Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
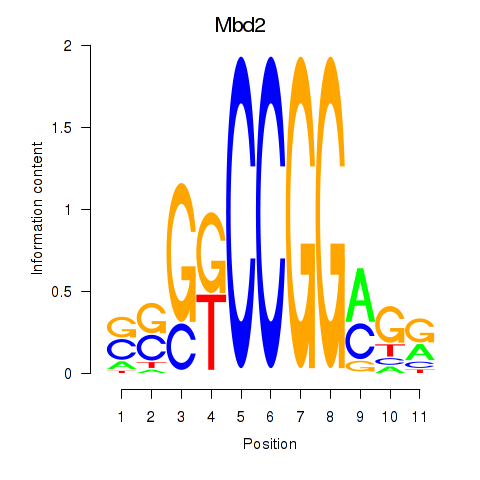
Results for Mbd2
Z-value: 1.11
Transcription factors associated with Mbd2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Mbd2
|
ENSMUSG00000024513.17 | Mbd2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Mbd2 | mm39_v1_chr18_+_70701260_70701469 | -0.14 | 2.3e-01 | Click! |
Activity profile of Mbd2 motif
Sorted Z-values of Mbd2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Mbd2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_+_37025810 | 10.48 |
ENSMUST00000031003.11
|
Ppp2r2c
|
protein phosphatase 2, regulatory subunit B, gamma |
| chr5_-_115332343 | 10.42 |
ENSMUST00000112113.8
|
Cabp1
|
calcium binding protein 1 |
| chr4_+_129878890 | 8.24 |
ENSMUST00000106017.8
ENSMUST00000121049.8 |
Adgrb2
|
adhesion G protein-coupled receptor B2 |
| chr5_+_37025926 | 8.03 |
ENSMUST00000201156.2
|
Ppp2r2c
|
protein phosphatase 2, regulatory subunit B, gamma |
| chr11_-_97944239 | 7.70 |
ENSMUST00000017544.9
|
Stac2
|
SH3 and cysteine rich domain 2 |
| chr12_-_107969673 | 7.34 |
ENSMUST00000109887.8
ENSMUST00000109891.3 |
Bcl11b
|
B cell leukemia/lymphoma 11B |
| chr15_+_74435587 | 6.40 |
ENSMUST00000185682.7
ENSMUST00000170845.8 ENSMUST00000187599.2 |
Adgrb1
|
adhesion G protein-coupled receptor B1 |
| chr15_-_31367872 | 6.33 |
ENSMUST00000123325.9
|
Ankrd33b
|
ankyrin repeat domain 33B |
| chr5_+_137286535 | 5.95 |
ENSMUST00000024099.11
ENSMUST00000196208.5 ENSMUST00000085934.4 |
Ache
|
acetylcholinesterase |
| chr5_-_108697857 | 5.93 |
ENSMUST00000129040.2
ENSMUST00000046892.10 |
Cplx1
|
complexin 1 |
| chr1_+_74894069 | 5.83 |
ENSMUST00000160379.4
|
Cdk5r2
|
cyclin-dependent kinase 5, regulatory subunit 2 (p39) |
| chr7_+_4693759 | 5.80 |
ENSMUST00000048248.9
|
Brsk1
|
BR serine/threonine kinase 1 |
| chr9_+_26645024 | 5.67 |
ENSMUST00000160899.8
ENSMUST00000161431.3 ENSMUST00000159799.8 |
B3gat1
|
beta-1,3-glucuronyltransferase 1 (glucuronosyltransferase P) |
| chr7_+_4693603 | 5.67 |
ENSMUST00000120836.8
|
Brsk1
|
BR serine/threonine kinase 1 |
| chr2_+_157756535 | 5.64 |
ENSMUST00000109523.2
|
Vstm2l
|
V-set and transmembrane domain containing 2-like |
| chr2_+_28082943 | 5.47 |
ENSMUST00000113920.8
|
Olfm1
|
olfactomedin 1 |
| chr2_+_28083105 | 5.42 |
ENSMUST00000100244.10
|
Olfm1
|
olfactomedin 1 |
| chr10_+_126914755 | 5.29 |
ENSMUST00000039259.7
ENSMUST00000217941.2 |
Agap2
|
ArfGAP with GTPase domain, ankyrin repeat and PH domain 2 |
| chr7_+_25005510 | 5.10 |
ENSMUST00000119703.8
ENSMUST00000205639.3 ENSMUST00000108409.2 |
Tmem145
|
transmembrane protein 145 |
| chr8_+_112263632 | 5.07 |
ENSMUST00000173506.8
|
Znrf1
|
zinc and ring finger 1 |
| chr8_-_125161061 | 4.98 |
ENSMUST00000140012.8
|
Pgbd5
|
piggyBac transposable element derived 5 |
| chr7_-_125681577 | 4.92 |
ENSMUST00000073935.7
|
Gsg1l
|
GSG1-like |
| chr6_-_87958611 | 4.83 |
ENSMUST00000056403.7
|
H1f10
|
H1.10 linker histone |
| chr7_-_46782448 | 4.58 |
ENSMUST00000033142.13
|
Ptpn5
|
protein tyrosine phosphatase, non-receptor type 5 |
| chr15_-_31367668 | 4.57 |
ENSMUST00000110410.10
ENSMUST00000076942.5 |
Ankrd33b
|
ankyrin repeat domain 33B |
| chr12_-_72455708 | 4.51 |
ENSMUST00000078505.14
|
Rtn1
|
reticulon 1 |
| chr14_-_68362284 | 4.40 |
ENSMUST00000111089.8
ENSMUST00000022638.6 |
Nefm
|
neurofilament, medium polypeptide |
| chr4_+_155819257 | 4.39 |
ENSMUST00000147721.8
ENSMUST00000127188.3 |
Tmem240
|
transmembrane protein 240 |
| chr4_+_129878627 | 4.31 |
ENSMUST00000120204.8
|
Adgrb2
|
adhesion G protein-coupled receptor B2 |
| chr7_-_64806164 | 4.30 |
ENSMUST00000148459.3
ENSMUST00000119118.8 |
Fam189a1
|
family with sequence similarity 189, member A1 |
| chr13_+_55097200 | 4.26 |
ENSMUST00000026994.14
ENSMUST00000109994.9 |
Unc5a
|
unc-5 netrin receptor A |
| chr11_+_116809669 | 4.25 |
ENSMUST00000103027.10
|
Mgat5b
|
mannoside acetylglucosaminyltransferase 5, isoenzyme B |
| chr8_+_112263465 | 4.16 |
ENSMUST00000095176.12
|
Znrf1
|
zinc and ring finger 1 |
| chr11_-_94364914 | 4.12 |
ENSMUST00000107786.8
ENSMUST00000107791.8 ENSMUST00000103166.9 ENSMUST00000107792.8 ENSMUST00000100561.10 ENSMUST00000107793.8 ENSMUST00000107788.8 ENSMUST00000107790.8 ENSMUST00000107789.8 ENSMUST00000107785.2 ENSMUST00000021234.15 |
Cacna1g
|
calcium channel, voltage-dependent, T type, alpha 1G subunit |
| chr15_-_75438457 | 4.09 |
ENSMUST00000163116.8
ENSMUST00000023241.12 |
Ly6h
|
lymphocyte antigen 6 complex, locus H |
| chr4_-_117146624 | 3.98 |
ENSMUST00000221654.2
|
Rnf220
|
ring finger protein 220 |
| chr15_-_75438660 | 3.90 |
ENSMUST00000065417.15
|
Ly6h
|
lymphocyte antigen 6 complex, locus H |
| chr7_+_141503411 | 3.79 |
ENSMUST00000078200.12
ENSMUST00000018971.15 |
Brsk2
|
BR serine/threonine kinase 2 |
| chr2_+_83642910 | 3.74 |
ENSMUST00000051454.4
|
Fam171b
|
family with sequence similarity 171, member B |
| chr9_+_26645141 | 3.71 |
ENSMUST00000115269.9
|
B3gat1
|
beta-1,3-glucuronyltransferase 1 (glucuronosyltransferase P) |
| chr5_+_146321757 | 3.70 |
ENSMUST00000016143.9
|
Wasf3
|
WASP family, member 3 |
| chr11_-_107685383 | 3.70 |
ENSMUST00000021066.4
|
Cacng4
|
calcium channel, voltage-dependent, gamma subunit 4 |
| chr19_-_36034740 | 3.64 |
ENSMUST00000164639.8
ENSMUST00000166074.2 ENSMUST00000099505.4 |
Htr7
|
5-hydroxytryptamine (serotonin) receptor 7 |
| chr7_+_120991489 | 3.62 |
ENSMUST00000084628.5
|
Hs3st2
|
heparan sulfate (glucosamine) 3-O-sulfotransferase 2 |
| chr7_+_29003363 | 3.61 |
ENSMUST00000108231.8
|
Dpf1
|
D4, zinc and double PHD fingers family 1 |
| chr11_+_7013422 | 3.61 |
ENSMUST00000020706.5
|
Adcy1
|
adenylate cyclase 1 |
| chr5_-_108515740 | 3.55 |
ENSMUST00000197216.3
|
Gm42517
|
predicted gene 42517 |
| chr7_+_141503719 | 3.53 |
ENSMUST00000105989.9
ENSMUST00000075528.12 ENSMUST00000174499.8 |
Brsk2
|
BR serine/threonine kinase 2 |
| chr14_-_29443792 | 3.50 |
ENSMUST00000022567.9
|
Cacna2d3
|
calcium channel, voltage-dependent, alpha2/delta subunit 3 |
| chr11_-_97909134 | 3.46 |
ENSMUST00000107561.9
|
Cacnb1
|
calcium channel, voltage-dependent, beta 1 subunit |
| chr9_-_108067552 | 3.46 |
ENSMUST00000035208.14
|
Bsn
|
bassoon |
| chr1_+_75351914 | 3.46 |
ENSMUST00000087122.12
|
Speg
|
SPEG complex locus |
| chr11_+_103061905 | 3.45 |
ENSMUST00000042286.12
ENSMUST00000218163.2 |
Fmnl1
|
formin-like 1 |
| chr15_+_99600149 | 3.43 |
ENSMUST00000229236.2
|
Smarcd1
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 1 |
| chr11_-_102338473 | 3.38 |
ENSMUST00000049057.5
|
Fam171a2
|
family with sequence similarity 171, member A2 |
| chr2_-_180596469 | 3.34 |
ENSMUST00000148905.8
ENSMUST00000103053.10 ENSMUST00000108873.9 |
Nkain4
|
Na+/K+ transporting ATPase interacting 4 |
| chr9_+_95441652 | 3.30 |
ENSMUST00000079597.7
|
Paqr9
|
progestin and adipoQ receptor family member IX |
| chr2_-_84717036 | 3.25 |
ENSMUST00000054514.6
ENSMUST00000151799.8 |
Rtn4rl2
|
reticulon 4 receptor-like 2 |
| chr1_+_91468409 | 3.23 |
ENSMUST00000027538.9
ENSMUST00000190484.7 ENSMUST00000186068.2 |
Asb1
|
ankyrin repeat and SOCS box-containing 1 |
| chr9_-_20638233 | 3.20 |
ENSMUST00000217198.2
|
Olfm2
|
olfactomedin 2 |
| chr11_-_97913420 | 3.19 |
ENSMUST00000103144.10
ENSMUST00000017552.13 ENSMUST00000092736.11 ENSMUST00000107562.2 |
Cacnb1
|
calcium channel, voltage-dependent, beta 1 subunit |
| chr7_+_29003382 | 3.19 |
ENSMUST00000049977.13
|
Dpf1
|
D4, zinc and double PHD fingers family 1 |
| chr15_+_99599978 | 3.16 |
ENSMUST00000023759.6
|
Smarcd1
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 1 |
| chr15_+_99122742 | 3.08 |
ENSMUST00000041415.5
|
Kcnh3
|
potassium voltage-gated channel, subfamily H (eag-related), member 3 |
| chr4_+_149671012 | 3.07 |
ENSMUST00000039144.7
|
Clstn1
|
calsyntenin 1 |
| chr1_+_91468266 | 3.07 |
ENSMUST00000086843.11
|
Asb1
|
ankyrin repeat and SOCS box-containing 1 |
| chr15_+_89407954 | 3.05 |
ENSMUST00000230807.2
|
Shank3
|
SH3 and multiple ankyrin repeat domains 3 |
| chr6_-_122463316 | 3.02 |
ENSMUST00000205114.2
|
Rimklb
|
ribosomal modification protein rimK-like family member B |
| chr18_-_35836161 | 3.01 |
ENSMUST00000025208.7
|
Dnajc18
|
DnaJ heat shock protein family (Hsp40) member C18 |
| chr15_-_91457383 | 3.00 |
ENSMUST00000109283.2
|
Slc2a13
|
solute carrier family 2 (facilitated glucose transporter), member 13 |
| chr11_+_77928736 | 2.97 |
ENSMUST00000072289.12
ENSMUST00000100784.9 ENSMUST00000073660.7 |
Flot2
|
flotillin 2 |
| chr4_+_152423075 | 2.94 |
ENSMUST00000030775.12
ENSMUST00000164662.8 |
Chd5
|
chromodomain helicase DNA binding protein 5 |
| chr4_-_148372384 | 2.93 |
ENSMUST00000047720.9
|
Disp3
|
dispatched RND transporter family member 3 |
| chr2_-_180596413 | 2.92 |
ENSMUST00000139929.8
|
Nkain4
|
Na+/K+ transporting ATPase interacting 4 |
| chr4_-_139974062 | 2.91 |
ENSMUST00000039331.9
|
Igsf21
|
immunoglobulin superfamily, member 21 |
| chr5_+_37185673 | 2.90 |
ENSMUST00000173836.8
|
Jakmip1
|
janus kinase and microtubule interacting protein 1 |
| chr10_+_78616304 | 2.88 |
ENSMUST00000005490.10
|
Slc1a6
|
solute carrier family 1 (high affinity aspartate/glutamate transporter), member 6 |
| chr1_+_172168764 | 2.88 |
ENSMUST00000056136.4
|
Kcnj10
|
potassium inwardly-rectifying channel, subfamily J, member 10 |
| chr1_+_91468796 | 2.87 |
ENSMUST00000188081.7
ENSMUST00000188879.2 |
Asb1
|
ankyrin repeat and SOCS box-containing 1 |
| chr11_-_106050724 | 2.85 |
ENSMUST00000064545.11
|
Limd2
|
LIM domain containing 2 |
| chr14_-_39194782 | 2.83 |
ENSMUST00000168810.9
ENSMUST00000173780.2 ENSMUST00000166968.9 |
Nrg3
|
neuregulin 3 |
| chr5_-_24556602 | 2.79 |
ENSMUST00000036092.10
|
Kcnh2
|
potassium voltage-gated channel, subfamily H (eag-related), member 2 |
| chr15_+_99600475 | 2.79 |
ENSMUST00000228984.2
ENSMUST00000229845.2 |
Smarcd1
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 1 |
| chr11_+_77384234 | 2.79 |
ENSMUST00000037285.10
ENSMUST00000100812.4 |
Git1
|
GIT ArfGAP 1 |
| chr10_+_13841819 | 2.77 |
ENSMUST00000187083.7
|
Hivep2
|
human immunodeficiency virus type I enhancer binding protein 2 |
| chr4_+_42950367 | 2.77 |
ENSMUST00000084662.12
|
Dnajb5
|
DnaJ heat shock protein family (Hsp40) member B5 |
| chr12_-_79054050 | 2.71 |
ENSMUST00000056660.13
ENSMUST00000174721.8 |
Tmem229b
|
transmembrane protein 229B |
| chr7_+_141503583 | 2.69 |
ENSMUST00000172652.8
|
Brsk2
|
BR serine/threonine kinase 2 |
| chr4_+_152423344 | 2.69 |
ENSMUST00000005175.5
|
Chd5
|
chromodomain helicase DNA binding protein 5 |
| chr15_+_74388044 | 2.67 |
ENSMUST00000042035.16
|
Adgrb1
|
adhesion G protein-coupled receptor B1 |
| chr15_+_87509413 | 2.67 |
ENSMUST00000068088.8
|
Tafa5
|
TAFA chemokine like family member 5 |
| chr4_-_32950812 | 2.65 |
ENSMUST00000084750.8
ENSMUST00000084748.9 |
Ankrd6
|
ankyrin repeat domain 6 |
| chr10_+_127216459 | 2.58 |
ENSMUST00000166820.8
|
R3hdm2
|
R3H domain containing 2 |
| chr2_+_156455583 | 2.55 |
ENSMUST00000109567.10
ENSMUST00000169464.9 |
Dlgap4
|
DLG associated protein 4 |
| chr9_+_58489523 | 2.53 |
ENSMUST00000177292.8
ENSMUST00000085651.12 ENSMUST00000176557.8 ENSMUST00000114121.11 ENSMUST00000177064.8 |
Nptn
|
neuroplastin |
| chr4_+_149670889 | 2.51 |
ENSMUST00000105691.8
|
Clstn1
|
calsyntenin 1 |
| chr7_+_3381434 | 2.50 |
ENSMUST00000092891.6
|
Cacng7
|
calcium channel, voltage-dependent, gamma subunit 7 |
| chr15_+_34837501 | 2.50 |
ENSMUST00000072868.5
|
Kcns2
|
K+ voltage-gated channel, subfamily S, 2 |
| chr11_-_77380492 | 2.44 |
ENSMUST00000037593.14
ENSMUST00000092892.10 |
Ankrd13b
|
ankyrin repeat domain 13b |
| chr11_-_97464866 | 2.43 |
ENSMUST00000207653.2
ENSMUST00000107593.8 |
Srcin1
|
SRC kinase signaling inhibitor 1 |
| chr16_-_18052937 | 2.37 |
ENSMUST00000076957.7
|
Zdhhc8
|
zinc finger, DHHC domain containing 8 |
| chr13_+_54651592 | 2.36 |
ENSMUST00000121401.8
ENSMUST00000118072.8 ENSMUST00000159721.2 |
Simc1
|
SUMO-interacting motifs containing 1 |
| chr8_-_88198992 | 2.36 |
ENSMUST00000169693.2
|
Cbln1
|
cerebellin 1 precursor protein |
| chr10_+_75152705 | 2.30 |
ENSMUST00000105420.3
|
Adora2a
|
adenosine A2a receptor |
| chr6_-_120470768 | 2.29 |
ENSMUST00000178687.2
|
Tmem121b
|
transmembrane protein 121B |
| chr15_+_34838195 | 2.27 |
ENSMUST00000228725.2
|
Kcns2
|
K+ voltage-gated channel, subfamily S, 2 |
| chr6_+_4747298 | 2.26 |
ENSMUST00000166678.2
ENSMUST00000176204.8 |
Peg10
|
paternally expressed 10 |
| chr7_-_45016138 | 2.26 |
ENSMUST00000211067.2
ENSMUST00000003961.16 |
Ppfia3
|
protein tyrosine phosphatase, receptor type, f polypeptide (PTPRF), interacting protein (liprin), alpha 3 |
| chr14_+_122712809 | 2.25 |
ENSMUST00000075888.6
|
Zic2
|
zinc finger protein of the cerebellum 2 |
| chr12_+_24622274 | 2.25 |
ENSMUST00000085553.13
|
Grhl1
|
grainyhead like transcription factor 1 |
| chr7_+_5054514 | 2.22 |
ENSMUST00000069324.7
|
Zfp580
|
zinc finger protein 580 |
| chr13_-_43457626 | 2.19 |
ENSMUST00000055341.7
|
Gfod1
|
glucose-fructose oxidoreductase domain containing 1 |
| chr12_-_107969853 | 2.13 |
ENSMUST00000066060.11
|
Bcl11b
|
B cell leukemia/lymphoma 11B |
| chr8_+_112264095 | 2.13 |
ENSMUST00000173726.8
ENSMUST00000174454.8 |
Znrf1
|
zinc and ring finger 1 |
| chr19_+_56710570 | 2.12 |
ENSMUST00000038949.6
|
Adrb1
|
adrenergic receptor, beta 1 |
| chr17_-_66826661 | 2.10 |
ENSMUST00000167962.2
ENSMUST00000070538.12 |
Rab12
|
RAB12, member RAS oncogene family |
| chrX_+_118836893 | 2.09 |
ENSMUST00000040961.3
ENSMUST00000113366.2 |
Pabpc5
|
poly(A) binding protein, cytoplasmic 5 |
| chr2_+_26518456 | 2.08 |
ENSMUST00000074240.4
|
Dipk1b
|
divergent protein kinase domain 1B |
| chr18_-_16942289 | 2.07 |
ENSMUST00000025166.14
|
Cdh2
|
cadherin 2 |
| chr11_+_94881861 | 2.05 |
ENSMUST00000038696.12
|
Ppp1r9b
|
protein phosphatase 1, regulatory subunit 9B |
| chr4_+_130297132 | 2.00 |
ENSMUST00000105993.4
|
Nkain1
|
Na+/K+ transporting ATPase interacting 1 |
| chr8_+_112263255 | 1.99 |
ENSMUST00000171182.8
ENSMUST00000168428.8 |
Znrf1
|
zinc and ring finger 1 |
| chr15_-_83609127 | 1.98 |
ENSMUST00000171496.9
ENSMUST00000043634.12 ENSMUST00000076060.12 ENSMUST00000016907.8 |
Scube1
|
signal peptide, CUB domain, EGF-like 1 |
| chr11_-_89193158 | 1.97 |
ENSMUST00000061728.5
|
Nog
|
noggin |
| chr7_+_135253659 | 1.96 |
ENSMUST00000209979.2
|
Ptpre
|
protein tyrosine phosphatase, receptor type, E |
| chr1_-_188740023 | 1.94 |
ENSMUST00000085678.8
|
Kctd3
|
potassium channel tetramerisation domain containing 3 |
| chr8_-_88199231 | 1.91 |
ENSMUST00000034076.16
|
Cbln1
|
cerebellin 1 precursor protein |
| chr15_-_99355623 | 1.88 |
ENSMUST00000023747.14
|
Nckap5l
|
NCK-associated protein 5-like |
| chr18_+_69478893 | 1.87 |
ENSMUST00000202354.4
|
Tcf4
|
transcription factor 4 |
| chr19_-_4356207 | 1.87 |
ENSMUST00000088737.11
|
Grk2
|
G protein-coupled receptor kinase 2 |
| chr18_-_25302064 | 1.86 |
ENSMUST00000115817.3
|
Tpgs2
|
tubulin polyglutamylase complex subunit 2 |
| chr6_+_4747356 | 1.83 |
ENSMUST00000176551.3
|
Peg10
|
paternally expressed 10 |
| chr11_-_97464755 | 1.83 |
ENSMUST00000126287.2
ENSMUST00000107590.9 |
Srcin1
|
SRC kinase signaling inhibitor 1 |
| chr4_+_152181155 | 1.82 |
ENSMUST00000105661.10
ENSMUST00000084115.4 |
Plekhg5
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 5 |
| chr8_+_95807814 | 1.82 |
ENSMUST00000034239.9
|
Katnb1
|
katanin p80 (WD40-containing) subunit B 1 |
| chr17_+_69144053 | 1.77 |
ENSMUST00000178545.3
|
Tmem200c
|
transmembrane protein 200C |
| chr3_-_90416757 | 1.77 |
ENSMUST00000107343.8
ENSMUST00000001043.14 ENSMUST00000107344.8 ENSMUST00000076639.11 ENSMUST00000107346.8 ENSMUST00000146740.8 ENSMUST00000107342.2 ENSMUST00000049937.13 |
Chtop
|
chromatin target of PRMT1 |
| chr19_-_4355983 | 1.77 |
ENSMUST00000025791.12
|
Grk2
|
G protein-coupled receptor kinase 2 |
| chr4_-_155430153 | 1.75 |
ENSMUST00000103178.11
|
Prkcz
|
protein kinase C, zeta |
| chr2_-_167032068 | 1.74 |
ENSMUST00000059826.10
|
Kcnb1
|
potassium voltage gated channel, Shab-related subfamily, member 1 |
| chr15_+_89383799 | 1.73 |
ENSMUST00000109309.9
|
Shank3
|
SH3 and multiple ankyrin repeat domains 3 |
| chr5_-_147337162 | 1.70 |
ENSMUST00000049324.13
|
Flt3
|
FMS-like tyrosine kinase 3 |
| chr2_+_116951855 | 1.69 |
ENSMUST00000028829.13
|
Spred1
|
sprouty protein with EVH-1 domain 1, related sequence |
| chr7_-_30850429 | 1.68 |
ENSMUST00000085636.13
ENSMUST00000001280.14 |
Gramd1a
|
GRAM domain containing 1A |
| chrX_+_98864627 | 1.66 |
ENSMUST00000096363.3
|
Tmem28
|
transmembrane protein 28 |
| chr7_+_26958150 | 1.65 |
ENSMUST00000079258.7
|
Numbl
|
numb-like |
| chr4_-_153567221 | 1.65 |
ENSMUST00000105646.3
|
Ajap1
|
adherens junction associated protein 1 |
| chr3_-_36744269 | 1.63 |
ENSMUST00000029271.5
|
Trpc3
|
transient receptor potential cation channel, subfamily C, member 3 |
| chrX_+_8758614 | 1.63 |
ENSMUST00000064196.5
|
B630019K06Rik
|
RIKEN cDNA B630019K06 gene |
| chr11_-_97466035 | 1.61 |
ENSMUST00000107596.9
ENSMUST00000238314.2 ENSMUST00000238597.2 ENSMUST00000238342.2 |
Srcin1
|
SRC kinase signaling inhibitor 1 |
| chr13_-_55635851 | 1.58 |
ENSMUST00000109921.9
ENSMUST00000109923.9 ENSMUST00000021950.15 |
Dbn1
|
drebrin 1 |
| chr17_-_10538253 | 1.58 |
ENSMUST00000233828.2
ENSMUST00000233645.2 ENSMUST00000042296.9 |
Qk
|
quaking, KH domain containing RNA binding |
| chr7_+_100355910 | 1.58 |
ENSMUST00000207875.2
ENSMUST00000208013.2 |
Fam168a
|
family with sequence similarity 168, member A |
| chr16_-_91485591 | 1.58 |
ENSMUST00000138560.2
ENSMUST00000117159.8 ENSMUST00000114031.8 ENSMUST00000023682.12 |
Donson
|
downstream neighbor of SON |
| chr10_-_127024641 | 1.57 |
ENSMUST00000218654.2
|
Arhgef25
|
Rho guanine nucleotide exchange factor (GEF) 25 |
| chr10_+_75152908 | 1.56 |
ENSMUST00000219044.2
|
Adora2a
|
adenosine A2a receptor |
| chr7_-_139162706 | 1.55 |
ENSMUST00000106095.3
|
Nkx6-2
|
NK6 homeobox 2 |
| chr2_+_30872291 | 1.54 |
ENSMUST00000102849.11
|
Usp20
|
ubiquitin specific peptidase 20 |
| chr2_+_30872362 | 1.49 |
ENSMUST00000061544.11
ENSMUST00000138161.8 ENSMUST00000142232.2 |
Usp20
|
ubiquitin specific peptidase 20 |
| chr15_-_93234681 | 1.49 |
ENSMUST00000080299.7
|
Yaf2
|
YY1 associated factor 2 |
| chr16_-_20245071 | 1.46 |
ENSMUST00000115547.9
ENSMUST00000096199.5 |
Abcc5
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 5 |
| chrX_-_166906307 | 1.44 |
ENSMUST00000112149.9
|
Frmpd4
|
FERM and PDZ domain containing 4 |
| chr2_-_29142965 | 1.42 |
ENSMUST00000155949.2
ENSMUST00000154682.8 ENSMUST00000028141.6 ENSMUST00000071201.5 |
6530402F18Rik
Ntng2
|
RIKEN cDNA 6530402F18 gene netrin G2 |
| chr15_-_85466009 | 1.41 |
ENSMUST00000023015.15
|
Wnt7b
|
wingless-type MMTV integration site family, member 7B |
| chr17_+_29833760 | 1.40 |
ENSMUST00000024817.15
ENSMUST00000162588.3 |
Rnf8
|
ring finger protein 8 |
| chr4_+_42949814 | 1.40 |
ENSMUST00000037872.10
ENSMUST00000098112.9 |
Dnajb5
|
DnaJ heat shock protein family (Hsp40) member B5 |
| chr8_-_123980825 | 1.37 |
ENSMUST00000118279.2
|
Vps9d1
|
VPS9 domain containing 1 |
| chr6_-_115971914 | 1.36 |
ENSMUST00000015511.15
|
Plxnd1
|
plexin D1 |
| chr7_+_18910340 | 1.35 |
ENSMUST00000117338.8
|
Eml2
|
echinoderm microtubule associated protein like 2 |
| chr16_-_20245138 | 1.35 |
ENSMUST00000079158.13
|
Abcc5
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 5 |
| chr15_-_79658608 | 1.34 |
ENSMUST00000229644.2
ENSMUST00000023055.8 |
Dnal4
|
dynein, axonemal, light chain 4 |
| chr13_-_105430932 | 1.33 |
ENSMUST00000224662.2
|
Rnf180
|
ring finger protein 180 |
| chr10_-_31321793 | 1.31 |
ENSMUST00000213639.2
ENSMUST00000215515.2 ENSMUST00000214644.2 ENSMUST00000213528.2 |
Tpd52l1
|
tumor protein D52-like 1 |
| chr15_+_68800546 | 1.31 |
ENSMUST00000230847.2
|
Khdrbs3
|
KH domain containing, RNA binding, signal transduction associated 3 |
| chr4_-_149858694 | 1.30 |
ENSMUST00000105686.3
|
Slc25a33
|
solute carrier family 25, member 33 |
| chr5_-_9211689 | 1.30 |
ENSMUST00000183973.8
ENSMUST00000184372.8 ENSMUST00000095017.11 ENSMUST00000071921.13 |
Dmtf1
|
cyclin D binding myb-like transcription factor 1 |
| chr4_+_137004793 | 1.29 |
ENSMUST00000045747.5
|
Wnt4
|
wingless-type MMTV integration site family, member 4 |
| chr13_-_120252259 | 1.29 |
ENSMUST00000223813.2
ENSMUST00000224946.2 |
Zfp131
|
zinc finger protein 131 |
| chr16_-_32065972 | 1.28 |
ENSMUST00000042732.6
|
Fbxo45
|
F-box protein 45 |
| chr8_+_85763780 | 1.27 |
ENSMUST00000211601.2
ENSMUST00000166592.2 |
Tnpo2
|
transportin 2 (importin 3, karyopherin beta 2b) |
| chr14_+_45457168 | 1.27 |
ENSMUST00000227086.2
ENSMUST00000147957.2 |
Gpr137c
|
G protein-coupled receptor 137C |
| chr11_-_119977609 | 1.26 |
ENSMUST00000106227.8
ENSMUST00000106229.8 ENSMUST00000180242.2 |
Cep131
|
centrosomal protein 131 |
| chr2_-_181335697 | 1.23 |
ENSMUST00000108779.8
ENSMUST00000108769.8 ENSMUST00000108772.8 |
Rgs19
|
regulator of G-protein signaling 19 |
| chr13_-_105430889 | 1.22 |
ENSMUST00000226044.2
|
Rnf180
|
ring finger protein 180 |
| chr14_-_106134253 | 1.21 |
ENSMUST00000022709.6
|
Spry2
|
sprouty RTK signaling antagonist 2 |
| chr11_-_86884507 | 1.21 |
ENSMUST00000018571.5
|
Ypel2
|
yippee like 2 |
| chr8_+_85763534 | 1.21 |
ENSMUST00000093360.12
|
Tnpo2
|
transportin 2 (importin 3, karyopherin beta 2b) |
| chrX_+_59591614 | 1.20 |
ENSMUST00000117865.3
|
Gm715
|
predicted gene 715 |
| chr15_+_97682210 | 1.19 |
ENSMUST00000117892.2
ENSMUST00000229084.2 |
Slc48a1
|
solute carrier family 48 (heme transporter), member 1 |
| chr9_+_121548469 | 1.17 |
ENSMUST00000182225.8
|
Nktr
|
natural killer tumor recognition sequence |
| chr4_+_43669266 | 1.17 |
ENSMUST00000107864.8
|
Tmem8b
|
transmembrane protein 8B |
| chr12_+_112978051 | 1.17 |
ENSMUST00000223502.2
ENSMUST00000084891.5 ENSMUST00000220541.2 |
Pacs2
|
phosphofurin acidic cluster sorting protein 2 |
| chr15_-_79658584 | 1.15 |
ENSMUST00000069877.12
|
Dnal4
|
dynein, axonemal, light chain 4 |
| chr4_+_44756553 | 1.15 |
ENSMUST00000107824.9
|
Zcchc7
|
zinc finger, CCHC domain containing 7 |
| chr14_+_75521783 | 1.15 |
ENSMUST00000022577.6
ENSMUST00000227049.2 |
Zc3h13
|
zinc finger CCCH type containing 13 |
| chr9_+_121548237 | 1.12 |
ENSMUST00000035112.13
ENSMUST00000182311.8 |
Nktr
|
natural killer tumor recognition sequence |
| chr19_+_4242064 | 1.11 |
ENSMUST00000046094.6
|
Ppp1ca
|
protein phosphatase 1 catalytic subunit alpha |
| chr1_+_106099482 | 1.11 |
ENSMUST00000061047.7
|
Phlpp1
|
PH domain and leucine rich repeat protein phosphatase 1 |
| chr14_-_20844074 | 1.11 |
ENSMUST00000080440.14
ENSMUST00000100837.11 ENSMUST00000071816.7 |
Camk2g
|
calcium/calmodulin-dependent protein kinase II gamma |
| chr8_-_123980908 | 1.10 |
ENSMUST00000122363.8
|
Vps9d1
|
VPS9 domain containing 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.2 | 9.5 | GO:0097534 | lymphoid lineage cell migration(GO:0097534) lymphoid lineage cell migration into thymus(GO:0097535) |
| 2.0 | 6.0 | GO:0045212 | neurotransmitter receptor biosynthetic process(GO:0045212) |
| 1.5 | 4.4 | GO:0033693 | neurofilament bundle assembly(GO:0033693) |
| 1.5 | 5.8 | GO:0021586 | pons maturation(GO:0021586) |
| 1.2 | 3.6 | GO:0051385 | response to mineralocorticoid(GO:0051385) |
| 1.2 | 3.5 | GO:0099526 | presynaptic signal transduction(GO:0098928) presynapse to nucleus signaling pathway(GO:0099526) |
| 1.1 | 4.6 | GO:2001025 | positive regulation of response to drug(GO:2001025) |
| 1.0 | 5.9 | GO:0000160 | phosphorelay signal transduction system(GO:0000160) |
| 0.9 | 10.9 | GO:0023041 | neuronal signal transduction(GO:0023041) |
| 0.8 | 4.1 | GO:0086046 | membrane depolarization during SA node cell action potential(GO:0086046) |
| 0.8 | 18.5 | GO:0070262 | peptidyl-serine dephosphorylation(GO:0070262) |
| 0.8 | 4.8 | GO:0021773 | striatal medium spiny neuron differentiation(GO:0021773) |
| 0.8 | 3.9 | GO:0014057 | positive regulation of acetylcholine secretion, neurotransmission(GO:0014057) |
| 0.7 | 2.9 | GO:0051935 | amino acid neurotransmitter reuptake(GO:0051933) glutamate reuptake(GO:0051935) |
| 0.7 | 2.1 | GO:1900135 | positive regulation of renin secretion into blood stream(GO:1900135) |
| 0.7 | 2.1 | GO:0071315 | cellular response to morphine(GO:0071315) regulation of opioid receptor signaling pathway(GO:2000474) |
| 0.7 | 7.9 | GO:0035093 | spermatogenesis, exchange of chromosomal proteins(GO:0035093) |
| 0.7 | 2.0 | GO:0060825 | regulation of cytokine activity(GO:0060300) fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:0060825) negative regulation of epithelial to mesenchymal transition involved in endocardial cushion formation(GO:1905006) regulation of fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:2000313) |
| 0.6 | 2.5 | GO:1902683 | regulation of receptor localization to synapse(GO:1902683) |
| 0.6 | 1.7 | GO:0036145 | dendritic cell homeostasis(GO:0036145) |
| 0.6 | 2.3 | GO:1900224 | positive regulation of nodal signaling pathway involved in determination of lateral mesoderm left/right asymmetry(GO:1900224) |
| 0.6 | 2.2 | GO:1905051 | regulation of base-excision repair(GO:1905051) positive regulation of base-excision repair(GO:1905053) |
| 0.5 | 1.6 | GO:0030860 | regulation of polarized epithelial cell differentiation(GO:0030860) |
| 0.5 | 1.6 | GO:1903244 | positive regulation of cardiac muscle adaptation(GO:0010615) positive regulation of cardiac muscle hypertrophy in response to stress(GO:1903244) |
| 0.5 | 3.7 | GO:1902897 | regulation of postsynaptic density protein 95 clustering(GO:1902897) |
| 0.5 | 3.0 | GO:0044860 | protein localization to plasma membrane raft(GO:0044860) regulation of establishment of T cell polarity(GO:1903903) |
| 0.5 | 2.0 | GO:0007185 | transmembrane receptor protein tyrosine phosphatase signaling pathway(GO:0007185) |
| 0.5 | 2.8 | GO:0060596 | mammary placode formation(GO:0060596) |
| 0.5 | 8.6 | GO:0006337 | nucleosome disassembly(GO:0006337) |
| 0.5 | 3.6 | GO:0007217 | tachykinin receptor signaling pathway(GO:0007217) |
| 0.5 | 1.8 | GO:0031117 | positive regulation of microtubule depolymerization(GO:0031117) |
| 0.4 | 1.3 | GO:2000019 | positive regulation of dermatome development(GO:0061184) renal vesicle induction(GO:0072034) negative regulation of male gonad development(GO:2000019) |
| 0.4 | 4.3 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
| 0.4 | 6.4 | GO:1900454 | positive regulation of long term synaptic depression(GO:1900454) |
| 0.4 | 2.9 | GO:1903802 | L-glutamate import across plasma membrane(GO:0098712) L-glutamate(1-) import into cell(GO:1903802) L-glutamate import into cell(GO:1990123) |
| 0.4 | 1.4 | GO:0060535 | trachea cartilage morphogenesis(GO:0060535) |
| 0.3 | 8.8 | GO:0050650 | chondroitin sulfate proteoglycan biosynthetic process(GO:0050650) |
| 0.3 | 1.2 | GO:0060437 | lung growth(GO:0060437) |
| 0.3 | 3.2 | GO:0006610 | ribosomal protein import into nucleus(GO:0006610) |
| 0.3 | 1.4 | GO:0051835 | positive regulation of synapse structural plasticity(GO:0051835) |
| 0.3 | 9.2 | GO:0030539 | male genitalia development(GO:0030539) |
| 0.3 | 1.6 | GO:1990314 | cellular response to insulin-like growth factor stimulus(GO:1990314) |
| 0.3 | 11.1 | GO:2000311 | regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000311) |
| 0.3 | 1.5 | GO:0021912 | regulation of transcription from RNA polymerase II promoter involved in spinal cord motor neuron fate specification(GO:0021912) |
| 0.2 | 1.7 | GO:2000664 | positive regulation of interleukin-5 secretion(GO:2000664) positive regulation of interleukin-13 secretion(GO:2000667) |
| 0.2 | 1.7 | GO:0061084 | negative regulation of protein refolding(GO:0061084) |
| 0.2 | 1.2 | GO:0036324 | vascular endothelial growth factor receptor-2 signaling pathway(GO:0036324) |
| 0.2 | 7.8 | GO:0095500 | acetylcholine receptor signaling pathway(GO:0095500) signal transduction involved in cellular response to ammonium ion(GO:1903831) response to acetylcholine(GO:1905144) cellular response to acetylcholine(GO:1905145) |
| 0.2 | 0.7 | GO:0061537 | glycine secretion(GO:0061536) glycine secretion, neurotransmission(GO:0061537) |
| 0.2 | 2.5 | GO:2000580 | regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000580) positive regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000582) |
| 0.2 | 1.4 | GO:0010968 | regulation of microtubule nucleation(GO:0010968) |
| 0.2 | 0.9 | GO:0015766 | disaccharide transport(GO:0015766) sucrose transport(GO:0015770) oligosaccharide transport(GO:0015772) |
| 0.2 | 4.0 | GO:0033601 | positive regulation of mammary gland epithelial cell proliferation(GO:0033601) |
| 0.2 | 24.0 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.2 | 11.5 | GO:0031572 | G2 DNA damage checkpoint(GO:0031572) |
| 0.2 | 1.3 | GO:0035735 | intraciliary transport involved in cilium morphogenesis(GO:0035735) |
| 0.2 | 0.9 | GO:0051562 | negative regulation of mitochondrial calcium ion concentration(GO:0051562) |
| 0.2 | 0.7 | GO:0010510 | regulation of acetyl-CoA biosynthetic process from pyruvate(GO:0010510) |
| 0.2 | 2.8 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.2 | 0.7 | GO:0035854 | eosinophil fate commitment(GO:0035854) |
| 0.2 | 1.2 | GO:0015886 | heme transport(GO:0015886) |
| 0.2 | 1.8 | GO:0000042 | protein targeting to Golgi(GO:0000042) |
| 0.2 | 1.3 | GO:0030920 | N-terminal peptidyl-serine acetylation(GO:0017198) N-terminal peptidyl-glutamic acid acetylation(GO:0018002) peptidyl-serine acetylation(GO:0030920) |
| 0.2 | 3.0 | GO:0035428 | hexose transmembrane transport(GO:0035428) |
| 0.2 | 1.1 | GO:1903527 | positive regulation of membrane tubulation(GO:1903527) |
| 0.2 | 1.4 | GO:0060666 | dichotomous subdivision of terminal units involved in salivary gland branching(GO:0060666) |
| 0.2 | 3.6 | GO:0007190 | activation of adenylate cyclase activity(GO:0007190) |
| 0.1 | 0.7 | GO:0042631 | cellular response to water deprivation(GO:0042631) |
| 0.1 | 1.1 | GO:0090038 | negative regulation of protein kinase C signaling(GO:0090038) |
| 0.1 | 0.8 | GO:0071442 | positive regulation of histone H3-K14 acetylation(GO:0071442) |
| 0.1 | 2.7 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.1 | 1.3 | GO:0021960 | anterior commissure morphogenesis(GO:0021960) |
| 0.1 | 1.7 | GO:0098703 | calcium ion import across plasma membrane(GO:0098703) calcium ion import into cell(GO:1990035) |
| 0.1 | 0.8 | GO:0071955 | recycling endosome to Golgi transport(GO:0071955) |
| 0.1 | 2.6 | GO:0042415 | norepinephrine metabolic process(GO:0042415) |
| 0.1 | 3.3 | GO:0022038 | corpus callosum development(GO:0022038) |
| 0.1 | 4.8 | GO:1902259 | regulation of delayed rectifier potassium channel activity(GO:1902259) |
| 0.1 | 0.7 | GO:0001922 | B-1 B cell homeostasis(GO:0001922) |
| 0.1 | 2.6 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.1 | 1.1 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.1 | 5.9 | GO:0061099 | negative regulation of protein tyrosine kinase activity(GO:0061099) |
| 0.1 | 2.3 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.1 | 4.2 | GO:0031643 | positive regulation of myelination(GO:0031643) |
| 0.1 | 0.4 | GO:0048619 | embryonic genitalia morphogenesis(GO:0030538) embryonic hindgut morphogenesis(GO:0048619) |
| 0.1 | 2.3 | GO:0001771 | immunological synapse formation(GO:0001771) |
| 0.1 | 2.6 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.1 | 6.7 | GO:1901385 | regulation of voltage-gated calcium channel activity(GO:1901385) |
| 0.1 | 0.5 | GO:0070966 | nuclear-transcribed mRNA catabolic process, no-go decay(GO:0070966) |
| 0.1 | 10.8 | GO:0061178 | regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0061178) |
| 0.1 | 1.4 | GO:0002934 | desmosome organization(GO:0002934) |
| 0.1 | 0.4 | GO:2001287 | negative regulation of caveolin-mediated endocytosis(GO:2001287) |
| 0.1 | 0.7 | GO:0018344 | protein geranylgeranylation(GO:0018344) |
| 0.1 | 0.3 | GO:0031339 | negative regulation of vesicle fusion(GO:0031339) |
| 0.1 | 1.6 | GO:0061158 | 3'-UTR-mediated mRNA destabilization(GO:0061158) |
| 0.1 | 8.0 | GO:0070936 | protein K48-linked ubiquitination(GO:0070936) |
| 0.1 | 0.8 | GO:0070197 | meiotic telomere tethering at nuclear periphery(GO:0044821) meiotic attachment of telomere to nuclear envelope(GO:0070197) chromosome attachment to the nuclear envelope(GO:0097240) |
| 0.1 | 1.8 | GO:0035767 | endothelial cell chemotaxis(GO:0035767) |
| 0.1 | 1.1 | GO:0005981 | regulation of glycogen catabolic process(GO:0005981) |
| 0.1 | 2.3 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.1 | 0.3 | GO:0060024 | rhythmic synaptic transmission(GO:0060024) |
| 0.1 | 2.4 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.1 | 0.5 | GO:0045196 | establishment or maintenance of neuroblast polarity(GO:0045196) establishment of neuroblast polarity(GO:0045200) |
| 0.1 | 1.2 | GO:0021670 | lateral ventricle development(GO:0021670) |
| 0.1 | 1.2 | GO:0034497 | protein localization to pre-autophagosomal structure(GO:0034497) |
| 0.1 | 0.9 | GO:2000009 | negative regulation of protein localization to cell surface(GO:2000009) |
| 0.1 | 1.1 | GO:1901897 | regulation of relaxation of cardiac muscle(GO:1901897) |
| 0.1 | 3.5 | GO:0051865 | protein autoubiquitination(GO:0051865) |
| 0.1 | 1.5 | GO:0042118 | endothelial cell activation(GO:0042118) |
| 0.1 | 0.9 | GO:0090141 | positive regulation of mitochondrial fission(GO:0090141) |
| 0.1 | 0.6 | GO:0006398 | mRNA 3'-end processing by stem-loop binding and cleavage(GO:0006398) |
| 0.1 | 5.2 | GO:0016079 | synaptic vesicle exocytosis(GO:0016079) |
| 0.1 | 0.8 | GO:0006336 | DNA replication-independent nucleosome assembly(GO:0006336) |
| 0.0 | 0.6 | GO:0051152 | positive regulation of smooth muscle cell differentiation(GO:0051152) |
| 0.0 | 1.6 | GO:0033260 | nuclear DNA replication(GO:0033260) |
| 0.0 | 0.8 | GO:0044331 | cell-cell adhesion mediated by cadherin(GO:0044331) |
| 0.0 | 0.2 | GO:0070602 | regulation of centromeric sister chromatid cohesion(GO:0070602) |
| 0.0 | 0.8 | GO:0071880 | adenylate cyclase-activating adrenergic receptor signaling pathway(GO:0071880) |
| 0.0 | 0.6 | GO:0001574 | ganglioside biosynthetic process(GO:0001574) |
| 0.0 | 2.2 | GO:0032757 | positive regulation of interleukin-8 production(GO:0032757) |
| 0.0 | 2.9 | GO:0007019 | microtubule depolymerization(GO:0007019) |
| 0.0 | 0.9 | GO:0021895 | cerebral cortex neuron differentiation(GO:0021895) |
| 0.0 | 2.1 | GO:0008333 | endosome to lysosome transport(GO:0008333) |
| 0.0 | 0.1 | GO:0072356 | chromosome passenger complex localization to kinetochore(GO:0072356) |
| 0.0 | 0.5 | GO:0031468 | nuclear envelope reassembly(GO:0031468) |
| 0.0 | 0.9 | GO:0032703 | negative regulation of interleukin-2 production(GO:0032703) |
| 0.0 | 0.5 | GO:0001672 | regulation of chromatin assembly or disassembly(GO:0001672) |
| 0.0 | 0.4 | GO:0006370 | 7-methylguanosine mRNA capping(GO:0006370) |
| 0.0 | 5.1 | GO:0019236 | response to pheromone(GO:0019236) |
| 0.0 | 0.2 | GO:0072344 | rescue of stalled ribosome(GO:0072344) |
| 0.0 | 4.2 | GO:0002028 | regulation of sodium ion transport(GO:0002028) |
| 0.0 | 1.1 | GO:0021511 | spinal cord patterning(GO:0021511) |
| 0.0 | 0.5 | GO:0033148 | positive regulation of intracellular estrogen receptor signaling pathway(GO:0033148) |
| 0.0 | 0.3 | GO:0051967 | negative regulation of synaptic transmission, glutamatergic(GO:0051967) |
| 0.0 | 1.0 | GO:0035024 | negative regulation of Rho protein signal transduction(GO:0035024) |
| 0.0 | 0.5 | GO:0042533 | tumor necrosis factor biosynthetic process(GO:0042533) regulation of tumor necrosis factor biosynthetic process(GO:0042534) |
| 0.0 | 0.9 | GO:0090382 | phagosome maturation(GO:0090382) |
| 0.0 | 0.3 | GO:0000963 | mitochondrial RNA processing(GO:0000963) |
| 0.0 | 1.0 | GO:0071539 | protein localization to centrosome(GO:0071539) |
| 0.0 | 0.2 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 0.0 | 1.8 | GO:2000179 | positive regulation of neural precursor cell proliferation(GO:2000179) |
| 0.0 | 0.7 | GO:0042711 | maternal behavior(GO:0042711) |
| 0.0 | 0.9 | GO:0071353 | cellular response to interleukin-4(GO:0071353) |
| 0.0 | 0.9 | GO:0034142 | toll-like receptor 4 signaling pathway(GO:0034142) |
| 0.0 | 0.1 | GO:2000182 | regulation of progesterone biosynthetic process(GO:2000182) |
| 0.0 | 0.3 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 0.0 | 0.2 | GO:0006283 | transcription-coupled nucleotide-excision repair(GO:0006283) |
| 0.0 | 0.2 | GO:0060179 | male mating behavior(GO:0060179) |
| 0.0 | 0.5 | GO:0040015 | negative regulation of multicellular organism growth(GO:0040015) |
| 0.0 | 0.6 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.0 | 0.3 | GO:0044804 | nucleophagy(GO:0044804) |
| 0.0 | 1.4 | GO:0045669 | positive regulation of osteoblast differentiation(GO:0045669) |
| 0.0 | 0.8 | GO:0035176 | social behavior(GO:0035176) intraspecies interaction between organisms(GO:0051703) |
| 0.0 | 3.3 | GO:0006457 | protein folding(GO:0006457) |
| 0.0 | 0.2 | GO:0043482 | pigment accumulation(GO:0043476) cellular pigment accumulation(GO:0043482) |
| 0.0 | 0.1 | GO:1903033 | regulation of microtubule plus-end binding(GO:1903031) positive regulation of microtubule plus-end binding(GO:1903033) |
| 0.0 | 0.8 | GO:0061512 | protein localization to cilium(GO:0061512) |
| 0.0 | 2.0 | GO:0035023 | regulation of Rho protein signal transduction(GO:0035023) |
| 0.0 | 0.3 | GO:0043968 | histone H2A acetylation(GO:0043968) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 5.8 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 1.2 | 5.9 | GO:0070032 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin I complex(GO:0070032) |
| 1.2 | 3.5 | GO:1990257 | piccolo-bassoon transport vesicle(GO:1990257) |
| 0.6 | 2.8 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.5 | 16.2 | GO:0071565 | nBAF complex(GO:0071565) |
| 0.5 | 18.1 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.5 | 10.4 | GO:0032279 | asymmetric synapse(GO:0032279) |
| 0.5 | 20.3 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.4 | 6.0 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.3 | 4.4 | GO:0005883 | neurofilament(GO:0005883) |
| 0.3 | 1.2 | GO:1990032 | climbing fiber(GO:0044301) parallel fiber(GO:1990032) |
| 0.3 | 0.9 | GO:0060187 | cell pole(GO:0060187) |
| 0.3 | 4.6 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.3 | 2.1 | GO:0032591 | dendritic spine membrane(GO:0032591) dendritic spine neck(GO:0044326) |
| 0.3 | 0.8 | GO:0098592 | cytoplasmic side of apical plasma membrane(GO:0098592) |
| 0.2 | 0.7 | GO:0005967 | mitochondrial pyruvate dehydrogenase complex(GO:0005967) |
| 0.2 | 0.7 | GO:1990730 | VCP-NSFL1C complex(GO:1990730) |
| 0.2 | 3.0 | GO:0001931 | uropod(GO:0001931) cell trailing edge(GO:0031254) |
| 0.2 | 5.2 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.2 | 11.1 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.2 | 4.4 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.2 | 1.6 | GO:0044214 | spanning component of plasma membrane(GO:0044214) spanning component of membrane(GO:0089717) |
| 0.2 | 1.7 | GO:0045179 | apical cortex(GO:0045179) |
| 0.1 | 1.1 | GO:0072357 | PTW/PP1 phosphatase complex(GO:0072357) |
| 0.1 | 0.7 | GO:0005968 | Rab-protein geranylgeranyltransferase complex(GO:0005968) |
| 0.1 | 6.0 | GO:0032809 | neuronal cell body membrane(GO:0032809) |
| 0.1 | 2.9 | GO:0097449 | astrocyte projection(GO:0097449) |
| 0.1 | 0.8 | GO:0070381 | endosome to plasma membrane transport vesicle(GO:0070381) |
| 0.1 | 1.3 | GO:0031415 | NatA complex(GO:0031415) |
| 0.1 | 0.9 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
| 0.1 | 1.1 | GO:0030137 | COPI-coated vesicle(GO:0030137) |
| 0.1 | 1.1 | GO:0002177 | manchette(GO:0002177) |
| 0.1 | 2.1 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.1 | 3.6 | GO:0005790 | smooth endoplasmic reticulum(GO:0005790) |
| 0.1 | 4.8 | GO:0060170 | ciliary membrane(GO:0060170) |
| 0.1 | 0.5 | GO:0097550 | transcriptional preinitiation complex(GO:0097550) |
| 0.1 | 3.6 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.1 | 4.1 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.1 | 1.9 | GO:0005849 | mRNA cleavage factor complex(GO:0005849) |
| 0.1 | 4.3 | GO:0045335 | phagocytic vesicle(GO:0045335) |
| 0.1 | 2.1 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.1 | 2.8 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.1 | 2.3 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.1 | 21.3 | GO:0014069 | postsynaptic density(GO:0014069) |
| 0.1 | 13.0 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.1 | 8.3 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 0.8 | GO:0000124 | SAGA complex(GO:0000124) |
| 0.0 | 0.8 | GO:0005639 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) |
| 0.0 | 17.1 | GO:0000139 | Golgi membrane(GO:0000139) |
| 0.0 | 0.9 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.0 | 2.5 | GO:0030286 | dynein complex(GO:0030286) |
| 0.0 | 7.3 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 3.2 | GO:0030136 | clathrin-coated vesicle(GO:0030136) |
| 0.0 | 3.5 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) |
| 0.0 | 0.3 | GO:0035867 | alphav-beta3 integrin-IGF-1-IGF1R complex(GO:0035867) |
| 0.0 | 0.1 | GO:0031372 | UBC13-MMS2 complex(GO:0031372) |
| 0.0 | 1.0 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.0 | 5.4 | GO:0045111 | intermediate filament cytoskeleton(GO:0045111) |
| 0.0 | 9.9 | GO:0097060 | synaptic membrane(GO:0097060) |
| 0.0 | 15.5 | GO:0045202 | synapse(GO:0045202) |
| 0.0 | 0.4 | GO:0051233 | spindle midzone(GO:0051233) |
| 0.0 | 2.4 | GO:0030426 | growth cone(GO:0030426) |
| 0.0 | 1.5 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.0 | 2.5 | GO:0005769 | early endosome(GO:0005769) |
| 0.0 | 1.2 | GO:0055037 | recycling endosome(GO:0055037) |
| 0.0 | 0.1 | GO:0070847 | core mediator complex(GO:0070847) |
| 0.0 | 1.2 | GO:0010008 | endosome membrane(GO:0010008) |
| 0.0 | 0.3 | GO:0035267 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.0 | 0.4 | GO:0005719 | nuclear euchromatin(GO:0005719) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.0 | 6.0 | GO:0003990 | acetylcholinesterase activity(GO:0003990) |
| 1.6 | 9.4 | GO:0015018 | galactosylgalactosylxylosylprotein 3-beta-glucuronosyltransferase activity(GO:0015018) |
| 1.4 | 5.6 | GO:0061628 | H3K27me3 modified histone binding(GO:0061628) |
| 1.4 | 4.1 | GO:0086059 | voltage-gated calcium channel activity involved SA node cell action potential(GO:0086059) |
| 1.2 | 3.6 | GO:0031755 | endothelial differentiation G-protein coupled receptor binding(GO:0031753) Edg-2 lysophosphatidic acid receptor binding(GO:0031755) |
| 1.1 | 21.5 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 1.1 | 4.2 | GO:0030144 | alpha-1,6-mannosylglycoprotein 6-beta-N-acetylglucosaminyltransferase activity(GO:0030144) |
| 1.0 | 5.9 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.8 | 8.0 | GO:0030550 | acetylcholine receptor inhibitor activity(GO:0030550) |
| 0.8 | 3.0 | GO:0072591 | citrate-L-glutamate ligase activity(GO:0072591) |
| 0.7 | 4.8 | GO:0030160 | GKAP/Homer scaffold activity(GO:0030160) |
| 0.6 | 1.7 | GO:0015275 | stretch-activated, cation-selective, calcium channel activity(GO:0015275) |
| 0.5 | 4.3 | GO:0005042 | netrin receptor activity(GO:0005042) |
| 0.5 | 2.1 | GO:0004939 | beta-adrenergic receptor activity(GO:0004939) alpha-2A adrenergic receptor binding(GO:0031694) |
| 0.5 | 3.6 | GO:0008294 | calcium- and calmodulin-responsive adenylate cyclase activity(GO:0008294) |
| 0.4 | 2.9 | GO:0015272 | ATP-activated inward rectifier potassium channel activity(GO:0015272) |
| 0.4 | 1.4 | GO:1902379 | chemoattractant activity involved in axon guidance(GO:1902379) |
| 0.3 | 1.7 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.3 | 5.3 | GO:0035014 | phosphatidylinositol 3-kinase regulator activity(GO:0035014) |
| 0.3 | 3.9 | GO:0001609 | G-protein coupled adenosine receptor activity(GO:0001609) |
| 0.3 | 2.4 | GO:0032184 | SUMO polymer binding(GO:0032184) |
| 0.3 | 1.7 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 0.3 | 6.7 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.3 | 3.6 | GO:0051378 | serotonin binding(GO:0051378) |
| 0.2 | 10.0 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.2 | 0.9 | GO:0047389 | glycerophosphocholine phosphodiesterase activity(GO:0047389) |
| 0.2 | 0.9 | GO:0035276 | calcium-independent protein kinase C activity(GO:0004699) ethanol binding(GO:0035276) |
| 0.2 | 4.4 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.2 | 1.1 | GO:0043559 | insulin binding(GO:0043559) |
| 0.2 | 0.9 | GO:0015157 | sucrose:proton symporter activity(GO:0008506) sucrose transmembrane transporter activity(GO:0008515) disaccharide transmembrane transporter activity(GO:0015154) oligosaccharide transmembrane transporter activity(GO:0015157) |
| 0.2 | 15.3 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.2 | 1.2 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.2 | 16.3 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.2 | 2.1 | GO:0031749 | D2 dopamine receptor binding(GO:0031749) |
| 0.2 | 3.1 | GO:0005522 | profilin binding(GO:0005522) |
| 0.2 | 2.9 | GO:0005314 | high-affinity glutamate transmembrane transporter activity(GO:0005314) |
| 0.2 | 5.9 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.2 | 0.7 | GO:0004740 | pyruvate dehydrogenase (acetyl-transferring) kinase activity(GO:0004740) |
| 0.2 | 1.4 | GO:0030375 | thyroid hormone receptor coactivator activity(GO:0030375) |
| 0.2 | 1.3 | GO:1990190 | peptide-glutamate-N-acetyltransferase activity(GO:1990190) |
| 0.2 | 3.3 | GO:0034483 | heparan sulfate sulfotransferase activity(GO:0034483) |
| 0.2 | 2.3 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.1 | 1.2 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 0.1 | 0.9 | GO:0043533 | inositol 1,3,4,5 tetrakisphosphate binding(GO:0043533) |
| 0.1 | 3.0 | GO:0005351 | sugar:proton symporter activity(GO:0005351) |
| 0.1 | 2.1 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.1 | 2.8 | GO:0045499 | chemorepellent activity(GO:0045499) |
| 0.1 | 5.9 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.1 | 3.0 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.1 | 0.7 | GO:0008269 | JAK pathway signal transduction adaptor activity(GO:0008269) |
| 0.1 | 2.9 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.1 | 1.3 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.1 | 0.8 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.1 | 1.4 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.1 | 1.5 | GO:0001087 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.1 | 1.7 | GO:0015271 | outward rectifier potassium channel activity(GO:0015271) |
| 0.1 | 1.7 | GO:0004697 | protein kinase C activity(GO:0004697) |
| 0.1 | 2.5 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.1 | 3.2 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.1 | 0.5 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.1 | 7.1 | GO:0035254 | glutamate receptor binding(GO:0035254) |
| 0.1 | 2.4 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.1 | 8.2 | GO:0032947 | protein complex scaffold(GO:0032947) |
| 0.1 | 0.7 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.1 | 4.4 | GO:0019003 | GDP binding(GO:0019003) |
| 0.1 | 0.5 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.1 | 0.7 | GO:0005283 | sodium:amino acid symporter activity(GO:0005283) |
| 0.1 | 0.6 | GO:0035374 | chondroitin sulfate binding(GO:0035374) |
| 0.1 | 0.8 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.1 | 0.9 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.0 | 1.3 | GO:0015215 | nucleotide transmembrane transporter activity(GO:0015215) |
| 0.0 | 0.9 | GO:0031078 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.0 | 2.2 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 5.9 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.0 | 2.3 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.0 | 0.8 | GO:0005234 | extracellular-glutamate-gated ion channel activity(GO:0005234) |
| 0.0 | 0.9 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.0 | 1.9 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.0 | 1.7 | GO:0043548 | phosphatidylinositol 3-kinase binding(GO:0043548) |
| 0.0 | 0.8 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) phospholipase C activity(GO:0004629) |
| 0.0 | 3.3 | GO:0005249 | voltage-gated potassium channel activity(GO:0005249) |
| 0.0 | 0.1 | GO:0030297 | transmembrane receptor protein tyrosine kinase activator activity(GO:0030297) |
| 0.0 | 2.5 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.0 | 0.7 | GO:0070742 | C2H2 zinc finger domain binding(GO:0070742) |
| 0.0 | 4.3 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.0 | 0.1 | GO:0032296 | ribonuclease III activity(GO:0004525) double-stranded RNA-specific ribonuclease activity(GO:0032296) |
| 0.0 | 1.1 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.0 | 3.3 | GO:0005496 | steroid binding(GO:0005496) |
| 0.0 | 0.8 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.4 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.0 | 0.8 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.0 | 11.1 | GO:0004930 | G-protein coupled receptor activity(GO:0004930) |
| 0.0 | 0.4 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.0 | 0.2 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
| 0.0 | 0.2 | GO:0070411 | I-SMAD binding(GO:0070411) |
| 0.0 | 0.9 | GO:0031491 | nucleosome binding(GO:0031491) |
| 0.0 | 3.8 | GO:0061630 | ubiquitin protein ligase activity(GO:0061630) |
| 0.0 | 0.3 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.0 | 0.6 | GO:0030295 | protein kinase activator activity(GO:0030295) |
| 0.0 | 0.2 | GO:0004791 | thioredoxin-disulfide reductase activity(GO:0004791) |
| 0.0 | 0.4 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 21.0 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.2 | 5.2 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.1 | 5.9 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.1 | 2.8 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.1 | 2.1 | PID IGF1 PATHWAY | IGF1 pathway |
| 0.1 | 5.2 | PID PI3K PLC TRK PATHWAY | Trk receptor signaling mediated by PI3K and PLC-gamma |
| 0.1 | 4.3 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.1 | 6.8 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.1 | 3.2 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.1 | 2.6 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.1 | 2.4 | PID ATM PATHWAY | ATM pathway |
| 0.1 | 2.9 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.1 | 3.0 | SIG PIP3 SIGNALING IN B LYMPHOCYTES | Genes related to PIP3 signaling in B lymphocytes |
| 0.1 | 2.4 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.0 | 1.3 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.0 | 2.4 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.0 | 1.4 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.0 | 2.5 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.0 | 1.1 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.0 | 3.9 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 1.0 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.0 | 1.2 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 1.4 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 1.6 | PID HDAC CLASSII PATHWAY | Signaling events mediated by HDAC Class II |
| 0.0 | 1.2 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.0 | 1.0 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.0 | 0.3 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.0 | 1.2 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 0.3 | PID PI3KCI AKT PATHWAY | Class I PI3K signaling events mediated by Akt |
| 0.0 | 0.8 | PID CD8 TCR PATHWAY | TCR signaling in naïve CD8+ T cells |
| 0.0 | 0.8 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.0 | 0.7 | PID PDGFRB PATHWAY | PDGFR-beta signaling pathway |
| 0.0 | 1.2 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 1.3 | PID P73PATHWAY | p73 transcription factor network |
| 0.0 | 0.7 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.0 | 0.3 | PID ALK1 PATHWAY | ALK1 signaling events |
| 0.0 | 0.3 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 3.6 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.3 | 9.4 | REACTOME A TETRASACCHARIDE LINKER SEQUENCE IS REQUIRED FOR GAG SYNTHESIS | Genes involved in A tetrasaccharide linker sequence is required for GAG synthesis |
| 0.3 | 4.3 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.3 | 4.4 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.2 | 11.3 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.2 | 6.0 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.2 | 11.1 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.2 | 1.6 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.2 | 2.8 | REACTOME HYALURONAN METABOLISM | Genes involved in Hyaluronan metabolism |
| 0.1 | 3.9 | REACTOME NUCLEOTIDE LIKE PURINERGIC RECEPTORS | Genes involved in Nucleotide-like (purinergic) receptors |
| 0.1 | 2.5 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.1 | 2.9 | REACTOME GLUTAMATE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Glutamate Neurotransmitter Release Cycle |
| 0.1 | 2.9 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.1 | 1.9 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.1 | 2.1 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.1 | 1.1 | REACTOME NEF MEDIATED DOWNREGULATION OF MHC CLASS I COMPLEX CELL SURFACE EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.1 | 0.7 | REACTOME OPSINS | Genes involved in Opsins |
| 0.1 | 0.7 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.1 | 1.6 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.1 | 2.1 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.1 | 3.9 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.1 | 0.7 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.1 | 1.1 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.0 | 1.2 | REACTOME SPRY REGULATION OF FGF SIGNALING | Genes involved in Spry regulation of FGF signaling |
| 0.0 | 1.4 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 0.7 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.0 | 1.1 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.0 | 1.7 | REACTOME GPVI MEDIATED ACTIVATION CASCADE | Genes involved in GPVI-mediated activation cascade |
| 0.0 | 0.8 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.0 | 0.6 | REACTOME CRMPS IN SEMA3A SIGNALING | Genes involved in CRMPs in Sema3A signaling |
| 0.0 | 0.8 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.0 | 1.5 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 2.8 | REACTOME TRANSPORT OF GLUCOSE AND OTHER SUGARS BILE SALTS AND ORGANIC ACIDS METAL IONS AND AMINE COMPOUNDS | Genes involved in Transport of glucose and other sugars, bile salts and organic acids, metal ions and amine compounds |
| 0.0 | 1.3 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 0.7 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.0 | 4.1 | REACTOME TRANSMISSION ACROSS CHEMICAL SYNAPSES | Genes involved in Transmission across Chemical Synapses |
| 0.0 | 0.8 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.0 | 1.0 | REACTOME G ALPHA1213 SIGNALLING EVENTS | Genes involved in G alpha (12/13) signalling events |
| 0.0 | 1.7 | REACTOME G ALPHA Q SIGNALLING EVENTS | Genes involved in G alpha (q) signalling events |