Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
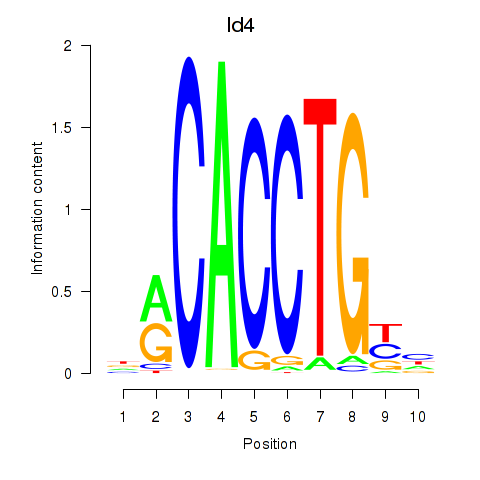
Results for Id4
Z-value: 1.99
Transcription factors associated with Id4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Id4
|
ENSMUSG00000021379.3 | Id4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Id4 | mm39_v1_chr13_+_48414582_48414704 | 0.04 | 7.1e-01 | Click! |
Activity profile of Id4 motif
Sorted Z-values of Id4 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Id4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chrX_+_100427331 | 23.56 |
ENSMUST00000119190.2
|
Gjb1
|
gap junction protein, beta 1 |
| chr17_+_26036893 | 23.01 |
ENSMUST00000235694.2
|
Fbxl16
|
F-box and leucine-rich repeat protein 16 |
| chr15_-_98575332 | 18.47 |
ENSMUST00000120997.2
ENSMUST00000109149.9 ENSMUST00000003451.11 |
Rnd1
|
Rho family GTPase 1 |
| chr5_+_35915217 | 16.07 |
ENSMUST00000101280.10
ENSMUST00000054598.12 ENSMUST00000114205.8 ENSMUST00000114206.9 |
Ablim2
|
actin-binding LIM protein 2 |
| chr14_+_40827317 | 14.98 |
ENSMUST00000047286.7
|
Mat1a
|
methionine adenosyltransferase I, alpha |
| chr15_+_54434576 | 14.70 |
ENSMUST00000025356.4
|
Mal2
|
mal, T cell differentiation protein 2 |
| chr5_+_35915290 | 14.39 |
ENSMUST00000114204.8
ENSMUST00000129347.8 |
Ablim2
|
actin-binding LIM protein 2 |
| chr12_+_82663785 | 14.30 |
ENSMUST00000200911.4
|
Rgs6
|
regulator of G-protein signaling 6 |
| chr7_-_30623592 | 13.43 |
ENSMUST00000217812.2
ENSMUST00000074671.9 |
Hamp2
|
hepcidin antimicrobial peptide 2 |
| chr2_+_71811526 | 13.33 |
ENSMUST00000090826.12
ENSMUST00000102698.10 |
Rapgef4
|
Rap guanine nucleotide exchange factor (GEF) 4 |
| chr14_+_40826970 | 13.06 |
ENSMUST00000225720.2
|
Mat1a
|
methionine adenosyltransferase I, alpha |
| chr17_-_26420300 | 12.94 |
ENSMUST00000025019.9
|
Arhgdig
|
Rho GDP dissociation inhibitor (GDI) gamma |
| chr15_+_99615396 | 12.61 |
ENSMUST00000023760.13
ENSMUST00000162194.2 |
Gpd1
|
glycerol-3-phosphate dehydrogenase 1 (soluble) |
| chr14_-_30645503 | 12.58 |
ENSMUST00000227995.2
|
Itih3
|
inter-alpha trypsin inhibitor, heavy chain 3 |
| chr14_-_57342150 | 12.42 |
ENSMUST00000055698.8
|
Gjb2
|
gap junction protein, beta 2 |
| chr14_+_40827108 | 12.39 |
ENSMUST00000224514.2
|
Mat1a
|
methionine adenosyltransferase I, alpha |
| chr9_+_108174052 | 12.38 |
ENSMUST00000035230.7
|
Amt
|
aminomethyltransferase |
| chr11_-_119438569 | 11.60 |
ENSMUST00000026670.5
|
Nptx1
|
neuronal pentraxin 1 |
| chr6_+_118043307 | 11.49 |
ENSMUST00000203804.3
ENSMUST00000203482.2 |
Rasgef1a
|
RasGEF domain family, member 1A |
| chr17_-_26420332 | 11.23 |
ENSMUST00000121959.3
|
Arhgdig
|
Rho GDP dissociation inhibitor (GDI) gamma |
| chr18_-_80133177 | 11.22 |
ENSMUST00000178391.2
|
Gm21886
|
predicted gene, 21886 |
| chrX_-_161426542 | 11.16 |
ENSMUST00000101102.2
|
Reps2
|
RALBP1 associated Eps domain containing protein 2 |
| chr2_+_126394327 | 11.05 |
ENSMUST00000061491.14
|
Slc27a2
|
solute carrier family 27 (fatty acid transporter), member 2 |
| chr2_-_121637505 | 11.04 |
ENSMUST00000138157.8
|
Frmd5
|
FERM domain containing 5 |
| chr19_-_30152814 | 10.97 |
ENSMUST00000025778.9
|
Gldc
|
glycine decarboxylase |
| chr7_+_127400016 | 10.79 |
ENSMUST00000106271.2
ENSMUST00000138432.2 |
Hsd3b7
|
hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 7 |
| chr2_+_102488985 | 10.71 |
ENSMUST00000080210.10
|
Slc1a2
|
solute carrier family 1 (glial high affinity glutamate transporter), member 2 |
| chr19_-_38113056 | 10.39 |
ENSMUST00000236283.2
|
Rbp4
|
retinol binding protein 4, plasma |
| chr11_-_35871300 | 10.36 |
ENSMUST00000018993.7
|
Wwc1
|
WW, C2 and coiled-coil domain containing 1 |
| chr11_+_120421496 | 10.04 |
ENSMUST00000026119.8
|
Gcgr
|
glucagon receptor |
| chr15_+_81695615 | 10.03 |
ENSMUST00000023024.8
|
Tef
|
thyrotroph embryonic factor |
| chr10_-_24803336 | 9.86 |
ENSMUST00000020161.10
|
Arg1
|
arginase, liver |
| chr19_-_36714053 | 9.80 |
ENSMUST00000087321.4
|
Ppp1r3c
|
protein phosphatase 1, regulatory subunit 3C |
| chr12_+_82663660 | 9.60 |
ENSMUST00000161801.8
ENSMUST00000185665.7 |
Rgs6
|
regulator of G-protein signaling 6 |
| chr6_+_47221293 | 9.25 |
ENSMUST00000199100.5
|
Cntnap2
|
contactin associated protein-like 2 |
| chr18_-_43032535 | 9.15 |
ENSMUST00000120632.2
|
Ppp2r2b
|
protein phosphatase 2, regulatory subunit B, beta |
| chr8_-_4267260 | 9.14 |
ENSMUST00000168386.9
|
Prr36
|
proline rich 36 |
| chr7_-_45750153 | 8.99 |
ENSMUST00000180081.3
|
Kcnj11
|
potassium inwardly rectifying channel, subfamily J, member 11 |
| chr11_+_42310557 | 8.96 |
ENSMUST00000007797.10
|
Gabrb2
|
gamma-aminobutyric acid (GABA) A receptor, subunit beta 2 |
| chr11_-_5900019 | 8.92 |
ENSMUST00000102920.4
|
Gck
|
glucokinase |
| chr2_-_121637469 | 8.90 |
ENSMUST00000110592.2
|
Frmd5
|
FERM domain containing 5 |
| chr14_+_55173696 | 8.89 |
ENSMUST00000037814.8
|
Cmtm5
|
CKLF-like MARVEL transmembrane domain containing 5 |
| chr18_-_43032514 | 8.87 |
ENSMUST00000236238.2
|
Ppp2r2b
|
protein phosphatase 2, regulatory subunit B, beta |
| chr7_+_140343652 | 8.86 |
ENSMUST00000026552.9
ENSMUST00000209253.2 ENSMUST00000210235.2 |
Cyp2e1
|
cytochrome P450, family 2, subfamily e, polypeptide 1 |
| chr19_-_40175709 | 8.85 |
ENSMUST00000051846.13
|
Cyp2c70
|
cytochrome P450, family 2, subfamily c, polypeptide 70 |
| chr2_+_102489558 | 8.85 |
ENSMUST00000111213.8
|
Slc1a2
|
solute carrier family 1 (glial high affinity glutamate transporter), member 2 |
| chr11_+_69983459 | 8.85 |
ENSMUST00000102572.8
|
Asgr2
|
asialoglycoprotein receptor 2 |
| chr11_+_69983531 | 8.71 |
ENSMUST00000124721.2
|
Asgr2
|
asialoglycoprotein receptor 2 |
| chr12_+_82663347 | 8.70 |
ENSMUST00000186848.5
|
Rgs6
|
regulator of G-protein signaling 6 |
| chr7_+_3352159 | 8.56 |
ENSMUST00000172109.4
|
Prkcg
|
protein kinase C, gamma |
| chr3_-_121056944 | 8.52 |
ENSMUST00000128909.8
ENSMUST00000029777.14 |
Tlcd4
|
TLC domain containing 4 |
| chr2_+_70392351 | 8.48 |
ENSMUST00000094934.11
|
Gad1
|
glutamate decarboxylase 1 |
| chr3_+_75464837 | 8.42 |
ENSMUST00000161776.8
ENSMUST00000029423.9 |
Serpini1
|
serine (or cysteine) peptidase inhibitor, clade I, member 1 |
| chr12_+_112688597 | 8.38 |
ENSMUST00000101018.11
ENSMUST00000092279.7 ENSMUST00000179041.8 ENSMUST00000222711.2 |
Cep170b
|
centrosomal protein 170B |
| chr1_-_87438027 | 8.37 |
ENSMUST00000027477.15
|
Ngef
|
neuronal guanine nucleotide exchange factor |
| chr8_-_70892204 | 8.36 |
ENSMUST00000076615.6
|
Crtc1
|
CREB regulated transcription coactivator 1 |
| chr14_-_57370706 | 8.34 |
ENSMUST00000160703.2
|
Gjb6
|
gap junction protein, beta 6 |
| chr1_+_182591425 | 8.28 |
ENSMUST00000155229.7
ENSMUST00000153348.8 |
Susd4
|
sushi domain containing 4 |
| chr16_+_16031182 | 8.22 |
ENSMUST00000039408.3
|
Pkp2
|
plakophilin 2 |
| chr2_+_31360219 | 8.22 |
ENSMUST00000102840.5
|
Ass1
|
argininosuccinate synthetase 1 |
| chr4_+_133280680 | 8.10 |
ENSMUST00000042706.3
|
Nr0b2
|
nuclear receptor subfamily 0, group B, member 2 |
| chr17_-_57394718 | 8.05 |
ENSMUST00000071135.6
|
Tubb4a
|
tubulin, beta 4A class IVA |
| chr1_+_74830675 | 7.99 |
ENSMUST00000006718.15
|
Wnt10a
|
wingless-type MMTV integration site family, member 10A |
| chr11_+_69983479 | 7.95 |
ENSMUST00000143772.8
|
Asgr2
|
asialoglycoprotein receptor 2 |
| chr17_+_81251997 | 7.92 |
ENSMUST00000025092.5
|
Tmem178
|
transmembrane protein 178 |
| chr3_+_145464413 | 7.84 |
ENSMUST00000029845.15
|
Ddah1
|
dimethylarginine dimethylaminohydrolase 1 |
| chr8_-_41087793 | 7.84 |
ENSMUST00000173957.2
ENSMUST00000048898.17 ENSMUST00000174205.8 |
Mtmr7
|
myotubularin related protein 7 |
| chr11_-_61344818 | 7.82 |
ENSMUST00000060255.14
ENSMUST00000054927.14 ENSMUST00000102661.4 |
Rnf112
|
ring finger protein 112 |
| chr10_-_80096842 | 7.71 |
ENSMUST00000105363.8
|
Gamt
|
guanidinoacetate methyltransferase |
| chr2_-_113659360 | 7.48 |
ENSMUST00000024005.8
|
Scg5
|
secretogranin V |
| chr10_+_123099945 | 7.46 |
ENSMUST00000238972.2
ENSMUST00000050756.8 |
Tafa2
|
TAFA chemokine like family member 2 |
| chr14_-_31362835 | 7.45 |
ENSMUST00000167066.8
ENSMUST00000127204.9 |
Hacl1
|
2-hydroxyacyl-CoA lyase 1 |
| chr7_+_3352019 | 7.45 |
ENSMUST00000100301.11
|
Prkcg
|
protein kinase C, gamma |
| chr2_-_64853083 | 7.38 |
ENSMUST00000028252.14
|
Grb14
|
growth factor receptor bound protein 14 |
| chr8_+_120121612 | 7.32 |
ENSMUST00000098367.5
|
Mlycd
|
malonyl-CoA decarboxylase |
| chr18_+_61058684 | 7.32 |
ENSMUST00000102888.10
ENSMUST00000025519.11 |
Camk2a
|
calcium/calmodulin-dependent protein kinase II alpha |
| chr2_+_49509288 | 7.30 |
ENSMUST00000028102.14
|
Kif5c
|
kinesin family member 5C |
| chr14_-_31362909 | 7.29 |
ENSMUST00000022437.16
|
Hacl1
|
2-hydroxyacyl-CoA lyase 1 |
| chr9_-_83688294 | 7.20 |
ENSMUST00000034796.14
ENSMUST00000183614.2 |
Elovl4
|
elongation of very long chain fatty acids (FEN1/Elo2, SUR4/Elo3, yeast)-like 4 |
| chr15_-_75438660 | 7.19 |
ENSMUST00000065417.15
|
Ly6h
|
lymphocyte antigen 6 complex, locus H |
| chr10_-_80096793 | 7.17 |
ENSMUST00000020359.7
|
Gamt
|
guanidinoacetate methyltransferase |
| chr18_-_38345010 | 7.16 |
ENSMUST00000159405.3
ENSMUST00000160721.8 |
Pcdh1
|
protocadherin 1 |
| chr6_+_21215472 | 7.14 |
ENSMUST00000081542.6
|
Kcnd2
|
potassium voltage-gated channel, Shal-related family, member 2 |
| chr17_-_45996899 | 7.09 |
ENSMUST00000145873.8
|
Tmem63b
|
transmembrane protein 63b |
| chr14_-_57371041 | 7.06 |
ENSMUST00000039380.9
|
Gjb6
|
gap junction protein, beta 6 |
| chr16_-_17906886 | 7.06 |
ENSMUST00000132241.2
ENSMUST00000139861.2 ENSMUST00000003620.13 |
Prodh
|
proline dehydrogenase |
| chr17_-_45997132 | 7.06 |
ENSMUST00000113523.9
|
Tmem63b
|
transmembrane protein 63b |
| chr7_-_45750050 | 7.03 |
ENSMUST00000209291.2
|
Kcnj11
|
potassium inwardly rectifying channel, subfamily J, member 11 |
| chr19_-_10282218 | 7.02 |
ENSMUST00000039327.11
|
Dagla
|
diacylglycerol lipase, alpha |
| chr16_-_45830575 | 7.01 |
ENSMUST00000130481.2
|
Plcxd2
|
phosphatidylinositol-specific phospholipase C, X domain containing 2 |
| chr18_-_43032359 | 6.97 |
ENSMUST00000117687.8
|
Ppp2r2b
|
protein phosphatase 2, regulatory subunit B, beta |
| chr2_-_52225763 | 6.93 |
ENSMUST00000238288.2
ENSMUST00000238749.2 |
Neb
|
nebulin |
| chr16_-_60425608 | 6.92 |
ENSMUST00000068860.13
|
Epha6
|
Eph receptor A6 |
| chr12_+_113106407 | 6.91 |
ENSMUST00000196015.5
|
Crip2
|
cysteine rich protein 2 |
| chr6_+_72074545 | 6.90 |
ENSMUST00000069994.11
ENSMUST00000114112.4 |
St3gal5
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 5 |
| chr14_+_27344385 | 6.90 |
ENSMUST00000210135.2
ENSMUST00000090302.6 ENSMUST00000211087.2 |
Erc2
|
ELKS/RAB6-interacting/CAST family member 2 |
| chrX_-_161426624 | 6.86 |
ENSMUST00000112334.8
|
Reps2
|
RALBP1 associated Eps domain containing protein 2 |
| chr13_-_49301407 | 6.83 |
ENSMUST00000162581.8
ENSMUST00000110097.9 ENSMUST00000049265.15 ENSMUST00000035538.13 ENSMUST00000110096.8 ENSMUST00000091623.10 |
Wnk2
|
WNK lysine deficient protein kinase 2 |
| chr16_+_20408886 | 6.80 |
ENSMUST00000232279.2
ENSMUST00000232474.2 |
Vwa5b2
|
von Willebrand factor A domain containing 5B2 |
| chrX_+_10581248 | 6.75 |
ENSMUST00000144356.8
|
Mid1ip1
|
Mid1 interacting protein 1 (gastrulation specific G12-like (zebrafish)) |
| chr1_-_125839897 | 6.66 |
ENSMUST00000159417.2
|
Lypd1
|
Ly6/Plaur domain containing 1 |
| chr6_+_82018604 | 6.65 |
ENSMUST00000042974.15
|
Eva1a
|
eva-1 homolog A (C. elegans) |
| chr7_+_57069417 | 6.64 |
ENSMUST00000085240.11
|
Gabrb3
|
gamma-aminobutyric acid (GABA) A receptor, subunit beta 3 |
| chr7_+_141503719 | 6.61 |
ENSMUST00000105989.9
ENSMUST00000075528.12 ENSMUST00000174499.8 |
Brsk2
|
BR serine/threonine kinase 2 |
| chr13_+_104246259 | 6.56 |
ENSMUST00000160322.8
ENSMUST00000159574.2 |
Sgtb
|
small glutamine-rich tetratricopeptide repeat (TPR)-containing, beta |
| chr5_+_37403098 | 6.56 |
ENSMUST00000031004.11
|
Crmp1
|
collapsin response mediator protein 1 |
| chr2_+_70392491 | 6.52 |
ENSMUST00000148210.8
|
Gad1
|
glutamate decarboxylase 1 |
| chr13_-_110417421 | 6.51 |
ENSMUST00000223922.2
|
Rab3c
|
RAB3C, member RAS oncogene family |
| chr7_-_46782448 | 6.45 |
ENSMUST00000033142.13
|
Ptpn5
|
protein tyrosine phosphatase, non-receptor type 5 |
| chr15_-_75438457 | 6.42 |
ENSMUST00000163116.8
ENSMUST00000023241.12 |
Ly6h
|
lymphocyte antigen 6 complex, locus H |
| chr9_-_114673158 | 6.41 |
ENSMUST00000047013.4
|
Cmtm8
|
CKLF-like MARVEL transmembrane domain containing 8 |
| chr10_+_75768964 | 6.40 |
ENSMUST00000219839.2
|
Chchd10
|
coiled-coil-helix-coiled-coil-helix domain containing 10 |
| chr8_-_4266912 | 6.39 |
ENSMUST00000177491.8
|
Prr36
|
proline rich 36 |
| chr10_+_127702326 | 6.37 |
ENSMUST00000092058.4
|
Rdh16f2
|
RDH16 family member 2 |
| chr8_-_85663976 | 6.36 |
ENSMUST00000109741.9
ENSMUST00000119820.2 |
Mast1
|
microtubule associated serine/threonine kinase 1 |
| chr19_-_58443593 | 6.30 |
ENSMUST00000135730.2
ENSMUST00000152507.8 |
Gfra1
|
glial cell line derived neurotrophic factor family receptor alpha 1 |
| chr7_+_4925781 | 6.22 |
ENSMUST00000207527.2
ENSMUST00000207687.2 ENSMUST00000208754.2 |
Nat14
|
N-acetyltransferase 14 |
| chr13_+_104246245 | 6.22 |
ENSMUST00000044385.14
|
Sgtb
|
small glutamine-rich tetratricopeptide repeat (TPR)-containing, beta |
| chr1_+_153616090 | 6.21 |
ENSMUST00000027748.8
|
Rgs16
|
regulator of G-protein signaling 16 |
| chrX_+_10118544 | 6.15 |
ENSMUST00000049910.13
|
Otc
|
ornithine transcarbamylase |
| chr6_+_121277186 | 6.15 |
ENSMUST00000064580.14
|
Slc6a13
|
solute carrier family 6 (neurotransmitter transporter, GABA), member 13 |
| chr1_+_93062962 | 6.14 |
ENSMUST00000027491.7
|
Agxt
|
alanine-glyoxylate aminotransferase |
| chr6_+_105654706 | 6.09 |
ENSMUST00000113261.9
ENSMUST00000113264.9 |
Cntn4
|
contactin 4 |
| chr11_+_78394273 | 6.09 |
ENSMUST00000001130.8
ENSMUST00000125670.3 |
Sebox
|
SEBOX homeobox |
| chr6_+_96092230 | 6.05 |
ENSMUST00000075080.6
|
Tafa1
|
TAFA chemokine like family member 1 |
| chr1_-_184615415 | 6.04 |
ENSMUST00000048308.6
|
C130074G19Rik
|
RIKEN cDNA C130074G19 gene |
| chr17_+_35455532 | 6.02 |
ENSMUST00000068261.9
|
Atp6v1g2
|
ATPase, H+ transporting, lysosomal V1 subunit G2 |
| chr6_+_113460258 | 5.98 |
ENSMUST00000032422.6
|
Creld1
|
cysteine-rich with EGF-like domains 1 |
| chr13_+_25240138 | 5.98 |
ENSMUST00000069614.7
|
Dcdc2a
|
doublecortin domain containing 2a |
| chr10_-_127724557 | 5.98 |
ENSMUST00000047199.5
|
Rdh7
|
retinol dehydrogenase 7 |
| chr9_-_20657643 | 5.96 |
ENSMUST00000215999.2
|
Olfm2
|
olfactomedin 2 |
| chr2_+_107120934 | 5.95 |
ENSMUST00000037012.3
|
Kcna4
|
potassium voltage-gated channel, shaker-related subfamily, member 4 |
| chrX_+_135567124 | 5.94 |
ENSMUST00000060904.11
ENSMUST00000113100.2 ENSMUST00000128040.2 |
Tceal3
|
transcription elongation factor A (SII)-like 3 |
| chr6_+_48514518 | 5.94 |
ENSMUST00000040361.8
|
Atp6v0e2
|
ATPase, H+ transporting, lysosomal V0 subunit E2 |
| chr11_+_101046708 | 5.94 |
ENSMUST00000043654.10
|
Tubg2
|
tubulin, gamma 2 |
| chr7_+_114344920 | 5.93 |
ENSMUST00000136645.8
ENSMUST00000169913.8 |
Insc
|
INSC spindle orientation adaptor protein |
| chr1_-_75195889 | 5.88 |
ENSMUST00000186213.7
|
Tuba4a
|
tubulin, alpha 4A |
| chr9_-_70048766 | 5.84 |
ENSMUST00000034749.16
|
Fam81a
|
family with sequence similarity 81, member A |
| chr18_+_61058716 | 5.80 |
ENSMUST00000115297.8
|
Camk2a
|
calcium/calmodulin-dependent protein kinase II alpha |
| chrX_+_10118600 | 5.79 |
ENSMUST00000115528.3
|
Otc
|
ornithine transcarbamylase |
| chr4_+_108736350 | 5.78 |
ENSMUST00000106651.9
|
Rab3b
|
RAB3B, member RAS oncogene family |
| chr6_+_105654729 | 5.76 |
ENSMUST00000089208.9
|
Cntn4
|
contactin 4 |
| chr13_+_91889626 | 5.74 |
ENSMUST00000022120.5
|
Acot12
|
acyl-CoA thioesterase 12 |
| chr7_-_84059170 | 5.72 |
ENSMUST00000208995.2
|
Arnt2
|
aryl hydrocarbon receptor nuclear translocator 2 |
| chr1_+_85973585 | 5.72 |
ENSMUST00000027429.11
ENSMUST00000165824.3 |
2810459M11Rik
|
RIKEN cDNA 2810459M11 gene |
| chr7_+_126447080 | 5.70 |
ENSMUST00000147257.2
ENSMUST00000139174.2 |
Doc2a
|
double C2, alpha |
| chr9_+_107812873 | 5.70 |
ENSMUST00000035700.14
|
Camkv
|
CaM kinase-like vesicle-associated |
| chr16_-_4950285 | 5.69 |
ENSMUST00000035672.5
|
Ppl
|
periplakin |
| chr2_-_30364219 | 5.69 |
ENSMUST00000065134.4
|
Ier5l
|
immediate early response 5-like |
| chr8_+_36924702 | 5.68 |
ENSMUST00000135373.8
ENSMUST00000147525.9 |
Trmt9b
|
tRNA methyltransferase 9B |
| chr6_+_72074718 | 5.66 |
ENSMUST00000187007.3
|
St3gal5
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 5 |
| chr7_+_87233554 | 5.65 |
ENSMUST00000125009.9
|
Grm5
|
glutamate receptor, metabotropic 5 |
| chr6_-_24956296 | 5.64 |
ENSMUST00000127247.4
|
Tmem229a
|
transmembrane protein 229A |
| chr16_-_34083315 | 5.61 |
ENSMUST00000114953.8
|
Kalrn
|
kalirin, RhoGEF kinase |
| chr6_-_92683136 | 5.58 |
ENSMUST00000032093.12
|
Prickle2
|
prickle planar cell polarity protein 2 |
| chr18_-_16942289 | 5.56 |
ENSMUST00000025166.14
|
Cdh2
|
cadherin 2 |
| chr7_-_44465043 | 5.55 |
ENSMUST00000107893.9
|
Atf5
|
activating transcription factor 5 |
| chr9_-_56703422 | 5.53 |
ENSMUST00000210032.2
|
Lingo1
|
leucine rich repeat and Ig domain containing 1 |
| chr14_+_55173936 | 5.52 |
ENSMUST00000227441.2
|
Cmtm5
|
CKLF-like MARVEL transmembrane domain containing 5 |
| chr14_+_21102662 | 5.51 |
ENSMUST00000223915.2
|
Adk
|
adenosine kinase |
| chr1_-_180310894 | 5.51 |
ENSMUST00000211561.2
ENSMUST00000136521.2 ENSMUST00000179826.2 |
Stum
|
mechanosensory transduction mediator |
| chr5_-_139115914 | 5.51 |
ENSMUST00000129851.8
|
Prkar1b
|
protein kinase, cAMP dependent regulatory, type I beta |
| chr18_-_62044871 | 5.50 |
ENSMUST00000166783.3
ENSMUST00000049378.15 |
Ablim3
|
actin binding LIM protein family, member 3 |
| chr1_+_66214445 | 5.48 |
ENSMUST00000114017.8
ENSMUST00000114015.8 |
Map2
|
microtubule-associated protein 2 |
| chr2_-_53975501 | 5.44 |
ENSMUST00000100089.3
|
Rprm
|
reprimo, TP53 dependent G2 arrest mediator candidate |
| chr4_-_118401185 | 5.44 |
ENSMUST00000128098.8
|
Tmem125
|
transmembrane protein 125 |
| chr8_-_4267131 | 5.42 |
ENSMUST00000175906.2
|
Prr36
|
proline rich 36 |
| chr19_-_58443830 | 5.41 |
ENSMUST00000026076.14
|
Gfra1
|
glial cell line derived neurotrophic factor family receptor alpha 1 |
| chr1_+_158190090 | 5.39 |
ENSMUST00000194369.6
ENSMUST00000195311.6 |
Astn1
|
astrotactin 1 |
| chr9_+_36743980 | 5.38 |
ENSMUST00000034630.15
|
Fez1
|
fasciculation and elongation protein zeta 1 (zygin I) |
| chr6_+_48514578 | 5.38 |
ENSMUST00000203011.2
|
Atp6v0e2
|
ATPase, H+ transporting, lysosomal V0 subunit E2 |
| chr16_-_34083549 | 5.36 |
ENSMUST00000114949.8
ENSMUST00000114954.8 |
Kalrn
|
kalirin, RhoGEF kinase |
| chr19_+_6547790 | 5.31 |
ENSMUST00000113458.8
ENSMUST00000113459.2 |
Nrxn2
|
neurexin II |
| chr14_-_104081119 | 5.31 |
ENSMUST00000227824.2
ENSMUST00000172237.2 |
Ednrb
|
endothelin receptor type B |
| chrX_+_139243012 | 5.30 |
ENSMUST00000208130.2
|
Frmpd3
|
FERM and PDZ domain containing 3 |
| chr16_-_34083200 | 5.30 |
ENSMUST00000114947.2
|
Kalrn
|
kalirin, RhoGEF kinase |
| chr6_+_85164420 | 5.27 |
ENSMUST00000045942.9
|
Emx1
|
empty spiracles homeobox 1 |
| chr4_+_58943574 | 5.26 |
ENSMUST00000107554.2
|
Zkscan16
|
zinc finger with KRAB and SCAN domains 16 |
| chr4_+_139350152 | 5.25 |
ENSMUST00000039818.10
|
Aldh4a1
|
aldehyde dehydrogenase 4 family, member A1 |
| chr15_+_41652777 | 5.22 |
ENSMUST00000230778.2
ENSMUST00000022918.15 ENSMUST00000090095.13 |
Oxr1
|
oxidation resistance 1 |
| chr8_-_112120442 | 5.18 |
ENSMUST00000038475.9
|
Fa2h
|
fatty acid 2-hydroxylase |
| chr14_-_29443792 | 5.18 |
ENSMUST00000022567.9
|
Cacna2d3
|
calcium channel, voltage-dependent, alpha2/delta subunit 3 |
| chr4_+_141473983 | 5.17 |
ENSMUST00000038161.5
|
Agmat
|
agmatine ureohydrolase (agmatinase) |
| chr15_-_64794139 | 5.17 |
ENSMUST00000023007.7
ENSMUST00000228014.2 |
Adcy8
|
adenylate cyclase 8 |
| chr11_+_113510135 | 5.17 |
ENSMUST00000146390.3
|
Sstr2
|
somatostatin receptor 2 |
| chr19_+_5100475 | 5.16 |
ENSMUST00000225427.2
|
Rin1
|
Ras and Rab interactor 1 |
| chr14_+_21102642 | 5.16 |
ENSMUST00000045376.11
|
Adk
|
adenosine kinase |
| chr14_-_24054927 | 5.13 |
ENSMUST00000145596.3
|
Kcnma1
|
potassium large conductance calcium-activated channel, subfamily M, alpha member 1 |
| chr13_+_19132375 | 5.12 |
ENSMUST00000239207.2
ENSMUST00000003345.10 ENSMUST00000200466.5 |
Amph
|
amphiphysin |
| chr10_-_79973210 | 5.12 |
ENSMUST00000170219.9
ENSMUST00000169546.9 |
Cbarp
|
calcium channel, voltage-dependent, beta subunit associated regulatory protein |
| chr15_-_89726063 | 5.11 |
ENSMUST00000029441.4
|
Syt10
|
synaptotagmin X |
| chr9_+_32027335 | 5.10 |
ENSMUST00000174641.8
|
Arhgap32
|
Rho GTPase activating protein 32 |
| chr9_-_81515865 | 5.10 |
ENSMUST00000183482.2
|
Htr1b
|
5-hydroxytryptamine (serotonin) receptor 1B |
| chr4_-_46991842 | 5.09 |
ENSMUST00000107749.4
|
Gabbr2
|
gamma-aminobutyric acid (GABA) B receptor, 2 |
| chr4_-_155445818 | 5.09 |
ENSMUST00000030922.15
|
Prkcz
|
protein kinase C, zeta |
| chr8_-_4267459 | 5.08 |
ENSMUST00000176227.2
|
Prr36
|
proline rich 36 |
| chr11_+_16207705 | 5.07 |
ENSMUST00000109645.9
ENSMUST00000109647.3 |
Vstm2a
|
V-set and transmembrane domain containing 2A |
| chr3_+_13536696 | 5.05 |
ENSMUST00000191806.3
ENSMUST00000193117.3 |
Ralyl
|
RALY RNA binding protein-like |
| chr1_-_86598286 | 5.05 |
ENSMUST00000027449.6
|
Nppc
|
natriuretic peptide type C |
| chr17_+_46608842 | 5.05 |
ENSMUST00000166617.8
ENSMUST00000170271.2 |
Dlk2
|
delta like non-canonical Notch ligand 2 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 13.5 | 40.4 | GO:0009087 | methionine catabolic process(GO:0009087) |
| 7.8 | 23.4 | GO:0019464 | glycine catabolic process(GO:0006546) glycine decarboxylation via glycine cleavage system(GO:0019464) |
| 5.4 | 16.3 | GO:0006601 | creatine biosynthetic process(GO:0006601) |
| 4.4 | 13.1 | GO:0046168 | glycerol-3-phosphate catabolic process(GO:0046168) |
| 4.1 | 12.3 | GO:0010133 | proline catabolic process to glutamate(GO:0010133) |
| 4.0 | 11.9 | GO:0042450 | arginine biosynthetic process via ornithine(GO:0042450) |
| 3.7 | 25.8 | GO:0001561 | fatty acid alpha-oxidation(GO:0001561) |
| 3.6 | 10.8 | GO:0097494 | regulation of vesicle size(GO:0097494) |
| 3.6 | 10.7 | GO:0006169 | adenosine salvage(GO:0006169) dATP biosynthetic process(GO:0006175) |
| 3.4 | 13.4 | GO:0034757 | negative regulation of iron ion transport(GO:0034757) negative regulation of iron ion transmembrane transport(GO:0034760) |
| 3.2 | 12.7 | GO:0099541 | trans-synaptic signaling by lipid(GO:0099541) trans-synaptic signaling by endocannabinoid(GO:0099542) |
| 3.1 | 9.2 | GO:0071874 | cellular response to norepinephrine stimulus(GO:0071874) |
| 3.0 | 15.0 | GO:0009449 | gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 2.9 | 11.6 | GO:0030450 | regulation of complement activation, classical pathway(GO:0030450) |
| 2.8 | 25.5 | GO:0098885 | modification of postsynaptic actin cytoskeleton(GO:0098885) |
| 2.8 | 19.6 | GO:0070777 | D-aspartate transport(GO:0070777) D-aspartate import(GO:0070779) |
| 2.8 | 8.4 | GO:1902630 | regulation of membrane hyperpolarization(GO:1902630) |
| 2.8 | 8.3 | GO:0021682 | nerve maturation(GO:0021682) |
| 2.7 | 16.0 | GO:0032423 | regulation of mismatch repair(GO:0032423) |
| 2.6 | 7.8 | GO:1904274 | tricellular tight junction assembly(GO:1904274) |
| 2.6 | 12.9 | GO:0009730 | detection of carbohydrate stimulus(GO:0009730) detection of hexose stimulus(GO:0009732) detection of monosaccharide stimulus(GO:0034287) detection of glucose(GO:0051594) |
| 2.6 | 10.3 | GO:2000984 | regulation of ATP citrate synthase activity(GO:2000983) negative regulation of ATP citrate synthase activity(GO:2000984) |
| 2.5 | 7.4 | GO:1903537 | meiotic cell cycle process involved in oocyte maturation(GO:1903537) regulation of meiotic cell cycle process involved in oocyte maturation(GO:1903538) |
| 2.4 | 7.3 | GO:2001293 | malonyl-CoA metabolic process(GO:2001293) |
| 2.3 | 11.7 | GO:1904457 | positive regulation of neuronal action potential(GO:1904457) |
| 2.3 | 16.4 | GO:0045163 | clustering of voltage-gated potassium channels(GO:0045163) |
| 2.1 | 10.7 | GO:0043091 | L-arginine import(GO:0043091) arginine import(GO:0090467) |
| 2.1 | 10.4 | GO:0048807 | female genitalia morphogenesis(GO:0048807) |
| 2.1 | 8.2 | GO:0010046 | response to mycotoxin(GO:0010046) |
| 2.0 | 22.5 | GO:0035754 | B cell chemotaxis(GO:0035754) |
| 2.0 | 6.1 | GO:0019265 | glycine biosynthetic process, by transamination of glyoxylate(GO:0019265) L-alanine catabolic process(GO:0042853) oxalic acid secretion(GO:0046724) |
| 2.0 | 36.6 | GO:0043653 | mitochondrial fragmentation involved in apoptotic process(GO:0043653) |
| 2.0 | 24.1 | GO:0015868 | purine ribonucleotide transport(GO:0015868) |
| 1.8 | 7.3 | GO:0098964 | dendritic transport of ribonucleoprotein complex(GO:0098961) dendritic transport of messenger ribonucleoprotein complex(GO:0098963) anterograde dendritic transport of messenger ribonucleoprotein complex(GO:0098964) |
| 1.8 | 9.0 | GO:0007207 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) phospholipase C-activating G-protein coupled acetylcholine receptor signaling pathway(GO:0007207) |
| 1.8 | 5.3 | GO:0003165 | Purkinje myocyte development(GO:0003165) |
| 1.6 | 6.6 | GO:1904616 | regulation of actin filament binding(GO:1904529) regulation of actin binding(GO:1904616) |
| 1.6 | 4.9 | GO:0019085 | early viral transcription(GO:0019085) |
| 1.6 | 6.5 | GO:2001025 | positive regulation of response to drug(GO:2001025) |
| 1.5 | 15.1 | GO:0031580 | membrane raft polarization(GO:0001766) membrane raft distribution(GO:0031580) |
| 1.5 | 12.0 | GO:1903286 | regulation of potassium ion import(GO:1903286) |
| 1.5 | 9.0 | GO:1903278 | positive regulation of sodium ion export(GO:1903275) positive regulation of sodium ion export from cell(GO:1903278) |
| 1.5 | 1.5 | GO:0006597 | spermine biosynthetic process(GO:0006597) |
| 1.5 | 1.5 | GO:0070650 | actin filament bundle distribution(GO:0070650) |
| 1.4 | 8.7 | GO:0032439 | endosome localization(GO:0032439) |
| 1.4 | 7.0 | GO:0061642 | chemoattraction of axon(GO:0061642) |
| 1.4 | 4.2 | GO:0015705 | iodide transport(GO:0015705) |
| 1.4 | 2.8 | GO:0051582 | positive regulation of neurotransmitter uptake(GO:0051582) positive regulation of dopamine uptake involved in synaptic transmission(GO:0051586) positive regulation of catecholamine uptake involved in synaptic transmission(GO:0051944) |
| 1.4 | 8.2 | GO:0034334 | adherens junction maintenance(GO:0034334) |
| 1.3 | 13.1 | GO:0098989 | NMDA selective glutamate receptor signaling pathway(GO:0098989) |
| 1.3 | 3.9 | GO:0006850 | mitochondrial pyruvate transport(GO:0006850) mitochondrial pyruvate transmembrane transport(GO:1902361) |
| 1.3 | 3.9 | GO:0035928 | rRNA import into mitochondrion(GO:0035928) |
| 1.3 | 9.1 | GO:1904075 | regulation of oligodendrocyte apoptotic process(GO:1900141) negative regulation of oligodendrocyte apoptotic process(GO:1900142) regulation of trophectodermal cell proliferation(GO:1904073) positive regulation of trophectodermal cell proliferation(GO:1904075) |
| 1.3 | 7.6 | GO:0032902 | nerve growth factor production(GO:0032902) |
| 1.3 | 5.1 | GO:0070346 | positive regulation of fat cell proliferation(GO:0070346) |
| 1.3 | 10.0 | GO:0033762 | response to glucagon(GO:0033762) |
| 1.2 | 3.7 | GO:0001982 | baroreceptor response to decreased systemic arterial blood pressure(GO:0001982) |
| 1.2 | 8.6 | GO:0071502 | cellular response to temperature stimulus(GO:0071502) |
| 1.2 | 3.7 | GO:1904024 | response to cobalt ion(GO:0032025) negative regulation of NAD metabolic process(GO:1902689) negative regulation of glucose catabolic process to lactate via pyruvate(GO:1904024) |
| 1.2 | 3.7 | GO:1990697 | protein depalmitoleylation(GO:1990697) |
| 1.2 | 4.8 | GO:0015755 | fructose transport(GO:0015755) |
| 1.2 | 1.2 | GO:0061646 | positive regulation of glutamate neurotransmitter secretion in response to membrane depolarization(GO:0061646) |
| 1.2 | 4.7 | GO:0097477 | lateral motor column neuron migration(GO:0097477) |
| 1.2 | 11.8 | GO:0035331 | negative regulation of hippo signaling(GO:0035331) |
| 1.2 | 3.5 | GO:0021627 | olfactory nerve morphogenesis(GO:0021627) olfactory nerve structural organization(GO:0021629) |
| 1.2 | 10.5 | GO:0007196 | adenylate cyclase-inhibiting G-protein coupled glutamate receptor signaling pathway(GO:0007196) |
| 1.1 | 3.4 | GO:0002541 | activation of plasma proteins involved in acute inflammatory response(GO:0002541) |
| 1.1 | 5.6 | GO:0001830 | trophectodermal cell fate commitment(GO:0001830) |
| 1.1 | 3.3 | GO:0007509 | mesoderm migration involved in gastrulation(GO:0007509) |
| 1.1 | 4.3 | GO:0021679 | cerebellar molecular layer development(GO:0021679) vestibular nucleus development(GO:0021750) |
| 1.1 | 11.8 | GO:2000601 | positive regulation of Arp2/3 complex-mediated actin nucleation(GO:2000601) |
| 1.1 | 3.2 | GO:0070904 | L-ascorbic acid transport(GO:0015882) transepithelial L-ascorbic acid transport(GO:0070904) |
| 1.1 | 3.2 | GO:0051182 | coenzyme transport(GO:0051182) |
| 1.1 | 3.2 | GO:0097105 | presynaptic membrane assembly(GO:0097105) |
| 1.1 | 5.3 | GO:0007494 | midgut development(GO:0007494) |
| 1.0 | 4.2 | GO:0031959 | mineralocorticoid receptor signaling pathway(GO:0031959) |
| 1.0 | 3.1 | GO:1903920 | positive regulation of actin filament severing(GO:1903920) |
| 1.0 | 5.0 | GO:0061622 | glycolytic process through glucose-1-phosphate(GO:0061622) |
| 1.0 | 4.0 | GO:1904428 | negative regulation of tubulin deacetylation(GO:1904428) |
| 1.0 | 3.8 | GO:0072365 | regulation of cellular ketone metabolic process by negative regulation of transcription from RNA polymerase II promoter(GO:0072365) |
| 0.9 | 4.7 | GO:0050917 | sensory perception of umami taste(GO:0050917) |
| 0.9 | 4.6 | GO:0035359 | negative regulation of peroxisome proliferator activated receptor signaling pathway(GO:0035359) |
| 0.9 | 0.9 | GO:0051542 | elastin biosynthetic process(GO:0051542) |
| 0.9 | 10.1 | GO:0006527 | arginine catabolic process(GO:0006527) |
| 0.9 | 2.7 | GO:0090076 | relaxation of skeletal muscle(GO:0090076) |
| 0.9 | 4.4 | GO:0071985 | multivesicular body sorting pathway(GO:0071985) |
| 0.9 | 12.4 | GO:0031914 | negative regulation of synaptic plasticity(GO:0031914) |
| 0.9 | 2.6 | GO:2000503 | positive regulation of natural killer cell chemotaxis(GO:2000503) |
| 0.9 | 2.6 | GO:1904633 | glomerular visceral epithelial cell apoptotic process(GO:1903210) regulation of glomerular visceral epithelial cell apoptotic process(GO:1904633) positive regulation of glomerular visceral epithelial cell apoptotic process(GO:1904635) positive regulation of progesterone biosynthetic process(GO:2000184) |
| 0.9 | 5.2 | GO:0008295 | spermidine biosynthetic process(GO:0008295) |
| 0.9 | 5.1 | GO:0060082 | response to carbon monoxide(GO:0034465) eye blink reflex(GO:0060082) |
| 0.8 | 3.4 | GO:0051866 | general adaptation syndrome(GO:0051866) |
| 0.8 | 10.0 | GO:0034625 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.8 | 1.6 | GO:0060741 | prostate gland stromal morphogenesis(GO:0060741) |
| 0.8 | 2.4 | GO:1900673 | olefin metabolic process(GO:1900673) |
| 0.8 | 3.2 | GO:1904879 | positive regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1904879) |
| 0.8 | 5.6 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 0.8 | 1.6 | GO:0031179 | peptide modification(GO:0031179) |
| 0.8 | 9.4 | GO:0007158 | neuron cell-cell adhesion(GO:0007158) |
| 0.8 | 3.9 | GO:0060596 | mammary placode formation(GO:0060596) |
| 0.8 | 11.5 | GO:0051901 | positive regulation of mitochondrial depolarization(GO:0051901) |
| 0.8 | 9.2 | GO:0030432 | peristalsis(GO:0030432) |
| 0.8 | 4.5 | GO:0001757 | somite specification(GO:0001757) |
| 0.7 | 2.2 | GO:0060060 | post-embryonic retina morphogenesis in camera-type eye(GO:0060060) |
| 0.7 | 7.4 | GO:0061727 | methylglyoxal catabolic process to D-lactate via S-lactoyl-glutathione(GO:0019243) methylglyoxal catabolic process(GO:0051596) methylglyoxal catabolic process to lactate(GO:0061727) |
| 0.7 | 2.9 | GO:0046959 | habituation(GO:0046959) |
| 0.7 | 10.8 | GO:0060019 | radial glial cell differentiation(GO:0060019) oligodendrocyte progenitor proliferation(GO:0070444) regulation of oligodendrocyte progenitor proliferation(GO:0070445) |
| 0.7 | 2.9 | GO:0051121 | hepoxilin metabolic process(GO:0051121) hepoxilin biosynthetic process(GO:0051122) |
| 0.7 | 10.0 | GO:0006751 | glutathione catabolic process(GO:0006751) |
| 0.7 | 6.4 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.7 | 2.8 | GO:0010726 | positive regulation of hydrogen peroxide metabolic process(GO:0010726) |
| 0.7 | 20.3 | GO:0095500 | acetylcholine receptor signaling pathway(GO:0095500) signal transduction involved in cellular response to ammonium ion(GO:1903831) response to acetylcholine(GO:1905144) cellular response to acetylcholine(GO:1905145) |
| 0.7 | 2.8 | GO:0070368 | positive regulation of hepatocyte differentiation(GO:0070368) |
| 0.7 | 2.1 | GO:0061536 | glycine secretion(GO:0061536) glycine secretion, neurotransmission(GO:0061537) |
| 0.7 | 15.6 | GO:0071420 | cellular response to histamine(GO:0071420) |
| 0.7 | 9.5 | GO:0009650 | UV protection(GO:0009650) |
| 0.7 | 2.7 | GO:0009826 | unidimensional cell growth(GO:0009826) |
| 0.7 | 4.7 | GO:0035166 | post-embryonic hemopoiesis(GO:0035166) |
| 0.6 | 1.9 | GO:0007181 | transforming growth factor beta receptor complex assembly(GO:0007181) |
| 0.6 | 5.1 | GO:0003419 | growth plate cartilage chondrocyte proliferation(GO:0003419) |
| 0.6 | 3.8 | GO:0010890 | positive regulation of sequestering of triglyceride(GO:0010890) |
| 0.6 | 10.5 | GO:0090177 | establishment of planar polarity involved in neural tube closure(GO:0090177) |
| 0.6 | 14.1 | GO:0016322 | neuron remodeling(GO:0016322) |
| 0.6 | 1.8 | GO:0061763 | multivesicular body-lysosome fusion(GO:0061763) |
| 0.6 | 4.8 | GO:2001184 | myoblast differentiation involved in skeletal muscle regeneration(GO:0014835) positive regulation of interleukin-12 secretion(GO:2001184) |
| 0.6 | 9.6 | GO:1902902 | negative regulation of autophagosome assembly(GO:1902902) |
| 0.6 | 4.2 | GO:0001705 | ectoderm formation(GO:0001705) ectodermal cell fate commitment(GO:0001712) |
| 0.6 | 3.5 | GO:0021633 | optic nerve structural organization(GO:0021633) |
| 0.6 | 1.2 | GO:0003051 | angiotensin-mediated drinking behavior(GO:0003051) |
| 0.6 | 2.4 | GO:0034552 | respiratory chain complex II assembly(GO:0034552) mitochondrial respiratory chain complex II assembly(GO:0034553) mitochondrial respiratory chain complex II biogenesis(GO:0097032) |
| 0.6 | 1.7 | GO:0061743 | motor learning(GO:0061743) |
| 0.6 | 2.9 | GO:0014042 | positive regulation of neuron maturation(GO:0014042) |
| 0.6 | 4.0 | GO:0060373 | regulation of ventricular cardiac muscle cell membrane depolarization(GO:0060373) |
| 0.6 | 5.2 | GO:0032286 | central nervous system myelin maintenance(GO:0032286) |
| 0.6 | 3.4 | GO:0071677 | positive regulation of mononuclear cell migration(GO:0071677) |
| 0.6 | 24.0 | GO:0010107 | potassium ion import(GO:0010107) |
| 0.6 | 4.0 | GO:2000664 | positive regulation of interleukin-5 secretion(GO:2000664) positive regulation of interleukin-13 secretion(GO:2000667) |
| 0.6 | 4.0 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
| 0.6 | 3.9 | GO:1902474 | positive regulation of protein localization to synapse(GO:1902474) |
| 0.6 | 8.4 | GO:0061002 | negative regulation of dendritic spine morphogenesis(GO:0061002) |
| 0.6 | 2.2 | GO:0070537 | histone H2A K63-linked deubiquitination(GO:0070537) |
| 0.5 | 2.7 | GO:0002225 | positive regulation of antimicrobial peptide production(GO:0002225) positive regulation of antibacterial peptide production(GO:0002803) |
| 0.5 | 7.7 | GO:1902083 | negative regulation of peptidyl-cysteine S-nitrosylation(GO:1902083) |
| 0.5 | 18.1 | GO:2000311 | regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000311) |
| 0.5 | 4.3 | GO:0009313 | ganglioside catabolic process(GO:0006689) oligosaccharide catabolic process(GO:0009313) |
| 0.5 | 2.7 | GO:0032849 | regulation of cellular pH reduction(GO:0032847) positive regulation of cellular pH reduction(GO:0032849) |
| 0.5 | 3.7 | GO:0045200 | establishment or maintenance of neuroblast polarity(GO:0045196) establishment of neuroblast polarity(GO:0045200) |
| 0.5 | 2.1 | GO:0042662 | cardiogenic plate morphogenesis(GO:0003142) negative regulation of mesodermal cell fate specification(GO:0042662) regulation of transcription from RNA polymerase II promoter involved in definitive endodermal cell fate specification(GO:0060807) |
| 0.5 | 0.5 | GO:1902462 | regulation of mesenchymal stem cell proliferation(GO:1902460) positive regulation of mesenchymal stem cell proliferation(GO:1902462) |
| 0.5 | 3.1 | GO:0035585 | calcium-mediated signaling using extracellular calcium source(GO:0035585) |
| 0.5 | 3.1 | GO:0070294 | renal sodium ion absorption(GO:0070294) |
| 0.5 | 2.5 | GO:0098957 | anterograde axonal transport of mitochondrion(GO:0098957) |
| 0.5 | 2.5 | GO:0010747 | positive regulation of plasma membrane long-chain fatty acid transport(GO:0010747) |
| 0.5 | 5.5 | GO:0048715 | negative regulation of oligodendrocyte differentiation(GO:0048715) |
| 0.5 | 4.5 | GO:0060054 | positive regulation of epithelial cell proliferation involved in wound healing(GO:0060054) |
| 0.5 | 8.4 | GO:0070673 | response to interleukin-18(GO:0070673) |
| 0.5 | 1.0 | GO:0061107 | seminal vesicle development(GO:0061107) |
| 0.5 | 1.9 | GO:0046951 | ketone body biosynthetic process(GO:0046951) |
| 0.5 | 8.1 | GO:0031340 | positive regulation of vesicle fusion(GO:0031340) |
| 0.5 | 3.3 | GO:0016259 | selenocysteine metabolic process(GO:0016259) |
| 0.5 | 1.9 | GO:0072086 | specification of loop of Henle identity(GO:0072086) |
| 0.5 | 14.9 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 0.5 | 5.6 | GO:0021891 | olfactory bulb interneuron development(GO:0021891) |
| 0.5 | 2.3 | GO:0035694 | mitochondrial protein catabolic process(GO:0035694) |
| 0.5 | 1.4 | GO:1900477 | negative regulation of G1/S transition of mitotic cell cycle by negative regulation of transcription from RNA polymerase II promoter(GO:1900477) |
| 0.5 | 1.4 | GO:0035524 | proline transmembrane transport(GO:0035524) |
| 0.5 | 3.2 | GO:0001880 | Mullerian duct regression(GO:0001880) |
| 0.4 | 2.2 | GO:0031394 | positive regulation of prostaglandin biosynthetic process(GO:0031394) positive regulation of unsaturated fatty acid biosynthetic process(GO:2001280) |
| 0.4 | 1.3 | GO:0035880 | embryonic nail plate morphogenesis(GO:0035880) |
| 0.4 | 13.4 | GO:0045662 | negative regulation of myoblast differentiation(GO:0045662) |
| 0.4 | 7.1 | GO:0071257 | cellular response to electrical stimulus(GO:0071257) |
| 0.4 | 2.2 | GO:0090080 | positive regulation of MAPKKK cascade by fibroblast growth factor receptor signaling pathway(GO:0090080) |
| 0.4 | 1.3 | GO:0010615 | positive regulation of cardiac muscle adaptation(GO:0010615) positive regulation of cardiac muscle hypertrophy in response to stress(GO:1903244) |
| 0.4 | 4.9 | GO:0033227 | dsRNA transport(GO:0033227) |
| 0.4 | 2.7 | GO:0033563 | dorsal/ventral axon guidance(GO:0033563) |
| 0.4 | 4.9 | GO:0002328 | pro-B cell differentiation(GO:0002328) |
| 0.4 | 14.8 | GO:0097320 | membrane tubulation(GO:0097320) |
| 0.4 | 6.4 | GO:0070166 | enamel mineralization(GO:0070166) |
| 0.4 | 2.1 | GO:0048069 | eye pigmentation(GO:0048069) |
| 0.4 | 8.8 | GO:2000651 | positive regulation of sodium ion transmembrane transporter activity(GO:2000651) |
| 0.4 | 3.7 | GO:0098528 | skeletal muscle fiber differentiation(GO:0098528) |
| 0.4 | 0.4 | GO:1902993 | positive regulation of beta-amyloid formation(GO:1902004) positive regulation of amyloid precursor protein catabolic process(GO:1902993) |
| 0.4 | 4.2 | GO:0034058 | endosomal vesicle fusion(GO:0034058) |
| 0.4 | 1.2 | GO:0014916 | regulation of lung blood pressure(GO:0014916) |
| 0.4 | 7.0 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.4 | 3.3 | GO:0048170 | positive regulation of long-term neuronal synaptic plasticity(GO:0048170) |
| 0.4 | 9.7 | GO:0009251 | glycogen catabolic process(GO:0005980) glucan catabolic process(GO:0009251) cellular polysaccharide catabolic process(GO:0044247) |
| 0.4 | 1.2 | GO:0060157 | urinary bladder development(GO:0060157) |
| 0.4 | 9.6 | GO:0031987 | locomotion involved in locomotory behavior(GO:0031987) |
| 0.4 | 3.2 | GO:0038042 | dimeric G-protein coupled receptor signaling pathway(GO:0038042) calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.4 | 4.4 | GO:0006013 | mannose metabolic process(GO:0006013) |
| 0.4 | 9.6 | GO:0045475 | locomotor rhythm(GO:0045475) |
| 0.4 | 7.2 | GO:0055089 | fatty acid homeostasis(GO:0055089) |
| 0.4 | 6.0 | GO:0034331 | cell junction maintenance(GO:0034331) cell-cell junction maintenance(GO:0045217) |
| 0.4 | 3.6 | GO:0042904 | 9-cis-retinoic acid biosynthetic process(GO:0042904) 9-cis-retinoic acid metabolic process(GO:0042905) |
| 0.4 | 2.4 | GO:0002528 | regulation of vascular permeability involved in acute inflammatory response(GO:0002528) |
| 0.4 | 6.7 | GO:0043586 | tongue development(GO:0043586) |
| 0.4 | 4.3 | GO:1900383 | regulation of synaptic plasticity by receptor localization to synapse(GO:1900383) |
| 0.4 | 1.2 | GO:0006589 | octopamine biosynthetic process(GO:0006589) octopamine metabolic process(GO:0046333) |
| 0.4 | 0.8 | GO:0036363 | transforming growth factor beta activation(GO:0036363) |
| 0.4 | 3.5 | GO:0006102 | isocitrate metabolic process(GO:0006102) |
| 0.4 | 2.7 | GO:2000323 | negative regulation of glucocorticoid receptor signaling pathway(GO:2000323) |
| 0.4 | 1.9 | GO:0072383 | plus-end-directed vesicle transport along microtubule(GO:0072383) plus-end-directed organelle transport along microtubule(GO:0072386) |
| 0.4 | 1.1 | GO:0039663 | fusion of virus membrane with host plasma membrane(GO:0019064) membrane fusion involved in viral entry into host cell(GO:0039663) multi-organism membrane fusion(GO:0044800) |
| 0.4 | 3.8 | GO:1902669 | positive regulation of axon guidance(GO:1902669) |
| 0.4 | 1.5 | GO:0035262 | gonad morphogenesis(GO:0035262) |
| 0.4 | 7.5 | GO:2000480 | negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
| 0.4 | 6.4 | GO:1904262 | negative regulation of TORC1 signaling(GO:1904262) |
| 0.4 | 2.6 | GO:0030050 | vesicle transport along actin filament(GO:0030050) |
| 0.4 | 7.8 | GO:0046838 | phosphorylated carbohydrate dephosphorylation(GO:0046838) inositol phosphate dephosphorylation(GO:0046855) |
| 0.4 | 2.6 | GO:1990034 | calcium ion export from cell(GO:1990034) |
| 0.4 | 8.1 | GO:0031115 | negative regulation of microtubule polymerization(GO:0031115) |
| 0.4 | 0.7 | GO:1903860 | negative regulation of dendrite extension(GO:1903860) |
| 0.4 | 7.5 | GO:0016486 | peptide hormone processing(GO:0016486) |
| 0.4 | 2.8 | GO:0048227 | plasma membrane to endosome transport(GO:0048227) |
| 0.4 | 1.8 | GO:1990592 | protein polyufmylation(GO:1990564) protein K69-linked ufmylation(GO:1990592) |
| 0.3 | 4.2 | GO:2001288 | positive regulation of caveolin-mediated endocytosis(GO:2001288) |
| 0.3 | 4.2 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.3 | 2.4 | GO:0045077 | negative regulation of interferon-gamma biosynthetic process(GO:0045077) |
| 0.3 | 5.1 | GO:1901386 | negative regulation of voltage-gated calcium channel activity(GO:1901386) |
| 0.3 | 2.0 | GO:0010499 | proteasomal ubiquitin-independent protein catabolic process(GO:0010499) |
| 0.3 | 2.4 | GO:0090292 | nuclear matrix organization(GO:0043578) nuclear matrix anchoring at nuclear membrane(GO:0090292) |
| 0.3 | 2.0 | GO:0071947 | protein deubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0071947) |
| 0.3 | 2.7 | GO:2000766 | negative regulation of cytoplasmic translation(GO:2000766) |
| 0.3 | 3.0 | GO:0030828 | positive regulation of cGMP metabolic process(GO:0030825) positive regulation of cGMP biosynthetic process(GO:0030828) |
| 0.3 | 7.9 | GO:0071625 | vocalization behavior(GO:0071625) |
| 0.3 | 4.9 | GO:0045714 | regulation of low-density lipoprotein particle receptor biosynthetic process(GO:0045714) |
| 0.3 | 1.3 | GO:0044860 | protein localization to plasma membrane raft(GO:0044860) regulation of establishment of T cell polarity(GO:1903903) |
| 0.3 | 1.6 | GO:0014005 | microglia differentiation(GO:0014004) microglia development(GO:0014005) |
| 0.3 | 1.6 | GO:0010644 | cell communication by electrical coupling(GO:0010644) |
| 0.3 | 5.7 | GO:0000338 | protein deneddylation(GO:0000338) |
| 0.3 | 3.5 | GO:0051152 | positive regulation of smooth muscle cell differentiation(GO:0051152) |
| 0.3 | 8.8 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.3 | 10.8 | GO:0006084 | acetyl-CoA metabolic process(GO:0006084) |
| 0.3 | 0.9 | GO:0045226 | extracellular polysaccharide biosynthetic process(GO:0045226) extracellular polysaccharide metabolic process(GO:0046379) |
| 0.3 | 10.8 | GO:0014003 | oligodendrocyte development(GO:0014003) |
| 0.3 | 6.7 | GO:0007250 | activation of NF-kappaB-inducing kinase activity(GO:0007250) |
| 0.3 | 2.8 | GO:0021840 | directional guidance of interneurons involved in migration from the subpallium to the cortex(GO:0021840) chemorepulsion involved in interneuron migration from the subpallium to the cortex(GO:0021842) |
| 0.3 | 10.3 | GO:0007616 | long-term memory(GO:0007616) |
| 0.3 | 2.6 | GO:0003062 | regulation of heart rate by chemical signal(GO:0003062) |
| 0.3 | 2.9 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.3 | 0.9 | GO:1903251 | multi-ciliated epithelial cell differentiation(GO:1903251) |
| 0.3 | 7.5 | GO:0045956 | positive regulation of calcium ion-dependent exocytosis(GO:0045956) |
| 0.3 | 2.3 | GO:0006777 | Mo-molybdopterin cofactor biosynthetic process(GO:0006777) Mo-molybdopterin cofactor metabolic process(GO:0019720) |
| 0.3 | 2.6 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.3 | 0.9 | GO:0086046 | membrane depolarization during SA node cell action potential(GO:0086046) |
| 0.3 | 1.7 | GO:0009235 | cobalamin metabolic process(GO:0009235) |
| 0.3 | 1.1 | GO:0007571 | age-dependent response to oxidative stress(GO:0001306) age-dependent general metabolic decline(GO:0007571) |
| 0.3 | 2.2 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.3 | 1.9 | GO:0042769 | DNA damage response, detection of DNA damage(GO:0042769) |
| 0.3 | 1.1 | GO:0018406 | protein C-linked glycosylation(GO:0018103) peptidyl-tryptophan modification(GO:0018211) protein C-linked glycosylation via tryptophan(GO:0018317) protein C-linked glycosylation via 2'-alpha-mannosyl-L-tryptophan(GO:0018406) |
| 0.3 | 5.9 | GO:0060314 | regulation of ryanodine-sensitive calcium-release channel activity(GO:0060314) |
| 0.3 | 9.4 | GO:0048488 | synaptic vesicle endocytosis(GO:0048488) |
| 0.3 | 2.9 | GO:0090394 | negative regulation of excitatory postsynaptic potential(GO:0090394) regulation of retina development in camera-type eye(GO:1902866) |
| 0.3 | 0.8 | GO:0046087 | cytidine catabolic process(GO:0006216) cytidine deamination(GO:0009972) cytidine metabolic process(GO:0046087) |
| 0.3 | 1.6 | GO:0060762 | regulation of branching involved in mammary gland duct morphogenesis(GO:0060762) |
| 0.3 | 7.6 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.3 | 2.8 | GO:0060484 | lung-associated mesenchyme development(GO:0060484) |
| 0.3 | 0.8 | GO:1903378 | positive regulation of oxidative stress-induced neuron intrinsic apoptotic signaling pathway(GO:1903378) |
| 0.3 | 4.8 | GO:0071578 | zinc II ion transmembrane import(GO:0071578) |
| 0.3 | 8.6 | GO:0061003 | positive regulation of dendritic spine morphogenesis(GO:0061003) |
| 0.2 | 7.0 | GO:0045671 | negative regulation of osteoclast differentiation(GO:0045671) |
| 0.2 | 1.0 | GO:0003199 | endocardial cushion to mesenchymal transition involved in heart valve formation(GO:0003199) |
| 0.2 | 2.5 | GO:2000346 | negative regulation of hepatocyte proliferation(GO:2000346) |
| 0.2 | 9.2 | GO:0015991 | ATP hydrolysis coupled proton transport(GO:0015991) ATP hydrolysis coupled transmembrane transport(GO:0090662) |
| 0.2 | 10.1 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.2 | 0.7 | GO:0015920 | lipopolysaccharide transport(GO:0015920) |
| 0.2 | 3.6 | GO:0036065 | fucosylation(GO:0036065) |
| 0.2 | 3.6 | GO:0010642 | negative regulation of platelet-derived growth factor receptor signaling pathway(GO:0010642) |
| 0.2 | 3.1 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.2 | 1.2 | GO:0098886 | modification of dendritic spine(GO:0098886) |
| 0.2 | 2.0 | GO:0071816 | tail-anchored membrane protein insertion into ER membrane(GO:0071816) |
| 0.2 | 0.2 | GO:0072034 | renal vesicle induction(GO:0072034) |
| 0.2 | 3.1 | GO:0045741 | positive regulation of epidermal growth factor-activated receptor activity(GO:0045741) |
| 0.2 | 8.0 | GO:0045746 | negative regulation of Notch signaling pathway(GO:0045746) |
| 0.2 | 2.0 | GO:0061001 | regulation of dendritic spine morphogenesis(GO:0061001) |
| 0.2 | 1.3 | GO:0050915 | sensory perception of sour taste(GO:0050915) |
| 0.2 | 1.3 | GO:0098838 | reduced folate transmembrane transport(GO:0098838) |
| 0.2 | 1.1 | GO:0007262 | STAT protein import into nucleus(GO:0007262) |
| 0.2 | 1.9 | GO:0036465 | synaptic vesicle recycling(GO:0036465) |
| 0.2 | 4.3 | GO:0071786 | endoplasmic reticulum tubular network organization(GO:0071786) |
| 0.2 | 1.5 | GO:0032380 | regulation of intracellular lipid transport(GO:0032377) regulation of intracellular sterol transport(GO:0032380) regulation of intracellular cholesterol transport(GO:0032383) |
| 0.2 | 2.8 | GO:1901898 | negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.2 | 3.8 | GO:2001223 | negative regulation of neuron migration(GO:2001223) |
| 0.2 | 1.5 | GO:1901552 | positive regulation of endothelial cell development(GO:1901552) positive regulation of establishment of endothelial barrier(GO:1903142) |
| 0.2 | 2.5 | GO:0051967 | negative regulation of synaptic transmission, glutamatergic(GO:0051967) |
| 0.2 | 1.0 | GO:0009240 | isopentenyl diphosphate biosynthetic process(GO:0009240) isopentenyl diphosphate metabolic process(GO:0046490) |
| 0.2 | 0.2 | GO:1905165 | regulation of lysosomal protein catabolic process(GO:1905165) |
| 0.2 | 2.0 | GO:0039536 | negative regulation of RIG-I signaling pathway(GO:0039536) |
| 0.2 | 4.7 | GO:0015936 | coenzyme A metabolic process(GO:0015936) |
| 0.2 | 0.6 | GO:0070318 | myoblast development(GO:0048627) positive regulation of G0 to G1 transition(GO:0070318) |
| 0.2 | 8.1 | GO:0051351 | positive regulation of ligase activity(GO:0051351) |
| 0.2 | 1.1 | GO:0010991 | regulation of SMAD protein complex assembly(GO:0010990) negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.2 | 7.4 | GO:0032024 | positive regulation of insulin secretion(GO:0032024) |
| 0.2 | 1.1 | GO:0090038 | negative regulation of protein kinase C signaling(GO:0090038) |
| 0.2 | 1.5 | GO:0044805 | late nucleophagy(GO:0044805) |
| 0.2 | 0.7 | GO:0097498 | endothelial tube lumen extension(GO:0097498) |
| 0.2 | 10.0 | GO:0005978 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.2 | 1.1 | GO:0010815 | bradykinin catabolic process(GO:0010815) |
| 0.2 | 2.7 | GO:0046498 | S-adenosylhomocysteine metabolic process(GO:0046498) |
| 0.2 | 6.5 | GO:0048013 | ephrin receptor signaling pathway(GO:0048013) |
| 0.2 | 7.1 | GO:0071158 | positive regulation of cell cycle arrest(GO:0071158) |
| 0.2 | 3.6 | GO:0006622 | protein targeting to lysosome(GO:0006622) |
| 0.2 | 1.2 | GO:0071910 | determination of pancreatic left/right asymmetry(GO:0035469) determination of liver left/right asymmetry(GO:0071910) |
| 0.2 | 6.2 | GO:0032012 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.2 | 4.1 | GO:0060396 | growth hormone receptor signaling pathway(GO:0060396) cellular response to growth hormone stimulus(GO:0071378) |
| 0.2 | 4.1 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.2 | 4.4 | GO:0051290 | protein heterotetramerization(GO:0051290) |
| 0.2 | 4.5 | GO:0046839 | phospholipid dephosphorylation(GO:0046839) |
| 0.2 | 2.7 | GO:0042749 | regulation of circadian sleep/wake cycle(GO:0042749) regulation of circadian sleep/wake cycle, sleep(GO:0045187) |
| 0.2 | 2.1 | GO:0031290 | retinal ganglion cell axon guidance(GO:0031290) |
| 0.2 | 18.8 | GO:0055088 | lipid homeostasis(GO:0055088) |
| 0.2 | 1.9 | GO:0036158 | outer dynein arm assembly(GO:0036158) |
| 0.2 | 1.7 | GO:1990440 | positive regulation of transcription from RNA polymerase II promoter in response to endoplasmic reticulum stress(GO:1990440) |
| 0.2 | 12.6 | GO:0009311 | oligosaccharide metabolic process(GO:0009311) |
| 0.2 | 1.1 | GO:0003350 | pulmonary myocardium development(GO:0003350) |
| 0.1 | 0.1 | GO:1903588 | negative regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903588) |
| 0.1 | 8.2 | GO:0070059 | intrinsic apoptotic signaling pathway in response to endoplasmic reticulum stress(GO:0070059) |
| 0.1 | 0.4 | GO:1905168 | positive regulation of double-strand break repair via homologous recombination(GO:1905168) |
| 0.1 | 6.6 | GO:0003298 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.1 | 0.9 | GO:0019627 | urea cycle(GO:0000050) urea metabolic process(GO:0019627) |
| 0.1 | 1.2 | GO:1900029 | positive regulation of ruffle assembly(GO:1900029) |
| 0.1 | 1.0 | GO:0043922 | negative regulation by host of viral transcription(GO:0043922) |
| 0.1 | 1.0 | GO:0002043 | blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:0002043) |
| 0.1 | 0.6 | GO:0019323 | pentose catabolic process(GO:0019323) |
| 0.1 | 0.8 | GO:0051464 | cortisol secretion(GO:0043400) regulation of cortisol secretion(GO:0051462) positive regulation of cortisol secretion(GO:0051464) positive regulation of glucocorticoid secretion(GO:2000851) |
| 0.1 | 0.4 | GO:0006207 | 'de novo' pyrimidine nucleobase biosynthetic process(GO:0006207) pyrimidine nucleobase biosynthetic process(GO:0019856) |
| 0.1 | 2.0 | GO:0045672 | positive regulation of osteoclast differentiation(GO:0045672) |
| 0.1 | 5.9 | GO:0007193 | adenylate cyclase-inhibiting G-protein coupled receptor signaling pathway(GO:0007193) |
| 0.1 | 7.6 | GO:0046579 | positive regulation of Ras protein signal transduction(GO:0046579) |
| 0.1 | 0.3 | GO:0000189 | MAPK import into nucleus(GO:0000189) |
| 0.1 | 2.0 | GO:0043517 | positive regulation of DNA damage response, signal transduction by p53 class mediator(GO:0043517) |
| 0.1 | 3.8 | GO:0010758 | regulation of macrophage chemotaxis(GO:0010758) |
| 0.1 | 3.9 | GO:0046326 | positive regulation of glucose import(GO:0046326) |
| 0.1 | 16.1 | GO:0007416 | synapse assembly(GO:0007416) |
| 0.1 | 1.8 | GO:2000191 | regulation of fatty acid transport(GO:2000191) |
| 0.1 | 2.5 | GO:0048490 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.1 | 0.9 | GO:0015886 | heme transport(GO:0015886) |
| 0.1 | 1.3 | GO:0060732 | positive regulation of inositol phosphate biosynthetic process(GO:0060732) |
| 0.1 | 0.8 | GO:0010996 | response to auditory stimulus(GO:0010996) |
| 0.1 | 0.5 | GO:0070268 | cornification(GO:0070268) |
| 0.1 | 3.9 | GO:0031424 | keratinization(GO:0031424) |
| 0.1 | 2.6 | GO:2000171 | negative regulation of dendrite development(GO:2000171) |
| 0.1 | 1.6 | GO:0016264 | gap junction assembly(GO:0016264) |
| 0.1 | 2.6 | GO:0043552 | positive regulation of phosphatidylinositol 3-kinase activity(GO:0043552) |
| 0.1 | 2.9 | GO:0071480 | cellular response to gamma radiation(GO:0071480) |
| 0.1 | 7.9 | GO:0002028 | regulation of sodium ion transport(GO:0002028) |
| 0.1 | 0.2 | GO:0035526 | retrograde transport, plasma membrane to Golgi(GO:0035526) |
| 0.1 | 2.8 | GO:0035563 | positive regulation of chromatin binding(GO:0035563) |
| 0.1 | 2.2 | GO:0006878 | cellular copper ion homeostasis(GO:0006878) |
| 0.1 | 1.9 | GO:0009649 | entrainment of circadian clock(GO:0009649) |
| 0.1 | 0.5 | GO:1902269 | positive regulation of polyamine transmembrane transport(GO:1902269) |
| 0.1 | 1.6 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.1 | 2.0 | GO:0007202 | activation of phospholipase C activity(GO:0007202) |
| 0.1 | 4.1 | GO:0046677 | response to antibiotic(GO:0046677) |
| 0.1 | 0.8 | GO:2001269 | positive regulation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:2001269) |
| 0.1 | 1.8 | GO:0043162 | ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043162) |
| 0.1 | 0.3 | GO:0006500 | N-terminal protein palmitoylation(GO:0006500) negative regulation of lipoprotein metabolic process(GO:0050748) |
| 0.1 | 1.6 | GO:0042551 | neuron maturation(GO:0042551) |
| 0.1 | 0.2 | GO:0006649 | phospholipid transfer to membrane(GO:0006649) |
| 0.1 | 0.5 | GO:0033326 | cerebrospinal fluid secretion(GO:0033326) |
| 0.1 | 3.3 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.1 | 5.4 | GO:0008542 | visual learning(GO:0008542) |
| 0.1 | 2.1 | GO:0000729 | DNA double-strand break processing(GO:0000729) |
| 0.1 | 0.3 | GO:0014908 | myotube differentiation involved in skeletal muscle regeneration(GO:0014908) |
| 0.1 | 2.6 | GO:0008333 | endosome to lysosome transport(GO:0008333) |
| 0.1 | 0.3 | GO:0030327 | prenylated protein catabolic process(GO:0030327) |
| 0.1 | 3.8 | GO:0007257 | activation of JUN kinase activity(GO:0007257) |
| 0.1 | 0.6 | GO:0045198 | establishment of epithelial cell apical/basal polarity(GO:0045198) |
| 0.1 | 0.3 | GO:0098749 | cerebellar neuron development(GO:0098749) |
| 0.1 | 4.0 | GO:0071300 | cellular response to retinoic acid(GO:0071300) |
| 0.1 | 1.8 | GO:0008272 | sulfate transport(GO:0008272) |
| 0.1 | 2.0 | GO:0006152 | purine nucleoside catabolic process(GO:0006152) purine ribonucleoside catabolic process(GO:0046130) |
| 0.1 | 0.9 | GO:0048207 | vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) |
| 0.1 | 0.8 | GO:0061709 | reticulophagy(GO:0061709) |
| 0.1 | 6.5 | GO:0034605 | cellular response to heat(GO:0034605) |
| 0.1 | 6.7 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.1 | 0.6 | GO:0070213 | protein auto-ADP-ribosylation(GO:0070213) |
| 0.1 | 0.5 | GO:0006420 | arginyl-tRNA aminoacylation(GO:0006420) |
| 0.1 | 0.3 | GO:0006517 | protein deglycosylation(GO:0006517) |
| 0.1 | 0.8 | GO:0090070 | positive regulation of ribosome biogenesis(GO:0090070) positive regulation of rRNA processing(GO:2000234) |
| 0.1 | 0.3 | GO:2001106 | regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001106) |
| 0.1 | 0.9 | GO:0097186 | amelogenesis(GO:0097186) |
| 0.1 | 0.2 | GO:0035522 | monoubiquitinated histone deubiquitination(GO:0035521) monoubiquitinated histone H2A deubiquitination(GO:0035522) |
| 0.1 | 1.1 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.1 | 3.8 | GO:0042775 | mitochondrial ATP synthesis coupled electron transport(GO:0042775) |
| 0.1 | 0.6 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.1 | 0.6 | GO:0031848 | protection from non-homologous end joining at telomere(GO:0031848) |
| 0.1 | 1.8 | GO:2000785 | regulation of autophagosome assembly(GO:2000785) |
| 0.1 | 0.9 | GO:0043496 | regulation of protein homodimerization activity(GO:0043496) |
| 0.1 | 12.5 | GO:0042787 | protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0042787) |
| 0.1 | 1.9 | GO:0035329 | hippo signaling(GO:0035329) |
| 0.1 | 2.2 | GO:0043949 | regulation of cAMP-mediated signaling(GO:0043949) |
| 0.1 | 3.2 | GO:0010761 | fibroblast migration(GO:0010761) |
| 0.1 | 2.0 | GO:0006767 | water-soluble vitamin metabolic process(GO:0006767) |
| 0.1 | 0.6 | GO:1903975 | regulation of glial cell migration(GO:1903975) |
| 0.1 | 0.4 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 0.1 | 2.4 | GO:0042596 | behavioral fear response(GO:0001662) fear response(GO:0042596) |
| 0.1 | 0.6 | GO:0048630 | skeletal muscle tissue growth(GO:0048630) |
| 0.1 | 1.8 | GO:0040020 | regulation of meiotic nuclear division(GO:0040020) |
| 0.1 | 0.7 | GO:0034982 | mitochondrial protein processing(GO:0034982) |
| 0.1 | 1.0 | GO:0042982 | amyloid precursor protein metabolic process(GO:0042982) |
| 0.1 | 0.3 | GO:0044727 | DNA demethylation of male pronucleus(GO:0044727) |
| 0.1 | 0.4 | GO:1903691 | positive regulation of wound healing, spreading of epidermal cells(GO:1903691) |
| 0.1 | 1.8 | GO:0006829 | zinc II ion transport(GO:0006829) |
| 0.1 | 1.3 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.1 | 1.3 | GO:0030575 | nuclear body organization(GO:0030575) |
| 0.1 | 1.0 | GO:2000052 | positive regulation of non-canonical Wnt signaling pathway(GO:2000052) |
| 0.1 | 1.4 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.1 | 0.8 | GO:1904355 | positive regulation of telomere capping(GO:1904355) |
| 0.1 | 0.1 | GO:0030035 | microspike assembly(GO:0030035) positive regulation of endocytic recycling(GO:2001137) |
| 0.1 | 0.9 | GO:0019933 | cAMP-mediated signaling(GO:0019933) |
| 0.1 | 0.2 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.1 | 1.5 | GO:0003016 | respiratory system process(GO:0003016) |
| 0.1 | 0.3 | GO:0035735 | intraciliary transport involved in cilium morphogenesis(GO:0035735) |
| 0.1 | 1.4 | GO:0007212 | dopamine receptor signaling pathway(GO:0007212) |
| 0.1 | 1.2 | GO:0034063 | stress granule assembly(GO:0034063) |
| 0.1 | 2.0 | GO:0035418 | protein localization to synapse(GO:0035418) |
| 0.1 | 10.1 | GO:0006457 | protein folding(GO:0006457) |
| 0.1 | 5.1 | GO:0001578 | microtubule bundle formation(GO:0001578) |
| 0.1 | 2.9 | GO:0060428 | lung epithelium development(GO:0060428) |
| 0.0 | 1.2 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.0 | 0.7 | GO:0043983 | histone H4-K12 acetylation(GO:0043983) |
| 0.0 | 0.5 | GO:0070475 | rRNA base methylation(GO:0070475) |
| 0.0 | 1.5 | GO:0021680 | cerebellar Purkinje cell layer development(GO:0021680) |
| 0.0 | 0.2 | GO:0030043 | actin filament fragmentation(GO:0030043) |
| 0.0 | 1.7 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 0.0 | 2.7 | GO:0015992 | proton transport(GO:0015992) |
| 0.0 | 3.4 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.0 | 0.5 | GO:0031468 | nuclear envelope reassembly(GO:0031468) |
| 0.0 | 4.7 | GO:0042471 | ear morphogenesis(GO:0042471) |
| 0.0 | 3.3 | GO:1901379 | regulation of potassium ion transmembrane transport(GO:1901379) |
| 0.0 | 1.2 | GO:0033141 | positive regulation of peptidyl-serine phosphorylation of STAT protein(GO:0033141) |
| 0.0 | 1.2 | GO:0019228 | neuronal action potential(GO:0019228) |
| 0.0 | 2.2 | GO:0055013 | cardiac muscle cell development(GO:0055013) |
| 0.0 | 1.4 | GO:0006687 | glycosphingolipid metabolic process(GO:0006687) |
| 0.0 | 0.8 | GO:0022027 | interkinetic nuclear migration(GO:0022027) |
| 0.0 | 1.5 | GO:0031365 | N-terminal protein amino acid modification(GO:0031365) |
| 0.0 | 1.4 | GO:0007602 | phototransduction(GO:0007602) |
| 0.0 | 4.4 | GO:0000910 | cytokinesis(GO:0000910) |
| 0.0 | 0.5 | GO:0007220 | Notch receptor processing(GO:0007220) |
| 0.0 | 12.6 | GO:0045666 | positive regulation of neuron differentiation(GO:0045666) |
| 0.0 | 1.8 | GO:0051865 | protein autoubiquitination(GO:0051865) |
| 0.0 | 1.0 | GO:0007020 | microtubule nucleation(GO:0007020) |
| 0.0 | 2.3 | GO:0032543 | mitochondrial translation(GO:0032543) |
| 0.0 | 1.9 | GO:0000045 | autophagosome assembly(GO:0000045) |
| 0.0 | 0.5 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.0 | 0.6 | GO:0032967 | positive regulation of collagen biosynthetic process(GO:0032967) |
| 0.0 | 0.9 | GO:0070266 | necroptotic process(GO:0070266) |
| 0.0 | 0.3 | GO:0036444 | calcium ion transmembrane import into mitochondrion(GO:0036444) |
| 0.0 | 0.6 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 0.0 | 0.4 | GO:0014887 | muscle hypertrophy in response to stress(GO:0003299) cardiac muscle adaptation(GO:0014887) cardiac muscle hypertrophy in response to stress(GO:0014898) |
| 0.0 | 0.2 | GO:0019262 | N-acetylneuraminate catabolic process(GO:0019262) |
| 0.0 | 0.1 | GO:1903575 | cornified envelope assembly(GO:1903575) |
| 0.0 | 4.1 | GO:0019236 | response to pheromone(GO:0019236) |
| 0.0 | 1.0 | GO:0070936 | protein K48-linked ubiquitination(GO:0070936) |
| 0.0 | 0.8 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.0 | 0.2 | GO:0075525 | viral translational termination-reinitiation(GO:0075525) |
| 0.0 | 1.0 | GO:2000279 | negative regulation of DNA biosynthetic process(GO:2000279) |
| 0.0 | 1.0 | GO:0001825 | blastocyst formation(GO:0001825) |
| 0.0 | 1.5 | GO:0016525 | negative regulation of angiogenesis(GO:0016525) |
| 0.0 | 0.5 | GO:0043620 | regulation of DNA-templated transcription in response to stress(GO:0043620) |
| 0.0 | 1.1 | GO:0050909 | sensory perception of taste(GO:0050909) |
| 0.0 | 1.7 | GO:0035914 | skeletal muscle cell differentiation(GO:0035914) |
| 0.0 | 0.1 | GO:0006012 | galactose metabolic process(GO:0006012) |
| 0.0 | 0.5 | GO:0043113 | receptor clustering(GO:0043113) |
| 0.0 | 4.5 | GO:0007409 | axonogenesis(GO:0007409) |
| 0.0 | 0.5 | GO:0042531 | positive regulation of tyrosine phosphorylation of STAT protein(GO:0042531) |
| 0.0 | 1.3 | GO:0042439 | ethanolamine-containing compound metabolic process(GO:0042439) |
| 0.0 | 0.7 | GO:0019217 | regulation of fatty acid metabolic process(GO:0019217) |
| 0.0 | 0.3 | GO:0043507 | positive regulation of JUN kinase activity(GO:0043507) |
| 0.0 | 0.1 | GO:0006481 | C-terminal protein methylation(GO:0006481) |
| 0.0 | 2.2 | GO:0031032 | actomyosin structure organization(GO:0031032) |
| 0.0 | 0.3 | GO:0016126 | sterol biosynthetic process(GO:0016126) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.3 | 13.1 | GO:0099573 | glutamatergic postsynaptic density(GO:0099573) |
| 2.8 | 8.4 | GO:0034774 | secretory granule lumen(GO:0034774) cytoplasmic membrane-bounded vesicle lumen(GO:0060205) |
| 2.7 | 13.3 | GO:0044316 | cone cell pedicle(GO:0044316) |
| 2.6 | 13.1 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 2.3 | 20.6 | GO:0008282 | ATP-sensitive potassium channel complex(GO:0008282) |
| 1.9 | 52.2 | GO:0005922 | connexon complex(GO:0005922) |
| 1.7 | 5.2 | GO:0097233 | lamellar body membrane(GO:0097232) alveolar lamellar body membrane(GO:0097233) |
| 1.3 | 9.2 | GO:0097513 | myosin II filament(GO:0097513) |
| 1.1 | 12.5 | GO:0045179 | apical cortex(GO:0045179) |
| 1.1 | 3.4 | GO:1904602 | serotonin-activated cation-selective channel complex(GO:1904602) |
| 1.1 | 11.8 | GO:0042587 | glycogen granule(GO:0042587) |
| 1.0 | 5.1 | GO:0038039 | G-protein coupled receptor heterodimeric complex(GO:0038039) |
| 1.0 | 5.0 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 1.0 | 8.1 | GO:0033269 | internode region of axon(GO:0033269) |
| 0.9 | 8.5 | GO:0097442 | CA3 pyramidal cell dendrite(GO:0097442) |
| 0.9 | 4.5 | GO:0000308 | cytoplasmic cyclin-dependent protein kinase holoenzyme complex(GO:0000308) |
| 0.9 | 13.5 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.9 | 2.6 | GO:1990844 | subsarcolemmal mitochondrion(GO:1990843) interfibrillar mitochondrion(GO:1990844) |
| 0.8 | 8.2 | GO:0042567 | insulin-like growth factor ternary complex(GO:0042567) |
| 0.8 | 4.9 | GO:0005958 | DNA-dependent protein kinase-DNA ligase 4 complex(GO:0005958) DNA ligase IV complex(GO:0032807) |
| 0.8 | 17.6 | GO:0032279 | asymmetric synapse(GO:0032279) |
| 0.8 | 3.2 | GO:0097447 | dendritic tree(GO:0097447) |
| 0.8 | 36.2 | GO:0030673 | axolemma(GO:0030673) |
| 0.8 | 11.5 | GO:0033179 | proton-transporting V-type ATPase, V0 domain(GO:0033179) |
| 0.7 | 45.4 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.7 | 25.0 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.7 | 5.8 | GO:0097025 | MPP7-DLG1-LIN7 complex(GO:0097025) |
| 0.7 | 8.7 | GO:0043203 | axon hillock(GO:0043203) |
| 0.7 | 19.9 | GO:1902711 | GABA receptor complex(GO:1902710) GABA-A receptor complex(GO:1902711) |
| 0.7 | 5.6 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.7 | 2.0 | GO:0036501 | UFD1-NPL4 complex(GO:0036501) |
| 0.7 | 1.3 | GO:0031673 | H zone(GO:0031673) |
| 0.6 | 1.9 | GO:0048179 | activin receptor complex(GO:0048179) |
| 0.6 | 9.4 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.6 | 6.2 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.6 | 42.2 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.6 | 7.1 | GO:0016011 | dystroglycan complex(GO:0016011) |
| 0.6 | 14.1 | GO:0098839 | postsynaptic density membrane(GO:0098839) |
| 0.6 | 6.1 | GO:0030314 | junctional membrane complex(GO:0030314) |
| 0.5 | 2.7 | GO:0097209 | epidermal lamellar body(GO:0097209) |
| 0.5 | 3.2 | GO:0097648 | G-protein coupled receptor dimeric complex(GO:0038037) G-protein coupled receptor complex(GO:0097648) calcitonin family receptor complex(GO:1903439) amylin receptor complex(GO:1903440) |
| 0.5 | 3.7 | GO:0070695 | FHF complex(GO:0070695) |
| 0.5 | 4.2 | GO:0033263 | CORVET complex(GO:0033263) |
| 0.5 | 14.5 | GO:0005782 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.5 | 6.8 | GO:0060091 | kinocilium(GO:0060091) |
| 0.5 | 3.9 | GO:0017071 | intracellular cyclic nucleotide activated cation channel complex(GO:0017071) |
| 0.5 | 7.5 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.5 | 2.8 | GO:0090498 | extrinsic component of Golgi membrane(GO:0090498) |
| 0.5 | 8.9 | GO:0045180 | basal cortex(GO:0045180) |
| 0.4 | 3.6 | GO:0031313 | extrinsic component of endosome membrane(GO:0031313) |
| 0.4 | 1.8 | GO:0097637 | intrinsic component of autophagosome membrane(GO:0097636) integral component of autophagosome membrane(GO:0097637) |
| 0.4 | 1.3 | GO:0019008 | molybdopterin synthase complex(GO:0019008) |
| 0.4 | 3.7 | GO:0061689 | tricellular tight junction(GO:0061689) |
| 0.4 | 16.9 | GO:0032809 | neuronal cell body membrane(GO:0032809) |
| 0.4 | 12.8 | GO:0032839 | dendrite cytoplasm(GO:0032839) |
| 0.4 | 3.3 | GO:0032983 | kainate selective glutamate receptor complex(GO:0032983) |
| 0.4 | 4.3 | GO:1990761 | growth cone lamellipodium(GO:1990761) growth cone filopodium(GO:1990812) |
| 0.4 | 7.9 | GO:0030285 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.4 | 11.0 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) |
| 0.3 | 11.5 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.3 | 1.4 | GO:0005594 | collagen type IX trimer(GO:0005594) |
| 0.3 | 9.6 | GO:0030137 | COPI-coated vesicle(GO:0030137) |
| 0.3 | 2.2 | GO:0070552 | BRISC complex(GO:0070552) |
| 0.3 | 10.4 | GO:0045334 | clathrin-coated endocytic vesicle(GO:0045334) |
| 0.3 | 1.2 | GO:0097444 | spine apparatus(GO:0097444) |
| 0.3 | 0.9 | GO:0036488 | CHOP-C/EBP complex(GO:0036488) |
| 0.3 | 7.0 | GO:0043196 | varicosity(GO:0043196) |
| 0.3 | 3.4 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.3 | 2.6 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.3 | 3.7 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.3 | 8.7 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.3 | 5.1 | GO:0012510 | trans-Golgi network transport vesicle membrane(GO:0012510) |
| 0.3 | 0.8 | GO:0017109 | glutamate-cysteine ligase complex(GO:0017109) |
| 0.3 | 2.7 | GO:0072687 | meiotic spindle(GO:0072687) |
| 0.3 | 2.4 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.3 | 5.0 | GO:0036057 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.3 | 24.1 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.3 | 6.7 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.3 | 4.7 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.3 | 3.1 | GO:0033270 | paranode region of axon(GO:0033270) |
| 0.3 | 1.3 | GO:0071439 | clathrin complex(GO:0071439) |
| 0.2 | 3.5 | GO:0044294 | dendritic growth cone(GO:0044294) |
| 0.2 | 7.2 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.2 | 2.4 | GO:0030478 | actin cap(GO:0030478) |
| 0.2 | 3.6 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.2 | 5.0 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.2 | 1.8 | GO:0014731 | spectrin-associated cytoskeleton(GO:0014731) |
| 0.2 | 2.0 | GO:0072379 | BAT3 complex(GO:0071818) ER membrane insertion complex(GO:0072379) |
| 0.2 | 4.0 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.2 | 26.1 | GO:0014704 | intercalated disc(GO:0014704) |
| 0.2 | 2.8 | GO:0032584 | growth cone membrane(GO:0032584) |
| 0.2 | 2.4 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.2 | 6.7 | GO:0071782 | endoplasmic reticulum tubular network(GO:0071782) |
| 0.2 | 1.5 | GO:0005964 | phosphorylase kinase complex(GO:0005964) |
| 0.2 | 2.1 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.2 | 46.7 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.2 | 45.7 | GO:0070382 | exocytic vesicle(GO:0070382) |
| 0.2 | 2.6 | GO:0031045 | dense core granule(GO:0031045) |
| 0.2 | 10.1 | GO:1990204 | oxidoreductase complex(GO:1990204) |
| 0.2 | 0.2 | GO:0031310 | integral component of vacuolar membrane(GO:0031166) intrinsic component of vacuolar membrane(GO:0031310) |
| 0.2 | 2.4 | GO:0034993 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.2 | 1.3 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.2 | 2.4 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.2 | 0.9 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.2 | 0.7 | GO:0005745 | m-AAA complex(GO:0005745) |
