Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
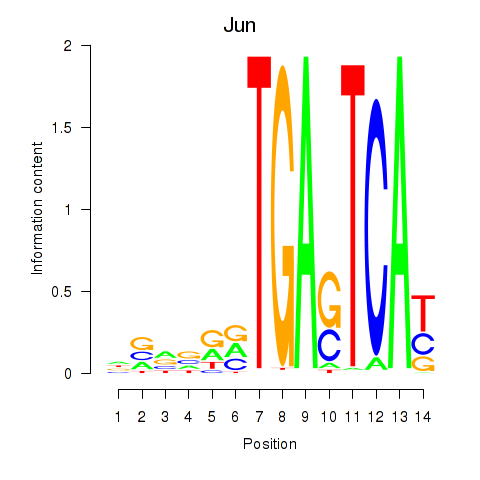
Results for Jun
Z-value: 1.30
Transcription factors associated with Jun
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Jun
|
ENSMUSG00000052684.5 | Jun |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Jun | mm39_v1_chr4_-_94940425_94940459 | 0.51 | 5.2e-06 | Click! |
Activity profile of Jun motif
Sorted Z-values of Jun motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Jun
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_-_92393193 | 13.51 |
ENSMUST00000054599.8
|
Sprr1a
|
small proline-rich protein 1A |
| chr10_+_18720760 | 12.24 |
ENSMUST00000019998.9
|
Perp
|
PERP, TP53 apoptosis effector |
| chr6_-_87327885 | 11.05 |
ENSMUST00000032129.3
|
Gkn1
|
gastrokine 1 |
| chr7_+_30487322 | 11.04 |
ENSMUST00000189673.7
ENSMUST00000190990.7 ENSMUST00000189962.7 ENSMUST00000187493.7 ENSMUST00000098559.3 |
Krtdap
|
keratinocyte differentiation associated protein |
| chr15_-_74618533 | 10.94 |
ENSMUST00000057932.8
|
Slurp2
|
secreted Ly6/Plaur domain containing 2 |
| chr5_+_36361360 | 10.89 |
ENSMUST00000052224.6
|
Psapl1
|
prosaposin-like 1 |
| chr4_+_128999325 | 10.61 |
ENSMUST00000106064.10
ENSMUST00000030575.15 ENSMUST00000030577.11 |
Tmem54
|
transmembrane protein 54 |
| chr7_+_140918793 | 9.46 |
ENSMUST00000026577.13
|
Eps8l2
|
EPS8-like 2 |
| chr19_+_8966641 | 9.33 |
ENSMUST00000092956.4
ENSMUST00000092955.11 |
Ahnak
|
AHNAK nucleoprotein (desmoyokin) |
| chr17_-_31383976 | 8.81 |
ENSMUST00000235870.2
|
Tff1
|
trefoil factor 1 |
| chr11_+_76795346 | 8.77 |
ENSMUST00000072633.4
|
Tmigd1
|
transmembrane and immunoglobulin domain containing 1 |
| chr7_+_30463175 | 8.65 |
ENSMUST00000165887.8
ENSMUST00000085691.11 ENSMUST00000054427.13 ENSMUST00000085688.11 |
Dmkn
|
dermokine |
| chr7_+_18817767 | 8.56 |
ENSMUST00000032568.14
ENSMUST00000122999.8 ENSMUST00000108473.10 ENSMUST00000108474.2 ENSMUST00000238982.2 |
Dmpk
|
dystrophia myotonica-protein kinase |
| chr18_+_12637217 | 8.46 |
ENSMUST00000188815.2
|
Lama3
|
laminin, alpha 3 |
| chr4_+_43506966 | 8.45 |
ENSMUST00000030183.10
|
Car9
|
carbonic anhydrase 9 |
| chr15_-_74599860 | 8.37 |
ENSMUST00000023261.4
ENSMUST00000190433.2 |
Slurp1
|
secreted Ly6/Plaur domain containing 1 |
| chr5_-_134975773 | 8.16 |
ENSMUST00000051401.4
|
Cldn4
|
claudin 4 |
| chr7_+_140918876 | 7.99 |
ENSMUST00000143633.4
|
Eps8l2
|
EPS8-like 2 |
| chr4_+_118384426 | 7.77 |
ENSMUST00000030261.6
|
2610528J11Rik
|
RIKEN cDNA 2610528J11 gene |
| chr2_+_151414524 | 7.69 |
ENSMUST00000028950.9
|
Sdcbp2
|
syndecan binding protein (syntenin) 2 |
| chr2_-_25129863 | 7.61 |
ENSMUST00000186719.2
ENSMUST00000043379.5 |
Cysrt1
|
cysteine rich tail 1 |
| chr17_-_25973288 | 7.58 |
ENSMUST00000075884.8
ENSMUST00000238120.2 ENSMUST00000236137.2 ENSMUST00000237359.2 |
Msln
|
mesothelin |
| chr11_-_100139728 | 7.56 |
ENSMUST00000007280.9
|
Krt16
|
keratin 16 |
| chr7_+_30475819 | 7.53 |
ENSMUST00000041703.10
|
Dmkn
|
dermokine |
| chr4_-_133329479 | 7.52 |
ENSMUST00000057311.4
|
Sfn
|
stratifin |
| chr7_+_28563255 | 7.50 |
ENSMUST00000138272.8
|
Lgals7
|
lectin, galactose binding, soluble 7 |
| chr7_+_140659038 | 7.42 |
ENSMUST00000159375.8
|
Pkp3
|
plakophilin 3 |
| chr14_+_102078038 | 7.32 |
ENSMUST00000159314.8
|
Lmo7
|
LIM domain only 7 |
| chr19_-_32080496 | 7.21 |
ENSMUST00000235213.2
ENSMUST00000236504.2 |
Asah2
|
N-acylsphingosine amidohydrolase 2 |
| chr2_+_69210775 | 7.21 |
ENSMUST00000063690.4
|
Dhrs9
|
dehydrogenase/reductase (SDR family) member 9 |
| chr11_+_9068098 | 7.12 |
ENSMUST00000020677.8
ENSMUST00000101525.9 |
Upp1
|
uridine phosphorylase 1 |
| chr7_-_44320244 | 7.11 |
ENSMUST00000048102.15
|
Myh14
|
myosin, heavy polypeptide 14 |
| chr17_+_87944110 | 6.73 |
ENSMUST00000234623.2
|
Epcam
|
epithelial cell adhesion molecule |
| chr3_+_93427791 | 6.72 |
ENSMUST00000029515.5
|
S100a11
|
S100 calcium binding protein A11 |
| chr16_-_56652241 | 6.72 |
ENSMUST00000135672.2
|
Tmem45a
|
transmembrane protein 45a |
| chr5_-_76511634 | 6.71 |
ENSMUST00000031146.3
|
Nmu
|
neuromedin U |
| chr11_+_76795292 | 6.71 |
ENSMUST00000142166.8
|
Tmigd1
|
transmembrane and immunoglobulin domain containing 1 |
| chr8_-_110765983 | 6.68 |
ENSMUST00000109222.4
|
Chst4
|
carbohydrate sulfotransferase 4 |
| chr3_+_137923521 | 6.66 |
ENSMUST00000090171.7
|
Adh7
|
alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide |
| chr1_+_172328768 | 6.34 |
ENSMUST00000111228.2
|
Tagln2
|
transgelin 2 |
| chr6_+_49013517 | 6.17 |
ENSMUST00000031840.10
|
Gpnmb
|
glycoprotein (transmembrane) nmb |
| chr1_+_182392577 | 6.06 |
ENSMUST00000048941.14
|
Capn8
|
calpain 8 |
| chr2_+_155593030 | 6.02 |
ENSMUST00000029140.12
ENSMUST00000132608.2 |
Procr
|
protein C receptor, endothelial |
| chr4_+_118384183 | 6.00 |
ENSMUST00000106367.8
|
2610528J11Rik
|
RIKEN cDNA 2610528J11 gene |
| chr6_+_49013601 | 5.96 |
ENSMUST00000204260.2
|
Gpnmb
|
glycoprotein (transmembrane) nmb |
| chr2_+_143757193 | 5.91 |
ENSMUST00000103172.4
|
Dstn
|
destrin |
| chr11_+_9068134 | 5.90 |
ENSMUST00000170444.8
|
Upp1
|
uridine phosphorylase 1 |
| chr12_+_75355082 | 5.65 |
ENSMUST00000118602.8
ENSMUST00000118966.8 ENSMUST00000055390.6 |
Rhoj
|
ras homolog family member J |
| chr6_-_113411718 | 5.57 |
ENSMUST00000113091.8
|
Cidec
|
cell death-inducing DFFA-like effector c |
| chr3_+_92315290 | 5.50 |
ENSMUST00000047264.3
|
Sprr2i
|
small proline-rich protein 2I |
| chr1_+_182392559 | 5.49 |
ENSMUST00000168514.7
|
Capn8
|
calpain 8 |
| chr12_-_103423472 | 5.45 |
ENSMUST00000044687.7
|
Ifi27l2b
|
interferon, alpha-inducible protein 27 like 2B |
| chr3_+_14706781 | 5.41 |
ENSMUST00000029071.9
|
Car13
|
carbonic anhydrase 13 |
| chr8_-_62044164 | 5.34 |
ENSMUST00000135439.2
ENSMUST00000121200.9 |
Palld
|
palladin, cytoskeletal associated protein |
| chr14_+_102077937 | 5.32 |
ENSMUST00000159026.8
|
Lmo7
|
LIM domain only 7 |
| chrX_+_72719098 | 5.26 |
ENSMUST00000171398.2
|
Slc6a8
|
solute carrier family 6 (neurotransmitter transporter, creatine), member 8 |
| chr7_-_28947882 | 5.23 |
ENSMUST00000032808.6
|
2200002D01Rik
|
RIKEN cDNA 2200002D01 gene |
| chr7_+_24335969 | 5.21 |
ENSMUST00000080718.6
|
Lypd3
|
Ly6/Plaur domain containing 3 |
| chr7_-_105131407 | 5.17 |
ENSMUST00000047040.4
|
Cavin3
|
caveolae associated 3 |
| chr10_-_76562002 | 5.13 |
ENSMUST00000001147.5
|
Col6a1
|
collagen, type VI, alpha 1 |
| chr3_-_88410495 | 5.02 |
ENSMUST00000120377.8
ENSMUST00000029699.13 |
Lmna
|
lamin A |
| chr12_+_31315270 | 4.98 |
ENSMUST00000002979.16
ENSMUST00000239496.2 ENSMUST00000170495.3 |
Lamb1
|
laminin B1 |
| chr3_-_95158457 | 4.93 |
ENSMUST00000125515.3
ENSMUST00000107195.9 |
Bnipl
|
BCL2/adenovirus E1B 19kD interacting protein like |
| chrX_-_7537580 | 4.90 |
ENSMUST00000033486.6
|
Plp2
|
proteolipid protein 2 |
| chr19_-_11796282 | 4.90 |
ENSMUST00000069285.6
|
Stx3
|
syntaxin 3 |
| chr3_-_95646856 | 4.89 |
ENSMUST00000153026.8
ENSMUST00000123143.8 ENSMUST00000137912.8 ENSMUST00000029753.14 ENSMUST00000131376.8 ENSMUST00000117507.10 ENSMUST00000128885.8 ENSMUST00000147217.2 |
Ecm1
|
extracellular matrix protein 1 |
| chr3_-_59102517 | 4.89 |
ENSMUST00000200095.2
|
Gpr87
|
G protein-coupled receptor 87 |
| chr10_-_43880353 | 4.88 |
ENSMUST00000020017.14
|
Crybg1
|
crystallin beta-gamma domain containing 1 |
| chr13_+_75987987 | 4.82 |
ENSMUST00000022082.8
ENSMUST00000223120.2 ENSMUST00000220523.2 |
Glrx
|
glutaredoxin |
| chr11_-_69771797 | 4.77 |
ENSMUST00000238978.2
|
Kctd11
|
potassium channel tetramerisation domain containing 11 |
| chr7_+_127728712 | 4.65 |
ENSMUST00000033053.8
ENSMUST00000205460.2 |
Itgax
|
integrin alpha X |
| chr5_+_32293145 | 4.65 |
ENSMUST00000031017.11
|
Fosl2
|
fos-like antigen 2 |
| chr8_-_110766009 | 4.65 |
ENSMUST00000212934.2
|
Chst4
|
carbohydrate sulfotransferase 4 |
| chr3_-_95158360 | 4.65 |
ENSMUST00000098871.11
ENSMUST00000137250.9 |
Bnipl
|
BCL2/adenovirus E1B 19kD interacting protein like |
| chr19_+_8975249 | 4.61 |
ENSMUST00000236390.2
|
Ahnak
|
AHNAK nucleoprotein (desmoyokin) |
| chr7_+_30450896 | 4.60 |
ENSMUST00000182229.8
ENSMUST00000080518.14 ENSMUST00000182227.8 ENSMUST00000182721.8 |
Sbsn
|
suprabasin |
| chr6_+_30568366 | 4.60 |
ENSMUST00000049251.6
|
Cpa4
|
carboxypeptidase A4 |
| chr7_+_141988714 | 4.52 |
ENSMUST00000118276.8
ENSMUST00000105976.8 ENSMUST00000097939.9 |
Syt8
|
synaptotagmin VIII |
| chr3_-_92481033 | 4.50 |
ENSMUST00000053107.6
|
Ivl
|
involucrin |
| chr7_-_4607040 | 4.44 |
ENSMUST00000166650.3
|
Ptprh
|
protein tyrosine phosphatase, receptor type, H |
| chr12_+_31315227 | 4.43 |
ENSMUST00000169088.8
|
Lamb1
|
laminin B1 |
| chr7_+_141048722 | 4.43 |
ENSMUST00000058746.7
|
Cd151
|
CD151 antigen |
| chr8_-_62576140 | 4.34 |
ENSMUST00000034052.14
ENSMUST00000034054.9 |
Anxa10
|
annexin A10 |
| chr2_+_24226857 | 4.32 |
ENSMUST00000114487.9
ENSMUST00000142093.7 |
Il1rn
|
interleukin 1 receptor antagonist |
| chr5_+_93004330 | 4.31 |
ENSMUST00000225438.2
|
Shroom3
|
shroom family member 3 |
| chr17_-_24863956 | 4.25 |
ENSMUST00000019684.13
|
Slc9a3r2
|
solute carrier family 9 (sodium/hydrogen exchanger), member 3 regulator 2 |
| chr7_+_18573879 | 4.22 |
ENSMUST00000072415.9
ENSMUST00000072386.11 ENSMUST00000228493.2 ENSMUST00000227379.2 |
Mill2
|
MHC I like leukocyte 2 |
| chr9_+_44953723 | 4.18 |
ENSMUST00000034600.5
|
Mpzl2
|
myelin protein zero-like 2 |
| chr7_-_140856642 | 4.14 |
ENSMUST00000080654.7
ENSMUST00000167263.9 |
Cdhr5
|
cadherin-related family member 5 |
| chr2_+_103242027 | 4.08 |
ENSMUST00000239273.2
ENSMUST00000164172.8 |
Elf5
|
E74-like factor 5 |
| chr3_+_142326363 | 4.06 |
ENSMUST00000165774.8
|
Gbp2
|
guanylate binding protein 2 |
| chr17_+_35268942 | 4.02 |
ENSMUST00000007257.10
|
Clic1
|
chloride intracellular channel 1 |
| chr11_+_61575245 | 3.99 |
ENSMUST00000093019.6
|
Fam83g
|
family with sequence similarity 83, member G |
| chr14_-_70415117 | 3.98 |
ENSMUST00000022681.11
|
Pdlim2
|
PDZ and LIM domain 2 |
| chr7_+_126365506 | 3.97 |
ENSMUST00000032944.9
|
Gdpd3
|
glycerophosphodiester phosphodiesterase domain containing 3 |
| chr18_-_38131766 | 3.91 |
ENSMUST00000236588.2
ENSMUST00000237272.2 ENSMUST00000236134.2 |
Arap3
|
ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 3 |
| chr14_+_14210394 | 3.76 |
ENSMUST00000164598.8
|
Acox2
|
acyl-Coenzyme A oxidase 2, branched chain |
| chr14_-_34310602 | 3.69 |
ENSMUST00000064098.14
ENSMUST00000090040.12 ENSMUST00000022330.9 ENSMUST00000022327.13 |
Ldb3
|
LIM domain binding 3 |
| chr1_+_107517726 | 3.68 |
ENSMUST00000000514.11
ENSMUST00000112706.4 |
Serpinb8
|
serine (or cysteine) peptidase inhibitor, clade B, member 8 |
| chr13_-_95754987 | 3.66 |
ENSMUST00000059193.7
|
F2r
|
coagulation factor II (thrombin) receptor |
| chr16_+_17379749 | 3.62 |
ENSMUST00000171002.10
ENSMUST00000023441.11 |
P2rx6
|
purinergic receptor P2X, ligand-gated ion channel, 6 |
| chr5_-_66238313 | 3.59 |
ENSMUST00000202700.4
ENSMUST00000094757.9 ENSMUST00000113724.6 |
Rbm47
|
RNA binding motif protein 47 |
| chr7_+_30252687 | 3.57 |
ENSMUST00000044048.8
|
Hspb6
|
heat shock protein, alpha-crystallin-related, B6 |
| chr14_+_14210932 | 3.51 |
ENSMUST00000022271.14
|
Acox2
|
acyl-Coenzyme A oxidase 2, branched chain |
| chr6_+_17463925 | 3.50 |
ENSMUST00000115442.8
|
Met
|
met proto-oncogene |
| chr7_-_80051455 | 3.42 |
ENSMUST00000120753.3
|
Furin
|
furin (paired basic amino acid cleaving enzyme) |
| chr7_-_144305705 | 3.40 |
ENSMUST00000155175.8
|
Ano1
|
anoctamin 1, calcium activated chloride channel |
| chr9_-_116004386 | 3.37 |
ENSMUST00000035014.8
|
Tgfbr2
|
transforming growth factor, beta receptor II |
| chr1_+_171265103 | 3.35 |
ENSMUST00000043839.5
|
F11r
|
F11 receptor |
| chr9_+_7445822 | 3.35 |
ENSMUST00000034497.8
|
Mmp3
|
matrix metallopeptidase 3 |
| chr2_+_110551927 | 3.33 |
ENSMUST00000111017.9
|
Muc15
|
mucin 15 |
| chr18_+_42186713 | 3.32 |
ENSMUST00000072008.11
|
Sh3rf2
|
SH3 domain containing ring finger 2 |
| chr7_-_143056252 | 3.30 |
ENSMUST00000010904.5
|
Phlda2
|
pleckstrin homology like domain, family A, member 2 |
| chr17_-_24863907 | 3.29 |
ENSMUST00000234505.2
|
Slc9a3r2
|
solute carrier family 9 (sodium/hydrogen exchanger), member 3 regulator 2 |
| chr8_+_72889073 | 3.23 |
ENSMUST00000003575.11
|
Tpm4
|
tropomyosin 4 |
| chr3_-_88243455 | 3.23 |
ENSMUST00000193872.2
|
Tmem79
|
transmembrane protein 79 |
| chr1_-_190915441 | 3.18 |
ENSMUST00000027941.14
|
Atf3
|
activating transcription factor 3 |
| chr10_+_12966532 | 3.16 |
ENSMUST00000121646.8
ENSMUST00000121325.8 ENSMUST00000121766.8 |
Plagl1
|
pleiomorphic adenoma gene-like 1 |
| chr16_+_33614715 | 3.15 |
ENSMUST00000023520.7
|
Muc13
|
mucin 13, epithelial transmembrane |
| chr9_-_116004265 | 3.14 |
ENSMUST00000061101.12
|
Tgfbr2
|
transforming growth factor, beta receptor II |
| chr17_+_44263890 | 3.09 |
ENSMUST00000177857.9
ENSMUST00000044792.6 |
Rcan2
|
regulator of calcineurin 2 |
| chr8_-_25581354 | 3.08 |
ENSMUST00000125466.2
|
Plekha2
|
pleckstrin homology domain-containing, family A (phosphoinositide binding specific) member 2 |
| chr3_+_27371168 | 3.07 |
ENSMUST00000046383.12
|
Tnfsf10
|
tumor necrosis factor (ligand) superfamily, member 10 |
| chr2_+_174602412 | 3.02 |
ENSMUST00000029030.9
|
Edn3
|
endothelin 3 |
| chr7_-_45391879 | 2.99 |
ENSMUST00000210754.2
ENSMUST00000210147.2 |
Sult2b1
|
sulfotransferase family, cytosolic, 2B, member 1 |
| chr6_+_17463748 | 2.96 |
ENSMUST00000115443.8
|
Met
|
met proto-oncogene |
| chr18_+_42186757 | 2.95 |
ENSMUST00000074679.4
|
Sh3rf2
|
SH3 domain containing ring finger 2 |
| chr2_+_70948267 | 2.95 |
ENSMUST00000028403.3
|
Cybrd1
|
cytochrome b reductase 1 |
| chr2_+_110551685 | 2.94 |
ENSMUST00000111016.9
|
Muc15
|
mucin 15 |
| chr19_-_4092218 | 2.93 |
ENSMUST00000237999.2
ENSMUST00000042700.12 |
Gstp2
|
glutathione S-transferase, pi 2 |
| chrX_-_7606445 | 2.90 |
ENSMUST00000128289.8
|
Ccdc120
|
coiled-coil domain containing 120 |
| chr6_-_113411489 | 2.88 |
ENSMUST00000133348.2
|
Cidec
|
cell death-inducing DFFA-like effector c |
| chr1_+_74164700 | 2.88 |
ENSMUST00000080167.11
ENSMUST00000127134.2 |
Rufy4
|
RUN and FYVE domain containing 4 |
| chr2_+_84891281 | 2.85 |
ENSMUST00000238769.2
|
Tnks1bp1
|
tankyrase 1 binding protein 1 |
| chr2_-_17735847 | 2.85 |
ENSMUST00000028080.12
|
Nebl
|
nebulette |
| chr8_+_21868531 | 2.85 |
ENSMUST00000170275.4
|
Defa2
|
defensin, alpha, 2 |
| chr3_+_87855973 | 2.81 |
ENSMUST00000005019.6
|
Crabp2
|
cellular retinoic acid binding protein II |
| chr14_-_34310438 | 2.81 |
ENSMUST00000228044.2
ENSMUST00000022328.14 |
Ldb3
|
LIM domain binding 3 |
| chr2_+_110551976 | 2.79 |
ENSMUST00000090332.5
|
Muc15
|
mucin 15 |
| chr2_+_160722562 | 2.78 |
ENSMUST00000109456.9
|
Lpin3
|
lipin 3 |
| chr10_+_94386714 | 2.77 |
ENSMUST00000148910.3
ENSMUST00000117460.8 |
Tmcc3
|
transmembrane and coiled coil domains 3 |
| chr3_+_27371206 | 2.76 |
ENSMUST00000174840.2
|
Tnfsf10
|
tumor necrosis factor (ligand) superfamily, member 10 |
| chr2_+_25285878 | 2.76 |
ENSMUST00000028328.3
|
Entpd2
|
ectonucleoside triphosphate diphosphohydrolase 2 |
| chr9_-_75448979 | 2.74 |
ENSMUST00000214171.2
|
Tmod3
|
tropomodulin 3 |
| chr4_-_45489794 | 2.73 |
ENSMUST00000146236.8
|
Shb
|
src homology 2 domain-containing transforming protein B |
| chr6_+_17463819 | 2.67 |
ENSMUST00000140070.8
|
Met
|
met proto-oncogene |
| chr11_-_69906171 | 2.65 |
ENSMUST00000018718.8
ENSMUST00000102574.10 |
Acadvl
|
acyl-Coenzyme A dehydrogenase, very long chain |
| chr3_-_92346078 | 2.52 |
ENSMUST00000062160.4
|
Sprr1b
|
small proline-rich protein 1B |
| chr7_-_100613579 | 2.47 |
ENSMUST00000060174.6
|
P2ry6
|
pyrimidinergic receptor P2Y, G-protein coupled, 6 |
| chrX_+_132809189 | 2.46 |
ENSMUST00000113304.2
|
Srpx2
|
sushi-repeat-containing protein, X-linked 2 |
| chr14_-_52011035 | 2.46 |
ENSMUST00000073860.6
|
Ang4
|
angiogenin, ribonuclease A family, member 4 |
| chr6_-_122833109 | 2.46 |
ENSMUST00000042081.9
|
C3ar1
|
complement component 3a receptor 1 |
| chr1_-_37575313 | 2.43 |
ENSMUST00000042161.15
|
Mgat4a
|
mannoside acetylglucosaminyltransferase 4, isoenzyme A |
| chrX_-_165992145 | 2.38 |
ENSMUST00000112176.8
|
Tmsb4x
|
thymosin, beta 4, X chromosome |
| chrX_+_132809166 | 2.37 |
ENSMUST00000033606.15
|
Srpx2
|
sushi-repeat-containing protein, X-linked 2 |
| chr8_-_78244412 | 2.34 |
ENSMUST00000210922.2
ENSMUST00000210519.2 |
Arhgap10
|
Rho GTPase activating protein 10 |
| chrX_+_163052367 | 2.32 |
ENSMUST00000145412.8
ENSMUST00000033749.9 |
Pir
|
pirin |
| chr2_+_145627900 | 2.28 |
ENSMUST00000110005.8
ENSMUST00000094480.11 |
Rin2
|
Ras and Rab interactor 2 |
| chr1_+_180978491 | 2.21 |
ENSMUST00000134115.8
ENSMUST00000111059.2 |
Cnih4
|
cornichon family AMPA receptor auxiliary protein 4 |
| chr15_-_75886166 | 2.20 |
ENSMUST00000060807.12
|
Fam83h
|
family with sequence similarity 83, member H |
| chr3_-_122828592 | 2.20 |
ENSMUST00000029761.14
|
Myoz2
|
myozenin 2 |
| chr10_+_128769642 | 2.19 |
ENSMUST00000099112.4
ENSMUST00000218290.2 |
Itga7
|
integrin alpha 7 |
| chr3_-_131196213 | 2.18 |
ENSMUST00000197057.2
|
Sgms2
|
sphingomyelin synthase 2 |
| chr19_-_4087940 | 2.17 |
ENSMUST00000237893.2
ENSMUST00000169613.4 |
Gstp1
|
glutathione S-transferase, pi 1 |
| chr11_+_76836545 | 2.16 |
ENSMUST00000125145.8
|
Blmh
|
bleomycin hydrolase |
| chr7_-_19005721 | 2.16 |
ENSMUST00000032561.9
|
Vasp
|
vasodilator-stimulated phosphoprotein |
| chr13_-_24175141 | 2.15 |
ENSMUST00000021770.8
|
Scgn
|
secretagogin, EF-hand calcium binding protein |
| chr15_-_102165740 | 2.13 |
ENSMUST00000135466.2
|
Rarg
|
retinoic acid receptor, gamma |
| chrX_-_73067351 | 2.11 |
ENSMUST00000114353.10
ENSMUST00000101458.9 |
Irak1
|
interleukin-1 receptor-associated kinase 1 |
| chr16_+_33575920 | 2.06 |
ENSMUST00000128105.2
|
Heg1
|
heart development protein with EGF-like domains 1 |
| chr9_+_7347369 | 2.04 |
ENSMUST00000005950.12
ENSMUST00000065079.6 |
Mmp12
|
matrix metallopeptidase 12 |
| chr9_+_7272514 | 2.02 |
ENSMUST00000015394.10
|
Mmp13
|
matrix metallopeptidase 13 |
| chr14_-_55995912 | 2.02 |
ENSMUST00000001497.9
|
Cideb
|
cell death-inducing DNA fragmentation factor, alpha subunit-like effector B |
| chrX_-_73067514 | 2.00 |
ENSMUST00000033769.15
ENSMUST00000114352.8 ENSMUST00000068286.12 ENSMUST00000114360.10 ENSMUST00000114354.10 |
Irak1
|
interleukin-1 receptor-associated kinase 1 |
| chr9_-_86577940 | 2.00 |
ENSMUST00000034989.15
|
Me1
|
malic enzyme 1, NADP(+)-dependent, cytosolic |
| chr7_+_43418321 | 1.97 |
ENSMUST00000107970.8
|
Klk12
|
kallikrein related-peptidase 12 |
| chr19_-_4087907 | 1.95 |
ENSMUST00000237982.2
|
Gstp1
|
glutathione S-transferase, pi 1 |
| chr5_-_86780277 | 1.94 |
ENSMUST00000116553.9
|
Tmprss11f
|
transmembrane protease, serine 11f |
| chr18_+_61178211 | 1.92 |
ENSMUST00000025522.11
ENSMUST00000115274.2 |
Pdgfrb
|
platelet derived growth factor receptor, beta polypeptide |
| chr19_-_11796085 | 1.91 |
ENSMUST00000211047.2
ENSMUST00000075304.14 ENSMUST00000211641.2 |
Stx3
|
syntaxin 3 |
| chr19_+_29229147 | 1.91 |
ENSMUST00000025705.7
ENSMUST00000065796.10 ENSMUST00000236990.2 |
Jak2
|
Janus kinase 2 |
| chr6_+_48849804 | 1.90 |
ENSMUST00000204856.2
|
Aoc1
|
amine oxidase, copper-containing 1 |
| chr9_+_78522783 | 1.90 |
ENSMUST00000093812.5
|
Cd109
|
CD109 antigen |
| chr7_+_4463686 | 1.88 |
ENSMUST00000167298.2
ENSMUST00000171445.8 |
Eps8l1
|
EPS8-like 1 |
| chr3_+_79793237 | 1.85 |
ENSMUST00000029567.9
|
Gask1b
|
golgi associated kinase 1B |
| chr2_-_113883285 | 1.85 |
ENSMUST00000090269.7
|
Actc1
|
actin, alpha, cardiac muscle 1 |
| chr6_-_115735935 | 1.84 |
ENSMUST00000072933.13
|
Tmem40
|
transmembrane protein 40 |
| chr11_-_83957889 | 1.83 |
ENSMUST00000108101.8
|
Dusp14
|
dual specificity phosphatase 14 |
| chr11_+_73158214 | 1.79 |
ENSMUST00000049676.3
|
Trpv3
|
transient receptor potential cation channel, subfamily V, member 3 |
| chr18_-_35760260 | 1.77 |
ENSMUST00000025212.8
|
Slc23a1
|
solute carrier family 23 (nucleobase transporters), member 1 |
| chr2_+_162829250 | 1.77 |
ENSMUST00000018012.14
|
Sgk2
|
serum/glucocorticoid regulated kinase 2 |
| chr16_+_58548273 | 1.74 |
ENSMUST00000023426.12
ENSMUST00000162057.8 ENSMUST00000162191.2 |
Cldnd1
|
claudin domain containing 1 |
| chr9_-_106353571 | 1.73 |
ENSMUST00000123555.8
ENSMUST00000125850.2 |
Parp3
|
poly (ADP-ribose) polymerase family, member 3 |
| chr14_-_56499690 | 1.72 |
ENSMUST00000015581.6
|
Gzmb
|
granzyme B |
| chr16_-_30102070 | 1.71 |
ENSMUST00000064606.8
|
Lrrc15
|
leucine rich repeat containing 15 |
| chr7_-_133384449 | 1.71 |
ENSMUST00000063669.8
|
Dhx32
|
DEAH (Asp-Glu-Ala-His) box polypeptide 32 |
| chr9_+_106247943 | 1.71 |
ENSMUST00000173748.2
|
Dusp7
|
dual specificity phosphatase 7 |
| chr15_-_74650673 | 1.69 |
ENSMUST00000186014.2
ENSMUST00000191407.2 |
Ly6g6g
|
lymphocyte antigen 6 complex, locus G6G |
| chr1_+_170104889 | 1.68 |
ENSMUST00000179976.3
|
Sh2d1b1
|
SH2 domain containing 1B1 |
| chr11_+_76836330 | 1.64 |
ENSMUST00000021197.10
|
Blmh
|
bleomycin hydrolase |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.3 | 13.0 | GO:0046108 | uridine catabolic process(GO:0006218) uridine metabolic process(GO:0046108) |
| 4.0 | 16.2 | GO:1903575 | cornified envelope assembly(GO:1903575) |
| 3.8 | 11.3 | GO:0018146 | keratan sulfate biosynthetic process(GO:0018146) |
| 2.5 | 7.5 | GO:0010482 | epidermal cell division(GO:0010481) regulation of epidermal cell division(GO:0010482) |
| 2.4 | 9.4 | GO:0021812 | neuronal-glial interaction involved in cerebral cortex radial glia guided migration(GO:0021812) |
| 2.2 | 6.5 | GO:0002514 | B cell tolerance induction(GO:0002514) regulation of tolerance induction to self antigen(GO:0002649) regulation of B cell tolerance induction(GO:0002661) positive regulation of B cell tolerance induction(GO:0002663) |
| 1.8 | 5.3 | GO:0015881 | creatine transport(GO:0015881) |
| 1.7 | 8.6 | GO:0014722 | regulation of skeletal muscle contraction by calcium ion signaling(GO:0014722) |
| 1.7 | 6.7 | GO:0006069 | ethanol oxidation(GO:0006069) |
| 1.4 | 7.0 | GO:2000048 | negative regulation of cell-cell adhesion mediated by cadherin(GO:2000048) |
| 1.4 | 4.1 | GO:2000469 | negative regulation of peroxidase activity(GO:2000469) |
| 1.3 | 19.7 | GO:0002934 | desmosome organization(GO:0002934) |
| 1.3 | 9.1 | GO:0070495 | regulation of thrombin receptor signaling pathway(GO:0070494) negative regulation of thrombin receptor signaling pathway(GO:0070495) |
| 1.3 | 20.9 | GO:1900029 | positive regulation of ruffle assembly(GO:1900029) |
| 1.3 | 5.2 | GO:1901003 | negative regulation of fermentation(GO:1901003) |
| 1.2 | 3.7 | GO:0099553 | trans-synaptic signaling by lipid, modulating synaptic transmission(GO:0099552) trans-synaptic signaling by endocannabinoid, modulating synaptic transmission(GO:0099553) |
| 1.2 | 7.3 | GO:0033540 | fatty acid beta-oxidation using acyl-CoA oxidase(GO:0033540) |
| 1.0 | 5.9 | GO:0030043 | actin filament fragmentation(GO:0030043) |
| 1.0 | 2.9 | GO:0051563 | smooth endoplasmic reticulum calcium ion homeostasis(GO:0051563) cellular response to norepinephrine stimulus(GO:0071874) |
| 0.9 | 3.8 | GO:0043418 | homocysteine catabolic process(GO:0043418) |
| 0.9 | 18.9 | GO:0061436 | establishment of skin barrier(GO:0061436) |
| 0.9 | 8.4 | GO:0042904 | 9-cis-retinoic acid biosynthetic process(GO:0042904) 9-cis-retinoic acid metabolic process(GO:0042905) |
| 0.9 | 9.6 | GO:0040009 | regulation of growth rate(GO:0040009) |
| 0.9 | 3.4 | GO:0090472 | dibasic protein processing(GO:0090472) |
| 0.8 | 5.0 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 0.8 | 4.1 | GO:1904970 | brush border assembly(GO:1904970) |
| 0.8 | 4.8 | GO:2000587 | negative regulation of platelet-derived growth factor receptor-beta signaling pathway(GO:2000587) |
| 0.8 | 3.2 | GO:1903984 | positive regulation of TRAIL-activated apoptotic signaling pathway(GO:1903984) |
| 0.8 | 10.3 | GO:0010839 | negative regulation of keratinocyte proliferation(GO:0010839) |
| 0.7 | 4.4 | GO:0014826 | regulation of systemic arterial blood pressure by endothelin(GO:0003100) vein smooth muscle contraction(GO:0014826) |
| 0.7 | 2.1 | GO:0003430 | growth plate cartilage chondrocyte growth(GO:0003430) |
| 0.7 | 4.2 | GO:0050689 | negative regulation of defense response to virus by host(GO:0050689) |
| 0.7 | 21.5 | GO:0031424 | keratinization(GO:0031424) |
| 0.7 | 6.8 | GO:0016081 | synaptic vesicle docking(GO:0016081) |
| 0.6 | 1.9 | GO:0035441 | cell migration involved in vasculogenesis(GO:0035441) |
| 0.6 | 3.2 | GO:2000660 | negative regulation of interleukin-1-mediated signaling pathway(GO:2000660) |
| 0.6 | 1.9 | GO:0035722 | interleukin-12-mediated signaling pathway(GO:0035722) cellular response to interleukin-12(GO:0071349) |
| 0.6 | 1.9 | GO:0097185 | amiloride transport(GO:0015898) cellular response to copper ion starvation(GO:0035874) response to azide(GO:0097184) cellular response to azide(GO:0097185) |
| 0.6 | 1.8 | GO:0070904 | L-ascorbic acid transport(GO:0015882) transepithelial L-ascorbic acid transport(GO:0070904) |
| 0.6 | 4.1 | GO:0001712 | ectodermal cell fate commitment(GO:0001712) |
| 0.6 | 8.5 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.5 | 4.9 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 0.5 | 3.2 | GO:1990166 | protein localization to site of double-strand break(GO:1990166) |
| 0.5 | 2.1 | GO:0003017 | lymph circulation(GO:0003017) |
| 0.5 | 4.1 | GO:0090370 | negative regulation of cholesterol efflux(GO:0090370) |
| 0.5 | 2.0 | GO:0090187 | positive regulation of pancreatic juice secretion(GO:0090187) |
| 0.5 | 10.0 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.4 | 5.7 | GO:0061299 | retina vasculature morphogenesis in camera-type eye(GO:0061299) |
| 0.4 | 3.9 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) |
| 0.4 | 4.8 | GO:0007406 | negative regulation of neuroblast proliferation(GO:0007406) |
| 0.4 | 1.3 | GO:0001983 | baroreceptor response to increased systemic arterial blood pressure(GO:0001983) |
| 0.4 | 3.0 | GO:0000103 | sulfate assimilation(GO:0000103) |
| 0.4 | 1.7 | GO:0002767 | immune response-inhibiting cell surface receptor signaling pathway(GO:0002767) |
| 0.4 | 10.9 | GO:0060742 | epithelial cell differentiation involved in prostate gland development(GO:0060742) |
| 0.4 | 2.5 | GO:0030321 | transepithelial chloride transport(GO:0030321) |
| 0.4 | 6.7 | GO:2000252 | negative regulation of feeding behavior(GO:2000252) |
| 0.4 | 14.9 | GO:0031954 | positive regulation of protein autophosphorylation(GO:0031954) |
| 0.4 | 1.1 | GO:0006435 | threonyl-tRNA aminoacylation(GO:0006435) |
| 0.4 | 3.2 | GO:0006108 | malate metabolic process(GO:0006108) |
| 0.4 | 2.5 | GO:0002461 | complement receptor mediated signaling pathway(GO:0002430) tolerance induction dependent upon immune response(GO:0002461) positive regulation of glomerular mesangial cell proliferation(GO:0072126) |
| 0.3 | 4.8 | GO:0090050 | positive regulation of cell migration involved in sprouting angiogenesis(GO:0090050) |
| 0.3 | 1.7 | GO:0008626 | granzyme-mediated apoptotic signaling pathway(GO:0008626) |
| 0.3 | 8.4 | GO:0034389 | lipid particle organization(GO:0034389) |
| 0.3 | 3.4 | GO:0010727 | negative regulation of hydrogen peroxide metabolic process(GO:0010727) |
| 0.3 | 10.9 | GO:0043616 | keratinocyte proliferation(GO:0043616) |
| 0.3 | 0.9 | GO:0002541 | activation of plasma proteins involved in acute inflammatory response(GO:0002541) |
| 0.3 | 3.4 | GO:0015705 | iodide transport(GO:0015705) |
| 0.3 | 2.8 | GO:0009181 | purine nucleoside diphosphate catabolic process(GO:0009137) purine ribonucleoside diphosphate catabolic process(GO:0009181) |
| 0.3 | 0.9 | GO:0010808 | positive regulation of synaptic vesicle priming(GO:0010808) |
| 0.3 | 4.0 | GO:0051152 | positive regulation of smooth muscle cell differentiation(GO:0051152) |
| 0.3 | 1.7 | GO:0046813 | receptor-mediated virion attachment to host cell(GO:0046813) |
| 0.3 | 3.6 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.3 | 0.5 | GO:0061152 | trachea submucosa development(GO:0061152) trachea gland development(GO:0061153) |
| 0.3 | 2.7 | GO:0042760 | very long-chain fatty acid catabolic process(GO:0042760) |
| 0.3 | 1.3 | GO:0052041 | negative regulation by symbiont of host apoptotic process(GO:0033668) negative regulation by symbiont of host programmed cell death(GO:0052041) negative regulation by organism of programmed cell death in other organism involved in symbiotic interaction(GO:0052490) |
| 0.3 | 1.6 | GO:0038128 | ERBB2 signaling pathway(GO:0038128) |
| 0.3 | 1.5 | GO:0019371 | cyclooxygenase pathway(GO:0019371) |
| 0.3 | 2.0 | GO:0060339 | negative regulation of type I interferon-mediated signaling pathway(GO:0060339) |
| 0.3 | 2.8 | GO:0042573 | retinoic acid metabolic process(GO:0042573) |
| 0.3 | 1.8 | GO:0042636 | negative regulation of hair cycle(GO:0042636) |
| 0.3 | 7.5 | GO:0014067 | negative regulation of phosphatidylinositol 3-kinase signaling(GO:0014067) |
| 0.2 | 2.7 | GO:1900194 | negative regulation of oocyte maturation(GO:1900194) |
| 0.2 | 13.9 | GO:0006730 | one-carbon metabolic process(GO:0006730) |
| 0.2 | 3.7 | GO:0090136 | epithelial cell-cell adhesion(GO:0090136) |
| 0.2 | 0.7 | GO:0036145 | dendritic cell homeostasis(GO:0036145) |
| 0.2 | 1.2 | GO:0000117 | regulation of transcription involved in G2/M transition of mitotic cell cycle(GO:0000117) |
| 0.2 | 1.2 | GO:1905161 | protein localization to phagocytic vesicle(GO:1905161) regulation of protein localization to phagocytic vesicle(GO:1905169) positive regulation of protein localization to phagocytic vesicle(GO:1905171) |
| 0.2 | 2.7 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.2 | 3.4 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.2 | 1.3 | GO:1901674 | histone H3-K27 acetylation(GO:0043974) spongiotrophoblast differentiation(GO:0060708) regulation of histone H3-K27 acetylation(GO:1901674) |
| 0.2 | 2.4 | GO:0030240 | skeletal muscle thin filament assembly(GO:0030240) |
| 0.2 | 0.8 | GO:0009838 | abscission(GO:0009838) |
| 0.2 | 4.1 | GO:0044406 | adhesion of symbiont to host(GO:0044406) |
| 0.2 | 7.1 | GO:0070584 | mitochondrion morphogenesis(GO:0070584) |
| 0.2 | 5.9 | GO:0090200 | positive regulation of release of cytochrome c from mitochondria(GO:0090200) |
| 0.2 | 1.4 | GO:0010815 | bradykinin catabolic process(GO:0010815) |
| 0.2 | 1.4 | GO:2000286 | regulation of endosome size(GO:0051036) receptor internalization involved in canonical Wnt signaling pathway(GO:2000286) |
| 0.2 | 10.5 | GO:1901385 | regulation of voltage-gated calcium channel activity(GO:1901385) |
| 0.2 | 5.1 | GO:0070208 | protein heterotrimerization(GO:0070208) |
| 0.2 | 1.6 | GO:0072602 | interleukin-4 secretion(GO:0072602) |
| 0.2 | 0.8 | GO:0002268 | follicular dendritic cell activation(GO:0002266) follicular dendritic cell differentiation(GO:0002268) |
| 0.2 | 3.3 | GO:1900120 | regulation of receptor binding(GO:1900120) |
| 0.1 | 4.9 | GO:0030502 | negative regulation of bone mineralization(GO:0030502) |
| 0.1 | 1.9 | GO:2001288 | positive regulation of caveolin-mediated endocytosis(GO:2001288) |
| 0.1 | 1.2 | GO:0071883 | activation of MAPK activity by adrenergic receptor signaling pathway(GO:0071883) |
| 0.1 | 21.8 | GO:0007586 | digestion(GO:0007586) |
| 0.1 | 1.0 | GO:0035726 | common myeloid progenitor cell proliferation(GO:0035726) |
| 0.1 | 2.9 | GO:0010039 | response to iron ion(GO:0010039) |
| 0.1 | 11.1 | GO:0030239 | myofibril assembly(GO:0030239) |
| 0.1 | 1.0 | GO:1900147 | Schwann cell migration(GO:0036135) regulation of Schwann cell migration(GO:1900147) |
| 0.1 | 0.9 | GO:0044805 | late nucleophagy(GO:0044805) |
| 0.1 | 0.6 | GO:1902167 | positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator(GO:1902167) |
| 0.1 | 0.6 | GO:0006538 | glutamate catabolic process(GO:0006538) |
| 0.1 | 3.6 | GO:0033198 | response to ATP(GO:0033198) |
| 0.1 | 5.4 | GO:0050819 | negative regulation of coagulation(GO:0050819) |
| 0.1 | 1.2 | GO:0016540 | protein autoprocessing(GO:0016540) |
| 0.1 | 0.8 | GO:0018243 | protein O-linked glycosylation via threonine(GO:0018243) |
| 0.1 | 2.5 | GO:0019731 | antibacterial humoral response(GO:0019731) |
| 0.1 | 7.2 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.1 | 0.8 | GO:1903912 | negative regulation of endoplasmic reticulum stress-induced eIF2 alpha phosphorylation(GO:1903912) |
| 0.1 | 1.3 | GO:0006228 | UTP biosynthetic process(GO:0006228) |
| 0.1 | 5.4 | GO:0034113 | heterotypic cell-cell adhesion(GO:0034113) |
| 0.1 | 0.8 | GO:0071394 | cellular response to testosterone stimulus(GO:0071394) |
| 0.1 | 0.4 | GO:0097368 | establishment of Sertoli cell barrier(GO:0097368) |
| 0.1 | 3.7 | GO:0019432 | triglyceride biosynthetic process(GO:0019432) |
| 0.1 | 3.5 | GO:0035335 | peptidyl-tyrosine dephosphorylation(GO:0035335) |
| 0.1 | 0.6 | GO:0035815 | positive regulation of renal sodium excretion(GO:0035815) |
| 0.1 | 4.2 | GO:0007340 | acrosome reaction(GO:0007340) |
| 0.1 | 1.0 | GO:1901857 | positive regulation of cellular respiration(GO:1901857) |
| 0.1 | 1.4 | GO:0006027 | glycosaminoglycan catabolic process(GO:0006027) |
| 0.1 | 1.6 | GO:0045589 | regulation of regulatory T cell differentiation(GO:0045589) |
| 0.1 | 0.6 | GO:0032876 | negative regulation of DNA endoreduplication(GO:0032876) |
| 0.1 | 1.1 | GO:0048484 | enteric nervous system development(GO:0048484) |
| 0.1 | 0.5 | GO:0072093 | ureteric bud invasion(GO:0072092) metanephric renal vesicle formation(GO:0072093) |
| 0.1 | 0.8 | GO:0018095 | protein polyglutamylation(GO:0018095) |
| 0.1 | 1.4 | GO:0031987 | locomotion involved in locomotory behavior(GO:0031987) |
| 0.1 | 3.5 | GO:0044380 | protein localization to cytoskeleton(GO:0044380) |
| 0.1 | 2.1 | GO:0030224 | monocyte differentiation(GO:0030224) |
| 0.1 | 0.3 | GO:0021506 | anterior neuropore closure(GO:0021506) neuropore closure(GO:0021995) |
| 0.1 | 0.3 | GO:0060332 | positive regulation of response to interferon-gamma(GO:0060332) positive regulation of interferon-gamma-mediated signaling pathway(GO:0060335) |
| 0.1 | 3.1 | GO:0001954 | positive regulation of cell-matrix adhesion(GO:0001954) |
| 0.1 | 10.1 | GO:0007160 | cell-matrix adhesion(GO:0007160) |
| 0.1 | 3.4 | GO:1902476 | chloride transmembrane transport(GO:1902476) |
| 0.1 | 0.6 | GO:0071361 | cellular response to ethanol(GO:0071361) |
| 0.0 | 5.2 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 0.0 | 0.2 | GO:1902269 | positive regulation of polyamine transmembrane transport(GO:1902269) |
| 0.0 | 1.0 | GO:0060707 | trophoblast giant cell differentiation(GO:0060707) |
| 0.0 | 3.3 | GO:0002066 | columnar/cuboidal epithelial cell development(GO:0002066) |
| 0.0 | 0.8 | GO:0043552 | positive regulation of phosphatidylinositol 3-kinase activity(GO:0043552) |
| 0.0 | 3.3 | GO:0071594 | T cell differentiation in thymus(GO:0033077) thymocyte aggregation(GO:0071594) |
| 0.0 | 0.9 | GO:0039703 | viral RNA genome replication(GO:0039694) RNA replication(GO:0039703) |
| 0.0 | 2.7 | GO:0050830 | defense response to Gram-positive bacterium(GO:0050830) |
| 0.0 | 8.6 | GO:0008544 | epidermis development(GO:0008544) |
| 0.0 | 4.1 | GO:0006888 | ER to Golgi vesicle-mediated transport(GO:0006888) |
| 0.0 | 2.9 | GO:0006749 | glutathione metabolic process(GO:0006749) |
| 0.0 | 3.2 | GO:0035914 | skeletal muscle cell differentiation(GO:0035914) |
| 0.0 | 0.7 | GO:0008090 | retrograde axonal transport(GO:0008090) |
| 0.0 | 0.3 | GO:0061469 | regulation of type B pancreatic cell proliferation(GO:0061469) |
| 0.0 | 4.0 | GO:0030509 | BMP signaling pathway(GO:0030509) |
| 0.0 | 0.3 | GO:0036444 | calcium ion transmembrane import into mitochondrion(GO:0036444) |
| 0.0 | 0.6 | GO:0001893 | maternal placenta development(GO:0001893) |
| 0.0 | 2.2 | GO:0051289 | protein homotetramerization(GO:0051289) |
| 0.0 | 1.1 | GO:0009250 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.0 | 0.3 | GO:0002716 | negative regulation of leukocyte mediated cytotoxicity(GO:0001911) negative regulation of natural killer cell mediated immunity(GO:0002716) negative regulation of natural killer cell mediated cytotoxicity(GO:0045953) |
| 0.0 | 0.5 | GO:0006516 | glycoprotein catabolic process(GO:0006516) |
| 0.0 | 2.6 | GO:0051781 | positive regulation of cell division(GO:0051781) |
| 0.0 | 0.5 | GO:0008608 | attachment of spindle microtubules to kinetochore(GO:0008608) |
| 0.0 | 0.3 | GO:0032516 | positive regulation of phosphoprotein phosphatase activity(GO:0032516) |
| 0.0 | 0.2 | GO:0042226 | interleukin-6 biosynthetic process(GO:0042226) |
| 0.0 | 1.0 | GO:0046513 | ceramide biosynthetic process(GO:0046513) |
| 0.0 | 1.0 | GO:0043124 | negative regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043124) |
| 0.0 | 0.3 | GO:0010667 | negative regulation of cardiac muscle cell apoptotic process(GO:0010667) |
| 0.0 | 1.3 | GO:0016202 | regulation of striated muscle tissue development(GO:0016202) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.1 | 9.4 | GO:0005607 | laminin-2 complex(GO:0005607) |
| 2.1 | 8.5 | GO:0005608 | laminin-3 complex(GO:0005608) |
| 1.5 | 7.4 | GO:0005914 | spot adherens junction(GO:0005914) |
| 1.4 | 7.0 | GO:0097057 | TRAF2-GSTP1 complex(GO:0097057) |
| 1.0 | 7.1 | GO:0097513 | myosin II filament(GO:0097513) |
| 0.7 | 2.2 | GO:0034677 | integrin alpha7-beta1 complex(GO:0034677) |
| 0.7 | 5.0 | GO:0005638 | lamin filament(GO:0005638) |
| 0.6 | 1.8 | GO:0042643 | actomyosin, actin portion(GO:0042643) |
| 0.6 | 3.6 | GO:1990796 | photoreceptor cell terminal bouton(GO:1990796) |
| 0.6 | 11.0 | GO:0042599 | lamellar body(GO:0042599) |
| 0.5 | 26.0 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.5 | 1.4 | GO:0098842 | postsynaptic early endosome(GO:0098842) |
| 0.4 | 1.7 | GO:0044194 | cytolytic granule(GO:0044194) |
| 0.4 | 4.1 | GO:0089717 | spanning component of plasma membrane(GO:0044214) spanning component of membrane(GO:0089717) |
| 0.3 | 3.7 | GO:0031094 | platelet dense tubular network(GO:0031094) |
| 0.3 | 0.9 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 0.3 | 4.1 | GO:0020005 | symbiont-containing vacuole membrane(GO:0020005) |
| 0.3 | 8.8 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.3 | 2.9 | GO:1990812 | growth cone lamellipodium(GO:1990761) growth cone filopodium(GO:1990812) |
| 0.2 | 7.7 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.2 | 3.2 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 0.2 | 3.4 | GO:0012510 | trans-Golgi network transport vesicle membrane(GO:0012510) |
| 0.2 | 3.4 | GO:0036056 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.2 | 3.6 | GO:0031229 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) |
| 0.2 | 1.3 | GO:0005827 | polar microtubule(GO:0005827) |
| 0.2 | 4.2 | GO:0031362 | anchored component of external side of plasma membrane(GO:0031362) |
| 0.2 | 0.8 | GO:0033257 | Bcl3/NF-kappaB2 complex(GO:0033257) |
| 0.1 | 0.9 | GO:0098831 | presynaptic active zone cytoplasmic component(GO:0098831) |
| 0.1 | 2.8 | GO:0005605 | basal lamina(GO:0005605) |
| 0.1 | 1.7 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.1 | 2.0 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.1 | 4.6 | GO:0008305 | integrin complex(GO:0008305) |
| 0.1 | 15.2 | GO:0030315 | T-tubule(GO:0030315) |
| 0.1 | 11.8 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.1 | 13.5 | GO:0005811 | lipid particle(GO:0005811) |
| 0.1 | 21.0 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.1 | 6.7 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.1 | 17.9 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.1 | 0.7 | GO:0031673 | H zone(GO:0031673) |
| 0.1 | 1.1 | GO:0042587 | glycogen granule(GO:0042587) |
| 0.1 | 7.4 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.1 | 7.5 | GO:0032420 | stereocilium(GO:0032420) |
| 0.1 | 1.0 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.1 | 1.2 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.1 | 10.9 | GO:0005901 | caveola(GO:0005901) |
| 0.1 | 3.2 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.1 | 7.0 | GO:0030864 | cortical actin cytoskeleton(GO:0030864) |
| 0.1 | 4.3 | GO:0005902 | microvillus(GO:0005902) |
| 0.1 | 0.3 | GO:1990246 | uniplex complex(GO:1990246) |
| 0.1 | 0.8 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.1 | 7.3 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.1 | 6.0 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.1 | 4.5 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.1 | 4.8 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.1 | 1.9 | GO:0070971 | endoplasmic reticulum exit site(GO:0070971) |
| 0.1 | 4.4 | GO:0005604 | basement membrane(GO:0005604) |
| 0.0 | 5.8 | GO:0005884 | actin filament(GO:0005884) |
| 0.0 | 0.8 | GO:0090543 | Flemming body(GO:0090543) |
| 0.0 | 6.7 | GO:0043296 | apical junction complex(GO:0043296) |
| 0.0 | 0.3 | GO:0044754 | autolysosome(GO:0044754) |
| 0.0 | 2.5 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 8.0 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 0.5 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.0 | 2.0 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.0 | 1.2 | GO:0031985 | Golgi cisterna(GO:0031985) |
| 0.0 | 11.3 | GO:0000139 | Golgi membrane(GO:0000139) |
| 0.0 | 73.9 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 0.8 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.0 | 5.6 | GO:0005777 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.0 | 0.9 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.0 | 1.1 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.0 | 0.6 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.0 | 0.6 | GO:0005771 | multivesicular body(GO:0005771) |
| 0.0 | 0.5 | GO:0005859 | muscle myosin complex(GO:0005859) |
| 0.0 | 2.6 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 2.5 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 60.4 | GO:0005576 | extracellular region(GO:0005576) |
| 0.0 | 1.7 | GO:0016324 | apical plasma membrane(GO:0016324) |
| 0.0 | 1.0 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 0.5 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.0 | 2.3 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.0 | 15.3 | GO:0005887 | integral component of plasma membrane(GO:0005887) |
| 0.0 | 2.3 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.0 | 0.3 | GO:0090545 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.0 | 1.0 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.3 | 7.0 | GO:0035731 | S-nitrosoglutathione binding(GO:0035730) dinitrosyl-iron complex binding(GO:0035731) |
| 2.3 | 13.9 | GO:0004022 | alcohol dehydrogenase (NAD) activity(GO:0004022) |
| 2.3 | 9.1 | GO:0005008 | hepatocyte growth factor-activated receptor activity(GO:0005008) |
| 2.3 | 13.5 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 2.2 | 6.5 | GO:0005026 | transforming growth factor beta receptor activity, type II(GO:0005026) |
| 2.2 | 13.0 | GO:0004850 | uridine phosphorylase activity(GO:0004850) |
| 1.8 | 5.3 | GO:0005308 | creatine transmembrane transporter activity(GO:0005308) |
| 1.6 | 11.3 | GO:0001517 | N-acetylglucosamine 6-O-sulfotransferase activity(GO:0001517) |
| 1.6 | 4.8 | GO:0019153 | protein-disulfide reductase (glutathione) activity(GO:0019153) |
| 1.2 | 12.1 | GO:0045545 | syndecan binding(GO:0045545) |
| 1.2 | 8.4 | GO:0030549 | acetylcholine receptor activator activity(GO:0030549) |
| 1.1 | 4.4 | GO:0031708 | endothelin B receptor binding(GO:0031708) |
| 1.1 | 4.3 | GO:0005152 | interleukin-1 receptor antagonist activity(GO:0005152) |
| 1.0 | 9.4 | GO:0043208 | glycosphingolipid binding(GO:0043208) |
| 1.0 | 4.9 | GO:0005134 | interleukin-2 receptor binding(GO:0005134) |
| 0.9 | 3.7 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.9 | 7.2 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.9 | 2.7 | GO:0017099 | very-long-chain-acyl-CoA dehydrogenase activity(GO:0017099) |
| 0.8 | 11.7 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.8 | 2.3 | GO:0008127 | quercetin 2,3-dioxygenase activity(GO:0008127) |
| 0.8 | 13.9 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.7 | 14.5 | GO:0097493 | structural molecule activity conferring elasticity(GO:0097493) |
| 0.7 | 2.0 | GO:0004473 | malic enzyme activity(GO:0004470) malate dehydrogenase (decarboxylating) (NAD+) activity(GO:0004471) malate dehydrogenase (decarboxylating) (NADP+) activity(GO:0004473) |
| 0.6 | 16.9 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.6 | 1.9 | GO:0005017 | platelet-derived growth factor-activated receptor activity(GO:0005017) |
| 0.6 | 1.9 | GO:0052597 | diamine oxidase activity(GO:0052597) histamine oxidase activity(GO:0052598) methylputrescine oxidase activity(GO:0052599) propane-1,3-diamine oxidase activity(GO:0052600) |
| 0.6 | 2.5 | GO:0071553 | uridine nucleotide receptor activity(GO:0015065) G-protein coupled pyrimidinergic nucleotide receptor activity(GO:0071553) |
| 0.6 | 7.3 | GO:0003997 | acyl-CoA oxidase activity(GO:0003997) |
| 0.6 | 1.8 | GO:0008520 | L-ascorbate:sodium symporter activity(GO:0008520) L-ascorbic acid transporter activity(GO:0015229) sodium-dependent L-ascorbate transmembrane transporter activity(GO:0070890) |
| 0.6 | 3.4 | GO:0015111 | iodide transmembrane transporter activity(GO:0015111) |
| 0.5 | 3.2 | GO:1990254 | keratin filament binding(GO:1990254) |
| 0.5 | 19.4 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.4 | 3.5 | GO:0000293 | ferric-chelate reductase activity(GO:0000293) |
| 0.4 | 2.2 | GO:0051373 | FATZ binding(GO:0051373) |
| 0.4 | 3.0 | GO:0050294 | alcohol sulfotransferase activity(GO:0004027) steroid sulfotransferase activity(GO:0050294) |
| 0.4 | 13.4 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.4 | 2.9 | GO:0071532 | ankyrin repeat binding(GO:0071532) |
| 0.4 | 3.6 | GO:0004931 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.4 | 1.9 | GO:0005143 | interleukin-12 receptor binding(GO:0005143) |
| 0.4 | 6.8 | GO:0050544 | arachidonic acid binding(GO:0050544) |
| 0.4 | 5.1 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.4 | 2.9 | GO:0051425 | PTB domain binding(GO:0051425) |
| 0.4 | 1.1 | GO:0004829 | threonine-tRNA ligase activity(GO:0004829) |
| 0.3 | 4.4 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.3 | 1.3 | GO:0005146 | leukemia inhibitory factor receptor binding(GO:0005146) |
| 0.3 | 4.2 | GO:0046703 | natural killer cell lectin-like receptor binding(GO:0046703) |
| 0.3 | 1.3 | GO:0004937 | alpha1-adrenergic receptor activity(GO:0004937) |
| 0.3 | 3.2 | GO:0003909 | DNA ligase activity(GO:0003909) DNA ligase (ATP) activity(GO:0003910) |
| 0.3 | 2.8 | GO:0043262 | adenosine-diphosphatase activity(GO:0043262) |
| 0.3 | 4.7 | GO:0045028 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.3 | 12.2 | GO:0043236 | laminin binding(GO:0043236) |
| 0.3 | 7.1 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.3 | 2.5 | GO:0004875 | complement receptor activity(GO:0004875) |
| 0.3 | 3.0 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.3 | 11.3 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.2 | 1.2 | GO:0030060 | L-malate dehydrogenase activity(GO:0030060) |
| 0.2 | 12.6 | GO:0042805 | actinin binding(GO:0042805) |
| 0.2 | 3.7 | GO:0005522 | profilin binding(GO:0005522) |
| 0.2 | 0.8 | GO:0070740 | tubulin-glutamic acid ligase activity(GO:0070740) |
| 0.2 | 3.5 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.2 | 3.4 | GO:0048406 | nerve growth factor binding(GO:0048406) |
| 0.2 | 2.2 | GO:0031730 | CCR5 chemokine receptor binding(GO:0031730) |
| 0.2 | 1.0 | GO:1990050 | phosphatidic acid transporter activity(GO:1990050) |
| 0.2 | 1.5 | GO:0017050 | sphinganine kinase activity(GO:0008481) D-erythro-sphingosine kinase activity(GO:0017050) |
| 0.2 | 1.0 | GO:0047493 | sphingomyelin synthase activity(GO:0033188) ceramide cholinephosphotransferase activity(GO:0047493) |
| 0.2 | 4.9 | GO:0019956 | chemokine binding(GO:0019956) |
| 0.2 | 1.4 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.2 | 2.1 | GO:0003708 | retinoic acid receptor activity(GO:0003708) |
| 0.1 | 3.0 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.1 | 4.0 | GO:0004622 | lysophospholipase activity(GO:0004622) |
| 0.1 | 1.4 | GO:0004415 | hyalurononglucosaminidase activity(GO:0004415) |
| 0.1 | 1.4 | GO:0051021 | GDP-dissociation inhibitor binding(GO:0051021) |
| 0.1 | 2.8 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.1 | 0.7 | GO:0008142 | oxysterol binding(GO:0008142) |
| 0.1 | 5.7 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.1 | 10.4 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
| 0.1 | 0.9 | GO:0004366 | glycerol-3-phosphate O-acyltransferase activity(GO:0004366) |
| 0.1 | 5.8 | GO:0050750 | low-density lipoprotein particle receptor binding(GO:0050750) |
| 0.1 | 2.8 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.1 | 2.8 | GO:0017081 | chloride channel regulator activity(GO:0017081) |
| 0.1 | 1.2 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.1 | 7.8 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.1 | 2.5 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.1 | 3.9 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.1 | 1.3 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.1 | 0.6 | GO:0070728 | leucine binding(GO:0070728) |
| 0.1 | 1.3 | GO:0005041 | low-density lipoprotein receptor activity(GO:0005041) |
| 0.1 | 1.9 | GO:0019871 | potassium channel inhibitor activity(GO:0019870) sodium channel inhibitor activity(GO:0019871) |
| 0.1 | 16.1 | GO:0008083 | growth factor activity(GO:0008083) |
| 0.1 | 1.3 | GO:0002161 | aminoacyl-tRNA editing activity(GO:0002161) |
| 0.1 | 0.6 | GO:0042030 | ATPase inhibitor activity(GO:0042030) |
| 0.1 | 4.0 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 5.2 | GO:0004177 | aminopeptidase activity(GO:0004177) |
| 0.1 | 2.3 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.1 | 1.9 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.1 | 1.2 | GO:0001206 | transcriptional repressor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001206) |
| 0.1 | 2.0 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.1 | 1.2 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
| 0.1 | 0.9 | GO:0031005 | filamin binding(GO:0031005) |
| 0.1 | 0.9 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.1 | 2.1 | GO:0071855 | neuropeptide receptor binding(GO:0071855) |
| 0.1 | 12.8 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.1 | 0.6 | GO:0051428 | peptide hormone receptor binding(GO:0051428) |
| 0.0 | 0.2 | GO:0042978 | ornithine decarboxylase activator activity(GO:0042978) |
| 0.0 | 3.6 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.0 | 4.7 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 7.6 | GO:0051219 | phosphoprotein binding(GO:0051219) |
| 0.0 | 0.6 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.0 | 4.6 | GO:0019838 | growth factor binding(GO:0019838) |
| 0.0 | 0.8 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.0 | 0.6 | GO:0031210 | phosphatidylcholine binding(GO:0031210) |
| 0.0 | 3.9 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.0 | 1.3 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.0 | 0.5 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.0 | 2.0 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.3 | GO:0019855 | calcium channel inhibitor activity(GO:0019855) |
| 0.0 | 1.5 | GO:0016667 | oxidoreductase activity, acting on a sulfur group of donors(GO:0016667) |
| 0.0 | 0.4 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
| 0.0 | 6.1 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.0 | 0.6 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.0 | 5.2 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.0 | 1.0 | GO:0031593 | polyubiquitin binding(GO:0031593) |
| 0.0 | 1.8 | GO:0005262 | calcium channel activity(GO:0005262) |
| 0.0 | 0.2 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.0 | 0.2 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.0 | 0.3 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.0 | 1.7 | GO:0004519 | endonuclease activity(GO:0004519) |
| 0.0 | 7.4 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 17.9 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.3 | 24.4 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.3 | 13.3 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.2 | 7.4 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.2 | 6.8 | PID INTEGRIN CS PATHWAY | Integrin family cell surface interactions |
| 0.2 | 15.0 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.2 | 6.3 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.1 | 1.9 | ST INTERFERON GAMMA PATHWAY | Interferon gamma pathway. |
| 0.1 | 4.7 | PID TRAIL PATHWAY | TRAIL signaling pathway |
| 0.1 | 24.0 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.1 | 4.6 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
| 0.1 | 6.9 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.1 | 10.6 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.1 | 3.0 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.1 | 2.7 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.1 | 1.6 | PID AVB3 OPN PATHWAY | Osteopontin-mediated events |
| 0.1 | 4.2 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.1 | 1.4 | PID IL8 CXCR1 PATHWAY | IL8- and CXCR1-mediated signaling events |
| 0.1 | 2.1 | PID THROMBIN PAR1 PATHWAY | PAR1-mediated thrombin signaling events |
| 0.1 | 1.1 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.1 | 1.3 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.1 | 5.0 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.1 | 3.8 | PID PI3KCI PATHWAY | Class I PI3K signaling events |
| 0.1 | 2.1 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.1 | 3.4 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 1.3 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.0 | 2.7 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 1.4 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.0 | 9.0 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 6.5 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 1.4 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.0 | 1.9 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 0.9 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.0 | 0.6 | PID CONE PATHWAY | Visual signal transduction: Cones |
| 0.0 | 0.7 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.0 | 0.2 | PID IL23 PATHWAY | IL23-mediated signaling events |
| 0.0 | 0.8 | PID IL12 2PATHWAY | IL12-mediated signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 13.1 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.6 | 6.7 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.5 | 12.2 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.4 | 9.9 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.4 | 17.0 | REACTOME SEMA4D IN SEMAPHORIN SIGNALING | Genes involved in Sema4D in semaphorin signaling |
| 0.3 | 5.8 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.3 | 6.7 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 7ALPHA HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 7alpha-hydroxycholesterol |
| 0.2 | 3.4 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.2 | 8.2 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.2 | 3.0 | REACTOME IRAK1 RECRUITS IKK COMPLEX | Genes involved in IRAK1 recruits IKK complex |
| 0.2 | 3.0 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.2 | 1.9 | REACTOME DOWNSTREAM SIGNAL TRANSDUCTION | Genes involved in Downstream signal transduction |
| 0.2 | 6.1 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.2 | 3.7 | REACTOME RIP MEDIATED NFKB ACTIVATION VIA DAI | Genes involved in RIP-mediated NFkB activation via DAI |
| 0.2 | 15.0 | REACTOME CELL JUNCTION ORGANIZATION | Genes involved in Cell junction organization |
| 0.2 | 2.7 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.1 | 2.5 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.1 | 2.4 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.1 | 3.0 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.1 | 5.1 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.1 | 7.8 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.1 | 2.9 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.1 | 5.6 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.1 | 1.7 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.1 | 0.9 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.1 | 2.9 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.1 | 4.1 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 4.2 | REACTOME THROMBIN SIGNALLING THROUGH PROTEINASE ACTIVATED RECEPTORS PARS | Genes involved in Thrombin signalling through proteinase activated receptors (PARs) |
| 0.1 | 1.9 | REACTOME REGULATION OF IFNG SIGNALING | Genes involved in Regulation of IFNG signaling |
| 0.1 | 4.2 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.1 | 1.3 | REACTOME G ALPHA1213 SIGNALLING EVENTS | Genes involved in G alpha (12/13) signalling events |
| 0.1 | 3.5 | REACTOME IRON UPTAKE AND TRANSPORT | Genes involved in Iron uptake and transport |
| 0.1 | 3.0 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.1 | 12.3 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.1 | 2.3 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 5.5 | REACTOME CELL SURFACE INTERACTIONS AT THE VASCULAR WALL | Genes involved in Cell surface interactions at the vascular wall |
| 0.0 | 1.0 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.0 | 0.9 | REACTOME SYNTHESIS OF PIPS AT THE GOLGI MEMBRANE | Genes involved in Synthesis of PIPs at the Golgi membrane |
| 0.0 | 1.1 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.0 | 0.6 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.0 | 6.6 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 0.8 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.0 | 0.9 | REACTOME FORMATION OF FIBRIN CLOT CLOTTING CASCADE | Genes involved in Formation of Fibrin Clot (Clotting Cascade) |
| 0.0 | 1.7 | REACTOME INTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Intrinsic Pathway for Apoptosis |
| 0.0 | 2.8 | REACTOME TRIGLYCERIDE BIOSYNTHESIS | Genes involved in Triglyceride Biosynthesis |
| 0.0 | 0.6 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GLP1 | Genes involved in Synthesis, Secretion, and Inactivation of Glucagon-like Peptide-1 (GLP-1) |
| 0.0 | 1.0 | REACTOME SPHINGOLIPID METABOLISM | Genes involved in Sphingolipid metabolism |
| 0.0 | 0.6 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.0 | 1.1 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.0 | 2.1 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 2.7 | REACTOME L1CAM INTERACTIONS | Genes involved in L1CAM interactions |
| 0.0 | 0.8 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 0.9 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.0 | 1.0 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |