Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
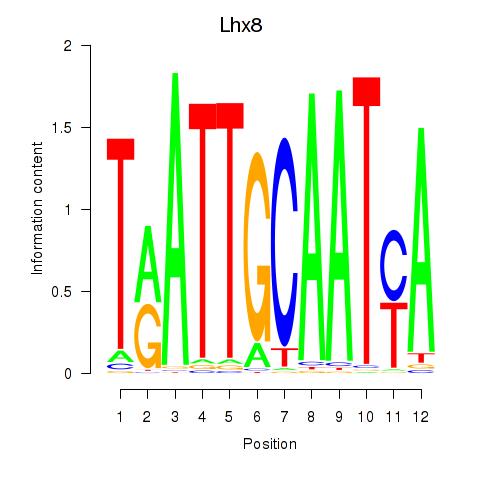
Results for Lhx8
Z-value: 1.12
Transcription factors associated with Lhx8
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Lhx8
|
ENSMUSG00000096225.8 | Lhx8 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Lhx8 | mm39_v1_chr3_-_154036180_154036296 | 0.04 | 7.4e-01 | Click! |
Activity profile of Lhx8 motif
Sorted Z-values of Lhx8 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Lhx8
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_+_137551774 | 9.38 |
ENSMUST00000136088.8
ENSMUST00000139395.8 |
Actl6b
|
actin-like 6B |
| chr11_+_110858842 | 9.05 |
ENSMUST00000180023.8
ENSMUST00000106636.8 |
Kcnj16
|
potassium inwardly-rectifying channel, subfamily J, member 16 |
| chr2_-_52448552 | 8.62 |
ENSMUST00000102760.10
ENSMUST00000102761.9 |
Cacnb4
|
calcium channel, voltage-dependent, beta 4 subunit |
| chr15_+_54609011 | 8.61 |
ENSMUST00000050027.9
|
Ccn3
|
cellular communication network factor 3 |
| chr7_-_59654797 | 8.15 |
ENSMUST00000194059.2
|
Snurf
|
SNRPN upstream reading frame |
| chr7_-_59654849 | 8.01 |
ENSMUST00000059305.17
|
Snrpn
|
small nuclear ribonucleoprotein N |
| chr4_-_96479793 | 7.61 |
ENSMUST00000055693.9
|
Cyp2j9
|
cytochrome P450, family 2, subfamily j, polypeptide 9 |
| chr3_+_130411097 | 7.53 |
ENSMUST00000166187.8
ENSMUST00000072271.13 |
Etnppl
|
ethanolamine phosphate phospholyase |
| chr4_-_129015682 | 7.52 |
ENSMUST00000139450.8
ENSMUST00000125931.9 ENSMUST00000116444.10 |
Hpca
|
hippocalcin |
| chr5_+_137551790 | 7.45 |
ENSMUST00000136565.8
ENSMUST00000149292.8 ENSMUST00000125489.2 |
Actl6b
|
actin-like 6B |
| chr8_-_65146079 | 6.52 |
ENSMUST00000048967.9
|
Cpe
|
carboxypeptidase E |
| chr5_-_87572060 | 6.31 |
ENSMUST00000072818.6
|
Ugt2b38
|
UDP glucuronosyltransferase 2 family, polypeptide B38 |
| chr3_+_130411294 | 5.70 |
ENSMUST00000163620.8
|
Etnppl
|
ethanolamine phosphate phospholyase |
| chr3_-_72964276 | 5.58 |
ENSMUST00000192477.2
|
Slitrk3
|
SLIT and NTRK-like family, member 3 |
| chr11_+_110858882 | 5.57 |
ENSMUST00000125692.2
|
Kcnj16
|
potassium inwardly-rectifying channel, subfamily J, member 16 |
| chr5_-_139115417 | 5.50 |
ENSMUST00000026973.14
|
Prkar1b
|
protein kinase, cAMP dependent regulatory, type I beta |
| chr17_-_90395771 | 5.47 |
ENSMUST00000197268.5
ENSMUST00000173917.8 |
Nrxn1
|
neurexin I |
| chr3_-_113368407 | 5.19 |
ENSMUST00000106540.8
|
Amy1
|
amylase 1, salivary |
| chr2_+_65451100 | 5.06 |
ENSMUST00000144254.6
ENSMUST00000028377.14 |
Scn2a
|
sodium channel, voltage-gated, type II, alpha |
| chr10_+_5589210 | 4.85 |
ENSMUST00000019906.6
|
Vip
|
vasoactive intestinal polypeptide |
| chr7_+_90739904 | 4.57 |
ENSMUST00000107196.10
ENSMUST00000074273.10 |
Dlg2
|
discs large MAGUK scaffold protein 2 |
| chr1_+_180878797 | 4.47 |
ENSMUST00000036819.7
|
9130409I23Rik
|
RIKEN cDNA 9130409I23 gene |
| chr1_+_17215581 | 4.36 |
ENSMUST00000026879.8
|
Gdap1
|
ganglioside-induced differentiation-associated-protein 1 |
| chr3_+_106020545 | 4.27 |
ENSMUST00000079132.12
ENSMUST00000139086.2 |
Chia1
|
chitinase, acidic 1 |
| chr18_+_23548534 | 4.21 |
ENSMUST00000221880.2
ENSMUST00000220904.2 ENSMUST00000047954.15 |
Dtna
|
dystrobrevin alpha |
| chr8_+_46080746 | 4.18 |
ENSMUST00000145458.9
ENSMUST00000134321.8 |
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr19_+_39980868 | 4.11 |
ENSMUST00000049178.3
|
Cyp2c37
|
cytochrome P450, family 2. subfamily c, polypeptide 37 |
| chr8_+_124169707 | 4.07 |
ENSMUST00000093049.10
ENSMUST00000065534.10 ENSMUST00000001522.10 ENSMUST00000124741.8 ENSMUST00000108832.8 ENSMUST00000132063.8 ENSMUST00000128424.8 |
Def8
|
differentially expressed in FDCP 8 |
| chr11_+_16207705 | 3.98 |
ENSMUST00000109645.9
ENSMUST00000109647.3 |
Vstm2a
|
V-set and transmembrane domain containing 2A |
| chr2_+_177200363 | 3.94 |
ENSMUST00000108940.3
|
Gm14403
|
predicted gene 14403 |
| chr1_+_72323526 | 3.93 |
ENSMUST00000027380.12
ENSMUST00000141783.2 |
Tmem169
|
transmembrane protein 169 |
| chr11_-_109886601 | 3.91 |
ENSMUST00000020948.15
|
Abca8b
|
ATP-binding cassette, sub-family A (ABC1), member 8b |
| chr1_+_21310803 | 3.75 |
ENSMUST00000027067.15
|
Gsta3
|
glutathione S-transferase, alpha 3 |
| chr3_-_82957104 | 3.59 |
ENSMUST00000048246.5
|
Fgb
|
fibrinogen beta chain |
| chr8_+_124170027 | 3.38 |
ENSMUST00000108830.2
|
Def8
|
differentially expressed in FDCP 8 |
| chr2_-_174973657 | 3.36 |
ENSMUST00000108929.3
|
Gm14399
|
predicted gene 14399 |
| chr1_+_107289659 | 3.33 |
ENSMUST00000027566.3
|
Serpinb11
|
serine (or cysteine) peptidase inhibitor, clade B (ovalbumin), member 11 |
| chr1_-_72323407 | 3.30 |
ENSMUST00000097698.5
|
Pecr
|
peroxisomal trans-2-enoyl-CoA reductase |
| chr19_-_4092218 | 3.27 |
ENSMUST00000237999.2
ENSMUST00000042700.12 |
Gstp2
|
glutathione S-transferase, pi 2 |
| chr5_-_87402659 | 3.25 |
ENSMUST00000075858.4
|
Ugt2b37
|
UDP glucuronosyltransferase 2 family, polypeptide B37 |
| chr8_+_110806390 | 3.20 |
ENSMUST00000212754.2
ENSMUST00000058804.10 |
Zfp612
|
zinc finger protein 612 |
| chr1_+_21310821 | 3.19 |
ENSMUST00000121676.8
ENSMUST00000124990.3 |
Gsta3
|
glutathione S-transferase, alpha 3 |
| chr1_+_6557455 | 3.08 |
ENSMUST00000140079.8
ENSMUST00000131494.8 |
St18
|
suppression of tumorigenicity 18 |
| chr12_+_84034628 | 3.05 |
ENSMUST00000021649.8
|
Acot2
|
acyl-CoA thioesterase 2 |
| chr18_+_23548192 | 3.03 |
ENSMUST00000222515.2
|
Dtna
|
dystrobrevin alpha |
| chr4_-_95965767 | 2.94 |
ENSMUST00000030305.13
ENSMUST00000107078.9 |
Cyp2j13
|
cytochrome P450, family 2, subfamily j, polypeptide 13 |
| chr12_+_38833501 | 2.93 |
ENSMUST00000159334.8
|
Etv1
|
ets variant 1 |
| chr17_-_47143214 | 2.88 |
ENSMUST00000233537.2
|
Bicral
|
BRD4 interacting chromatin remodeling complex associated protein like |
| chr1_-_72323464 | 2.77 |
ENSMUST00000027381.13
|
Pecr
|
peroxisomal trans-2-enoyl-CoA reductase |
| chr4_-_95965747 | 2.77 |
ENSMUST00000097973.3
|
Cyp2j13
|
cytochrome P450, family 2, subfamily j, polypeptide 13 |
| chr5_+_20433169 | 2.70 |
ENSMUST00000197553.5
ENSMUST00000208219.2 |
Magi2
|
membrane associated guanylate kinase, WW and PDZ domain containing 2 |
| chr12_+_38833454 | 2.60 |
ENSMUST00000161980.8
ENSMUST00000160701.8 |
Etv1
|
ets variant 1 |
| chr3_+_75982890 | 2.60 |
ENSMUST00000160261.8
|
Fstl5
|
follistatin-like 5 |
| chr7_+_89951444 | 2.58 |
ENSMUST00000207578.2
|
Sytl2
|
synaptotagmin-like 2 |
| chr18_+_23548455 | 2.58 |
ENSMUST00000115832.4
|
Dtna
|
dystrobrevin alpha |
| chr11_+_104122341 | 2.58 |
ENSMUST00000106993.10
|
Mapt
|
microtubule-associated protein tau |
| chrM_+_7758 | 2.51 |
ENSMUST00000082407.1
|
mt-Atp8
|
mitochondrially encoded ATP synthase 8 |
| chr8_-_95929420 | 2.51 |
ENSMUST00000212842.2
|
Kifc3
|
kinesin family member C3 |
| chr18_-_43820759 | 2.50 |
ENSMUST00000082254.8
|
Jakmip2
|
janus kinase and microtubule interacting protein 2 |
| chr1_-_130589349 | 2.46 |
ENSMUST00000027657.14
|
C4bp
|
complement component 4 binding protein |
| chr1_+_107288928 | 2.45 |
ENSMUST00000191425.7
|
Serpinb11
|
serine (or cysteine) peptidase inhibitor, clade B (ovalbumin), member 11 |
| chr11_-_12414850 | 2.43 |
ENSMUST00000109650.8
|
Cobl
|
cordon-bleu WH2 repeat |
| chr12_+_112940086 | 2.40 |
ENSMUST00000165079.8
ENSMUST00000221500.2 ENSMUST00000221104.2 ENSMUST00000002880.7 ENSMUST00000222209.2 |
Btbd6
|
BTB (POZ) domain containing 6 |
| chr5_+_25964985 | 2.36 |
ENSMUST00000128727.8
ENSMUST00000088244.6 |
Actr3b
|
ARP3 actin-related protein 3B |
| chr6_-_136150491 | 2.29 |
ENSMUST00000111905.8
ENSMUST00000152012.8 ENSMUST00000143943.8 ENSMUST00000125905.2 |
Grin2b
|
glutamate receptor, ionotropic, NMDA2B (epsilon 2) |
| chr8_+_72973560 | 2.17 |
ENSMUST00000003123.10
|
Fam32a
|
family with sequence similarity 32, member A |
| chr15_-_98118858 | 2.14 |
ENSMUST00000142443.8
ENSMUST00000170618.8 |
Gm44579
Olfr287
|
predicted gene 44579 olfactory receptor 287 |
| chr17_+_35150229 | 2.10 |
ENSMUST00000007253.6
|
Neu1
|
neuraminidase 1 |
| chr5_-_145128376 | 2.09 |
ENSMUST00000037056.9
|
Atp5j2
|
ATP synthase, H+ transporting, mitochondrial F0 complex, subunit F2 |
| chr5_-_76511634 | 2.06 |
ENSMUST00000031146.3
|
Nmu
|
neuromedin U |
| chrM_+_7779 | 2.05 |
ENSMUST00000082408.1
|
mt-Atp6
|
mitochondrially encoded ATP synthase 6 |
| chr5_+_67464284 | 2.03 |
ENSMUST00000113676.6
ENSMUST00000162372.8 |
Slc30a9
|
solute carrier family 30 (zinc transporter), member 9 |
| chr2_+_172994841 | 2.00 |
ENSMUST00000029017.6
|
Pck1
|
phosphoenolpyruvate carboxykinase 1, cytosolic |
| chr8_+_67933573 | 1.99 |
ENSMUST00000212171.2
|
Nat1
|
N-acetyl transferase 1 |
| chr9_-_43151179 | 1.98 |
ENSMUST00000034512.7
|
Oaf
|
out at first homolog |
| chr17_-_24851549 | 1.88 |
ENSMUST00000227607.2
ENSMUST00000227509.2 ENSMUST00000227745.2 |
Tsc2
|
TSC complex subunit 2 |
| chr2_+_73142945 | 1.87 |
ENSMUST00000090811.11
ENSMUST00000112050.2 |
Scrn3
|
secernin 3 |
| chr16_-_58895441 | 1.85 |
ENSMUST00000071243.4
|
Olfr190
|
olfactory receptor 190 |
| chr7_-_19093383 | 1.81 |
ENSMUST00000047036.10
|
Cd3eap
|
CD3E antigen, epsilon polypeptide associated protein |
| chr5_-_145128321 | 1.78 |
ENSMUST00000161845.2
|
Atp5j2
|
ATP synthase, H+ transporting, mitochondrial F0 complex, subunit F2 |
| chr12_-_108859123 | 1.78 |
ENSMUST00000161154.2
ENSMUST00000161410.8 |
Wars
|
tryptophanyl-tRNA synthetase |
| chr3_+_107198528 | 1.75 |
ENSMUST00000029502.14
|
Slc16a4
|
solute carrier family 16 (monocarboxylic acid transporters), member 4 |
| chr2_-_110136074 | 1.75 |
ENSMUST00000046233.9
|
Bbox1
|
butyrobetaine (gamma), 2-oxoglutarate dioxygenase 1 (gamma-butyrobetaine hydroxylase) |
| chr8_+_105558204 | 1.67 |
ENSMUST00000059449.7
|
Ces2b
|
carboxyesterase 2B |
| chr2_-_113844100 | 1.66 |
ENSMUST00000090275.5
|
Gjd2
|
gap junction protein, delta 2 |
| chr9_+_38179147 | 1.64 |
ENSMUST00000212156.3
|
Olfr895
|
olfactory receptor 895 |
| chr11_-_102469896 | 1.64 |
ENSMUST00000107080.2
|
Gm11627
|
predicted gene 11627 |
| chr6_+_15185399 | 1.63 |
ENSMUST00000115474.8
ENSMUST00000115472.8 ENSMUST00000031545.14 |
Foxp2
|
forkhead box P2 |
| chr3_+_116653113 | 1.60 |
ENSMUST00000040260.11
|
Frrs1
|
ferric-chelate reductase 1 |
| chr9_+_39818632 | 1.60 |
ENSMUST00000214904.2
|
Olfr229
|
olfactory receptor 229 |
| chr5_+_90666791 | 1.56 |
ENSMUST00000113179.9
ENSMUST00000128740.2 |
Afm
|
afamin |
| chr2_-_89675315 | 1.55 |
ENSMUST00000099762.2
|
Olfr48
|
olfactory receptor 48 |
| chr4_-_144447974 | 1.51 |
ENSMUST00000036876.8
|
AAdacl4fm3
|
AADACL4 family member 3 |
| chr15_+_21111428 | 1.50 |
ENSMUST00000075132.8
|
Cdh12
|
cadherin 12 |
| chr13_+_22462487 | 1.47 |
ENSMUST00000091736.5
|
Vmn1r195
|
vomeronasal 1 receptor 195 |
| chr3_-_94490023 | 1.46 |
ENSMUST00000029783.16
|
Snx27
|
sorting nexin family member 27 |
| chr3_-_57599956 | 1.45 |
ENSMUST00000238789.2
ENSMUST00000197088.5 ENSMUST00000099091.4 |
Ankub1
|
ankrin repeat and ubiquitin domain containing 1 |
| chr8_+_89309408 | 1.42 |
ENSMUST00000211113.2
|
Nkd1
|
naked cuticle 1 |
| chr18_-_32044877 | 1.42 |
ENSMUST00000054984.8
|
Sft2d3
|
SFT2 domain containing 3 |
| chr2_-_34760960 | 1.41 |
ENSMUST00000028225.12
|
Psmd5
|
proteasome (prosome, macropain) 26S subunit, non-ATPase, 5 |
| chr7_+_88079534 | 1.41 |
ENSMUST00000208478.2
|
Rab38
|
RAB38, member RAS oncogene family |
| chr17_+_17594808 | 1.32 |
ENSMUST00000232396.2
ENSMUST00000232199.2 |
Riok2
|
RIO kinase 2 |
| chr10_-_4382311 | 1.31 |
ENSMUST00000126102.8
ENSMUST00000131853.2 ENSMUST00000042251.11 |
Rmnd1
|
required for meiotic nuclear division 1 homolog |
| chr2_+_85519775 | 1.25 |
ENSMUST00000054868.2
|
Olfr1008
|
olfactory receptor 1008 |
| chr14_-_30213408 | 1.23 |
ENSMUST00000112250.6
|
Cacna1d
|
calcium channel, voltage-dependent, L type, alpha 1D subunit |
| chrX_-_166907286 | 1.23 |
ENSMUST00000239138.2
|
Frmpd4
|
FERM and PDZ domain containing 4 |
| chr17_-_24851581 | 1.20 |
ENSMUST00000226398.2
ENSMUST00000226284.2 ENSMUST00000097373.2 ENSMUST00000228412.2 |
Tsc2
|
TSC complex subunit 2 |
| chr8_-_49296915 | 1.19 |
ENSMUST00000211812.2
|
Tenm3
|
teneurin transmembrane protein 3 |
| chr1_-_170695328 | 1.17 |
ENSMUST00000027974.7
|
Atf6
|
activating transcription factor 6 |
| chr14_+_48683581 | 1.15 |
ENSMUST00000227440.2
ENSMUST00000124720.8 ENSMUST00000226422.2 ENSMUST00000226400.2 |
Tmem260
|
transmembrane protein 260 |
| chr5_-_77053310 | 1.10 |
ENSMUST00000146570.8
ENSMUST00000142450.2 ENSMUST00000120963.8 |
Aasdh
|
aminoadipate-semialdehyde dehydrogenase |
| chrX_+_100683662 | 1.09 |
ENSMUST00000119299.8
ENSMUST00000044475.5 |
Ogt
|
O-linked N-acetylglucosamine (GlcNAc) transferase (UDP-N-acetylglucosamine:polypeptide-N-acetylglucosaminyl transferase) |
| chr8_+_105048199 | 1.08 |
ENSMUST00000109447.2
|
Gm11020
|
predicted gene 11020 |
| chr2_+_111136546 | 1.06 |
ENSMUST00000090329.2
|
Olfr1279
|
olfactory receptor 1279 |
| chr8_-_41507808 | 1.03 |
ENSMUST00000093534.11
|
Mtus1
|
mitochondrial tumor suppressor 1 |
| chr4_+_118522716 | 1.03 |
ENSMUST00000102666.5
|
Olfr62
|
olfactory receptor 62 |
| chr7_-_12096691 | 1.03 |
ENSMUST00000086228.3
|
Vmn1r84
|
vomeronasal 1 receptor 84 |
| chr10_-_4382283 | 1.00 |
ENSMUST00000155172.8
|
Rmnd1
|
required for meiotic nuclear division 1 homolog |
| chr10_-_129448439 | 0.97 |
ENSMUST00000214584.3
|
Olfr796
|
olfactory receptor 796 |
| chr2_+_89842475 | 0.97 |
ENSMUST00000214382.2
ENSMUST00000217065.3 |
Olfr1263
|
olfactory receptor 1263 |
| chr10_+_129219952 | 0.96 |
ENSMUST00000214064.2
|
Olfr784
|
olfactory receptor 784 |
| chr1_-_59200718 | 0.96 |
ENSMUST00000078874.14
ENSMUST00000066374.14 |
Mpp4
|
membrane protein, palmitoylated 4 (MAGUK p55 subfamily member 4) |
| chr3_+_20011405 | 0.96 |
ENSMUST00000108325.9
|
Cp
|
ceruloplasmin |
| chr18_+_44960813 | 0.95 |
ENSMUST00000037763.11
|
Ythdc2
|
YTH domain containing 2 |
| chr13_+_23433408 | 0.93 |
ENSMUST00000091719.2
|
Vmn1r223
|
vomeronasal 1 receptor 223 |
| chr3_-_59118293 | 0.93 |
ENSMUST00000040622.3
|
P2ry13
|
purinergic receptor P2Y, G-protein coupled 13 |
| chr6_-_119085508 | 0.92 |
ENSMUST00000188522.7
|
Cacna1c
|
calcium channel, voltage-dependent, L type, alpha 1C subunit |
| chr17_-_37481536 | 0.91 |
ENSMUST00000174673.3
|
Olfr753-ps1
|
olfactory receptor 753, pseudogene 1 |
| chr4_-_113856697 | 0.88 |
ENSMUST00000169631.8
ENSMUST00000170105.8 ENSMUST00000171627.9 |
Skint5
|
selection and upkeep of intraepithelial T cells 5 |
| chr4_+_19818718 | 0.87 |
ENSMUST00000035890.8
|
Slc7a13
|
solute carrier family 7, (cationic amino acid transporter, y+ system) member 13 |
| chr2_-_13798843 | 0.86 |
ENSMUST00000003509.10
|
St8sia6
|
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 6 |
| chr9_+_65102635 | 0.85 |
ENSMUST00000216702.2
|
Parp16
|
poly (ADP-ribose) polymerase family, member 16 |
| chr5_-_77053226 | 0.81 |
ENSMUST00000135954.2
|
Aasdh
|
aminoadipate-semialdehyde dehydrogenase |
| chr19_-_24938909 | 0.78 |
ENSMUST00000025815.10
|
Cbwd1
|
COBW domain containing 1 |
| chr6_+_132739094 | 0.76 |
ENSMUST00000069268.3
|
Tas2r102
|
taste receptor, type 2, member 102 |
| chr16_-_58647137 | 0.74 |
ENSMUST00000215069.2
ENSMUST00000079955.6 |
Olfr175
|
olfactory receptor 175 |
| chr16_-_19390500 | 0.74 |
ENSMUST00000215040.2
ENSMUST00000216070.2 ENSMUST00000215476.2 |
Olfr169
|
olfactory receptor 169 |
| chr2_-_111251294 | 0.74 |
ENSMUST00000099617.2
|
Olfr1286
|
olfactory receptor 1286 |
| chr19_-_12302465 | 0.71 |
ENSMUST00000207241.3
|
Olfr1437
|
olfactory receptor 1437 |
| chr5_+_138159333 | 0.71 |
ENSMUST00000019638.15
ENSMUST00000110951.8 |
Cops6
|
COP9 signalosome subunit 6 |
| chr19_+_13208692 | 0.70 |
ENSMUST00000207246.4
|
Olfr1463
|
olfactory receptor 1463 |
| chr7_-_48493388 | 0.69 |
ENSMUST00000167786.4
|
Csrp3
|
cysteine and glycine-rich protein 3 |
| chr5_+_125080504 | 0.69 |
ENSMUST00000197746.2
|
Rflna
|
refilin A |
| chr2_-_104680104 | 0.68 |
ENSMUST00000028593.11
|
Prrg4
|
proline rich Gla (G-carboxyglutamic acid) 4 (transmembrane) |
| chr6_+_40941688 | 0.68 |
ENSMUST00000076638.7
|
1810009J06Rik
|
RIKEN cDNA 1810009J06 gene |
| chr11_+_102175985 | 0.65 |
ENSMUST00000156326.2
|
Tmub2
|
transmembrane and ubiquitin-like domain containing 2 |
| chr7_+_88079465 | 0.64 |
ENSMUST00000107256.4
|
Rab38
|
RAB38, member RAS oncogene family |
| chr7_-_104628324 | 0.61 |
ENSMUST00000217091.2
ENSMUST00000210963.3 |
Olfr671
|
olfactory receptor 671 |
| chr2_-_89273344 | 0.61 |
ENSMUST00000216123.2
|
Olfr1240
|
olfactory receptor 1240 |
| chr16_+_23247883 | 0.60 |
ENSMUST00000038730.7
|
Rtp1
|
receptor transporter protein 1 |
| chr10_-_129577771 | 0.60 |
ENSMUST00000215142.3
ENSMUST00000213239.2 |
Olfr806
|
olfactory receptor 806 |
| chr17_+_33848054 | 0.57 |
ENSMUST00000166627.8
ENSMUST00000073570.12 ENSMUST00000170225.3 |
Zfp414
|
zinc finger protein 414 |
| chr2_+_21210527 | 0.57 |
ENSMUST00000054591.10
ENSMUST00000102952.8 ENSMUST00000138965.8 ENSMUST00000138914.8 ENSMUST00000102951.2 |
Thnsl1
|
threonine synthase-like 1 (bacterial) |
| chr9_+_23285228 | 0.57 |
ENSMUST00000214050.2
|
Bmper
|
BMP-binding endothelial regulator |
| chr6_-_131664458 | 0.56 |
ENSMUST00000053652.6
|
Tas2r105
|
taste receptor, type 2, member 105 |
| chr2_-_88854443 | 0.55 |
ENSMUST00000099799.5
|
Olfr1217
|
olfactory receptor 1217 |
| chr6_+_11926757 | 0.54 |
ENSMUST00000133776.2
|
Phf14
|
PHD finger protein 14 |
| chr13_-_62771948 | 0.53 |
ENSMUST00000223338.2
ENSMUST00000201563.4 |
6720489N17Rik
|
RIKEN cDNA 6720489N17 gene |
| chr19_-_57227742 | 0.51 |
ENSMUST00000111559.8
|
Ablim1
|
actin-binding LIM protein 1 |
| chr3_-_100592747 | 0.50 |
ENSMUST00000008907.14
|
Man1a2
|
mannosidase, alpha, class 1A, member 2 |
| chr11_+_101137786 | 0.49 |
ENSMUST00000107282.4
|
Ramp2
|
receptor (calcitonin) activity modifying protein 2 |
| chr17_-_37422872 | 0.48 |
ENSMUST00000168659.2
|
Olfr92
|
olfactory receptor 92 |
| chr6_+_67736650 | 0.43 |
ENSMUST00000103305.2
|
Igkv1-132
|
immunoglobulin kappa variable 1-132 |
| chr6_-_129890129 | 0.43 |
ENSMUST00000169901.7
ENSMUST00000014683.13 |
Klra5
|
killer cell lectin-like receptor, subfamily A, member 5 |
| chr11_-_75828551 | 0.42 |
ENSMUST00000121287.8
|
Rph3al
|
rabphilin 3A-like (without C2 domains) |
| chr6_+_65502292 | 0.42 |
ENSMUST00000212402.2
|
Tnip3
|
TNFAIP3 interacting protein 3 |
| chr10_+_103987773 | 0.38 |
ENSMUST00000180692.3
|
Gm6763
|
predicted gene 6763 |
| chr10_+_103996282 | 0.38 |
ENSMUST00000181166.3
|
Gm8764
|
predicted gene 8764 |
| chr2_+_85420854 | 0.37 |
ENSMUST00000052307.5
|
Olfr998
|
olfactory receptor 998 |
| chr10_+_104030318 | 0.36 |
ENSMUST00000181036.2
|
Gm4340
|
predicted gene 4340 |
| chr10_+_103979263 | 0.36 |
ENSMUST00000180664.2
|
Gm21293
|
predicted gene, 21293 |
| chr10_+_104004791 | 0.36 |
ENSMUST00000180568.2
|
Gm21304
|
predicted gene, 21304 |
| chr10_+_104013300 | 0.36 |
ENSMUST00000181239.2
|
Gm21312
|
predicted gene, 21312 |
| chr10_+_104021809 | 0.36 |
ENSMUST00000181707.2
|
Gm20765
|
predicted gene, 20765 |
| chr3_-_100591906 | 0.36 |
ENSMUST00000130066.2
|
Man1a2
|
mannosidase, alpha, class 1A, member 2 |
| chr3_+_115681788 | 0.35 |
ENSMUST00000196804.5
ENSMUST00000106505.8 ENSMUST00000043342.10 |
Dph5
|
diphthamide biosynthesis 5 |
| chr17_+_17594526 | 0.35 |
ENSMUST00000024620.8
|
Riok2
|
RIO kinase 2 |
| chr10_+_129184719 | 0.35 |
ENSMUST00000213970.2
|
Olfr782
|
olfactory receptor 782 |
| chr10_+_129493563 | 0.34 |
ENSMUST00000217094.2
|
Olfr800
|
olfactory receptor 800 |
| chr10_+_104029903 | 0.34 |
ENSMUST00000181615.8
|
Gm4340
|
predicted gene 4340 |
| chr2_-_88562710 | 0.34 |
ENSMUST00000213118.2
|
Olfr1197
|
olfactory receptor 1197 |
| chr10_+_103978848 | 0.34 |
ENSMUST00000181634.8
|
Gm21293
|
predicted gene, 21293 |
| chr10_+_103987358 | 0.34 |
ENSMUST00000181287.5
|
Gm6763
|
predicted gene 6763 |
| chr10_+_103995867 | 0.34 |
ENSMUST00000181179.5
|
Gm8764
|
predicted gene 8764 |
| chr10_+_104004376 | 0.34 |
ENSMUST00000181059.8
|
Gm21304
|
predicted gene, 21304 |
| chr10_+_104012885 | 0.34 |
ENSMUST00000180889.8
|
Gm21312
|
predicted gene, 21312 |
| chr10_+_104021394 | 0.34 |
ENSMUST00000181703.8
|
Gm20765
|
predicted gene, 20765 |
| chr7_+_114318746 | 0.34 |
ENSMUST00000182044.2
|
Calcb
|
calcitonin-related polypeptide, beta |
| chr17_+_24851647 | 0.34 |
ENSMUST00000047611.4
|
Nthl1
|
nth (endonuclease III)-like 1 (E.coli) |
| chr2_-_88577230 | 0.32 |
ENSMUST00000099814.2
|
Olfr1198
|
olfactory receptor 1198 |
| chr7_+_12711760 | 0.32 |
ENSMUST00000108535.8
ENSMUST00000108537.8 ENSMUST00000045810.15 |
Zfp446
|
zinc finger protein 446 |
| chr7_-_104651236 | 0.32 |
ENSMUST00000213942.3
|
Olfr672
|
olfactory receptor 672 |
| chr13_+_95460110 | 0.32 |
ENSMUST00000221807.2
|
Zbed3
|
zinc finger, BED type containing 3 |
| chr4_-_155729865 | 0.31 |
ENSMUST00000115821.3
|
Gm10563
|
predicted gene 10563 |
| chr14_+_53743184 | 0.30 |
ENSMUST00000103583.5
|
Trav10
|
T cell receptor alpha variable 10 |
| chr16_-_88548523 | 0.29 |
ENSMUST00000053149.4
|
Krtap13
|
keratin associated protein 13 |
| chr7_+_6200089 | 0.29 |
ENSMUST00000037553.6
|
Galp
|
galanin-like peptide |
| chr6_-_3763618 | 0.29 |
ENSMUST00000171613.8
|
Calcr
|
calcitonin receptor |
| chr17_-_22127073 | 0.25 |
ENSMUST00000106026.3
|
Zfp995
|
zinc finger protein 995 |
| chr4_+_53011880 | 0.24 |
ENSMUST00000015391.10
|
Nipsnap3b
|
nipsnap homolog 3B |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.5 | 7.5 | GO:0030827 | negative regulation of cGMP metabolic process(GO:0030824) negative regulation of cGMP biosynthetic process(GO:0030827) negative regulation of guanylate cyclase activity(GO:0031283) |
| 2.3 | 6.9 | GO:0043387 | mycotoxin catabolic process(GO:0043387) aflatoxin catabolic process(GO:0046223) organic heteropentacyclic compound catabolic process(GO:1901377) |
| 2.2 | 6.5 | GO:0030070 | insulin processing(GO:0030070) |
| 2.0 | 6.1 | GO:0033306 | phytol metabolic process(GO:0033306) fatty alcohol metabolic process(GO:1903173) |
| 0.9 | 5.5 | GO:0090126 | protein complex assembly involved in synapse maturation(GO:0090126) |
| 0.9 | 3.6 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.8 | 4.0 | GO:0070346 | positive regulation of fat cell proliferation(GO:0070346) |
| 0.6 | 8.6 | GO:0019227 | neuronal action potential propagation(GO:0019227) action potential propagation(GO:0098870) |
| 0.6 | 3.1 | GO:0044861 | protein transport into plasma membrane raft(GO:0044861) |
| 0.5 | 2.0 | GO:0006114 | glycerol biosynthetic process(GO:0006114) positive regulation of transcription from RNA polymerase II promoter in response to acidic pH(GO:0061402) |
| 0.5 | 4.3 | GO:0006030 | chitin metabolic process(GO:0006030) chitin catabolic process(GO:0006032) |
| 0.5 | 1.4 | GO:0070682 | proteasome regulatory particle assembly(GO:0070682) |
| 0.5 | 5.1 | GO:0008627 | intrinsic apoptotic signaling pathway in response to osmotic stress(GO:0008627) |
| 0.5 | 2.3 | GO:0099566 | regulation of postsynaptic cytosolic calcium ion concentration(GO:0099566) |
| 0.5 | 2.7 | GO:0097118 | neuroligin clustering involved in postsynaptic membrane assembly(GO:0097118) |
| 0.4 | 1.8 | GO:0006436 | tryptophanyl-tRNA aminoacylation(GO:0006436) |
| 0.4 | 4.9 | GO:0060406 | positive regulation of penile erection(GO:0060406) |
| 0.4 | 1.7 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 0.4 | 1.9 | GO:0019482 | beta-alanine metabolic process(GO:0019482) |
| 0.4 | 1.1 | GO:0090526 | regulation of gluconeogenesis involved in cellular glucose homeostasis(GO:0090526) |
| 0.4 | 2.5 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
| 0.3 | 2.4 | GO:0001757 | somite specification(GO:0001757) |
| 0.3 | 1.2 | GO:0086046 | membrane depolarization during SA node cell action potential(GO:0086046) |
| 0.3 | 3.1 | GO:0042760 | very long-chain fatty acid catabolic process(GO:0042760) |
| 0.3 | 2.1 | GO:0009313 | oligosaccharide catabolic process(GO:0009313) |
| 0.3 | 2.0 | GO:1903232 | platelet dense granule organization(GO:0060155) phagosome acidification(GO:0090383) melanosome assembly(GO:1903232) |
| 0.3 | 5.5 | GO:0048934 | peripheral nervous system neuron differentiation(GO:0048934) peripheral nervous system neuron development(GO:0048935) |
| 0.3 | 13.7 | GO:0010107 | potassium ion import(GO:0010107) |
| 0.3 | 5.5 | GO:2000480 | negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
| 0.3 | 2.6 | GO:1900034 | regulation of cellular response to heat(GO:1900034) |
| 0.2 | 1.2 | GO:0051835 | positive regulation of synapse structural plasticity(GO:0051835) |
| 0.2 | 0.7 | GO:1903920 | positive regulation of actin filament severing(GO:1903920) |
| 0.2 | 4.6 | GO:0045161 | neuronal ion channel clustering(GO:0045161) |
| 0.2 | 1.7 | GO:0090070 | positive regulation of ribosome biogenesis(GO:0090070) positive regulation of rRNA processing(GO:2000234) |
| 0.2 | 1.4 | GO:0038031 | non-canonical Wnt signaling pathway via JNK cascade(GO:0038031) |
| 0.2 | 2.3 | GO:0070131 | positive regulation of mitochondrial translation(GO:0070131) |
| 0.2 | 2.6 | GO:0070257 | positive regulation of mucus secretion(GO:0070257) |
| 0.2 | 0.7 | GO:1900158 | negative regulation of bone mineralization involved in bone maturation(GO:1900158) |
| 0.2 | 3.1 | GO:2001269 | positive regulation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:2001269) |
| 0.2 | 0.9 | GO:0098912 | membrane depolarization during atrial cardiac muscle cell action potential(GO:0098912) |
| 0.1 | 2.1 | GO:2000821 | regulation of grooming behavior(GO:2000821) |
| 0.1 | 0.8 | GO:0070213 | protein auto-ADP-ribosylation(GO:0070213) |
| 0.1 | 4.4 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 0.1 | 4.6 | GO:0015985 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.1 | 4.1 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 0.1 | 0.6 | GO:0017183 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 0.1 | 0.3 | GO:0006296 | base-excision repair, AP site formation(GO:0006285) nucleotide-excision repair, DNA incision, 5'-to lesion(GO:0006296) |
| 0.1 | 0.5 | GO:2000583 | regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000583) negative regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000584) |
| 0.1 | 0.8 | GO:0038042 | dimeric G-protein coupled receptor signaling pathway(GO:0038042) calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.1 | 0.9 | GO:0044829 | positive regulation by host of viral genome replication(GO:0044829) |
| 0.1 | 3.9 | GO:0006754 | ATP biosynthetic process(GO:0006754) |
| 0.1 | 11.5 | GO:0006333 | chromatin assembly or disassembly(GO:0006333) |
| 0.1 | 5.3 | GO:0046513 | ceramide biosynthetic process(GO:0046513) |
| 0.1 | 1.2 | GO:1990440 | positive regulation of transcription from RNA polymerase II promoter in response to endoplasmic reticulum stress(GO:1990440) |
| 0.1 | 4.2 | GO:0003301 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.1 | 1.8 | GO:0009303 | rRNA transcription(GO:0009303) |
| 0.1 | 1.2 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.1 | 1.5 | GO:1990126 | retrograde transport, endosome to plasma membrane(GO:1990126) |
| 0.0 | 1.9 | GO:0001580 | detection of chemical stimulus involved in sensory perception of bitter taste(GO:0001580) |
| 0.0 | 5.6 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.0 | 2.4 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.0 | 2.0 | GO:0006829 | zinc II ion transport(GO:0006829) |
| 0.0 | 0.7 | GO:0000338 | protein deneddylation(GO:0000338) |
| 0.0 | 3.3 | GO:0006749 | glutathione metabolic process(GO:0006749) |
| 0.0 | 5.2 | GO:0016052 | carbohydrate catabolic process(GO:0016052) |
| 0.0 | 0.6 | GO:0002043 | blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:0002043) |
| 0.0 | 0.9 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.0 | 1.0 | GO:0046688 | response to copper ion(GO:0046688) |
| 0.0 | 0.9 | GO:0031280 | negative regulation of cyclase activity(GO:0031280) |
| 0.0 | 1.6 | GO:0051180 | vitamin transport(GO:0051180) |
| 0.0 | 1.0 | GO:0010758 | regulation of macrophage chemotaxis(GO:0010758) |
| 0.0 | 6.0 | GO:0010466 | negative regulation of peptidase activity(GO:0010466) |
| 0.0 | 6.6 | GO:0000377 | RNA splicing, via transesterification reactions with bulged adenosine as nucleophile(GO:0000377) mRNA splicing, via spliceosome(GO:0000398) |
| 0.0 | 0.9 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.0 | 0.1 | GO:0042706 | eye photoreceptor cell fate commitment(GO:0042706) photoreceptor cell fate commitment(GO:0046552) |
| 0.0 | 0.3 | GO:0032098 | regulation of appetite(GO:0032098) |
| 0.0 | 1.0 | GO:0035418 | protein localization to synapse(GO:0035418) |
| 0.0 | 3.3 | GO:0050907 | detection of chemical stimulus involved in sensory perception(GO:0050907) |
| 0.0 | 1.5 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.0 | 0.6 | GO:0045744 | negative regulation of G-protein coupled receptor protein signaling pathway(GO:0045744) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.3 | 9.8 | GO:0016014 | dystrobrevin complex(GO:0016014) |
| 0.8 | 3.1 | GO:0033596 | TSC1-TSC2 complex(GO:0033596) |
| 0.7 | 8.0 | GO:0005687 | U4 snRNP(GO:0005687) |
| 0.6 | 16.8 | GO:0016514 | SWI/SNF complex(GO:0016514) |
| 0.6 | 2.9 | GO:0097057 | TRAF2-GSTP1 complex(GO:0097057) |
| 0.5 | 7.5 | GO:0044327 | dendritic spine head(GO:0044327) |
| 0.5 | 8.4 | GO:0045263 | proton-transporting ATP synthase complex, coupling factor F(o)(GO:0045263) |
| 0.4 | 8.2 | GO:0036056 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.4 | 3.6 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.3 | 5.5 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.3 | 6.5 | GO:0031045 | dense core granule(GO:0031045) |
| 0.3 | 0.9 | GO:0002095 | caveolar macromolecular signaling complex(GO:0002095) |
| 0.3 | 2.6 | GO:0045298 | tubulin complex(GO:0045298) |
| 0.3 | 2.4 | GO:1990357 | terminal web(GO:1990357) |
| 0.3 | 2.0 | GO:0045009 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.2 | 5.1 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.2 | 2.5 | GO:0005915 | zonula adherens(GO:0005915) |
| 0.2 | 9.9 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.1 | 2.3 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.1 | 4.6 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 0.1 | 0.8 | GO:1903439 | calcitonin family receptor complex(GO:1903439) amylin receptor complex(GO:1903440) |
| 0.1 | 1.8 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.1 | 1.7 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.1 | 4.4 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.1 | 1.6 | GO:0008540 | proteasome regulatory particle, base subcomplex(GO:0008540) |
| 0.1 | 2.4 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.1 | 1.5 | GO:0071203 | WASH complex(GO:0071203) |
| 0.1 | 2.6 | GO:0042470 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.1 | 6.1 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.1 | 1.7 | GO:0005922 | connexon complex(GO:0005922) |
| 0.0 | 1.4 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.0 | 2.4 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 0.8 | GO:0071782 | endoplasmic reticulum tubular network(GO:0071782) |
| 0.0 | 8.1 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.0 | 1.1 | GO:0000791 | euchromatin(GO:0000791) |
| 0.0 | 3.1 | GO:0032993 | protein-DNA complex(GO:0032993) |
| 0.0 | 0.1 | GO:0031673 | H zone(GO:0031673) |
| 0.0 | 0.2 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.0 | 7.2 | GO:0005743 | mitochondrial inner membrane(GO:0005743) |
| 0.0 | 1.0 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.0 | 1.0 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.0 | 2.1 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 0.7 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 1.2 | GO:0043197 | dendritic spine(GO:0043197) |
| 0.0 | 4.7 | GO:0014069 | postsynaptic density(GO:0014069) |
| 0.0 | 20.0 | GO:0005739 | mitochondrion(GO:0005739) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.4 | 13.2 | GO:0016838 | carbon-oxygen lyase activity, acting on phosphates(GO:0016838) |
| 1.5 | 6.1 | GO:0019166 | trans-2-enoyl-CoA reductase (NADPH) activity(GO:0019166) |
| 1.1 | 4.5 | GO:0042284 | sphingolipid delta-4 desaturase activity(GO:0042284) |
| 1.1 | 3.3 | GO:0035731 | S-nitrosoglutathione binding(GO:0035730) dinitrosyl-iron complex binding(GO:0035731) |
| 0.7 | 5.2 | GO:0004556 | alpha-amylase activity(GO:0004556) |
| 0.7 | 2.0 | GO:0036461 | BLOC-2 complex binding(GO:0036461) |
| 0.7 | 2.0 | GO:0004611 | phosphoenolpyruvate carboxykinase activity(GO:0004611) phosphoenolpyruvate carboxykinase (GTP) activity(GO:0004613) |
| 0.5 | 6.5 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.5 | 1.6 | GO:0008431 | vitamin E binding(GO:0008431) |
| 0.5 | 2.6 | GO:0099609 | microtubule lateral binding(GO:0099609) |
| 0.5 | 4.3 | GO:0004568 | chitinase activity(GO:0004568) |
| 0.5 | 2.7 | GO:0031697 | beta-1 adrenergic receptor binding(GO:0031697) |
| 0.4 | 1.8 | GO:0004830 | tryptophan-tRNA ligase activity(GO:0004830) |
| 0.4 | 2.1 | GO:0052796 | exo-alpha-sialidase activity(GO:0004308) alpha-sialidase activity(GO:0016997) exo-alpha-(2->3)-sialidase activity(GO:0052794) exo-alpha-(2->6)-sialidase activity(GO:0052795) exo-alpha-(2->8)-sialidase activity(GO:0052796) |
| 0.4 | 10.8 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.4 | 2.0 | GO:0004060 | arylamine N-acetyltransferase activity(GO:0004060) |
| 0.4 | 5.5 | GO:0097109 | neuroligin family protein binding(GO:0097109) |
| 0.4 | 14.6 | GO:0005242 | inward rectifier potassium channel activity(GO:0005242) |
| 0.4 | 5.5 | GO:0008603 | cAMP-dependent protein kinase regulator activity(GO:0008603) |
| 0.4 | 1.1 | GO:0016262 | protein N-acetylglucosaminyltransferase activity(GO:0016262) |
| 0.3 | 1.9 | GO:0004043 | L-aminoadipate-semialdehyde dehydrogenase activity(GO:0004043) |
| 0.3 | 5.1 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.3 | 2.3 | GO:0099583 | neurotransmitter receptor activity involved in regulation of postsynaptic cytosolic calcium ion concentration(GO:0099583) |
| 0.3 | 4.6 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.2 | 9.6 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.2 | 11.3 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.2 | 1.6 | GO:0000293 | ferric-chelate reductase activity(GO:0000293) |
| 0.2 | 2.6 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.2 | 3.1 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
| 0.1 | 0.8 | GO:0097643 | amylin receptor activity(GO:0097643) |
| 0.1 | 4.1 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.1 | 0.7 | GO:0061629 | RNA polymerase II sequence-specific DNA binding transcription factor binding(GO:0061629) |
| 0.1 | 0.9 | GO:1990247 | N6-methyladenosine-containing RNA binding(GO:1990247) |
| 0.1 | 0.3 | GO:0000702 | oxidized base lesion DNA N-glycosylase activity(GO:0000702) |
| 0.1 | 1.9 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.1 | 2.5 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) |
| 0.1 | 1.0 | GO:0004322 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.1 | 11.2 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.1 | 0.6 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.1 | 0.9 | GO:0004571 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) |
| 0.1 | 1.3 | GO:0033038 | bitter taste receptor activity(GO:0033038) |
| 0.1 | 3.1 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.1 | 1.2 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.0 | 2.0 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.0 | 2.1 | GO:0071855 | neuropeptide receptor binding(GO:0071855) |
| 0.0 | 0.9 | GO:0045028 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.0 | 8.1 | GO:0051117 | ATPase binding(GO:0051117) |
| 0.0 | 0.6 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.0 | 3.9 | GO:0042626 | ATPase activity, coupled to transmembrane movement of substances(GO:0042626) |
| 0.0 | 4.6 | GO:0015078 | hydrogen ion transmembrane transporter activity(GO:0015078) |
| 0.0 | 1.8 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 6.0 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 3.6 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.0 | 4.0 | GO:0005179 | hormone activity(GO:0005179) |
| 0.0 | 1.8 | GO:0008028 | monocarboxylic acid transmembrane transporter activity(GO:0008028) |
| 0.0 | 0.7 | GO:0031005 | filamin binding(GO:0031005) |
| 0.0 | 0.8 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 0.1 | GO:0004977 | melanocortin receptor activity(GO:0004977) |
| 0.0 | 1.5 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.0 | 1.7 | GO:0016706 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, 2-oxoglutarate as one donor, and incorporation of one atom each of oxygen into both donors(GO:0016706) |
| 0.0 | 1.2 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.0 | 0.9 | GO:0008373 | sialyltransferase activity(GO:0008373) |
| 0.0 | 1.3 | GO:0005550 | pheromone binding(GO:0005550) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 8.6 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.2 | 2.6 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.1 | 3.6 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.1 | 5.5 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.1 | 6.5 | PID ARF6 TRAFFICKING PATHWAY | Arf6 trafficking events |
| 0.1 | 8.1 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.0 | 2.3 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.0 | 2.0 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.0 | 1.4 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.0 | 1.2 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.0 | 5.6 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 14.6 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.3 | 6.5 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.2 | 3.1 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.2 | 3.6 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.2 | 3.9 | REACTOME FORMATION OF ATP BY CHEMIOSMOTIC COUPLING | Genes involved in Formation of ATP by chemiosmotic coupling |
| 0.2 | 5.5 | REACTOME PKA MEDIATED PHOSPHORYLATION OF CREB | Genes involved in PKA-mediated phosphorylation of CREB |
| 0.1 | 4.9 | REACTOME GLUCAGON TYPE LIGAND RECEPTORS | Genes involved in Glucagon-type ligand receptors |
| 0.1 | 6.9 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 8.6 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.1 | 2.0 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.1 | 1.2 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.1 | 5.1 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.1 | 2.3 | REACTOME CREB PHOSPHORYLATION THROUGH THE ACTIVATION OF CAMKII | Genes involved in CREB phosphorylation through the activation of CaMKII |
| 0.1 | 2.7 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.1 | 2.6 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.1 | 1.7 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.1 | 1.8 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.1 | 0.9 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.0 | 2.5 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 2.1 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.0 | 1.5 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 0.9 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.0 | 0.9 | REACTOME TRANSPORT TO THE GOLGI AND SUBSEQUENT MODIFICATION | Genes involved in Transport to the Golgi and subsequent modification |
| 0.0 | 1.0 | REACTOME METAL ION SLC TRANSPORTERS | Genes involved in Metal ion SLC transporters |
| 0.0 | 0.5 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |
| 0.0 | 0.8 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |