Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
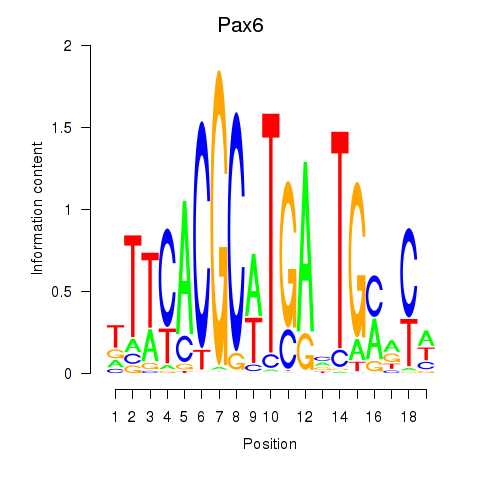
Results for Pax6
Z-value: 0.70
Transcription factors associated with Pax6
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Pax6
|
ENSMUSG00000027168.22 | Pax6 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Pax6 | mm39_v1_chr2_+_105499280_105499297 | 0.18 | 1.4e-01 | Click! |
Activity profile of Pax6 motif
Sorted Z-values of Pax6 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Pax6
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr7_-_86425016 | 5.07 |
ENSMUST00000107271.10
|
Folh1
|
folate hydrolase 1 |
| chr7_-_86425147 | 4.30 |
ENSMUST00000001824.7
|
Folh1
|
folate hydrolase 1 |
| chr11_+_72326337 | 4.25 |
ENSMUST00000076443.10
|
Ggt6
|
gamma-glutamyltransferase 6 |
| chr15_-_101759212 | 3.89 |
ENSMUST00000023790.5
|
Krt1
|
keratin 1 |
| chrX_+_102465616 | 3.65 |
ENSMUST00000182089.2
|
Gm26992
|
predicted gene, 26992 |
| chr1_+_106866678 | 3.58 |
ENSMUST00000112724.3
|
Serpinb12
|
serine (or cysteine) peptidase inhibitor, clade B (ovalbumin), member 12 |
| chr4_-_59915035 | 3.21 |
ENSMUST00000030081.2
|
Slc46a2
|
solute carrier family 46, member 2 |
| chr11_+_72326358 | 3.17 |
ENSMUST00000108499.2
|
Ggt6
|
gamma-glutamyltransferase 6 |
| chr11_+_72326391 | 3.15 |
ENSMUST00000100903.3
|
Ggt6
|
gamma-glutamyltransferase 6 |
| chr13_-_110416637 | 2.80 |
ENSMUST00000167824.3
ENSMUST00000224180.2 |
Rab3c
|
RAB3C, member RAS oncogene family |
| chr4_+_85123654 | 2.79 |
ENSMUST00000030212.15
ENSMUST00000107189.8 ENSMUST00000107184.8 |
Sh3gl2
|
SH3-domain GRB2-like 2 |
| chr3_-_98471301 | 2.72 |
ENSMUST00000058728.10
|
Gm10681
|
predicted gene 10681 |
| chr11_-_121410152 | 2.69 |
ENSMUST00000092298.6
|
Zfp750
|
zinc finger protein 750 |
| chr10_+_75409282 | 2.53 |
ENSMUST00000006508.10
|
Ggt1
|
gamma-glutamyltransferase 1 |
| chr3_-_98660781 | 2.48 |
ENSMUST00000094050.11
ENSMUST00000090743.13 |
Hsd3b3
|
hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 3 |
| chr5_+_110248276 | 2.42 |
ENSMUST00000141066.8
|
Plcxd1
|
phosphatidylinositol-specific phospholipase C, X domain containing 1 |
| chr15_+_6329278 | 2.23 |
ENSMUST00000159046.2
ENSMUST00000161040.8 |
Dab2
|
disabled 2, mitogen-responsive phosphoprotein |
| chr5_-_74838461 | 2.21 |
ENSMUST00000117525.8
ENSMUST00000113531.9 ENSMUST00000039744.13 ENSMUST00000121690.8 |
Lnx1
|
ligand of numb-protein X 1 |
| chr6_-_144994327 | 2.20 |
ENSMUST00000204138.3
|
Bcat1
|
branched chain aminotransferase 1, cytosolic |
| chr5_-_86893645 | 2.18 |
ENSMUST00000161306.2
|
Tmprss11e
|
transmembrane protease, serine 11e |
| chr3_-_98364359 | 2.09 |
ENSMUST00000188356.3
ENSMUST00000167753.8 |
Gm4450
|
predicted gene 4450 |
| chr7_-_101518217 | 2.08 |
ENSMUST00000123321.8
|
Folr1
|
folate receptor 1 (adult) |
| chr15_+_6329263 | 2.03 |
ENSMUST00000078019.13
|
Dab2
|
disabled 2, mitogen-responsive phosphoprotein |
| chr6_-_144994534 | 1.91 |
ENSMUST00000032402.12
|
Bcat1
|
branched chain aminotransferase 1, cytosolic |
| chr4_+_85123358 | 1.86 |
ENSMUST00000107188.10
|
Sh3gl2
|
SH3-domain GRB2-like 2 |
| chr7_-_101517874 | 1.85 |
ENSMUST00000150184.2
|
Folr1
|
folate receptor 1 (adult) |
| chr11_+_69016722 | 1.85 |
ENSMUST00000021268.9
|
Aloxe3
|
arachidonate lipoxygenase 3 |
| chrX_+_36059274 | 1.80 |
ENSMUST00000016463.4
|
Slc25a5
|
solute carrier family 25 (mitochondrial carrier, adenine nucleotide translocator), member 5 |
| chrX_+_16485937 | 1.79 |
ENSMUST00000026013.6
|
Maoa
|
monoamine oxidase A |
| chr9_+_15150341 | 1.72 |
ENSMUST00000034413.8
|
Vstm5
|
V-set and transmembrane domain containing 5 |
| chr17_-_57366795 | 1.51 |
ENSMUST00000040280.14
|
Slc25a23
|
solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 23 |
| chr9_-_54554483 | 1.44 |
ENSMUST00000128163.8
|
Acsbg1
|
acyl-CoA synthetase bubblegum family member 1 |
| chr12_-_11200306 | 1.43 |
ENSMUST00000055673.2
|
Kcns3
|
potassium voltage-gated channel, delayed-rectifier, subfamily S, member 3 |
| chr16_+_34605282 | 1.38 |
ENSMUST00000023538.9
|
Mylk
|
myosin, light polypeptide kinase |
| chr1_-_120047868 | 1.37 |
ENSMUST00000112648.8
|
Dbi
|
diazepam binding inhibitor |
| chr3_-_89152320 | 1.31 |
ENSMUST00000107464.8
ENSMUST00000090924.13 |
Trim46
|
tripartite motif-containing 46 |
| chr13_-_110417421 | 1.30 |
ENSMUST00000223922.2
|
Rab3c
|
RAB3C, member RAS oncogene family |
| chr6_+_83119368 | 1.25 |
ENSMUST00000130622.8
ENSMUST00000129316.2 |
Rtkn
|
rhotekin |
| chr19_-_57107330 | 1.24 |
ENSMUST00000111526.8
|
Ablim1
|
actin-binding LIM protein 1 |
| chr17_+_25585255 | 1.04 |
ENSMUST00000234477.2
|
Tpsb2
|
tryptase beta 2 |
| chr13_-_25121568 | 0.95 |
ENSMUST00000037615.7
|
Aldh5a1
|
aldhehyde dehydrogenase family 5, subfamily A1 |
| chr7_+_12631727 | 0.95 |
ENSMUST00000055528.11
ENSMUST00000117189.2 ENSMUST00000120809.2 ENSMUST00000119989.3 |
Zscan22
|
zinc finger and SCAN domain containing 22 |
| chr10_-_127025851 | 0.93 |
ENSMUST00000222006.2
ENSMUST00000019611.15 ENSMUST00000219245.2 |
Arhgef25
|
Rho guanine nucleotide exchange factor (GEF) 25 |
| chr2_+_119857961 | 0.93 |
ENSMUST00000044675.5
ENSMUST00000129685.8 ENSMUST00000156805.2 |
Jmjd7
Gm28042
|
jumonji domain containing 7 predicted gene, 28042 |
| chr19_-_57107413 | 0.91 |
ENSMUST00000111528.8
ENSMUST00000111529.8 ENSMUST00000104902.9 |
Ablim1
|
actin-binding LIM protein 1 |
| chr9_-_22218934 | 0.87 |
ENSMUST00000086278.7
ENSMUST00000215202.2 |
Zfp810
|
zinc finger protein 810 |
| chr13_-_4573312 | 0.86 |
ENSMUST00000221564.2
ENSMUST00000078239.5 ENSMUST00000080361.13 |
Akr1c20
|
aldo-keto reductase family 1, member C20 |
| chr2_-_117173190 | 0.85 |
ENSMUST00000173541.8
ENSMUST00000172901.8 ENSMUST00000173252.2 |
Rasgrp1
|
RAS guanyl releasing protein 1 |
| chr10_+_94386714 | 0.81 |
ENSMUST00000148910.3
ENSMUST00000117460.8 |
Tmcc3
|
transmembrane and coiled coil domains 3 |
| chr9_+_50686647 | 0.75 |
ENSMUST00000159576.2
|
Alg9
|
asparagine-linked glycosylation 9 (alpha 1,2 mannosyltransferase) |
| chr4_-_140787852 | 0.71 |
ENSMUST00000144196.2
ENSMUST00000097816.9 |
Crocc
|
ciliary rootlet coiled-coil, rootletin |
| chr1_+_155916655 | 0.70 |
ENSMUST00000065648.15
ENSMUST00000097526.3 |
Tor1aip2
|
torsin A interacting protein 2 |
| chr11_-_6150411 | 0.69 |
ENSMUST00000066496.10
|
Nudcd3
|
NudC domain containing 3 |
| chr2_-_117173312 | 0.69 |
ENSMUST00000178884.8
|
Rasgrp1
|
RAS guanyl releasing protein 1 |
| chr5_+_87148697 | 0.68 |
ENSMUST00000031186.9
|
Ugt2b35
|
UDP glucuronosyltransferase 2 family, polypeptide B35 |
| chr15_+_95698574 | 0.67 |
ENSMUST00000226793.2
|
Ano6
|
anoctamin 6 |
| chr11_+_67857268 | 0.66 |
ENSMUST00000021286.11
ENSMUST00000108675.2 |
Stx8
|
syntaxin 8 |
| chr10_-_17898977 | 0.64 |
ENSMUST00000020002.9
|
Abracl
|
ABRA C-terminal like |
| chr1_+_21310803 | 0.62 |
ENSMUST00000027067.15
|
Gsta3
|
glutathione S-transferase, alpha 3 |
| chr2_-_75812311 | 0.60 |
ENSMUST00000099994.5
|
Ttc30a1
|
tetratricopeptide repeat domain 30A1 |
| chr6_-_122833109 | 0.59 |
ENSMUST00000042081.9
|
C3ar1
|
complement component 3a receptor 1 |
| chr5_+_124045552 | 0.58 |
ENSMUST00000166233.2
|
Denr
|
density-regulated protein |
| chr6_-_124718316 | 0.57 |
ENSMUST00000004389.6
|
Grcc10
|
gene rich cluster, C10 gene |
| chr7_+_5054514 | 0.55 |
ENSMUST00000069324.7
|
Zfp580
|
zinc finger protein 580 |
| chr5_-_38637624 | 0.54 |
ENSMUST00000067886.12
|
Slc2a9
|
solute carrier family 2 (facilitated glucose transporter), member 9 |
| chr1_+_21310821 | 0.51 |
ENSMUST00000121676.8
ENSMUST00000124990.3 |
Gsta3
|
glutathione S-transferase, alpha 3 |
| chr5_+_142615292 | 0.50 |
ENSMUST00000036872.16
ENSMUST00000110778.2 |
Wipi2
|
WD repeat domain, phosphoinositide interacting 2 |
| chr16_-_52272828 | 0.49 |
ENSMUST00000170035.8
ENSMUST00000164728.8 ENSMUST00000168071.2 |
Alcam
|
activated leukocyte cell adhesion molecule |
| chr4_+_155575792 | 0.49 |
ENSMUST00000165335.8
ENSMUST00000105616.10 ENSMUST00000030940.14 |
Gnb1
|
guanine nucleotide binding protein (G protein), beta 1 |
| chr7_+_25760922 | 0.49 |
ENSMUST00000005669.9
|
Cyp2b13
|
cytochrome P450, family 2, subfamily b, polypeptide 13 |
| chr6_-_88021999 | 0.42 |
ENSMUST00000113598.8
|
Rab7
|
RAB7, member RAS oncogene family |
| chr6_+_29468067 | 0.42 |
ENSMUST00000143101.4
ENSMUST00000149646.3 |
Atp6v1f
|
ATPase, H+ transporting, lysosomal V1 subunit F |
| chr9_+_105520154 | 0.42 |
ENSMUST00000190358.2
ENSMUST00000191268.7 ENSMUST00000065778.13 ENSMUST00000188784.2 |
Pik3r4
|
phosphoinositide-3-kinase regulatory subunit 4 |
| chr19_+_34560922 | 0.41 |
ENSMUST00000102825.4
|
Ifit3
|
interferon-induced protein with tetratricopeptide repeats 3 |
| chr13_-_24098981 | 0.41 |
ENSMUST00000110407.4
|
Slc17a4
|
solute carrier family 17 (sodium phosphate), member 4 |
| chr14_-_103583785 | 0.39 |
ENSMUST00000160758.8
|
Mycbp2
|
MYC binding protein 2, E3 ubiquitin protein ligase |
| chr8_-_47866869 | 0.38 |
ENSMUST00000211882.2
|
Stox2
|
storkhead box 2 |
| chr13_+_63963054 | 0.35 |
ENSMUST00000021926.13
ENSMUST00000067821.13 ENSMUST00000144763.2 ENSMUST00000021925.14 ENSMUST00000238465.2 |
Ercc6l2
|
excision repair cross-complementing rodent repair deficiency, complementation group 6 like 2 |
| chr1_+_59802543 | 0.35 |
ENSMUST00000087435.7
|
Bmpr2
|
bone morphogenetic protein receptor, type II (serine/threonine kinase) |
| chr16_+_20317544 | 0.35 |
ENSMUST00000003320.14
|
Eif2b5
|
eukaryotic translation initiation factor 2B, subunit 5 epsilon |
| chr4_-_123507494 | 0.34 |
ENSMUST00000238866.2
|
Macf1
|
microtubule-actin crosslinking factor 1 |
| chr5_-_38637474 | 0.33 |
ENSMUST00000143758.8
ENSMUST00000156272.8 |
Slc2a9
|
solute carrier family 2 (facilitated glucose transporter), member 9 |
| chr8_-_27718522 | 0.32 |
ENSMUST00000117565.2
|
Adrb3
|
adrenergic receptor, beta 3 |
| chr12_+_76593799 | 0.32 |
ENSMUST00000218380.2
ENSMUST00000219751.2 |
Plekhg3
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 3 |
| chr7_+_86444235 | 0.30 |
ENSMUST00000233099.2
ENSMUST00000164996.2 |
Vmn2r77
|
vomeronasal 2, receptor 77 |
| chr2_-_105734829 | 0.26 |
ENSMUST00000122965.8
|
Elp4
|
elongator acetyltransferase complex subunit 4 |
| chr13_-_24098951 | 0.25 |
ENSMUST00000021769.16
|
Slc17a4
|
solute carrier family 17 (sodium phosphate), member 4 |
| chr10_-_126866658 | 0.25 |
ENSMUST00000120547.2
ENSMUST00000152054.8 |
Tsfm
|
Ts translation elongation factor, mitochondrial |
| chr18_-_80133177 | 0.25 |
ENSMUST00000178391.2
|
Gm21886
|
predicted gene, 21886 |
| chr1_+_155916629 | 0.25 |
ENSMUST00000060404.5
|
Tor1aip2
|
torsin A interacting protein 2 |
| chr7_-_85956319 | 0.24 |
ENSMUST00000055690.3
|
Olfr309
|
olfactory receptor 309 |
| chr15_-_35938155 | 0.24 |
ENSMUST00000156915.3
|
Cox6c
|
cytochrome c oxidase subunit 6C |
| chr2_-_131987008 | 0.23 |
ENSMUST00000028815.15
|
Slc23a2
|
solute carrier family 23 (nucleobase transporters), member 2 |
| chr11_+_78219241 | 0.23 |
ENSMUST00000048073.9
|
Pigs
|
phosphatidylinositol glycan anchor biosynthesis, class S |
| chr11_-_72097821 | 0.22 |
ENSMUST00000204457.3
|
Gm43951
|
predicted gene, 43951 |
| chr11_+_60590584 | 0.21 |
ENSMUST00000108719.4
|
Llgl1
|
LLGL1 scribble cell polarity complex component |
| chr1_-_192812509 | 0.21 |
ENSMUST00000085555.7
|
Utp25
|
UTP25 small subunit processome component |
| chr11_+_60590498 | 0.21 |
ENSMUST00000052346.10
|
Llgl1
|
LLGL1 scribble cell polarity complex component |
| chr1_-_138547403 | 0.19 |
ENSMUST00000027642.5
ENSMUST00000186017.7 |
Nek7
|
NIMA (never in mitosis gene a)-related expressed kinase 7 |
| chr5_-_134668152 | 0.17 |
ENSMUST00000036125.10
ENSMUST00000202622.4 |
Eif4h
|
eukaryotic translation initiation factor 4H |
| chr9_+_64086553 | 0.16 |
ENSMUST00000034965.8
|
Snapc5
|
small nuclear RNA activating complex, polypeptide 5 |
| chr10_+_80192293 | 0.15 |
ENSMUST00000039836.15
ENSMUST00000105351.2 |
Plk5
|
polo like kinase 5 |
| chr1_+_173877941 | 0.14 |
ENSMUST00000062665.4
|
Olfr432
|
olfactory receptor 432 |
| chr2_-_33777874 | 0.14 |
ENSMUST00000041555.10
|
Mvb12b
|
multivesicular body subunit 12B |
| chr9_-_117701613 | 0.13 |
ENSMUST00000239475.2
|
Rbms3
|
RNA binding motif, single stranded interacting protein |
| chr13_+_73775001 | 0.10 |
ENSMUST00000022104.9
|
Tert
|
telomerase reverse transcriptase |
| chr7_+_75259778 | 0.07 |
ENSMUST00000207923.2
|
Akap13
|
A kinase (PRKA) anchor protein 13 |
| chr16_+_3702604 | 0.03 |
ENSMUST00000115860.8
|
Naa60
|
N(alpha)-acetyltransferase 60, NatF catalytic subunit |
| chr11_+_102159558 | 0.03 |
ENSMUST00000036467.5
|
Asb16
|
ankyrin repeat and SOCS box-containing 16 |
| chr4_+_148123554 | 0.02 |
ENSMUST00000141283.8
|
Mthfr
|
methylenetetrahydrofolate reductase |
| chr6_+_56933451 | 0.02 |
ENSMUST00000096612.4
|
Vmn1r4
|
vomeronasal 1 receptor 4 |
| chr1_+_63215976 | 0.01 |
ENSMUST00000129339.8
|
Eef1b2
|
eukaryotic translation elongation factor 1 beta 2 |
| chr5_+_124045238 | 0.01 |
ENSMUST00000023869.15
|
Denr
|
density-regulated protein |
| chr16_+_3702523 | 0.01 |
ENSMUST00000176625.8
ENSMUST00000186375.8 |
Naa60
|
N(alpha)-acetyltransferase 60, NatF catalytic subunit |
| chr15_-_35938328 | 0.01 |
ENSMUST00000014457.15
|
Cox6c
|
cytochrome c oxidase subunit 6C |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 13.1 | GO:0006751 | glutathione catabolic process(GO:0006751) |
| 0.7 | 4.3 | GO:0035026 | leading edge cell differentiation(GO:0035026) |
| 0.5 | 4.1 | GO:0009099 | branched-chain amino acid biosynthetic process(GO:0009082) leucine biosynthetic process(GO:0009098) valine biosynthetic process(GO:0009099) |
| 0.5 | 4.7 | GO:1990416 | cellular response to brain-derived neurotrophic factor stimulus(GO:1990416) |
| 0.4 | 3.9 | GO:0061713 | neural crest cell migration involved in heart formation(GO:0003147) anterior neural tube closure(GO:0061713) cellular response to folic acid(GO:0071231) |
| 0.4 | 1.5 | GO:0097274 | urea homeostasis(GO:0097274) |
| 0.4 | 1.1 | GO:1901377 | mycotoxin catabolic process(GO:0043387) aflatoxin catabolic process(GO:0046223) organic heteropentacyclic compound catabolic process(GO:1901377) |
| 0.4 | 1.8 | GO:0051122 | hepoxilin metabolic process(GO:0051121) hepoxilin biosynthetic process(GO:0051122) |
| 0.3 | 3.9 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.3 | 1.4 | GO:0060414 | aorta smooth muscle tissue morphogenesis(GO:0060414) |
| 0.3 | 1.4 | GO:0036151 | phosphatidylcholine acyl-chain remodeling(GO:0036151) |
| 0.3 | 1.8 | GO:0042420 | dopamine catabolic process(GO:0042420) |
| 0.2 | 9.4 | GO:0006760 | folic acid-containing compound metabolic process(GO:0006760) |
| 0.2 | 0.7 | GO:0002543 | activation of blood coagulation via clotting cascade(GO:0002543) phosphatidylserine exposure on blood platelet(GO:0097045) |
| 0.2 | 1.8 | GO:1901029 | negative regulation of mitochondrial outer membrane permeabilization involved in apoptotic signaling pathway(GO:1901029) |
| 0.2 | 0.6 | GO:0035720 | intraciliary anterograde transport(GO:0035720) |
| 0.2 | 0.9 | GO:0006538 | glutamate catabolic process(GO:0006538) |
| 0.1 | 0.5 | GO:0061739 | protein lipidation involved in autophagosome assembly(GO:0061739) |
| 0.1 | 0.7 | GO:0032053 | ciliary basal body organization(GO:0032053) positive regulation of protein localization to cilium(GO:1903566) |
| 0.1 | 0.3 | GO:2000137 | regulation of lung blood pressure(GO:0014916) negative regulation of cell proliferation involved in heart morphogenesis(GO:2000137) cell proliferation involved in heart valve development(GO:2000793) |
| 0.1 | 0.6 | GO:0036343 | psychomotor behavior(GO:0036343) |
| 0.1 | 0.4 | GO:1903542 | negative regulation of exosomal secretion(GO:1903542) |
| 0.1 | 1.5 | GO:1902715 | positive regulation of interferon-gamma secretion(GO:1902715) |
| 0.1 | 0.6 | GO:0002188 | translation reinitiation(GO:0002188) |
| 0.1 | 0.6 | GO:0002430 | complement receptor mediated signaling pathway(GO:0002430) tolerance induction dependent upon immune response(GO:0002461) |
| 0.1 | 0.3 | GO:0002025 | vasodilation by norepinephrine-epinephrine involved in regulation of systemic arterial blood pressure(GO:0002025) |
| 0.1 | 0.2 | GO:0015882 | L-ascorbic acid transport(GO:0015882) transepithelial L-ascorbic acid transport(GO:0070904) |
| 0.1 | 1.3 | GO:0048490 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.0 | 1.3 | GO:0070233 | negative regulation of T cell apoptotic process(GO:0070233) |
| 0.0 | 0.9 | GO:0046415 | urate metabolic process(GO:0046415) |
| 0.0 | 0.4 | GO:0030242 | pexophagy(GO:0030242) |
| 0.0 | 0.2 | GO:0016255 | attachment of GPI anchor to protein(GO:0016255) |
| 0.0 | 0.4 | GO:0021785 | branchiomotor neuron axon guidance(GO:0021785) |
| 0.0 | 0.1 | GO:1900369 | negative regulation of RNA interference(GO:1900369) |
| 0.0 | 4.0 | GO:0019882 | antigen processing and presentation(GO:0019882) |
| 0.0 | 0.4 | GO:0035457 | cellular response to interferon-alpha(GO:0035457) |
| 0.0 | 0.5 | GO:0090435 | protein localization to nuclear envelope(GO:0090435) |
| 0.0 | 0.5 | GO:0007603 | phototransduction, visible light(GO:0007603) |
| 0.0 | 1.4 | GO:0000038 | very long-chain fatty acid metabolic process(GO:0000038) |
| 0.0 | 0.2 | GO:0006384 | transcription initiation from RNA polymerase III promoter(GO:0006384) |
| 0.0 | 0.3 | GO:0006620 | posttranslational protein targeting to membrane(GO:0006620) |
| 0.0 | 0.4 | GO:0035090 | maintenance of apical/basal cell polarity(GO:0035090) |
| 0.0 | 0.4 | GO:0036297 | interstrand cross-link repair(GO:0036297) |
| 0.0 | 0.5 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 0.0 | 0.1 | GO:1900169 | regulation of glucocorticoid mediated signaling pathway(GO:1900169) |
| 0.0 | 3.3 | GO:0006694 | steroid biosynthetic process(GO:0006694) |
| 0.0 | 0.7 | GO:0045022 | early endosome to late endosome transport(GO:0045022) |
| 0.0 | 3.6 | GO:0002244 | hematopoietic progenitor cell differentiation(GO:0002244) |
| 0.0 | 0.2 | GO:0001731 | formation of translation preinitiation complex(GO:0001731) |
| 0.0 | 1.5 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.0 | 0.5 | GO:0008045 | motor neuron axon guidance(GO:0008045) |
| 0.0 | 0.6 | GO:0032757 | positive regulation of interleukin-8 production(GO:0032757) |
| 0.0 | 0.4 | GO:0090662 | ATP hydrolysis coupled proton transport(GO:0015991) ATP hydrolysis coupled transmembrane transport(GO:0090662) |
| 0.0 | 0.3 | GO:0014002 | astrocyte development(GO:0014002) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 4.3 | GO:0070022 | transforming growth factor beta receptor homodimeric complex(GO:0070022) |
| 0.4 | 1.8 | GO:0071817 | MMXD complex(GO:0071817) |
| 0.3 | 1.3 | GO:1990769 | proximal neuron projection(GO:1990769) |
| 0.2 | 0.6 | GO:0070992 | translation initiation complex(GO:0070992) |
| 0.1 | 3.9 | GO:0031362 | anchored component of external side of plasma membrane(GO:0031362) |
| 0.1 | 3.9 | GO:0045095 | keratin filament(GO:0045095) |
| 0.1 | 0.4 | GO:0034271 | phosphatidylinositol 3-kinase complex, class III, type I(GO:0034271) phosphatidylinositol 3-kinase complex, class III, type II(GO:0034272) |
| 0.1 | 0.9 | GO:0034045 | pre-autophagosomal structure membrane(GO:0034045) |
| 0.0 | 0.3 | GO:0005851 | eukaryotic translation initiation factor 2B complex(GO:0005851) |
| 0.0 | 1.4 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.0 | 0.3 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.0 | 0.4 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.0 | 0.2 | GO:0042765 | GPI-anchor transamidase complex(GO:0042765) |
| 0.0 | 0.4 | GO:0000137 | Golgi cis cisterna(GO:0000137) |
| 0.0 | 0.7 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.0 | 2.5 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.0 | 0.5 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.0 | 3.5 | GO:0097517 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.0 | 0.7 | GO:0005868 | cytoplasmic dynein complex(GO:0005868) |
| 0.0 | 4.1 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.0 | 0.6 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.0 | 0.5 | GO:0042101 | T cell receptor complex(GO:0042101) |
| 0.0 | 5.5 | GO:0005769 | early endosome(GO:0005769) |
| 0.0 | 0.2 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 13.1 | GO:0036374 | glutathione hydrolase activity(GO:0036374) |
| 0.6 | 3.9 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.5 | 4.1 | GO:0004084 | branched-chain-amino-acid transaminase activity(GO:0004084) L-leucine transaminase activity(GO:0052654) L-valine transaminase activity(GO:0052655) L-isoleucine transaminase activity(GO:0052656) |
| 0.4 | 9.4 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.4 | 3.9 | GO:0051870 | methotrexate binding(GO:0051870) |
| 0.4 | 2.5 | GO:0004769 | steroid delta-isomerase activity(GO:0004769) |
| 0.3 | 1.4 | GO:0030156 | benzodiazepine receptor binding(GO:0030156) |
| 0.3 | 3.3 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) ADP transmembrane transporter activity(GO:0015217) |
| 0.2 | 4.3 | GO:0098748 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
| 0.2 | 1.8 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.2 | 0.7 | GO:0000026 | alpha-1,2-mannosyltransferase activity(GO:0000026) |
| 0.2 | 1.2 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.2 | 1.4 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.1 | 4.1 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.1 | 1.5 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.1 | 1.4 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.1 | 0.3 | GO:0004939 | beta-adrenergic receptor activity(GO:0004939) beta-3 adrenergic receptor binding(GO:0031699) |
| 0.1 | 0.2 | GO:0008520 | L-ascorbate:sodium symporter activity(GO:0008520) L-ascorbic acid transporter activity(GO:0015229) sodium-dependent L-ascorbate transmembrane transporter activity(GO:0070890) |
| 0.1 | 0.6 | GO:0004875 | complement receptor activity(GO:0004875) |
| 0.1 | 2.7 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.1 | 0.9 | GO:0015143 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.0 | 0.7 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 0.0 | 0.7 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) |
| 0.0 | 0.3 | GO:0008607 | phosphorylase kinase regulator activity(GO:0008607) |
| 0.0 | 0.3 | GO:0098821 | BMP binding(GO:0036122) BMP receptor activity(GO:0098821) |
| 0.0 | 0.5 | GO:0047391 | alkylglycerophosphoethanolamine phosphodiesterase activity(GO:0047391) |
| 0.0 | 0.2 | GO:0003923 | GPI-anchor transamidase activity(GO:0003923) |
| 0.0 | 0.9 | GO:0004033 | aldo-keto reductase (NADP) activity(GO:0004033) |
| 0.0 | 0.7 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.0 | 0.5 | GO:0010314 | phosphatidylinositol-5-phosphate binding(GO:0010314) |
| 0.0 | 0.8 | GO:0016701 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen(GO:0016701) oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.0 | 0.9 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.0 | 0.2 | GO:0033592 | RNA strand annealing activity(GO:0033592) |
| 0.0 | 1.1 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.0 | 2.4 | GO:0008081 | phosphoric diester hydrolase activity(GO:0008081) |
| 0.0 | 2.8 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.7 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.0 | 0.9 | GO:0016620 | oxidoreductase activity, acting on the aldehyde or oxo group of donors, NAD or NADP as acceptor(GO:0016620) |
| 0.0 | 0.9 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.5 | GO:0008392 | arachidonic acid epoxygenase activity(GO:0008392) |
| 0.0 | 3.0 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 0.4 | GO:0046961 | proton-transporting ATPase activity, rotational mechanism(GO:0046961) |
| 0.0 | 2.2 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 1.4 | GO:0005249 | voltage-gated potassium channel activity(GO:0005249) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 4.7 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.0 | 4.3 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 0.3 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.0 | 1.5 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 0.5 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.0 | 4.2 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 1.4 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.0 | 3.9 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 2.2 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 0.9 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 3.5 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 4.3 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 0.2 | 4.7 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.2 | 4.1 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.1 | 1.8 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.1 | 3.7 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 0.5 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.1 | 2.2 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |
| 0.1 | 0.9 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.1 | 0.9 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.0 | 0.7 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.0 | 1.8 | REACTOME INTERACTIONS OF VPR WITH HOST CELLULAR PROTEINS | Genes involved in Interactions of Vpr with host cellular proteins |
| 0.0 | 1.5 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 1.4 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.0 | 1.4 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.0 | 0.7 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.0 | 0.4 | REACTOME INSULIN RECEPTOR RECYCLING | Genes involved in Insulin receptor recycling |
| 0.0 | 0.3 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |