Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
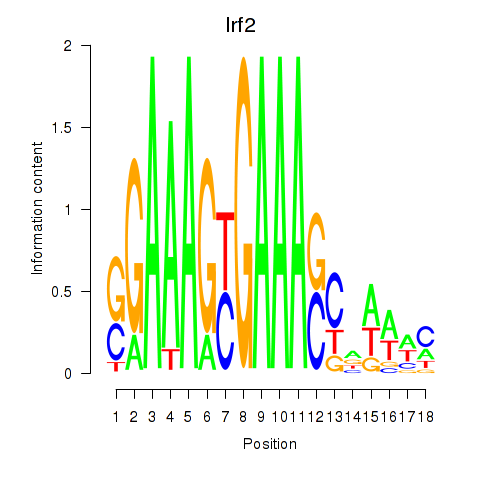
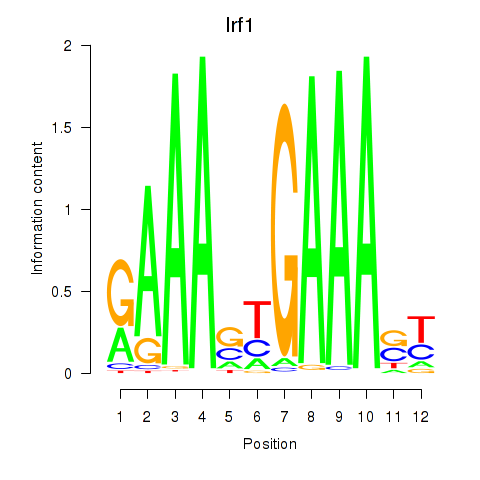
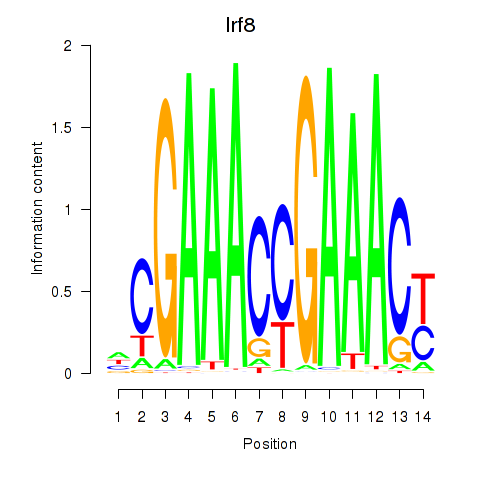
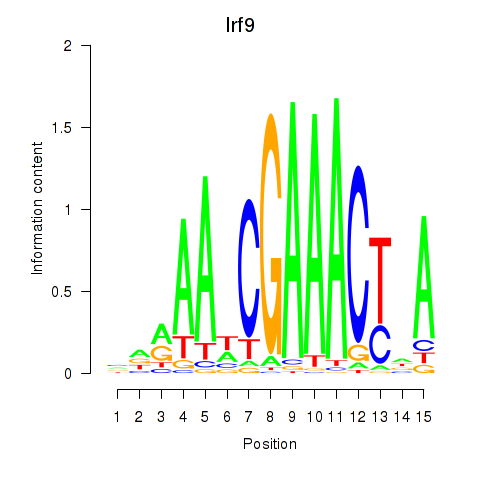
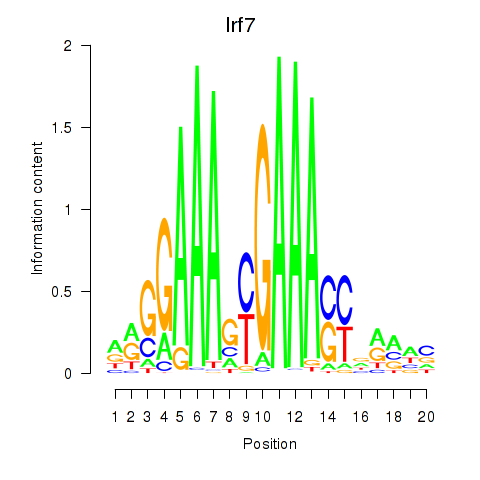
Results for Irf2_Irf1_Irf8_Irf9_Irf7
Z-value: 4.12
Transcription factors associated with Irf2_Irf1_Irf8_Irf9_Irf7
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Irf2
|
ENSMUSG00000031627.10 | Irf2 |
|
Irf1
|
ENSMUSG00000018899.18 | Irf1 |
|
Irf8
|
ENSMUSG00000041515.11 | Irf8 |
|
Irf9
|
ENSMUSG00000002325.16 | Irf9 |
|
Irf7
|
ENSMUSG00000025498.16 | Irf7 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Irf7 | mm39_v1_chr7_-_140846328_140846394 | 0.91 | 3.7e-29 | Click! |
| Irf1 | mm39_v1_chr11_+_53660834_53660886 | 0.88 | 3.4e-24 | Click! |
| Irf9 | mm39_v1_chr14_+_55842002_55842036 | 0.83 | 1.1e-19 | Click! |
| Irf8 | mm39_v1_chr8_+_121463090_121463124 | 0.72 | 6.2e-13 | Click! |
| Irf2 | mm39_v1_chr8_+_47192767_47192829 | 0.56 | 2.9e-07 | Click! |
Activity profile of Irf2_Irf1_Irf8_Irf9_Irf7 motif
Sorted Z-values of Irf2_Irf1_Irf8_Irf9_Irf7 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Irf2_Irf1_Irf8_Irf9_Irf7
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr11_+_48977852 | 51.69 |
ENSMUST00000046704.7
ENSMUST00000203810.3 ENSMUST00000203149.3 |
Ifi47
Olfr56
|
interferon gamma inducible protein 47 olfactory receptor 56 |
| chr4_-_156285247 | 48.34 |
ENSMUST00000085425.6
|
Isg15
|
ISG15 ubiquitin-like modifier |
| chr6_+_121222865 | 45.89 |
ENSMUST00000032198.11
|
Usp18
|
ubiquitin specific peptidase 18 |
| chr11_+_48977888 | 45.28 |
ENSMUST00000214804.2
|
Ifi47
|
interferon gamma inducible protein 47 |
| chr13_-_113237505 | 45.16 |
ENSMUST00000224282.2
ENSMUST00000023897.7 |
Gzma
|
granzyme A |
| chr2_-_173060647 | 44.53 |
ENSMUST00000109116.3
ENSMUST00000029018.14 |
Zbp1
|
Z-DNA binding protein 1 |
| chr7_-_140846328 | 44.23 |
ENSMUST00000106023.8
ENSMUST00000097952.9 ENSMUST00000026571.11 |
Irf7
|
interferon regulatory factor 7 |
| chr3_+_142326363 | 42.34 |
ENSMUST00000165774.8
|
Gbp2
|
guanylate binding protein 2 |
| chr14_+_78141679 | 37.71 |
ENSMUST00000022591.16
ENSMUST00000169978.2 ENSMUST00000227903.2 |
Epsti1
|
epithelial stromal interaction 1 (breast) |
| chr17_-_34406193 | 35.27 |
ENSMUST00000173831.3
|
Psmb9
|
proteasome (prosome, macropain) subunit, beta type 9 (large multifunctional peptidase 2) |
| chr1_-_173426640 | 35.16 |
ENSMUST00000150649.9
ENSMUST00000180215.2 ENSMUST00000097462.9 |
Ifi213
|
interferon activated gene 213 |
| chr11_-_70350783 | 34.97 |
ENSMUST00000019064.9
|
Cxcl16
|
chemokine (C-X-C motif) ligand 16 |
| chr11_+_119283887 | 34.26 |
ENSMUST00000093902.12
ENSMUST00000131035.10 |
Rnf213
|
ring finger protein 213 |
| chr16_+_35758836 | 33.93 |
ENSMUST00000114878.8
|
Parp9
|
poly (ADP-ribose) polymerase family, member 9 |
| chr1_+_173501215 | 33.75 |
ENSMUST00000085876.12
|
Ifi208
|
interferon activated gene 208 |
| chr1_-_173770010 | 33.42 |
ENSMUST00000042228.15
ENSMUST00000081216.12 ENSMUST00000129829.8 ENSMUST00000123708.8 |
Ifi203
|
interferon activated gene 203 |
| chr15_-_77417512 | 33.20 |
ENSMUST00000062562.7
ENSMUST00000230863.2 |
Apol7c
|
apolipoprotein L 7c |
| chr2_+_121978156 | 32.73 |
ENSMUST00000102476.5
|
B2m
|
beta-2 microglobulin |
| chr1_+_85577709 | 31.70 |
ENSMUST00000155094.8
ENSMUST00000054279.15 ENSMUST00000147552.8 ENSMUST00000153574.8 ENSMUST00000150967.8 |
Sp100
|
nuclear antigen Sp100 |
| chr19_+_34618271 | 31.37 |
ENSMUST00000102824.4
|
Ifit1
|
interferon-induced protein with tetratricopeptide repeats 1 |
| chr1_-_173318607 | 30.95 |
ENSMUST00000160565.4
|
Ifi206
|
interferon activated gene 206 |
| chr1_+_85577766 | 30.61 |
ENSMUST00000066427.11
|
Sp100
|
nuclear antigen Sp100 |
| chr16_+_35759346 | 30.57 |
ENSMUST00000023622.13
|
Parp9
|
poly (ADP-ribose) polymerase family, member 9 |
| chr1_-_173859321 | 30.15 |
ENSMUST00000059226.7
|
Ifi205
|
interferon activated gene 205 |
| chr1_-_173707677 | 29.85 |
ENSMUST00000190651.4
ENSMUST00000188804.7 |
Mndal
|
myeloid nuclear differentiation antigen like |
| chr6_+_145067457 | 28.95 |
ENSMUST00000032396.13
|
Lrmp
|
lymphoid-restricted membrane protein |
| chr18_+_60515755 | 28.37 |
ENSMUST00000237185.2
|
Iigp1
|
interferon inducible GTPase 1 |
| chr9_+_107852733 | 27.82 |
ENSMUST00000035216.11
|
Uba7
|
ubiquitin-like modifier activating enzyme 7 |
| chr19_+_29344846 | 27.45 |
ENSMUST00000016640.8
|
Cd274
|
CD274 antigen |
| chr3_+_27371168 | 27.39 |
ENSMUST00000046383.12
|
Tnfsf10
|
tumor necrosis factor (ligand) superfamily, member 10 |
| chr3_-_151455514 | 27.35 |
ENSMUST00000029671.9
|
Ifi44
|
interferon-induced protein 44 |
| chr11_-_78875689 | 27.33 |
ENSMUST00000108269.10
ENSMUST00000108268.10 |
Lgals9
|
lectin, galactose binding, soluble 9 |
| chr19_+_56385531 | 26.95 |
ENSMUST00000026062.10
|
Casp7
|
caspase 7 |
| chr11_-_48955028 | 26.14 |
ENSMUST00000046745.7
|
Tgtp2
|
T cell specific GTPase 2 |
| chr7_+_103893658 | 26.11 |
ENSMUST00000106849.9
ENSMUST00000060315.12 |
Trim34a
|
tripartite motif-containing 34A |
| chr11_-_78875657 | 25.88 |
ENSMUST00000073001.5
|
Lgals9
|
lectin, galactose binding, soluble 9 |
| chr1_-_85526517 | 25.45 |
ENSMUST00000093508.7
|
Sp110
|
Sp110 nuclear body protein |
| chr8_-_71990085 | 25.13 |
ENSMUST00000051672.9
|
Bst2
|
bone marrow stromal cell antigen 2 |
| chr15_-_76127600 | 24.75 |
ENSMUST00000165738.8
ENSMUST00000075689.7 |
Parp10
|
poly (ADP-ribose) polymerase family, member 10 |
| chr16_-_35759461 | 24.75 |
ENSMUST00000081933.14
ENSMUST00000114885.3 |
Dtx3l
|
deltex 3-like, E3 ubiquitin ligase |
| chr4_-_40239700 | 24.38 |
ENSMUST00000142055.2
|
Ddx58
|
DEAD/H box helicase 58 |
| chr7_+_103893672 | 24.01 |
ENSMUST00000106848.8
|
Trim34a
|
tripartite motif-containing 34A |
| chr16_+_23428650 | 23.46 |
ENSMUST00000038423.6
ENSMUST00000211349.2 |
Rtp4
|
receptor transporter protein 4 |
| chr17_+_34406523 | 23.37 |
ENSMUST00000170086.8
|
Tap1
|
transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) |
| chr11_+_46701619 | 23.21 |
ENSMUST00000068877.7
|
Timd4
|
T cell immunoglobulin and mucin domain containing 4 |
| chr11_-_48883053 | 23.08 |
ENSMUST00000059930.3
ENSMUST00000068063.4 |
Gm12185
Tgtp1
|
predicted gene 12185 T cell specific GTPase 1 |
| chr1_-_173363523 | 22.98 |
ENSMUST00000139092.8
|
Ifi214
|
interferon activated gene 214 |
| chr7_-_140590605 | 22.90 |
ENSMUST00000026565.7
|
Ifitm3
|
interferon induced transmembrane protein 3 |
| chr4_-_40239778 | 22.83 |
ENSMUST00000037907.13
|
Ddx58
|
DEAD/H box helicase 58 |
| chr15_+_75734159 | 22.68 |
ENSMUST00000023238.6
ENSMUST00000230514.2 ENSMUST00000229331.2 |
Gsdmd
|
gasdermin D |
| chr18_+_60345152 | 22.30 |
ENSMUST00000031549.6
|
Gm4951
|
predicted gene 4951 |
| chr4_-_155013002 | 22.29 |
ENSMUST00000152687.8
ENSMUST00000137803.8 ENSMUST00000145296.2 |
Tnfrsf14
|
tumor necrosis factor receptor superfamily, member 14 (herpesvirus entry mediator) |
| chrX_-_9335525 | 22.06 |
ENSMUST00000015484.10
|
Cybb
|
cytochrome b-245, beta polypeptide |
| chr17_-_35081129 | 21.96 |
ENSMUST00000154526.8
|
Cfb
|
complement factor B |
| chr4_-_42773987 | 21.89 |
ENSMUST00000095114.5
|
Ccl21a
|
chemokine (C-C motif) ligand 21A (serine) |
| chr17_-_36353582 | 21.53 |
ENSMUST00000058801.15
ENSMUST00000080015.12 ENSMUST00000077960.7 |
H2-T22
|
histocompatibility 2, T region locus 22 |
| chr5_-_92496730 | 21.34 |
ENSMUST00000038816.13
ENSMUST00000118006.3 |
Cxcl10
|
chemokine (C-X-C motif) ligand 10 |
| chr8_-_71219299 | 21.26 |
ENSMUST00000222087.4
|
Ifi30
|
interferon gamma inducible protein 30 |
| chr7_-_102214636 | 21.24 |
ENSMUST00000106913.3
ENSMUST00000033264.12 |
Trim21
|
tripartite motif-containing 21 |
| chr2_-_51863203 | 21.03 |
ENSMUST00000028314.9
|
Nmi
|
N-myc (and STAT) interactor |
| chr4_-_46536088 | 20.94 |
ENSMUST00000102924.3
ENSMUST00000046897.13 |
Trim14
|
tripartite motif-containing 14 |
| chr17_+_34406762 | 20.85 |
ENSMUST00000041633.15
|
Tap1
|
transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) |
| chr11_-_48762170 | 20.77 |
ENSMUST00000049519.4
ENSMUST00000097271.4 |
Irgm1
|
immunity-related GTPase family M member 1 |
| chr1_+_175459559 | 20.68 |
ENSMUST00000040250.15
|
Kmo
|
kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) |
| chr12_+_111383864 | 20.47 |
ENSMUST00000220537.2
ENSMUST00000223050.2 ENSMUST00000072646.8 ENSMUST00000223431.2 ENSMUST00000221144.2 ENSMUST00000222437.2 |
Exoc3l4
|
exocyst complex component 3-like 4 |
| chr7_+_78563513 | 20.14 |
ENSMUST00000038142.15
|
Isg20
|
interferon-stimulated protein |
| chr11_+_101473062 | 20.03 |
ENSMUST00000039581.14
ENSMUST00000100403.9 ENSMUST00000107194.8 ENSMUST00000128614.2 |
Tmem106a
|
transmembrane protein 106A |
| chr9_+_5298669 | 19.87 |
ENSMUST00000238505.2
|
Casp1
|
caspase 1 |
| chr5_+_115034997 | 19.84 |
ENSMUST00000031542.13
ENSMUST00000146072.8 ENSMUST00000150361.2 |
Oasl2
|
2'-5' oligoadenylate synthetase-like 2 |
| chr12_-_26506422 | 19.49 |
ENSMUST00000020970.10
|
Rsad2
|
radical S-adenosyl methionine domain containing 2 |
| chr6_-_38331187 | 19.45 |
ENSMUST00000114900.8
ENSMUST00000143702.5 |
Zc3hav1
|
zinc finger CCCH type, antiviral 1 |
| chr15_+_77613239 | 19.39 |
ENSMUST00000230979.2
ENSMUST00000109775.4 |
Apol9b
|
apolipoprotein L 9b |
| chr5_-_105287405 | 19.13 |
ENSMUST00000100961.5
ENSMUST00000031235.13 ENSMUST00000197799.2 ENSMUST00000199629.2 ENSMUST00000196677.5 ENSMUST00000100962.8 |
Gbp9
Gbp8
Gbp4
|
guanylate-binding protein 9 guanylate-binding protein 8 guanylate binding protein 4 |
| chr1_+_130754413 | 18.86 |
ENSMUST00000027675.14
ENSMUST00000133792.8 |
Pigr
|
polymeric immunoglobulin receptor |
| chr5_-_105441554 | 18.50 |
ENSMUST00000050011.10
ENSMUST00000196520.2 |
Gm43302
Gbp6
|
predicted gene 43302 guanylate binding protein 6 |
| chr4_-_155012534 | 18.45 |
ENSMUST00000219534.3
|
Tnfrsf14
|
tumor necrosis factor receptor superfamily, member 14 (herpesvirus entry mediator) |
| chr2_-_51862941 | 18.34 |
ENSMUST00000145481.8
ENSMUST00000112705.9 |
Nmi
|
N-myc (and STAT) interactor |
| chr9_-_88613967 | 18.30 |
ENSMUST00000098486.4
|
Bcl2a1d
|
B cell leukemia/lymphoma 2 related protein A1d |
| chr4_-_155012643 | 18.17 |
ENSMUST00000123514.8
|
Tnfrsf14
|
tumor necrosis factor receptor superfamily, member 14 (herpesvirus entry mediator) |
| chr11_+_88890202 | 18.05 |
ENSMUST00000100627.9
ENSMUST00000107896.10 ENSMUST00000000284.7 |
Trim25
|
tripartite motif-containing 25 |
| chr1_-_173740467 | 17.17 |
ENSMUST00000009340.10
|
Ifi211
|
interferon activated gene 211 |
| chr19_+_29388312 | 16.97 |
ENSMUST00000112576.4
|
Pdcd1lg2
|
programmed cell death 1 ligand 2 |
| chr6_-_125208738 | 16.96 |
ENSMUST00000043422.8
|
Tapbpl
|
TAP binding protein-like |
| chr1_-_173594475 | 16.65 |
ENSMUST00000111214.4
|
Ifi204
|
interferon activated gene 204 |
| chr1_+_175459735 | 16.58 |
ENSMUST00000097458.4
|
Kmo
|
kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) |
| chr8_-_45786226 | 16.41 |
ENSMUST00000095328.6
|
Cyp4v3
|
cytochrome P450, family 4, subfamily v, polypeptide 3 |
| chr1_-_156501860 | 16.33 |
ENSMUST00000188964.7
ENSMUST00000190607.2 ENSMUST00000079625.11 |
Tor3a
|
torsin family 3, member A |
| chr3_+_27371206 | 16.10 |
ENSMUST00000174840.2
|
Tnfsf10
|
tumor necrosis factor (ligand) superfamily, member 10 |
| chr5_-_105258142 | 15.90 |
ENSMUST00000031238.13
|
Gbp9
|
guanylate-binding protein 9 |
| chr17_-_35081456 | 15.65 |
ENSMUST00000025229.11
ENSMUST00000176203.9 ENSMUST00000128767.8 |
Cfb
|
complement factor B |
| chr17_+_34573760 | 15.62 |
ENSMUST00000178562.2
ENSMUST00000025198.15 |
Btnl2
|
butyrophilin-like 2 |
| chr13_+_120761861 | 15.56 |
ENSMUST00000225029.2
|
BC147527
|
cDNA sequence BC147527 |
| chr14_+_14475188 | 15.43 |
ENSMUST00000026315.8
|
Dnase1l3
|
deoxyribonuclease 1-like 3 |
| chr9_+_88838953 | 15.39 |
ENSMUST00000098485.4
|
Bcl2a1a
|
B cell leukemia/lymphoma 2 related protein A1a |
| chr2_+_84629172 | 15.34 |
ENSMUST00000102642.9
ENSMUST00000150325.2 |
Ube2l6
|
ubiquitin-conjugating enzyme E2L 6 |
| chr5_-_120915693 | 15.26 |
ENSMUST00000044833.9
|
Oas3
|
2'-5' oligoadenylate synthetase 3 |
| chr7_-_104002534 | 15.02 |
ENSMUST00000130139.3
ENSMUST00000059037.15 |
Trim12c
|
tripartite motif-containing 12C |
| chr13_-_100454182 | 14.85 |
ENSMUST00000118574.8
|
Naip6
|
NLR family, apoptosis inhibitory protein 6 |
| chr11_+_48977495 | 14.67 |
ENSMUST00000152914.2
|
Ifi47
|
interferon gamma inducible protein 47 |
| chr18_+_60936910 | 14.63 |
ENSMUST00000097563.9
ENSMUST00000050487.16 ENSMUST00000167610.2 |
Cd74
|
CD74 antigen (invariant polypeptide of major histocompatibility complex, class II antigen-associated) |
| chr6_+_70703409 | 14.54 |
ENSMUST00000103410.3
|
Igkc
|
immunoglobulin kappa constant |
| chr7_+_78563964 | 14.54 |
ENSMUST00000120331.4
|
Isg20
|
interferon-stimulated protein |
| chr6_+_39358036 | 14.51 |
ENSMUST00000031986.5
|
Rab19
|
RAB19, member RAS oncogene family |
| chr18_+_60509101 | 14.44 |
ENSMUST00000032473.7
ENSMUST00000066912.13 |
Iigp1
|
interferon inducible GTPase 1 |
| chr5_-_134258435 | 14.27 |
ENSMUST00000016094.13
ENSMUST00000111275.8 ENSMUST00000144086.2 |
Ncf1
|
neutrophil cytosolic factor 1 |
| chr11_-_53750016 | 14.20 |
ENSMUST00000117316.8
ENSMUST00000120776.8 ENSMUST00000121435.2 |
Gm12216
|
predicted gene 12216 |
| chr19_+_6214416 | 13.98 |
ENSMUST00000045042.8
ENSMUST00000237511.2 |
Batf2
|
basic leucine zipper transcription factor, ATF-like 2 |
| chr5_-_120887864 | 13.77 |
ENSMUST00000053909.13
ENSMUST00000081491.13 |
Oas2
|
2'-5' oligoadenylate synthetase 2 |
| chr17_+_35658131 | 13.62 |
ENSMUST00000071951.14
ENSMUST00000116598.10 ENSMUST00000078205.14 ENSMUST00000076256.8 |
H2-Q7
|
histocompatibility 2, Q region locus 7 |
| chr16_-_10603389 | 13.61 |
ENSMUST00000229866.2
ENSMUST00000038099.6 |
Socs1
|
suppressor of cytokine signaling 1 |
| chr5_+_115061293 | 13.33 |
ENSMUST00000031540.11
ENSMUST00000112143.4 |
Oasl1
|
2'-5' oligoadenylate synthetase-like 1 |
| chr2_+_24275321 | 13.33 |
ENSMUST00000056641.15
ENSMUST00000142522.8 ENSMUST00000131930.2 |
Psd4
|
pleckstrin and Sec7 domain containing 4 |
| chr9_-_106353571 | 13.32 |
ENSMUST00000123555.8
ENSMUST00000125850.2 |
Parp3
|
poly (ADP-ribose) polymerase family, member 3 |
| chr10_-_34003933 | 13.31 |
ENSMUST00000062784.8
|
Calhm6
|
calcium homeostasis modulator family member 6 |
| chr19_+_34560922 | 13.27 |
ENSMUST00000102825.4
|
Ifit3
|
interferon-induced protein with tetratricopeptide repeats 3 |
| chr19_-_29344694 | 13.15 |
ENSMUST00000138051.2
|
Plgrkt
|
plasminogen receptor, C-terminal lysine transmembrane protein |
| chr11_-_100595019 | 13.11 |
ENSMUST00000017974.13
|
Dhx58
|
DEXH (Asp-Glu-X-His) box polypeptide 58 |
| chr8_-_106665060 | 12.94 |
ENSMUST00000034369.10
|
Psmb10
|
proteasome (prosome, macropain) subunit, beta type 10 |
| chr8_-_112064783 | 12.90 |
ENSMUST00000056157.14
ENSMUST00000120432.3 |
Mlkl
|
mixed lineage kinase domain-like |
| chr17_+_34138699 | 12.77 |
ENSMUST00000234320.2
|
Tapbp
|
TAP binding protein |
| chr7_-_104157006 | 12.75 |
ENSMUST00000033211.14
ENSMUST00000071069.13 |
Trim30d
|
tripartite motif-containing 30D |
| chr16_-_35691914 | 12.69 |
ENSMUST00000042665.9
|
Parp14
|
poly (ADP-ribose) polymerase family, member 14 |
| chr13_+_56757389 | 12.66 |
ENSMUST00000045173.10
|
Tgfbi
|
transforming growth factor, beta induced |
| chr4_-_144973423 | 12.61 |
ENSMUST00000030336.11
|
Tnfrsf1b
|
tumor necrosis factor receptor superfamily, member 1b |
| chr5_-_105387395 | 12.60 |
ENSMUST00000065588.7
|
Gbp10
|
guanylate-binding protein 10 |
| chr3_+_95434093 | 12.58 |
ENSMUST00000015667.9
ENSMUST00000116304.3 |
Ctss
|
cathepsin S |
| chr7_+_78564062 | 12.52 |
ENSMUST00000205981.2
|
Isg20
|
interferon-stimulated protein |
| chr1_-_184464810 | 12.49 |
ENSMUST00000048572.7
|
Hlx
|
H2.0-like homeobox |
| chr14_+_55815999 | 12.36 |
ENSMUST00000172738.2
ENSMUST00000089619.13 |
Psme1
|
proteasome (prosome, macropain) activator subunit 1 (PA28 alpha) |
| chr3_+_142236086 | 12.34 |
ENSMUST00000171263.8
ENSMUST00000045097.11 |
Gbp7
|
guanylate binding protein 7 |
| chr11_+_58090382 | 12.26 |
ENSMUST00000035266.11
ENSMUST00000094169.11 ENSMUST00000168280.2 ENSMUST00000058704.9 |
Igtp
Irgm2
|
interferon gamma induced GTPase immunity-related GTPase family M member 2 |
| chr2_+_84818538 | 12.17 |
ENSMUST00000028466.12
|
Prg3
|
proteoglycan 3 |
| chr6_-_38331482 | 12.09 |
ENSMUST00000031850.10
ENSMUST00000114898.3 |
Zc3hav1
|
zinc finger CCCH type, antiviral 1 |
| chr13_+_74787952 | 12.03 |
ENSMUST00000221822.2
ENSMUST00000221526.2 |
Erap1
|
endoplasmic reticulum aminopeptidase 1 |
| chr14_-_66071412 | 12.01 |
ENSMUST00000022613.10
|
Esco2
|
establishment of sister chromatid cohesion N-acetyltransferase 2 |
| chr17_+_36432553 | 11.96 |
ENSMUST00000173128.2
|
Gm19684
|
predicted gene, 19684 |
| chr3_-_7678785 | 11.93 |
ENSMUST00000194279.6
|
Il7
|
interleukin 7 |
| chr9_+_5345405 | 11.87 |
ENSMUST00000027009.11
|
Casp12
|
caspase 12 |
| chr1_+_52158599 | 11.87 |
ENSMUST00000186574.7
ENSMUST00000070968.14 ENSMUST00000191435.7 ENSMUST00000186857.7 ENSMUST00000188681.7 |
Stat1
|
signal transducer and activator of transcription 1 |
| chr7_-_140535899 | 11.83 |
ENSMUST00000081649.10
|
Ifitm2
|
interferon induced transmembrane protein 2 |
| chr17_-_34219225 | 11.79 |
ENSMUST00000238098.2
ENSMUST00000087189.7 ENSMUST00000173075.3 ENSMUST00000172912.8 ENSMUST00000236740.2 ENSMUST00000025181.18 |
H2-K1
|
histocompatibility 2, K1, K region |
| chr14_+_55815879 | 11.74 |
ENSMUST00000174563.8
|
Psme1
|
proteasome (prosome, macropain) activator subunit 1 (PA28 alpha) |
| chr1_+_85454323 | 11.52 |
ENSMUST00000239236.2
|
Gm7592
|
predicted gene 7592 |
| chr1_+_153627692 | 11.51 |
ENSMUST00000183241.8
|
Rnasel
|
ribonuclease L (2', 5'-oligoisoadenylate synthetase-dependent) |
| chr17_+_34138611 | 11.50 |
ENSMUST00000234247.2
|
Tapbp
|
TAP binding protein |
| chr3_-_7678796 | 11.48 |
ENSMUST00000192202.6
|
Il7
|
interleukin 7 |
| chr17_+_37504783 | 11.34 |
ENSMUST00000038844.7
|
Ubd
|
ubiquitin D |
| chr1_+_52158693 | 11.31 |
ENSMUST00000189347.7
|
Stat1
|
signal transducer and activator of transcription 1 |
| chr11_+_101339233 | 11.23 |
ENSMUST00000010502.13
|
Ifi35
|
interferon-induced protein 35 |
| chr3_+_142266113 | 11.20 |
ENSMUST00000106221.8
|
Gbp3
|
guanylate binding protein 3 |
| chr9_+_5345450 | 10.96 |
ENSMUST00000151332.2
|
Casp12
|
caspase 12 |
| chr6_+_40619913 | 10.95 |
ENSMUST00000238599.2
|
Mgam
|
maltase-glucoamylase |
| chr17_-_79190002 | 10.94 |
ENSMUST00000024884.5
|
Eif2ak2
|
eukaryotic translation initiation factor 2-alpha kinase 2 |
| chr14_+_55815817 | 10.91 |
ENSMUST00000174259.8
|
Psme1
|
proteasome (prosome, macropain) activator subunit 1 (PA28 alpha) |
| chr7_+_142014546 | 10.87 |
ENSMUST00000105968.8
ENSMUST00000018963.11 ENSMUST00000105967.8 |
Lsp1
|
lymphocyte specific 1 |
| chr4_+_134847949 | 10.87 |
ENSMUST00000056977.14
|
Runx3
|
runt related transcription factor 3 |
| chr19_-_34579356 | 10.85 |
ENSMUST00000168254.3
|
Ifit1bl1
|
interferon induced protein with tetratricpeptide repeats 1B like 1 |
| chr15_+_79776597 | 10.81 |
ENSMUST00000175714.8
ENSMUST00000109620.10 ENSMUST00000165537.8 ENSMUST00000175752.8 ENSMUST00000176325.8 |
Apobec3
|
apolipoprotein B mRNA editing enzyme, catalytic polypeptide 3 |
| chr15_-_77480311 | 10.75 |
ENSMUST00000089465.6
|
Apol10b
|
apolipoprotein L 10B |
| chr7_-_44785815 | 10.70 |
ENSMUST00000146760.7
|
Flt3l
|
FMS-like tyrosine kinase 3 ligand |
| chr3_+_142265904 | 10.44 |
ENSMUST00000128609.8
ENSMUST00000029935.14 |
Gbp3
|
guanylate binding protein 3 |
| chr2_-_103114105 | 10.40 |
ENSMUST00000111174.8
|
Ehf
|
ets homologous factor |
| chr5_-_137145030 | 10.37 |
ENSMUST00000054384.6
ENSMUST00000152207.2 |
Trim56
|
tripartite motif-containing 56 |
| chr1_-_172418058 | 10.36 |
ENSMUST00000065679.8
|
Slamf8
|
SLAM family member 8 |
| chr19_+_10819896 | 10.34 |
ENSMUST00000025646.3
|
Slc15a3
|
solute carrier family 15, member 3 |
| chr16_-_97264099 | 10.28 |
ENSMUST00000023655.13
|
Mx1
|
MX dynamin-like GTPase 1 |
| chr1_-_69725045 | 10.16 |
ENSMUST00000190771.7
|
Ikzf2
|
IKAROS family zinc finger 2 |
| chr12_-_79054050 | 10.04 |
ENSMUST00000056660.13
ENSMUST00000174721.8 |
Tmem229b
|
transmembrane protein 229B |
| chr19_-_3955230 | 10.02 |
ENSMUST00000145791.8
|
Tcirg1
|
T cell, immune regulator 1, ATPase, H+ transporting, lysosomal V0 protein A3 |
| chr3_+_142265787 | 9.91 |
ENSMUST00000199325.5
ENSMUST00000106222.9 |
Gbp3
|
guanylate binding protein 3 |
| chr7_-_44785709 | 9.90 |
ENSMUST00000211429.2
|
Flt3l
|
FMS-like tyrosine kinase 3 ligand |
| chr17_+_35481702 | 9.88 |
ENSMUST00000172785.8
|
H2-D1
|
histocompatibility 2, D region locus 1 |
| chr9_-_106353303 | 9.87 |
ENSMUST00000156426.8
|
Parp3
|
poly (ADP-ribose) polymerase family, member 3 |
| chr7_-_140535828 | 9.82 |
ENSMUST00000211129.2
|
Gm45717
|
predicted gene 45717 |
| chr7_-_44785480 | 9.74 |
ENSMUST00000211246.2
ENSMUST00000210197.2 |
Flt3l
|
FMS-like tyrosine kinase 3 ligand |
| chr14_+_55815580 | 9.72 |
ENSMUST00000174484.8
|
Psme1
|
proteasome (prosome, macropain) activator subunit 1 (PA28 alpha) |
| chr7_-_130148984 | 9.70 |
ENSMUST00000160289.9
|
Nsmce4a
|
NSE4 homolog A, SMC5-SMC6 complex component |
| chr2_+_118428690 | 9.66 |
ENSMUST00000038341.8
|
Bub1b
|
BUB1B, mitotic checkpoint serine/threonine kinase |
| chr19_+_34585322 | 9.64 |
ENSMUST00000076249.6
|
Ifit3b
|
interferon-induced protein with tetratricopeptide repeats 3B |
| chr17_-_34218301 | 9.58 |
ENSMUST00000235463.2
|
H2-K1
|
histocompatibility 2, K1, K region |
| chr16_+_97337275 | 9.49 |
ENSMUST00000024112.8
|
Mx2
|
MX dynamin-like GTPase 2 |
| chr11_+_83066009 | 9.46 |
ENSMUST00000000208.13
ENSMUST00000167596.2 |
Slfn4
|
schlafen 4 |
| chr15_-_77295234 | 9.41 |
ENSMUST00000089452.6
ENSMUST00000081776.11 |
Apol9a
|
apolipoprotein L 9a |
| chr11_-_48870105 | 9.34 |
ENSMUST00000141200.2
ENSMUST00000097494.9 ENSMUST00000093153.2 |
9930111J21Rik1
|
RIKEN cDNA 9930111J21 gene 1 |
| chr19_-_11243530 | 9.33 |
ENSMUST00000169159.3
|
Ms4a1
|
membrane-spanning 4-domains, subfamily A, member 1 |
| chr6_-_69626340 | 9.31 |
ENSMUST00000198328.2
|
Igkv4-53
|
immunoglobulin kappa variable 4-53 |
| chr1_-_144651157 | 9.22 |
ENSMUST00000027603.4
|
Rgs18
|
regulator of G-protein signaling 18 |
| chr1_-_69724939 | 9.18 |
ENSMUST00000027146.9
|
Ikzf2
|
IKAROS family zinc finger 2 |
| chr1_+_58750647 | 9.14 |
ENSMUST00000097722.9
ENSMUST00000114313.8 |
Cflar
|
CASP8 and FADD-like apoptosis regulator |
| chr6_+_34722926 | 9.10 |
ENSMUST00000126181.8
|
Cald1
|
caldesmon 1 |
| chr1_+_52158721 | 9.06 |
ENSMUST00000186057.7
|
Stat1
|
signal transducer and activator of transcription 1 |
| chr8_+_95161006 | 8.95 |
ENSMUST00000211816.2
|
Nlrc5
|
NLR family, CARD domain containing 5 |
| chr10_-_88192852 | 8.87 |
ENSMUST00000020249.2
|
Dram1
|
DNA-damage regulated autophagy modulator 1 |
| chr2_+_164897547 | 8.85 |
ENSMUST00000017799.12
ENSMUST00000073707.9 |
Cd40
|
CD40 antigen |
| chr12_-_103409912 | 8.78 |
ENSMUST00000055071.9
|
Ifi27l2a
|
interferon, alpha-inducible protein 27 like 2A |
| chr6_-_54949587 | 8.74 |
ENSMUST00000060655.15
|
Nod1
|
nucleotide-binding oligomerization domain containing 1 |
| chr4_+_52439237 | 8.73 |
ENSMUST00000102915.10
ENSMUST00000117280.8 ENSMUST00000142227.3 |
Smc2
|
structural maintenance of chromosomes 2 |
| chr3_+_142300601 | 8.68 |
ENSMUST00000029936.5
|
Gbp2b
|
guanylate binding protein 2b |
| chr17_+_35598583 | 8.67 |
ENSMUST00000081435.5
|
H2-Q4
|
histocompatibility 2, Q region locus 4 |
| chr14_+_55842002 | 8.60 |
ENSMUST00000138037.2
|
Irf9
|
interferon regulatory factor 9 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 19.6 | 39.1 | GO:0002481 | antigen processing and presentation of exogenous peptide antigen via MHC class Ib(GO:0002477) antigen processing and presentation of exogenous protein antigen via MHC class Ib, TAP-dependent(GO:0002481) |
| 16.7 | 50.1 | GO:0046967 | cytosol to ER transport(GO:0046967) |
| 13.1 | 39.4 | GO:1902524 | positive regulation of protein K48-linked ubiquitination(GO:1902524) |
| 12.0 | 12.0 | GO:0002483 | antigen processing and presentation of endogenous peptide antigen(GO:0002483) |
| 11.5 | 91.7 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 11.1 | 44.2 | GO:0034124 | regulation of MyD88-dependent toll-like receptor signaling pathway(GO:0034124) |
| 10.7 | 32.2 | GO:0034240 | negative regulation of macrophage fusion(GO:0034240) |
| 10.6 | 53.2 | GO:0043321 | regulation of natural killer cell degranulation(GO:0043321) |
| 10.1 | 533.1 | GO:0035458 | cellular response to interferon-beta(GO:0035458) |
| 9.6 | 28.7 | GO:2000547 | regulation of dendritic cell dendrite assembly(GO:2000547) |
| 9.5 | 56.9 | GO:0035549 | interferon-beta secretion(GO:0035546) regulation of interferon-beta secretion(GO:0035547) positive regulation of interferon-beta secretion(GO:0035549) |
| 9.4 | 37.4 | GO:0070212 | protein poly-ADP-ribosylation(GO:0070212) |
| 8.5 | 51.1 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 7.8 | 31.2 | GO:0002484 | antigen processing and presentation of endogenous peptide antigen via MHC class I via ER pathway(GO:0002484) antigen processing and presentation of endogenous peptide antigen via MHC class I via ER pathway, TAP-dependent(GO:0002485) |
| 7.6 | 15.3 | GO:0060700 | regulation of ribonuclease activity(GO:0060700) |
| 7.6 | 30.5 | GO:1901740 | negative regulation of myoblast fusion(GO:1901740) |
| 7.4 | 44.2 | GO:0019805 | quinolinate biosynthetic process(GO:0019805) |
| 6.8 | 6.8 | GO:0002428 | antigen processing and presentation of peptide antigen via MHC class Ib(GO:0002428) antigen processing and presentation of endogenous peptide antigen via MHC class Ib(GO:0002476) |
| 6.5 | 19.5 | GO:0034165 | positive regulation of toll-like receptor 9 signaling pathway(GO:0034165) |
| 6.3 | 18.9 | GO:0002414 | immune response in mucosal-associated lymphoid tissue(GO:0002386) immunoglobulin transcytosis in epithelial cells(GO:0002414) |
| 6.1 | 24.3 | GO:0050822 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-dependent(GO:0002479) peptide stabilization(GO:0050822) peptide antigen stabilization(GO:0050823) |
| 5.9 | 70.7 | GO:0035456 | response to interferon-beta(GO:0035456) |
| 5.9 | 17.6 | GO:0002590 | regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002589) negative regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002590) |
| 5.9 | 29.3 | GO:0090290 | positive regulation of osteoclast proliferation(GO:0090290) |
| 5.8 | 87.0 | GO:0060330 | regulation of response to interferon-gamma(GO:0060330) |
| 5.7 | 22.7 | GO:0051673 | membrane disruption in other organism(GO:0051673) |
| 5.4 | 27.0 | GO:0072733 | response to staurosporine(GO:0072733) cellular response to staurosporine(GO:0072734) |
| 5.1 | 35.7 | GO:1903265 | positive regulation of tumor necrosis factor-mediated signaling pathway(GO:1903265) |
| 5.1 | 50.8 | GO:0032074 | negative regulation of nuclease activity(GO:0032074) |
| 4.9 | 44.5 | GO:0060340 | positive regulation of type I interferon-mediated signaling pathway(GO:0060340) |
| 4.9 | 29.5 | GO:1990166 | protein localization to site of double-strand break(GO:1990166) |
| 4.8 | 14.3 | GO:0070947 | neutrophil mediated killing of fungus(GO:0070947) |
| 4.7 | 23.4 | GO:0002606 | positive regulation of dendritic cell antigen processing and presentation(GO:0002606) |
| 4.6 | 13.9 | GO:0045083 | negative regulation of interleukin-12 biosynthetic process(GO:0045083) response to diterpene(GO:1904629) cellular response to diterpene(GO:1904630) response to glucoside(GO:1904631) cellular response to glucoside(GO:1904632) |
| 4.3 | 21.3 | GO:0042590 | antigen processing and presentation of exogenous peptide antigen via MHC class I(GO:0042590) |
| 4.2 | 12.6 | GO:0034769 | basement membrane disassembly(GO:0034769) |
| 4.1 | 61.2 | GO:0046642 | negative regulation of alpha-beta T cell proliferation(GO:0046642) |
| 3.9 | 27.5 | GO:2001181 | positive regulation of interleukin-10 secretion(GO:2001181) |
| 3.9 | 15.4 | GO:0042270 | protection from natural killer cell mediated cytotoxicity(GO:0042270) |
| 3.8 | 11.3 | GO:0030887 | positive regulation of myeloid dendritic cell activation(GO:0030887) |
| 3.7 | 7.5 | GO:0002649 | regulation of tolerance induction to self antigen(GO:0002649) |
| 3.7 | 18.7 | GO:0070543 | response to linoleic acid(GO:0070543) |
| 3.7 | 18.7 | GO:0010760 | negative regulation of macrophage chemotaxis(GO:0010760) |
| 3.3 | 10.0 | GO:0045404 | positive regulation of interleukin-4 biosynthetic process(GO:0045404) |
| 3.3 | 9.9 | GO:2001200 | positive regulation of dendritic cell differentiation(GO:2001200) |
| 3.1 | 15.4 | GO:0033634 | positive regulation of cell-cell adhesion mediated by integrin(GO:0033634) |
| 3.0 | 23.7 | GO:0097411 | hypoxia-inducible factor-1alpha signaling pathway(GO:0097411) |
| 3.0 | 47.3 | GO:1900246 | positive regulation of RIG-I signaling pathway(GO:1900246) |
| 2.9 | 14.7 | GO:0034421 | post-translational protein acetylation(GO:0034421) |
| 2.9 | 14.6 | GO:0071350 | interleukin-15-mediated signaling pathway(GO:0035723) cellular response to interleukin-15(GO:0071350) |
| 2.8 | 31.1 | GO:2000051 | negative regulation of non-canonical Wnt signaling pathway(GO:2000051) |
| 2.8 | 97.6 | GO:0042832 | defense response to protozoan(GO:0042832) |
| 2.7 | 30.2 | GO:0045341 | MHC class I biosynthetic process(GO:0045341) regulation of MHC class I biosynthetic process(GO:0045343) positive regulation of MHC class I biosynthetic process(GO:0045345) |
| 2.7 | 16.4 | GO:0060369 | positive regulation of natural killer cell cytokine production(GO:0002729) positive regulation of Fc receptor mediated stimulatory signaling pathway(GO:0060369) |
| 2.7 | 26.5 | GO:0046598 | positive regulation of viral entry into host cell(GO:0046598) |
| 2.6 | 82.5 | GO:0002474 | antigen processing and presentation of peptide antigen via MHC class I(GO:0002474) |
| 2.5 | 12.5 | GO:0045629 | negative regulation of T-helper 2 cell differentiation(GO:0045629) |
| 2.4 | 7.1 | GO:0036275 | response to 5-fluorouracil(GO:0036275) |
| 2.4 | 37.6 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 2.3 | 7.0 | GO:0031104 | dendrite regeneration(GO:0031104) |
| 2.1 | 6.3 | GO:0060809 | CD8-positive, alpha-beta T cell differentiation involved in immune response(GO:0002302) mesodermal to mesenchymal transition involved in gastrulation(GO:0060809) |
| 2.1 | 6.3 | GO:0002632 | regulation of granuloma formation(GO:0002631) negative regulation of granuloma formation(GO:0002632) regulation of toll-like receptor 5 signaling pathway(GO:0034147) negative regulation of toll-like receptor 5 signaling pathway(GO:0034148) negative regulation of nucleotide-binding oligomerization domain containing 1 signaling pathway(GO:0070429) tolerance induction to lipopolysaccharide(GO:0072573) |
| 2.1 | 22.9 | GO:0002903 | negative regulation of B cell apoptotic process(GO:0002903) |
| 2.0 | 48.9 | GO:0019884 | antigen processing and presentation of exogenous antigen(GO:0019884) |
| 1.9 | 7.7 | GO:0035279 | mRNA cleavage involved in gene silencing by miRNA(GO:0035279) mRNA cleavage involved in gene silencing(GO:0098795) |
| 1.9 | 22.8 | GO:0016540 | protein autoprocessing(GO:0016540) |
| 1.9 | 13.1 | GO:0010756 | positive regulation of plasminogen activation(GO:0010756) |
| 1.9 | 9.4 | GO:0097309 | cap1 mRNA methylation(GO:0097309) |
| 1.9 | 50.0 | GO:0035634 | response to stilbenoid(GO:0035634) |
| 1.8 | 35.0 | GO:0010818 | T cell chemotaxis(GO:0010818) |
| 1.8 | 5.5 | GO:0060220 | camera-type eye photoreceptor cell fate commitment(GO:0060220) negative regulation of neural retina development(GO:0061076) negative regulation of retina development in camera-type eye(GO:1902867) negative regulation of amacrine cell differentiation(GO:1902870) |
| 1.8 | 9.2 | GO:0001692 | histamine metabolic process(GO:0001692) |
| 1.8 | 5.5 | GO:0045590 | negative regulation of regulatory T cell differentiation(GO:0045590) |
| 1.8 | 5.3 | GO:1902202 | regulation of hepatocyte growth factor receptor signaling pathway(GO:1902202) |
| 1.7 | 13.8 | GO:0018377 | protein myristoylation(GO:0018377) |
| 1.6 | 6.5 | GO:0006272 | leading strand elongation(GO:0006272) mitotic telomere maintenance via semi-conservative replication(GO:1902990) |
| 1.6 | 8.1 | GO:0051754 | meiotic sister chromatid cohesion, centromeric(GO:0051754) |
| 1.6 | 4.8 | GO:0042450 | arginine biosynthetic process via ornithine(GO:0042450) |
| 1.5 | 4.6 | GO:0071288 | cellular response to mercury ion(GO:0071288) |
| 1.5 | 14.8 | GO:0070269 | pyroptosis(GO:0070269) |
| 1.5 | 4.4 | GO:2000769 | positive regulation of NMDA glutamate receptor activity(GO:1904783) regulation of establishment or maintenance of cell polarity regulating cell shape(GO:2000769) positive regulation of establishment or maintenance of cell polarity regulating cell shape(GO:2000771) |
| 1.5 | 17.5 | GO:1900225 | regulation of NLRP3 inflammasome complex assembly(GO:1900225) |
| 1.3 | 5.3 | GO:0034499 | late endosome to Golgi transport(GO:0034499) |
| 1.3 | 4.0 | GO:0032701 | negative regulation of granulocyte macrophage colony-stimulating factor production(GO:0032685) negative regulation of interleukin-18 production(GO:0032701) |
| 1.3 | 23.7 | GO:1902187 | negative regulation of viral release from host cell(GO:1902187) |
| 1.3 | 9.2 | GO:1900264 | regulation of DNA-directed DNA polymerase activity(GO:1900262) positive regulation of DNA-directed DNA polymerase activity(GO:1900264) |
| 1.3 | 5.2 | GO:0051329 | interphase(GO:0051325) mitotic interphase(GO:0051329) |
| 1.3 | 3.9 | GO:1903377 | negative regulation of oxidative stress-induced neuron intrinsic apoptotic signaling pathway(GO:1903377) |
| 1.3 | 10.3 | GO:0003374 | dynamin polymerization involved in membrane fission(GO:0003373) dynamin polymerization involved in mitochondrial fission(GO:0003374) |
| 1.3 | 17.6 | GO:0045351 | type I interferon biosynthetic process(GO:0045351) |
| 1.2 | 1.2 | GO:0038018 | Wnt receptor catabolic process(GO:0038018) |
| 1.2 | 20.9 | GO:0032897 | negative regulation of viral transcription(GO:0032897) |
| 1.2 | 30.6 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 1.2 | 2.4 | GO:0034146 | toll-like receptor 5 signaling pathway(GO:0034146) |
| 1.2 | 12.1 | GO:0032876 | negative regulation of DNA endoreduplication(GO:0032876) |
| 1.2 | 20.5 | GO:0051601 | exocyst localization(GO:0051601) |
| 1.2 | 14.2 | GO:0007039 | protein catabolic process in the vacuole(GO:0007039) |
| 1.2 | 15.0 | GO:0036462 | TRAIL-activated apoptotic signaling pathway(GO:0036462) |
| 1.2 | 5.8 | GO:0000454 | snoRNA guided rRNA pseudouridine synthesis(GO:0000454) telomerase RNA stabilization(GO:0090669) |
| 1.1 | 4.6 | GO:0090096 | lactic acid secretion(GO:0046722) regulation of metanephric cap mesenchymal cell proliferation(GO:0090095) positive regulation of metanephric cap mesenchymal cell proliferation(GO:0090096) |
| 1.1 | 4.5 | GO:0010751 | negative regulation of nitric oxide mediated signal transduction(GO:0010751) |
| 1.1 | 18.0 | GO:0032825 | positive regulation of natural killer cell differentiation(GO:0032825) |
| 1.1 | 15.6 | GO:1900042 | positive regulation of interleukin-2 secretion(GO:1900042) |
| 1.1 | 7.7 | GO:0010940 | positive regulation of necrotic cell death(GO:0010940) |
| 1.1 | 4.4 | GO:0002838 | negative regulation of response to tumor cell(GO:0002835) negative regulation of immune response to tumor cell(GO:0002838) |
| 1.1 | 6.3 | GO:0019364 | pyridine nucleotide catabolic process(GO:0019364) |
| 1.0 | 3.1 | GO:1903976 | negative regulation of glial cell migration(GO:1903976) |
| 1.0 | 40.4 | GO:0090200 | positive regulation of release of cytochrome c from mitochondria(GO:0090200) |
| 1.0 | 5.1 | GO:0001783 | B cell apoptotic process(GO:0001783) |
| 1.0 | 3.1 | GO:1903632 | positive regulation of aminoacyl-tRNA ligase activity(GO:1903632) |
| 1.0 | 8.1 | GO:0035726 | common myeloid progenitor cell proliferation(GO:0035726) |
| 1.0 | 2.9 | GO:0000320 | re-entry into mitotic cell cycle(GO:0000320) |
| 1.0 | 2.9 | GO:0045226 | extracellular polysaccharide biosynthetic process(GO:0045226) extracellular polysaccharide metabolic process(GO:0046379) regulation of monocyte aggregation(GO:1900623) positive regulation of monocyte aggregation(GO:1900625) |
| 1.0 | 35.3 | GO:0019882 | antigen processing and presentation(GO:0019882) |
| 0.9 | 3.7 | GO:0006227 | dUDP biosynthetic process(GO:0006227) pyrimidine nucleoside diphosphate metabolic process(GO:0009138) pyrimidine nucleoside diphosphate biosynthetic process(GO:0009139) pyrimidine deoxyribonucleoside diphosphate metabolic process(GO:0009196) pyrimidine deoxyribonucleoside diphosphate biosynthetic process(GO:0009197) dUDP metabolic process(GO:0046077) |
| 0.9 | 3.7 | GO:1903237 | negative regulation of leukocyte tethering or rolling(GO:1903237) |
| 0.9 | 4.5 | GO:1904975 | response to bleomycin(GO:1904975) cellular response to bleomycin(GO:1904976) |
| 0.9 | 0.9 | GO:1990009 | retinal cell apoptotic process(GO:1990009) |
| 0.9 | 16.0 | GO:0006309 | apoptotic DNA fragmentation(GO:0006309) |
| 0.9 | 8.7 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 0.9 | 5.2 | GO:0006116 | NADH oxidation(GO:0006116) |
| 0.9 | 35.0 | GO:0045071 | negative regulation of viral genome replication(GO:0045071) |
| 0.8 | 3.3 | GO:2000383 | regulation of ectoderm development(GO:2000383) negative regulation of ectoderm development(GO:2000384) |
| 0.8 | 2.5 | GO:0014739 | positive regulation of muscle hyperplasia(GO:0014739) |
| 0.8 | 2.5 | GO:1990918 | meiotic DNA double-strand break processing involved in reciprocal meiotic recombination(GO:0010705) double-strand break repair involved in meiotic recombination(GO:1990918) |
| 0.8 | 24.0 | GO:0002360 | T cell lineage commitment(GO:0002360) |
| 0.8 | 6.4 | GO:0034242 | negative regulation of syncytium formation by plasma membrane fusion(GO:0034242) |
| 0.8 | 3.2 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.8 | 3.2 | GO:0006447 | regulation of translational initiation by iron(GO:0006447) positive regulation of translational initiation by iron(GO:0045994) |
| 0.8 | 3.2 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 0.8 | 2.4 | GO:0030221 | basophil differentiation(GO:0030221) |
| 0.8 | 3.1 | GO:0072086 | specification of loop of Henle identity(GO:0072086) |
| 0.8 | 14.0 | GO:0043011 | myeloid dendritic cell differentiation(GO:0043011) |
| 0.8 | 2.3 | GO:0032348 | negative regulation of aldosterone metabolic process(GO:0032345) negative regulation of aldosterone biosynthetic process(GO:0032348) negative regulation of cortisol biosynthetic process(GO:2000065) |
| 0.8 | 3.1 | GO:0072355 | histone H3-T3 phosphorylation(GO:0072355) |
| 0.8 | 3.1 | GO:0043328 | protein targeting to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043328) |
| 0.8 | 0.8 | GO:0061763 | multivesicular body-lysosome fusion(GO:0061763) |
| 0.8 | 6.1 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 0.7 | 6.0 | GO:2000659 | regulation of interleukin-1-mediated signaling pathway(GO:2000659) |
| 0.7 | 5.2 | GO:0002827 | positive regulation of T-helper 1 type immune response(GO:0002827) |
| 0.7 | 2.2 | GO:0018900 | dichloromethane metabolic process(GO:0018900) |
| 0.7 | 2.2 | GO:0021558 | trochlear nerve development(GO:0021558) auditory receptor cell fate determination(GO:0042668) negative regulation of forebrain neuron differentiation(GO:2000978) |
| 0.7 | 0.7 | GO:0002884 | negative regulation of hypersensitivity(GO:0002884) |
| 0.7 | 6.5 | GO:0033184 | positive regulation of histone ubiquitination(GO:0033184) |
| 0.7 | 2.2 | GO:0090367 | negative regulation of mRNA modification(GO:0090367) |
| 0.7 | 5.0 | GO:2000623 | regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000622) negative regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000623) |
| 0.7 | 2.1 | GO:0015688 | iron chelate transport(GO:0015688) siderophore transport(GO:0015891) |
| 0.7 | 4.9 | GO:1900170 | negative regulation of glucocorticoid mediated signaling pathway(GO:1900170) |
| 0.7 | 18.7 | GO:0051205 | protein insertion into membrane(GO:0051205) |
| 0.7 | 3.4 | GO:0018101 | protein citrullination(GO:0018101) histone citrullination(GO:0036414) |
| 0.7 | 3.4 | GO:0070245 | positive regulation of thymocyte apoptotic process(GO:0070245) |
| 0.7 | 3.4 | GO:0046947 | hydroxylysine metabolic process(GO:0046946) hydroxylysine biosynthetic process(GO:0046947) |
| 0.7 | 2.7 | GO:0006982 | response to lipid hydroperoxide(GO:0006982) cellular response to lipid hydroperoxide(GO:0071449) |
| 0.7 | 4.1 | GO:0071918 | urea transmembrane transport(GO:0071918) |
| 0.7 | 5.3 | GO:0001957 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 0.7 | 2.0 | GO:0098583 | mastication(GO:0071626) learned vocalization behavior(GO:0098583) |
| 0.7 | 5.9 | GO:0031293 | membrane protein intracellular domain proteolysis(GO:0031293) |
| 0.6 | 7.1 | GO:0090503 | RNA phosphodiester bond hydrolysis, exonucleolytic(GO:0090503) |
| 0.6 | 3.8 | GO:0071357 | type I interferon signaling pathway(GO:0060337) cellular response to type I interferon(GO:0071357) |
| 0.6 | 1.9 | GO:0002014 | vasoconstriction of artery involved in ischemic response to lowering of systemic arterial blood pressure(GO:0002014) |
| 0.6 | 4.4 | GO:0044351 | macropinocytosis(GO:0044351) |
| 0.6 | 4.9 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.6 | 17.3 | GO:0042789 | mRNA transcription from RNA polymerase II promoter(GO:0042789) |
| 0.6 | 2.5 | GO:0038094 | Fc-gamma receptor signaling pathway(GO:0038094) |
| 0.6 | 2.4 | GO:0061386 | closure of optic fissure(GO:0061386) |
| 0.6 | 10.3 | GO:0006857 | oligopeptide transport(GO:0006857) |
| 0.6 | 4.2 | GO:0031584 | activation of phospholipase D activity(GO:0031584) |
| 0.6 | 6.6 | GO:0060158 | phospholipase C-activating dopamine receptor signaling pathway(GO:0060158) |
| 0.6 | 2.4 | GO:0016332 | establishment or maintenance of polarity of embryonic epithelium(GO:0016332) |
| 0.6 | 3.6 | GO:0000821 | regulation of arginine metabolic process(GO:0000821) |
| 0.6 | 4.2 | GO:0046826 | negative regulation of protein export from nucleus(GO:0046826) |
| 0.6 | 1.8 | GO:0019626 | short-chain fatty acid catabolic process(GO:0019626) |
| 0.6 | 3.5 | GO:0070164 | negative regulation of adiponectin secretion(GO:0070164) |
| 0.6 | 1.8 | GO:0035441 | cell migration involved in vasculogenesis(GO:0035441) |
| 0.6 | 1.7 | GO:1903904 | negative regulation of establishment of T cell polarity(GO:1903904) |
| 0.6 | 1.7 | GO:0046061 | DNA protection(GO:0042262) dATP catabolic process(GO:0046061) |
| 0.6 | 3.5 | GO:0031642 | negative regulation of myelination(GO:0031642) |
| 0.6 | 2.3 | GO:0038156 | interleukin-3-mediated signaling pathway(GO:0038156) |
| 0.6 | 7.9 | GO:0051490 | negative regulation of filopodium assembly(GO:0051490) |
| 0.6 | 5.6 | GO:0031666 | positive regulation of lipopolysaccharide-mediated signaling pathway(GO:0031666) |
| 0.6 | 1.1 | GO:1904742 | regulation of telomeric DNA binding(GO:1904742) |
| 0.6 | 2.2 | GO:0009816 | defense response, incompatible interaction(GO:0009814) defense response to bacterium, incompatible interaction(GO:0009816) regulation of defense response to bacterium, incompatible interaction(GO:1902477) |
| 0.5 | 8.2 | GO:0051315 | attachment of mitotic spindle microtubules to kinetochore(GO:0051315) |
| 0.5 | 12.6 | GO:0051044 | positive regulation of membrane protein ectodomain proteolysis(GO:0051044) |
| 0.5 | 2.7 | GO:0060355 | positive regulation of cell adhesion molecule production(GO:0060355) |
| 0.5 | 3.2 | GO:0061511 | centriole elongation(GO:0061511) |
| 0.5 | 7.0 | GO:0006968 | cellular defense response(GO:0006968) |
| 0.5 | 2.1 | GO:1902896 | terminal web assembly(GO:1902896) |
| 0.5 | 2.6 | GO:0036023 | limb joint morphogenesis(GO:0036022) embryonic skeletal limb joint morphogenesis(GO:0036023) |
| 0.5 | 3.0 | GO:0003199 | endocardial cushion to mesenchymal transition involved in heart valve formation(GO:0003199) |
| 0.5 | 2.5 | GO:0021622 | optic cup structural organization(GO:0003409) oculomotor nerve development(GO:0021557) oculomotor nerve morphogenesis(GO:0021622) oculomotor nerve formation(GO:0021623) |
| 0.5 | 3.5 | GO:0016266 | O-glycan processing(GO:0016266) |
| 0.5 | 2.0 | GO:0015766 | disaccharide transport(GO:0015766) sucrose transport(GO:0015770) oligosaccharide transport(GO:0015772) |
| 0.5 | 6.8 | GO:0032693 | negative regulation of interleukin-10 production(GO:0032693) |
| 0.5 | 1.5 | GO:0045013 | carbon catabolite repression of transcription(GO:0045013) negative regulation of transcription by glucose(GO:0045014) |
| 0.5 | 2.4 | GO:0006041 | glucosamine metabolic process(GO:0006041) |
| 0.5 | 3.8 | GO:0032494 | response to peptidoglycan(GO:0032494) |
| 0.5 | 1.4 | GO:0045348 | positive regulation of MHC class II biosynthetic process(GO:0045348) |
| 0.5 | 9.8 | GO:0070266 | necroptotic process(GO:0070266) |
| 0.5 | 1.4 | GO:0006553 | lysine metabolic process(GO:0006553) |
| 0.5 | 0.5 | GO:2000642 | negative regulation of early endosome to late endosome transport(GO:2000642) |
| 0.5 | 2.8 | GO:0007296 | vitellogenesis(GO:0007296) |
| 0.5 | 13.1 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.4 | 10.6 | GO:0050869 | negative regulation of B cell activation(GO:0050869) |
| 0.4 | 5.3 | GO:1990001 | inhibition of cysteine-type endopeptidase activity involved in apoptotic process(GO:1990001) |
| 0.4 | 11.4 | GO:0019731 | antibacterial humoral response(GO:0019731) |
| 0.4 | 6.1 | GO:0048525 | negative regulation of viral process(GO:0048525) |
| 0.4 | 6.1 | GO:0019985 | translesion synthesis(GO:0019985) |
| 0.4 | 3.0 | GO:0070494 | regulation of thrombin receptor signaling pathway(GO:0070494) negative regulation of thrombin receptor signaling pathway(GO:0070495) |
| 0.4 | 3.4 | GO:0032488 | Cdc42 protein signal transduction(GO:0032488) |
| 0.4 | 68.2 | GO:0051607 | defense response to virus(GO:0051607) |
| 0.4 | 2.6 | GO:0006438 | valyl-tRNA aminoacylation(GO:0006438) |
| 0.4 | 2.1 | GO:0017185 | peptidyl-lysine hydroxylation(GO:0017185) |
| 0.4 | 3.8 | GO:0043031 | negative regulation of macrophage activation(GO:0043031) |
| 0.4 | 6.8 | GO:0045779 | negative regulation of bone resorption(GO:0045779) |
| 0.4 | 60.4 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.4 | 2.9 | GO:0045654 | positive regulation of megakaryocyte differentiation(GO:0045654) |
| 0.4 | 3.7 | GO:0009143 | nucleoside triphosphate catabolic process(GO:0009143) |
| 0.4 | 4.5 | GO:0032463 | negative regulation of protein homooligomerization(GO:0032463) |
| 0.4 | 1.2 | GO:1902626 | assembly of large subunit precursor of preribosome(GO:1902626) |
| 0.4 | 4.8 | GO:0019367 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.4 | 0.4 | GO:0002314 | germinal center B cell differentiation(GO:0002314) |
| 0.4 | 1.2 | GO:0036090 | cleavage furrow ingression(GO:0036090) |
| 0.4 | 3.5 | GO:1901223 | negative regulation of NIK/NF-kappaB signaling(GO:1901223) |
| 0.4 | 3.5 | GO:0006307 | DNA dealkylation involved in DNA repair(GO:0006307) |
| 0.4 | 14.1 | GO:0010390 | histone monoubiquitination(GO:0010390) |
| 0.4 | 8.1 | GO:0031953 | negative regulation of protein autophosphorylation(GO:0031953) |
| 0.4 | 4.0 | GO:0001672 | regulation of chromatin assembly or disassembly(GO:0001672) |
| 0.4 | 1.1 | GO:0046501 | protoporphyrinogen IX metabolic process(GO:0046501) |
| 0.4 | 11.5 | GO:0035855 | megakaryocyte development(GO:0035855) |
| 0.4 | 1.1 | GO:0072488 | ammonium transmembrane transport(GO:0072488) |
| 0.4 | 1.1 | GO:0006583 | melanin biosynthetic process from tyrosine(GO:0006583) |
| 0.4 | 1.4 | GO:2001245 | regulation of phosphatidylcholine biosynthetic process(GO:2001245) |
| 0.4 | 4.6 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.4 | 3.2 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 0.3 | 1.7 | GO:0010990 | regulation of SMAD protein complex assembly(GO:0010990) negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.3 | 2.8 | GO:0090344 | negative regulation of cell aging(GO:0090344) |
| 0.3 | 5.5 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.3 | 1.4 | GO:0051771 | negative regulation of nitric-oxide synthase biosynthetic process(GO:0051771) |
| 0.3 | 1.3 | GO:2000348 | regulation of CD40 signaling pathway(GO:2000348) |
| 0.3 | 7.6 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.3 | 2.3 | GO:0008655 | pyrimidine-containing compound salvage(GO:0008655) pyrimidine nucleoside salvage(GO:0043097) |
| 0.3 | 1.3 | GO:1902916 | positive regulation of protein polyubiquitination(GO:1902916) |
| 0.3 | 2.8 | GO:0015760 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.3 | 2.2 | GO:0038165 | oncostatin-M-mediated signaling pathway(GO:0038165) |
| 0.3 | 2.5 | GO:0042427 | serotonin biosynthetic process(GO:0042427) primary amino compound biosynthetic process(GO:1901162) |
| 0.3 | 6.8 | GO:0051923 | sulfation(GO:0051923) |
| 0.3 | 1.5 | GO:0044375 | regulation of peroxisome size(GO:0044375) |
| 0.3 | 3.8 | GO:0002347 | response to tumor cell(GO:0002347) |
| 0.3 | 0.9 | GO:1904735 | negative regulation of electron carrier activity(GO:1904733) regulation of fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:1904735) negative regulation of fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:1904736) |
| 0.3 | 3.7 | GO:0045648 | positive regulation of erythrocyte differentiation(GO:0045648) |
| 0.3 | 1.7 | GO:0006663 | platelet activating factor biosynthetic process(GO:0006663) |
| 0.3 | 3.6 | GO:0042541 | hemoglobin biosynthetic process(GO:0042541) |
| 0.3 | 1.6 | GO:0032483 | regulation of Rab protein signal transduction(GO:0032483) |
| 0.3 | 4.4 | GO:0043248 | proteasome assembly(GO:0043248) |
| 0.3 | 1.4 | GO:0031639 | plasminogen activation(GO:0031639) |
| 0.3 | 3.8 | GO:0032000 | positive regulation of fatty acid beta-oxidation(GO:0032000) |
| 0.3 | 3.0 | GO:0005981 | regulation of glycogen catabolic process(GO:0005981) |
| 0.3 | 2.7 | GO:0042407 | cristae formation(GO:0042407) |
| 0.3 | 1.0 | GO:0001732 | formation of cytoplasmic translation initiation complex(GO:0001732) |
| 0.3 | 2.4 | GO:0072677 | eosinophil chemotaxis(GO:0048245) eosinophil migration(GO:0072677) |
| 0.3 | 22.7 | GO:0006940 | regulation of smooth muscle contraction(GO:0006940) |
| 0.3 | 5.9 | GO:0042119 | neutrophil activation(GO:0042119) |
| 0.3 | 2.0 | GO:0051095 | regulation of helicase activity(GO:0051095) positive regulation of helicase activity(GO:0051096) |
| 0.3 | 6.8 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.3 | 1.0 | GO:0060744 | thelarche(GO:0042695) mammary gland branching involved in thelarche(GO:0060744) |
| 0.3 | 0.8 | GO:0035037 | sperm entry(GO:0035037) |
| 0.2 | 9.1 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.2 | 2.7 | GO:0035878 | nail development(GO:0035878) |
| 0.2 | 1.2 | GO:0006420 | arginyl-tRNA aminoacylation(GO:0006420) |
| 0.2 | 6.6 | GO:0017144 | drug metabolic process(GO:0017144) |
| 0.2 | 2.2 | GO:0006528 | asparagine metabolic process(GO:0006528) |
| 0.2 | 0.9 | GO:1902965 | regulation of protein localization to early endosome(GO:1902965) positive regulation of protein localization to early endosome(GO:1902966) |
| 0.2 | 2.1 | GO:0046884 | follicle-stimulating hormone secretion(GO:0046884) |
| 0.2 | 1.8 | GO:0046533 | regulation of photoreceptor cell differentiation(GO:0046532) negative regulation of photoreceptor cell differentiation(GO:0046533) |
| 0.2 | 3.3 | GO:0090527 | actin filament reorganization(GO:0090527) |
| 0.2 | 4.6 | GO:0016322 | neuron remodeling(GO:0016322) |
| 0.2 | 1.3 | GO:0036112 | medium-chain fatty-acyl-CoA metabolic process(GO:0036112) |
| 0.2 | 5.5 | GO:0070208 | protein heterotrimerization(GO:0070208) |
| 0.2 | 0.6 | GO:0048850 | hypophysis morphogenesis(GO:0048850) |
| 0.2 | 1.3 | GO:0090160 | positive regulation of pinocytosis(GO:0048549) Golgi to lysosome transport(GO:0090160) |
| 0.2 | 1.4 | GO:0000707 | meiotic DNA recombinase assembly(GO:0000707) |
| 0.2 | 1.4 | GO:0061502 | early endosome to recycling endosome transport(GO:0061502) |
| 0.2 | 1.4 | GO:0032377 | regulation of intracellular lipid transport(GO:0032377) regulation of intracellular sterol transport(GO:0032380) regulation of intracellular cholesterol transport(GO:0032383) |
| 0.2 | 0.8 | GO:0019853 | L-ascorbic acid biosynthetic process(GO:0019853) |
| 0.2 | 4.2 | GO:0031000 | response to caffeine(GO:0031000) |
| 0.2 | 0.4 | GO:0061357 | positive regulation of Wnt protein secretion(GO:0061357) |
| 0.2 | 1.9 | GO:1904707 | positive regulation of vascular smooth muscle cell proliferation(GO:1904707) |
| 0.2 | 1.7 | GO:0044336 | canonical Wnt signaling pathway involved in negative regulation of apoptotic process(GO:0044336) |
| 0.2 | 0.8 | GO:0032887 | regulation of spindle elongation(GO:0032887) regulation of mitotic spindle elongation(GO:0032888) anastral spindle assembly(GO:0055048) protein localization to spindle pole body(GO:0071988) regulation of protein localization to spindle pole body(GO:1902363) positive regulation of protein localization to spindle pole body(GO:1902365) positive regulation of mitotic spindle elongation(GO:1902846) |
| 0.2 | 6.3 | GO:0051482 | positive regulation of cytosolic calcium ion concentration involved in phospholipase C-activating G-protein coupled signaling pathway(GO:0051482) |
| 0.2 | 4.6 | GO:1903441 | protein localization to ciliary membrane(GO:1903441) |
| 0.2 | 3.1 | GO:0006622 | protein targeting to lysosome(GO:0006622) |
| 0.2 | 5.0 | GO:0043984 | histone H4-K16 acetylation(GO:0043984) |
| 0.2 | 14.7 | GO:1900046 | regulation of blood coagulation(GO:0030193) regulation of hemostasis(GO:1900046) |
| 0.2 | 1.5 | GO:0090074 | negative regulation of protein homodimerization activity(GO:0090074) |
| 0.2 | 3.7 | GO:1903427 | negative regulation of reactive oxygen species biosynthetic process(GO:1903427) |
| 0.2 | 0.4 | GO:2001176 | mediator complex assembly(GO:0036034) regulation of mediator complex assembly(GO:2001176) positive regulation of mediator complex assembly(GO:2001178) |
| 0.2 | 1.4 | GO:0097428 | protein maturation by iron-sulfur cluster transfer(GO:0097428) |
| 0.2 | 34.2 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.2 | 0.9 | GO:0018242 | protein O-linked glycosylation via serine(GO:0018242) |
| 0.2 | 3.3 | GO:0042088 | T-helper 1 type immune response(GO:0042088) |
| 0.2 | 1.6 | GO:0006271 | DNA strand elongation involved in DNA replication(GO:0006271) |
| 0.2 | 0.8 | GO:0034334 | adherens junction maintenance(GO:0034334) |
| 0.2 | 0.8 | GO:0006189 | 'de novo' IMP biosynthetic process(GO:0006189) |
| 0.2 | 4.1 | GO:0009072 | aromatic amino acid family metabolic process(GO:0009072) |
| 0.2 | 0.8 | GO:0016075 | rRNA catabolic process(GO:0016075) |
| 0.2 | 1.1 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.2 | 2.0 | GO:0048386 | positive regulation of retinoic acid receptor signaling pathway(GO:0048386) |
| 0.2 | 16.1 | GO:0019395 | fatty acid oxidation(GO:0019395) |
| 0.1 | 4.0 | GO:0045116 | protein neddylation(GO:0045116) |
| 0.1 | 7.2 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.1 | 1.8 | GO:0090091 | positive regulation of extracellular matrix disassembly(GO:0090091) |
| 0.1 | 2.8 | GO:0006465 | signal peptide processing(GO:0006465) |
| 0.1 | 5.2 | GO:0006767 | water-soluble vitamin metabolic process(GO:0006767) |
| 0.1 | 8.9 | GO:0002224 | toll-like receptor signaling pathway(GO:0002224) |
| 0.1 | 2.4 | GO:0060510 | Type II pneumocyte differentiation(GO:0060510) |
| 0.1 | 2.7 | GO:0000460 | maturation of 5.8S rRNA(GO:0000460) |
| 0.1 | 1.9 | GO:1904659 | glucose transmembrane transport(GO:1904659) |
| 0.1 | 12.1 | GO:0038083 | peptidyl-tyrosine autophosphorylation(GO:0038083) |
| 0.1 | 2.1 | GO:0021702 | cerebellar Purkinje cell differentiation(GO:0021702) |
| 0.1 | 0.4 | GO:0032218 | riboflavin transport(GO:0032218) |
| 0.1 | 5.9 | GO:0050829 | defense response to Gram-negative bacterium(GO:0050829) |
| 0.1 | 1.7 | GO:0001731 | formation of translation preinitiation complex(GO:0001731) |
| 0.1 | 1.7 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.1 | 1.7 | GO:0008300 | isoprenoid catabolic process(GO:0008300) |
| 0.1 | 1.4 | GO:0010606 | positive regulation of cytoplasmic mRNA processing body assembly(GO:0010606) |
| 0.1 | 1.4 | GO:1902231 | positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage(GO:1902231) |
| 0.1 | 0.5 | GO:0032298 | positive regulation of DNA-dependent DNA replication initiation(GO:0032298) |
| 0.1 | 1.7 | GO:1904778 | regulation of protein localization to cell cortex(GO:1904776) positive regulation of protein localization to cell cortex(GO:1904778) |
| 0.1 | 3.2 | GO:0001780 | neutrophil homeostasis(GO:0001780) |
| 0.1 | 1.0 | GO:0000291 | nuclear-transcribed mRNA catabolic process, exonucleolytic(GO:0000291) |
| 0.1 | 0.7 | GO:0045905 | translational frameshifting(GO:0006452) positive regulation of translational termination(GO:0045905) |
| 0.1 | 3.8 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.1 | 3.2 | GO:0006911 | phagocytosis, engulfment(GO:0006911) |
| 0.1 | 0.7 | GO:0008611 | ether lipid biosynthetic process(GO:0008611) glycerol ether biosynthetic process(GO:0046504) ether biosynthetic process(GO:1901503) |
| 0.1 | 0.4 | GO:0002036 | regulation of L-glutamate transport(GO:0002036) |
| 0.1 | 0.4 | GO:0006669 | sphinganine-1-phosphate biosynthetic process(GO:0006669) |
| 0.1 | 0.3 | GO:0051344 | negative regulation of cyclic-nucleotide phosphodiesterase activity(GO:0051344) |
| 0.1 | 0.9 | GO:0006032 | chitin metabolic process(GO:0006030) chitin catabolic process(GO:0006032) |
| 0.1 | 1.4 | GO:0015838 | amino-acid betaine transport(GO:0015838) |
| 0.1 | 1.7 | GO:0060628 | regulation of ER to Golgi vesicle-mediated transport(GO:0060628) |
| 0.1 | 4.2 | GO:1901998 | toxin transport(GO:1901998) |
| 0.1 | 1.4 | GO:0032801 | receptor catabolic process(GO:0032801) |
| 0.1 | 3.5 | GO:0046825 | regulation of protein export from nucleus(GO:0046825) |
| 0.1 | 2.7 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.1 | 7.8 | GO:0042632 | cholesterol homeostasis(GO:0042632) sterol homeostasis(GO:0055092) |
| 0.1 | 0.4 | GO:0042985 | negative regulation of amyloid precursor protein biosynthetic process(GO:0042985) |
| 0.1 | 0.1 | GO:0001914 | regulation of T cell mediated cytotoxicity(GO:0001914) |
| 0.1 | 1.5 | GO:0050832 | defense response to fungus(GO:0050832) |
| 0.1 | 0.4 | GO:1901837 | negative regulation of transcription of nuclear large rRNA transcript from RNA polymerase I promoter(GO:1901837) |
| 0.1 | 3.8 | GO:0000132 | establishment of mitotic spindle orientation(GO:0000132) |
| 0.1 | 8.8 | GO:0006487 | protein N-linked glycosylation(GO:0006487) |
| 0.1 | 4.9 | GO:0006695 | cholesterol biosynthetic process(GO:0006695) |
| 0.1 | 0.8 | GO:0032836 | glomerular basement membrane development(GO:0032836) |
| 0.1 | 2.1 | GO:0060712 | spongiotrophoblast layer development(GO:0060712) |
| 0.1 | 1.6 | GO:0071392 | cellular response to estradiol stimulus(GO:0071392) |
| 0.1 | 3.6 | GO:0002260 | lymphocyte homeostasis(GO:0002260) |
| 0.1 | 1.6 | GO:0035162 | embryonic hemopoiesis(GO:0035162) |
| 0.1 | 4.1 | GO:0008203 | cholesterol metabolic process(GO:0008203) |
| 0.1 | 0.8 | GO:0021842 | directional guidance of interneurons involved in migration from the subpallium to the cortex(GO:0021840) chemorepulsion involved in interneuron migration from the subpallium to the cortex(GO:0021842) |
| 0.1 | 1.4 | GO:0006516 | glycoprotein catabolic process(GO:0006516) |
| 0.1 | 0.6 | GO:0061462 | protein localization to lysosome(GO:0061462) |
| 0.1 | 0.2 | GO:0060467 | negative regulation of fertilization(GO:0060467) |
| 0.1 | 0.5 | GO:0090234 | regulation of kinetochore assembly(GO:0090234) |
| 0.1 | 1.0 | GO:0006054 | N-acetylneuraminate metabolic process(GO:0006054) |
| 0.1 | 0.2 | GO:0002034 | regulation of blood vessel size by renin-angiotensin(GO:0002034) renal control of peripheral vascular resistance involved in regulation of systemic arterial blood pressure(GO:0003072) |
| 0.1 | 5.1 | GO:0002250 | adaptive immune response(GO:0002250) |
| 0.1 | 0.7 | GO:0061727 | methylglyoxal metabolic process(GO:0009438) methylglyoxal catabolic process to D-lactate via S-lactoyl-glutathione(GO:0019243) methylglyoxal catabolic process(GO:0051596) methylglyoxal catabolic process to lactate(GO:0061727) |
| 0.1 | 7.5 | GO:0034446 | substrate adhesion-dependent cell spreading(GO:0034446) |
| 0.1 | 16.1 | GO:0009617 | response to bacterium(GO:0009617) |
| 0.1 | 3.7 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.1 | 0.4 | GO:0036337 | Fas signaling pathway(GO:0036337) |
| 0.1 | 0.1 | GO:0071169 | establishment of protein localization to chromatin(GO:0071169) |
| 0.1 | 0.5 | GO:0035729 | cellular response to hepatocyte growth factor stimulus(GO:0035729) |
| 0.1 | 1.0 | GO:0006388 | tRNA splicing, via endonucleolytic cleavage and ligation(GO:0006388) |
| 0.1 | 2.6 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.1 | 0.8 | GO:0045721 | negative regulation of gluconeogenesis(GO:0045721) |
| 0.1 | 0.5 | GO:0002155 | thyroid hormone mediated signaling pathway(GO:0002154) regulation of thyroid hormone mediated signaling pathway(GO:0002155) |
| 0.1 | 0.2 | GO:0006391 | transcription initiation from mitochondrial promoter(GO:0006391) |
| 0.1 | 1.9 | GO:0071526 | semaphorin-plexin signaling pathway(GO:0071526) |
| 0.1 | 0.8 | GO:0007220 | Notch receptor processing(GO:0007220) |
| 0.1 | 0.5 | GO:0046520 | sphingoid biosynthetic process(GO:0046520) |
| 0.1 | 2.3 | GO:0010718 | positive regulation of epithelial to mesenchymal transition(GO:0010718) |
| 0.1 | 0.1 | GO:0045085 | negative regulation of interleukin-2 biosynthetic process(GO:0045085) |
| 0.1 | 0.7 | GO:0052696 | flavonoid metabolic process(GO:0009812) flavonoid glucuronidation(GO:0052696) xenobiotic glucuronidation(GO:0052697) |
| 0.1 | 0.1 | GO:0006207 | 'de novo' pyrimidine nucleobase biosynthetic process(GO:0006207) pyrimidine nucleobase biosynthetic process(GO:0019856) |
| 0.1 | 0.3 | GO:0070914 | UV-damage excision repair(GO:0070914) |
| 0.1 | 1.4 | GO:0000305 | response to oxygen radical(GO:0000305) |
| 0.1 | 1.2 | GO:0001893 | maternal placenta development(GO:0001893) |
| 0.1 | 1.8 | GO:0032088 | negative regulation of NF-kappaB transcription factor activity(GO:0032088) |
| 0.1 | 0.4 | GO:0030214 | hyaluronan catabolic process(GO:0030214) |
| 0.1 | 2.9 | GO:0006939 | smooth muscle contraction(GO:0006939) |
| 0.1 | 1.1 | GO:0030033 | microvillus assembly(GO:0030033) |
| 0.1 | 5.8 | GO:0042098 | T cell proliferation(GO:0042098) |
| 0.1 | 0.7 | GO:0051418 | interphase microtubule nucleation by interphase microtubule organizing center(GO:0051415) microtubule nucleation by microtubule organizing center(GO:0051418) |
| 0.1 | 1.2 | GO:0044783 | mitotic G1 DNA damage checkpoint(GO:0031571) G1 DNA damage checkpoint(GO:0044783) |
| 0.1 | 0.4 | GO:1900037 | regulation of cellular response to hypoxia(GO:1900037) |
| 0.1 | 2.8 | GO:0071277 | cellular response to calcium ion(GO:0071277) |
| 0.1 | 0.8 | GO:0033962 | cytoplasmic mRNA processing body assembly(GO:0033962) |
| 0.1 | 1.8 | GO:0048025 | negative regulation of mRNA splicing, via spliceosome(GO:0048025) |
| 0.1 | 2.1 | GO:0003170 | heart valve development(GO:0003170) |
| 0.1 | 4.0 | GO:0045766 | positive regulation of angiogenesis(GO:0045766) |
| 0.0 | 0.4 | GO:0007250 | activation of NF-kappaB-inducing kinase activity(GO:0007250) |
| 0.0 | 0.9 | GO:0030238 | male sex determination(GO:0030238) |
| 0.0 | 0.4 | GO:0035067 | negative regulation of histone acetylation(GO:0035067) |
| 0.0 | 1.5 | GO:0032467 | positive regulation of cytokinesis(GO:0032467) |
| 0.0 | 1.5 | GO:0001702 | gastrulation with mouth forming second(GO:0001702) |
| 0.0 | 1.0 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 0.0 | 0.6 | GO:0042761 | very long-chain fatty acid biosynthetic process(GO:0042761) |
| 0.0 | 0.4 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.0 | 2.1 | GO:0051972 | regulation of telomerase activity(GO:0051972) |
| 0.0 | 0.4 | GO:0015961 | diadenosine polyphosphate catabolic process(GO:0015961) |
| 0.0 | 0.3 | GO:0018094 | protein polyglycylation(GO:0018094) |
| 0.0 | 0.3 | GO:0043277 | apoptotic cell clearance(GO:0043277) |
| 0.0 | 1.7 | GO:0044380 | protein localization to cytoskeleton(GO:0044380) |
| 0.0 | 2.9 | GO:0071774 | response to fibroblast growth factor(GO:0071774) |
| 0.0 | 0.7 | GO:0045056 | transcytosis(GO:0045056) |
| 0.0 | 0.8 | GO:0045898 | regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045898) |
| 0.0 | 2.0 | GO:0006909 | phagocytosis(GO:0006909) |
| 0.0 | 0.4 | GO:0006105 | succinate metabolic process(GO:0006105) |
| 0.0 | 2.4 | GO:0006637 | acyl-CoA metabolic process(GO:0006637) thioester metabolic process(GO:0035383) |
| 0.0 | 1.2 | GO:0007339 | binding of sperm to zona pellucida(GO:0007339) |
| 0.0 | 0.5 | GO:1902042 | negative regulation of extrinsic apoptotic signaling pathway via death domain receptors(GO:1902042) |
| 0.0 | 0.3 | GO:0036265 | RNA (guanine-N7)-methylation(GO:0036265) |
| 0.0 | 1.0 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.0 | 5.1 | GO:0030308 | negative regulation of cell growth(GO:0030308) |
| 0.0 | 0.6 | GO:0030490 | maturation of SSU-rRNA(GO:0030490) |
| 0.0 | 0.5 | GO:0001886 | endothelial cell morphogenesis(GO:0001886) |
| 0.0 | 1.8 | GO:0006513 | protein monoubiquitination(GO:0006513) |
| 0.0 | 0.1 | GO:0061090 | positive regulation of sequestering of zinc ion(GO:0061090) |
| 0.0 | 0.2 | GO:0040033 | miRNA mediated inhibition of translation(GO:0035278) negative regulation of translation, ncRNA-mediated(GO:0040033) regulation of translation, ncRNA-mediated(GO:0045974) |
| 0.0 | 3.0 | GO:0009791 | post-embryonic development(GO:0009791) |
| 0.0 | 0.4 | GO:0046415 | urate metabolic process(GO:0046415) |
| 0.0 | 2.5 | GO:0050821 | protein stabilization(GO:0050821) |
| 0.0 | 0.1 | GO:2000535 | exocyst assembly(GO:0001927) entry of bacterium into host cell(GO:0035635) regulation of entry of bacterium into host cell(GO:2000535) |
| 0.0 | 0.2 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.0 | 1.7 | GO:0034101 | erythrocyte homeostasis(GO:0034101) |
| 0.0 | 0.1 | GO:1990928 | response to amino acid starvation(GO:1990928) |
| 0.0 | 1.6 | GO:0007098 | centrosome cycle(GO:0007098) |
| 0.0 | 0.7 | GO:0043001 | Golgi to plasma membrane protein transport(GO:0043001) |
| 0.0 | 1.0 | GO:0042113 | B cell activation(GO:0042113) |
| 0.0 | 0.7 | GO:0042158 | protein lipidation(GO:0006497) lipoprotein biosynthetic process(GO:0042158) |
| 0.0 | 0.9 | GO:0008033 | tRNA processing(GO:0008033) |
| 0.0 | 0.2 | GO:0060325 | face morphogenesis(GO:0060325) |
| 0.0 | 0.2 | GO:0045943 | positive regulation of transcription from RNA polymerase I promoter(GO:0045943) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 13.7 | 191.3 | GO:0020005 | symbiont-containing vacuole membrane(GO:0020005) |
| 9.3 | 74.4 | GO:0042825 | TAP complex(GO:0042825) |
| 8.4 | 118.1 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 8.4 | 16.9 | GO:0020003 | symbiont-containing vacuole(GO:0020003) |
| 8.0 | 48.2 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 7.7 | 30.8 | GO:0097169 | AIM2 inflammasome complex(GO:0097169) |
| 7.5 | 44.7 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 5.8 | 23.2 | GO:0072559 | NLRP3 inflammasome complex(GO:0072559) |
| 3.0 | 27.3 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) |
| 3.0 | 14.8 | GO:0072557 | IPAF inflammasome complex(GO:0072557) |
| 2.5 | 7.6 | GO:0032127 | dense core granule membrane(GO:0032127) |
| 2.4 | 7.1 | GO:0097132 | cyclin D2-CDK6 complex(GO:0097132) |
| 2.3 | 22.7 | GO:0030478 | actin cap(GO:0030478) |
| 2.2 | 6.5 | GO:0070557 | PCNA-p21 complex(GO:0070557) |
| 2.1 | 14.8 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 2.1 | 33.9 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 2.0 | 20.3 | GO:1990462 | omegasome(GO:1990462) |
| 1.8 | 30.1 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 1.7 | 8.7 | GO:0030915 | Smc5-Smc6 complex(GO:0030915) |
| 1.6 | 21.0 | GO:0042611 | MHC protein complex(GO:0042611) |
| 1.5 | 9.2 | GO:0005663 | DNA replication factor C complex(GO:0005663) |
| 1.4 | 4.3 | GO:0097224 | sperm connecting piece(GO:0097224) |
| 1.4 | 5.8 | GO:0090661 | box H/ACA telomerase RNP complex(GO:0090661) |
| 1.4 | 21.9 | GO:0044754 | autolysosome(GO:0044754) |
| 1.3 | 3.9 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 1.2 | 14.7 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 1.2 | 4.8 | GO:0097447 | dendritic tree(GO:0097447) |
| 1.1 | 36.4 | GO:0031233 | intrinsic component of external side of plasma membrane(GO:0031233) |
| 1.1 | 13.9 | GO:0033256 | I-kappaB/NF-kappaB complex(GO:0033256) |
| 1.1 | 3.2 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 1.0 | 3.1 | GO:0000814 | ESCRT II complex(GO:0000814) |
| 1.0 | 13.5 | GO:0000778 | condensed nuclear chromosome kinetochore(GO:0000778) |
| 0.9 | 129.7 | GO:0016605 | PML body(GO:0016605) |
| 0.9 | 4.6 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.9 | 8.7 | GO:0000796 | condensin complex(GO:0000796) |
| 0.8 | 5.9 | GO:0071458 | integral component of cytoplasmic side of endoplasmic reticulum membrane(GO:0071458) integral component of lumenal side of endoplasmic reticulum membrane(GO:0071556) lumenal side of endoplasmic reticulum membrane(GO:0098553) |
| 0.8 | 0.8 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.8 | 20.6 | GO:0000145 | exocyst(GO:0000145) |
| 0.8 | 2.4 | GO:1990666 | PCSK9-LDLR complex(GO:1990666) |
| 0.8 | 29.9 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.8 | 3.1 | GO:0043202 | lysosomal lumen(GO:0043202) |
| 0.8 | 4.5 | GO:0031390 | Ctf18 RFC-like complex(GO:0031390) |
| 0.7 | 2.2 | GO:0005900 | oncostatin-M receptor complex(GO:0005900) |
| 0.7 | 2.2 | GO:0070992 | translation initiation complex(GO:0070992) |
| 0.7 | 5.0 | GO:0072487 | MSL complex(GO:0072487) |
| 0.7 | 9.3 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.7 | 51.8 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.6 | 6.5 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 0.5 | 2.7 | GO:0017177 | glucosidase II complex(GO:0017177) |
| 0.5 | 279.3 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.5 | 30.7 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.5 | 4.0 | GO:0045293 | mRNA editing complex(GO:0045293) |
| 0.5 | 12.6 | GO:0043196 | varicosity(GO:0043196) |
| 0.5 | 15.0 | GO:0005868 | cytoplasmic dynein complex(GO:0005868) |
| 0.4 | 2.7 | GO:0061617 | MICOS complex(GO:0061617) |
| 0.4 | 1.7 | GO:0060171 | stereocilium membrane(GO:0060171) |
| 0.4 | 4.3 | GO:0005677 | chromatin silencing complex(GO:0005677) |
| 0.4 | 2.1 | GO:0000839 | Hrd1p ubiquitin ligase ERAD-L complex(GO:0000839) |
| 0.4 | 3.9 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.4 | 0.4 | GO:0008024 | cyclin/CDK positive transcription elongation factor complex(GO:0008024) |
| 0.4 | 16.0 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.4 | 4.2 | GO:0070531 | BRCA1-A complex(GO:0070531) |
| 0.4 | 3.3 | GO:0045298 | tubulin complex(GO:0045298) |
| 0.4 | 212.3 | GO:0005789 | endoplasmic reticulum membrane(GO:0005789) |
| 0.4 | 48.7 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.4 | 1.8 | GO:0097058 | CRLF-CLCF1 complex(GO:0097058) |
| 0.4 | 9.1 | GO:0016471 | vacuolar proton-transporting V-type ATPase complex(GO:0016471) |
| 0.3 | 2.7 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.3 | 3.0 | GO:0042587 | glycogen granule(GO:0042587) |
| 0.3 | 1.6 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 0.3 | 3.4 | GO:0061700 | GATOR2 complex(GO:0061700) |
| 0.3 | 2.1 | GO:1990357 | terminal web(GO:1990357) |
| 0.3 | 0.9 | GO:0030896 | checkpoint clamp complex(GO:0030896) |
| 0.3 | 1.1 | GO:0002142 | stereocilia ankle link complex(GO:0002142) USH2 complex(GO:1990696) |
| 0.3 | 1.4 | GO:0031251 | PAN complex(GO:0031251) |
| 0.3 | 334.6 | GO:0005730 | nucleolus(GO:0005730) |
| 0.3 | 1.9 | GO:0036157 | outer dynein arm(GO:0036157) |
| 0.3 | 4.8 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.3 | 2.4 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.3 | 1.5 | GO:0033503 | HULC complex(GO:0033503) |
| 0.2 | 29.7 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.2 | 6.3 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.2 | 2.4 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.2 | 2.3 | GO:0044354 | pinosome(GO:0044352) macropinosome(GO:0044354) |
| 0.2 | 1.4 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.2 | 1.6 | GO:0043657 | host(GO:0018995) host cell part(GO:0033643) host cell(GO:0043657) |
| 0.2 | 4.1 | GO:0034362 | low-density lipoprotein particle(GO:0034362) |
| 0.2 | 15.9 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.2 | 1.4 | GO:0033063 | Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 0.2 | 45.6 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.2 | 1.0 | GO:0005914 | spot adherens junction(GO:0005914) |
| 0.2 | 11.6 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.2 | 4.4 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.2 | 2.8 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 0.2 | 1.1 | GO:0045009 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.2 | 1.7 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.2 | 5.4 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.2 | 2.6 | GO:0042555 | MCM complex(GO:0042555) |
| 0.2 | 7.7 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.1 | 11.9 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.1 | 2.2 | GO:0005697 | telomerase holoenzyme complex(GO:0005697) |
| 0.1 | 2.7 | GO:0070971 | endoplasmic reticulum exit site(GO:0070971) |
| 0.1 | 1.6 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.1 | 1.8 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.1 | 2.9 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.1 | 4.0 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.1 | 0.6 | GO:0044307 | dendritic branch(GO:0044307) |
| 0.1 | 0.4 | GO:0045257 | mitochondrial respiratory chain complex II, succinate dehydrogenase complex (ubiquinone)(GO:0005749) succinate dehydrogenase complex (ubiquinone)(GO:0045257) respiratory chain complex II(GO:0045273) succinate dehydrogenase complex(GO:0045281) fumarate reductase complex(GO:0045283) |
| 0.1 | 9.7 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.1 | 1.7 | GO:0031462 | Cul2-RING ubiquitin ligase complex(GO:0031462) |
| 0.1 | 70.4 | GO:0005764 | lytic vacuole(GO:0000323) lysosome(GO:0005764) |
| 0.1 | 2.5 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.1 | 0.2 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.1 | 0.4 | GO:0043540 | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase complex(GO:0043540) |
| 0.1 | 0.6 | GO:0043293 | apoptosome(GO:0043293) |
| 0.1 | 1.4 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.1 | 3.0 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.1 | 1.2 | GO:0070938 | contractile ring(GO:0070938) |
| 0.1 | 13.8 | GO:0000775 | chromosome, centromeric region(GO:0000775) |
| 0.1 | 3.9 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.1 | 0.8 | GO:0098645 | collagen type IV trimer(GO:0005587) network-forming collagen trimer(GO:0098642) collagen network(GO:0098645) basement membrane collagen trimer(GO:0098651) |
| 0.1 | 14.0 | GO:0005604 | basement membrane(GO:0005604) |
| 0.1 | 0.3 | GO:0017071 | intracellular cyclic nucleotide activated cation channel complex(GO:0017071) |
| 0.1 | 0.9 | GO:0051286 | cell tip(GO:0051286) |
| 0.1 | 0.3 | GO:0034751 | aryl hydrocarbon receptor complex(GO:0034751) |
| 0.1 | 2.0 | GO:0097470 | ribbon synapse(GO:0097470) |
| 0.1 | 0.9 | GO:0097197 | tetraspanin-enriched microdomain(GO:0097197) |
| 0.1 | 0.6 | GO:0044327 | dendritic spine head(GO:0044327) |
| 0.1 | 0.7 | GO:0000931 | gamma-tubulin large complex(GO:0000931) gamma-tubulin ring complex(GO:0008274) |
| 0.1 | 0.7 | GO:0005642 | annulate lamellae(GO:0005642) |
| 0.1 | 1.1 | GO:0034045 | pre-autophagosomal structure membrane(GO:0034045) |
| 0.1 | 6.1 | GO:0030175 | filopodium(GO:0030175) |
| 0.1 | 1.0 | GO:0005852 | eukaryotic translation initiation factor 3 complex(GO:0005852) |
| 0.1 | 23.0 | GO:0009898 | cytoplasmic side of plasma membrane(GO:0009898) |
| 0.1 | 3.2 | GO:1990391 | DNA repair complex(GO:1990391) |
| 0.1 | 315.2 | GO:0005829 | cytosol(GO:0005829) |
| 0.1 | 3.3 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.1 | 0.7 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.1 | 2.1 | GO:0015935 | small ribosomal subunit(GO:0015935) |
| 0.1 | 0.8 | GO:0070765 | gamma-secretase complex(GO:0070765) |
| 0.1 | 0.9 | GO:0030122 | AP-2 adaptor complex(GO:0030122) |
| 0.1 | 4.7 | GO:0005884 | actin filament(GO:0005884) |
| 0.1 | 1.1 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.1 | 1.8 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.1 | 2.3 | GO:0045095 | keratin filament(GO:0045095) |
| 0.1 | 1.9 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.0 | 1.9 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 23.1 | GO:0009986 | cell surface(GO:0009986) |
| 0.0 | 1.0 | GO:0005669 | transcription factor TFIID complex(GO:0005669) |
| 0.0 | 6.1 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.0 | 1.5 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.0 | 2.6 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.0 | 0.6 | GO:0031010 | ISWI-type complex(GO:0031010) |
| 0.0 | 3.6 | GO:0005635 | nuclear envelope(GO:0005635) |
| 0.0 | 0.9 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.0 | 136.0 | GO:0005576 | extracellular region(GO:0005576) |
| 0.0 | 1.4 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 0.3 | GO:0031011 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
| 0.0 | 0.3 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.0 | 1.0 | GO:0001725 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.0 | 2.2 | GO:0005903 | brush border(GO:0005903) |
| 0.0 | 0.6 | GO:0001726 | ruffle(GO:0001726) |
| 0.0 | 0.5 | GO:0030008 | TRAPP complex(GO:0030008) |
| 0.0 | 0.2 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.0 | 0.8 | GO:0031970 | organelle envelope lumen(GO:0031970) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 16.7 | 50.1 | GO:0015433 | peptide antigen-transporting ATPase activity(GO:0015433) tapasin binding(GO:0046980) |
| 12.4 | 37.3 | GO:0003692 | left-handed Z-DNA binding(GO:0003692) |
| 10.2 | 51.1 | GO:0008859 | exoribonuclease II activity(GO:0008859) |
| 6.4 | 95.8 | GO:0046977 | TAP binding(GO:0046977) |
| 6.4 | 76.4 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 5.9 | 53.2 | GO:0016936 | galactoside binding(GO:0016936) |
| 5.8 | 174.6 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 5.6 | 27.8 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 5.1 | 96.9 | GO:0097153 | cysteine-type endopeptidase activity involved in apoptotic process(GO:0097153) |
| 4.5 | 13.5 | GO:0005174 | CD40 receptor binding(GO:0005174) |
| 4.4 | 48.9 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 3.8 | 11.3 | GO:0032394 | MHC class Ib receptor activity(GO:0032394) |
| 3.7 | 18.7 | GO:0070892 | lipoteichoic acid receptor activity(GO:0070892) |
| 3.6 | 21.3 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 3.4 | 37.9 | GO:0016174 | NAD(P)H oxidase activity(GO:0016174) |
| 3.4 | 6.8 | GO:0003953 | NAD+ nucleosidase activity(GO:0003953) |
| 3.3 | 39.9 | GO:0031730 | CCR5 chemokine receptor binding(GO:0031730) |
| 2.9 | 47.0 | GO:0031386 | protein tag(GO:0031386) |
| 2.9 | 8.7 | GO:0019002 | GMP binding(GO:0019002) |
| 2.9 | 40.4 | GO:0005041 | low-density lipoprotein receptor activity(GO:0005041) |
| 2.9 | 17.1 | GO:0047844 | deoxycytidine deaminase activity(GO:0047844) |
| 2.8 | 33.9 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 2.8 | 94.5 | GO:0005031 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 2.7 | 10.9 | GO:0004694 | eukaryotic translation initiation factor 2alpha kinase activity(GO:0004694) |
| 2.6 | 15.5 | GO:0042296 | ISG15 transferase activity(GO:0042296) |
| 2.6 | 7.7 | GO:0030337 | DNA polymerase processivity factor activity(GO:0030337) |
| 2.5 | 7.5 | GO:0005026 | transforming growth factor beta receptor activity, type II(GO:0005026) |
| 2.5 | 22.4 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 2.4 | 14.7 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 2.4 | 23.9 | GO:0019763 | immunoglobulin receptor activity(GO:0019763) |
| 2.3 | 13.6 | GO:0030881 | beta-2-microglobulin binding(GO:0030881) |
| 2.3 | 6.8 | GO:0031735 | CCR10 chemokine receptor binding(GO:0031735) |
| 2.3 | 22.7 | GO:1901612 | cardiolipin binding(GO:1901612) |
| 2.2 | 6.6 | GO:0042015 | interleukin-20 binding(GO:0042015) |
| 2.1 | 21.0 | GO:0050700 | CARD domain binding(GO:0050700) |
| 2.1 | 12.4 | GO:0032450 | maltose alpha-glucosidase activity(GO:0032450) |
| 2.0 | 46.6 | GO:0070628 | proteasome binding(GO:0070628) |
| 1.8 | 7.1 | GO:0098770 | FBXO family protein binding(GO:0098770) |
| 1.8 | 8.8 | GO:0008853 | exodeoxyribonuclease III activity(GO:0008853) |
| 1.8 | 5.3 | GO:0005171 | hepatocyte growth factor receptor binding(GO:0005171) |
| 1.8 | 5.3 | GO:0050785 | advanced glycation end-product receptor activity(GO:0050785) |
| 1.7 | 41.8 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 1.7 | 42.9 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 1.7 | 8.3 | GO:0004974 | leukotriene receptor activity(GO:0004974) |
| 1.6 | 6.6 | GO:0016823 | hydrolase activity, acting on acid carbon-carbon bonds(GO:0016822) hydrolase activity, acting on acid carbon-carbon bonds, in ketonic substances(GO:0016823) |
| 1.6 | 9.4 | GO:0004483 | mRNA (nucleoside-2'-O-)-methyltransferase activity(GO:0004483) |
| 1.5 | 13.8 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 1.5 | 4.5 | GO:0045142 | triplex DNA binding(GO:0045142) |
| 1.4 | 4.3 | GO:0001716 | L-amino-acid oxidase activity(GO:0001716) |
| 1.4 | 7.1 | GO:0004534 | 5'-3' exoribonuclease activity(GO:0004534) |
| 1.4 | 35.2 | GO:0051400 | BH domain binding(GO:0051400) |
| 1.4 | 1.4 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) |
| 1.4 | 4.2 | GO:0016749 | 5-aminolevulinate synthase activity(GO:0003870) N-succinyltransferase activity(GO:0016749) |
| 1.4 | 6.8 | GO:0031996 | thioesterase binding(GO:0031996) |
| 1.3 | 21.5 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 1.3 | 14.7 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 1.3 | 5.3 | GO:0070976 | TIR domain binding(GO:0070976) |
| 1.3 | 3.8 | GO:1990698 | palmitoleoyltransferase activity(GO:1990698) |
| 1.2 | 22.4 | GO:0008329 | signaling pattern recognition receptor activity(GO:0008329) pattern recognition receptor activity(GO:0038187) |
| 1.2 | 4.8 | GO:0016743 | carboxyl- or carbamoyltransferase activity(GO:0016743) |
| 1.1 | 8.0 | GO:0061513 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 1.1 | 25.1 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 1.1 | 45.6 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 1.1 | 4.2 | GO:0031752 | D5 dopamine receptor binding(GO:0031752) |
| 1.0 | 4.1 | GO:0030229 | very-low-density lipoprotein particle receptor activity(GO:0030229) |
| 1.0 | 3.1 | GO:0035175 | histone kinase activity (H3-S10 specific)(GO:0035175) |
| 1.0 | 8.3 | GO:0000293 | ferric-chelate reductase activity(GO:0000293) |
| 1.0 | 372.9 | GO:0003924 | GTPase activity(GO:0003924) |
| 1.0 | 14.4 | GO:0050897 | cobalt ion binding(GO:0050897) |
| 1.0 | 276.0 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 1.0 | 5.0 | GO:0045322 | unmethylated CpG binding(GO:0045322) |
| 1.0 | 4.8 | GO:0004060 | arylamine N-acetyltransferase activity(GO:0004060) |
| 0.9 | 5.7 | GO:0060698 | endoribonuclease inhibitor activity(GO:0060698) |
| 0.9 | 3.7 | GO:0004127 | cytidylate kinase activity(GO:0004127) uridylate kinase activity(GO:0009041) |
| 0.9 | 4.6 | GO:0045340 | mercury ion binding(GO:0045340) |
| 0.9 | 6.9 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.8 | 10.1 | GO:0070181 | small ribosomal subunit rRNA binding(GO:0070181) |
| 0.8 | 9.2 | GO:0033170 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.8 | 20.9 | GO:0005537 | mannose binding(GO:0005537) |
| 0.8 | 6.7 | GO:0004726 | non-membrane spanning protein tyrosine phosphatase activity(GO:0004726) |
| 0.8 | 5.8 | GO:0034513 | box H/ACA snoRNA binding(GO:0034513) |
| 0.8 | 4.9 | GO:0038051 | glucocorticoid receptor activity(GO:0004883) transcription factor activity, ligand-activated RNA polymerase II transcription factor binding(GO:0038049) glucocorticoid-activated RNA polymerase II transcription factor binding transcription factor activity(GO:0038051) |
| 0.8 | 2.5 | GO:0004510 | tryptophan 5-monooxygenase activity(GO:0004510) |
| 0.8 | 7.0 | GO:0048019 | receptor antagonist activity(GO:0048019) |
| 0.8 | 4.7 | GO:0034618 | arginine binding(GO:0034618) |
| 0.8 | 3.0 | GO:0005008 | hepatocyte growth factor-activated receptor activity(GO:0005008) |
| 0.8 | 2.3 | GO:0004912 | interleukin-3 receptor activity(GO:0004912) interleukin-3 binding(GO:0019978) |
| 0.7 | 2.2 | GO:0047651 | hydrolase activity, acting on acid halide bonds(GO:0016824) hydrolase activity, acting on acid halide bonds, in C-halide compounds(GO:0019120) alkylhalidase activity(GO:0047651) |
| 0.7 | 3.7 | GO:0004528 | phosphodiesterase I activity(GO:0004528) |
| 0.7 | 2.9 | GO:0050501 | hyaluronan synthase activity(GO:0050501) |
| 0.7 | 7.7 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.7 | 4.8 | GO:0004769 | steroid delta-isomerase activity(GO:0004769) |
| 0.7 | 3.4 | GO:0008475 | procollagen-lysine 5-dioxygenase activity(GO:0008475) |
| 0.7 | 13.0 | GO:0046703 | natural killer cell lectin-like receptor binding(GO:0046703) |
| 0.6 | 8.4 | GO:0001069 | regulatory region RNA binding(GO:0001069) |
| 0.6 | 4.3 | GO:0032027 | myosin light chain binding(GO:0032027) |
| 0.6 | 1.8 | GO:0008775 | acetate CoA-transferase activity(GO:0008775) |
| 0.6 | 5.9 | GO:0042500 | aspartic endopeptidase activity, intramembrane cleaving(GO:0042500) |
| 0.6 | 2.9 | GO:0072345 | NAADP-sensitive calcium-release channel activity(GO:0072345) |
| 0.6 | 4.1 | GO:0015265 | urea channel activity(GO:0015265) |
| 0.6 | 1.8 | GO:0005017 | platelet-derived growth factor-activated receptor activity(GO:0005017) |
| 0.6 | 6.9 | GO:0008381 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.6 | 3.3 | GO:0004614 | phosphoglucomutase activity(GO:0004614) |
| 0.5 | 52.1 | GO:0003725 | double-stranded RNA binding(GO:0003725) |
| 0.5 | 1.6 | GO:0052658 | inositol-1,4,5-trisphosphate 5-phosphatase activity(GO:0052658) |
| 0.5 | 6.6 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.5 | 3.8 | GO:0043515 | kinetochore binding(GO:0043515) |
| 0.5 | 2.7 | GO:0090599 | alpha-glucosidase activity(GO:0090599) |
| 0.5 | 2.2 | GO:0004924 | oncostatin-M receptor activity(GO:0004924) |
| 0.5 | 4.3 | GO:0097157 | pre-mRNA intronic binding(GO:0097157) |
| 0.5 | 3.2 | GO:0070644 | vitamin D response element binding(GO:0070644) |
| 0.5 | 63.0 | GO:0000979 | RNA polymerase II core promoter sequence-specific DNA binding(GO:0000979) |
| 0.5 | 5.5 | GO:0003956 | NAD(P)+-protein-arginine ADP-ribosyltransferase activity(GO:0003956) |
| 0.5 | 4.0 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 0.5 | 2.0 | GO:0008311 | double-stranded DNA 3'-5' exodeoxyribonuclease activity(GO:0008311) |
| 0.5 | 2.0 | GO:0008515 | sucrose:proton symporter activity(GO:0008506) sucrose transmembrane transporter activity(GO:0008515) disaccharide transmembrane transporter activity(GO:0015154) oligosaccharide transmembrane transporter activity(GO:0015157) |
| 0.5 | 25.0 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.5 | 1.9 | GO:0001512 | dihydronicotinamide riboside quinone reductase activity(GO:0001512) melatonin binding(GO:1904408) |
| 0.5 | 3.8 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.5 | 2.4 | GO:0005143 | interleukin-12 receptor binding(GO:0005143) |
| 0.5 | 1.4 | GO:0047192 | 1-alkylglycerophosphocholine O-acetyltransferase activity(GO:0047192) |
| 0.5 | 17.9 | GO:0001968 | fibronectin binding(GO:0001968) |
| 0.5 | 22.6 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.5 | 3.7 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.5 | 8.7 | GO:0045028 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.5 | 22.3 | GO:0051539 | 4 iron, 4 sulfur cluster binding(GO:0051539) |
| 0.5 | 24.8 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.4 | 1.3 | GO:0016508 | long-chain-enoyl-CoA hydratase activity(GO:0016508) |
| 0.4 | 4.4 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.4 | 2.6 | GO:0004832 | valine-tRNA ligase activity(GO:0004832) |
| 0.4 | 9.3 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 0.4 | 55.2 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.4 | 5.4 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.4 | 13.6 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
| 0.4 | 2.9 | GO:0004791 | thioredoxin-disulfide reductase activity(GO:0004791) |
| 0.4 | 90.9 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.4 | 31.3 | GO:0030971 | receptor tyrosine kinase binding(GO:0030971) |
| 0.4 | 4.8 | GO:0102338 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.4 | 0.4 | GO:0061628 | H3K27me3 modified histone binding(GO:0061628) |
| 0.4 | 5.2 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 0.4 | 1.4 | GO:0003844 | 1,4-alpha-glucan branching enzyme activity(GO:0003844) |
| 0.3 | 3.5 | GO:0043140 | ATP-dependent 3'-5' DNA helicase activity(GO:0043140) |
| 0.3 | 3.1 | GO:0055104 | ligase inhibitor activity(GO:0055104) ubiquitin ligase inhibitor activity(GO:1990948) |
| 0.3 | 18.7 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.3 | 3.3 | GO:0070513 | death domain binding(GO:0070513) |
| 0.3 | 1.0 | GO:0016426 | tRNA (adenine) methyltransferase activity(GO:0016426) |
| 0.3 | 2.3 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.3 | 3.1 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.3 | 9.4 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.3 | 3.4 | GO:0015217 | ATP transmembrane transporter activity(GO:0005347) ADP transmembrane transporter activity(GO:0015217) |
| 0.3 | 2.4 | GO:0003996 | acyl-CoA ligase activity(GO:0003996) |
| 0.3 | 16.1 | GO:0004520 | endodeoxyribonuclease activity(GO:0004520) |
| 0.3 | 12.4 | GO:0015036 | disulfide oxidoreductase activity(GO:0015036) |
| 0.3 | 11.8 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.3 | 1.4 | GO:0004661 | protein geranylgeranyltransferase activity(GO:0004661) |
| 0.3 | 16.7 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.3 | 2.2 | GO:1990247 | N6-methyladenosine-containing RNA binding(GO:1990247) |
| 0.3 | 2.2 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.3 | 0.8 | GO:0004637 | phosphoribosylamine-glycine ligase activity(GO:0004637) |
| 0.3 | 3.1 | GO:0003756 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.2 | 16.3 | GO:0016836 | hydro-lyase activity(GO:0016836) |
| 0.2 | 27.9 | GO:0036459 | thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.2 | 1.4 | GO:0016647 | oxidoreductase activity, acting on the CH-NH group of donors, oxygen as acceptor(GO:0016647) |
| 0.2 | 1.6 | GO:0042328 | heparan sulfate N-acetylglucosaminyltransferase activity(GO:0042328) |
| 0.2 | 2.3 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.2 | 0.2 | GO:0070815 | peptidyl-lysine 5-dioxygenase activity(GO:0070815) |
| 0.2 | 3.9 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.2 | 12.7 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.2 | 5.4 | GO:0070003 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.2 | 2.4 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 0.2 | 1.3 | GO:0047023 | androsterone dehydrogenase activity(GO:0047023) |
| 0.2 | 3.3 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.2 | 2.0 | GO:0030267 | hydroxypyruvate reductase activity(GO:0016618) glyoxylate reductase (NADP) activity(GO:0030267) |
| 0.2 | 0.8 | GO:0030629 | U6 snRNA 3'-end binding(GO:0030629) |
| 0.2 | 3.9 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 0.2 | 2.4 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.2 | 1.4 | GO:0050694 | galactose 3-O-sulfotransferase activity(GO:0050694) |
| 0.2 | 9.6 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.2 | 17.5 | GO:0042805 | actinin binding(GO:0042805) |
| 0.2 | 1.5 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.2 | 1.0 | GO:0008761 | UDP-N-acetylglucosamine 2-epimerase activity(GO:0008761) |
| 0.2 | 22.1 | GO:0033613 | activating transcription factor binding(GO:0033613) |
| 0.2 | 1.7 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.2 | 1.1 | GO:0010521 | telomerase inhibitor activity(GO:0010521) |
| 0.2 | 0.4 | GO:0046965 | retinoid X receptor binding(GO:0046965) |
| 0.2 | 1.1 | GO:0016716 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, another compound as one donor, and incorporation of one atom of oxygen(GO:0016716) |
| 0.2 | 1.2 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.2 | 0.4 | GO:0033906 | hyaluronoglucuronidase activity(GO:0033906) |
| 0.2 | 0.3 | GO:0043142 | single-stranded DNA-dependent ATPase activity(GO:0043142) |
| 0.2 | 3.6 | GO:0015174 | basic amino acid transmembrane transporter activity(GO:0015174) |
| 0.2 | 0.5 | GO:0031531 | thyrotropin-releasing hormone receptor binding(GO:0031531) |
| 0.2 | 1.5 | GO:0030368 | interleukin-17 receptor activity(GO:0030368) |
| 0.2 | 0.5 | GO:0070698 | type I activin receptor binding(GO:0070698) |
| 0.2 | 2.2 | GO:0005225 | volume-sensitive anion channel activity(GO:0005225) |
| 0.2 | 24.7 | GO:0004386 | helicase activity(GO:0004386) |
| 0.2 | 1.4 | GO:0035368 | selenocysteine insertion sequence binding(GO:0035368) |
| 0.2 | 0.3 | GO:0008113 | peptide-methionine (S)-S-oxide reductase activity(GO:0008113) |
| 0.2 | 1.9 | GO:0005412 | glucose:sodium symporter activity(GO:0005412) |
| 0.1 | 0.6 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.1 | 1.9 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.1 | 3.1 | GO:0005527 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.1 | 1.2 | GO:0003705 | transcription factor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0003705) |
| 0.1 | 1.1 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.1 | 3.2 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.1 | 3.6 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.1 | 0.9 | GO:0004568 | chitinase activity(GO:0004568) |
| 0.1 | 2.8 | GO:0070300 | phosphatidic acid binding(GO:0070300) |
| 0.1 | 0.4 | GO:0032217 | riboflavin transporter activity(GO:0032217) |
| 0.1 | 0.9 | GO:0005166 | neurotrophin p75 receptor binding(GO:0005166) |
| 0.1 | 1.7 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.1 | 1.0 | GO:0032184 | SUMO polymer binding(GO:0032184) |
| 0.1 | 2.2 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
| 0.1 | 1.0 | GO:0030375 | thyroid hormone receptor coactivator activity(GO:0030375) |
| 0.1 | 33.0 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.1 | 0.5 | GO:0047961 | glycine N-acyltransferase activity(GO:0047961) |
| 0.1 | 2.5 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.1 | 0.9 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.1 | 6.5 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.1 | 1.2 | GO:0016634 | oxidoreductase activity, acting on the CH-CH group of donors, oxygen as acceptor(GO:0016634) |
| 0.1 | 11.2 | GO:0004540 | ribonuclease activity(GO:0004540) |
| 0.1 | 1.0 | GO:0016886 | ligase activity, forming phosphoric ester bonds(GO:0016886) |
| 0.1 | 1.0 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.1 | 16.0 | GO:0070851 | growth factor receptor binding(GO:0070851) |
| 0.1 | 0.3 | GO:0036004 | GAF domain binding(GO:0036004) |
| 0.1 | 0.6 | GO:0005223 | intracellular cGMP activated cation channel activity(GO:0005223) |
| 0.1 | 20.3 | GO:0042393 | histone binding(GO:0042393) |
| 0.1 | 0.4 | GO:0047710 | bis(5'-adenosyl)-triphosphatase activity(GO:0047710) |
| 0.1 | 2.7 | GO:0043295 | glutathione binding(GO:0043295) oligopeptide binding(GO:1900750) |
| 0.1 | 11.5 | GO:0005178 | integrin binding(GO:0005178) |
| 0.1 | 1.5 | GO:0048273 | mitogen-activated protein kinase p38 binding(GO:0048273) |
| 0.1 | 0.9 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.1 | 1.0 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.1 | 5.3 | GO:0000978 | RNA polymerase II core promoter proximal region sequence-specific DNA binding(GO:0000978) |
| 0.1 | 0.2 | GO:0070573 | metallodipeptidase activity(GO:0070573) |
| 0.1 | 0.8 | GO:0000217 | DNA secondary structure binding(GO:0000217) |
| 0.1 | 0.3 | GO:0089720 | caspase binding(GO:0089720) |
| 0.1 | 2.4 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.1 | 3.4 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.1 | 0.4 | GO:0070095 | fructose-6-phosphate binding(GO:0070095) |
| 0.1 | 1.5 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.1 | 1.8 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.1 | 1.7 | GO:0000980 | RNA polymerase II distal enhancer sequence-specific DNA binding(GO:0000980) |
| 0.1 | 0.3 | GO:0004095 | carnitine O-palmitoyltransferase activity(GO:0004095) O-palmitoyltransferase activity(GO:0016416) |
| 0.1 | 0.6 | GO:0015093 | ferrous iron transmembrane transporter activity(GO:0015093) |
| 0.1 | 3.5 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) |
| 0.1 | 0.3 | GO:0070736 | protein-glycine ligase activity, initiating(GO:0070736) |
| 0.1 | 0.9 | GO:0008392 | arachidonic acid epoxygenase activity(GO:0008392) |
| 0.1 | 2.4 | GO:0018024 | histone-lysine N-methyltransferase activity(GO:0018024) |
| 0.1 | 2.9 | GO:0046332 | SMAD binding(GO:0046332) |
| 0.1 | 1.6 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.1 | 0.6 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.1 | 0.3 | GO:0008176 | tRNA (guanine-N7-)-methyltransferase activity(GO:0008176) |
| 0.1 | 35.7 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.1 | 15.3 | GO:0003712 | transcription cofactor activity(GO:0003712) |
| 0.1 | 0.1 | GO:0016635 | oxidoreductase activity, acting on the CH-CH group of donors, quinone or related compound as acceptor(GO:0016635) |
| 0.1 | 1.1 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.1 | 2.7 | GO:0043531 | ADP binding(GO:0043531) |
| 0.1 | 1.5 | GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.1 | 0.3 | GO:0008545 | JUN kinase kinase activity(GO:0008545) |
| 0.1 | 2.3 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.1 | 0.7 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.1 | 0.2 | GO:0033814 | propanoyl-CoA C-acyltransferase activity(GO:0033814) propionyl-CoA C2-trimethyltridecanoyltransferase activity(GO:0050632) phosphatidylethanolamine transporter activity(GO:1904121) |
| 0.1 | 1.7 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 0.1 | 0.4 | GO:0004032 | alditol:NADP+ 1-oxidoreductase activity(GO:0004032) |
| 0.1 | 3.1 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.1 | 1.1 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) |
| 0.1 | 0.7 | GO:0016846 | carbon-sulfur lyase activity(GO:0016846) |
| 0.1 | 1.5 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 3.2 | GO:0016811 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amides(GO:0016811) |
| 0.0 | 1.6 | GO:0070491 | repressing transcription factor binding(GO:0070491) |
| 0.0 | 0.4 | GO:0008481 | sphinganine kinase activity(GO:0008481) D-erythro-sphingosine kinase activity(GO:0017050) |
| 0.0 | 1.0 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.0 | 4.9 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 65.9 | GO:0003677 | DNA binding(GO:0003677) |
| 0.0 | 0.5 | GO:0005186 | pheromone activity(GO:0005186) |
| 0.0 | 1.2 | GO:0016706 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, 2-oxoglutarate as one donor, and incorporation of one atom each of oxygen into both donors(GO:0016706) |
| 0.0 | 0.7 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
| 0.0 | 5.7 | GO:0020037 | heme binding(GO:0020037) |
| 0.0 | 0.2 | GO:0019966 | interleukin-1 binding(GO:0019966) |
| 0.0 | 0.1 | GO:1990254 | keratin filament binding(GO:1990254) |
| 0.0 | 1.1 | GO:0008519 | ammonium transmembrane transporter activity(GO:0008519) |
| 0.0 | 7.5 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.0 | 0.2 | GO:0004945 | angiotensin type II receptor activity(GO:0004945) |
| 0.0 | 1.5 | GO:0008080 | N-acetyltransferase activity(GO:0008080) |
| 0.0 | 3.7 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.0 | 5.2 | GO:0061135 | endopeptidase regulator activity(GO:0061135) |
| 0.0 | 0.3 | GO:0016278 | lysine N-methyltransferase activity(GO:0016278) protein-lysine N-methyltransferase activity(GO:0016279) |
| 0.0 | 0.3 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.0 | 0.9 | GO:0017022 | myosin binding(GO:0017022) |
| 0.0 | 0.2 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.0 | 0.3 | GO:0019789 | SUMO transferase activity(GO:0019789) |
| 0.0 | 0.4 | GO:0015149 | glucose transmembrane transporter activity(GO:0005355) hexose transmembrane transporter activity(GO:0015149) |
| 0.0 | 0.3 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.0 | 0.8 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.0 | 0.7 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.0 | 0.5 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.0 | 0.2 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.0 | 0.3 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.0 | 0.3 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 0.7 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.0 | GO:0016833 | oxo-acid-lyase activity(GO:0016833) |
| 0.0 | 0.0 | GO:0045131 | pre-mRNA branch point binding(GO:0045131) |
| 0.0 | 0.3 | GO:0001099 | basal transcription machinery binding(GO:0001098) basal RNA polymerase II transcription machinery binding(GO:0001099) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.9 | 42.8 | ST TYPE I INTERFERON PATHWAY | Type I Interferon (alpha/beta IFN) Pathway |
| 2.0 | 31.7 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 1.4 | 127.7 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 1.3 | 20.2 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 1.1 | 52.3 | PID TRAIL PATHWAY | TRAIL signaling pathway |
| 1.1 | 27.6 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 1.0 | 37.6 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.9 | 18.9 | ST TUMOR NECROSIS FACTOR PATHWAY | Tumor Necrosis Factor Pathway. |
| 0.7 | 28.4 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.6 | 17.3 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.6 | 22.3 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 0.5 | 10.9 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.5 | 6.4 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 0.5 | 3.7 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.5 | 20.7 | PID MYC PATHWAY | C-MYC pathway |
| 0.5 | 31.0 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.5 | 5.3 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.4 | 91.4 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.4 | 2.9 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.4 | 28.9 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.4 | 55.2 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.4 | 31.6 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.4 | 66.5 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.4 | 7.2 | SIG CD40PATHWAYMAP | Genes related to CD40 signaling |
| 0.3 | 8.7 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.3 | 8.8 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.3 | 10.5 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.3 | 14.6 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.3 | 7.1 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.3 | 17.6 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.3 | 6.8 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.3 | 9.3 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.3 | 6.1 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.2 | 6.2 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.2 | 4.7 | PID ATM PATHWAY | ATM pathway |
| 0.2 | 10.0 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.2 | 2.3 | PID IL3 PATHWAY | IL3-mediated signaling events |
| 0.2 | 8.1 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.2 | 8.3 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.2 | 3.9 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.2 | 4.0 | ST PHOSPHOINOSITIDE 3 KINASE PATHWAY | PI3K Pathway |
| 0.2 | 4.4 | SIG PIP3 SIGNALING IN CARDIAC MYOCTES | Genes related to PIP3 signaling in cardiac myocytes |
| 0.1 | 6.7 | PID E2F PATHWAY | E2F transcription factor network |
| 0.1 | 4.4 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.1 | 1.8 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.1 | 6.1 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.1 | 5.9 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.1 | 2.7 | PID THROMBIN PAR1 PATHWAY | PAR1-mediated thrombin signaling events |
| 0.1 | 33.5 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.1 | 1.9 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.1 | 3.5 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.1 | 0.9 | ST GA13 PATHWAY | G alpha 13 Pathway |
| 0.1 | 3.6 | PID TCR PATHWAY | TCR signaling in naïve CD4+ T cells |
| 0.1 | 2.9 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.1 | 1.0 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.1 | 1.7 | PID VEGFR1 2 PATHWAY | Signaling events mediated by VEGFR1 and VEGFR2 |
| 0.1 | 2.0 | ST FAS SIGNALING PATHWAY | Fas Signaling Pathway |
| 0.1 | 0.6 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.1 | 3.2 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 1.2 | PID INTEGRIN1 PATHWAY | Beta1 integrin cell surface interactions |
| 0.1 | 0.7 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.0 | 1.9 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.0 | 0.9 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.0 | 2.7 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 1.7 | PID P53 REGULATION PATHWAY | p53 pathway |
| 0.0 | 1.6 | PID ARF6 TRAFFICKING PATHWAY | Arf6 trafficking events |
| 0.0 | 1.5 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 0.8 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.0 | 1.4 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 8.0 | NABA MATRISOME ASSOCIATED | Ensemble of genes encoding ECM-associated proteins including ECM-affilaited proteins, ECM regulators and secreted factors |
| 0.0 | 0.7 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.0 | 0.4 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 15.1 | 45.3 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 6.1 | 448.6 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 5.7 | 68.4 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 4.7 | 80.5 | REACTOME NFKB ACTIVATION THROUGH FADD RIP1 PATHWAY MEDIATED BY CASPASE 8 AND10 | Genes involved in NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 |
| 3.7 | 151.1 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 3.4 | 63.8 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 3.2 | 44.4 | REACTOME PD1 SIGNALING | Genes involved in PD-1 signaling |
| 2.7 | 10.9 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 2.7 | 37.6 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 2.7 | 53.4 | REACTOME RIP MEDIATED NFKB ACTIVATION VIA DAI | Genes involved in RIP-mediated NFkB activation via DAI |
| 2.4 | 33.5 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 1.6 | 22.9 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 1.5 | 127.7 | REACTOME ANTIGEN PROCESSING CROSS PRESENTATION | Genes involved in Antigen processing-Cross presentation |
| 1.5 | 82.1 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 1.5 | 63.8 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 1.2 | 13.1 | REACTOME RIG I MDA5 MEDIATED INDUCTION OF IFN ALPHA BETA PATHWAYS | Genes involved in RIG-I/MDA5 mediated induction of IFN-alpha/beta pathways |
| 1.1 | 26.4 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 1.0 | 17.4 | REACTOME POL SWITCHING | Genes involved in Polymerase switching |
| 0.9 | 46.9 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.9 | 60.6 | REACTOME COSTIMULATION BY THE CD28 FAMILY | Genes involved in Costimulation by the CD28 family |
| 0.7 | 5.2 | REACTOME P53 INDEPENDENT G1 S DNA DAMAGE CHECKPOINT | Genes involved in p53-Independent G1/S DNA damage checkpoint |
| 0.7 | 8.7 | REACTOME TAK1 ACTIVATES NFKB BY PHOSPHORYLATION AND ACTIVATION OF IKKS COMPLEX | Genes involved in TAK1 activates NFkB by phosphorylation and activation of IKKs complex |
| 0.7 | 22.6 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.6 | 4.0 | REACTOME INFLAMMASOMES | Genes involved in Inflammasomes |
| 0.6 | 8.3 | REACTOME EICOSANOID LIGAND BINDING RECEPTORS | Genes involved in Eicosanoid ligand-binding receptors |
| 0.6 | 10.5 | REACTOME DESTABILIZATION OF MRNA BY BRF1 | Genes involved in Destabilization of mRNA by Butyrate Response Factor 1 (BRF1) |
| 0.5 | 9.1 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.5 | 4.2 | REACTOME DEFENSINS | Genes involved in Defensins |
| 0.5 | 6.6 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.5 | 4.0 | REACTOME MRNA PROCESSING | Genes involved in mRNA Processing |
| 0.5 | 1.4 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.5 | 3.2 | REACTOME P75NTR RECRUITS SIGNALLING COMPLEXES | Genes involved in p75NTR recruits signalling complexes |
| 0.4 | 3.4 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.4 | 10.9 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.4 | 5.8 | REACTOME EXTENSION OF TELOMERES | Genes involved in Extension of Telomeres |
| 0.4 | 5.7 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.4 | 7.0 | REACTOME NOD1 2 SIGNALING PATHWAY | Genes involved in NOD1/2 Signaling Pathway |
| 0.4 | 12.4 | REACTOME PRE NOTCH TRANSCRIPTION AND TRANSLATION | Genes involved in Pre-NOTCH Transcription and Translation |
| 0.3 | 4.6 | REACTOME GRB2 SOS PROVIDES LINKAGE TO MAPK SIGNALING FOR INTERGRINS | Genes involved in GRB2:SOS provides linkage to MAPK signaling for Intergrins |
| 0.3 | 7.3 | REACTOME INHIBITION OF THE PROTEOLYTIC ACTIVITY OF APC C REQUIRED FOR THE ONSET OF ANAPHASE BY MITOTIC SPINDLE CHECKPOINT COMPONENTS | Genes involved in Inhibition of the proteolytic activity of APC/C required for the onset of anaphase by mitotic spindle checkpoint components |
| 0.3 | 4.1 | REACTOME ACTIVATED AMPK STIMULATES FATTY ACID OXIDATION IN MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.3 | 3.2 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.3 | 8.7 | REACTOME IL RECEPTOR SHC SIGNALING | Genes involved in Interleukin receptor SHC signaling |
| 0.3 | 7.7 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.3 | 14.1 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.3 | 3.9 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.3 | 7.6 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.3 | 2.9 | REACTOME DOWNSTREAM SIGNALING EVENTS OF B CELL RECEPTOR BCR | Genes involved in Downstream Signaling Events Of B Cell Receptor (BCR) |
| 0.3 | 3.6 | REACTOME DESTABILIZATION OF MRNA BY AUF1 HNRNP D0 | Genes involved in Destabilization of mRNA by AUF1 (hnRNP D0) |
| 0.3 | 5.3 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.2 | 5.3 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.2 | 21.1 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.2 | 13.1 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.2 | 2.7 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 0.2 | 2.4 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.2 | 4.4 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.2 | 6.2 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.2 | 2.1 | REACTOME HORMONE LIGAND BINDING RECEPTORS | Genes involved in Hormone ligand-binding receptors |
| 0.2 | 4.8 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.2 | 3.2 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.2 | 1.7 | REACTOME ACTIVATION OF THE MRNA UPON BINDING OF THE CAP BINDING COMPLEX AND EIFS AND SUBSEQUENT BINDING TO 43S | Genes involved in Activation of the mRNA upon binding of the cap-binding complex and eIFs, and subsequent binding to 43S |
| 0.2 | 5.7 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.2 | 5.9 | REACTOME CYCLIN E ASSOCIATED EVENTS DURING G1 S TRANSITION | Genes involved in Cyclin E associated events during G1/S transition |
| 0.2 | 7.8 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.2 | 2.9 | REACTOME HYALURONAN METABOLISM | Genes involved in Hyaluronan metabolism |
| 0.2 | 1.9 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.2 | 9.9 | REACTOME INTERFERON SIGNALING | Genes involved in Interferon Signaling |
| 0.2 | 3.9 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.2 | 2.6 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.2 | 5.4 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.2 | 32.7 | REACTOME CLASS I MHC MEDIATED ANTIGEN PROCESSING PRESENTATION | Genes involved in Class I MHC mediated antigen processing & presentation |
| 0.1 | 10.6 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.1 | 3.1 | REACTOME ACYL CHAIN REMODELLING OF PG | Genes involved in Acyl chain remodelling of PG |
| 0.1 | 2.5 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.1 | 6.8 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.1 | 2.8 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.1 | 1.6 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.1 | 2.2 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.1 | 4.4 | REACTOME TRANSPORT OF MATURE TRANSCRIPT TO CYTOPLASM | Genes involved in Transport of Mature Transcript to Cytoplasm |
| 0.1 | 2.8 | REACTOME THE ROLE OF NEF IN HIV1 REPLICATION AND DISEASE PATHOGENESIS | Genes involved in The role of Nef in HIV-1 replication and disease pathogenesis |
| 0.1 | 6.6 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.1 | 2.8 | REACTOME SIGNALING BY EGFR IN CANCER | Genes involved in Signaling by EGFR in Cancer |
| 0.1 | 2.4 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.1 | 1.0 | REACTOME RAF MAP KINASE CASCADE | Genes involved in RAF/MAP kinase cascade |
| 0.1 | 1.9 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.1 | 1.5 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.1 | 1.8 | REACTOME SEMA3A PAK DEPENDENT AXON REPULSION | Genes involved in Sema3A PAK dependent Axon repulsion |
| 0.1 | 3.0 | REACTOME SEMA4D IN SEMAPHORIN SIGNALING | Genes involved in Sema4D in semaphorin signaling |
| 0.1 | 1.4 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.1 | 3.5 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.1 | 6.3 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.1 | 2.5 | REACTOME THROMBOXANE SIGNALLING THROUGH TP RECEPTOR | Genes involved in Thromboxane signalling through TP receptor |
| 0.1 | 3.9 | REACTOME RECRUITMENT OF MITOTIC CENTROSOME PROTEINS AND COMPLEXES | Genes involved in Recruitment of mitotic centrosome proteins and complexes |
| 0.1 | 5.4 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.1 | 0.8 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.1 | 0.8 | REACTOME NRIF SIGNALS CELL DEATH FROM THE NUCLEUS | Genes involved in NRIF signals cell death from the nucleus |
| 0.1 | 1.4 | REACTOME ACTIVATION OF ATR IN RESPONSE TO REPLICATION STRESS | Genes involved in Activation of ATR in response to replication stress |
| 0.1 | 1.8 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.1 | 1.4 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.0 | 5.0 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.0 | 1.4 | REACTOME RECYCLING PATHWAY OF L1 | Genes involved in Recycling pathway of L1 |
| 0.0 | 0.8 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.0 | 0.5 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.0 | 1.1 | REACTOME INFLUENZA VIRAL RNA TRANSCRIPTION AND REPLICATION | Genes involved in Influenza Viral RNA Transcription and Replication |
| 0.0 | 0.9 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.0 | 0.6 | REACTOME REGULATION OF BETA CELL DEVELOPMENT | Genes involved in Regulation of beta-cell development |
| 0.0 | 4.1 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.0 | 0.6 | REACTOME INTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Intrinsic Pathway for Apoptosis |
| 0.0 | 0.4 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.0 | 2.7 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.0 | 0.7 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.0 | 0.5 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.0 | 1.7 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.0 | 0.1 | REACTOME REGULATION OF THE FANCONI ANEMIA PATHWAY | Genes involved in Regulation of the Fanconi anemia pathway |
| 0.0 | 0.5 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |
| 0.0 | 0.3 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.0 | 1.2 | REACTOME SIGNALING BY PDGF | Genes involved in Signaling by PDGF |
| 0.0 | 0.5 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |