Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
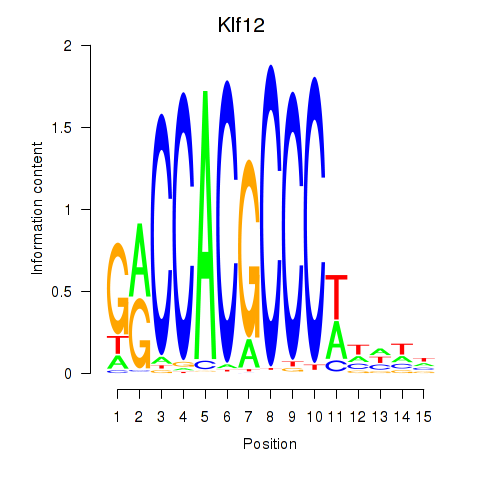
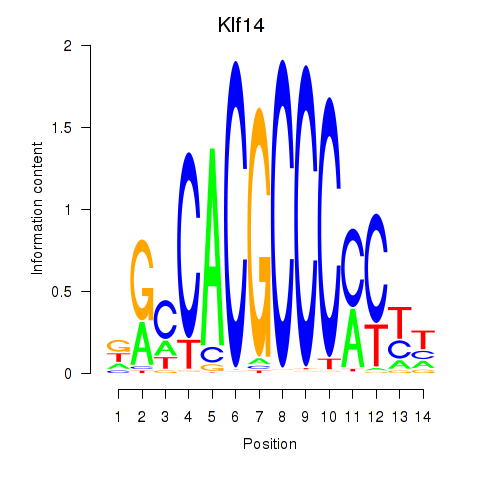
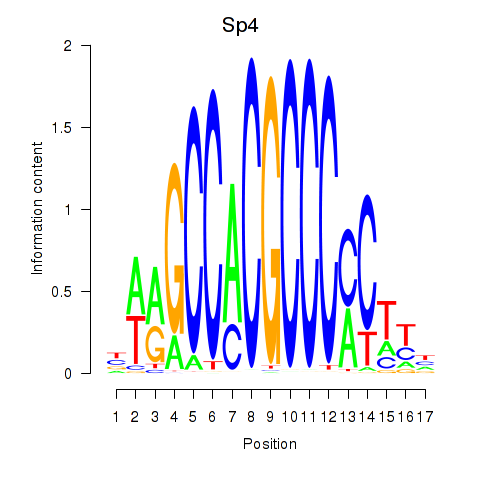
Results for Klf12_Klf14_Sp4
Z-value: 1.67
Transcription factors associated with Klf12_Klf14_Sp4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Klf12
|
ENSMUSG00000072294.6 | Klf12 |
|
Klf14
|
ENSMUSG00000073209.5 | Klf14 |
|
Sp4
|
ENSMUSG00000025323.11 | Sp4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Klf14 | mm39_v1_chr6_-_30936013_30936013 | -0.49 | 1.0e-05 | Click! |
| Klf12 | mm39_v1_chr14_-_100521888_100521934 | -0.47 | 3.8e-05 | Click! |
| Sp4 | mm39_v1_chr12_-_118265163_118265193 | 0.17 | 1.6e-01 | Click! |
Activity profile of Klf12_Klf14_Sp4 motif
Sorted Z-values of Klf12_Klf14_Sp4 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Klf12_Klf14_Sp4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_-_169973076 | 10.18 |
ENSMUST00000063710.13
|
Zfp217
|
zinc finger protein 217 |
| chr9_+_7764042 | 9.88 |
ENSMUST00000052865.16
|
Tmem123
|
transmembrane protein 123 |
| chr13_+_35925296 | 9.62 |
ENSMUST00000163595.3
|
Cdyl
|
chromodomain protein, Y chromosome-like |
| chr8_-_72011515 | 9.59 |
ENSMUST00000052072.8
|
Tmem221
|
transmembrane protein 221 |
| chr14_+_60872167 | 9.05 |
ENSMUST00000022566.14
ENSMUST00000159729.2 |
Spata13
|
spermatogenesis associated 13 |
| chr4_+_117109148 | 8.77 |
ENSMUST00000062824.12
|
Tmem53
|
transmembrane protein 53 |
| chr7_-_19483389 | 8.72 |
ENSMUST00000108450.5
ENSMUST00000075447.14 |
Nectin2
|
nectin cell adhesion molecule 2 |
| chr7_+_24310738 | 8.66 |
ENSMUST00000073325.6
|
Phldb3
|
pleckstrin homology like domain, family B, member 3 |
| chr4_+_117109204 | 8.46 |
ENSMUST00000125943.8
ENSMUST00000106434.8 |
Tmem53
|
transmembrane protein 53 |
| chr7_+_101467512 | 7.52 |
ENSMUST00000008090.11
|
Phox2a
|
paired-like homeobox 2a |
| chr7_+_46700349 | 7.36 |
ENSMUST00000010451.8
|
Tmem86a
|
transmembrane protein 86A |
| chr10_+_79855454 | 7.34 |
ENSMUST00000043311.7
|
Arhgap45
|
Rho GTPase activating protein 45 |
| chr4_+_137196080 | 7.18 |
ENSMUST00000030547.15
ENSMUST00000171332.2 |
Hspg2
|
perlecan (heparan sulfate proteoglycan 2) |
| chr11_-_52174129 | 6.98 |
ENSMUST00000109071.3
|
Tcf7
|
transcription factor 7, T cell specific |
| chr18_+_50112580 | 6.77 |
ENSMUST00000179937.2
|
Tnfaip8
|
tumor necrosis factor, alpha-induced protein 8 |
| chr5_-_144294854 | 6.73 |
ENSMUST00000055190.8
|
Baiap2l1
|
BAI1-associated protein 2-like 1 |
| chr8_+_117648474 | 6.70 |
ENSMUST00000034205.5
ENSMUST00000212775.2 |
Cenpn
|
centromere protein N |
| chr3_-_100396635 | 6.49 |
ENSMUST00000061455.9
|
Tent5c
|
terminal nucleotidyltransferase 5C |
| chr18_+_50112494 | 6.33 |
ENSMUST00000148989.3
|
Tnfaip8
|
tumor necrosis factor, alpha-induced protein 8 |
| chr7_-_30579686 | 6.24 |
ENSMUST00000188157.7
ENSMUST00000190753.2 ENSMUST00000186154.7 ENSMUST00000190617.7 |
Cd22
|
CD22 antigen |
| chr7_+_27173187 | 6.13 |
ENSMUST00000068641.8
|
Sertad3
|
SERTA domain containing 3 |
| chr11_-_70242850 | 6.00 |
ENSMUST00000019068.7
|
Alox15
|
arachidonate 15-lipoxygenase |
| chr3_+_68912302 | 5.99 |
ENSMUST00000136502.8
ENSMUST00000107803.7 |
Smc4
|
structural maintenance of chromosomes 4 |
| chr4_-_117109074 | 5.92 |
ENSMUST00000165128.9
|
Armh1
|
armadillo-like helical domain containing 1 |
| chr2_+_3714492 | 5.86 |
ENSMUST00000027965.11
|
Fam107b
|
family with sequence similarity 107, member B |
| chr8_+_106786190 | 5.85 |
ENSMUST00000109308.3
|
Nfatc3
|
nuclear factor of activated T cells, cytoplasmic, calcineurin dependent 3 |
| chr12_+_108758871 | 5.84 |
ENSMUST00000021692.9
|
Yy1
|
YY1 transcription factor |
| chr15_+_25622611 | 5.83 |
ENSMUST00000110457.8
ENSMUST00000137601.8 |
Myo10
|
myosin X |
| chr5_-_24235646 | 5.80 |
ENSMUST00000197617.5
ENSMUST00000030849.13 |
Fam126a
|
family with sequence similarity 126, member A |
| chr7_+_16515265 | 5.79 |
ENSMUST00000108496.9
|
Slc1a5
|
solute carrier family 1 (neutral amino acid transporter), member 5 |
| chr4_-_150093435 | 5.69 |
ENSMUST00000030830.4
|
H6pd
|
hexose-6-phosphate dehydrogenase (glucose 1-dehydrogenase) |
| chr19_+_6952580 | 5.69 |
ENSMUST00000237084.2
ENSMUST00000236218.2 ENSMUST00000237235.2 |
Ppp1r14b
|
protein phosphatase 1, regulatory inhibitor subunit 14B |
| chr2_+_3714515 | 5.59 |
ENSMUST00000115053.9
|
Fam107b
|
family with sequence similarity 107, member B |
| chr3_+_68912043 | 5.58 |
ENSMUST00000042901.15
|
Smc4
|
structural maintenance of chromosomes 4 |
| chr11_-_116515026 | 5.54 |
ENSMUST00000103029.10
|
Rhbdf2
|
rhomboid 5 homolog 2 |
| chr16_-_10603389 | 5.54 |
ENSMUST00000229866.2
ENSMUST00000038099.6 |
Socs1
|
suppressor of cytokine signaling 1 |
| chr5_+_139777263 | 5.41 |
ENSMUST00000018287.10
|
Mafk
|
v-maf musculoaponeurotic fibrosarcoma oncogene family, protein K (avian) |
| chr4_-_43499608 | 5.36 |
ENSMUST00000136005.3
ENSMUST00000054538.13 |
Arhgef39
|
Rho guanine nucleotide exchange factor (GEF) 39 |
| chr18_-_35795233 | 5.34 |
ENSMUST00000025209.12
ENSMUST00000096573.4 |
Spata24
|
spermatogenesis associated 24 |
| chr17_+_64907697 | 5.33 |
ENSMUST00000086723.10
|
Man2a1
|
mannosidase 2, alpha 1 |
| chr4_+_111577382 | 5.28 |
ENSMUST00000084354.4
|
Spata6
|
spermatogenesis associated 6 |
| chr7_-_80453033 | 5.27 |
ENSMUST00000167377.3
|
Iqgap1
|
IQ motif containing GTPase activating protein 1 |
| chr6_-_51989456 | 5.24 |
ENSMUST00000078214.8
ENSMUST00000204778.3 |
Skap2
|
src family associated phosphoprotein 2 |
| chr3_+_79536378 | 5.18 |
ENSMUST00000029388.10
|
4930579G24Rik
|
RIKEN cDNA 4930579G24 gene |
| chr8_-_93924426 | 5.16 |
ENSMUST00000034172.8
|
Ces1d
|
carboxylesterase 1D |
| chr7_-_127837154 | 5.15 |
ENSMUST00000078816.5
|
9130023H24Rik
|
RIKEN cDNA 9130023H24 gene |
| chr17_-_34247016 | 5.10 |
ENSMUST00000236627.2
ENSMUST00000237759.2 ENSMUST00000045467.14 ENSMUST00000114303.4 |
H2-Ke6
|
H2-K region expressed gene 6 |
| chr19_+_6952319 | 5.08 |
ENSMUST00000070850.8
|
Ppp1r14b
|
protein phosphatase 1, regulatory inhibitor subunit 14B |
| chr6_+_89620956 | 5.08 |
ENSMUST00000000828.14
ENSMUST00000101171.3 |
Txnrd3
|
thioredoxin reductase 3 |
| chr13_+_73911797 | 5.04 |
ENSMUST00000017900.9
|
Slc12a7
|
solute carrier family 12, member 7 |
| chr18_-_35795175 | 5.03 |
ENSMUST00000236574.2
ENSMUST00000236971.2 |
Spata24
|
spermatogenesis associated 24 |
| chr11_+_75358866 | 5.02 |
ENSMUST00000043598.14
ENSMUST00000108435.2 |
Tlcd2
|
TLC domain containing 2 |
| chr11_-_103158190 | 4.99 |
ENSMUST00000021324.3
|
Map3k14
|
mitogen-activated protein kinase kinase kinase 14 |
| chr8_-_47070201 | 4.88 |
ENSMUST00000210264.2
ENSMUST00000040468.16 ENSMUST00000209787.2 |
Primpol
|
primase and polymerase (DNA-directed) |
| chr1_-_183766195 | 4.88 |
ENSMUST00000050306.8
|
1700056E22Rik
|
RIKEN cDNA 1700056E22 gene |
| chr4_+_86792633 | 4.87 |
ENSMUST00000045224.14
ENSMUST00000084433.5 |
Acer2
|
alkaline ceramidase 2 |
| chr4_-_134495234 | 4.77 |
ENSMUST00000037828.8
|
Ldlrap1
|
low density lipoprotein receptor adaptor protein 1 |
| chr5_-_73025775 | 4.74 |
ENSMUST00000071944.13
ENSMUST00000073843.13 ENSMUST00000113594.8 |
Tec
|
tec protein tyrosine kinase |
| chr7_-_4755971 | 4.70 |
ENSMUST00000183971.8
ENSMUST00000182173.2 ENSMUST00000182738.8 ENSMUST00000182111.8 ENSMUST00000184143.8 ENSMUST00000182048.2 ENSMUST00000063324.14 |
Cox6b2
|
cytochrome c oxidase subunit 6B2 |
| chr17_+_31783708 | 4.67 |
ENSMUST00000097352.11
ENSMUST00000237248.2 ENSMUST00000235869.2 ENSMUST00000175806.9 |
Pknox1
|
Pbx/knotted 1 homeobox |
| chr19_+_6135013 | 4.66 |
ENSMUST00000025704.3
|
Cdca5
|
cell division cycle associated 5 |
| chr9_-_61854050 | 4.60 |
ENSMUST00000034815.9
|
Kif23
|
kinesin family member 23 |
| chr19_+_3438852 | 4.59 |
ENSMUST00000025840.16
ENSMUST00000151341.8 |
Tesmin
|
testis expressed metallothionein like |
| chr9_-_21709796 | 4.51 |
ENSMUST00000213691.2
|
Kank2
|
KN motif and ankyrin repeat domains 2 |
| chr6_+_120643323 | 4.46 |
ENSMUST00000112686.8
|
Cecr2
|
CECR2, histone acetyl-lysine reader |
| chr5_-_76139107 | 4.44 |
ENSMUST00000113516.2
|
Kdr
|
kinase insert domain protein receptor |
| chr10_-_80165257 | 4.43 |
ENSMUST00000020340.15
|
Pcsk4
|
proprotein convertase subtilisin/kexin type 4 |
| chr8_+_70275079 | 4.42 |
ENSMUST00000164890.8
ENSMUST00000034325.6 ENSMUST00000238452.2 |
Lpar2
|
lysophosphatidic acid receptor 2 |
| chr8_-_22675773 | 4.40 |
ENSMUST00000046916.9
|
Ckap2
|
cytoskeleton associated protein 2 |
| chr14_+_99283807 | 4.40 |
ENSMUST00000022656.8
|
Bora
|
bora, aurora kinase A activator |
| chr4_+_47353217 | 4.40 |
ENSMUST00000007757.15
|
Tgfbr1
|
transforming growth factor, beta receptor I |
| chr15_-_97806142 | 4.39 |
ENSMUST00000023119.15
|
Vdr
|
vitamin D (1,25-dihydroxyvitamin D3) receptor |
| chr2_-_164197987 | 4.33 |
ENSMUST00000165980.2
|
Slpi
|
secretory leukocyte peptidase inhibitor |
| chr7_+_65759198 | 4.33 |
ENSMUST00000036372.8
|
Chsy1
|
chondroitin sulfate synthase 1 |
| chr17_+_34808772 | 4.31 |
ENSMUST00000038244.15
|
Gpsm3
|
G-protein signalling modulator 3 (AGS3-like, C. elegans) |
| chr4_+_111577172 | 4.29 |
ENSMUST00000038868.14
ENSMUST00000070513.13 ENSMUST00000153746.8 |
Spata6
|
spermatogenesis associated 6 |
| chr17_-_45906428 | 4.25 |
ENSMUST00000171081.8
ENSMUST00000172301.8 ENSMUST00000167332.8 ENSMUST00000170488.8 ENSMUST00000167195.8 ENSMUST00000064889.13 ENSMUST00000051574.13 ENSMUST00000164217.8 |
Slc29a1
|
solute carrier family 29 (nucleoside transporters), member 1 |
| chr5_-_121974913 | 4.25 |
ENSMUST00000040308.14
ENSMUST00000086310.8 |
Sh2b3
|
SH2B adaptor protein 3 |
| chr6_+_91661034 | 4.24 |
ENSMUST00000032185.9
|
Slc6a6
|
solute carrier family 6 (neurotransmitter transporter, taurine), member 6 |
| chr17_-_57137898 | 4.23 |
ENSMUST00000233000.2
ENSMUST00000002444.15 ENSMUST00000086801.7 |
Rfx2
|
regulatory factor X, 2 (influences HLA class II expression) |
| chr6_+_49344673 | 4.21 |
ENSMUST00000060561.15
ENSMUST00000121903.2 ENSMUST00000134786.2 |
Fam221a
|
family with sequence similarity 221, member A |
| chr17_-_45906768 | 4.21 |
ENSMUST00000164618.8
ENSMUST00000097317.10 ENSMUST00000170113.8 |
Slc29a1
|
solute carrier family 29 (nucleoside transporters), member 1 |
| chr1_+_58035130 | 4.16 |
ENSMUST00000027202.9
|
Sgo2a
|
shugoshin 2A |
| chr8_+_46924074 | 4.16 |
ENSMUST00000034046.13
ENSMUST00000211644.2 |
Acsl1
|
acyl-CoA synthetase long-chain family member 1 |
| chr8_-_86159398 | 4.16 |
ENSMUST00000047749.7
|
4921524J17Rik
|
RIKEN cDNA 4921524J17 gene |
| chr10_-_128540847 | 4.15 |
ENSMUST00000026415.9
ENSMUST00000026416.15 |
Cdk2
|
cyclin-dependent kinase 2 |
| chr7_+_27147403 | 4.10 |
ENSMUST00000037399.16
ENSMUST00000108358.8 |
Blvrb
|
biliverdin reductase B (flavin reductase (NADPH)) |
| chr1_+_93731662 | 4.09 |
ENSMUST00000027505.13
|
Ing5
|
inhibitor of growth family, member 5 |
| chr18_+_65564009 | 4.07 |
ENSMUST00000224056.3
ENSMUST00000049248.7 |
Malt1
|
MALT1 paracaspase |
| chr13_+_17869727 | 4.06 |
ENSMUST00000221480.2
|
Mplkip
|
M-phase specific PLK1 intereacting protein |
| chr10_-_80374941 | 4.03 |
ENSMUST00000020383.6
|
Atp8b3
|
ATPase, class I, type 8B, member 3 |
| chr17_+_29020064 | 4.01 |
ENSMUST00000004985.11
|
Brpf3
|
bromodomain and PHD finger containing, 3 |
| chr17_-_68311073 | 4.00 |
ENSMUST00000024840.12
|
Arhgap28
|
Rho GTPase activating protein 28 |
| chr18_+_64473091 | 3.99 |
ENSMUST00000175965.10
|
Onecut2
|
one cut domain, family member 2 |
| chr5_+_145051090 | 3.99 |
ENSMUST00000196111.5
ENSMUST00000141602.2 |
Arpc1b
|
actin related protein 2/3 complex, subunit 1B |
| chr2_+_153334710 | 3.96 |
ENSMUST00000109783.2
|
4930404H24Rik
|
RIKEN cDNA 4930404H24 gene |
| chr1_-_191776840 | 3.95 |
ENSMUST00000195815.3
|
Traf5
|
TNF receptor-associated factor 5 |
| chr10_-_117681864 | 3.95 |
ENSMUST00000064667.9
|
Rap1b
|
RAS related protein 1b |
| chr3_-_142101418 | 3.95 |
ENSMUST00000029941.16
ENSMUST00000058626.9 |
Pdlim5
|
PDZ and LIM domain 5 |
| chr19_+_6276408 | 3.95 |
ENSMUST00000025698.14
ENSMUST00000113526.2 |
Gpha2
|
glycoprotein hormone alpha 2 |
| chr5_-_115257336 | 3.90 |
ENSMUST00000031524.11
|
Acads
|
acyl-Coenzyme A dehydrogenase, short chain |
| chr1_-_21149392 | 3.89 |
ENSMUST00000037998.6
|
Tram2
|
translocating chain-associating membrane protein 2 |
| chr15_+_81469538 | 3.87 |
ENSMUST00000068387.11
|
Ep300
|
E1A binding protein p300 |
| chr9_-_61854036 | 3.85 |
ENSMUST00000214295.2
|
Kif23
|
kinesin family member 23 |
| chr7_+_13012735 | 3.84 |
ENSMUST00000098814.13
ENSMUST00000146998.9 |
Lig1
|
ligase I, DNA, ATP-dependent |
| chr8_-_106140106 | 3.82 |
ENSMUST00000167294.8
ENSMUST00000063071.13 |
Kctd19
|
potassium channel tetramerisation domain containing 19 |
| chr17_+_34809132 | 3.81 |
ENSMUST00000173772.2
|
Gpsm3
|
G-protein signalling modulator 3 (AGS3-like, C. elegans) |
| chr2_+_31360219 | 3.80 |
ENSMUST00000102840.5
|
Ass1
|
argininosuccinate synthetase 1 |
| chr9_-_21671571 | 3.75 |
ENSMUST00000217382.2
ENSMUST00000214149.2 ENSMUST00000098942.6 ENSMUST00000216057.2 |
Spc24
|
SPC24, NDC80 kinetochore complex component, homolog (S. cerevisiae) |
| chr9_-_105022272 | 3.74 |
ENSMUST00000190661.2
ENSMUST00000035180.5 |
Nudt16l2
|
nudix (nucleoside diphosphate linked moiety X)-type motif 16-like 2 |
| chr17_+_57556449 | 3.73 |
ENSMUST00000224947.2
ENSMUST00000019631.11 ENSMUST00000224885.2 ENSMUST00000224152.2 |
Trip10
|
thyroid hormone receptor interactor 10 |
| chr17_+_6697511 | 3.71 |
ENSMUST00000179569.3
|
Dynlt1b
|
dynein light chain Tctex-type 1B |
| chr1_+_87254729 | 3.68 |
ENSMUST00000172794.8
ENSMUST00000164992.9 ENSMUST00000173173.8 |
Gigyf2
|
GRB10 interacting GYF protein 2 |
| chr1_-_9770434 | 3.61 |
ENSMUST00000088658.11
|
Mybl1
|
myeloblastosis oncogene-like 1 |
| chr16_-_90607251 | 3.61 |
ENSMUST00000140920.2
|
Urb1
|
URB1 ribosome biogenesis 1 homolog (S. cerevisiae) |
| chr11_-_55075855 | 3.60 |
ENSMUST00000039305.6
|
Slc36a2
|
solute carrier family 36 (proton/amino acid symporter), member 2 |
| chr15_-_99268311 | 3.59 |
ENSMUST00000081224.14
ENSMUST00000120633.2 ENSMUST00000088233.13 |
Fmnl3
|
formin-like 3 |
| chr11_-_106163753 | 3.56 |
ENSMUST00000021052.16
|
Smarcd2
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 2 |
| chr2_-_164198427 | 3.56 |
ENSMUST00000109367.10
|
Slpi
|
secretory leukocyte peptidase inhibitor |
| chr4_-_129590372 | 3.54 |
ENSMUST00000137640.3
|
Tmem39b
|
transmembrane protein 39b |
| chr6_+_91661074 | 3.54 |
ENSMUST00000205480.2
ENSMUST00000206545.2 |
Slc6a6
|
solute carrier family 6 (neurotransmitter transporter, taurine), member 6 |
| chr5_+_145051025 | 3.53 |
ENSMUST00000085679.13
|
Arpc1b
|
actin related protein 2/3 complex, subunit 1B |
| chr4_+_47353277 | 3.51 |
ENSMUST00000044234.14
|
Tgfbr1
|
transforming growth factor, beta receptor I |
| chr1_+_165591315 | 3.51 |
ENSMUST00000111432.10
|
Creg1
|
cellular repressor of E1A-stimulated genes 1 |
| chr8_+_57964921 | 3.49 |
ENSMUST00000067925.8
|
Hmgb2
|
high mobility group box 2 |
| chr8_+_57964956 | 3.48 |
ENSMUST00000210871.2
|
Hmgb2
|
high mobility group box 2 |
| chr4_+_52439237 | 3.48 |
ENSMUST00000102915.10
ENSMUST00000117280.8 ENSMUST00000142227.3 |
Smc2
|
structural maintenance of chromosomes 2 |
| chr12_-_69274936 | 3.47 |
ENSMUST00000221411.2
ENSMUST00000021359.7 |
Pole2
|
polymerase (DNA directed), epsilon 2 (p59 subunit) |
| chr15_+_73384407 | 3.47 |
ENSMUST00000043414.12
|
Dennd3
|
DENN/MADD domain containing 3 |
| chr3_+_121746862 | 3.44 |
ENSMUST00000037958.14
ENSMUST00000196904.5 |
Arhgap29
|
Rho GTPase activating protein 29 |
| chr9_+_69361348 | 3.40 |
ENSMUST00000134907.8
|
Anxa2
|
annexin A2 |
| chr10_+_79690452 | 3.39 |
ENSMUST00000165704.8
|
Ptbp1
|
polypyrimidine tract binding protein 1 |
| chr12_+_113115632 | 3.38 |
ENSMUST00000006523.12
ENSMUST00000200553.2 |
Crip1
|
cysteine-rich protein 1 (intestinal) |
| chr1_-_71142305 | 3.38 |
ENSMUST00000027393.8
|
Bard1
|
BRCA1 associated RING domain 1 |
| chr6_-_34887487 | 3.37 |
ENSMUST00000152488.8
|
Wdr91
|
WD repeat domain 91 |
| chr8_+_84682136 | 3.36 |
ENSMUST00000005607.9
|
Asf1b
|
anti-silencing function 1B histone chaperone |
| chr9_-_107971729 | 3.34 |
ENSMUST00000193254.6
|
Apeh
|
acylpeptide hydrolase |
| chr9_-_21710029 | 3.31 |
ENSMUST00000034717.7
|
Kank2
|
KN motif and ankyrin repeat domains 2 |
| chr2_-_35226981 | 3.30 |
ENSMUST00000028241.7
|
Stom
|
stomatin |
| chr9_-_107971640 | 3.30 |
ENSMUST00000081309.13
ENSMUST00000191985.2 |
Apeh
|
acylpeptide hydrolase |
| chr1_+_131890679 | 3.30 |
ENSMUST00000191034.2
ENSMUST00000177943.8 |
Gm29103
Slc45a3
|
predicted gene 29103 solute carrier family 45, member 3 |
| chr10_-_80512117 | 3.29 |
ENSMUST00000200082.5
|
Mknk2
|
MAP kinase-interacting serine/threonine kinase 2 |
| chr5_-_92822984 | 3.28 |
ENSMUST00000060930.10
|
Ccdc158
|
coiled-coil domain containing 158 |
| chr1_+_132243849 | 3.26 |
ENSMUST00000072177.14
ENSMUST00000082125.6 |
Nuak2
|
NUAK family, SNF1-like kinase, 2 |
| chr9_-_21829385 | 3.26 |
ENSMUST00000128442.2
ENSMUST00000119055.8 ENSMUST00000122211.8 ENSMUST00000115351.10 |
Rab3d
|
RAB3D, member RAS oncogene family |
| chr12_+_117807224 | 3.26 |
ENSMUST00000021592.16
|
Cdca7l
|
cell division cycle associated 7 like |
| chr17_+_35278011 | 3.26 |
ENSMUST00000007255.13
ENSMUST00000174493.8 |
Ddah2
|
dimethylarginine dimethylaminohydrolase 2 |
| chr8_+_120301974 | 3.25 |
ENSMUST00000093100.3
|
Dnaaf1
|
dynein, axonemal assembly factor 1 |
| chr11_-_102298141 | 3.25 |
ENSMUST00000149777.8
ENSMUST00000154001.8 |
Slc25a39
|
solute carrier family 25, member 39 |
| chr15_+_81900570 | 3.24 |
ENSMUST00000069530.13
ENSMUST00000168581.8 ENSMUST00000164779.2 |
Xrcc6
|
X-ray repair complementing defective repair in Chinese hamster cells 6 |
| chr11_-_78056347 | 3.23 |
ENSMUST00000017530.4
|
Traf4
|
TNF receptor associated factor 4 |
| chr13_-_120252337 | 3.21 |
ENSMUST00000177916.8
ENSMUST00000178271.3 ENSMUST00000223722.2 |
Zfp131
|
zinc finger protein 131 |
| chr10_+_26648473 | 3.21 |
ENSMUST00000039557.9
|
Arhgap18
|
Rho GTPase activating protein 18 |
| chr13_-_119545479 | 3.19 |
ENSMUST00000223268.2
|
Nnt
|
nicotinamide nucleotide transhydrogenase |
| chr16_+_91855158 | 3.18 |
ENSMUST00000047429.9
ENSMUST00000232677.2 ENSMUST00000113975.3 |
Mrps6
Gm49711
Slc5a3
|
mitochondrial ribosomal protein S6 predicted gene, 49711 solute carrier family 5 (inositol transporters), member 3 |
| chr10_+_58159288 | 3.18 |
ENSMUST00000020078.14
|
Lims1
|
LIM and senescent cell antigen-like domains 1 |
| chr2_+_3285240 | 3.17 |
ENSMUST00000081932.13
|
Nmt2
|
N-myristoyltransferase 2 |
| chr7_+_100142544 | 3.16 |
ENSMUST00000126534.8
ENSMUST00000207748.2 |
Ucp2
|
uncoupling protein 2 (mitochondrial, proton carrier) |
| chr8_+_46924206 | 3.15 |
ENSMUST00000135955.8
|
Acsl1
|
acyl-CoA synthetase long-chain family member 1 |
| chr15_+_78810919 | 3.15 |
ENSMUST00000089377.6
|
Lgals1
|
lectin, galactose binding, soluble 1 |
| chr19_+_38043446 | 3.13 |
ENSMUST00000236044.2
ENSMUST00000116506.8 ENSMUST00000096096.12 ENSMUST00000169673.3 |
Cep55
|
centrosomal protein 55 |
| chr2_+_109111083 | 3.13 |
ENSMUST00000028527.8
|
Kif18a
|
kinesin family member 18A |
| chr12_-_26506422 | 3.12 |
ENSMUST00000020970.10
|
Rsad2
|
radical S-adenosyl methionine domain containing 2 |
| chr7_-_19363280 | 3.11 |
ENSMUST00000094762.10
ENSMUST00000049912.15 ENSMUST00000098754.5 |
Relb
|
avian reticuloendotheliosis viral (v-rel) oncogene related B |
| chr5_-_92823111 | 3.08 |
ENSMUST00000151180.8
ENSMUST00000150359.2 |
Ccdc158
|
coiled-coil domain containing 158 |
| chr2_-_93292800 | 3.07 |
ENSMUST00000028644.11
|
Cd82
|
CD82 antigen |
| chr6_-_34887743 | 3.06 |
ENSMUST00000081214.12
|
Wdr91
|
WD repeat domain 91 |
| chr9_-_107167046 | 3.05 |
ENSMUST00000035194.8
|
Mapkapk3
|
mitogen-activated protein kinase-activated protein kinase 3 |
| chr2_+_26824040 | 3.05 |
ENSMUST00000153771.8
ENSMUST00000055406.9 |
Stkld1
|
serine/threonine kinase-like domain containing 1 |
| chr5_+_73071607 | 3.05 |
ENSMUST00000144843.8
|
Slain2
|
SLAIN motif family, member 2 |
| chr2_+_167380112 | 3.05 |
ENSMUST00000052631.8
|
Snai1
|
snail family zinc finger 1 |
| chr5_-_125418107 | 3.03 |
ENSMUST00000111390.8
ENSMUST00000086075.13 |
Scarb1
|
scavenger receptor class B, member 1 |
| chr12_+_111409087 | 3.03 |
ENSMUST00000109792.8
|
Tnfaip2
|
tumor necrosis factor, alpha-induced protein 2 |
| chr10_+_79690492 | 3.03 |
ENSMUST00000171599.8
ENSMUST00000095457.11 |
Ptbp1
|
polypyrimidine tract binding protein 1 |
| chr5_+_38377972 | 3.02 |
ENSMUST00000114106.8
|
Lyar
|
Ly1 antibody reactive clone |
| chr11_+_54988866 | 3.01 |
ENSMUST00000000608.8
|
Gm2a
|
GM2 ganglioside activator protein |
| chr5_+_38377814 | 3.01 |
ENSMUST00000087514.9
ENSMUST00000130721.8 ENSMUST00000123207.8 ENSMUST00000132190.8 ENSMUST00000202506.2 ENSMUST00000152066.8 ENSMUST00000155300.8 |
Lyar
|
Ly1 antibody reactive clone |
| chr9_-_110571645 | 3.01 |
ENSMUST00000006005.12
|
Pth1r
|
parathyroid hormone 1 receptor |
| chr8_+_120719177 | 2.99 |
ENSMUST00000132583.8
ENSMUST00000034282.16 |
Crispld2
|
cysteine-rich secretory protein LCCL domain containing 2 |
| chr2_+_34999497 | 2.99 |
ENSMUST00000028235.11
ENSMUST00000156933.8 ENSMUST00000028237.15 ENSMUST00000113032.8 |
Cntrl
|
centriolin |
| chr11_+_44508137 | 2.99 |
ENSMUST00000109268.2
ENSMUST00000101326.10 ENSMUST00000081265.12 |
Ebf1
|
early B cell factor 1 |
| chr14_+_24540745 | 2.99 |
ENSMUST00000112384.10
|
Rps24
|
ribosomal protein S24 |
| chr7_+_27147475 | 2.97 |
ENSMUST00000133750.8
|
Blvrb
|
biliverdin reductase B (flavin reductase (NADPH)) |
| chr1_-_191129223 | 2.96 |
ENSMUST00000067976.9
|
Ppp2r5a
|
protein phosphatase 2, regulatory subunit B', alpha |
| chr11_+_95603494 | 2.96 |
ENSMUST00000107717.8
|
Zfp652
|
zinc finger protein 652 |
| chr19_+_41970148 | 2.95 |
ENSMUST00000026170.3
|
Ubtd1
|
ubiquitin domain containing 1 |
| chr5_+_115644727 | 2.95 |
ENSMUST00000067268.15
ENSMUST00000086523.7 ENSMUST00000212819.3 |
Pxn
|
paxillin |
| chr10_+_86136236 | 2.94 |
ENSMUST00000020234.14
|
Timp3
|
tissue inhibitor of metalloproteinase 3 |
| chr7_-_44198157 | 2.94 |
ENSMUST00000145956.2
ENSMUST00000049343.15 |
Pold1
|
polymerase (DNA directed), delta 1, catalytic subunit |
| chr7_-_19138008 | 2.92 |
ENSMUST00000108458.10
|
Klc3
|
kinesin light chain 3 |
| chr5_+_77413282 | 2.90 |
ENSMUST00000080359.12
|
Rest
|
RE1-silencing transcription factor |
| chr10_-_80269436 | 2.89 |
ENSMUST00000105346.10
ENSMUST00000020377.13 ENSMUST00000105340.8 ENSMUST00000020379.13 ENSMUST00000105344.8 ENSMUST00000105342.8 ENSMUST00000105345.10 ENSMUST00000105343.8 |
Tcf3
|
transcription factor 3 |
| chr7_-_127307898 | 2.89 |
ENSMUST00000207019.2
|
Bcl7c
|
B cell CLL/lymphoma 7C |
| chr4_-_135749032 | 2.89 |
ENSMUST00000030427.6
|
Eloa
|
elongin A |
| chr14_-_70396859 | 2.88 |
ENSMUST00000058240.14
ENSMUST00000153871.2 |
9930012K11Rik
|
RIKEN cDNA 9930012K11 gene |
| chr5_+_112424549 | 2.88 |
ENSMUST00000031287.12
ENSMUST00000071455.7 ENSMUST00000239473.2 |
Tpst2
|
protein-tyrosine sulfotransferase 2 |
| chr4_-_132523657 | 2.88 |
ENSMUST00000045154.6
|
Themis2
|
thymocyte selection associated family member 2 |
| chr19_+_46294119 | 2.86 |
ENSMUST00000111881.4
|
Nfkb2
|
nuclear factor of kappa light polypeptide gene enhancer in B cells 2, p49/p100 |
| chr12_-_112792971 | 2.86 |
ENSMUST00000062092.7
ENSMUST00000220899.2 |
Cdca4
|
cell division cycle associated 4 |
| chr8_+_85598734 | 2.85 |
ENSMUST00000170296.2
ENSMUST00000136026.8 |
Syce2
|
synaptonemal complex central element protein 2 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.0 | 9.1 | GO:0019626 | short-chain fatty acid catabolic process(GO:0019626) |
| 2.6 | 7.9 | GO:1905223 | epicardium morphogenesis(GO:1905223) |
| 2.6 | 7.8 | GO:0070563 | negative regulation of vitamin D receptor signaling pathway(GO:0070563) |
| 2.6 | 7.8 | GO:0001762 | beta-alanine transport(GO:0001762) taurine transport(GO:0015734) |
| 2.5 | 7.5 | GO:0021558 | trochlear nerve development(GO:0021558) |
| 2.4 | 9.8 | GO:0039663 | fusion of virus membrane with host plasma membrane(GO:0019064) membrane fusion involved in viral entry into host cell(GO:0039663) multi-organism membrane fusion(GO:0044800) |
| 2.4 | 7.2 | GO:0006740 | NADPH regeneration(GO:0006740) |
| 2.2 | 6.6 | GO:1901074 | regulation of engulfment of apoptotic cell(GO:1901074) |
| 2.1 | 8.4 | GO:0000915 | assembly of actomyosin apparatus involved in cytokinesis(GO:0000912) actomyosin contractile ring assembly(GO:0000915) actomyosin contractile ring organization(GO:0044837) |
| 1.9 | 5.8 | GO:0071707 | immunoglobulin heavy chain V-D-J recombination(GO:0071707) |
| 1.9 | 5.8 | GO:0015825 | L-serine transport(GO:0015825) |
| 1.7 | 5.1 | GO:0018008 | N-terminal peptidyl-glycine N-myristoylation(GO:0018008) |
| 1.6 | 4.9 | GO:0090285 | negative regulation of protein glycosylation in Golgi(GO:0090285) |
| 1.6 | 15.9 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 1.5 | 4.6 | GO:0048627 | myoblast development(GO:0048627) |
| 1.5 | 3.0 | GO:0010899 | regulation of phosphatidylcholine catabolic process(GO:0010899) |
| 1.4 | 5.8 | GO:1901675 | negative regulation of histone H3-K27 acetylation(GO:1901675) |
| 1.3 | 2.7 | GO:0070318 | positive regulation of G0 to G1 transition(GO:0070318) |
| 1.3 | 5.3 | GO:0038156 | interleukin-3-mediated signaling pathway(GO:0038156) |
| 1.3 | 3.9 | GO:1990773 | regulation of matrix metallopeptidase secretion(GO:1904464) matrix metallopeptidase secretion(GO:1990773) |
| 1.3 | 3.9 | GO:0014737 | positive regulation of muscle atrophy(GO:0014737) |
| 1.3 | 3.9 | GO:0043973 | histone H3-K4 acetylation(GO:0043973) |
| 1.3 | 6.4 | GO:0042631 | cellular response to water deprivation(GO:0042631) |
| 1.2 | 3.7 | GO:1903778 | protein localization to vacuolar membrane(GO:1903778) |
| 1.2 | 3.7 | GO:1904826 | regulation of hydrogen sulfide biosynthetic process(GO:1904826) positive regulation of hydrogen sulfide biosynthetic process(GO:1904828) |
| 1.2 | 8.5 | GO:0015862 | uridine transport(GO:0015862) |
| 1.2 | 4.8 | GO:0090118 | receptor-mediated endocytosis of low-density lipoprotein particle involved in cholesterol transport(GO:0090118) |
| 1.2 | 2.4 | GO:0032911 | negative regulation of transforming growth factor beta1 production(GO:0032911) |
| 1.2 | 9.5 | GO:0031536 | positive regulation of exit from mitosis(GO:0031536) |
| 1.2 | 3.5 | GO:0006021 | inositol biosynthetic process(GO:0006021) |
| 1.2 | 3.5 | GO:0060821 | inactivation of X chromosome by DNA methylation(GO:0060821) |
| 1.1 | 4.5 | GO:0015772 | disaccharide transport(GO:0015766) sucrose transport(GO:0015770) oligosaccharide transport(GO:0015772) |
| 1.1 | 4.5 | GO:0060800 | regulation of cell differentiation involved in embryonic placenta development(GO:0060800) |
| 1.1 | 1.1 | GO:0060854 | patterning of lymph vessels(GO:0060854) |
| 1.1 | 4.4 | GO:0038183 | bile acid signaling pathway(GO:0038183) |
| 1.1 | 3.2 | GO:0038163 | thrombopoietin-mediated signaling pathway(GO:0038163) |
| 1.1 | 4.3 | GO:0097680 | double-strand break repair via classical nonhomologous end joining(GO:0097680) |
| 1.1 | 7.5 | GO:0048254 | snoRNA localization(GO:0048254) |
| 1.1 | 5.3 | GO:0002905 | mature B cell apoptotic process(GO:0002901) regulation of mature B cell apoptotic process(GO:0002905) negative regulation of mature B cell apoptotic process(GO:0002906) |
| 1.1 | 5.3 | GO:0019516 | lactate oxidation(GO:0019516) |
| 1.1 | 3.2 | GO:0034117 | erythrocyte aggregation(GO:0034117) regulation of erythrocyte aggregation(GO:0034118) |
| 1.0 | 4.1 | GO:0032298 | positive regulation of DNA-dependent DNA replication initiation(GO:0032298) |
| 1.0 | 1.0 | GO:1903753 | negative regulation of p38MAPK cascade(GO:1903753) |
| 1.0 | 1.0 | GO:0061723 | glycophagy(GO:0061723) |
| 1.0 | 6.9 | GO:0034616 | response to laminar fluid shear stress(GO:0034616) cellular response to laminar fluid shear stress(GO:0071499) |
| 1.0 | 2.9 | GO:0045004 | DNA replication proofreading(GO:0045004) |
| 1.0 | 2.9 | GO:0036275 | response to 5-fluorouracil(GO:0036275) |
| 1.0 | 1.0 | GO:0072717 | cellular response to actinomycin D(GO:0072717) |
| 1.0 | 4.9 | GO:0001923 | B-1 B cell differentiation(GO:0001923) |
| 1.0 | 2.9 | GO:0015680 | intracellular copper ion transport(GO:0015680) |
| 1.0 | 4.8 | GO:0072733 | response to staurosporine(GO:0072733) cellular response to staurosporine(GO:0072734) |
| 1.0 | 2.9 | GO:0006478 | peptidyl-tyrosine sulfation(GO:0006478) |
| 1.0 | 7.6 | GO:0033015 | porphyrin-containing compound catabolic process(GO:0006787) tetrapyrrole catabolic process(GO:0033015) heme catabolic process(GO:0042167) pigment catabolic process(GO:0046149) |
| 0.9 | 5.7 | GO:0042758 | long-chain fatty acid catabolic process(GO:0042758) |
| 0.9 | 10.3 | GO:0034723 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.9 | 2.7 | GO:0071579 | regulation of zinc ion transport(GO:0071579) |
| 0.9 | 9.1 | GO:0045654 | positive regulation of megakaryocyte differentiation(GO:0045654) |
| 0.9 | 9.0 | GO:0002568 | somatic diversification of T cell receptor genes(GO:0002568) somatic recombination of T cell receptor gene segments(GO:0002681) T cell receptor V(D)J recombination(GO:0033153) |
| 0.9 | 2.7 | GO:0006550 | isoleucine catabolic process(GO:0006550) |
| 0.9 | 0.9 | GO:2000286 | receptor internalization involved in canonical Wnt signaling pathway(GO:2000286) |
| 0.9 | 3.5 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 0.9 | 5.3 | GO:0032804 | negative regulation of low-density lipoprotein particle receptor catabolic process(GO:0032804) |
| 0.9 | 2.6 | GO:0038018 | Wnt receptor catabolic process(GO:0038018) |
| 0.9 | 3.5 | GO:0036233 | glycine import(GO:0036233) |
| 0.9 | 4.3 | GO:2000048 | negative regulation of cell-cell adhesion mediated by cadherin(GO:2000048) |
| 0.8 | 4.1 | GO:0015888 | thiamine transport(GO:0015888) |
| 0.8 | 0.8 | GO:0032329 | serine transport(GO:0032329) |
| 0.8 | 1.6 | GO:0006842 | tricarboxylic acid transport(GO:0006842) citrate transport(GO:0015746) |
| 0.8 | 3.2 | GO:1905077 | negative regulation of interleukin-17 secretion(GO:1905077) |
| 0.8 | 3.2 | GO:0060697 | positive regulation of phospholipid catabolic process(GO:0060697) |
| 0.8 | 2.4 | GO:0036446 | myofibroblast differentiation(GO:0036446) cardiac fibroblast cell differentiation(GO:0060935) cardiac fibroblast cell development(GO:0060936) epicardium-derived cardiac fibroblast cell differentiation(GO:0060938) epicardium-derived cardiac fibroblast cell development(GO:0060939) regulation of myofibroblast differentiation(GO:1904760) negative regulation of myofibroblast differentiation(GO:1904761) negative regulation of vascular smooth muscle cell differentiation(GO:1905064) |
| 0.8 | 3.1 | GO:0034165 | positive regulation of toll-like receptor 9 signaling pathway(GO:0034165) |
| 0.8 | 3.9 | GO:0035519 | protein K29-linked ubiquitination(GO:0035519) |
| 0.8 | 6.2 | GO:0046600 | negative regulation of centriole replication(GO:0046600) |
| 0.8 | 4.6 | GO:0006933 | negative regulation of cell adhesion involved in substrate-bound cell migration(GO:0006933) |
| 0.8 | 3.8 | GO:0000973 | posttranscriptional tethering of RNA polymerase II gene DNA at nuclear periphery(GO:0000973) |
| 0.7 | 2.9 | GO:0097298 | regulation of nucleus size(GO:0097298) |
| 0.7 | 2.2 | GO:0000715 | nucleotide-excision repair, DNA damage recognition(GO:0000715) |
| 0.7 | 2.9 | GO:2000705 | regulation of dense core granule biogenesis(GO:2000705) |
| 0.7 | 2.9 | GO:1903984 | positive regulation of TRAIL-activated apoptotic signaling pathway(GO:1903984) |
| 0.7 | 1.4 | GO:0015675 | nickel cation transport(GO:0015675) |
| 0.7 | 0.7 | GO:0048861 | leukemia inhibitory factor signaling pathway(GO:0048861) |
| 0.7 | 2.9 | GO:0050822 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-dependent(GO:0002479) peptide stabilization(GO:0050822) peptide antigen stabilization(GO:0050823) |
| 0.7 | 2.9 | GO:0060398 | regulation of growth hormone receptor signaling pathway(GO:0060398) |
| 0.7 | 2.9 | GO:0002266 | follicular dendritic cell activation(GO:0002266) follicular dendritic cell differentiation(GO:0002268) |
| 0.7 | 3.5 | GO:0042276 | error-prone translesion synthesis(GO:0042276) |
| 0.7 | 1.4 | GO:0071211 | protein targeting to vacuole involved in autophagy(GO:0071211) |
| 0.7 | 5.5 | GO:0010533 | regulation of activation of Janus kinase activity(GO:0010533) |
| 0.7 | 3.4 | GO:1902728 | positive regulation of skeletal muscle satellite cell proliferation(GO:1902724) positive regulation of growth factor dependent skeletal muscle satellite cell proliferation(GO:1902728) |
| 0.7 | 4.1 | GO:0030423 | targeting of mRNA for destruction involved in RNA interference(GO:0030423) |
| 0.7 | 4.1 | GO:0032954 | regulation of cytokinetic process(GO:0032954) regulation of mitotic cytokinetic process(GO:1903436) positive regulation of mitotic cytokinetic process(GO:1903438) positive regulation of mitotic cytokinesis(GO:1903490) |
| 0.7 | 3.4 | GO:0061086 | negative regulation of histone H3-K27 methylation(GO:0061086) |
| 0.7 | 2.7 | GO:0035938 | estradiol secretion(GO:0035938) regulation of estradiol secretion(GO:2000864) |
| 0.7 | 5.3 | GO:0048597 | post-embryonic camera-type eye morphogenesis(GO:0048597) |
| 0.7 | 4.6 | GO:0051697 | protein delipidation(GO:0051697) |
| 0.7 | 2.0 | GO:1904582 | positive regulation of intracellular mRNA localization(GO:1904582) |
| 0.7 | 2.6 | GO:1901536 | negative regulation of DNA demethylation(GO:1901536) |
| 0.7 | 3.3 | GO:0098763 | mitotic cell cycle phase(GO:0098763) |
| 0.7 | 2.6 | GO:0046351 | disaccharide biosynthetic process(GO:0046351) |
| 0.6 | 0.6 | GO:0035993 | deltoid tuberosity development(GO:0035993) |
| 0.6 | 6.4 | GO:0044806 | G-quadruplex DNA unwinding(GO:0044806) |
| 0.6 | 5.1 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.6 | 4.4 | GO:0090521 | glomerular visceral epithelial cell migration(GO:0090521) |
| 0.6 | 6.3 | GO:0045002 | double-strand break repair via single-strand annealing(GO:0045002) |
| 0.6 | 5.0 | GO:0033567 | DNA replication, Okazaki fragment processing(GO:0033567) |
| 0.6 | 3.1 | GO:1901165 | positive regulation of trophoblast cell migration(GO:1901165) |
| 0.6 | 4.3 | GO:0035469 | determination of pancreatic left/right asymmetry(GO:0035469) |
| 0.6 | 2.5 | GO:0036166 | phenotypic switching(GO:0036166) |
| 0.6 | 1.9 | GO:0045404 | interleukin-10 biosynthetic process(GO:0042091) regulation of interleukin-10 biosynthetic process(GO:0045074) positive regulation of interleukin-4 biosynthetic process(GO:0045404) |
| 0.6 | 4.3 | GO:0006741 | NADP biosynthetic process(GO:0006741) |
| 0.6 | 2.5 | GO:0090327 | negative regulation of locomotion involved in locomotory behavior(GO:0090327) |
| 0.6 | 1.8 | GO:0006097 | glyoxylate cycle(GO:0006097) |
| 0.6 | 3.6 | GO:0019276 | UDP-N-acetylgalactosamine metabolic process(GO:0019276) |
| 0.6 | 2.4 | GO:0090187 | positive regulation of pancreatic juice secretion(GO:0090187) |
| 0.6 | 1.8 | GO:0006391 | transcription initiation from mitochondrial promoter(GO:0006391) |
| 0.6 | 2.3 | GO:0018076 | N-terminal peptidyl-lysine acetylation(GO:0018076) |
| 0.6 | 1.7 | GO:0098974 | postsynaptic actin cytoskeleton organization(GO:0098974) |
| 0.6 | 5.2 | GO:0007144 | female meiosis I(GO:0007144) |
| 0.6 | 1.7 | GO:1902263 | apoptotic process involved in embryonic digit morphogenesis(GO:1902263) |
| 0.6 | 0.6 | GO:0098705 | plasma membrane copper ion transport(GO:0015679) copper ion import across plasma membrane(GO:0098705) copper ion import into cell(GO:1902861) |
| 0.6 | 4.6 | GO:0009313 | oligosaccharide catabolic process(GO:0009313) |
| 0.6 | 4.0 | GO:0001561 | fatty acid alpha-oxidation(GO:0001561) |
| 0.6 | 2.8 | GO:0033629 | negative regulation of cell adhesion mediated by integrin(GO:0033629) |
| 0.6 | 2.3 | GO:0031662 | regulation of cyclin-dependent protein serine/threonine kinase activity involved in G2/M transition of mitotic cell cycle(GO:0031660) positive regulation of cyclin-dependent protein serine/threonine kinase activity involved in G2/M transition of mitotic cell cycle(GO:0031662) |
| 0.6 | 0.6 | GO:1900110 | negative regulation of histone H3-K9 dimethylation(GO:1900110) |
| 0.6 | 3.9 | GO:0034093 | positive regulation of maintenance of sister chromatid cohesion(GO:0034093) |
| 0.6 | 6.2 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.6 | 1.7 | GO:2001200 | positive regulation of dendritic cell differentiation(GO:2001200) |
| 0.6 | 1.7 | GO:0060448 | dichotomous subdivision of terminal units involved in lung branching(GO:0060448) |
| 0.6 | 2.2 | GO:0040030 | regulation of molecular function, epigenetic(GO:0040030) |
| 0.5 | 2.7 | GO:0007066 | female meiosis sister chromatid cohesion(GO:0007066) |
| 0.5 | 1.6 | GO:0033082 | regulation of extrathymic T cell differentiation(GO:0033082) |
| 0.5 | 3.2 | GO:1904627 | response to phorbol 13-acetate 12-myristate(GO:1904627) cellular response to phorbol 13-acetate 12-myristate(GO:1904628) |
| 0.5 | 3.7 | GO:0042538 | hyperosmotic salinity response(GO:0042538) |
| 0.5 | 1.1 | GO:0018894 | dibenzo-p-dioxin metabolic process(GO:0018894) |
| 0.5 | 3.2 | GO:0034628 | nicotinamide nucleotide biosynthetic process from aspartate(GO:0019355) 'de novo' NAD biosynthetic process(GO:0034627) 'de novo' NAD biosynthetic process from aspartate(GO:0034628) |
| 0.5 | 1.6 | GO:0046125 | pyrimidine deoxyribonucleoside metabolic process(GO:0046125) |
| 0.5 | 1.6 | GO:1903334 | positive regulation of protein folding(GO:1903334) |
| 0.5 | 7.3 | GO:0044539 | long-chain fatty acid import(GO:0044539) |
| 0.5 | 1.6 | GO:0048320 | axial mesoderm formation(GO:0048320) |
| 0.5 | 2.1 | GO:0006269 | DNA replication, synthesis of RNA primer(GO:0006269) |
| 0.5 | 0.5 | GO:0070171 | negative regulation of tooth mineralization(GO:0070171) |
| 0.5 | 11.4 | GO:0045793 | positive regulation of cell size(GO:0045793) |
| 0.5 | 1.5 | GO:0046203 | spermidine catabolic process(GO:0046203) |
| 0.5 | 10.7 | GO:0070935 | 3'-UTR-mediated mRNA stabilization(GO:0070935) |
| 0.5 | 2.5 | GO:0009052 | pentose-phosphate shunt, non-oxidative branch(GO:0009052) |
| 0.5 | 4.5 | GO:2000301 | negative regulation of synaptic vesicle exocytosis(GO:2000301) |
| 0.5 | 2.0 | GO:0061085 | regulation of histone H3-K27 methylation(GO:0061085) positive regulation of histone H3-K27 methylation(GO:0061087) |
| 0.5 | 2.0 | GO:0055048 | regulation of spindle elongation(GO:0032887) regulation of mitotic spindle elongation(GO:0032888) anastral spindle assembly(GO:0055048) protein localization to spindle pole body(GO:0071988) regulation of protein localization to spindle pole body(GO:1902363) positive regulation of protein localization to spindle pole body(GO:1902365) positive regulation of mitotic spindle elongation(GO:1902846) |
| 0.5 | 2.0 | GO:0003050 | regulation of systemic arterial blood pressure by atrial natriuretic peptide(GO:0003050) |
| 0.5 | 3.4 | GO:0085020 | protein K6-linked ubiquitination(GO:0085020) |
| 0.5 | 2.9 | GO:0010836 | negative regulation of protein ADP-ribosylation(GO:0010836) |
| 0.5 | 4.9 | GO:0032485 | regulation of Ral protein signal transduction(GO:0032485) |
| 0.5 | 2.4 | GO:0046121 | deoxyribonucleoside catabolic process(GO:0046121) |
| 0.5 | 8.7 | GO:0036120 | cellular response to platelet-derived growth factor stimulus(GO:0036120) |
| 0.5 | 7.2 | GO:0001842 | neural fold formation(GO:0001842) |
| 0.5 | 3.8 | GO:0070294 | renal sodium ion absorption(GO:0070294) |
| 0.5 | 2.4 | GO:0007089 | traversing start control point of mitotic cell cycle(GO:0007089) |
| 0.5 | 1.9 | GO:0071608 | macrophage inflammatory protein-1 alpha production(GO:0071608) |
| 0.5 | 1.4 | GO:0048817 | negative regulation of hair follicle maturation(GO:0048817) |
| 0.5 | 2.3 | GO:1904502 | regulation of lipophagy(GO:1904502) positive regulation of lipophagy(GO:1904504) |
| 0.5 | 2.8 | GO:0061198 | fungiform papilla formation(GO:0061198) |
| 0.5 | 19.3 | GO:0000305 | response to oxygen radical(GO:0000305) |
| 0.5 | 1.8 | GO:0021526 | medial motor column neuron differentiation(GO:0021526) |
| 0.5 | 2.3 | GO:0031509 | telomeric heterochromatin assembly(GO:0031509) negative regulation of chromosome condensation(GO:1902340) |
| 0.5 | 4.5 | GO:0045048 | protein insertion into ER membrane(GO:0045048) |
| 0.5 | 1.4 | GO:1901355 | response to rapamycin(GO:1901355) |
| 0.4 | 3.1 | GO:0006177 | GMP biosynthetic process(GO:0006177) |
| 0.4 | 0.4 | GO:0031125 | rRNA 3'-end processing(GO:0031125) |
| 0.4 | 0.9 | GO:1903588 | negative regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903588) |
| 0.4 | 2.2 | GO:0010826 | negative regulation of centrosome duplication(GO:0010826) negative regulation of centrosome cycle(GO:0046606) |
| 0.4 | 4.9 | GO:0019985 | translesion synthesis(GO:0019985) |
| 0.4 | 6.2 | GO:0006098 | pentose-phosphate shunt(GO:0006098) |
| 0.4 | 3.5 | GO:2000051 | negative regulation of non-canonical Wnt signaling pathway(GO:2000051) |
| 0.4 | 1.3 | GO:0002380 | immunoglobulin secretion involved in immune response(GO:0002380) |
| 0.4 | 0.9 | GO:0009826 | unidimensional cell growth(GO:0009826) |
| 0.4 | 1.3 | GO:0030174 | regulation of DNA-dependent DNA replication initiation(GO:0030174) |
| 0.4 | 1.3 | GO:0042986 | positive regulation of amyloid precursor protein biosynthetic process(GO:0042986) |
| 0.4 | 8.1 | GO:1900017 | positive regulation of cytokine production involved in inflammatory response(GO:1900017) |
| 0.4 | 2.1 | GO:0000117 | regulation of transcription involved in G2/M transition of mitotic cell cycle(GO:0000117) |
| 0.4 | 6.4 | GO:2000035 | regulation of stem cell division(GO:2000035) |
| 0.4 | 1.3 | GO:0036245 | cellular response to menadione(GO:0036245) |
| 0.4 | 3.8 | GO:0060718 | chorionic trophoblast cell differentiation(GO:0060718) |
| 0.4 | 1.7 | GO:0044339 | canonical Wnt signaling pathway involved in osteoblast differentiation(GO:0044339) |
| 0.4 | 1.7 | GO:0034285 | response to sucrose(GO:0009744) response to disaccharide(GO:0034285) |
| 0.4 | 1.7 | GO:2000383 | regulation of ectoderm development(GO:2000383) negative regulation of ectoderm development(GO:2000384) |
| 0.4 | 3.3 | GO:1900748 | positive regulation of vascular endothelial growth factor signaling pathway(GO:1900748) |
| 0.4 | 1.7 | GO:0015744 | succinate transport(GO:0015744) |
| 0.4 | 2.9 | GO:0002326 | B cell lineage commitment(GO:0002326) |
| 0.4 | 2.9 | GO:0010519 | negative regulation of phospholipase activity(GO:0010519) |
| 0.4 | 0.4 | GO:0015864 | pyrimidine nucleoside transport(GO:0015864) |
| 0.4 | 0.4 | GO:0032241 | positive regulation of nucleobase-containing compound transport(GO:0032241) |
| 0.4 | 1.6 | GO:1901187 | regulation of ephrin receptor signaling pathway(GO:1901187) |
| 0.4 | 0.8 | GO:0035546 | interferon-beta secretion(GO:0035546) regulation of interferon-beta secretion(GO:0035547) positive regulation of interferon-beta secretion(GO:0035549) |
| 0.4 | 6.1 | GO:0006517 | protein deglycosylation(GO:0006517) |
| 0.4 | 10.6 | GO:0044458 | motile cilium assembly(GO:0044458) |
| 0.4 | 1.6 | GO:0072757 | cellular response to camptothecin(GO:0072757) |
| 0.4 | 3.6 | GO:0060903 | positive regulation of meiosis I(GO:0060903) |
| 0.4 | 1.6 | GO:1904425 | negative regulation of GTP binding(GO:1904425) |
| 0.4 | 2.8 | GO:0033277 | abortive mitotic cell cycle(GO:0033277) |
| 0.4 | 2.8 | GO:0046784 | viral mRNA export from host cell nucleus(GO:0046784) |
| 0.4 | 2.0 | GO:0006548 | histamine metabolic process(GO:0001692) histidine catabolic process(GO:0006548) |
| 0.4 | 7.8 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.4 | 1.2 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.4 | 1.9 | GO:0036493 | positive regulation of translation in response to endoplasmic reticulum stress(GO:0036493) |
| 0.4 | 1.9 | GO:0070966 | nuclear-transcribed mRNA catabolic process, no-go decay(GO:0070966) |
| 0.4 | 2.7 | GO:0036123 | histone H3-K9 dimethylation(GO:0036123) |
| 0.4 | 1.9 | GO:0061314 | Notch signaling involved in heart development(GO:0061314) |
| 0.4 | 0.7 | GO:1903921 | protein processing in phagocytic vesicle(GO:1900756) regulation of protein processing in phagocytic vesicle(GO:1903921) positive regulation of protein processing in phagocytic vesicle(GO:1903923) |
| 0.4 | 1.5 | GO:0071651 | positive regulation of chemokine (C-C motif) ligand 5 production(GO:0071651) |
| 0.4 | 0.4 | GO:0002424 | T cell mediated immune response to tumor cell(GO:0002424) regulation of T cell mediated immune response to tumor cell(GO:0002840) |
| 0.4 | 3.3 | GO:0044829 | positive regulation by host of viral genome replication(GO:0044829) |
| 0.4 | 4.0 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.4 | 0.7 | GO:0044340 | canonical Wnt signaling pathway involved in regulation of cell proliferation(GO:0044340) |
| 0.4 | 1.1 | GO:0021589 | hindbrain structural organization(GO:0021577) cerebellum structural organization(GO:0021589) |
| 0.4 | 1.4 | GO:0007037 | vacuolar phosphate transport(GO:0007037) positive regulation of parathyroid hormone secretion(GO:2000830) |
| 0.4 | 0.7 | GO:0010481 | epidermal cell division(GO:0010481) regulation of epidermal cell division(GO:0010482) |
| 0.4 | 1.1 | GO:0016240 | autophagosome docking(GO:0016240) |
| 0.4 | 0.4 | GO:0003383 | apical constriction(GO:0003383) |
| 0.4 | 2.1 | GO:0048633 | positive regulation of skeletal muscle tissue growth(GO:0048633) |
| 0.4 | 1.1 | GO:0021682 | nerve maturation(GO:0021682) |
| 0.3 | 19.1 | GO:0007339 | binding of sperm to zona pellucida(GO:0007339) |
| 0.3 | 1.0 | GO:1902220 | positive regulation of intrinsic apoptotic signaling pathway in response to osmotic stress(GO:1902220) |
| 0.3 | 2.1 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 0.3 | 1.0 | GO:1904742 | regulation of telomeric DNA binding(GO:1904742) |
| 0.3 | 3.1 | GO:0060666 | dichotomous subdivision of terminal units involved in salivary gland branching(GO:0060666) |
| 0.3 | 10.2 | GO:0032060 | bleb assembly(GO:0032060) |
| 0.3 | 1.0 | GO:0060729 | intestinal epithelial structure maintenance(GO:0060729) |
| 0.3 | 2.4 | GO:0003415 | chondrocyte hypertrophy(GO:0003415) |
| 0.3 | 1.0 | GO:0008358 | oocyte construction(GO:0007308) oocyte axis specification(GO:0007309) oocyte anterior/posterior axis specification(GO:0007314) pole plasm assembly(GO:0007315) maternal determination of anterior/posterior axis, embryo(GO:0008358) P granule organization(GO:0030719) |
| 0.3 | 1.7 | GO:0038032 | termination of G-protein coupled receptor signaling pathway(GO:0038032) |
| 0.3 | 2.3 | GO:0097503 | sialylation(GO:0097503) |
| 0.3 | 1.3 | GO:0010286 | heat acclimation(GO:0010286) |
| 0.3 | 1.3 | GO:1904158 | axonemal central apparatus assembly(GO:1904158) |
| 0.3 | 1.0 | GO:0010182 | carbohydrate mediated signaling(GO:0009756) hexose mediated signaling(GO:0009757) sugar mediated signaling pathway(GO:0010182) glucose mediated signaling pathway(GO:0010255) |
| 0.3 | 1.0 | GO:2000451 | positive regulation of CD8-positive, alpha-beta T cell extravasation(GO:2000451) |
| 0.3 | 4.6 | GO:1904776 | regulation of protein localization to cell cortex(GO:1904776) positive regulation of protein localization to cell cortex(GO:1904778) |
| 0.3 | 1.6 | GO:0097029 | mature conventional dendritic cell differentiation(GO:0097029) |
| 0.3 | 11.5 | GO:0019731 | antibacterial humoral response(GO:0019731) |
| 0.3 | 1.0 | GO:1990166 | protein localization to site of double-strand break(GO:1990166) |
| 0.3 | 2.8 | GO:0090070 | positive regulation of ribosome biogenesis(GO:0090070) positive regulation of rRNA processing(GO:2000234) |
| 0.3 | 1.3 | GO:0006297 | nucleotide-excision repair, DNA gap filling(GO:0006297) |
| 0.3 | 2.5 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 0.3 | 1.6 | GO:0009597 | detection of virus(GO:0009597) |
| 0.3 | 1.9 | GO:0071698 | olfactory placode formation(GO:0030910) olfactory placode development(GO:0071698) olfactory placode morphogenesis(GO:0071699) |
| 0.3 | 1.6 | GO:0007084 | mitotic nuclear envelope reassembly(GO:0007084) |
| 0.3 | 6.2 | GO:0051923 | sulfation(GO:0051923) |
| 0.3 | 7.4 | GO:0043011 | myeloid dendritic cell differentiation(GO:0043011) |
| 0.3 | 1.2 | GO:1903031 | regulation of microtubule plus-end binding(GO:1903031) positive regulation of microtubule plus-end binding(GO:1903033) |
| 0.3 | 0.9 | GO:0034729 | histone H3-K79 methylation(GO:0034729) |
| 0.3 | 3.4 | GO:0006398 | mRNA 3'-end processing by stem-loop binding and cleavage(GO:0006398) |
| 0.3 | 3.1 | GO:0044351 | macropinocytosis(GO:0044351) |
| 0.3 | 1.2 | GO:1902525 | regulation of protein monoubiquitination(GO:1902525) |
| 0.3 | 1.8 | GO:0034154 | toll-like receptor 7 signaling pathway(GO:0034154) |
| 0.3 | 1.8 | GO:1904354 | negative regulation of telomere capping(GO:1904354) |
| 0.3 | 1.5 | GO:0035616 | histone H2B conserved C-terminal lysine deubiquitination(GO:0035616) |
| 0.3 | 0.9 | GO:0018900 | dichloromethane metabolic process(GO:0018900) |
| 0.3 | 0.6 | GO:0010847 | regulation of chromatin assembly(GO:0010847) |
| 0.3 | 0.9 | GO:0046901 | tetrahydrofolylpolyglutamate biosynthetic process(GO:0046901) |
| 0.3 | 0.9 | GO:0032242 | regulation of nucleoside transport(GO:0032242) |
| 0.3 | 1.2 | GO:0072355 | histone H3-T3 phosphorylation(GO:0072355) |
| 0.3 | 7.9 | GO:0034508 | centromere complex assembly(GO:0034508) |
| 0.3 | 1.2 | GO:1902268 | negative regulation of polyamine transmembrane transport(GO:1902268) |
| 0.3 | 4.1 | GO:0034501 | protein localization to kinetochore(GO:0034501) |
| 0.3 | 3.2 | GO:2000346 | negative regulation of hepatocyte proliferation(GO:2000346) |
| 0.3 | 1.2 | GO:1902047 | polyamine transmembrane transport(GO:1902047) regulation of polyamine transmembrane transport(GO:1902267) |
| 0.3 | 10.4 | GO:0006783 | heme biosynthetic process(GO:0006783) |
| 0.3 | 1.4 | GO:0032290 | peripheral nervous system myelin formation(GO:0032290) |
| 0.3 | 0.9 | GO:1902174 | positive regulation of keratinocyte apoptotic process(GO:1902174) |
| 0.3 | 1.1 | GO:0030091 | protein repair(GO:0030091) |
| 0.3 | 1.1 | GO:0060084 | synaptic transmission involved in micturition(GO:0060084) |
| 0.3 | 1.4 | GO:0072513 | positive regulation of secondary heart field cardioblast proliferation(GO:0072513) |
| 0.3 | 3.1 | GO:0070914 | UV-damage excision repair(GO:0070914) |
| 0.3 | 0.8 | GO:1904491 | protein localization to ciliary transition zone(GO:1904491) |
| 0.3 | 0.3 | GO:2001181 | positive regulation of interleukin-10 secretion(GO:2001181) |
| 0.3 | 1.7 | GO:0002442 | serotonin production involved in inflammatory response(GO:0002351) serotonin secretion involved in inflammatory response(GO:0002442) serotonin secretion by platelet(GO:0002554) |
| 0.3 | 8.9 | GO:0050779 | RNA destabilization(GO:0050779) |
| 0.3 | 0.8 | GO:0042732 | D-xylose metabolic process(GO:0042732) |
| 0.3 | 1.9 | GO:0006616 | SRP-dependent cotranslational protein targeting to membrane, translocation(GO:0006616) |
| 0.3 | 2.2 | GO:0009435 | NAD biosynthetic process(GO:0009435) |
| 0.3 | 3.3 | GO:0006527 | arginine catabolic process(GO:0006527) |
| 0.3 | 0.8 | GO:0010760 | negative regulation of macrophage chemotaxis(GO:0010760) |
| 0.3 | 1.3 | GO:0040010 | positive regulation of growth rate(GO:0040010) |
| 0.3 | 0.8 | GO:0021524 | visceral motor neuron differentiation(GO:0021524) |
| 0.3 | 1.1 | GO:0000019 | regulation of mitotic recombination(GO:0000019) negative regulation of mitotic recombination(GO:0045950) |
| 0.3 | 1.3 | GO:0006189 | 'de novo' IMP biosynthetic process(GO:0006189) |
| 0.3 | 0.8 | GO:1901873 | regulation of post-translational protein modification(GO:1901873) negative regulation of post-translational protein modification(GO:1901874) |
| 0.3 | 4.5 | GO:0071243 | cellular response to arsenic-containing substance(GO:0071243) |
| 0.3 | 2.1 | GO:1902856 | negative regulation of nonmotile primary cilium assembly(GO:1902856) |
| 0.3 | 4.4 | GO:0007130 | synaptonemal complex assembly(GO:0007130) |
| 0.3 | 1.0 | GO:1904779 | regulation of protein localization to centrosome(GO:1904779) |
| 0.3 | 1.0 | GO:0061209 | cell proliferation involved in mesonephros development(GO:0061209) |
| 0.3 | 1.0 | GO:0071043 | CUT catabolic process(GO:0071034) CUT metabolic process(GO:0071043) |
| 0.3 | 0.8 | GO:0010814 | neuropeptide catabolic process(GO:0010813) substance P catabolic process(GO:0010814) calcitonin catabolic process(GO:0010816) endothelin maturation(GO:0034959) |
| 0.3 | 0.8 | GO:0072488 | ammonium transmembrane transport(GO:0072488) |
| 0.3 | 0.5 | GO:0001922 | B-1 B cell homeostasis(GO:0001922) |
| 0.3 | 4.3 | GO:0070986 | left/right axis specification(GO:0070986) |
| 0.3 | 4.9 | GO:0060670 | branching involved in labyrinthine layer morphogenesis(GO:0060670) |
| 0.3 | 0.5 | GO:1903862 | positive regulation of oxidative phosphorylation(GO:1903862) |
| 0.3 | 1.0 | GO:0046391 | 5-phosphoribose 1-diphosphate biosynthetic process(GO:0006015) 5-phosphoribose 1-diphosphate metabolic process(GO:0046391) |
| 0.3 | 0.8 | GO:1904016 | response to Thyroglobulin triiodothyronine(GO:1904016) cellular response to Thyroglobulin triiodothyronine(GO:1904017) |
| 0.3 | 6.4 | GO:0060396 | growth hormone receptor signaling pathway(GO:0060396) cellular response to growth hormone stimulus(GO:0071378) |
| 0.3 | 1.3 | GO:1900041 | negative regulation of interleukin-2 secretion(GO:1900041) |
| 0.3 | 2.8 | GO:1901030 | positive regulation of mitochondrial outer membrane permeabilization involved in apoptotic signaling pathway(GO:1901030) |
| 0.3 | 1.0 | GO:0019682 | glyceraldehyde-3-phosphate metabolic process(GO:0019682) |
| 0.3 | 2.8 | GO:0010891 | negative regulation of sequestering of triglyceride(GO:0010891) |
| 0.3 | 1.0 | GO:0035405 | histone-threonine phosphorylation(GO:0035405) |
| 0.3 | 2.0 | GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0000083) |
| 0.3 | 1.5 | GO:0042256 | mature ribosome assembly(GO:0042256) |
| 0.2 | 0.7 | GO:0001180 | transcription initiation from RNA polymerase I promoter for nuclear large rRNA transcript(GO:0001180) |
| 0.2 | 1.0 | GO:0072268 | pattern specification involved in metanephros development(GO:0072268) |
| 0.2 | 1.5 | GO:0035095 | behavioral response to nicotine(GO:0035095) |
| 0.2 | 2.2 | GO:0000730 | DNA recombinase assembly(GO:0000730) double-strand break repair via synthesis-dependent strand annealing(GO:0045003) |
| 0.2 | 0.2 | GO:0071459 | protein localization to chromosome, centromeric region(GO:0071459) |
| 0.2 | 2.2 | GO:2000381 | negative regulation of mesoderm development(GO:2000381) |
| 0.2 | 0.7 | GO:0070782 | phosphatidylserine exposure on apoptotic cell surface(GO:0070782) |
| 0.2 | 1.5 | GO:0045653 | negative regulation of megakaryocyte differentiation(GO:0045653) |
| 0.2 | 5.1 | GO:0051085 | chaperone mediated protein folding requiring cofactor(GO:0051085) |
| 0.2 | 6.8 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.2 | 2.7 | GO:0072553 | terminal button organization(GO:0072553) |
| 0.2 | 0.7 | GO:0016259 | selenocysteine metabolic process(GO:0016259) |
| 0.2 | 1.0 | GO:0006049 | UDP-N-acetylglucosamine catabolic process(GO:0006049) |
| 0.2 | 2.1 | GO:2000484 | positive regulation of interleukin-8 secretion(GO:2000484) |
| 0.2 | 1.7 | GO:2000271 | positive regulation of fibroblast apoptotic process(GO:2000271) |
| 0.2 | 3.1 | GO:0061299 | retina vasculature morphogenesis in camera-type eye(GO:0061299) |
| 0.2 | 2.4 | GO:0015879 | carnitine transport(GO:0015879) |
| 0.2 | 0.2 | GO:0031204 | posttranslational protein targeting to membrane, translocation(GO:0031204) |
| 0.2 | 0.7 | GO:0071460 | cellular response to cell-matrix adhesion(GO:0071460) |
| 0.2 | 3.7 | GO:0006337 | nucleosome disassembly(GO:0006337) |
| 0.2 | 0.7 | GO:0019427 | acetate biosynthetic process(GO:0019413) acetyl-CoA biosynthetic process from acetate(GO:0019427) propionate biosynthetic process(GO:0019542) |
| 0.2 | 2.8 | GO:0090219 | negative regulation of lipid kinase activity(GO:0090219) |
| 0.2 | 0.7 | GO:0002017 | regulation of blood volume by renal aldosterone(GO:0002017) |
| 0.2 | 0.7 | GO:0001743 | optic placode formation(GO:0001743) optic placode formation involved in camera-type eye formation(GO:0046619) |
| 0.2 | 0.2 | GO:0060796 | regulation of transcription involved in primary germ layer cell fate commitment(GO:0060796) |
| 0.2 | 0.9 | GO:0098795 | mRNA cleavage involved in gene silencing by miRNA(GO:0035279) mRNA cleavage involved in gene silencing(GO:0098795) |
| 0.2 | 7.0 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.2 | 1.4 | GO:0006451 | selenocysteine incorporation(GO:0001514) translational readthrough(GO:0006451) |
| 0.2 | 1.8 | GO:0051660 | establishment of centrosome localization(GO:0051660) |
| 0.2 | 0.5 | GO:0044789 | modulation by host of viral release from host cell(GO:0044789) positive regulation by host of viral release from host cell(GO:0044791) |
| 0.2 | 3.4 | GO:0060211 | regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060211) positive regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060213) |
| 0.2 | 3.4 | GO:0043248 | proteasome assembly(GO:0043248) |
| 0.2 | 2.0 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 0.2 | 0.9 | GO:0003017 | lymph circulation(GO:0003017) |
| 0.2 | 0.9 | GO:0070562 | regulation of vitamin D receptor signaling pathway(GO:0070562) |
| 0.2 | 15.2 | GO:0043966 | histone H3 acetylation(GO:0043966) |
| 0.2 | 2.9 | GO:0071493 | cellular response to UV-B(GO:0071493) |
| 0.2 | 2.2 | GO:1904667 | negative regulation of ubiquitin protein ligase activity(GO:1904667) |
| 0.2 | 0.2 | GO:0003134 | BMP signaling pathway involved in heart induction(GO:0003130) endodermal-mesodermal cell signaling(GO:0003133) endodermal-mesodermal cell signaling involved in heart induction(GO:0003134) cell-cell signaling involved in cell fate commitment(GO:0045168) |
| 0.2 | 15.0 | GO:0032611 | interleukin-1 beta production(GO:0032611) |
| 0.2 | 3.5 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.2 | 1.3 | GO:1903182 | regulation of SUMO transferase activity(GO:1903182) positive regulation of SUMO transferase activity(GO:1903755) |
| 0.2 | 0.7 | GO:2000158 | positive regulation of ubiquitin-specific protease activity(GO:2000158) |
| 0.2 | 0.7 | GO:0035928 | rRNA import into mitochondrion(GO:0035928) |
| 0.2 | 1.7 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.2 | 0.9 | GO:2001287 | negative regulation of caveolin-mediated endocytosis(GO:2001287) |
| 0.2 | 1.5 | GO:0008635 | activation of cysteine-type endopeptidase activity involved in apoptotic process by cytochrome c(GO:0008635) |
| 0.2 | 4.7 | GO:0038092 | nodal signaling pathway(GO:0038092) |
| 0.2 | 1.7 | GO:0045046 | peroxisomal membrane transport(GO:0015919) protein import into peroxisome membrane(GO:0045046) |
| 0.2 | 1.3 | GO:1900103 | positive regulation of endoplasmic reticulum unfolded protein response(GO:1900103) |
| 0.2 | 3.8 | GO:0070633 | transepithelial transport(GO:0070633) |
| 0.2 | 0.8 | GO:0023016 | signal transduction by trans-phosphorylation(GO:0023016) |
| 0.2 | 1.1 | GO:0070459 | prolactin secretion(GO:0070459) |
| 0.2 | 2.3 | GO:0034242 | negative regulation of syncytium formation by plasma membrane fusion(GO:0034242) |
| 0.2 | 0.6 | GO:0045084 | positive regulation of interleukin-12 biosynthetic process(GO:0045084) |
| 0.2 | 3.1 | GO:0032331 | negative regulation of chondrocyte differentiation(GO:0032331) |
| 0.2 | 2.5 | GO:0000244 | spliceosomal tri-snRNP complex assembly(GO:0000244) |
| 0.2 | 3.6 | GO:0043312 | neutrophil degranulation(GO:0043312) |
| 0.2 | 0.8 | GO:0017126 | nucleologenesis(GO:0017126) |
| 0.2 | 1.9 | GO:0006307 | DNA dealkylation involved in DNA repair(GO:0006307) |
| 0.2 | 0.8 | GO:0051292 | nuclear pore complex assembly(GO:0051292) |
| 0.2 | 2.3 | GO:0032367 | intracellular sterol transport(GO:0032366) intracellular cholesterol transport(GO:0032367) |
| 0.2 | 1.2 | GO:0010815 | bradykinin catabolic process(GO:0010815) |
| 0.2 | 1.6 | GO:0032570 | response to progesterone(GO:0032570) |
| 0.2 | 2.0 | GO:0050861 | positive regulation of B cell receptor signaling pathway(GO:0050861) |
| 0.2 | 3.7 | GO:0030213 | hyaluronan biosynthetic process(GO:0030213) |
| 0.2 | 0.8 | GO:0060100 | positive regulation of phagocytosis, engulfment(GO:0060100) positive regulation of membrane invagination(GO:1905155) |
| 0.2 | 2.4 | GO:0060766 | negative regulation of androgen receptor signaling pathway(GO:0060766) |
| 0.2 | 1.6 | GO:0033184 | positive regulation of histone ubiquitination(GO:0033184) |
| 0.2 | 3.2 | GO:0007250 | activation of NF-kappaB-inducing kinase activity(GO:0007250) |
| 0.2 | 0.6 | GO:0046967 | cytosol to ER transport(GO:0046967) |
| 0.2 | 1.0 | GO:0035660 | MyD88-dependent toll-like receptor 4 signaling pathway(GO:0035660) |
| 0.2 | 1.4 | GO:0021775 | smoothened signaling pathway involved in ventral spinal cord interneuron specification(GO:0021775) smoothened signaling pathway involved in spinal cord motor neuron cell fate specification(GO:0021776) |
| 0.2 | 0.2 | GO:0010982 | regulation of high-density lipoprotein particle clearance(GO:0010982) |
| 0.2 | 8.2 | GO:0000184 | nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:0000184) |
| 0.2 | 2.0 | GO:0034312 | diol biosynthetic process(GO:0034312) |
| 0.2 | 1.6 | GO:0060770 | negative regulation of epithelial cell proliferation involved in prostate gland development(GO:0060770) |
| 0.2 | 2.6 | GO:0035721 | intraciliary retrograde transport(GO:0035721) |
| 0.2 | 0.6 | GO:0060464 | lung lobe formation(GO:0060464) diaphragm morphogenesis(GO:0060540) negative regulation of connective tissue replacement(GO:1905204) |
| 0.2 | 5.0 | GO:0060252 | positive regulation of glial cell proliferation(GO:0060252) |
| 0.2 | 2.0 | GO:0015791 | polyol transport(GO:0015791) |
| 0.2 | 0.8 | GO:0070889 | platelet alpha granule organization(GO:0070889) |
| 0.2 | 1.4 | GO:0030263 | apoptotic chromosome condensation(GO:0030263) |
| 0.2 | 0.8 | GO:0090346 | nitrate catabolic process(GO:0043602) nitric oxide catabolic process(GO:0046210) cellular organohalogen metabolic process(GO:0090345) cellular organofluorine metabolic process(GO:0090346) |
| 0.2 | 1.2 | GO:0045607 | regulation of auditory receptor cell differentiation(GO:0045607) regulation of mechanoreceptor differentiation(GO:0045631) regulation of inner ear receptor cell differentiation(GO:2000980) |
| 0.2 | 3.5 | GO:0035518 | histone H2A monoubiquitination(GO:0035518) |
| 0.2 | 0.8 | GO:0060391 | positive regulation of SMAD protein import into nucleus(GO:0060391) |
| 0.2 | 1.0 | GO:0033313 | meiotic cell cycle checkpoint(GO:0033313) |
| 0.2 | 0.8 | GO:0030422 | production of siRNA involved in RNA interference(GO:0030422) |
| 0.2 | 4.3 | GO:0019884 | antigen processing and presentation of exogenous antigen(GO:0019884) |
| 0.2 | 0.6 | GO:0006059 | hexitol metabolic process(GO:0006059) |
| 0.2 | 8.0 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.2 | 0.6 | GO:0048865 | stem cell fate commitment(GO:0048865) taste bud development(GO:0061193) |
| 0.2 | 0.4 | GO:0070278 | extracellular matrix constituent secretion(GO:0070278) |
| 0.2 | 4.6 | GO:0071539 | protein localization to centrosome(GO:0071539) |
| 0.2 | 3.3 | GO:0016578 | histone deubiquitination(GO:0016578) |
| 0.2 | 0.6 | GO:0045053 | protein retention in Golgi apparatus(GO:0045053) |
| 0.2 | 2.1 | GO:0002361 | CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0002361) |
| 0.2 | 0.8 | GO:0045719 | negative regulation of glycogen biosynthetic process(GO:0045719) |
| 0.2 | 1.1 | GO:0038128 | ERBB2 signaling pathway(GO:0038128) |
| 0.2 | 1.3 | GO:0040016 | embryonic cleavage(GO:0040016) |
| 0.2 | 2.3 | GO:0009301 | snRNA transcription(GO:0009301) |
| 0.2 | 1.3 | GO:0038165 | oncostatin-M-mediated signaling pathway(GO:0038165) |
| 0.2 | 0.4 | GO:0036091 | positive regulation of transcription from RNA polymerase II promoter in response to oxidative stress(GO:0036091) |
| 0.2 | 0.9 | GO:0031022 | nuclear migration along microfilament(GO:0031022) |
| 0.2 | 0.9 | GO:1902809 | regulation of skeletal muscle fiber differentiation(GO:1902809) |
| 0.2 | 3.9 | GO:0071157 | negative regulation of cell cycle arrest(GO:0071157) |
| 0.2 | 1.1 | GO:0036066 | protein O-linked fucosylation(GO:0036066) |
| 0.2 | 0.7 | GO:0044208 | AMP biosynthetic process(GO:0006167) 'de novo' AMP biosynthetic process(GO:0044208) |
| 0.2 | 0.7 | GO:1902303 | negative regulation of potassium ion export(GO:1902303) |
| 0.2 | 3.1 | GO:0036093 | germ cell proliferation(GO:0036093) |
| 0.2 | 1.6 | GO:1900745 | positive regulation of p38MAPK cascade(GO:1900745) |
| 0.2 | 4.8 | GO:0045747 | positive regulation of Notch signaling pathway(GO:0045747) |
| 0.2 | 3.0 | GO:0051601 | exocyst localization(GO:0051601) |
| 0.2 | 8.8 | GO:0000281 | mitotic cytokinesis(GO:0000281) |
| 0.2 | 0.7 | GO:0051562 | negative regulation of mitochondrial calcium ion concentration(GO:0051562) |
| 0.2 | 1.8 | GO:0060732 | positive regulation of inositol phosphate biosynthetic process(GO:0060732) |
| 0.2 | 1.6 | GO:0046477 | glycosylceramide catabolic process(GO:0046477) |
| 0.2 | 0.9 | GO:2000325 | regulation of ligand-dependent nuclear receptor transcription coactivator activity(GO:2000325) positive regulation of ligand-dependent nuclear receptor transcription coactivator activity(GO:2000327) |
| 0.2 | 0.9 | GO:1900825 | regulation of membrane depolarization during cardiac muscle cell action potential(GO:1900825) |
| 0.2 | 2.6 | GO:0032959 | inositol trisphosphate biosynthetic process(GO:0032959) |
| 0.2 | 1.0 | GO:0031848 | protection from non-homologous end joining at telomere(GO:0031848) |
| 0.2 | 2.7 | GO:1902414 | protein localization to cell junction(GO:1902414) |
| 0.2 | 0.7 | GO:0006436 | tryptophanyl-tRNA aminoacylation(GO:0006436) |
| 0.2 | 2.2 | GO:0035330 | regulation of hippo signaling(GO:0035330) |
| 0.2 | 0.2 | GO:0034146 | toll-like receptor 5 signaling pathway(GO:0034146) |
| 0.2 | 0.3 | GO:0044878 | mitotic cytokinesis checkpoint(GO:0044878) |
| 0.2 | 0.3 | GO:0019087 | transformation of host cell by virus(GO:0019087) |
| 0.2 | 0.5 | GO:0006553 | lysine metabolic process(GO:0006553) |
| 0.2 | 3.8 | GO:0033962 | cytoplasmic mRNA processing body assembly(GO:0033962) |
| 0.2 | 4.3 | GO:0035162 | embryonic hemopoiesis(GO:0035162) |
| 0.2 | 1.2 | GO:0061502 | early endosome to recycling endosome transport(GO:0061502) |
| 0.2 | 4.4 | GO:0090224 | regulation of mitotic spindle organization(GO:0060236) regulation of spindle organization(GO:0090224) |
| 0.2 | 0.7 | GO:0022615 | protein to membrane docking(GO:0022615) |
| 0.2 | 0.8 | GO:0046826 | negative regulation of protein export from nucleus(GO:0046826) |
| 0.2 | 1.0 | GO:1990839 | response to endothelin(GO:1990839) |
| 0.2 | 1.0 | GO:0048702 | embryonic neurocranium morphogenesis(GO:0048702) |
| 0.2 | 0.8 | GO:0070272 | proton-transporting ATP synthase complex assembly(GO:0043461) proton-transporting ATP synthase complex biogenesis(GO:0070272) |
| 0.2 | 1.3 | GO:0046602 | regulation of mitotic centrosome separation(GO:0046602) |
| 0.2 | 0.6 | GO:0045627 | positive regulation of T-helper 1 cell differentiation(GO:0045627) |
| 0.2 | 0.5 | GO:0033306 | phytol metabolic process(GO:0033306) fatty alcohol metabolic process(GO:1903173) |
| 0.2 | 1.9 | GO:0003215 | cardiac right ventricle morphogenesis(GO:0003215) |
| 0.2 | 4.2 | GO:0070208 | protein heterotrimerization(GO:0070208) |
| 0.2 | 0.5 | GO:0035038 | female pronucleus assembly(GO:0035038) |
| 0.2 | 1.5 | GO:0045602 | negative regulation of endothelial cell differentiation(GO:0045602) |
| 0.2 | 0.3 | GO:0003218 | cardiac left ventricle formation(GO:0003218) |
| 0.2 | 0.6 | GO:0043922 | negative regulation by host of viral transcription(GO:0043922) |
| 0.2 | 0.5 | GO:0043686 | co-translational protein modification(GO:0043686) |
| 0.2 | 1.1 | GO:0097368 | establishment of Sertoli cell barrier(GO:0097368) |
| 0.2 | 0.2 | GO:0018312 | peptidyl-serine ADP-ribosylation(GO:0018312) |
| 0.2 | 1.1 | GO:0060158 | phospholipase C-activating dopamine receptor signaling pathway(GO:0060158) |
| 0.1 | 5.1 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.1 | 0.9 | GO:0070345 | negative regulation of fat cell proliferation(GO:0070345) |
| 0.1 | 0.9 | GO:0009396 | folic acid-containing compound biosynthetic process(GO:0009396) |
| 0.1 | 0.6 | GO:2001137 | positive regulation of endocytic recycling(GO:2001137) |
| 0.1 | 2.7 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.1 | 5.0 | GO:0060351 | cartilage development involved in endochondral bone morphogenesis(GO:0060351) |
| 0.1 | 1.3 | GO:0031274 | positive regulation of pseudopodium assembly(GO:0031274) |
| 0.1 | 2.8 | GO:0048934 | peripheral nervous system neuron differentiation(GO:0048934) peripheral nervous system neuron development(GO:0048935) |
| 0.1 | 1.3 | GO:0002115 | store-operated calcium entry(GO:0002115) |
| 0.1 | 0.9 | GO:0038161 | prolactin signaling pathway(GO:0038161) |
| 0.1 | 1.1 | GO:0097428 | protein maturation by iron-sulfur cluster transfer(GO:0097428) |
| 0.1 | 0.1 | GO:0045918 | negative regulation of cytolysis(GO:0045918) |
| 0.1 | 1.6 | GO:0015909 | long-chain fatty acid transport(GO:0015909) |
| 0.1 | 0.3 | GO:0033140 | negative regulation of peptidyl-serine phosphorylation of STAT protein(GO:0033140) |
| 0.1 | 2.2 | GO:0045947 | negative regulation of translational initiation(GO:0045947) |
| 0.1 | 3.7 | GO:0043153 | entrainment of circadian clock by photoperiod(GO:0043153) |
| 0.1 | 1.2 | GO:0016480 | negative regulation of transcription from RNA polymerase III promoter(GO:0016480) positive regulation of tau-protein kinase activity(GO:1902949) |
| 0.1 | 0.7 | GO:0031441 | negative regulation of mRNA 3'-end processing(GO:0031441) |
| 0.1 | 1.1 | GO:0006686 | sphingomyelin biosynthetic process(GO:0006686) |
| 0.1 | 3.5 | GO:0007140 | male meiosis(GO:0007140) |
| 0.1 | 1.6 | GO:0034472 | snRNA 3'-end processing(GO:0034472) |
| 0.1 | 1.1 | GO:0010838 | positive regulation of keratinocyte proliferation(GO:0010838) |
| 0.1 | 2.4 | GO:0009299 | mRNA transcription(GO:0009299) |
| 0.1 | 0.5 | GO:1904925 | positive regulation of macromitophagy(GO:1901526) positive regulation of mitophagy in response to mitochondrial depolarization(GO:1904925) |
| 0.1 | 0.4 | GO:0090272 | negative regulation of fibroblast growth factor production(GO:0090272) |
| 0.1 | 0.9 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 0.1 | 4.5 | GO:0010718 | positive regulation of epithelial to mesenchymal transition(GO:0010718) |
| 0.1 | 1.2 | GO:0038203 | TORC2 signaling(GO:0038203) |
| 0.1 | 0.3 | GO:0031440 | regulation of mRNA 3'-end processing(GO:0031440) |
| 0.1 | 1.4 | GO:0010739 | positive regulation of protein kinase A signaling(GO:0010739) |
| 0.1 | 2.0 | GO:0032967 | positive regulation of collagen biosynthetic process(GO:0032967) |
| 0.1 | 1.3 | GO:0007143 | female meiotic division(GO:0007143) |
| 0.1 | 0.6 | GO:0046073 | dTMP biosynthetic process(GO:0006231) dTMP metabolic process(GO:0046073) |
| 0.1 | 0.5 | GO:0033578 | protein glycosylation in Golgi(GO:0033578) |
| 0.1 | 1.1 | GO:0060179 | male mating behavior(GO:0060179) |
| 0.1 | 0.3 | GO:0060449 | bud elongation involved in lung branching(GO:0060449) |
| 0.1 | 0.5 | GO:0032342 | aldosterone biosynthetic process(GO:0032342) |
| 0.1 | 0.3 | GO:0034145 | positive regulation of toll-like receptor 4 signaling pathway(GO:0034145) |
| 0.1 | 1.2 | GO:0048304 | positive regulation of isotype switching to IgG isotypes(GO:0048304) |
| 0.1 | 1.2 | GO:0046685 | response to arsenic-containing substance(GO:0046685) |
| 0.1 | 4.0 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.1 | 0.4 | GO:2001293 | fatty-acyl-CoA biosynthetic process(GO:0046949) malonyl-CoA metabolic process(GO:2001293) |
| 0.1 | 1.4 | GO:0071361 | cellular response to ethanol(GO:0071361) |
| 0.1 | 2.0 | GO:0050434 | positive regulation of viral transcription(GO:0050434) |
| 0.1 | 0.4 | GO:2001034 | positive regulation of double-strand break repair via nonhomologous end joining(GO:2001034) |
| 0.1 | 1.8 | GO:0048757 | endosome to melanosome transport(GO:0035646) endosome to pigment granule transport(GO:0043485) pigment granule maturation(GO:0048757) |
| 0.1 | 0.7 | GO:0043102 | amino acid salvage(GO:0043102) L-methionine salvage(GO:0071267) |
| 0.1 | 0.2 | GO:0021912 | regulation of transcription from RNA polymerase II promoter involved in spinal cord motor neuron fate specification(GO:0021912) |
| 0.1 | 1.0 | GO:0033211 | adiponectin-activated signaling pathway(GO:0033211) |
| 0.1 | 0.8 | GO:0060309 | elastin catabolic process(GO:0060309) |
| 0.1 | 2.4 | GO:0043574 | protein targeting to peroxisome(GO:0006625) peroxisomal transport(GO:0043574) protein localization to peroxisome(GO:0072662) establishment of protein localization to peroxisome(GO:0072663) |
| 0.1 | 5.1 | GO:0006919 | activation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0006919) |
| 0.1 | 0.6 | GO:0043589 | skin morphogenesis(GO:0043589) |
| 0.1 | 2.1 | GO:0097284 | hepatocyte apoptotic process(GO:0097284) |
| 0.1 | 7.3 | GO:0007569 | cell aging(GO:0007569) |
| 0.1 | 0.9 | GO:2000353 | positive regulation of endothelial cell apoptotic process(GO:2000353) |
| 0.1 | 1.3 | GO:0006642 | triglyceride mobilization(GO:0006642) |
| 0.1 | 2.3 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.1 | 1.8 | GO:0030261 | chromosome condensation(GO:0030261) |
| 0.1 | 0.5 | GO:0001880 | Mullerian duct regression(GO:0001880) |
| 0.1 | 4.6 | GO:0042246 | tissue regeneration(GO:0042246) |
| 0.1 | 0.5 | GO:0009113 | adenine salvage(GO:0006168) purine nucleobase biosynthetic process(GO:0009113) adenine metabolic process(GO:0046083) adenine biosynthetic process(GO:0046084) |
| 0.1 | 0.8 | GO:0018027 | peptidyl-lysine dimethylation(GO:0018027) |
| 0.1 | 1.3 | GO:0071712 | ER-associated misfolded protein catabolic process(GO:0071712) |
| 0.1 | 0.3 | GO:0034031 | coenzyme A catabolic process(GO:0015938) nucleoside bisphosphate catabolic process(GO:0033869) ribonucleoside bisphosphate catabolic process(GO:0034031) purine nucleoside bisphosphate catabolic process(GO:0034034) acetyl-CoA catabolic process(GO:0046356) |
| 0.1 | 0.7 | GO:0051534 | negative regulation of NFAT protein import into nucleus(GO:0051534) |
| 0.1 | 0.9 | GO:0048642 | negative regulation of skeletal muscle tissue development(GO:0048642) |
| 0.1 | 0.5 | GO:0035617 | stress granule disassembly(GO:0035617) |
| 0.1 | 1.4 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.1 | 3.0 | GO:0032801 | receptor catabolic process(GO:0032801) |
| 0.1 | 0.8 | GO:0003351 | epithelial cilium movement(GO:0003351) |
| 0.1 | 0.4 | GO:0060708 | spongiotrophoblast differentiation(GO:0060708) |
| 0.1 | 0.1 | GO:0070358 | actin polymerization-dependent cell motility(GO:0070358) |
| 0.1 | 1.4 | GO:0070072 | vacuolar proton-transporting V-type ATPase complex assembly(GO:0070072) |
| 0.1 | 0.5 | GO:0071894 | histone H2B conserved C-terminal lysine ubiquitination(GO:0071894) |
| 0.1 | 3.9 | GO:0006284 | base-excision repair(GO:0006284) |
| 0.1 | 0.3 | GO:1990168 | protein K27-linked deubiquitination(GO:1990167) protein K33-linked deubiquitination(GO:1990168) |
| 0.1 | 0.2 | GO:0021698 | cerebellar cortex structural organization(GO:0021698) |
| 0.1 | 1.4 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.1 | 0.6 | GO:1904872 | regulation of telomerase RNA localization to Cajal body(GO:1904872) |
| 0.1 | 0.3 | GO:0006106 | fumarate metabolic process(GO:0006106) |
| 0.1 | 0.4 | GO:0071680 | response to indole-3-methanol(GO:0071680) cellular response to indole-3-methanol(GO:0071681) |
| 0.1 | 2.2 | GO:0042481 | regulation of odontogenesis(GO:0042481) |
| 0.1 | 0.4 | GO:0043569 | negative regulation of insulin-like growth factor receptor signaling pathway(GO:0043569) |
| 0.1 | 0.2 | GO:0061153 | trachea submucosa development(GO:0061152) trachea gland development(GO:0061153) |
| 0.1 | 0.4 | GO:0097202 | activation of cysteine-type endopeptidase activity(GO:0097202) |
| 0.1 | 0.2 | GO:0030961 | peptidyl-arginine hydroxylation(GO:0030961) |
| 0.1 | 2.4 | GO:0032467 | positive regulation of cytokinesis(GO:0032467) |
| 0.1 | 2.0 | GO:0032516 | positive regulation of phosphoprotein phosphatase activity(GO:0032516) |
| 0.1 | 1.8 | GO:0016973 | poly(A)+ mRNA export from nucleus(GO:0016973) |
| 0.1 | 0.9 | GO:0009437 | carnitine metabolic process(GO:0009437) |
| 0.1 | 0.2 | GO:1903410 | nitric oxide production involved in inflammatory response(GO:0002537) lysine import(GO:0034226) L-lysine import(GO:0061461) L-lysine import into cell(GO:1903410) |
| 0.1 | 2.3 | GO:0071470 | cellular response to osmotic stress(GO:0071470) |
| 0.1 | 0.7 | GO:0060613 | fat pad development(GO:0060613) |
| 0.1 | 1.1 | GO:0016180 | snRNA processing(GO:0016180) |
| 0.1 | 2.8 | GO:0035313 | wound healing, spreading of epidermal cells(GO:0035313) |
| 0.1 | 3.5 | GO:0090162 | establishment of epithelial cell polarity(GO:0090162) |
| 0.1 | 0.8 | GO:0000712 | resolution of meiotic recombination intermediates(GO:0000712) |
| 0.1 | 0.6 | GO:1905214 | regulation of mRNA binding(GO:1902415) positive regulation of mRNA binding(GO:1902416) regulation of RNA binding(GO:1905214) positive regulation of RNA binding(GO:1905216) |
| 0.1 | 0.8 | GO:0060712 | spongiotrophoblast layer development(GO:0060712) |
| 0.1 | 1.0 | GO:0016576 | histone dephosphorylation(GO:0016576) |
| 0.1 | 1.0 | GO:0070236 | negative regulation of activation-induced cell death of T cells(GO:0070236) |
| 0.1 | 0.6 | GO:0035063 | nuclear speck organization(GO:0035063) |
| 0.1 | 0.3 | GO:0031642 | negative regulation of myelination(GO:0031642) |
| 0.1 | 2.7 | GO:0030947 | regulation of vascular endothelial growth factor receptor signaling pathway(GO:0030947) |
| 0.1 | 0.4 | GO:0030382 | sperm mitochondrion organization(GO:0030382) |
| 0.1 | 1.0 | GO:0019730 | antimicrobial humoral response(GO:0019730) |
| 0.1 | 0.5 | GO:0090527 | actin filament reorganization(GO:0090527) |
| 0.1 | 0.8 | GO:0042637 | catagen(GO:0042637) |
| 0.1 | 3.2 | GO:0050435 | beta-amyloid metabolic process(GO:0050435) |
| 0.1 | 0.2 | GO:0061300 | cerebellum vasculature development(GO:0061300) |
| 0.1 | 0.6 | GO:0060017 | parathyroid gland development(GO:0060017) |
| 0.1 | 1.2 | GO:0033750 | ribosomal subunit export from nucleus(GO:0000054) ribosome localization(GO:0033750) establishment of ribosome localization(GO:0033753) |
| 0.1 | 0.6 | GO:0006283 | transcription-coupled nucleotide-excision repair(GO:0006283) |
| 0.1 | 2.6 | GO:1903963 | arachidonic acid secretion(GO:0050482) arachidonate transport(GO:1903963) |
| 0.1 | 0.5 | GO:0090214 | spongiotrophoblast layer developmental growth(GO:0090214) |
| 0.1 | 0.5 | GO:0033314 | mitotic DNA replication checkpoint(GO:0033314) |
| 0.1 | 4.7 | GO:0007566 | embryo implantation(GO:0007566) |
| 0.1 | 2.3 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.1 | 0.4 | GO:0045086 | positive regulation of interleukin-2 biosynthetic process(GO:0045086) |
| 0.1 | 0.4 | GO:0060136 | embryonic process involved in female pregnancy(GO:0060136) |
| 0.1 | 0.5 | GO:2000562 | regulation of CD4-positive, alpha-beta T cell proliferation(GO:2000561) negative regulation of CD4-positive, alpha-beta T cell proliferation(GO:2000562) |
| 0.1 | 0.2 | GO:0036230 | granulocyte activation(GO:0036230) |
| 0.1 | 0.9 | GO:0031119 | tRNA pseudouridine synthesis(GO:0031119) |
| 0.1 | 0.9 | GO:0031163 | iron-sulfur cluster assembly(GO:0016226) metallo-sulfur cluster assembly(GO:0031163) |
| 0.1 | 0.6 | GO:0061029 | eyelid development in camera-type eye(GO:0061029) |
| 0.1 | 0.8 | GO:0009950 | dorsal/ventral axis specification(GO:0009950) |
| 0.1 | 2.2 | GO:0000132 | establishment of mitotic spindle orientation(GO:0000132) |
| 0.1 | 2.9 | GO:0003281 | ventricular septum development(GO:0003281) |
| 0.1 | 0.7 | GO:0061484 | hematopoietic stem cell homeostasis(GO:0061484) |
| 0.1 | 0.3 | GO:0002949 | tRNA threonylcarbamoyladenosine modification(GO:0002949) |
| 0.1 | 0.7 | GO:2000394 | positive regulation of lamellipodium morphogenesis(GO:2000394) |
| 0.1 | 0.8 | GO:0006458 | 'de novo' protein folding(GO:0006458) |
| 0.1 | 0.6 | GO:0035360 | positive regulation of peroxisome proliferator activated receptor signaling pathway(GO:0035360) |
| 0.1 | 0.5 | GO:0001731 | formation of translation preinitiation complex(GO:0001731) |
| 0.1 | 2.0 | GO:0060065 | uterus development(GO:0060065) |
| 0.1 | 0.2 | GO:0071500 | cellular response to nitrosative stress(GO:0071500) |
| 0.1 | 1.3 | GO:1902230 | negative regulation of intrinsic apoptotic signaling pathway in response to DNA damage(GO:1902230) |
| 0.1 | 0.2 | GO:0000076 | DNA replication checkpoint(GO:0000076) |
| 0.1 | 0.8 | GO:0006353 | DNA-templated transcription, termination(GO:0006353) |
| 0.1 | 0.2 | GO:0045212 | neurotransmitter receptor biosynthetic process(GO:0045212) |
| 0.1 | 0.6 | GO:0045200 | establishment or maintenance of neuroblast polarity(GO:0045196) establishment of neuroblast polarity(GO:0045200) |
| 0.1 | 0.5 | GO:0033762 | response to glucagon(GO:0033762) |
| 0.1 | 1.3 | GO:0060644 | mammary gland epithelial cell differentiation(GO:0060644) |
| 0.1 | 0.5 | GO:0046643 | regulation of gamma-delta T cell differentiation(GO:0045586) regulation of gamma-delta T cell activation(GO:0046643) |
| 0.1 | 1.2 | GO:0002446 | neutrophil mediated immunity(GO:0002446) |
| 0.1 | 0.2 | GO:2000313 | fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:0060825) regulation of fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:2000313) |
| 0.1 | 0.4 | GO:0038031 | non-canonical Wnt signaling pathway via MAPK cascade(GO:0038030) non-canonical Wnt signaling pathway via JNK cascade(GO:0038031) |
| 0.1 | 0.9 | GO:0051490 | negative regulation of filopodium assembly(GO:0051490) |
| 0.1 | 0.5 | GO:0007256 | activation of JNKK activity(GO:0007256) negative regulation of interleukin-8 secretion(GO:2000483) |
| 0.1 | 0.6 | GO:0006907 | pinocytosis(GO:0006907) |
| 0.1 | 0.4 | GO:0034720 | histone H3-K4 demethylation(GO:0034720) |
| 0.1 | 0.3 | GO:0032506 | cytokinetic process(GO:0032506) |
| 0.1 | 0.2 | GO:0019919 | peptidyl-arginine methylation, to asymmetrical-dimethyl arginine(GO:0019919) |
| 0.1 | 3.1 | GO:0035025 | positive regulation of Rho protein signal transduction(GO:0035025) |
| 0.1 | 1.4 | GO:0048745 | smooth muscle tissue development(GO:0048745) |
| 0.1 | 0.8 | GO:0015074 | DNA integration(GO:0015074) |
| 0.1 | 1.9 | GO:0042092 | type 2 immune response(GO:0042092) |
| 0.1 | 0.6 | GO:0043435 | response to corticotropin-releasing hormone(GO:0043435) cellular response to corticotropin-releasing hormone stimulus(GO:0071376) |
| 0.1 | 0.2 | GO:0034134 | toll-like receptor 2 signaling pathway(GO:0034134) |
| 0.1 | 0.4 | GO:2000491 | positive regulation of hepatic stellate cell activation(GO:2000491) |
| 0.1 | 2.9 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.1 | 0.4 | GO:0000731 | DNA synthesis involved in DNA repair(GO:0000731) |
| 0.1 | 1.2 | GO:0030903 | notochord development(GO:0030903) |
| 0.1 | 1.2 | GO:0060441 | epithelial tube branching involved in lung morphogenesis(GO:0060441) |
| 0.1 | 2.2 | GO:0060325 | face morphogenesis(GO:0060325) |
| 0.1 | 0.5 | GO:1903297 | regulation of hypoxia-induced intrinsic apoptotic signaling pathway(GO:1903297) negative regulation of hypoxia-induced intrinsic apoptotic signaling pathway(GO:1903298) |
| 0.1 | 0.9 | GO:0072675 | osteoclast fusion(GO:0072675) |
| 0.1 | 0.9 | GO:1901223 | negative regulation of NIK/NF-kappaB signaling(GO:1901223) |
| 0.1 | 1.6 | GO:0071260 | cellular response to mechanical stimulus(GO:0071260) |
| 0.1 | 0.6 | GO:0034982 | mitochondrial protein processing(GO:0034982) |
| 0.1 | 0.8 | GO:0043278 | response to morphine(GO:0043278) |
| 0.1 | 0.7 | GO:0046322 | negative regulation of fatty acid oxidation(GO:0046322) |
| 0.1 | 0.5 | GO:0071850 | mitotic cell cycle arrest(GO:0071850) |
| 0.1 | 0.3 | GO:0030576 | Cajal body organization(GO:0030576) |
| 0.1 | 2.4 | GO:0006368 | transcription elongation from RNA polymerase II promoter(GO:0006368) |
| 0.1 | 2.2 | GO:0009072 | aromatic amino acid family metabolic process(GO:0009072) |
| 0.1 | 2.4 | GO:0001825 | blastocyst formation(GO:0001825) |
| 0.1 | 0.2 | GO:0007225 | patched ligand maturation(GO:0007225) signal maturation(GO:0035638) |
| 0.1 | 1.9 | GO:0035634 | response to stilbenoid(GO:0035634) |
| 0.1 | 0.4 | GO:0018158 | protein oxidation(GO:0018158) |
| 0.1 | 0.5 | GO:0061084 | regulation of protein refolding(GO:0061083) negative regulation of protein refolding(GO:0061084) |
| 0.1 | 1.8 | GO:0051443 | positive regulation of ubiquitin-protein transferase activity(GO:0051443) |
| 0.1 | 0.5 | GO:0032486 | Rap protein signal transduction(GO:0032486) |
| 0.1 | 1.0 | GO:0001706 | endoderm formation(GO:0001706) |
| 0.1 | 0.5 | GO:0000289 | nuclear-transcribed mRNA poly(A) tail shortening(GO:0000289) |
| 0.1 | 0.7 | GO:0006228 | UTP biosynthetic process(GO:0006228) |
| 0.1 | 31.9 | GO:0048232 | spermatogenesis(GO:0007283) male gamete generation(GO:0048232) |
| 0.1 | 0.1 | GO:0002884 | regulation of type IV hypersensitivity(GO:0001807) negative regulation of hypersensitivity(GO:0002884) |
| 0.1 | 0.8 | GO:0007099 | centriole replication(GO:0007099) |
| 0.1 | 0.8 | GO:1902187 | negative regulation of viral release from host cell(GO:1902187) |
| 0.1 | 0.1 | GO:0010040 | response to iron(II) ion(GO:0010040) |
| 0.1 | 0.6 | GO:0015858 | nucleoside transport(GO:0015858) |
| 0.1 | 1.7 | GO:0048536 | spleen development(GO:0048536) |
| 0.1 | 3.4 | GO:0009062 | fatty acid catabolic process(GO:0009062) |
| 0.1 | 2.0 | GO:0046854 | phosphatidylinositol phosphorylation(GO:0046854) |
| 0.1 | 1.3 | GO:0001913 | T cell mediated cytotoxicity(GO:0001913) |
| 0.1 | 0.2 | GO:1901301 | regulation of cargo loading into COPII-coated vesicle(GO:1901301) |
| 0.1 | 0.4 | GO:0042635 | positive regulation of hair cycle(GO:0042635) positive regulation of hair follicle development(GO:0051798) |
| 0.1 | 0.2 | GO:0006419 | alanyl-tRNA aminoacylation(GO:0006419) |
| 0.1 | 0.5 | GO:0060510 | Type II pneumocyte differentiation(GO:0060510) |
| 0.1 | 0.8 | GO:0051894 | positive regulation of focal adhesion assembly(GO:0051894) |
| 0.1 | 0.9 | GO:0016998 | cell wall macromolecule catabolic process(GO:0016998) |
| 0.1 | 3.6 | GO:0042058 | regulation of epidermal growth factor receptor signaling pathway(GO:0042058) |
| 0.1 | 1.7 | GO:0006767 | water-soluble vitamin metabolic process(GO:0006767) |
| 0.1 | 1.0 | GO:0090022 | regulation of neutrophil chemotaxis(GO:0090022) |
| 0.1 | 0.4 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.1 | 0.1 | GO:0030043 | actin filament fragmentation(GO:0030043) |
| 0.1 | 0.4 | GO:1902751 | positive regulation of cell cycle G2/M phase transition(GO:1902751) |
| 0.1 | 0.3 | GO:0070189 | kynurenine metabolic process(GO:0070189) |
| 0.1 | 0.4 | GO:0038063 | collagen-activated tyrosine kinase receptor signaling pathway(GO:0038063) |
| 0.1 | 0.8 | GO:0006622 | protein targeting to lysosome(GO:0006622) |
| 0.1 | 0.4 | GO:0090043 | regulation of tubulin deacetylation(GO:0090043) |
| 0.1 | 0.6 | GO:0006610 | ribosomal protein import into nucleus(GO:0006610) |
| 0.1 | 1.4 | GO:0050962 | detection of light stimulus involved in visual perception(GO:0050908) detection of light stimulus involved in sensory perception(GO:0050962) |
| 0.1 | 0.3 | GO:0032483 | regulation of Rab protein signal transduction(GO:0032483) |
| 0.1 | 2.2 | GO:0001580 | detection of chemical stimulus involved in sensory perception of bitter taste(GO:0001580) |
| 0.1 | 0.7 | GO:0045722 | positive regulation of gluconeogenesis(GO:0045722) |
| 0.1 | 0.7 | GO:0090136 | epithelial cell-cell adhesion(GO:0090136) |
| 0.1 | 0.1 | GO:0070650 | actin filament bundle distribution(GO:0070650) |
| 0.1 | 0.5 | GO:1900028 | negative regulation of ruffle assembly(GO:1900028) |
| 0.1 | 0.5 | GO:1904030 | negative regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0045736) negative regulation of cyclin-dependent protein kinase activity(GO:1904030) |
| 0.1 | 0.2 | GO:0030656 | regulation of vitamin metabolic process(GO:0030656) |
| 0.1 | 0.7 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.1 | 0.7 | GO:2000351 | regulation of endothelial cell apoptotic process(GO:2000351) |
| 0.1 | 1.1 | GO:0018023 | peptidyl-lysine trimethylation(GO:0018023) |
| 0.1 | 0.3 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 0.1 | 0.4 | GO:0061709 | reticulophagy(GO:0061709) |
| 0.1 | 0.2 | GO:0061739 | protein lipidation involved in autophagosome assembly(GO:0061739) |
| 0.1 | 0.3 | GO:0000466 | maturation of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000466) |
| 0.1 | 0.2 | GO:1903608 | protein localization to cytoplasmic stress granule(GO:1903608) |
| 0.1 | 0.8 | GO:0019433 | triglyceride catabolic process(GO:0019433) |
| 0.1 | 0.7 | GO:0031936 | negative regulation of chromatin silencing(GO:0031936) |
| 0.1 | 0.2 | GO:0036367 | adaptation of rhodopsin mediated signaling(GO:0016062) light adaption(GO:0036367) |
| 0.1 | 0.2 | GO:0006530 | asparagine catabolic process(GO:0006530) |
| 0.1 | 0.6 | GO:0072673 | lamellipodium morphogenesis(GO:0072673) |
| 0.1 | 0.2 | GO:0045830 | positive regulation of isotype switching(GO:0045830) |
| 0.1 | 1.5 | GO:0010596 | negative regulation of endothelial cell migration(GO:0010596) |
| 0.0 | 0.3 | GO:0006348 | chromatin silencing at telomere(GO:0006348) |
| 0.0 | 0.1 | GO:2000232 | regulation of rRNA processing(GO:2000232) |
| 0.0 | 2.4 | GO:0045453 | bone resorption(GO:0045453) |
| 0.0 | 0.1 | GO:0016132 | brassinosteroid metabolic process(GO:0016131) brassinosteroid biosynthetic process(GO:0016132) |
| 0.0 | 0.0 | GO:1902445 | regulation of mitochondrial membrane permeability involved in programmed necrotic cell death(GO:1902445) |
| 0.0 | 0.2 | GO:0000185 | activation of MAPKKK activity(GO:0000185) |
| 0.0 | 0.1 | GO:0060346 | bone trabecula formation(GO:0060346) |
| 0.0 | 0.2 | GO:0009992 | cellular water homeostasis(GO:0009992) |
| 0.0 | 1.0 | GO:0045604 | regulation of epidermal cell differentiation(GO:0045604) |
| 0.0 | 0.1 | GO:0019417 | sulfur oxidation(GO:0019417) |
| 0.0 | 0.5 | GO:0033234 | negative regulation of protein sumoylation(GO:0033234) |
| 0.0 | 3.7 | GO:0046847 | filopodium assembly(GO:0046847) |
| 0.0 | 0.9 | GO:1903077 | negative regulation of protein localization to plasma membrane(GO:1903077) negative regulation of protein localization to cell periphery(GO:1904376) |
| 0.0 | 0.1 | GO:0060339 | negative regulation of type I interferon-mediated signaling pathway(GO:0060339) |
| 0.0 | 0.7 | GO:0021516 | dorsal spinal cord development(GO:0021516) |
| 0.0 | 0.4 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.0 | 0.1 | GO:1901069 | guanosine-containing compound catabolic process(GO:1901069) |
| 0.0 | 0.1 | GO:0006409 | tRNA export from nucleus(GO:0006409) tRNA-containing ribonucleoprotein complex export from nucleus(GO:0071431) |
| 0.0 | 0.1 | GO:0090320 | chylomicron assembly(GO:0034378) regulation of chylomicron remnant clearance(GO:0090320) |
| 0.0 | 2.4 | GO:0002181 | cytoplasmic translation(GO:0002181) |
| 0.0 | 0.5 | GO:0048148 | behavioral response to cocaine(GO:0048148) |
| 0.0 | 0.3 | GO:0032532 | regulation of microvillus length(GO:0032532) |
| 0.0 | 0.1 | GO:0030540 | female genitalia development(GO:0030540) |
| 0.0 | 0.2 | GO:0015886 | heme transport(GO:0015886) |
| 0.0 | 1.1 | GO:0042733 | embryonic digit morphogenesis(GO:0042733) |
| 0.0 | 2.2 | GO:0014020 | neural tube closure(GO:0001843) primary neural tube formation(GO:0014020) tube closure(GO:0060606) |
| 0.0 | 0.4 | GO:0046007 | negative regulation of activated T cell proliferation(GO:0046007) |
| 0.0 | 0.5 | GO:0060628 | regulation of ER to Golgi vesicle-mediated transport(GO:0060628) |
| 0.0 | 0.3 | GO:0010578 | regulation of adenylate cyclase activity involved in G-protein coupled receptor signaling pathway(GO:0010578) positive regulation of adenylate cyclase activity involved in G-protein coupled receptor signaling pathway(GO:0010579) |
| 0.0 | 1.8 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.0 | 0.3 | GO:0071763 | nuclear membrane organization(GO:0071763) |
| 0.0 | 0.5 | GO:0030728 | ovulation(GO:0030728) |
| 0.0 | 0.2 | GO:1903525 | regulation of membrane tubulation(GO:1903525) positive regulation of membrane tubulation(GO:1903527) |
| 0.0 | 0.0 | GO:0039689 | negative stranded viral RNA replication(GO:0039689) multi-organism biosynthetic process(GO:0044034) |
| 0.0 | 0.2 | GO:0007023 | post-chaperonin tubulin folding pathway(GO:0007023) |
| 0.0 | 1.1 | GO:0007212 | dopamine receptor signaling pathway(GO:0007212) |
| 0.0 | 0.3 | GO:0006620 | posttranslational protein targeting to membrane(GO:0006620) |
| 0.0 | 0.2 | GO:0003170 | heart valve development(GO:0003170) |
| 0.0 | 0.4 | GO:0002002 | regulation of angiotensin levels in blood(GO:0002002) |
| 0.0 | 0.3 | GO:0019262 | N-acetylneuraminate catabolic process(GO:0019262) |
| 0.0 | 0.4 | GO:0035067 | negative regulation of histone acetylation(GO:0035067) |
| 0.0 | 0.2 | GO:0051013 | microtubule severing(GO:0051013) |
| 0.0 | 0.9 | GO:0001824 | blastocyst development(GO:0001824) |
| 0.0 | 0.6 | GO:1901222 | regulation of NIK/NF-kappaB signaling(GO:1901222) |
| 0.0 | 0.9 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.0 | 0.1 | GO:0060133 | somatotropin secreting cell development(GO:0060133) |
| 0.0 | 0.2 | GO:0039694 | viral RNA genome replication(GO:0039694) RNA replication(GO:0039703) |
| 0.0 | 0.4 | GO:0050820 | positive regulation of coagulation(GO:0050820) |
| 0.0 | 0.6 | GO:0006107 | oxaloacetate metabolic process(GO:0006107) |
| 0.0 | 0.9 | GO:0071398 | cellular response to fatty acid(GO:0071398) |
| 0.0 | 0.6 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.0 | 0.1 | GO:0014029 | neural crest formation(GO:0014029) |
| 0.0 | 0.3 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.0 | 0.4 | GO:0022027 | interkinetic nuclear migration(GO:0022027) |
| 0.0 | 1.9 | GO:0007229 | integrin-mediated signaling pathway(GO:0007229) |
| 0.0 | 0.4 | GO:0042953 | lipoprotein transport(GO:0042953) lipoprotein localization(GO:0044872) |
| 0.0 | 0.7 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.0 | 0.1 | GO:1904766 | negative regulation of macroautophagy by TORC1 signaling(GO:1904766) |
| 0.0 | 0.5 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.0 | 0.3 | GO:0043457 | regulation of cellular respiration(GO:0043457) |
| 0.0 | 0.2 | GO:0070127 | tRNA aminoacylation for mitochondrial protein translation(GO:0070127) |
| 0.0 | 0.5 | GO:0071391 | cellular response to estrogen stimulus(GO:0071391) |
| 0.0 | 0.3 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 0.0 | 1.5 | GO:2000117 | negative regulation of cysteine-type endopeptidase activity(GO:2000117) |
| 0.0 | 0.1 | GO:2000189 | cellular response to lipoprotein particle stimulus(GO:0071402) cellular response to low-density lipoprotein particle stimulus(GO:0071404) positive regulation of cholesterol homeostasis(GO:2000189) |
| 0.0 | 0.3 | GO:0006104 | succinyl-CoA metabolic process(GO:0006104) |
| 0.0 | 0.3 | GO:0036159 | inner dynein arm assembly(GO:0036159) |
| 0.0 | 0.2 | GO:0060713 | labyrinthine layer morphogenesis(GO:0060713) |
| 0.0 | 1.1 | GO:0031122 | cytoplasmic microtubule organization(GO:0031122) |
| 0.0 | 0.3 | GO:0001522 | pseudouridine synthesis(GO:0001522) |
| 0.0 | 0.7 | GO:0006378 | mRNA polyadenylation(GO:0006378) RNA polyadenylation(GO:0043631) |
| 0.0 | 0.9 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.0 | 0.4 | GO:0030259 | lipid glycosylation(GO:0030259) |
| 0.0 | 0.1 | GO:0010587 | miRNA catabolic process(GO:0010587) |
| 0.0 | 0.4 | GO:0000470 | maturation of LSU-rRNA(GO:0000470) |
| 0.0 | 1.5 | GO:0010212 | response to ionizing radiation(GO:0010212) |
| 0.0 | 1.1 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.0 | 0.1 | GO:0051026 | chiasma assembly(GO:0051026) |
| 0.0 | 0.1 | GO:0006432 | phenylalanyl-tRNA aminoacylation(GO:0006432) |
| 0.0 | 0.1 | GO:0072344 | rescue of stalled ribosome(GO:0072344) |
| 0.0 | 0.7 | GO:0031529 | ruffle organization(GO:0031529) |
| 0.0 | 0.4 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.0 | 0.4 | GO:0045662 | negative regulation of myoblast differentiation(GO:0045662) |
| 0.0 | 0.6 | GO:0048025 | negative regulation of mRNA splicing, via spliceosome(GO:0048025) |
| 0.0 | 0.0 | GO:0002946 | tRNA C5-cytosine methylation(GO:0002946) |
| 0.0 | 0.4 | GO:0042044 | fluid transport(GO:0042044) |
| 0.0 | 0.3 | GO:0014850 | response to muscle activity(GO:0014850) |
| 0.0 | 0.0 | GO:0034086 | maintenance of sister chromatid cohesion(GO:0034086) maintenance of mitotic sister chromatid cohesion(GO:0034088) |
| 0.0 | 4.5 | GO:0009615 | response to virus(GO:0009615) |
| 0.0 | 0.2 | GO:2000060 | positive regulation of protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:2000060) |
| 0.0 | 0.1 | GO:0021691 | cerebellar Purkinje cell layer maturation(GO:0021691) |
| 0.0 | 0.3 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.0 | 1.4 | GO:0006364 | rRNA processing(GO:0006364) |
| 0.0 | 0.3 | GO:0048147 | negative regulation of fibroblast proliferation(GO:0048147) |
| 0.0 | 1.2 | GO:0045766 | positive regulation of angiogenesis(GO:0045766) |
| 0.0 | 0.1 | GO:0051315 | attachment of mitotic spindle microtubules to kinetochore(GO:0051315) |
| 0.0 | 0.0 | GO:0061355 | Wnt protein secretion(GO:0061355) regulation of Wnt protein secretion(GO:0061356) positive regulation of Wnt protein secretion(GO:0061357) |
| 0.0 | 0.1 | GO:2000427 | positive regulation of apoptotic cell clearance(GO:2000427) |
| 0.0 | 0.2 | GO:0019367 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.0 | 0.2 | GO:0090435 | protein localization to nuclear envelope(GO:0090435) |
| 0.0 | 0.3 | GO:0071578 | zinc II ion transmembrane import(GO:0071578) |
| 0.0 | 0.1 | GO:0071340 | skeletal muscle acetylcholine-gated channel clustering(GO:0071340) |
| 0.0 | 0.3 | GO:0006884 | cell volume homeostasis(GO:0006884) |
| 0.0 | 0.2 | GO:0033014 | porphyrin-containing compound biosynthetic process(GO:0006779) tetrapyrrole biosynthetic process(GO:0033014) |
| 0.0 | 0.2 | GO:0007320 | insemination(GO:0007320) |
| 0.0 | 0.1 | GO:2000270 | negative regulation of fibroblast apoptotic process(GO:2000270) |
| 0.0 | 0.3 | GO:0035082 | axoneme assembly(GO:0035082) |
| 0.0 | 0.2 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.0 | 3.1 | GO:0042113 | B cell activation(GO:0042113) |
| 0.0 | 0.2 | GO:0009812 | flavonoid metabolic process(GO:0009812) |
| 0.0 | 0.1 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.0 | 0.1 | GO:0032957 | inositol trisphosphate metabolic process(GO:0032957) |
| 0.0 | 0.2 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.0 | 0.8 | GO:0006367 | transcription initiation from RNA polymerase II promoter(GO:0006367) |
| 0.0 | 0.1 | GO:0048853 | forebrain morphogenesis(GO:0048853) |
| 0.0 | 0.2 | GO:1900077 | negative regulation of insulin receptor signaling pathway(GO:0046627) negative regulation of cellular response to insulin stimulus(GO:1900077) |
| 0.0 | 0.5 | GO:0007029 | endoplasmic reticulum organization(GO:0007029) |
| 0.0 | 0.3 | GO:1902041 | regulation of extrinsic apoptotic signaling pathway via death domain receptors(GO:1902041) |
| 0.0 | 0.2 | GO:0051568 | histone H3-K4 methylation(GO:0051568) |
| 0.0 | 0.3 | GO:0042035 | regulation of cytokine biosynthetic process(GO:0042035) |
| 0.0 | 0.0 | GO:0044030 | regulation of DNA methylation(GO:0044030) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.2 | 9.6 | GO:0097224 | sperm connecting piece(GO:0097224) |
| 1.9 | 7.5 | GO:0036284 | tubulobulbar complex(GO:0036284) |
| 1.8 | 10.9 | GO:0097149 | centralspindlin complex(GO:0097149) |
| 1.8 | 5.3 | GO:1990667 | PCSK9-AnxA2 complex(GO:1990667) |
| 1.4 | 4.1 | GO:0097135 | cyclin E2-CDK2 complex(GO:0097135) |
| 1.4 | 16.2 | GO:0000796 | condensin complex(GO:0000796) |
| 1.3 | 12.0 | GO:0070776 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 1.0 | 5.1 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 1.0 | 3.0 | GO:0090537 | CERF complex(GO:0090537) |
| 1.0 | 2.9 | GO:0097132 | cyclin D2-CDK6 complex(GO:0097132) |
| 1.0 | 4.8 | GO:0030121 | AP-1 adaptor complex(GO:0030121) |
| 0.9 | 3.8 | GO:0016035 | zeta DNA polymerase complex(GO:0016035) |
| 0.9 | 3.8 | GO:0001740 | Barr body(GO:0001740) |
| 0.9 | 3.7 | GO:0031680 | G-protein beta/gamma-subunit complex(GO:0031680) |
| 0.9 | 0.9 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.9 | 10.6 | GO:0008278 | cohesin complex(GO:0008278) |
| 0.8 | 3.1 | GO:1990590 | ATF1-ATF4 transcription factor complex(GO:1990590) |
| 0.8 | 7.1 | GO:0001651 | dense fibrillar component(GO:0001651) |
| 0.8 | 5.4 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
| 0.8 | 4.6 | GO:0032133 | chromosome passenger complex(GO:0032133) |
| 0.8 | 3.1 | GO:0033186 | CAF-1 complex(GO:0033186) |
| 0.7 | 2.8 | GO:0097543 | ciliary inversin compartment(GO:0097543) |
| 0.7 | 9.2 | GO:0033256 | I-kappaB/NF-kappaB complex(GO:0033256) |
| 0.7 | 2.7 | GO:0005971 | ribonucleoside-diphosphate reductase complex(GO:0005971) |
| 0.7 | 3.3 | GO:0045160 | myosin I complex(GO:0045160) |
| 0.7 | 3.3 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 0.6 | 2.6 | GO:0045251 | mitochondrial electron transfer flavoprotein complex(GO:0017133) electron transfer flavoprotein complex(GO:0045251) |
| 0.6 | 1.9 | GO:1990423 | RZZ complex(GO:1990423) |
| 0.6 | 3.8 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.6 | 3.8 | GO:0031262 | Ndc80 complex(GO:0031262) |
| 0.6 | 4.2 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.6 | 2.9 | GO:0044614 | nuclear pore cytoplasmic filaments(GO:0044614) |
| 0.6 | 1.7 | GO:0020016 | ciliary pocket(GO:0020016) ciliary pocket membrane(GO:0020018) |
| 0.6 | 5.6 | GO:0072687 | meiotic spindle(GO:0072687) |
| 0.6 | 6.7 | GO:0031264 | death-inducing signaling complex(GO:0031264) |
| 0.5 | 2.2 | GO:0071942 | XPC complex(GO:0071942) |
| 0.5 | 3.2 | GO:0043564 | Ku70:Ku80 complex(GO:0043564) |
| 0.5 | 1.6 | GO:0032783 | ELL-EAF complex(GO:0032783) |
| 0.5 | 8.0 | GO:0005915 | zonula adherens(GO:0005915) |
| 0.5 | 4.7 | GO:0098576 | lumenal side of membrane(GO:0098576) |
| 0.5 | 5.5 | GO:0000808 | origin recognition complex(GO:0000808) nuclear origin of replication recognition complex(GO:0005664) |
| 0.5 | 3.0 | GO:0005672 | transcription factor TFIIA complex(GO:0005672) |
| 0.5 | 9.9 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.5 | 1.5 | GO:0070195 | growth hormone receptor complex(GO:0070195) |
| 0.5 | 2.4 | GO:0030312 | external encapsulating structure(GO:0030312) |
| 0.5 | 1.4 | GO:0000942 | condensed nuclear chromosome outer kinetochore(GO:0000942) |
| 0.5 | 1.4 | GO:0070435 | Shc-EGFR complex(GO:0070435) |
| 0.5 | 5.1 | GO:0070531 | BRCA1-A complex(GO:0070531) |
| 0.4 | 3.1 | GO:0097443 | sorting endosome(GO:0097443) |
| 0.4 | 1.3 | GO:0005900 | oncostatin-M receptor complex(GO:0005900) |
| 0.4 | 4.8 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.4 | 3.5 | GO:0042825 | TAP complex(GO:0042825) |
| 0.4 | 3.8 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 0.4 | 8.0 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.4 | 2.1 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.4 | 4.5 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.4 | 2.8 | GO:0099524 | region of cytosol(GO:0099522) postsynaptic cytosol(GO:0099524) |
| 0.4 | 0.8 | GO:0034679 | integrin alpha9-beta1 complex(GO:0034679) |
| 0.4 | 4.3 | GO:0070761 | pre-snoRNP complex(GO:0070761) |
| 0.4 | 3.9 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.4 | 3.4 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.4 | 8.7 | GO:1990023 | mitotic spindle midzone(GO:1990023) |
| 0.4 | 4.0 | GO:0070578 | RISC-loading complex(GO:0070578) |
| 0.4 | 5.5 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.3 | 2.0 | GO:0070449 | elongin complex(GO:0070449) |
| 0.3 | 1.7 | GO:0044194 | cytolytic granule(GO:0044194) |
| 0.3 | 1.3 | GO:1990716 | axonemal central apparatus(GO:1990716) |
| 0.3 | 1.6 | GO:0071953 | elastic fiber(GO:0071953) |
| 0.3 | 1.0 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 0.3 | 8.8 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.3 | 2.3 | GO:0005726 | perichromatin fibrils(GO:0005726) |
| 0.3 | 2.9 | GO:0033061 | DNA recombinase mediator complex(GO:0033061) |
| 0.3 | 4.5 | GO:0000138 | Golgi trans cisterna(GO:0000138) |
| 0.3 | 1.0 | GO:1990630 | IRE1-RACK1-PP2A complex(GO:1990630) |
| 0.3 | 7.0 | GO:0090545 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.3 | 0.9 | GO:0032156 | septin cytoskeleton(GO:0032156) |
| 0.3 | 6.9 | GO:0031011 | Ino80 complex(GO:0031011) |
| 0.3 | 1.9 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.3 | 3.7 | GO:0000801 | central element(GO:0000801) |
| 0.3 | 0.9 | GO:0034455 | t-UTP complex(GO:0034455) |
| 0.3 | 2.1 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.3 | 1.5 | GO:0035363 | histone locus body(GO:0035363) |
| 0.3 | 1.5 | GO:0010370 | perinucleolar chromocenter(GO:0010370) |
| 0.3 | 0.9 | GO:0097361 | CIA complex(GO:0097361) |
| 0.3 | 5.3 | GO:0005797 | Golgi medial cisterna(GO:0005797) |
| 0.3 | 0.8 | GO:0060187 | cell pole(GO:0060187) |
| 0.3 | 0.8 | GO:0034666 | integrin alpha2-beta1 complex(GO:0034666) |
| 0.3 | 4.3 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.3 | 0.8 | GO:0005668 | RNA polymerase transcription factor SL1 complex(GO:0005668) |
| 0.3 | 4.3 | GO:0090543 | Flemming body(GO:0090543) |
| 0.3 | 1.8 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.3 | 4.7 | GO:0000346 | transcription export complex(GO:0000346) |
| 0.3 | 2.6 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.3 | 4.1 | GO:0097431 | mitotic spindle pole(GO:0097431) |
| 0.3 | 4.3 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.3 | 0.5 | GO:0090661 | box H/ACA telomerase RNP complex(GO:0090661) |
| 0.2 | 8.8 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.2 | 1.5 | GO:0042583 | chromaffin granule(GO:0042583) |
| 0.2 | 3.1 | GO:0097524 | sperm plasma membrane(GO:0097524) |
| 0.2 | 1.7 | GO:0005638 | lamin filament(GO:0005638) |
| 0.2 | 5.5 | GO:0000777 | condensed chromosome kinetochore(GO:0000777) |
| 0.2 | 1.7 | GO:0005785 | signal recognition particle receptor complex(GO:0005785) |
| 0.2 | 2.4 | GO:0000812 | Swr1 complex(GO:0000812) |
| 0.2 | 1.7 | GO:0070522 | ERCC4-ERCC1 complex(GO:0070522) |
| 0.2 | 0.9 | GO:0035061 | interchromatin granule(GO:0035061) |
| 0.2 | 1.2 | GO:0043293 | apoptosome(GO:0043293) |
| 0.2 | 2.8 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 0.2 | 13.6 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.2 | 2.9 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.2 | 3.3 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.2 | 1.8 | GO:0071204 | histone pre-mRNA 3'end processing complex(GO:0071204) |
| 0.2 | 0.4 | GO:0071817 | MMXD complex(GO:0071817) |
| 0.2 | 1.3 | GO:1990356 | sumoylated E2 ligase complex(GO:1990356) |
| 0.2 | 0.6 | GO:0032311 | angiogenin-PRI complex(GO:0032311) |
| 0.2 | 0.9 | GO:1990794 | lateral part of cell(GO:0097574) basolateral part of cell(GO:1990794) rod bipolar cell terminal bouton(GO:1990795) |
| 0.2 | 4.1 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.2 | 3.2 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 0.2 | 1.7 | GO:0033162 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.2 | 1.1 | GO:0097425 | smooth endoplasmic reticulum membrane(GO:0030868) smooth endoplasmic reticulum part(GO:0097425) |
| 0.2 | 0.8 | GO:0043541 | UDP-N-acetylglucosamine transferase complex(GO:0043541) |
| 0.2 | 1.5 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.2 | 3.3 | GO:0001673 | male germ cell nucleus(GO:0001673) |
| 0.2 | 1.2 | GO:0030981 | cortical microtubule cytoskeleton(GO:0030981) |
| 0.2 | 21.1 | GO:0005811 | lipid particle(GO:0005811) |
| 0.2 | 14.9 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.2 | 2.2 | GO:0097542 | ciliary tip(GO:0097542) |
| 0.2 | 2.8 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.2 | 3.4 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.2 | 2.2 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
| 0.2 | 3.7 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.2 | 0.8 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.2 | 4.8 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.2 | 3.3 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.2 | 7.9 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.2 | 1.7 | GO:0045298 | tubulin complex(GO:0045298) |
| 0.2 | 3.3 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.2 | 0.7 | GO:1990578 | perinuclear endoplasmic reticulum membrane(GO:1990578) |
| 0.2 | 3.1 | GO:0044815 | DNA packaging complex(GO:0044815) |
| 0.2 | 1.8 | GO:0031390 | Ctf18 RFC-like complex(GO:0031390) |
| 0.2 | 9.0 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.2 | 1.2 | GO:0031205 | endoplasmic reticulum Sec complex(GO:0031205) |
| 0.2 | 3.8 | GO:0016514 | SWI/SNF complex(GO:0016514) |
| 0.2 | 4.3 | GO:0070938 | contractile ring(GO:0070938) |
| 0.2 | 2.1 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.2 | 1.2 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.2 | 0.3 | GO:0034750 | Scrib-APC-beta-catenin complex(GO:0034750) |
| 0.2 | 0.3 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.2 | 4.2 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.2 | 0.5 | GO:0030905 | retromer, tubulation complex(GO:0030905) |
| 0.2 | 1.0 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.2 | 4.4 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.2 | 2.1 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.2 | 4.1 | GO:0005849 | mRNA cleavage factor complex(GO:0005849) |
| 0.2 | 4.2 | GO:0071339 | MLL1/2 complex(GO:0044665) MLL1 complex(GO:0071339) |
| 0.2 | 2.3 | GO:1902562 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.2 | 1.1 | GO:0016342 | catenin complex(GO:0016342) |
| 0.2 | 0.3 | GO:0031314 | extrinsic component of mitochondrial inner membrane(GO:0031314) |
| 0.2 | 0.5 | GO:0017109 | glutamate-cysteine ligase complex(GO:0017109) |
| 0.2 | 0.6 | GO:0071149 | TEAD-2-YAP complex(GO:0071149) |
| 0.1 | 2.3 | GO:0031091 | platelet alpha granule(GO:0031091) |
| 0.1 | 0.7 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.1 | 2.6 | GO:0036038 | MKS complex(GO:0036038) |
| 0.1 | 1.3 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.1 | 4.4 | GO:0031231 | intrinsic component of peroxisomal membrane(GO:0031231) |
| 0.1 | 2.8 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.1 | 1.7 | GO:0036128 | CatSper complex(GO:0036128) |
| 0.1 | 3.2 | GO:0035686 | sperm fibrous sheath(GO:0035686) |
| 0.1 | 24.2 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.1 | 2.9 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.1 | 0.3 | GO:0043224 | nuclear SCF ubiquitin ligase complex(GO:0043224) |
| 0.1 | 2.6 | GO:0030430 | host cell cytoplasm(GO:0030430) host cell cytoplasm part(GO:0033655) |
| 0.1 | 3.8 | GO:0005605 | basal lamina(GO:0005605) |
| 0.1 | 0.5 | GO:0002111 | BRCA2-BRAF35 complex(GO:0002111) |
| 0.1 | 2.0 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.1 | 5.1 | GO:0005657 | replication fork(GO:0005657) |
| 0.1 | 1.8 | GO:0042555 | MCM complex(GO:0042555) |
| 0.1 | 50.0 | GO:0000790 | nuclear chromatin(GO:0000790) |
| 0.1 | 9.6 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.1 | 1.0 | GO:0097129 | cyclin D2-CDK4 complex(GO:0097129) |
| 0.1 | 0.5 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.1 | 1.1 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.1 | 0.2 | GO:0034684 | integrin alphav-beta5 complex(GO:0034684) |
| 0.1 | 2.6 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.1 | 0.3 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.1 | 3.7 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.1 | 3.1 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.1 | 1.8 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.1 | 1.7 | GO:0071203 | WASH complex(GO:0071203) |
| 0.1 | 1.7 | GO:0044232 | organelle membrane contact site(GO:0044232) |
| 0.1 | 0.8 | GO:0097441 | basilar dendrite(GO:0097441) |
| 0.1 | 0.3 | GO:0043291 | RAVE complex(GO:0043291) |
| 0.1 | 0.8 | GO:0008540 | proteasome regulatory particle, base subcomplex(GO:0008540) |
| 0.1 | 2.0 | GO:0032039 | integrator complex(GO:0032039) |
| 0.1 | 1.7 | GO:0030914 | STAGA complex(GO:0030914) |
| 0.1 | 4.6 | GO:0000123 | histone acetyltransferase complex(GO:0000123) |
| 0.1 | 2.3 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.1 | 0.6 | GO:0019773 | proteasome core complex, alpha-subunit complex(GO:0019773) |
| 0.1 | 1.4 | GO:0043186 | P granule(GO:0043186) pole plasm(GO:0045495) germ plasm(GO:0060293) |
| 0.1 | 1.0 | GO:0002177 | manchette(GO:0002177) |
| 0.1 | 1.6 | GO:0001741 | XY body(GO:0001741) |
| 0.1 | 11.3 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.1 | 2.3 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.1 | 0.7 | GO:0030289 | protein phosphatase 4 complex(GO:0030289) |
| 0.1 | 28.2 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.1 | 0.3 | GO:0034066 | RIC1-RGP1 guanyl-nucleotide exchange factor complex(GO:0034066) |
| 0.1 | 0.3 | GO:0071001 | U4/U6 snRNP(GO:0071001) |
| 0.1 | 0.8 | GO:0010369 | chromocenter(GO:0010369) |
| 0.1 | 3.7 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.1 | 2.8 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.1 | 0.4 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.1 | 3.0 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.1 | 2.1 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.1 | 1.1 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.1 | 0.3 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.1 | 0.6 | GO:0072357 | PTW/PP1 phosphatase complex(GO:0072357) |
| 0.1 | 3.0 | GO:0015030 | Cajal body(GO:0015030) |
| 0.1 | 4.4 | GO:0044439 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) microbody part(GO:0044438) peroxisomal part(GO:0044439) |
| 0.1 | 0.7 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.1 | 1.8 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.1 | 0.6 | GO:0097504 | Gemini of coiled bodies(GO:0097504) |
| 0.1 | 0.8 | GO:0097546 | ciliary base(GO:0097546) |
| 0.1 | 2.2 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.1 | 1.2 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.1 | 0.8 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 0.1 | 0.3 | GO:0098835 | presynaptic endocytic zone(GO:0098833) presynaptic endocytic zone membrane(GO:0098835) |
| 0.1 | 1.5 | GO:0051233 | spindle midzone(GO:0051233) |
| 0.1 | 4.6 | GO:0016459 | myosin complex(GO:0016459) |
| 0.1 | 23.9 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.1 | 8.4 | GO:0030496 | midbody(GO:0030496) |
| 0.1 | 5.8 | GO:0016605 | PML body(GO:0016605) |
| 0.1 | 10.6 | GO:0031514 | motile cilium(GO:0031514) |
| 0.1 | 1.2 | GO:0032993 | protein-DNA complex(GO:0032993) |
| 0.1 | 0.4 | GO:0000308 | cytoplasmic cyclin-dependent protein kinase holoenzyme complex(GO:0000308) |
| 0.1 | 0.4 | GO:0005587 | collagen type IV trimer(GO:0005587) network-forming collagen trimer(GO:0098642) collagen network(GO:0098645) basement membrane collagen trimer(GO:0098651) |
| 0.1 | 3.3 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.1 | 0.6 | GO:0071818 | BAT3 complex(GO:0071818) ER membrane insertion complex(GO:0072379) |
| 0.1 | 1.3 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
| 0.1 | 0.7 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.1 | 1.0 | GO:0000792 | heterochromatin(GO:0000792) |
| 0.1 | 1.0 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 0.1 | 1.0 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.1 | 10.3 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.1 | 0.3 | GO:0000839 | Hrd1p ubiquitin ligase ERAD-L complex(GO:0000839) |
| 0.1 | 1.0 | GO:1990391 | DNA repair complex(GO:1990391) |
| 0.1 | 0.4 | GO:0005827 | polar microtubule(GO:0005827) |
| 0.1 | 0.2 | GO:0042720 | mitochondrial inner membrane peptidase complex(GO:0042720) |
| 0.1 | 4.1 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.1 | 0.1 | GO:0071598 | neuronal ribonucleoprotein granule(GO:0071598) |
| 0.1 | 4.2 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.1 | 0.3 | GO:0032584 | growth cone membrane(GO:0032584) |
| 0.1 | 0.3 | GO:1990130 | Iml1 complex(GO:1990130) |
| 0.1 | 6.1 | GO:0005777 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.1 | 0.5 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.1 | 3.0 | GO:0031430 | M band(GO:0031430) |
| 0.1 | 0.6 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.1 | 0.3 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.1 | 0.7 | GO:0097136 | Bcl-2 family protein complex(GO:0097136) |
| 0.1 | 2.3 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.1 | 0.3 | GO:0034709 | methylosome(GO:0034709) |
| 0.1 | 0.1 | GO:0034457 | Mpp10 complex(GO:0034457) |
| 0.1 | 2.2 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.1 | 14.2 | GO:0016604 | nuclear body(GO:0016604) |
| 0.1 | 2.0 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.1 | 0.5 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.1 | 4.1 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.1 | 25.2 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.1 | 0.7 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.0 | 0.2 | GO:0030896 | checkpoint clamp complex(GO:0030896) |
| 0.0 | 0.4 | GO:0005839 | proteasome core complex(GO:0005839) |
| 0.0 | 5.5 | GO:0097223 | sperm part(GO:0097223) |
| 0.0 | 9.2 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 0.1 | GO:0031467 | Cul7-RING ubiquitin ligase complex(GO:0031467) |
| 0.0 | 0.2 | GO:0097197 | tetraspanin-enriched microdomain(GO:0097197) |
| 0.0 | 0.3 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 0.0 | 1.4 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.0 | 2.5 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.0 | 0.2 | GO:0045273 | mitochondrial respiratory chain complex II, succinate dehydrogenase complex (ubiquinone)(GO:0005749) succinate dehydrogenase complex (ubiquinone)(GO:0045257) respiratory chain complex II(GO:0045273) succinate dehydrogenase complex(GO:0045281) fumarate reductase complex(GO:0045283) |
| 0.0 | 1.7 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 5.0 | GO:0097517 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.0 | 1.0 | GO:0000145 | exocyst(GO:0000145) |
| 0.0 | 0.2 | GO:0070032 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin I complex(GO:0070032) synaptobrevin 2-SNAP-25-syntaxin-1a-complexin II complex(GO:0070033) |
| 0.0 | 1.1 | GO:0034385 | very-low-density lipoprotein particle(GO:0034361) triglyceride-rich lipoprotein particle(GO:0034385) |
| 0.0 | 3.1 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.0 | 0.2 | GO:0033179 | proton-transporting V-type ATPase, V0 domain(GO:0033179) |
| 0.0 | 2.2 | GO:0044291 | cell-cell contact zone(GO:0044291) |
| 0.0 | 0.7 | GO:0002102 | podosome(GO:0002102) |
| 0.0 | 0.4 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.0 | 0.2 | GO:0005745 | m-AAA complex(GO:0005745) |
| 0.0 | 1.7 | GO:0000781 | chromosome, telomeric region(GO:0000781) |
| 0.0 | 1.7 | GO:0000502 | proteasome complex(GO:0000502) |
| 0.0 | 0.6 | GO:0045120 | pronucleus(GO:0045120) |
| 0.0 | 0.8 | GO:0030057 | desmosome(GO:0030057) |
| 0.0 | 67.4 | GO:0005654 | nucleoplasm(GO:0005654) |
| 0.0 | 0.3 | GO:0097422 | tubular endosome(GO:0097422) |
| 0.0 | 0.5 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 0.0 | 0.3 | GO:0036156 | inner dynein arm(GO:0036156) |
| 0.0 | 0.2 | GO:0017177 | glucosidase II complex(GO:0017177) |
| 0.0 | 2.8 | GO:0005901 | caveola(GO:0005901) |
| 0.0 | 0.3 | GO:0045239 | tricarboxylic acid cycle enzyme complex(GO:0045239) |
| 0.0 | 1.8 | GO:0005930 | axoneme(GO:0005930) |
| 0.0 | 2.3 | GO:0032420 | stereocilium(GO:0032420) |
| 0.0 | 0.4 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.0 | 1.4 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.0 | 0.0 | GO:0055087 | Ski complex(GO:0055087) |
| 0.0 | 3.0 | GO:0030018 | Z disc(GO:0030018) |
| 0.0 | 0.5 | GO:0030665 | clathrin-coated vesicle membrane(GO:0030665) |
| 0.0 | 0.1 | GO:0036020 | endolysosome membrane(GO:0036020) |
| 0.0 | 0.4 | GO:0035869 | ciliary transition zone(GO:0035869) |
| 0.0 | 2.7 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 1.0 | GO:0030864 | cortical actin cytoskeleton(GO:0030864) |
| 0.0 | 0.2 | GO:0044754 | autolysosome(GO:0044754) |
| 0.0 | 7.2 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 0.8 | GO:0005903 | brush border(GO:0005903) |
| 0.0 | 0.4 | GO:0016471 | vacuolar proton-transporting V-type ATPase complex(GO:0016471) |
| 0.0 | 0.2 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.0 | 0.5 | GO:0005922 | connexon complex(GO:0005922) |
| 0.0 | 0.1 | GO:0071004 | U2-type prespliceosome(GO:0071004) |
| 0.0 | 0.3 | GO:0005852 | eukaryotic translation initiation factor 3 complex(GO:0005852) |
| 0.0 | 0.4 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.0 | 57.0 | GO:0005634 | nucleus(GO:0005634) |
| 0.0 | 0.0 | GO:0045254 | pyruvate dehydrogenase complex(GO:0045254) |
| 0.0 | 0.1 | GO:0032391 | photoreceptor connecting cilium(GO:0032391) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.6 | 7.8 | GO:0005369 | beta-alanine transmembrane transporter activity(GO:0001761) taurine transmembrane transporter activity(GO:0005368) taurine:sodium symporter activity(GO:0005369) |
| 2.4 | 7.1 | GO:0042602 | riboflavin reductase (NADPH) activity(GO:0042602) |
| 1.9 | 5.7 | GO:0047936 | glucose 1-dehydrogenase [NAD(P)] activity(GO:0047936) |
| 1.8 | 5.3 | GO:0004572 | mannosyl-oligosaccharide 1,3-1,6-alpha-mannosidase activity(GO:0004572) |
| 1.8 | 5.3 | GO:0019978 | interleukin-3 receptor activity(GO:0004912) interleukin-3 binding(GO:0019978) |
| 1.7 | 5.2 | GO:0004771 | sterol esterase activity(GO:0004771) |
| 1.7 | 5.1 | GO:0019107 | glycylpeptide N-tetradecanoyltransferase activity(GO:0004379) myristoyltransferase activity(GO:0019107) |
| 1.6 | 4.9 | GO:0003896 | DNA primase activity(GO:0003896) |
| 1.6 | 4.8 | GO:0016005 | phospholipase A2 activator activity(GO:0016005) |
| 1.6 | 4.7 | GO:0033680 | ATP-dependent DNA/RNA helicase activity(GO:0033680) |
| 1.5 | 6.0 | GO:0004052 | arachidonate 12-lipoxygenase activity(GO:0004052) |
| 1.5 | 4.4 | GO:0050510 | N-acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase activity(GO:0050510) |
| 1.5 | 4.4 | GO:1902121 | lithocholic acid binding(GO:1902121) |
| 1.5 | 7.3 | GO:0043758 | acetate-CoA ligase (ADP-forming) activity(GO:0043758) |
| 1.4 | 4.2 | GO:0035851 | Krueppel-associated box domain binding(GO:0035851) |
| 1.4 | 9.6 | GO:0032027 | myosin light chain binding(GO:0032027) |
| 1.2 | 3.6 | GO:0005302 | L-tyrosine transmembrane transporter activity(GO:0005302) |
| 1.2 | 3.5 | GO:0016872 | inositol-3-phosphate synthase activity(GO:0004512) intramolecular lyase activity(GO:0016872) |
| 1.2 | 3.5 | GO:0031531 | thyrotropin-releasing hormone receptor binding(GO:0031531) |
| 1.1 | 4.5 | GO:0015157 | sucrose:proton symporter activity(GO:0008506) sucrose transmembrane transporter activity(GO:0008515) disaccharide transmembrane transporter activity(GO:0015154) oligosaccharide transmembrane transporter activity(GO:0015157) |
| 1.1 | 6.7 | GO:0017077 | oxidative phosphorylation uncoupler activity(GO:0017077) |
| 1.0 | 4.1 | GO:0015403 | thiamine uptake transmembrane transporter activity(GO:0015403) |
| 1.0 | 2.0 | GO:0045142 | triplex DNA binding(GO:0045142) |
| 1.0 | 2.9 | GO:0036009 | protein-glutamine N-methyltransferase activity(GO:0036009) histone-glutamine methyltransferase activity(GO:1990259) |
| 1.0 | 9.6 | GO:0005114 | type II transforming growth factor beta receptor binding(GO:0005114) |
| 1.0 | 2.9 | GO:0008476 | protein-tyrosine sulfotransferase activity(GO:0008476) |
| 0.9 | 2.6 | GO:0070773 | protein-N-terminal glutamine amidohydrolase activity(GO:0070773) |
| 0.8 | 1.6 | GO:0015142 | citrate transmembrane transporter activity(GO:0015137) tricarboxylic acid transmembrane transporter activity(GO:0015142) |
| 0.8 | 3.3 | GO:0030629 | U6 snRNA 3'-end binding(GO:0030629) |
| 0.8 | 3.2 | GO:0070976 | calcium-independent protein kinase C activity(GO:0004699) TIR domain binding(GO:0070976) |
| 0.8 | 3.2 | GO:0048030 | disaccharide binding(GO:0048030) |
| 0.8 | 3.1 | GO:0034647 | histone demethylase activity (H3-trimethyl-K4 specific)(GO:0034647) |
| 0.8 | 7.0 | GO:0019834 | phospholipase A2 inhibitor activity(GO:0019834) |
| 0.8 | 1.5 | GO:0015173 | aromatic amino acid transmembrane transporter activity(GO:0015173) |
| 0.8 | 3.1 | GO:0086038 | calcium:sodium antiporter activity involved in regulation of cardiac muscle cell membrane potential(GO:0086038) |
| 0.8 | 3.0 | GO:0070506 | high-density lipoprotein particle receptor activity(GO:0070506) |
| 0.7 | 6.7 | GO:0071535 | RING-like zinc finger domain binding(GO:0071535) |
| 0.7 | 4.4 | GO:0038085 | vascular endothelial growth factor binding(GO:0038085) |
| 0.7 | 2.9 | GO:0051435 | BH4 domain binding(GO:0051435) |
| 0.7 | 2.2 | GO:0001716 | L-amino-acid oxidase activity(GO:0001716) |
| 0.7 | 2.9 | GO:0098770 | FBXO family protein binding(GO:0098770) |
| 0.7 | 2.2 | GO:0004514 | nicotinate-nucleotide diphosphorylase (carboxylating) activity(GO:0004514) |
| 0.7 | 10.1 | GO:0000014 | single-stranded DNA endodeoxyribonuclease activity(GO:0000014) |
| 0.7 | 5.0 | GO:0004704 | NF-kappaB-inducing kinase activity(GO:0004704) |
| 0.7 | 0.7 | GO:0017099 | very-long-chain-acyl-CoA dehydrogenase activity(GO:0017099) |
| 0.7 | 5.6 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.7 | 7.0 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.7 | 2.1 | GO:0017116 | single-stranded DNA-dependent ATP-dependent DNA helicase activity(GO:0017116) |
| 0.7 | 4.8 | GO:0042134 | rRNA primary transcript binding(GO:0042134) |
| 0.7 | 0.7 | GO:0005128 | erythropoietin receptor binding(GO:0005128) |
| 0.6 | 3.2 | GO:0008427 | calcium-dependent protein kinase inhibitor activity(GO:0008427) |
| 0.6 | 4.5 | GO:0070404 | NADH binding(GO:0070404) |
| 0.6 | 3.1 | GO:0016649 | electron-transferring-flavoprotein dehydrogenase activity(GO:0004174) oxidoreductase activity, acting on the CH-NH group of donors, quinone or similar compound as acceptor(GO:0016649) |
| 0.6 | 1.9 | GO:0001160 | transcription termination site sequence-specific DNA binding(GO:0001147) transcription termination site DNA binding(GO:0001160) |
| 0.6 | 2.5 | GO:0031699 | beta-3 adrenergic receptor binding(GO:0031699) |
| 0.6 | 2.5 | GO:0004450 | isocitrate dehydrogenase (NADP+) activity(GO:0004450) |
| 0.6 | 2.4 | GO:0071207 | histone pre-mRNA stem-loop binding(GO:0071207) |
| 0.6 | 1.2 | GO:0004345 | glucose-6-phosphate dehydrogenase activity(GO:0004345) |
| 0.6 | 3.6 | GO:0016416 | carnitine O-palmitoyltransferase activity(GO:0004095) O-palmitoyltransferase activity(GO:0016416) |
| 0.6 | 2.4 | GO:0070991 | medium-chain-acyl-CoA dehydrogenase activity(GO:0070991) |
| 0.6 | 13.5 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.6 | 4.7 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.6 | 1.7 | GO:0098918 | structural constituent of synapse(GO:0098918) structural constituent of postsynaptic actin cytoskeleton(GO:0098973) |
| 0.6 | 2.3 | GO:0004176 | ATP-dependent peptidase activity(GO:0004176) |
| 0.6 | 1.7 | GO:0030348 | syntaxin-3 binding(GO:0030348) |
| 0.6 | 9.0 | GO:1990405 | protein antigen binding(GO:1990405) |
| 0.6 | 1.7 | GO:0004454 | ketohexokinase activity(GO:0004454) |
| 0.5 | 9.7 | GO:0004791 | thioredoxin-disulfide reductase activity(GO:0004791) |
| 0.5 | 2.1 | GO:0004103 | choline kinase activity(GO:0004103) |
| 0.5 | 5.8 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 0.5 | 5.8 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 0.5 | 3.2 | GO:0004515 | nicotinamide-nucleotide adenylyltransferase activity(GO:0000309) nicotinate-nucleotide adenylyltransferase activity(GO:0004515) |
| 0.5 | 1.6 | GO:0008456 | alpha-N-acetylgalactosaminidase activity(GO:0008456) |
| 0.5 | 1.6 | GO:0004461 | lactose synthase activity(GO:0004461) |
| 0.5 | 3.1 | GO:0003938 | IMP dehydrogenase activity(GO:0003938) |
| 0.5 | 2.1 | GO:0016744 | transferase activity, transferring aldehyde or ketonic groups(GO:0016744) |
| 0.5 | 5.2 | GO:0003909 | DNA ligase activity(GO:0003909) DNA ligase (ATP) activity(GO:0003910) |
| 0.5 | 2.1 | GO:0031721 | haptoglobin binding(GO:0031720) hemoglobin alpha binding(GO:0031721) |
| 0.5 | 1.5 | GO:0032767 | copper-dependent protein binding(GO:0032767) |
| 0.5 | 2.5 | GO:0003829 | beta-1,3-galactosyl-O-glycosyl-glycoprotein beta-1,6-N-acetylglucosaminyltransferase activity(GO:0003829) |
| 0.5 | 3.0 | GO:0046404 | ATP-dependent polydeoxyribonucleotide 5'-hydroxyl-kinase activity(GO:0046404) polydeoxyribonucleotide kinase activity(GO:0051733) ATP-dependent polynucleotide kinase activity(GO:0051734) |
| 0.5 | 1.5 | GO:0061649 | ubiquitinated histone binding(GO:0061649) |
| 0.5 | 2.0 | GO:0004008 | copper-exporting ATPase activity(GO:0004008) copper-transporting ATPase activity(GO:0043682) |
| 0.5 | 3.5 | GO:0046979 | TAP1 binding(GO:0046978) TAP2 binding(GO:0046979) |
| 0.5 | 2.0 | GO:0031752 | D5 dopamine receptor binding(GO:0031752) |
| 0.5 | 1.5 | GO:0004903 | growth hormone receptor activity(GO:0004903) |
| 0.5 | 3.4 | GO:1990254 | keratin filament binding(GO:1990254) |
| 0.5 | 7.3 | GO:0031996 | thioesterase binding(GO:0031996) |
| 0.5 | 2.9 | GO:0070644 | vitamin D response element binding(GO:0070644) |
| 0.5 | 3.3 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.5 | 4.7 | GO:0008420 | CTD phosphatase activity(GO:0008420) |
| 0.5 | 3.2 | GO:0004957 | prostaglandin E receptor activity(GO:0004957) |
| 0.4 | 1.3 | GO:0004637 | phosphoribosylamine-glycine ligase activity(GO:0004637) |
| 0.4 | 3.9 | GO:0030368 | interleukin-17 receptor activity(GO:0030368) |
| 0.4 | 3.0 | GO:0004991 | parathyroid hormone receptor activity(GO:0004991) |
| 0.4 | 1.3 | GO:0016882 | cyclo-ligase activity(GO:0016882) 5-formyltetrahydrofolate cyclo-ligase activity(GO:0030272) |
| 0.4 | 1.3 | GO:0034188 | apolipoprotein receptor activity(GO:0030226) apolipoprotein A-I receptor activity(GO:0034188) phosphatidylserine-translocating ATPase activity(GO:0090556) |
| 0.4 | 1.3 | GO:0005087 | Ran guanyl-nucleotide exchange factor activity(GO:0005087) |
| 0.4 | 1.7 | GO:0015141 | succinate transmembrane transporter activity(GO:0015141) |
| 0.4 | 1.6 | GO:0000405 | bubble DNA binding(GO:0000405) |
| 0.4 | 6.1 | GO:0005168 | neurotrophin TRKA receptor binding(GO:0005168) |
| 0.4 | 2.4 | GO:0015199 | amino-acid betaine transmembrane transporter activity(GO:0015199) carnitine transmembrane transporter activity(GO:0015226) |
| 0.4 | 2.0 | GO:0045322 | unmethylated CpG binding(GO:0045322) |
| 0.4 | 5.2 | GO:0022889 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.4 | 6.3 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.4 | 1.6 | GO:0019136 | deoxynucleoside kinase activity(GO:0019136) |
| 0.4 | 1.2 | GO:0035175 | histone kinase activity (H3-S10 specific)(GO:0035175) |
| 0.4 | 1.9 | GO:0000700 | mismatch base pair DNA N-glycosylase activity(GO:0000700) |
| 0.4 | 3.1 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.4 | 1.1 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 0.4 | 0.8 | GO:0032357 | oxidized DNA binding(GO:0032356) oxidized purine DNA binding(GO:0032357) |
| 0.4 | 2.2 | GO:0031782 | melanocortin receptor binding(GO:0031779) type 3 melanocortin receptor binding(GO:0031781) type 4 melanocortin receptor binding(GO:0031782) |
| 0.4 | 4.4 | GO:0003691 | double-stranded telomeric DNA binding(GO:0003691) |
| 0.4 | 18.1 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.4 | 20.9 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.4 | 6.1 | GO:0070513 | death domain binding(GO:0070513) |
| 0.4 | 0.7 | GO:0010385 | double-stranded methylated DNA binding(GO:0010385) |
| 0.4 | 4.2 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.3 | 7.3 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.3 | 3.8 | GO:0030976 | thiamine pyrophosphate binding(GO:0030976) |
| 0.3 | 11.0 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.3 | 4.1 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.3 | 4.7 | GO:0008301 | DNA binding, bending(GO:0008301) |
| 0.3 | 1.0 | GO:0050354 | glycerone kinase activity(GO:0004371) FAD-AMP lyase (cyclizing) activity(GO:0034012) triokinase activity(GO:0050354) |
| 0.3 | 2.3 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.3 | 0.3 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 0.3 | 0.6 | GO:0042610 | CD8 receptor binding(GO:0042610) |
| 0.3 | 2.2 | GO:0001094 | TFIID-class transcription factor binding(GO:0001094) |
| 0.3 | 1.0 | GO:0004967 | glucagon receptor activity(GO:0004967) |
| 0.3 | 10.2 | GO:0042162 | telomeric DNA binding(GO:0042162) |
| 0.3 | 1.9 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 0.3 | 0.9 | GO:0034191 | apolipoprotein A-I receptor binding(GO:0034191) |
| 0.3 | 2.5 | GO:0004726 | non-membrane spanning protein tyrosine phosphatase activity(GO:0004726) |
| 0.3 | 2.5 | GO:0008310 | single-stranded DNA 3'-5' exodeoxyribonuclease activity(GO:0008310) |
| 0.3 | 1.5 | GO:0052901 | polyamine oxidase activity(GO:0046592) spermine:oxygen oxidoreductase (spermidine-forming) activity(GO:0052901) |
| 0.3 | 0.9 | GO:0016824 | hydrolase activity, acting on acid halide bonds(GO:0016824) hydrolase activity, acting on acid halide bonds, in C-halide compounds(GO:0019120) alkylhalidase activity(GO:0047651) |
| 0.3 | 5.4 | GO:0008242 | omega peptidase activity(GO:0008242) |
| 0.3 | 0.9 | GO:0004326 | tetrahydrofolylpolyglutamate synthase activity(GO:0004326) |
| 0.3 | 0.9 | GO:0098640 | integrin binding involved in cell-matrix adhesion(GO:0098640) |
| 0.3 | 1.8 | GO:0005134 | interleukin-2 receptor binding(GO:0005134) |
| 0.3 | 10.0 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.3 | 1.5 | GO:0004165 | dodecenoyl-CoA delta-isomerase activity(GO:0004165) |
| 0.3 | 2.3 | GO:0008172 | S-methyltransferase activity(GO:0008172) |
| 0.3 | 1.5 | GO:0004301 | epoxide hydrolase activity(GO:0004301) |
| 0.3 | 1.2 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
| 0.3 | 1.2 | GO:0042979 | ornithine decarboxylase regulator activity(GO:0042979) |
| 0.3 | 1.7 | GO:0008269 | JAK pathway signal transduction adaptor activity(GO:0008269) |
| 0.3 | 7.2 | GO:0000400 | four-way junction DNA binding(GO:0000400) |
| 0.3 | 2.0 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.3 | 0.8 | GO:0004127 | cytidylate kinase activity(GO:0004127) |
| 0.3 | 23.9 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.3 | 1.6 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.3 | 2.2 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.3 | 2.9 | GO:0070181 | small ribosomal subunit rRNA binding(GO:0070181) |
| 0.3 | 5.9 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.3 | 0.8 | GO:0003918 | DNA topoisomerase type II (ATP-hydrolyzing) activity(GO:0003918) DNA topoisomerase II activity(GO:0061505) |
| 0.3 | 0.8 | GO:0019797 | procollagen-proline 3-dioxygenase activity(GO:0019797) |
| 0.3 | 2.4 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.3 | 0.3 | GO:0045030 | UTP-activated nucleotide receptor activity(GO:0045030) |
| 0.3 | 1.8 | GO:0031685 | adenosine receptor binding(GO:0031685) |
| 0.3 | 1.1 | GO:0003844 | 1,4-alpha-glucan branching enzyme activity(GO:0003844) |
| 0.3 | 7.8 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.3 | 1.8 | GO:0047179 | platelet-activating factor acetyltransferase activity(GO:0047179) |
| 0.3 | 7.7 | GO:0005337 | nucleoside transmembrane transporter activity(GO:0005337) |
| 0.3 | 2.1 | GO:0070883 | pre-miRNA binding(GO:0070883) |
| 0.2 | 0.5 | GO:0004616 | phosphogluconate dehydrogenase (decarboxylating) activity(GO:0004616) |
| 0.2 | 5.9 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.2 | 1.2 | GO:0004060 | arylamine N-acetyltransferase activity(GO:0004060) |
| 0.2 | 0.7 | GO:0004019 | adenylosuccinate synthase activity(GO:0004019) |
| 0.2 | 3.4 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.2 | 1.7 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 0.2 | 0.7 | GO:0070279 | vitamin B6 binding(GO:0070279) |
| 0.2 | 1.0 | GO:0003827 | alpha-1,3-mannosylglycoprotein 2-beta-N-acetylglucosaminyltransferase activity(GO:0003827) |
| 0.2 | 0.7 | GO:0005146 | leukemia inhibitory factor receptor binding(GO:0005146) |
| 0.2 | 2.9 | GO:0015280 | ligand-gated sodium channel activity(GO:0015280) |
| 0.2 | 1.4 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 0.2 | 3.1 | GO:0000900 | translation repressor activity, nucleic acid binding(GO:0000900) |
| 0.2 | 0.7 | GO:0003845 | 11-beta-hydroxysteroid dehydrogenase [NAD(P)] activity(GO:0003845) |
| 0.2 | 1.4 | GO:0008422 | beta-glucosidase activity(GO:0008422) |
| 0.2 | 2.3 | GO:1990189 | peptide-serine-N-acetyltransferase activity(GO:1990189) |
| 0.2 | 1.6 | GO:0047184 | 1-acylglycerophosphocholine O-acyltransferase activity(GO:0047184) |
| 0.2 | 2.4 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.2 | 2.9 | GO:0003688 | DNA replication origin binding(GO:0003688) |
| 0.2 | 1.3 | GO:0004924 | oncostatin-M receptor activity(GO:0004924) |
| 0.2 | 0.4 | GO:0043139 | 5'-3' DNA helicase activity(GO:0043139) |
| 0.2 | 1.1 | GO:0031711 | bradykinin receptor binding(GO:0031711) |
| 0.2 | 0.7 | GO:0016838 | carbon-oxygen lyase activity, acting on phosphates(GO:0016838) |
| 0.2 | 1.5 | GO:0002135 | CTP binding(GO:0002135) |
| 0.2 | 2.8 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.2 | 4.7 | GO:0043422 | protein kinase B binding(GO:0043422) |
| 0.2 | 20.4 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.2 | 2.1 | GO:0005375 | copper ion transmembrane transporter activity(GO:0005375) |
| 0.2 | 0.4 | GO:0097100 | supercoiled DNA binding(GO:0097100) |
| 0.2 | 2.1 | GO:0003899 | DNA-directed RNA polymerase activity(GO:0003899) RNA polymerase activity(GO:0034062) |
| 0.2 | 0.6 | GO:1990948 | ligase inhibitor activity(GO:0055104) ubiquitin ligase inhibitor activity(GO:1990948) |
| 0.2 | 1.5 | GO:0002060 | purine nucleobase binding(GO:0002060) |
| 0.2 | 1.2 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 0.2 | 1.0 | GO:0004045 | aminoacyl-tRNA hydrolase activity(GO:0004045) |
| 0.2 | 0.8 | GO:0050145 | nucleoside phosphate kinase activity(GO:0050145) |
| 0.2 | 4.8 | GO:0031491 | nucleosome binding(GO:0031491) |
| 0.2 | 1.0 | GO:0005124 | scavenger receptor binding(GO:0005124) |
| 0.2 | 7.4 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.2 | 0.8 | GO:0016708 | nitric oxide dioxygenase activity(GO:0008941) oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of two atoms of oxygen into one donor(GO:0016708) iron-cytochrome-c reductase activity(GO:0047726) |
| 0.2 | 3.5 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.2 | 1.0 | GO:0035877 | death effector domain binding(GO:0035877) |
| 0.2 | 0.4 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.2 | 2.7 | GO:0008199 | ferric iron binding(GO:0008199) |
| 0.2 | 2.9 | GO:0070742 | C2H2 zinc finger domain binding(GO:0070742) |
| 0.2 | 5.0 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.2 | 1.1 | GO:0033743 | peptide-methionine (R)-S-oxide reductase activity(GO:0033743) |
| 0.2 | 1.0 | GO:0097003 | adipokinetic hormone receptor activity(GO:0097003) |
| 0.2 | 2.9 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
| 0.2 | 0.8 | GO:0042284 | sphingolipid delta-4 desaturase activity(GO:0042284) |
| 0.2 | 10.0 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.2 | 10.2 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.2 | 0.9 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.2 | 0.6 | GO:0004392 | heme oxygenase (decyclizing) activity(GO:0004392) |
| 0.2 | 4.7 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.2 | 0.9 | GO:0046923 | ER retention sequence binding(GO:0046923) |
| 0.2 | 1.1 | GO:0001055 | RNA polymerase II activity(GO:0001055) |
| 0.2 | 4.7 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.2 | 0.9 | GO:0016997 | exo-alpha-sialidase activity(GO:0004308) alpha-sialidase activity(GO:0016997) exo-alpha-(2->3)-sialidase activity(GO:0052794) exo-alpha-(2->6)-sialidase activity(GO:0052795) exo-alpha-(2->8)-sialidase activity(GO:0052796) |
| 0.2 | 0.5 | GO:0030350 | iron-responsive element binding(GO:0030350) |
| 0.2 | 0.9 | GO:0034056 | estrogen response element binding(GO:0034056) |
| 0.2 | 5.6 | GO:0035173 | histone kinase activity(GO:0035173) |
| 0.2 | 2.5 | GO:0001164 | RNA polymerase I regulatory region sequence-specific DNA binding(GO:0001163) RNA polymerase I CORE element sequence-specific DNA binding(GO:0001164) |
| 0.2 | 2.7 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.2 | 2.8 | GO:0046030 | inositol-polyphosphate 5-phosphatase activity(GO:0004445) inositol trisphosphate phosphatase activity(GO:0046030) |
| 0.2 | 3.1 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.2 | 0.7 | GO:0004830 | tryptophan-tRNA ligase activity(GO:0004830) |
| 0.2 | 0.9 | GO:2001070 | starch binding(GO:2001070) |
| 0.2 | 2.4 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.2 | 1.0 | GO:0033188 | sphingomyelin synthase activity(GO:0033188) ceramide cholinephosphotransferase activity(GO:0047493) |
| 0.2 | 8.9 | GO:0050661 | NADP binding(GO:0050661) |
| 0.2 | 0.7 | GO:0019166 | trans-2-enoyl-CoA reductase (NADPH) activity(GO:0019166) |
| 0.2 | 2.0 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.2 | 0.8 | GO:0043515 | kinetochore binding(GO:0043515) |
| 0.2 | 2.3 | GO:0004000 | adenosine deaminase activity(GO:0004000) |
| 0.2 | 1.2 | GO:0047288 | monosialoganglioside sialyltransferase activity(GO:0047288) |
| 0.2 | 1.0 | GO:0055131 | C3HC4-type RING finger domain binding(GO:0055131) |
| 0.2 | 1.6 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
| 0.2 | 0.7 | GO:0035800 | deubiquitinase activator activity(GO:0035800) |
| 0.2 | 0.2 | GO:0010314 | phosphatidylinositol-5-phosphate binding(GO:0010314) |
| 0.2 | 1.0 | GO:0000980 | RNA polymerase II distal enhancer sequence-specific DNA binding(GO:0000980) |
| 0.2 | 1.0 | GO:0010997 | anaphase-promoting complex binding(GO:0010997) |
| 0.2 | 2.5 | GO:0015037 | peptide disulfide oxidoreductase activity(GO:0015037) |
| 0.2 | 11.1 | GO:0035591 | signaling adaptor activity(GO:0035591) |
| 0.2 | 0.8 | GO:0001225 | RNA polymerase II transcription coactivator binding(GO:0001225) |
| 0.2 | 2.8 | GO:0008484 | sulfuric ester hydrolase activity(GO:0008484) |
| 0.2 | 2.0 | GO:0005172 | vascular endothelial growth factor receptor binding(GO:0005172) |
| 0.2 | 1.7 | GO:0030346 | protein phosphatase 2B binding(GO:0030346) |
| 0.2 | 1.9 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.2 | 2.2 | GO:0008097 | 5S rRNA binding(GO:0008097) |
| 0.2 | 2.5 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.2 | 1.5 | GO:0030371 | translation repressor activity(GO:0030371) |
| 0.2 | 3.4 | GO:0032794 | GTPase activating protein binding(GO:0032794) |
| 0.2 | 1.4 | GO:0019238 | cyclohydrolase activity(GO:0019238) |
| 0.2 | 0.8 | GO:0030984 | kininogen binding(GO:0030984) |
| 0.2 | 1.2 | GO:0043560 | insulin receptor substrate binding(GO:0043560) |
| 0.2 | 0.8 | GO:0004528 | phosphodiesterase I activity(GO:0004528) |
| 0.2 | 0.5 | GO:0004357 | glutamate-cysteine ligase activity(GO:0004357) |
| 0.2 | 0.9 | GO:0032810 | sterol response element binding(GO:0032810) |
| 0.2 | 0.6 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.1 | 0.4 | GO:0005171 | hepatocyte growth factor receptor binding(GO:0005171) |
| 0.1 | 3.1 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.1 | 0.6 | GO:1990715 | mRNA CDS binding(GO:1990715) |
| 0.1 | 1.0 | GO:0016428 | tRNA (cytosine-5-)-methyltransferase activity(GO:0016428) |
| 0.1 | 3.2 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.1 | 0.6 | GO:0004335 | galactokinase activity(GO:0004335) |
| 0.1 | 4.5 | GO:0000062 | fatty-acyl-CoA binding(GO:0000062) |
| 0.1 | 1.6 | GO:0001162 | RNA polymerase II intronic transcription regulatory region sequence-specific DNA binding(GO:0001162) |
| 0.1 | 2.7 | GO:0044548 | S100 protein binding(GO:0044548) |
| 0.1 | 1.9 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.1 | 0.4 | GO:0071936 | coreceptor activity involved in Wnt signaling pathway(GO:0071936) |
| 0.1 | 2.1 | GO:0008409 | 5'-3' exonuclease activity(GO:0008409) |
| 0.1 | 1.4 | GO:0001054 | RNA polymerase I activity(GO:0001054) |
| 0.1 | 0.7 | GO:0003987 | acetate-CoA ligase activity(GO:0003987) |
| 0.1 | 0.6 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.1 | 0.3 | GO:0010484 | H3 histone acetyltransferase activity(GO:0010484) |
| 0.1 | 2.6 | GO:0042166 | acetylcholine binding(GO:0042166) |
| 0.1 | 0.4 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 0.1 | 0.3 | GO:0001884 | pyrimidine nucleoside binding(GO:0001884) |
| 0.1 | 3.1 | GO:0005537 | mannose binding(GO:0005537) |
| 0.1 | 3.3 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.1 | 0.8 | GO:0098634 | protein binding involved in cell-matrix adhesion(GO:0098634) collagen binding involved in cell-matrix adhesion(GO:0098639) |
| 0.1 | 0.5 | GO:0004909 | interleukin-1, Type I, activating receptor activity(GO:0004909) |
| 0.1 | 0.8 | GO:1990050 | phosphatidic acid transporter activity(GO:1990050) |
| 0.1 | 1.3 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.1 | 1.7 | GO:0071253 | connexin binding(GO:0071253) |
| 0.1 | 23.8 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.1 | 1.4 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.1 | 0.5 | GO:0033592 | RNA strand annealing activity(GO:0033592) |
| 0.1 | 1.9 | GO:0004697 | protein kinase C activity(GO:0004697) |
| 0.1 | 0.1 | GO:0001727 | lipid kinase activity(GO:0001727) ceramide kinase activity(GO:0001729) |
| 0.1 | 1.9 | GO:0008143 | poly(A) binding(GO:0008143) |
| 0.1 | 1.3 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.1 | 1.6 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 0.1 | 0.5 | GO:0008332 | low voltage-gated calcium channel activity(GO:0008332) |
| 0.1 | 1.4 | GO:0016505 | peptidase activator activity involved in apoptotic process(GO:0016505) |
| 0.1 | 0.5 | GO:1990460 | leptin receptor binding(GO:1990460) |
| 0.1 | 4.1 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.1 | 42.5 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.1 | 0.2 | GO:0098518 | polynucleotide phosphatase activity(GO:0098518) |
| 0.1 | 3.9 | GO:0008432 | JUN kinase binding(GO:0008432) |
| 0.1 | 5.4 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.1 | 0.6 | GO:1990188 | euchromatin binding(GO:1990188) |
| 0.1 | 3.1 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.1 | 2.1 | GO:0008430 | selenium binding(GO:0008430) |
| 0.1 | 0.9 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.1 | 3.2 | GO:0031683 | G-protein beta/gamma-subunit complex binding(GO:0031683) |
| 0.1 | 1.9 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.1 | 1.1 | GO:0015165 | pyrimidine nucleotide-sugar transmembrane transporter activity(GO:0015165) |
| 0.1 | 0.7 | GO:0000832 | inositol hexakisphosphate kinase activity(GO:0000828) inositol hexakisphosphate 5-kinase activity(GO:0000832) inositol hexakisphosphate 1-kinase activity(GO:0052723) inositol hexakisphosphate 3-kinase activity(GO:0052724) |
| 0.1 | 0.4 | GO:0051916 | granulocyte colony-stimulating factor binding(GO:0051916) |
| 0.1 | 3.0 | GO:0001618 | virus receptor activity(GO:0001618) |
| 0.1 | 0.3 | GO:0061711 | N(6)-L-threonylcarbamoyladenine synthase(GO:0061711) |
| 0.1 | 0.9 | GO:0008428 | ribonuclease inhibitor activity(GO:0008428) |
| 0.1 | 3.6 | GO:0004532 | exoribonuclease activity(GO:0004532) |
| 0.1 | 2.9 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.1 | 0.3 | GO:0030519 | snoRNP binding(GO:0030519) |
| 0.1 | 0.4 | GO:0031208 | POZ domain binding(GO:0031208) |
| 0.1 | 2.7 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.1 | 0.7 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.1 | 0.4 | GO:0019828 | aspartic-type endopeptidase inhibitor activity(GO:0019828) |
| 0.1 | 1.4 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.1 | 0.2 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.1 | 3.5 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.1 | 1.0 | GO:0005025 | transforming growth factor beta receptor activity, type I(GO:0005025) |
| 0.1 | 1.7 | GO:0001104 | RNA polymerase II transcription cofactor activity(GO:0001104) |
| 0.1 | 6.1 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.1 | 2.1 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.1 | 2.5 | GO:0015026 | coreceptor activity(GO:0015026) |
| 0.1 | 11.3 | GO:0003705 | transcription factor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0003705) |
| 0.1 | 0.6 | GO:0005052 | peroxisome matrix targeting signal-1 binding(GO:0005052) |
| 0.1 | 0.8 | GO:0008410 | CoA-transferase activity(GO:0008410) |
| 0.1 | 1.6 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.1 | 0.2 | GO:0001034 | RNA polymerase III transcription factor activity, sequence-specific DNA binding(GO:0001034) |
| 0.1 | 0.9 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.1 | 0.3 | GO:0004370 | glycerol kinase activity(GO:0004370) |
| 0.1 | 0.5 | GO:0034513 | box H/ACA snoRNA binding(GO:0034513) |
| 0.1 | 0.3 | GO:0004774 | succinate-CoA ligase activity(GO:0004774) |
| 0.1 | 22.7 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.1 | 0.5 | GO:0030621 | U4 snRNA binding(GO:0030621) |
| 0.1 | 4.6 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.1 | 0.2 | GO:0047134 | protein-disulfide reductase activity(GO:0047134) |
| 0.1 | 2.4 | GO:0042171 | lysophosphatidic acid acyltransferase activity(GO:0042171) |
| 0.1 | 3.1 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.1 | 5.6 | GO:0070063 | RNA polymerase binding(GO:0070063) |
| 0.1 | 5.6 | GO:0016278 | lysine N-methyltransferase activity(GO:0016278) protein-lysine N-methyltransferase activity(GO:0016279) |
| 0.1 | 1.5 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.1 | 0.9 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.1 | 3.2 | GO:0008376 | acetylgalactosaminyltransferase activity(GO:0008376) |
| 0.1 | 0.3 | GO:0005136 | interleukin-4 receptor binding(GO:0005136) |
| 0.1 | 2.5 | GO:0005112 | Notch binding(GO:0005112) |
| 0.1 | 0.3 | GO:0045127 | N-acetylglucosamine kinase activity(GO:0045127) |
| 0.1 | 1.6 | GO:0005041 | low-density lipoprotein receptor activity(GO:0005041) |
| 0.1 | 1.3 | GO:0017176 | phosphatidylinositol N-acetylglucosaminyltransferase activity(GO:0017176) |
| 0.1 | 16.9 | GO:0001047 | core promoter binding(GO:0001047) |
| 0.1 | 0.3 | GO:0008124 | 4-alpha-hydroxytetrahydrobiopterin dehydratase activity(GO:0008124) |
| 0.1 | 2.6 | GO:0003951 | NAD+ kinase activity(GO:0003951) |
| 0.1 | 0.4 | GO:0016742 | hydroxymethyl-, formyl- and related transferase activity(GO:0016742) |
| 0.1 | 1.6 | GO:0005487 | nucleocytoplasmic transporter activity(GO:0005487) |
| 0.1 | 0.5 | GO:0034713 | type I transforming growth factor beta receptor binding(GO:0034713) |
| 0.1 | 2.7 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.1 | 0.2 | GO:0016034 | maleylacetoacetate isomerase activity(GO:0016034) |
| 0.1 | 1.1 | GO:0070411 | I-SMAD binding(GO:0070411) |
| 0.1 | 0.2 | GO:0004921 | interleukin-11 receptor activity(GO:0004921) interleukin-11 binding(GO:0019970) |
| 0.1 | 1.0 | GO:0005161 | platelet-derived growth factor receptor binding(GO:0005161) |
| 0.1 | 0.8 | GO:0004579 | oligosaccharyl transferase activity(GO:0004576) dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.1 | 0.7 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 0.1 | 0.6 | GO:0070034 | telomerase RNA binding(GO:0070034) |
| 0.1 | 0.7 | GO:0090079 | translation regulator activity, nucleic acid binding(GO:0090079) |
| 0.1 | 0.2 | GO:0004067 | asparaginase activity(GO:0004067) |
| 0.1 | 1.3 | GO:0031490 | chromatin DNA binding(GO:0031490) |
| 0.1 | 1.7 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.1 | 4.5 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.1 | 1.0 | GO:0008417 | fucosyltransferase activity(GO:0008417) |
| 0.1 | 2.6 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.1 | 0.7 | GO:0004568 | chitinase activity(GO:0004568) |
| 0.1 | 2.9 | GO:0043130 | ubiquitin binding(GO:0043130) |
| 0.1 | 0.4 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) |
| 0.1 | 0.4 | GO:0003989 | acetyl-CoA carboxylase activity(GO:0003989) |
| 0.1 | 1.7 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.1 | 0.4 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 0.1 | 2.5 | GO:0030742 | GTP-dependent protein binding(GO:0030742) |
| 0.1 | 0.9 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.1 | 2.2 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.1 | 0.6 | GO:0005166 | neurotrophin p75 receptor binding(GO:0005166) |
| 0.1 | 0.3 | GO:0034437 | glycoprotein transporter activity(GO:0034437) |
| 0.1 | 1.1 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.1 | 1.7 | GO:0032451 | demethylase activity(GO:0032451) |
| 0.1 | 0.8 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 0.1 | 1.4 | GO:0031418 | L-ascorbic acid binding(GO:0031418) |
| 0.1 | 1.5 | GO:0051861 | glycolipid binding(GO:0051861) |
| 0.1 | 0.2 | GO:0005292 | high-affinity basic amino acid transmembrane transporter activity(GO:0005287) high-affinity arginine transmembrane transporter activity(GO:0005289) high-affinity lysine transmembrane transporter activity(GO:0005292) |
| 0.1 | 1.0 | GO:0008649 | rRNA methyltransferase activity(GO:0008649) |
| 0.1 | 3.6 | GO:0000987 | core promoter proximal region sequence-specific DNA binding(GO:0000987) core promoter proximal region DNA binding(GO:0001159) |
| 0.1 | 0.3 | GO:0003986 | acetyl-CoA hydrolase activity(GO:0003986) |
| 0.1 | 0.5 | GO:0008865 | fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.1 | 0.7 | GO:0004565 | beta-galactosidase activity(GO:0004565) |
| 0.1 | 0.5 | GO:0019770 | IgG receptor activity(GO:0019770) |
| 0.1 | 0.8 | GO:0055106 | ubiquitin-protein transferase regulator activity(GO:0055106) |
| 0.1 | 1.6 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.1 | 1.4 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.1 | 1.1 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.1 | 1.2 | GO:0005351 | sugar:proton symporter activity(GO:0005351) |
| 0.1 | 0.4 | GO:0003988 | acetyl-CoA C-acyltransferase activity(GO:0003988) |
| 0.1 | 1.6 | GO:0070273 | phosphatidylinositol-4-phosphate binding(GO:0070273) |
| 0.1 | 0.2 | GO:0004813 | alanine-tRNA ligase activity(GO:0004813) |
| 0.1 | 0.7 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 0.1 | 2.8 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.1 | 0.2 | GO:0008177 | succinate dehydrogenase (ubiquinone) activity(GO:0008177) |
| 0.1 | 3.3 | GO:0031593 | polyubiquitin binding(GO:0031593) |
| 0.1 | 9.2 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 1.8 | GO:0005031 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.1 | 2.2 | GO:0004623 | phospholipase A2 activity(GO:0004623) |
| 0.1 | 5.8 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.1 | 0.3 | GO:0047804 | cysteine-S-conjugate beta-lyase activity(GO:0047804) |
| 0.1 | 0.5 | GO:0008479 | queuine tRNA-ribosyltransferase activity(GO:0008479) |
| 0.1 | 0.6 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.1 | 0.4 | GO:0016409 | palmitoyltransferase activity(GO:0016409) |
| 0.1 | 2.0 | GO:0042393 | histone binding(GO:0042393) |
| 0.1 | 0.6 | GO:0042500 | aspartic endopeptidase activity, intramembrane cleaving(GO:0042500) |
| 0.1 | 0.5 | GO:0004028 | 3-chloroallyl aldehyde dehydrogenase activity(GO:0004028) |
| 0.1 | 26.3 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.1 | 0.6 | GO:0015245 | fatty acid transporter activity(GO:0015245) |
| 0.1 | 0.7 | GO:0042974 | retinoic acid receptor binding(GO:0042974) |
| 0.1 | 0.6 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.1 | 0.7 | GO:0031386 | protein tag(GO:0031386) |
| 0.1 | 0.4 | GO:0047276 | N-acetyllactosaminide 3-alpha-galactosyltransferase activity(GO:0047276) |
| 0.1 | 0.4 | GO:0015252 | hydrogen ion channel activity(GO:0015252) |
| 0.1 | 3.4 | GO:0008186 | RNA-dependent ATPase activity(GO:0008186) |
| 0.1 | 1.2 | GO:0008198 | ferrous iron binding(GO:0008198) |
| 0.1 | 0.9 | GO:0003796 | lysozyme activity(GO:0003796) |
| 0.1 | 0.3 | GO:0004366 | glycerol-3-phosphate O-acyltransferase activity(GO:0004366) |
| 0.1 | 1.0 | GO:0030414 | peptidase inhibitor activity(GO:0030414) |
| 0.1 | 3.7 | GO:0004843 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) |
| 0.1 | 0.3 | GO:0042030 | ATPase inhibitor activity(GO:0042030) |
| 0.1 | 6.2 | GO:0000981 | RNA polymerase II transcription factor activity, sequence-specific DNA binding(GO:0000981) |
| 0.1 | 1.6 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) |
| 0.1 | 9.8 | GO:0003713 | transcription coactivator activity(GO:0003713) |
| 0.1 | 2.7 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.0 | 0.6 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.0 | 0.7 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.0 | 0.1 | GO:0009918 | sterol delta7 reductase activity(GO:0009918) 7-dehydrocholesterol reductase activity(GO:0047598) |
| 0.0 | 21.7 | GO:0003700 | nucleic acid binding transcription factor activity(GO:0001071) transcription factor activity, sequence-specific DNA binding(GO:0003700) |
| 0.0 | 0.3 | GO:0019966 | interleukin-1 binding(GO:0019966) |
| 0.0 | 0.8 | GO:0004029 | aldehyde dehydrogenase (NAD) activity(GO:0004029) |
| 0.0 | 2.3 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 0.1 | GO:0044715 | 8-oxo-dGDP phosphatase activity(GO:0044715) |
| 0.0 | 0.6 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.0 | 2.2 | GO:0004386 | helicase activity(GO:0004386) |
| 0.0 | 1.1 | GO:0043236 | laminin binding(GO:0043236) |
| 0.0 | 0.5 | GO:0001076 | transcription factor activity, RNA polymerase II transcription factor binding(GO:0001076) |
| 0.0 | 0.1 | GO:0043566 | structure-specific DNA binding(GO:0043566) |
| 0.0 | 34.2 | GO:0003677 | DNA binding(GO:0003677) |
| 0.0 | 1.6 | GO:0046332 | SMAD binding(GO:0046332) |
| 0.0 | 0.4 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.0 | 1.3 | GO:0016881 | acid-amino acid ligase activity(GO:0016881) |
| 0.0 | 0.4 | GO:0015035 | protein disulfide oxidoreductase activity(GO:0015035) |
| 0.0 | 0.1 | GO:0042015 | interleukin-20 binding(GO:0042015) |
| 0.0 | 0.4 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.0 | 0.2 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.0 | 1.9 | GO:0061650 | ubiquitin-like protein conjugating enzyme activity(GO:0061650) |
| 0.0 | 0.2 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.0 | 0.6 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 0.0 | 0.3 | GO:0048156 | tau protein binding(GO:0048156) |
| 0.0 | 0.4 | GO:0015288 | porin activity(GO:0015288) |
| 0.0 | 1.0 | GO:0008536 | Ran GTPase binding(GO:0008536) |
| 0.0 | 0.3 | GO:0015093 | ferrous iron transmembrane transporter activity(GO:0015093) |
| 0.0 | 0.2 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.0 | 1.6 | GO:0030971 | receptor tyrosine kinase binding(GO:0030971) |
| 0.0 | 0.3 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.0 | 0.9 | GO:0016675 | oxidoreductase activity, acting on a heme group of donors(GO:0016675) |
| 0.0 | 1.3 | GO:0000049 | tRNA binding(GO:0000049) |
| 0.0 | 0.2 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.0 | 0.2 | GO:0008408 | 3'-5' exonuclease activity(GO:0008408) |
| 0.0 | 0.2 | GO:0005332 | gamma-aminobutyric acid:sodium symporter activity(GO:0005332) |
| 0.0 | 3.3 | GO:0004860 | protein kinase inhibitor activity(GO:0004860) |
| 0.0 | 0.1 | GO:0033192 | calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.0 | 0.2 | GO:0008469 | histone-arginine N-methyltransferase activity(GO:0008469) protein-arginine omega-N asymmetric methyltransferase activity(GO:0035242) |
| 0.0 | 0.2 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.0 | 0.3 | GO:0098599 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 0.0 | 0.7 | GO:0016780 | phosphotransferase activity, for other substituted phosphate groups(GO:0016780) |
| 0.0 | 0.8 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.0 | 0.1 | GO:0004741 | [pyruvate dehydrogenase (lipoamide)] phosphatase activity(GO:0004741) |
| 0.0 | 0.5 | GO:0016504 | peptidase activator activity(GO:0016504) |
| 0.0 | 0.1 | GO:0003747 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.0 | 0.6 | GO:0015238 | drug transmembrane transporter activity(GO:0015238) |
| 0.0 | 0.7 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.0 | 0.1 | GO:0003923 | GPI-anchor transamidase activity(GO:0003923) |
| 0.0 | 0.2 | GO:0031419 | cobalamin binding(GO:0031419) |
| 0.0 | 0.2 | GO:0001602 | pancreatic polypeptide receptor activity(GO:0001602) |
| 0.0 | 2.4 | GO:0003682 | chromatin binding(GO:0003682) |
| 0.0 | 0.1 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.0 | 0.9 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.0 | 0.7 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.0 | 0.1 | GO:0044390 | ubiquitin-like protein conjugating enzyme binding(GO:0044390) |
| 0.0 | 0.7 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.3 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.0 | 0.2 | GO:0102336 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.0 | 0.4 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.0 | 2.0 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 0.1 | GO:0051765 | inositol tetrakisphosphate kinase activity(GO:0051765) |
| 0.0 | 1.1 | GO:0019955 | cytokine binding(GO:0019955) |
| 0.0 | 0.1 | GO:0047291 | lactosylceramide alpha-2,3-sialyltransferase activity(GO:0047291) |
| 0.0 | 0.2 | GO:0005522 | profilin binding(GO:0005522) |
| 0.0 | 0.1 | GO:0016520 | growth hormone-releasing hormone receptor activity(GO:0016520) |
| 0.0 | 0.4 | GO:0008527 | taste receptor activity(GO:0008527) |
| 0.0 | 0.2 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) serine-type exopeptidase activity(GO:0070008) |
| 0.0 | 0.1 | GO:0097108 | hedgehog family protein binding(GO:0097108) |
| 0.0 | 0.1 | GO:0004736 | pyruvate carboxylase activity(GO:0004736) |
| 0.0 | 0.1 | GO:1990247 | N6-methyladenosine-containing RNA binding(GO:1990247) |
| 0.0 | 0.8 | GO:0042805 | actinin binding(GO:0042805) |
| 0.0 | 0.8 | GO:0008565 | protein transporter activity(GO:0008565) |
| 0.0 | 0.2 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 40.6 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.7 | 0.7 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.6 | 13.9 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.4 | 4.1 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.4 | 22.0 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.3 | 6.8 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.3 | 4.8 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.3 | 6.9 | PID AVB3 OPN PATHWAY | Osteopontin-mediated events |
| 0.3 | 0.3 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.3 | 11.8 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.3 | 26.1 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.3 | 19.3 | PID EPO PATHWAY | EPO signaling pathway |
| 0.3 | 4.7 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.3 | 5.2 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.3 | 11.9 | PID IFNG PATHWAY | IFN-gamma pathway |
| 0.2 | 7.2 | ST TUMOR NECROSIS FACTOR PATHWAY | Tumor Necrosis Factor Pathway. |
| 0.2 | 1.4 | PID LYMPH ANGIOGENESIS PATHWAY | VEGFR3 signaling in lymphatic endothelium |
| 0.2 | 11.3 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.2 | 18.8 | PID E2F PATHWAY | E2F transcription factor network |
| 0.2 | 15.7 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.2 | 5.2 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.2 | 12.9 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.2 | 5.5 | ST PHOSPHOINOSITIDE 3 KINASE PATHWAY | PI3K Pathway |
| 0.2 | 1.2 | ST TYPE I INTERFERON PATHWAY | Type I Interferon (alpha/beta IFN) Pathway |
| 0.2 | 1.8 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.2 | 5.1 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.2 | 0.6 | SIG CHEMOTAXIS | Genes related to chemotaxis |
| 0.2 | 10.4 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.2 | 8.6 | ST P38 MAPK PATHWAY | p38 MAPK Pathway |
| 0.2 | 4.0 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.2 | 9.8 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.2 | 2.9 | SIG CD40PATHWAYMAP | Genes related to CD40 signaling |
| 0.2 | 3.9 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.2 | 1.3 | ST JAK STAT PATHWAY | Jak-STAT Pathway |
| 0.2 | 0.5 | SA TRKA RECEPTOR | The TrkA receptor binds nerve growth factor to activate MAP kinase pathways and promote cell growth. |
| 0.2 | 2.7 | PID MYC PATHWAY | C-MYC pathway |
| 0.2 | 5.0 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.2 | 2.5 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.2 | 2.0 | PID S1P S1P2 PATHWAY | S1P2 pathway |
| 0.1 | 0.7 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.1 | 2.3 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.1 | 1.2 | PID IL2 1PATHWAY | IL2-mediated signaling events |
| 0.1 | 6.3 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.1 | 2.6 | PID TCPTP PATHWAY | Signaling events mediated by TCPTP |
| 0.1 | 7.7 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.1 | 4.6 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.1 | 5.3 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.1 | 1.2 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.1 | 2.3 | PID IL3 PATHWAY | IL3-mediated signaling events |
| 0.1 | 1.8 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
| 0.1 | 7.0 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 11.2 | PID PDGFRB PATHWAY | PDGFR-beta signaling pathway |
| 0.1 | 1.7 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.1 | 7.1 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.1 | 1.0 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.1 | 2.7 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.1 | 10.6 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.1 | 2.3 | PID ATR PATHWAY | ATR signaling pathway |
| 0.1 | 5.1 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.1 | 11.5 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.1 | 3.8 | PID VEGFR1 2 PATHWAY | Signaling events mediated by VEGFR1 and VEGFR2 |
| 0.1 | 4.7 | PID P53 REGULATION PATHWAY | p53 pathway |
| 0.1 | 5.0 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.1 | 2.2 | ST FAS SIGNALING PATHWAY | Fas Signaling Pathway |
| 0.1 | 0.9 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.1 | 1.2 | PID FOXO PATHWAY | FoxO family signaling |
| 0.1 | 0.4 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.1 | 1.4 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.1 | 3.1 | PID BMP PATHWAY | BMP receptor signaling |
| 0.1 | 1.7 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.1 | 1.4 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.1 | 0.4 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.0 | 0.9 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.0 | 1.3 | PID CD8 TCR PATHWAY | TCR signaling in naïve CD8+ T cells |
| 0.0 | 1.5 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.0 | 0.5 | PID THROMBIN PAR1 PATHWAY | PAR1-mediated thrombin signaling events |
| 0.0 | 2.4 | PID AVB3 INTEGRIN PATHWAY | Integrins in angiogenesis |
| 0.0 | 1.2 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 0.5 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.0 | 1.1 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 0.2 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
| 0.0 | 1.2 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 0.6 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 0.9 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
| 0.0 | 2.2 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
| 0.0 | 0.8 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.0 | 0.4 | PID ATM PATHWAY | ATM pathway |
| 0.0 | 0.4 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.0 | 7.8 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.0 | PID ERBB1 DOWNSTREAM PATHWAY | ErbB1 downstream signaling |
| 0.0 | 1.2 | PID FGF PATHWAY | FGF signaling pathway |
| 0.0 | 0.9 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.0 | 0.4 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.0 | 0.4 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.0 | 0.9 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 0.2 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.0 | 0.2 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.0 | 0.3 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.0 | 0.5 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.0 | 0.2 | PID TNF PATHWAY | TNF receptor signaling pathway |
| 0.0 | 0.3 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.0 | 0.4 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 0.0 | 0.1 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.0 | 0.2 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 0.9 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.9 | 11.6 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.6 | 16.2 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.5 | 7.0 | REACTOME E2F ENABLED INHIBITION OF PRE REPLICATION COMPLEX FORMATION | Genes involved in E2F-enabled inhibition of pre-replication complex formation |
| 0.5 | 11.5 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.5 | 12.1 | REACTOME REGULATION OF IFNG SIGNALING | Genes involved in Regulation of IFNG signaling |
| 0.5 | 2.0 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.5 | 11.3 | REACTOME BASE EXCISION REPAIR | Genes involved in Base Excision Repair |
| 0.4 | 6.6 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.4 | 6.0 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 0.4 | 4.2 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.4 | 7.1 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.4 | 14.1 | REACTOME SMAD2 SMAD3 SMAD4 HETEROTRIMER REGULATES TRANSCRIPTION | Genes involved in SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription |
| 0.4 | 12.7 | REACTOME ACTIVATION OF THE PRE REPLICATIVE COMPLEX | Genes involved in Activation of the pre-replicative complex |
| 0.4 | 12.7 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.4 | 0.7 | REACTOME DESTABILIZATION OF MRNA BY KSRP | Genes involved in Destabilization of mRNA by KSRP |
| 0.4 | 13.0 | REACTOME DOWNREGULATION OF TGF BETA RECEPTOR SIGNALING | Genes involved in Downregulation of TGF-beta receptor signaling |
| 0.4 | 0.7 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.4 | 1.1 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 0.4 | 10.2 | REACTOME ACTIVATED AMPK STIMULATES FATTY ACID OXIDATION IN MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.3 | 8.4 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.3 | 5.2 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 0.3 | 5.5 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.3 | 3.2 | REACTOME PROSTANOID LIGAND RECEPTORS | Genes involved in Prostanoid ligand receptors |
| 0.3 | 3.2 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.3 | 20.8 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.3 | 1.9 | REACTOME NFKB ACTIVATION THROUGH FADD RIP1 PATHWAY MEDIATED BY CASPASE 8 AND10 | Genes involved in NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 |
| 0.3 | 1.6 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.3 | 5.0 | REACTOME GRB2 SOS PROVIDES LINKAGE TO MAPK SIGNALING FOR INTERGRINS | Genes involved in GRB2:SOS provides linkage to MAPK signaling for Intergrins |
| 0.3 | 2.7 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.3 | 3.0 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.3 | 1.5 | REACTOME INFLUENZA VIRAL RNA TRANSCRIPTION AND REPLICATION | Genes involved in Influenza Viral RNA Transcription and Replication |
| 0.3 | 2.9 | REACTOME REGULATION OF IFNA SIGNALING | Genes involved in Regulation of IFNA signaling |
| 0.3 | 7.8 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.3 | 4.8 | REACTOME IL 3 5 AND GM CSF SIGNALING | Genes involved in Interleukin-3, 5 and GM-CSF signaling |
| 0.3 | 2.8 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.2 | 3.0 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.2 | 37.3 | REACTOME NONSENSE MEDIATED DECAY ENHANCED BY THE EXON JUNCTION COMPLEX | Genes involved in Nonsense Mediated Decay Enhanced by the Exon Junction Complex |
| 0.2 | 0.2 | REACTOME E2F MEDIATED REGULATION OF DNA REPLICATION | Genes involved in E2F mediated regulation of DNA replication |
| 0.2 | 7.8 | REACTOME N GLYCAN ANTENNAE ELONGATION IN THE MEDIAL TRANS GOLGI | Genes involved in N-glycan antennae elongation in the medial/trans-Golgi |
| 0.2 | 7.2 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.2 | 3.7 | REACTOME INTEGRATION OF PROVIRUS | Genes involved in Integration of provirus |
| 0.2 | 4.6 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.2 | 4.3 | REACTOME ACTIVATED TAK1 MEDIATES P38 MAPK ACTIVATION | Genes involved in activated TAK1 mediates p38 MAPK activation |
| 0.2 | 2.9 | REACTOME CTLA4 INHIBITORY SIGNALING | Genes involved in CTLA4 inhibitory signaling |
| 0.2 | 0.4 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 0.2 | 9.7 | REACTOME IL RECEPTOR SHC SIGNALING | Genes involved in Interleukin receptor SHC signaling |
| 0.2 | 10.1 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.2 | 5.4 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.2 | 2.6 | REACTOME RECEPTOR LIGAND BINDING INITIATES THE SECOND PROTEOLYTIC CLEAVAGE OF NOTCH RECEPTOR | Genes involved in Receptor-ligand binding initiates the second proteolytic cleavage of Notch receptor |
| 0.2 | 5.1 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.2 | 1.5 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.2 | 7.0 | REACTOME TRANSPORT OF MATURE MRNA DERIVED FROM AN INTRONLESS TRANSCRIPT | Genes involved in Transport of Mature mRNA Derived from an Intronless Transcript |
| 0.2 | 6.9 | REACTOME MEIOSIS | Genes involved in Meiosis |
| 0.2 | 4.4 | REACTOME TRAF3 DEPENDENT IRF ACTIVATION PATHWAY | Genes involved in TRAF3-dependent IRF activation pathway |
| 0.2 | 5.1 | REACTOME CD28 DEPENDENT PI3K AKT SIGNALING | Genes involved in CD28 dependent PI3K/Akt signaling |
| 0.2 | 5.1 | REACTOME TRANSPORT OF MATURE TRANSCRIPT TO CYTOPLASM | Genes involved in Transport of Mature Transcript to Cytoplasm |
| 0.2 | 4.3 | REACTOME RNA POL II PRE TRANSCRIPTION EVENTS | Genes involved in RNA Polymerase II Pre-transcription Events |
| 0.2 | 4.3 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.2 | 3.4 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.2 | 5.8 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.2 | 2.4 | REACTOME LIPOPROTEIN METABOLISM | Genes involved in Lipoprotein metabolism |
| 0.2 | 16.4 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.2 | 4.1 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.2 | 4.8 | REACTOME DOWNSTREAM TCR SIGNALING | Genes involved in Downstream TCR signaling |
| 0.2 | 3.1 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.2 | 7.8 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.2 | 9.5 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.2 | 18.3 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.2 | 3.2 | REACTOME ACYL CHAIN REMODELLING OF PS | Genes involved in Acyl chain remodelling of PS |
| 0.2 | 3.8 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.2 | 1.7 | REACTOME TAK1 ACTIVATES NFKB BY PHOSPHORYLATION AND ACTIVATION OF IKKS COMPLEX | Genes involved in TAK1 activates NFkB by phosphorylation and activation of IKKs complex |
| 0.2 | 1.7 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.2 | 2.3 | REACTOME NEF MEDIATED DOWNREGULATION OF MHC CLASS I COMPLEX CELL SURFACE EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.2 | 0.5 | REACTOME P53 DEPENDENT G1 DNA DAMAGE RESPONSE | Genes involved in p53-Dependent G1 DNA Damage Response |
| 0.2 | 4.6 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.2 | 3.0 | REACTOME RNA POL I TRANSCRIPTION INITIATION | Genes involved in RNA Polymerase I Transcription Initiation |
| 0.2 | 2.3 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.1 | 2.5 | REACTOME DESTABILIZATION OF MRNA BY TRISTETRAPROLIN TTP | Genes involved in Destabilization of mRNA by Tristetraprolin (TTP) |
| 0.1 | 2.8 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.1 | 1.2 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION IN TLR7 8 OR 9 SIGNALING | Genes involved in TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling |
| 0.1 | 1.4 | REACTOME ACTIVATION OF NF KAPPAB IN B CELLS | Genes involved in Activation of NF-kappaB in B Cells |
| 0.1 | 2.0 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.1 | 1.3 | REACTOME MICRORNA MIRNA BIOGENESIS | Genes involved in MicroRNA (miRNA) Biogenesis |
| 0.1 | 3.7 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.1 | 3.8 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.1 | 1.2 | REACTOME GROWTH HORMONE RECEPTOR SIGNALING | Genes involved in Growth hormone receptor signaling |
| 0.1 | 3.7 | REACTOME ELONGATION ARREST AND RECOVERY | Genes involved in Elongation arrest and recovery |
| 0.1 | 8.0 | REACTOME DESTABILIZATION OF MRNA BY AUF1 HNRNP D0 | Genes involved in Destabilization of mRNA by AUF1 (hnRNP D0) |
| 0.1 | 0.7 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION | Genes involved in TRAF6 mediated IRF7 activation |
| 0.1 | 0.5 | REACTOME NFKB IS ACTIVATED AND SIGNALS SURVIVAL | Genes involved in NF-kB is activated and signals survival |
| 0.1 | 3.8 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.1 | 5.2 | REACTOME CREB PHOSPHORYLATION THROUGH THE ACTIVATION OF RAS | Genes involved in CREB phosphorylation through the activation of Ras |
| 0.1 | 0.7 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.1 | 7.9 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.1 | 10.7 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.1 | 0.5 | REACTOME SIGNALLING TO P38 VIA RIT AND RIN | Genes involved in Signalling to p38 via RIT and RIN |
| 0.1 | 0.5 | REACTOME G2 M CHECKPOINTS | Genes involved in G2/M Checkpoints |
| 0.1 | 0.9 | REACTOME TRAF6 MEDIATED INDUCTION OF TAK1 COMPLEX | Genes involved in TRAF6 mediated induction of TAK1 complex |
| 0.1 | 11.9 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.1 | 4.9 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.1 | 1.7 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 0.1 | 3.6 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.1 | 2.1 | REACTOME CELL SURFACE INTERACTIONS AT THE VASCULAR WALL | Genes involved in Cell surface interactions at the vascular wall |
| 0.1 | 0.9 | REACTOME REGULATION OF ORNITHINE DECARBOXYLASE ODC | Genes involved in Regulation of ornithine decarboxylase (ODC) |
| 0.1 | 1.4 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.1 | 2.4 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.1 | 3.8 | REACTOME DNA REPAIR | Genes involved in DNA Repair |
| 0.1 | 0.9 | REACTOME UNFOLDED PROTEIN RESPONSE | Genes involved in Unfolded Protein Response |
| 0.1 | 1.0 | REACTOME GLYCOPROTEIN HORMONES | Genes involved in Glycoprotein hormones |
| 0.1 | 0.9 | REACTOME CHROMOSOME MAINTENANCE | Genes involved in Chromosome Maintenance |
| 0.1 | 0.8 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.1 | 2.2 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.1 | 6.8 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.1 | 0.9 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.1 | 2.9 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.1 | 2.7 | REACTOME INTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Intrinsic Pathway for Apoptosis |
| 0.1 | 2.1 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.1 | 1.5 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.1 | 0.7 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.1 | 0.3 | REACTOME PD1 SIGNALING | Genes involved in PD-1 signaling |
| 0.1 | 1.1 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.1 | 0.6 | REACTOME ER PHAGOSOME PATHWAY | Genes involved in ER-Phagosome pathway |
| 0.1 | 1.1 | REACTOME FGFR4 LIGAND BINDING AND ACTIVATION | Genes involved in FGFR4 ligand binding and activation |
| 0.1 | 3.2 | REACTOME APOPTOTIC CLEAVAGE OF CELLULAR PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |
| 0.1 | 3.3 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 1.2 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.0 | 0.7 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.0 | 0.7 | REACTOME N GLYCAN TRIMMING IN THE ER AND CALNEXIN CALRETICULIN CYCLE | Genes involved in N-glycan trimming in the ER and Calnexin/Calreticulin cycle |
| 0.0 | 0.4 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.0 | 0.9 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |
| 0.0 | 0.3 | REACTOME TRANSCRIPTIONAL ACTIVITY OF SMAD2 SMAD3 SMAD4 HETEROTRIMER | Genes involved in Transcriptional activity of SMAD2/SMAD3:SMAD4 heterotrimer |
| 0.0 | 0.7 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.0 | 0.3 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.0 | 0.1 | REACTOME REGULATORY RNA PATHWAYS | Genes involved in Regulatory RNA pathways |
| 0.0 | 0.4 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.0 | 1.5 | REACTOME RNA POL I RNA POL III AND MITOCHONDRIAL TRANSCRIPTION | Genes involved in RNA Polymerase I, RNA Polymerase III, and Mitochondrial Transcription |
| 0.0 | 0.4 | REACTOME SIGNALING BY NOTCH4 | Genes involved in Signaling by NOTCH4 |
| 0.0 | 0.2 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.0 | 0.4 | REACTOME PRE NOTCH TRANSCRIPTION AND TRANSLATION | Genes involved in Pre-NOTCH Transcription and Translation |
| 0.0 | 0.4 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.0 | 0.7 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.0 | 0.5 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.0 | 0.1 | REACTOME DNA STRAND ELONGATION | Genes involved in DNA strand elongation |
| 0.0 | 1.1 | REACTOME REGULATION OF BETA CELL DEVELOPMENT | Genes involved in Regulation of beta-cell development |
| 0.0 | 0.8 | REACTOME NUCLEAR EVENTS KINASE AND TRANSCRIPTION FACTOR ACTIVATION | Genes involved in Nuclear Events (kinase and transcription factor activation) |
| 0.0 | 1.6 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.3 | REACTOME OPSINS | Genes involved in Opsins |
| 0.0 | 1.1 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 1.2 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.0 | 0.4 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.0 | 0.6 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.0 | 0.5 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.0 | 0.4 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |
| 0.0 | 0.3 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.0 | 0.6 | REACTOME FATTY ACYL COA BIOSYNTHESIS | Genes involved in Fatty Acyl-CoA Biosynthesis |
| 0.0 | 0.1 | REACTOME LATE PHASE OF HIV LIFE CYCLE | Genes involved in Late Phase of HIV Life Cycle |
| 0.0 | 0.5 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 1.2 | REACTOME GENERIC TRANSCRIPTION PATHWAY | Genes involved in Generic Transcription Pathway |
| 0.0 | 0.8 | REACTOME TRANSLATION | Genes involved in Translation |