GO:1904871) |
| 0.3 | 2.5 | GO:0003433 | chondrocyte development involved in endochondral bone morphogenesis(GO:0003433) |
| 0.3 | 1.9 | GO:0071492 | cellular response to UV-A(GO:0071492) |
| 0.3 | 2.0 | GO:0002457 | T cell antigen processing and presentation(GO:0002457) |
| 0.2 | 2.2 | GO:0042769 | DNA damage response, detection of DNA damage(GO:0042769) |
| 0.2 | 1.7 | GO:0090383 | phagosome acidification(GO:0090383) |
| 0.2 | 1.6 | GO:0061083 | regulation of protein refolding(GO:0061083) negative regulation of protein refolding(GO:0061084) |
| 0.2 | 2.0 | GO:0018401 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) |
| 0.2 | 0.6 | GO:0031554 | regulation of DNA-templated transcription, termination(GO:0031554) |
| 0.2 | 3.3 | GO:0071243 | cellular response to arsenic-containing substance(GO:0071243) |
| 0.2 | 2.3 | GO:0060837 | blood vessel endothelial cell differentiation(GO:0060837) |
| 0.1 | 3.1 | GO:0006027 | glycosaminoglycan catabolic process(GO:0006027) |
| 0.1 | 1.5 | GO:0006610 | ribosomal protein import into nucleus(GO:0006610) |
| 0.1 | 2.0 | GO:0071361 | cellular response to ethanol(GO:0071361) |
| 0.1 | 2.2 | GO:1902018 | negative regulation of cilium assembly(GO:1902018) |
| 0.1 | 0.4 | GO:2001160 | regulation of histone H3-K79 methylation(GO:2001160) |
| 0.1 | 0.4 | GO:1990428 | miRNA transport(GO:1990428) |
| 0.1 | 6.2 | GO:0007026 | negative regulation of microtubule depolymerization(GO:0007026) |
| 0.1 | 0.5 | GO:0071596 | ubiquitin-dependent protein catabolic process via the N-end rule pathway(GO:0071596) |
| 0.1 | 0.5 | GO:0009838 | abscission(GO:0009838) |
| 0.1 | 0.8 | GO:0038033 | positive regulation of endothelial cell chemotaxis by VEGF-activated vascular endothelial growth factor receptor signaling pathway(GO:0038033) |
| 0.1 | 2.9 | GO:0070286 | axonemal dynein complex assembly(GO:0070286) |
| 0.1 | 0.4 | GO:0036233 | glycine import(GO:0036233) |
| 0.1 | 6.7 | GO:0034605 | cellular response to heat(GO:0034605) |
| 0.1 | 0.5 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 0.1 | 1.4 | GO:0030948 | negative regulation of vascular endothelial growth factor receptor signaling pathway(GO:0030948) |
| 0.1 | 1.7 | GO:0007252 | I-kappaB phosphorylation(GO:0007252) |
| 0.1 | 0.5 | GO:0015676 | vanadium ion transport(GO:0015676) lead ion transport(GO:0015692) |
| 0.1 | 3.0 | GO:0051205 | protein insertion into membrane(GO:0051205) |
| 0.1 | 2.6 | GO:0007097 | nuclear migration(GO:0007097) establishment of nucleus localization(GO:0040023) |
| 0.1 | 1.4 | GO:0035988 | chondrocyte proliferation(GO:0035988) |
| 0.1 | 3.6 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.1 | 3.5 | GO:0046470 | phosphatidylcholine metabolic process(GO:0046470) |
| 0.1 | 0.6 | GO:1901164 | negative regulation of trophoblast cell migration(GO:1901164) |
| 0.1 | 0.2 | GO:0035544 | negative regulation of SNARE complex assembly(GO:0035544) |
| 0.1 | 1.8 | GO:0003094 | glomerular filtration(GO:0003094) |
| 0.1 | 6.6 | GO:0032024 | positive regulation of insulin secretion(GO:0032024) |
| 0.1 | 3.1 | GO:0046839 | phospholipid dephosphorylation(GO:0046839) |
| 0.1 | 0.8 | GO:0060965 | negative regulation of gene silencing by miRNA(GO:0060965) |
| 0.1 | 0.5 | GO:0021506 | anterior neuropore closure(GO:0021506) neuropore closure(GO:0021995) |
| 0.1 | 0.5 | GO:0090037 | positive regulation of protein kinase C signaling(GO:0090037) |
| 0.1 | 0.3 | GO:0071335 | regulation of enamel mineralization(GO:0070173) hair follicle cell proliferation(GO:0071335) |
| 0.0 | 0.7 | GO:0061299 | retina vasculature morphogenesis in camera-type eye(GO:0061299) |
| 0.0 | 2.3 | GO:0006284 | base-excision repair(GO:0006284) |
| 0.0 | 0.9 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.0 | 3.2 | GO:0071277 | cellular response to calcium ion(GO:0071277) |
| 0.0 | 0.3 | GO:0070307 | lens fiber cell development(GO:0070307) |
| 0.0 | 1.0 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.0 | 5.3 | GO:0030308 | negative regulation of cell growth(GO:0030308) |
| 0.0 | 0.5 | GO:0045953 | negative regulation of natural killer cell mediated cytotoxicity(GO:0045953) |
| 0.0 | 0.4 | GO:0060416 | response to growth hormone(GO:0060416) |
| 0.0 | 5.9 | GO:0015711 | organic anion transport(GO:0015711) |
| 0.0 | 0.0 | GO:0032468 | cellular manganese ion homeostasis(GO:0030026) Golgi calcium ion homeostasis(GO:0032468) manganese ion homeostasis(GO:0055071) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 5.1 | GO:0034751 | aryl hydrocarbon receptor complex(GO:0034751) |
| 0.7 | 6.2 | GO:0097442 | CA3 pyramidal cell dendrite(GO:0097442) |
| 0.5 | 9.6 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 0.5 | 3.0 | GO:0097226 | sperm mitochondrial sheath(GO:0097226) |
| 0.4 | 3.5 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
| 0.4 | 2.0 | GO:0016222 | procollagen-proline 4-dioxygenase complex(GO:0016222) |
| 0.3 | 1.5 | GO:0043202 | lysosomal lumen(GO:0043202) |
| 0.3 | 2.3 | GO:0035985 | senescence-associated heterochromatin focus(GO:0035985) |
| 0.3 | 4.5 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.2 | 0.5 | GO:0048269 | methionine adenosyltransferase complex(GO:0048269) |
| 0.2 | 2.5 | GO:0034366 | spherical high-density lipoprotein particle(GO:0034366) |
| 0.1 | 6.3 | GO:0005790 | smooth endoplasmic reticulum(GO:0005790) |
| 0.1 | 2.2 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.1 | 0.5 | GO:0070826 | paraferritin complex(GO:0070826) |
| 0.1 | 2.0 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.1 | 1.7 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.1 | 4.3 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 0.4 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.0 | 3.2 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.0 | 2.3 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.0 | 0.4 | GO:0071598 | neuronal ribonucleoprotein granule(GO:0071598) |
| 0.0 | 0.4 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.0 | 2.9 | GO:0000502 | proteasome complex(GO:0000502) |
| 0.0 | 2.8 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 2.5 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 0.2 | GO:0097422 | tubular endosome(GO:0097422) |
| 0.0 | 6.3 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 1.7 | GO:0045335 | phagocytic vesicle(GO:0045335) |
| 0.0 | 10.7 | GO:0005874 | microtubule(GO:0005874) |
| 0.0 | 0.5 | GO:0090545 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.0 | 5.9 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 1.8 | GO:0042641 | actomyosin(GO:0042641) |
| 0.0 | 0.6 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.0 | 8.1 | GO:0048471 | perinuclear region of cytoplasm(GO:0048471) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 8.1 | GO:0002135 | CTP binding(GO:0002135) |
| 1.1 | 11.9 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.9 | 3.5 | GO:0031694 | alpha-2A adrenergic receptor binding(GO:0031694) |
| 0.8 | 1.6 | GO:0032564 | dATP binding(GO:0032564) |
| 0.7 | 2.2 | GO:0005169 | neurotrophin TRKB receptor binding(GO:0005169) |
| 0.7 | 6.2 | GO:0005519 | cytoskeletal regulatory protein binding(GO:0005519) |
| 0.6 | 3.5 | GO:0034186 | apolipoprotein A-I binding(GO:0034186) |
| 0.5 | 3.1 | GO:0004647 | phosphoserine phosphatase activity(GO:0004647) |
| 0.4 | 2.2 | GO:0071987 | WD40-repeat domain binding(GO:0071987) |
| 0.4 | 3.7 | GO:0055131 | C3HC4-type RING finger domain binding(GO:0055131) |
| 0.4 | 2.4 | GO:0008970 | phosphatidylcholine 1-acylhydrolase activity(GO:0008970) |
| 0.3 | 2.0 | GO:0004656 | procollagen-proline 4-dioxygenase activity(GO:0004656) |
| 0.3 | 3.0 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.3 | 9.2 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.3 | 1.9 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.3 | 1.1 | GO:0004307 | ethanolaminephosphotransferase activity(GO:0004307) |
| 0.2 | 1.7 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 0.2 | 3.1 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 0.2 | 0.6 | GO:0001160 | transcription termination site sequence-specific DNA binding(GO:0001147) transcription termination site DNA binding(GO:0001160) |
| 0.2 | 2.5 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.2 | 9.9 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.2 | 8.9 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.1 | 2.3 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.1 | 2.0 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.1 | 0.5 | GO:0004478 | methionine adenosyltransferase activity(GO:0004478) |
| 0.1 | 2.3 | GO:0003906 | DNA-(apurinic or apyrimidinic site) lyase activity(GO:0003906) |
| 0.1 | 1.6 | GO:0042609 | CD4 receptor binding(GO:0042609) |
| 0.1 | 4.3 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.1 | 1.6 | GO:0036042 | long-chain fatty acyl-CoA binding(GO:0036042) |
| 0.1 | 3.1 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.1 | 4.3 | GO:0031683 | G-protein beta/gamma-subunit complex binding(GO:0031683) |
| 0.1 | 5.6 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.1 | 2.5 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.1 | 0.5 | GO:0015086 | cadmium ion transmembrane transporter activity(GO:0015086) lead ion transmembrane transporter activity(GO:0015094) vanadium ion transmembrane transporter activity(GO:0015100) ferrous iron uptake transmembrane transporter activity(GO:0015639) |
| 0.1 | 0.6 | GO:0005094 | Rho GDP-dissociation inhibitor activity(GO:0005094) |
| 0.1 | 0.5 | GO:0070728 | leucine binding(GO:0070728) |
| 0.1 | 2.2 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.1 | 0.5 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.1 | 3.5 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 1.4 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.1 | 0.3 | GO:0032296 | ribonuclease III activity(GO:0004525) double-stranded RNA-specific ribonuclease activity(GO:0032296) |
| 0.0 | 3.6 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 1.8 | GO:0004177 | aminopeptidase activity(GO:0004177) |
| 0.0 | 1.5 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.0 | 0.3 | GO:0047184 | 1-acylglycerophosphocholine O-acyltransferase activity(GO:0047184) |
| 0.0 | 6.0 | GO:0042626 | ATPase activity, coupled to transmembrane movement of substances(GO:0042626) |
| 0.0 | 0.4 | GO:1990247 | N6-methyladenosine-containing RNA binding(GO:1990247) |
| 0.0 | 0.4 | GO:0015187 | glycine transmembrane transporter activity(GO:0015187) |
| 0.0 | 1.4 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.0 | 2.0 | GO:0003725 | double-stranded RNA binding(GO:0003725) |
| 0.0 | 0.8 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.0 | 3.9 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.0 | 0.4 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 6.2 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.2 | 6.0 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.1 | 3.0 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.1 | 6.2 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.1 | 2.3 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.1 | 0.5 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.0 | 2.8 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.0 | 5.1 | PID VEGFR1 2 PATHWAY | Signaling events mediated by VEGFR1 and VEGFR2 |
| 0.0 | 3.5 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.0 | 0.8 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.0 | 3.2 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.0 | 1.4 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.0 | 2.2 | PID SHP2 PATHWAY | SHP2 signaling |
| 0.0 | 2.2 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 2.5 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 2.1 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 5.6 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.8 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.0 | 0.7 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 0.5 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.0 | 2.5 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 4.3 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.5 | 9.0 | REACTOME HYALURONAN METABOLISM | Genes involved in Hyaluronan metabolism |
| 0.3 | 2.8 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.3 | 5.1 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.2 | 2.5 | REACTOME IRAK1 RECRUITS IKK COMPLEX | Genes involved in IRAK1 recruits IKK complex |
| 0.2 | 4.5 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.2 | 2.0 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.1 | 3.0 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.1 | 3.1 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.1 | 3.6 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.1 | 3.5 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.1 | 1.5 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.1 | 1.1 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.1 | 0.5 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.0 | 2.3 | REACTOME RNA POL I TRANSCRIPTION | Genes involved in RNA Polymerase I Transcription |
| 0.0 | 1.4 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.0 | 2.5 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 2.5 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.0 | 2.7 | REACTOME L1CAM INTERACTIONS | Genes involved in L1CAM interactions |
| 0.0 | 0.5 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.0 | 0.3 | REACTOME MICRORNA MIRNA BIOGENESIS | Genes involved in MicroRNA (miRNA) Biogenesis |