| 0.2 | 25.1 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.2 | 12.4 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.2 | 3.0 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.2 | 5.4 | GO:0030118 | clathrin coat(GO:0030118) |
| 0.2 | 0.8 | GO:0034751 | aryl hydrocarbon receptor complex(GO:0034751) |
| 0.2 | 2.1 | GO:0030660 | Golgi-associated vesicle membrane(GO:0030660) |
| 0.1 | 42.9 | GO:0097060 | synaptic membrane(GO:0097060) |
| 0.1 | 11.9 | GO:0034705 | potassium channel complex(GO:0034705) |
| 0.1 | 0.4 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.1 | 5.9 | GO:0045271 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.1 | 0.8 | GO:0070876 | SOSS complex(GO:0070876) |
| 0.1 | 1.3 | GO:0097451 | glial limiting end-foot(GO:0097451) |
| 0.1 | 7.3 | GO:0016529 | sarcoplasmic reticulum(GO:0016529) |
| 0.1 | 1.8 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.1 | 13.9 | GO:0043204 | perikaryon(GO:0043204) |
| 0.1 | 4.2 | GO:0031304 | intrinsic component of mitochondrial inner membrane(GO:0031304) integral component of mitochondrial inner membrane(GO:0031305) |
| 0.1 | 7.5 | GO:0031301 | integral component of organelle membrane(GO:0031301) |
| 0.1 | 20.3 | GO:0031227 | intrinsic component of endoplasmic reticulum membrane(GO:0031227) |
| 0.1 | 0.5 | GO:0097058 | CRLF-CLCF1 complex(GO:0097058) |
| 0.1 | 9.5 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.1 | 18.4 | GO:0014069 | postsynaptic density(GO:0014069) postsynaptic specialization(GO:0099572) |
| 0.1 | 0.6 | GO:0071547 | piP-body(GO:0071547) |
| 0.1 | 1.2 | GO:0030140 | trans-Golgi network transport vesicle(GO:0030140) |
| 0.1 | 5.5 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.1 | 1.0 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 0.1 | 0.5 | GO:0034272 | phosphatidylinositol 3-kinase complex, class III, type I(GO:0034271) phosphatidylinositol 3-kinase complex, class III, type II(GO:0034272) |
| 0.1 | 2.3 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.1 | 2.0 | GO:0030008 | TRAPP complex(GO:0030008) |
| 0.1 | 38.2 | GO:0019866 | organelle inner membrane(GO:0019866) |
| 0.1 | 3.7 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.1 | 7.4 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.1 | 32.8 | GO:0030425 | dendrite(GO:0030425) |
| 0.1 | 16.9 | GO:0031966 | mitochondrial membrane(GO:0031966) |
| 0.1 | 0.9 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.1 | 1.3 | GO:0002080 | acrosomal membrane(GO:0002080) |
| 0.1 | 1.4 | GO:0032982 | myosin filament(GO:0032982) |
| 0.1 | 0.4 | GO:0030870 | Mre11 complex(GO:0030870) |
| 0.1 | 0.9 | GO:0034464 | BBSome(GO:0034464) |
| 0.1 | 0.6 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.1 | 0.4 | GO:0034706 | sodium channel complex(GO:0034706) |
| 0.1 | 3.2 | GO:0030427 | site of polarized growth(GO:0030427) |
| 0.1 | 0.7 | GO:0005921 | gap junction(GO:0005921) |
| 0.1 | 0.4 | GO:0000835 | ER ubiquitin ligase complex(GO:0000835) Hrd1p ubiquitin ligase complex(GO:0000836) |
| 0.1 | 11.7 | GO:0000139 | Golgi membrane(GO:0000139) |
| 0.1 | 2.5 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.1 | 0.5 | GO:0070765 | gamma-secretase complex(GO:0070765) |
| 0.0 | 0.9 | GO:0000421 | autophagosome membrane(GO:0000421) |
| 0.0 | 0.5 | GO:0000815 | ESCRT III complex(GO:0000815) |
| 0.0 | 2.0 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.0 | 2.5 | GO:0005776 | autophagosome(GO:0005776) |
| 0.0 | 0.2 | GO:0035339 | SPOTS complex(GO:0035339) |
| 0.0 | 1.1 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.0 | 1.5 | GO:0000314 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.0 | 0.7 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.0 | 0.2 | GO:0097208 | alveolar lamellar body(GO:0097208) |
| 0.0 | 4.5 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.0 | 5.3 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 3.0 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.0 | 1.3 | GO:0098793 | presynapse(GO:0098793) |
| 0.0 | 2.2 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 7.5 | GO:0045202 | synapse(GO:0045202) |
| 0.0 | 0.9 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.0 | 0.6 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.0 | 39.4 | GO:0005739 | mitochondrion(GO:0005739) |
| 0.0 | 0.3 | GO:0001939 | female pronucleus(GO:0001939) male pronucleus(GO:0001940) |
| 0.0 | 1.9 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 0.6 | GO:0005697 | telomerase holoenzyme complex(GO:0005697) |
| 0.0 | 0.5 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.0 | 0.2 | GO:0072669 | tRNA-splicing ligase complex(GO:0072669) |
| 0.0 | 0.2 | GO:0071986 | Ragulator complex(GO:0071986) |
| 0.0 | 0.6 | GO:0032040 | small-subunit processome(GO:0032040) |
| 0.0 | 0.2 | GO:0045495 | P granule(GO:0043186) pole plasm(GO:0045495) germ plasm(GO:0060293) |
| 0.0 | 100.4 | GO:0031224 | intrinsic component of membrane(GO:0031224) |
| 0.0 | 5.6 | GO:0045177 | apical part of cell(GO:0045177) |
| 0.0 | 0.2 | GO:0032039 | integrator complex(GO:0032039) |
| 0.0 | 0.5 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.0 | 2.3 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.0 | 0.3 | GO:0005671 | Ada2/Gcn5/Ada3 transcription activator complex(GO:0005671) |
| 0.0 | 0.2 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.0 | 0.1 | GO:1990131 | Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.0 | 0.2 | GO:0005852 | eukaryotic translation initiation factor 3 complex(GO:0005852) |
| 0.0 | 0.3 | GO:0032391 | photoreceptor connecting cilium(GO:0032391) |
| 0.0 | 1.5 | GO:0001650 | fibrillar center(GO:0001650) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 10.1 | 40.4 | GO:0004478 | methionine adenosyltransferase activity(GO:0004478) |
| 5.6 | 22.5 | GO:0047016 | cholest-5-ene-3-beta,7-alpha-diol 3-beta-dehydrogenase activity(GO:0047016) |
| 5.3 | 16.0 | GO:0004698 | calcium-dependent protein kinase C activity(GO:0004698) |
| 4.2 | 12.6 | GO:0016901 | glycerol-3-phosphate dehydrogenase activity(GO:0004368) oxidoreductase activity, acting on the CH-OH group of donors, quinone or similar compound as acceptor(GO:0016901) |
| 4.2 | 12.6 | GO:0047291 | lactosylceramide alpha-2,3-sialyltransferase activity(GO:0047291) |
| 3.6 | 10.7 | GO:0004001 | adenosine kinase activity(GO:0004001) |
| 3.5 | 10.4 | GO:0034632 | retinol transporter activity(GO:0034632) |
| 3.3 | 10.0 | GO:0004967 | glucagon receptor activity(GO:0004967) |
| 3.3 | 19.6 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 3.2 | 9.7 | GO:0004134 | glycogen debranching enzyme activity(GO:0004133) 4-alpha-glucanotransferase activity(GO:0004134) amylo-alpha-1,6-glucosidase activity(GO:0004135) |
| 3.0 | 11.9 | GO:0016743 | carboxyl- or carbamoyltransferase activity(GO:0016743) |
| 2.7 | 24.2 | GO:0005094 | Rho GDP-dissociation inhibitor activity(GO:0005094) |
| 2.6 | 20.6 | GO:0015272 | ATP-activated inward rectifier potassium channel activity(GO:0015272) |
| 2.6 | 12.8 | GO:0004021 | L-alanine:2-oxoglutarate aminotransferase activity(GO:0004021) alanine-oxo-acid transaminase activity(GO:0047635) |
| 2.4 | 9.5 | GO:0004096 | catalase activity(GO:0004096) |
| 2.1 | 6.4 | GO:0015152 | glucose-6-phosphate transmembrane transporter activity(GO:0015152) |
| 2.1 | 15.0 | GO:0004351 | glutamate decarboxylase activity(GO:0004351) |
| 2.1 | 22.9 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 2.0 | 36.7 | GO:0005243 | gap junction channel activity(GO:0005243) |
| 2.0 | 20.3 | GO:0030550 | acetylcholine receptor inhibitor activity(GO:0030550) |
| 1.9 | 9.7 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 1.9 | 5.7 | GO:0001639 | PLC activating G-protein coupled glutamate receptor activity(GO:0001639) G-protein coupled receptor activity involved in regulation of postsynaptic membrane potential(GO:0099530) |
| 1.8 | 9.0 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 1.8 | 5.3 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 1.7 | 5.2 | GO:0005289 | high-affinity basic amino acid transmembrane transporter activity(GO:0005287) high-affinity arginine transmembrane transporter activity(GO:0005289) high-affinity lysine transmembrane transporter activity(GO:0005292) |
| 1.6 | 4.8 | GO:0005353 | fructose transmembrane transporter activity(GO:0005353) uniporter activity(GO:0015292) |
| 1.5 | 9.2 | GO:0004994 | somatostatin receptor activity(GO:0004994) |
| 1.5 | 10.5 | GO:0001642 | group III metabotropic glutamate receptor activity(GO:0001642) |
| 1.4 | 21.1 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 1.4 | 5.6 | GO:1902379 | chemoattractant activity involved in axon guidance(GO:1902379) |
| 1.4 | 4.2 | GO:0017082 | mineralocorticoid receptor activity(GO:0017082) |
| 1.3 | 3.8 | GO:0001565 | phorbol ester receptor activity(GO:0001565) non-kinase phorbol ester receptor activity(GO:0001566) |
| 1.2 | 4.9 | GO:0031694 | alpha-2A adrenergic receptor binding(GO:0031694) |
| 1.2 | 14.7 | GO:0030976 | thiamine pyrophosphate binding(GO:0030976) |
| 1.2 | 3.7 | GO:1990699 | palmitoleyl hydrolase activity(GO:1990699) |
| 1.2 | 7.3 | GO:0034190 | apolipoprotein receptor binding(GO:0034190) |
| 1.2 | 7.1 | GO:0015111 | iodide transmembrane transporter activity(GO:0015111) |
| 1.2 | 3.5 | GO:0098782 | mechanically-gated potassium channel activity(GO:0098782) |
| 1.2 | 4.6 | GO:0071614 | linoleic acid epoxygenase activity(GO:0071614) |
| 1.2 | 3.5 | GO:0051538 | 3 iron, 4 sulfur cluster binding(GO:0051538) |
| 1.1 | 5.7 | GO:0003986 | acetyl-CoA hydrolase activity(GO:0003986) |
| 1.1 | 9.0 | GO:0005237 | inhibitory extracellular ligand-gated ion channel activity(GO:0005237) |
| 1.1 | 3.3 | GO:0003858 | 3-hydroxybutyrate dehydrogenase activity(GO:0003858) |
| 1.1 | 3.3 | GO:0070279 | vitamin B6 binding(GO:0070279) |
| 1.1 | 3.2 | GO:0015229 | L-ascorbate:sodium symporter activity(GO:0008520) L-ascorbic acid transporter activity(GO:0015229) sodium-dependent L-ascorbate transmembrane transporter activity(GO:0070890) |
| 1.0 | 8.3 | GO:0043237 | laminin-1 binding(GO:0043237) |
| 1.0 | 6.2 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 1.0 | 6.1 | GO:0008453 | alanine-glyoxylate transaminase activity(GO:0008453) |
| 1.0 | 5.0 | GO:0003872 | 6-phosphofructokinase activity(GO:0003872) |
| 1.0 | 7.8 | GO:0016312 | inositol bisphosphate phosphatase activity(GO:0016312) |
| 1.0 | 5.7 | GO:0015277 | kainate selective glutamate receptor activity(GO:0015277) |
| 0.9 | 8.5 | GO:0005519 | cytoskeletal regulatory protein binding(GO:0005519) |
| 0.9 | 7.4 | GO:0004416 | hydroxyacylglutathione hydrolase activity(GO:0004416) |
| 0.9 | 3.7 | GO:0004937 | alpha1-adrenergic receptor activity(GO:0004937) |
| 0.9 | 6.3 | GO:0008294 | calcium- and calmodulin-responsive adenylate cyclase activity(GO:0008294) |
| 0.9 | 8.9 | GO:0004396 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) |
| 0.9 | 5.3 | GO:0008269 | JAK pathway signal transduction adaptor activity(GO:0008269) |
| 0.9 | 6.1 | GO:0005332 | gamma-aminobutyric acid:sodium symporter activity(GO:0005332) |
| 0.9 | 4.3 | GO:0004308 | exo-alpha-sialidase activity(GO:0004308) alpha-sialidase activity(GO:0016997) exo-alpha-(2->3)-sialidase activity(GO:0052794) exo-alpha-(2->6)-sialidase activity(GO:0052795) exo-alpha-(2->8)-sialidase activity(GO:0052796) |
| 0.8 | 5.1 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.8 | 8.4 | GO:0008140 | cAMP response element binding protein binding(GO:0008140) |
| 0.8 | 5.8 | GO:0097016 | L27 domain binding(GO:0097016) |
| 0.8 | 10.0 | GO:0102338 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.8 | 2.5 | GO:0004968 | gonadotropin-releasing hormone receptor activity(GO:0004968) |
| 0.8 | 4.9 | GO:0003839 | gamma-glutamylcyclotransferase activity(GO:0003839) |
| 0.8 | 4.9 | GO:0051032 | nucleic acid transmembrane transporter activity(GO:0051032) RNA transmembrane transporter activity(GO:0051033) |
| 0.8 | 7.3 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.8 | 3.2 | GO:0045183 | translation factor activity, non-nucleic acid binding(GO:0045183) |
| 0.8 | 13.0 | GO:0016594 | glycine binding(GO:0016594) |
| 0.7 | 14.5 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.7 | 4.3 | GO:0008273 | calcium, potassium:sodium antiporter activity(GO:0008273) |
| 0.7 | 10.1 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.7 | 4.3 | GO:0004706 | JUN kinase kinase kinase activity(GO:0004706) |
| 0.7 | 0.7 | GO:0003696 | satellite DNA binding(GO:0003696) |
| 0.7 | 2.8 | GO:0031752 | D5 dopamine receptor binding(GO:0031752) |
| 0.7 | 13.6 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.7 | 6.1 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.7 | 2.0 | GO:0016297 | [acyl-carrier-protein] S-malonyltransferase activity(GO:0004314) oleoyl-[acyl-carrier-protein] hydrolase activity(GO:0004320) myristoyl-[acyl-carrier-protein] hydrolase activity(GO:0016295) palmitoyl-[acyl-carrier-protein] hydrolase activity(GO:0016296) acyl-[acyl-carrier-protein] hydrolase activity(GO:0016297) S-acetyltransferase activity(GO:0016418) S-malonyltransferase activity(GO:0016419) malonyltransferase activity(GO:0016420) phosphopantetheine binding(GO:0031177) |
| 0.7 | 3.4 | GO:0022850 | serotonin-gated cation channel activity(GO:0022850) |
| 0.7 | 6.6 | GO:0022851 | GABA-gated chloride ion channel activity(GO:0022851) |
| 0.6 | 3.9 | GO:0005222 | intracellular cAMP activated cation channel activity(GO:0005222) intracellular cGMP activated cation channel activity(GO:0005223) |
| 0.6 | 3.9 | GO:0004792 | thiosulfate sulfurtransferase activity(GO:0004792) |
| 0.6 | 3.9 | GO:0051425 | PTB domain binding(GO:0051425) |
| 0.6 | 5.1 | GO:0060072 | large conductance calcium-activated potassium channel activity(GO:0060072) |
| 0.6 | 3.2 | GO:0051185 | coenzyme transporter activity(GO:0051185) |
| 0.6 | 8.3 | GO:0008603 | cAMP-dependent protein kinase regulator activity(GO:0008603) |
| 0.6 | 5.7 | GO:0004768 | stearoyl-CoA 9-desaturase activity(GO:0004768) |
| 0.6 | 5.1 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.6 | 1.2 | GO:0019238 | cyclohydrolase activity(GO:0019238) |
| 0.6 | 3.7 | GO:0047035 | testosterone dehydrogenase (NAD+) activity(GO:0047035) |
| 0.6 | 17.5 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.6 | 3.6 | GO:0046920 | alpha-(1->3)-fucosyltransferase activity(GO:0046920) |
| 0.6 | 2.4 | GO:0016822 | oxaloacetate decarboxylase activity(GO:0008948) hydrolase activity, acting on acid carbon-carbon bonds(GO:0016822) hydrolase activity, acting on acid carbon-carbon bonds, in ketonic substances(GO:0016823) |
| 0.6 | 7.1 | GO:0030955 | potassium ion binding(GO:0030955) |
| 0.6 | 2.2 | GO:0004566 | beta-glucuronidase activity(GO:0004566) |
| 0.6 | 3.9 | GO:0050833 | pyruvate transmembrane transporter activity(GO:0050833) |
| 0.6 | 2.2 | GO:0034617 | tetrahydrobiopterin binding(GO:0034617) |
| 0.6 | 2.8 | GO:0016715 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced ascorbate as one donor, and incorporation of one atom of oxygen(GO:0016715) |
| 0.5 | 0.5 | GO:0030899 | calcium-dependent ATPase activity(GO:0030899) |
| 0.5 | 2.7 | GO:0004719 | protein-L-isoaspartate (D-aspartate) O-methyltransferase activity(GO:0004719) |
| 0.5 | 3.2 | GO:0097643 | amylin receptor activity(GO:0097643) |
| 0.5 | 2.7 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.5 | 3.1 | GO:1990599 | 3' overhang single-stranded DNA endodeoxyribonuclease activity(GO:1990599) |
| 0.5 | 8.2 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.5 | 4.0 | GO:0086006 | voltage-gated sodium channel activity involved in cardiac muscle cell action potential(GO:0086006) |
| 0.5 | 2.0 | GO:0047710 | bis(5'-adenosyl)-triphosphatase activity(GO:0047710) |
| 0.5 | 5.8 | GO:0051378 | serotonin binding(GO:0051378) |
| 0.5 | 1.9 | GO:0038100 | nodal binding(GO:0038100) |
| 0.5 | 1.9 | GO:0016833 | oxo-acid-lyase activity(GO:0016833) |
| 0.5 | 11.4 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
| 0.5 | 4.3 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.5 | 3.2 | GO:0086056 | voltage-gated calcium channel activity involved in AV node cell action potential(GO:0086056) |
| 0.5 | 4.1 | GO:0071253 | connexin binding(GO:0071253) |
| 0.5 | 14.9 | GO:0008392 | arachidonic acid epoxygenase activity(GO:0008392) |
| 0.4 | 2.2 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.4 | 3.6 | GO:0004366 | glycerol-3-phosphate O-acyltransferase activity(GO:0004366) |
| 0.4 | 3.1 | GO:0061629 | RNA polymerase II sequence-specific DNA binding transcription factor binding(GO:0061629) |
| 0.4 | 4.0 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.4 | 3.1 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.4 | 2.2 | GO:0016499 | orexin receptor activity(GO:0016499) |
| 0.4 | 15.1 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.4 | 1.7 | GO:0004609 | phosphatidylserine decarboxylase activity(GO:0004609) |
| 0.4 | 2.1 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.4 | 0.9 | GO:0086059 | voltage-gated calcium channel activity involved SA node cell action potential(GO:0086059) |
| 0.4 | 12.2 | GO:0016769 | transferase activity, transferring nitrogenous groups(GO:0016769) |
| 0.4 | 5.0 | GO:0036374 | glutathione hydrolase activity(GO:0036374) |
| 0.4 | 2.5 | GO:0033192 | calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.4 | 2.1 | GO:0005283 | sodium:amino acid symporter activity(GO:0005283) |
| 0.4 | 2.0 | GO:0008853 | exodeoxyribonuclease III activity(GO:0008853) |
| 0.4 | 3.8 | GO:0016421 | CoA carboxylase activity(GO:0016421) |
| 0.4 | 2.2 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.4 | 1.1 | GO:0003875 | ADP-ribosylarginine hydrolase activity(GO:0003875) |
| 0.4 | 14.8 | GO:0036442 | hydrogen-exporting ATPase activity(GO:0036442) |
| 0.4 | 12.5 | GO:0030552 | cAMP binding(GO:0030552) |
| 0.4 | 13.6 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.4 | 16.6 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.4 | 2.1 | GO:0046624 | sphingolipid transporter activity(GO:0046624) |
| 0.4 | 2.1 | GO:0070326 | very-low-density lipoprotein particle receptor binding(GO:0070326) |
| 0.4 | 8.1 | GO:0031078 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.4 | 7.0 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.3 | 5.9 | GO:0048406 | nerve growth factor binding(GO:0048406) |
| 0.3 | 8.6 | GO:0030169 | low-density lipoprotein particle binding(GO:0030169) |
| 0.3 | 1.4 | GO:0050436 | microfibril binding(GO:0050436) |
| 0.3 | 1.0 | GO:0008330 | protein tyrosine/threonine phosphatase activity(GO:0008330) |
| 0.3 | 4.5 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.3 | 4.8 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.3 | 5.1 | GO:0015271 | outward rectifier potassium channel activity(GO:0015271) |
| 0.3 | 15.8 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.3 | 6.3 | GO:0005003 | ephrin receptor activity(GO:0005003) |
| 0.3 | 6.6 | GO:0045499 | chemorepellent activity(GO:0045499) |
| 0.3 | 4.9 | GO:0019871 | sodium channel inhibitor activity(GO:0019871) |
| 0.3 | 0.9 | GO:0031871 | proteinase activated receptor binding(GO:0031871) |
| 0.3 | 1.2 | GO:0004995 | tachykinin receptor activity(GO:0004995) |
| 0.3 | 13.1 | GO:0016645 | oxidoreductase activity, acting on the CH-NH group of donors(GO:0016645) |
| 0.3 | 1.1 | GO:0033906 | hyaluronoglucuronidase activity(GO:0033906) |
| 0.3 | 4.4 | GO:0019855 | calcium channel inhibitor activity(GO:0019855) |
| 0.3 | 0.8 | GO:0004357 | glutamate-cysteine ligase activity(GO:0004357) |
| 0.3 | 6.3 | GO:0031210 | phosphatidylcholine binding(GO:0031210) |
| 0.3 | 22.0 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.3 | 2.2 | GO:0016653 | oxidoreductase activity, acting on NAD(P)H, heme protein as acceptor(GO:0016653) |
| 0.3 | 0.5 | GO:0004367 | glycerol-3-phosphate dehydrogenase [NAD+] activity(GO:0004367) |
| 0.3 | 5.0 | GO:0010181 | FMN binding(GO:0010181) |
| 0.3 | 5.8 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.3 | 13.1 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.3 | 5.3 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.2 | 4.2 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 0.2 | 2.2 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.2 | 6.1 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.2 | 5.6 | GO:0030296 | protein tyrosine kinase activator activity(GO:0030296) |
| 0.2 | 0.9 | GO:0050501 | hyaluronan synthase activity(GO:0050501) |
| 0.2 | 1.8 | GO:0031821 | G-protein coupled serotonin receptor binding(GO:0031821) |
| 0.2 | 2.7 | GO:0005025 | transforming growth factor beta receptor activity, type I(GO:0005025) |
| 0.2 | 18.2 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.2 | 3.9 | GO:0016247 | channel regulator activity(GO:0016247) |
| 0.2 | 9.1 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
| 0.2 | 11.0 | GO:0035255 | ionotropic glutamate receptor binding(GO:0035255) |
| 0.2 | 1.8 | GO:0045134 | uridine-diphosphatase activity(GO:0045134) |
| 0.2 | 4.4 | GO:0004559 | alpha-mannosidase activity(GO:0004559) |
| 0.2 | 2.8 | GO:0046790 | virion binding(GO:0046790) |
| 0.2 | 1.3 | GO:0008518 | reduced folate carrier activity(GO:0008518) |
| 0.2 | 1.1 | GO:0043559 | insulin binding(GO:0043559) |
| 0.2 | 5.2 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.2 | 11.8 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.2 | 4.0 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.2 | 2.9 | GO:0015093 | ferrous iron transmembrane transporter activity(GO:0015093) |
| 0.2 | 1.0 | GO:0004452 | isopentenyl-diphosphate delta-isomerase activity(GO:0004452) |
| 0.2 | 1.0 | GO:0030151 | molybdenum ion binding(GO:0030151) |
| 0.2 | 5.4 | GO:0008527 | taste receptor activity(GO:0008527) |
| 0.2 | 0.8 | GO:0047874 | dolichyldiphosphatase activity(GO:0047874) |
| 0.2 | 8.1 | GO:0046966 | thyroid hormone receptor binding(GO:0046966) |
| 0.2 | 2.5 | GO:0051428 | peptide hormone receptor binding(GO:0051428) |
| 0.2 | 1.9 | GO:0008889 | glycerophosphodiester phosphodiesterase activity(GO:0008889) |
| 0.2 | 2.8 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 0.2 | 7.1 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.2 | 7.7 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.2 | 7.0 | GO:0016709 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen(GO:0016709) |
| 0.2 | 1.3 | GO:0004991 | parathyroid hormone receptor activity(GO:0004991) |
| 0.2 | 6.4 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.2 | 1.2 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.2 | 1.5 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.2 | 4.1 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.2 | 8.8 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.2 | 5.3 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) |
| 0.2 | 2.6 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.2 | 6.9 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.2 | 1.0 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 0.2 | 0.6 | GO:0005006 | epidermal growth factor-activated receptor activity(GO:0005006) |
| 0.2 | 2.5 | GO:0008190 | eukaryotic initiation factor 4E binding(GO:0008190) |
| 0.2 | 0.5 | GO:0042978 | ornithine decarboxylase activator activity(GO:0042978) |
| 0.1 | 0.9 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.1 | 3.0 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.1 | 2.7 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.1 | 6.2 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.1 | 1.7 | GO:0015280 | ligand-gated sodium channel activity(GO:0015280) |
| 0.1 | 1.7 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.1 | 0.8 | GO:0004966 | galanin receptor activity(GO:0004966) |
| 0.1 | 1.5 | GO:0016004 | phospholipase activator activity(GO:0016004) |
| 0.1 | 1.6 | GO:0036435 | K48-linked polyubiquitin binding(GO:0036435) |
| 0.1 | 12.2 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.1 | 2.2 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.1 | 0.5 | GO:0050347 | trans-hexaprenyltranstransferase activity(GO:0000010) trans-octaprenyltranstransferase activity(GO:0050347) |
| 0.1 | 4.5 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.1 | 3.8 | GO:0015459 | potassium channel regulator activity(GO:0015459) |
| 0.1 | 8.3 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.1 | 0.4 | GO:0000179 | rRNA (adenine-N6,N6-)-dimethyltransferase activity(GO:0000179) |
| 0.1 | 0.4 | GO:0016520 | growth hormone-releasing hormone receptor activity(GO:0016520) |
| 0.1 | 6.5 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.1 | 4.5 | GO:0035254 | glutamate receptor binding(GO:0035254) |
| 0.1 | 1.0 | GO:0034711 | inhibin binding(GO:0034711) |
| 0.1 | 8.1 | GO:0016879 | ligase activity, forming carbon-nitrogen bonds(GO:0016879) |
| 0.1 | 4.0 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.1 | 37.7 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.1 | 0.7 | GO:0045322 | unmethylated CpG binding(GO:0045322) |
| 0.1 | 7.7 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.1 | 4.7 | GO:0000983 | transcription factor activity, RNA polymerase II core promoter sequence-specific(GO:0000983) |
| 0.1 | 3.8 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.1 | 4.8 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.1 | 2.4 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.1 | 2.8 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.1 | 0.5 | GO:0016434 | rRNA (cytosine) methyltransferase activity(GO:0016434) |
| 0.1 | 1.3 | GO:0097322 | 7SK snRNA binding(GO:0097322) |
| 0.1 | 61.0 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 0.1 | 1.6 | GO:0030742 | GTP-dependent protein binding(GO:0030742) |
| 0.1 | 14.7 | GO:0005516 | calmodulin binding(GO:0005516) |
| 0.1 | 3.9 | GO:0030332 | cyclin binding(GO:0030332) |
| 0.1 | 2.4 | GO:0005521 | lamin binding(GO:0005521) |
| 0.1 | 5.3 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.1 | 0.5 | GO:0004814 | arginine-tRNA ligase activity(GO:0004814) |
| 0.1 | 0.9 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.1 | 0.6 | GO:0097157 | pre-mRNA intronic binding(GO:0097157) |
| 0.1 | 0.8 | GO:0043024 | ribosomal small subunit binding(GO:0043024) |
| 0.1 | 0.4 | GO:0008502 | melatonin receptor activity(GO:0008502) |
| 0.1 | 3.2 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.1 | 1.5 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.1 | 1.4 | GO:0015250 | water channel activity(GO:0015250) |
| 0.1 | 3.8 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 7.0 | GO:0008081 | phosphoric diester hydrolase activity(GO:0008081) |
| 0.1 | 9.7 | GO:0051219 | phosphoprotein binding(GO:0051219) |
| 0.1 | 0.6 | GO:0035312 | 5'-3' exodeoxyribonuclease activity(GO:0035312) |
| 0.1 | 2.5 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 0.1 | 2.5 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.1 | 0.6 | GO:0097027 | ubiquitin-protein transferase activator activity(GO:0097027) |
| 0.1 | 2.8 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.1 | 2.3 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.1 | 1.4 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.1 | 0.6 | GO:0034235 | GPI anchor binding(GO:0034235) |
| 0.1 | 9.0 | GO:0005178 | integrin binding(GO:0005178) |
| 0.1 | 3.5 | GO:0008536 | Ran GTPase binding(GO:0008536) |
| 0.1 | 0.2 | GO:0045127 | N-acetylglucosamine kinase activity(GO:0045127) |
| 0.1 | 0.2 | GO:0090555 | phosphatidylethanolamine-translocating ATPase activity(GO:0090555) |
| 0.1 | 11.1 | GO:0005244 | voltage-gated ion channel activity(GO:0005244) voltage-gated channel activity(GO:0022832) |
| 0.1 | 2.9 | GO:0036002 | pre-mRNA binding(GO:0036002) |
| 0.1 | 4.8 | GO:0032947 | protein complex scaffold(GO:0032947) |
| 0.1 | 0.6 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.1 | 0.9 | GO:0001013 | RNA polymerase I regulatory region DNA binding(GO:0001013) |
| 0.1 | 0.8 | GO:0098505 | G-rich strand telomeric DNA binding(GO:0098505) |
| 0.1 | 0.1 | GO:0010861 | thyroid hormone receptor activator activity(GO:0010861) thyroid hormone receptor coactivator activity(GO:0030375) |
| 0.1 | 1.3 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.1 | 16.4 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.1 | 0.5 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.1 | 4.2 | GO:0005179 | hormone activity(GO:0005179) |
| 0.1 | 0.8 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.1 | 1.2 | GO:0005132 | type I interferon receptor binding(GO:0005132) |
| 0.1 | 0.5 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.1 | 2.8 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.0 | 2.0 | GO:0051539 | 4 iron, 4 sulfur cluster binding(GO:0051539) |
| 0.0 | 2.6 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.0 | 7.4 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 1.3 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.0 | 0.8 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.0 | 0.5 | GO:0005127 | ciliary neurotrophic factor receptor binding(GO:0005127) |
| 0.0 | 0.3 | GO:0015278 | calcium-release channel activity(GO:0015278) ligand-gated calcium channel activity(GO:0099604) |
| 0.0 | 0.7 | GO:0034185 | apolipoprotein binding(GO:0034185) |
| 0.0 | 0.7 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.0 | 0.9 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.0 | 0.1 | GO:0004716 | receptor signaling protein tyrosine kinase activity(GO:0004716) |
| 0.0 | 2.6 | GO:0042626 | ATPase activity, coupled to transmembrane movement of substances(GO:0042626) |
| 0.0 | 1.8 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.0 | 8.4 | GO:0008168 | methyltransferase activity(GO:0008168) |
| 0.0 | 0.3 | GO:0070579 | methylcytosine dioxygenase activity(GO:0070579) |
| 0.0 | 0.6 | GO:0015037 | peptide disulfide oxidoreductase activity(GO:0015037) |
| 0.0 | 12.3 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.0 | 0.1 | GO:0016635 | succinate dehydrogenase (ubiquinone) activity(GO:0008177) oxidoreductase activity, acting on the CH-CH group of donors, quinone or related compound as acceptor(GO:0016635) |
| 0.0 | 0.3 | GO:0016670 | oxidoreductase activity, acting on a sulfur group of donors, oxygen as acceptor(GO:0016670) |
| 0.0 | 1.1 | GO:0008188 | neuropeptide receptor activity(GO:0008188) |
| 0.0 | 1.7 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 2.1 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.0 | 1.4 | GO:0019213 | deacetylase activity(GO:0019213) |
| 0.0 | 0.4 | GO:0042288 | MHC class I protein binding(GO:0042288) |
| 0.0 | 0.1 | GO:0015440 | peptide-transporting ATPase activity(GO:0015440) |
| 0.0 | 4.1 | GO:0016503 | pheromone receptor activity(GO:0016503) |
| 0.0 | 2.0 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.0 | 0.6 | GO:0050750 | low-density lipoprotein particle receptor binding(GO:0050750) |
| 0.0 | 2.7 | GO:0031072 | heat shock protein binding(GO:0031072) |
| 0.0 | 0.7 | GO:0000062 | fatty-acyl-CoA binding(GO:0000062) |
| 0.0 | 0.9 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.0 | 0.4 | GO:0004383 | guanylate cyclase activity(GO:0004383) |
| 0.0 | 0.6 | GO:0015238 | drug transmembrane transporter activity(GO:0015238) |
| 0.0 | 1.8 | GO:0031490 | chromatin DNA binding(GO:0031490) |
| 0.0 | 0.7 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.5 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
| 0.0 | 0.2 | GO:0016413 | O-acetyltransferase activity(GO:0016413) |
| 0.0 | 1.8 | GO:0008080 | N-acetyltransferase activity(GO:0008080) |
| 0.0 | 0.4 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.0 | 0.5 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.0 | 1.4 | GO:0002020 | protease binding(GO:0002020) |
| 0.0 | 0.4 | GO:0051018 | protein kinase A binding(GO:0051018) |
| 0.0 | 1.0 | GO:0005261 | cation channel activity(GO:0005261) |
| 0.0 | 3.9 | GO:0005549 | odorant binding(GO:0005549) |
| 0.0 | 0.6 | GO:0005507 | copper ion binding(GO:0005507) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 11.9 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 0.8 | 9.5 | PID FOXO PATHWAY | FoxO family signaling |
| 0.6 | 20.4 | ST WNT CA2 CYCLIC GMP PATHWAY | Wnt/Ca2+/cyclic GMP signaling. |
| 0.5 | 42.1 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.5 | 16.5 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.5 | 10.9 | PID IGF1 PATHWAY | IGF1 pathway |
| 0.4 | 10.1 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.4 | 31.5 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.4 | 11.2 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.4 | 28.2 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.3 | 19.7 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.3 | 6.6 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.3 | 21.3 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.3 | 14.5 | PID ATR PATHWAY | ATR signaling pathway |
| 0.2 | 17.4 | PID HDAC CLASSII PATHWAY | Signaling events mediated by HDAC Class II |
| 0.2 | 4.7 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.2 | 19.6 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.2 | 8.0 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.2 | 2.2 | PID ER NONGENOMIC PATHWAY | Plasma membrane estrogen receptor signaling |
| 0.2 | 6.6 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.2 | 8.4 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.2 | 1.5 | PID S1P S1P2 PATHWAY | S1P2 pathway |
| 0.2 | 5.6 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.2 | 7.1 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.2 | 2.4 | PID ERBB1 RECEPTOR PROXIMAL PATHWAY | EGF receptor (ErbB1) signaling pathway |
| 0.2 | 9.3 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.2 | 1.0 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.2 | 2.0 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.2 | 6.8 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.2 | 4.2 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.2 | 2.3 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.2 | 7.6 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.2 | 2.8 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.2 | 4.6 | PID ATM PATHWAY | ATM pathway |
| 0.1 | 4.7 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.1 | 1.9 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.1 | 11.0 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.1 | 3.4 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.1 | 6.3 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.1 | 5.0 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.1 | 1.2 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.1 | 9.0 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.1 | 1.5 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.1 | 1.6 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.1 | 5.5 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.1 | 17.2 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.1 | 1.7 | PID BMP PATHWAY | BMP receptor signaling |
| 0.1 | 1.5 | PID IL2 1PATHWAY | IL2-mediated signaling events |
| 0.1 | 2.0 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.1 | 0.9 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.0 | 0.8 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.0 | 1.4 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 1.3 | SIG BCR SIGNALING PATHWAY | Members of the BCR signaling pathway |
| 0.0 | 1.9 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.0 | 2.8 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 0.7 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.0 | 0.7 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.0 | 0.7 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.0 | 1.0 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.0 | 0.7 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 2.8 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.0 | 8.9 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 2.3 | 52.2 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 2.2 | 33.6 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 2.2 | 4.3 | REACTOME CELL CELL COMMUNICATION | Genes involved in Cell-Cell communication |
| 1.3 | 45.1 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 1.0 | 20.9 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 1.0 | 23.4 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 1.0 | 20.2 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.9 | 18.5 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.8 | 41.3 | REACTOME TRAFFICKING OF AMPA RECEPTORS | Genes involved in Trafficking of AMPA receptors |
| 0.7 | 10.0 | REACTOME GLUCAGON SIGNALING IN METABOLIC REGULATION | Genes involved in Glucagon signaling in metabolic regulation |
| 0.6 | 9.8 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.6 | 8.5 | REACTOME REGULATION OF WATER BALANCE BY RENAL AQUAPORINS | Genes involved in Regulation of Water Balance by Renal Aquaporins |
| 0.6 | 15.0 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.6 | 6.7 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.5 | 6.6 | REACTOME CRMPS IN SEMA3A SIGNALING | Genes involved in CRMPs in Sema3A signaling |
| 0.5 | 11.1 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.5 | 9.5 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.5 | 5.8 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.5 | 39.6 | REACTOME TRIGLYCERIDE BIOSYNTHESIS | Genes involved in Triglyceride Biosynthesis |
| 0.5 | 11.9 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.5 | 12.8 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.4 | 15.7 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.4 | 13.9 | REACTOME PKA MEDIATED PHOSPHORYLATION OF CREB | Genes involved in PKA-mediated phosphorylation of CREB |
| 0.4 | 27.9 | REACTOME INWARDLY RECTIFYING K CHANNELS | Genes involved in Inwardly rectifying K+ channels |
| 0.4 | 5.7 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.4 | 14.7 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.4 | 12.7 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.4 | 2.8 | REACTOME THROMBIN SIGNALLING THROUGH PROTEINASE ACTIVATED RECEPTORS PARS | Genes involved in Thrombin signalling through proteinase activated receptors (PARs) |
| 0.4 | 8.8 | REACTOME REGULATION OF INSULIN SECRETION BY GLUCAGON LIKE PEPTIDE1 | Genes involved in Regulation of Insulin Secretion by Glucagon-like Peptide-1 |
| 0.4 | 8.9 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.4 | 9.9 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |
| 0.4 | 7.8 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.3 | 7.0 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.3 | 68.7 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.3 | 9.6 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.3 | 19.0 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.3 | 4.3 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.3 | 8.6 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.3 | 3.4 | REACTOME LIGAND GATED ION CHANNEL TRANSPORT | Genes involved in Ligand-gated ion channel transport |
| 0.3 | 6.5 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.3 | 6.8 | REACTOME RIP MEDIATED NFKB ACTIVATION VIA DAI | Genes involved in RIP-mediated NFkB activation via DAI |
| 0.3 | 1.8 | REACTOME G PROTEIN ACTIVATION | Genes involved in G-protein activation |
| 0.3 | 8.4 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.3 | 2.3 | REACTOME SIGNALLING TO P38 VIA RIT AND RIN | Genes involved in Signalling to p38 via RIT and RIN |
| 0.2 | 2.7 | REACTOME INHIBITION OF INSULIN SECRETION BY ADRENALINE NORADRENALINE | Genes involved in Inhibition of Insulin Secretion by Adrenaline/Noradrenaline |
| 0.2 | 5.0 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.2 | 40.7 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.2 | 3.7 | REACTOME SHC1 EVENTS IN ERBB4 SIGNALING | Genes involved in SHC1 events in ERBB4 signaling |
| 0.2 | 3.6 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.2 | 2.5 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.2 | 3.1 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.2 | 2.2 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.2 | 3.9 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.2 | 12.4 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.2 | 6.6 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.2 | 15.7 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.2 | 20.9 | REACTOME TRANSMISSION ACROSS CHEMICAL SYNAPSES | Genes involved in Transmission across Chemical Synapses |
| 0.2 | 5.5 | REACTOME ERK MAPK TARGETS | Genes involved in ERK/MAPK targets |
| 0.2 | 4.1 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.2 | 14.7 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.2 | 25.2 | REACTOME G ALPHA Q SIGNALLING EVENTS | Genes involved in G alpha (q) signalling events |
| 0.2 | 4.9 | REACTOME DOUBLE STRAND BREAK REPAIR | Genes involved in Double-Strand Break Repair |
| 0.2 | 1.6 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.2 | 4.6 | REACTOME RORA ACTIVATES CIRCADIAN EXPRESSION | Genes involved in RORA Activates Circadian Expression |
| 0.2 | 3.8 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.2 | 2.4 | REACTOME DESTABILIZATION OF MRNA BY AUF1 HNRNP D0 | Genes involved in Destabilization of mRNA by AUF1 (hnRNP D0) |
| 0.1 | 4.7 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.1 | 1.1 | REACTOME ENERGY DEPENDENT REGULATION OF MTOR BY LKB1 AMPK | Genes involved in Energy dependent regulation of mTOR by LKB1-AMPK |
| 0.1 | 1.9 | REACTOME NEF MEDIATED DOWNREGULATION OF MHC CLASS I COMPLEX CELL SURFACE EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.1 | 1.4 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.1 | 2.8 | REACTOME FGFR4 LIGAND BINDING AND ACTIVATION | Genes involved in FGFR4 ligand binding and activation |
| 0.1 | 2.2 | REACTOME SHC MEDIATED CASCADE | Genes involved in SHC-mediated cascade |
| 0.1 | 1.3 | REACTOME NCAM SIGNALING FOR NEURITE OUT GROWTH | Genes involved in NCAM signaling for neurite out-growth |
| 0.1 | 1.8 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.1 | 1.7 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 0.1 | 2.1 | REACTOME HYALURONAN METABOLISM | Genes involved in Hyaluronan metabolism |
| 0.1 | 3.8 | REACTOME INSULIN RECEPTOR RECYCLING | Genes involved in Insulin receptor recycling |
| 0.1 | 2.0 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.1 | 5.5 | REACTOME GROWTH HORMONE RECEPTOR SIGNALING | Genes involved in Growth hormone receptor signaling |
| 0.1 | 3.4 | REACTOME FORMATION OF FIBRIN CLOT CLOTTING CASCADE | Genes involved in Formation of Fibrin Clot (Clotting Cascade) |
| 0.1 | 1.8 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.1 | 19.4 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.1 | 1.8 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.1 | 3.3 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.1 | 5.9 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.1 | 3.6 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.1 | 6.8 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.1 | 2.7 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.1 | 2.7 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 3.4 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.1 | 1.0 | REACTOME CIRCADIAN CLOCK | Genes involved in Circadian Clock |
| 0.1 | 1.7 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.0 | 0.7 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.0 | 2.4 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.0 | 5.6 | REACTOME FATTY ACID TRIACYLGLYCEROL AND KETONE BODY METABOLISM | Genes involved in Fatty acid, triacylglycerol, and ketone body metabolism |
| 0.0 | 0.7 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 0.9 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.0 | 1.1 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 0.5 | REACTOME NUCLEAR SIGNALING BY ERBB4 | Genes involved in Nuclear signaling by ERBB4 |
| 0.0 | 0.5 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |
| 0.0 | 0.4 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |