Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
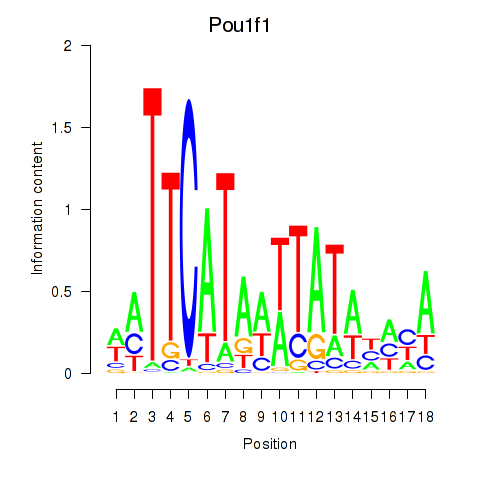
Results for Pou1f1
Z-value: 2.45
Transcription factors associated with Pou1f1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Pou1f1
|
ENSMUSG00000004842.19 | Pou1f1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Pou1f1 | mm39_v1_chr16_+_65317389_65317434 | -0.18 | 1.3e-01 | Click! |
Activity profile of Pou1f1 motif
Sorted Z-values of Pou1f1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Pou1f1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_-_87074380 | 34.28 |
ENSMUST00000031183.3
|
Ugt2b1
|
UDP glucuronosyltransferase 2 family, polypeptide B1 |
| chr5_-_87485023 | 27.96 |
ENSMUST00000031195.3
|
Ugt2a3
|
UDP glucuronosyltransferase 2 family, polypeptide A3 |
| chr5_-_87288177 | 26.38 |
ENSMUST00000067790.7
|
Ugt2b5
|
UDP glucuronosyltransferase 2 family, polypeptide B5 |
| chr13_+_4486105 | 24.66 |
ENSMUST00000156277.2
|
Akr1c6
|
aldo-keto reductase family 1, member C6 |
| chr5_-_87240405 | 22.65 |
ENSMUST00000132667.2
ENSMUST00000145617.8 ENSMUST00000094649.11 |
Ugt2b36
|
UDP glucuronosyltransferase 2 family, polypeptide B36 |
| chr19_-_39637489 | 22.04 |
ENSMUST00000067328.7
|
Cyp2c67
|
cytochrome P450, family 2, subfamily c, polypeptide 67 |
| chr19_+_39980868 | 18.97 |
ENSMUST00000049178.3
|
Cyp2c37
|
cytochrome P450, family 2. subfamily c, polypeptide 37 |
| chr10_-_127843377 | 16.39 |
ENSMUST00000219447.2
ENSMUST00000219780.2 ENSMUST00000219707.2 ENSMUST00000219953.2 ENSMUST00000219183.2 |
Hsd17b6
|
hydroxysteroid (17-beta) dehydrogenase 6 |
| chr5_-_145816774 | 16.17 |
ENSMUST00000035918.8
|
Cyp3a11
|
cytochrome P450, family 3, subfamily a, polypeptide 11 |
| chr4_-_96552349 | 15.91 |
ENSMUST00000030299.8
|
Cyp2j5
|
cytochrome P450, family 2, subfamily j, polypeptide 5 |
| chr5_-_87572060 | 14.98 |
ENSMUST00000072818.6
|
Ugt2b38
|
UDP glucuronosyltransferase 2 family, polypeptide B38 |
| chr4_-_60457902 | 14.84 |
ENSMUST00000084548.11
ENSMUST00000103012.10 ENSMUST00000107499.4 |
Mup1
|
major urinary protein 1 |
| chr8_+_110717062 | 14.81 |
ENSMUST00000001720.14
ENSMUST00000143741.2 |
Tat
|
tyrosine aminotransferase |
| chr10_+_87357657 | 14.80 |
ENSMUST00000020241.17
|
Pah
|
phenylalanine hydroxylase |
| chr2_-_110136074 | 14.69 |
ENSMUST00000046233.9
|
Bbox1
|
butyrobetaine (gamma), 2-oxoglutarate dioxygenase 1 (gamma-butyrobetaine hydroxylase) |
| chr4_-_60222580 | 14.06 |
ENSMUST00000095058.5
ENSMUST00000163931.8 |
Mup8
|
major urinary protein 8 |
| chr4_-_60697274 | 13.67 |
ENSMUST00000117932.2
|
Mup12
|
major urinary protein 12 |
| chr1_-_139708906 | 13.64 |
ENSMUST00000111986.8
ENSMUST00000027612.11 ENSMUST00000111989.9 |
Cfhr4
|
complement factor H-related 4 |
| chr4_-_60777462 | 13.64 |
ENSMUST00000211875.2
|
Mup22
|
major urinary protein 22 |
| chr5_-_89583469 | 13.46 |
ENSMUST00000200534.2
|
Gc
|
vitamin D binding protein |
| chr11_+_108271990 | 13.45 |
ENSMUST00000146050.2
ENSMUST00000152958.8 |
Apoh
|
apolipoprotein H |
| chr4_-_61259997 | 13.29 |
ENSMUST00000071005.9
ENSMUST00000075206.12 |
Mup14
|
major urinary protein 14 |
| chr6_+_40619913 | 12.32 |
ENSMUST00000238599.2
|
Mgam
|
maltase-glucoamylase |
| chr16_-_22848153 | 12.07 |
ENSMUST00000232459.2
|
Kng2
|
kininogen 2 |
| chr13_+_4484305 | 12.07 |
ENSMUST00000021630.15
|
Akr1c6
|
aldo-keto reductase family 1, member C6 |
| chr8_-_41668182 | 11.84 |
ENSMUST00000034003.5
|
Fgl1
|
fibrinogen-like protein 1 |
| chr6_+_121815473 | 11.64 |
ENSMUST00000032228.9
|
Mug1
|
murinoglobulin 1 |
| chr3_+_82915031 | 11.60 |
ENSMUST00000048486.13
ENSMUST00000194175.2 |
Fgg
|
fibrinogen gamma chain |
| chr1_-_184543367 | 11.22 |
ENSMUST00000048462.13
ENSMUST00000110992.9 |
Mtarc1
|
mitochondrial amidoxime reducing component 1 |
| chr7_-_46382003 | 11.09 |
ENSMUST00000006952.9
|
Saa4
|
serum amyloid A 4 |
| chr8_-_45747883 | 11.01 |
ENSMUST00000026907.6
|
Klkb1
|
kallikrein B, plasma 1 |
| chr3_+_138121245 | 10.73 |
ENSMUST00000161312.8
ENSMUST00000013458.9 |
Adh4
|
alcohol dehydrogenase 4 (class II), pi polypeptide |
| chr19_-_39875192 | 10.66 |
ENSMUST00000168838.3
|
Cyp2c69
|
cytochrome P450, family 2, subfamily c, polypeptide 69 |
| chrX_+_10118544 | 10.52 |
ENSMUST00000049910.13
|
Otc
|
ornithine transcarbamylase |
| chr19_-_40175709 | 10.35 |
ENSMUST00000051846.13
|
Cyp2c70
|
cytochrome P450, family 2, subfamily c, polypeptide 70 |
| chr8_-_93956143 | 10.29 |
ENSMUST00000176282.2
ENSMUST00000034173.14 |
Ces1e
|
carboxylesterase 1E |
| chr4_-_61972348 | 10.20 |
ENSMUST00000074018.4
|
Mup20
|
major urinary protein 20 |
| chr16_+_22877000 | 10.20 |
ENSMUST00000039492.14
ENSMUST00000023589.15 ENSMUST00000089902.8 |
Kng1
|
kininogen 1 |
| chr2_-_10135449 | 10.13 |
ENSMUST00000042290.14
|
Itih2
|
inter-alpha trypsin inhibitor, heavy chain 2 |
| chr1_+_21310803 | 10.12 |
ENSMUST00000027067.15
|
Gsta3
|
glutathione S-transferase, alpha 3 |
| chr4_-_61437704 | 10.10 |
ENSMUST00000095051.6
ENSMUST00000107483.8 |
Mup16
|
major urinary protein 16 |
| chr1_+_21310821 | 10.03 |
ENSMUST00000121676.8
ENSMUST00000124990.3 |
Gsta3
|
glutathione S-transferase, alpha 3 |
| chr4_-_61592331 | 9.92 |
ENSMUST00000098040.4
|
Mup18
|
major urinary protein 18 |
| chr19_-_40062174 | 9.78 |
ENSMUST00000048959.5
|
Cyp2c54
|
cytochrome P450, family 2, subfamily c, polypeptide 54 |
| chr6_+_71176811 | 9.68 |
ENSMUST00000067492.8
|
Fabp1
|
fatty acid binding protein 1, liver |
| chr4_-_60538151 | 9.64 |
ENSMUST00000098047.3
|
Mup10
|
major urinary protein 10 |
| chr19_+_40078132 | 9.48 |
ENSMUST00000068094.13
ENSMUST00000080171.3 |
Cyp2c50
|
cytochrome P450, family 2, subfamily c, polypeptide 50 |
| chr3_-_98660781 | 9.32 |
ENSMUST00000094050.11
ENSMUST00000090743.13 |
Hsd3b3
|
hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 3 |
| chr8_-_94006345 | 9.05 |
ENSMUST00000034178.9
|
Ces1f
|
carboxylesterase 1F |
| chrX_+_10118600 | 9.03 |
ENSMUST00000115528.3
|
Otc
|
ornithine transcarbamylase |
| chr1_+_58152295 | 8.98 |
ENSMUST00000040999.14
ENSMUST00000162011.3 |
Aox3
|
aldehyde oxidase 3 |
| chr4_-_60377932 | 8.97 |
ENSMUST00000107506.9
ENSMUST00000122381.8 ENSMUST00000118759.8 ENSMUST00000132829.3 |
Mup9
|
major urinary protein 9 |
| chr4_+_115375461 | 8.87 |
ENSMUST00000058785.10
ENSMUST00000094886.4 |
Cyp4a10
|
cytochrome P450, family 4, subfamily a, polypeptide 10 |
| chr19_-_39801188 | 8.85 |
ENSMUST00000162507.2
ENSMUST00000160476.9 ENSMUST00000239028.2 |
Cyp2c40
|
cytochrome P450, family 2, subfamily c, polypeptide 40 |
| chr19_+_39275518 | 8.40 |
ENSMUST00000003137.15
|
Cyp2c29
|
cytochrome P450, family 2, subfamily c, polypeptide 29 |
| chr19_+_12577317 | 8.23 |
ENSMUST00000181868.8
ENSMUST00000092931.7 |
Gm4952
|
predicted gene 4952 |
| chr5_-_87054796 | 8.14 |
ENSMUST00000031181.16
ENSMUST00000113333.2 |
Ugt2b34
|
UDP glucuronosyltransferase 2 family, polypeptide B34 |
| chr4_-_60139857 | 8.11 |
ENSMUST00000107490.5
ENSMUST00000074700.9 |
Mup2
|
major urinary protein 2 |
| chr4_-_61700450 | 7.94 |
ENSMUST00000107477.2
ENSMUST00000080606.9 |
Mup19
|
major urinary protein 19 |
| chr5_-_137919873 | 7.74 |
ENSMUST00000031741.8
|
Cyp3a13
|
cytochrome P450, family 3, subfamily a, polypeptide 13 |
| chr7_+_140343652 | 7.51 |
ENSMUST00000026552.9
ENSMUST00000209253.2 ENSMUST00000210235.2 |
Cyp2e1
|
cytochrome P450, family 2, subfamily e, polypeptide 1 |
| chrX_+_20416019 | 7.47 |
ENSMUST00000023832.7
|
Rgn
|
regucalcin |
| chr9_-_121745354 | 7.01 |
ENSMUST00000062474.5
|
Cyp8b1
|
cytochrome P450, family 8, subfamily b, polypeptide 1 |
| chr3_-_98537568 | 6.98 |
ENSMUST00000044094.6
|
Hsd3b5
|
hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 5 |
| chr11_+_101258368 | 6.91 |
ENSMUST00000019469.3
|
G6pc
|
glucose-6-phosphatase, catalytic |
| chr19_-_7943365 | 6.83 |
ENSMUST00000182102.8
ENSMUST00000075619.5 |
Slc22a27
|
solute carrier family 22, member 27 |
| chr4_-_61357980 | 6.82 |
ENSMUST00000095049.5
|
Mup15
|
major urinary protein 15 |
| chr11_-_59937302 | 6.80 |
ENSMUST00000000310.14
ENSMUST00000102693.9 ENSMUST00000148512.2 |
Pemt
|
phosphatidylethanolamine N-methyltransferase |
| chr7_+_13467422 | 6.78 |
ENSMUST00000086148.8
|
Sult2a2
|
sulfotransferase family 2A, dehydroepiandrosterone (DHEA)-preferring, member 2 |
| chr15_+_6474808 | 6.74 |
ENSMUST00000022749.17
ENSMUST00000239466.2 |
C9
|
complement component 9 |
| chr19_+_30210320 | 6.72 |
ENSMUST00000025797.7
|
Mbl2
|
mannose-binding lectin (protein C) 2 |
| chr8_-_45715049 | 6.70 |
ENSMUST00000034064.5
|
F11
|
coagulation factor XI |
| chrX_+_137986975 | 6.69 |
ENSMUST00000033625.2
|
4930513O06Rik
|
RIKEN cDNA 4930513O06 gene |
| chr5_-_147259245 | 6.60 |
ENSMUST00000100433.5
|
Urad
|
ureidoimidazoline (2-oxo-4-hydroxy-4-carboxy-5) decarboxylase |
| chr4_+_141473983 | 6.56 |
ENSMUST00000038161.5
|
Agmat
|
agmatine ureohydrolase (agmatinase) |
| chr6_+_121983720 | 6.53 |
ENSMUST00000081777.8
|
Mug2
|
murinoglobulin 2 |
| chr4_-_60618357 | 6.51 |
ENSMUST00000084544.5
ENSMUST00000098046.10 |
Mup11
|
major urinary protein 11 |
| chr10_+_75242745 | 6.34 |
ENSMUST00000039925.8
|
Upb1
|
ureidopropionase, beta |
| chr19_-_39451509 | 6.25 |
ENSMUST00000035488.3
|
Cyp2c38
|
cytochrome P450, family 2, subfamily c, polypeptide 38 |
| chr15_-_77283286 | 6.21 |
ENSMUST00000175919.8
ENSMUST00000176074.9 |
Apol7a
|
apolipoprotein L 7a |
| chr6_-_128503666 | 6.14 |
ENSMUST00000143664.2
ENSMUST00000112132.8 |
Pzp
|
PZP, alpha-2-macroglobulin like |
| chr3_+_94600863 | 6.09 |
ENSMUST00000090848.10
ENSMUST00000173981.8 ENSMUST00000173849.8 ENSMUST00000174223.2 |
Selenbp2
|
selenium binding protein 2 |
| chr12_-_103423472 | 5.72 |
ENSMUST00000044687.7
|
Ifi27l2b
|
interferon, alpha-inducible protein 27 like 2B |
| chr1_-_72323464 | 5.59 |
ENSMUST00000027381.13
|
Pecr
|
peroxisomal trans-2-enoyl-CoA reductase |
| chr12_-_103925197 | 5.58 |
ENSMUST00000122229.8
|
Serpina1e
|
serine (or cysteine) peptidase inhibitor, clade A, member 1E |
| chr3_+_129630380 | 5.57 |
ENSMUST00000077918.7
|
Cfi
|
complement component factor i |
| chr5_+_90666791 | 5.56 |
ENSMUST00000113179.9
ENSMUST00000128740.2 |
Afm
|
afamin |
| chr15_+_4756684 | 5.55 |
ENSMUST00000161997.8
ENSMUST00000022788.15 |
C6
|
complement component 6 |
| chr19_+_31846154 | 5.48 |
ENSMUST00000224564.2
ENSMUST00000224304.2 ENSMUST00000075838.8 ENSMUST00000224400.2 |
A1cf
|
APOBEC1 complementation factor |
| chr1_-_72323407 | 5.48 |
ENSMUST00000097698.5
|
Pecr
|
peroxisomal trans-2-enoyl-CoA reductase |
| chr10_+_87357782 | 5.46 |
ENSMUST00000219813.2
|
Pah
|
phenylalanine hydroxylase |
| chr10_+_87357816 | 5.43 |
ENSMUST00000218573.2
|
Pah
|
phenylalanine hydroxylase |
| chr8_-_94063823 | 5.41 |
ENSMUST00000044602.8
|
Ces1g
|
carboxylesterase 1G |
| chr3_+_59989282 | 5.39 |
ENSMUST00000029326.6
|
Sucnr1
|
succinate receptor 1 |
| chr1_+_88128323 | 5.36 |
ENSMUST00000049289.9
|
Ugt1a2
|
UDP glucuronosyltransferase 1 family, polypeptide A2 |
| chr17_-_32643067 | 5.26 |
ENSMUST00000237130.2
|
Pglyrp2
|
peptidoglycan recognition protein 2 |
| chr8_-_45786226 | 5.24 |
ENSMUST00000095328.6
|
Cyp4v3
|
cytochrome P450, family 4, subfamily v, polypeptide 3 |
| chr19_-_39729431 | 5.23 |
ENSMUST00000099472.4
|
Cyp2c68
|
cytochrome P450, family 2, subfamily c, polypeptide 68 |
| chr13_-_41373638 | 5.21 |
ENSMUST00000117096.2
|
Elovl2
|
elongation of very long chain fatty acids (FEN1/Elo2, SUR4/Elo3, yeast)-like 2 |
| chr19_-_7780025 | 4.85 |
ENSMUST00000065634.8
|
Slc22a26
|
solute carrier family 22 (organic cation transporter), member 26 |
| chr15_+_4756657 | 4.82 |
ENSMUST00000162585.8
|
C6
|
complement component 6 |
| chr5_+_45650716 | 4.75 |
ENSMUST00000046122.11
|
Lap3
|
leucine aminopeptidase 3 |
| chr5_+_115061293 | 4.74 |
ENSMUST00000031540.11
ENSMUST00000112143.4 |
Oasl1
|
2'-5' oligoadenylate synthetase-like 1 |
| chr3_+_20039775 | 4.74 |
ENSMUST00000172860.2
|
Cp
|
ceruloplasmin |
| chr7_-_19410749 | 4.54 |
ENSMUST00000003074.16
|
Apoc2
|
apolipoprotein C-II |
| chr7_-_13571334 | 4.50 |
ENSMUST00000108522.5
|
Sult2a1
|
sulfotransferase family 2A, dehydroepiandrosterone (DHEA)-preferring, member 1 |
| chr6_-_119365632 | 4.47 |
ENSMUST00000169744.8
|
Adipor2
|
adiponectin receptor 2 |
| chr6_-_42350188 | 4.41 |
ENSMUST00000073387.5
ENSMUST00000204357.2 |
Epha1
|
Eph receptor A1 |
| chr8_+_55053809 | 4.39 |
ENSMUST00000033917.7
|
Spata4
|
spermatogenesis associated 4 |
| chr6_-_141892517 | 4.37 |
ENSMUST00000168119.8
|
Slco1a1
|
solute carrier organic anion transporter family, member 1a1 |
| chr15_+_10358611 | 4.36 |
ENSMUST00000110541.8
ENSMUST00000110542.8 |
Agxt2
|
alanine-glyoxylate aminotransferase 2 |
| chr9_-_71075939 | 4.31 |
ENSMUST00000113570.8
|
Aqp9
|
aquaporin 9 |
| chr7_+_13357892 | 4.28 |
ENSMUST00000108525.4
|
Sult2a5
|
sulfotransferase family 2A, dehydroepiandrosterone (DHEA)-preferring, member 5 |
| chr6_-_3988835 | 4.25 |
ENSMUST00000203257.2
|
Tfpi2
|
tissue factor pathway inhibitor 2 |
| chr4_+_115458172 | 4.22 |
ENSMUST00000084342.6
|
Cyp4a32
|
cytochrome P450, family 4, subfamily a, polypeptide 32 |
| chr12_+_87204374 | 4.19 |
ENSMUST00000222222.2
|
Gstz1
|
glutathione transferase zeta 1 (maleylacetoacetate isomerase) |
| chrX_+_56454507 | 4.19 |
ENSMUST00000114725.3
|
Tm9sf5
|
transmembrane 9 superfamily member 5 |
| chr7_+_25597045 | 4.18 |
ENSMUST00000072438.13
ENSMUST00000005477.6 |
Cyp2b10
|
cytochrome P450, family 2, subfamily b, polypeptide 10 |
| chr1_-_155688551 | 4.14 |
ENSMUST00000194632.2
ENSMUST00000111764.8 |
Qsox1
|
quiescin Q6 sulfhydryl oxidase 1 |
| chr13_-_47196607 | 4.10 |
ENSMUST00000124948.2
|
Tpmt
|
thiopurine methyltransferase |
| chr1_-_90897329 | 4.08 |
ENSMUST00000130042.2
ENSMUST00000027529.12 |
Rab17
|
RAB17, member RAS oncogene family |
| chr11_+_66915969 | 4.08 |
ENSMUST00000079077.12
ENSMUST00000061786.6 |
Tmem220
|
transmembrane protein 220 |
| chr16_-_22847808 | 4.06 |
ENSMUST00000115349.9
|
Kng2
|
kininogen 2 |
| chr7_-_30810422 | 4.02 |
ENSMUST00000039435.15
|
Hpn
|
hepsin |
| chr19_-_7779943 | 3.97 |
ENSMUST00000120522.8
|
Slc22a26
|
solute carrier family 22 (organic cation transporter), member 26 |
| chr11_-_77784922 | 3.95 |
ENSMUST00000017597.5
|
Pipox
|
pipecolic acid oxidase |
| chr3_+_81906768 | 3.94 |
ENSMUST00000107736.2
|
Asic5
|
acid-sensing (proton-gated) ion channel family member 5 |
| chr19_+_39499288 | 3.92 |
ENSMUST00000025968.5
|
Cyp2c39
|
cytochrome P450, family 2, subfamily c, polypeptide 39 |
| chr16_-_22847760 | 3.91 |
ENSMUST00000039338.13
|
Kng2
|
kininogen 2 |
| chr4_+_115420876 | 3.87 |
ENSMUST00000126645.8
ENSMUST00000030480.4 |
Cyp4a31
|
cytochrome P450, family 4, subfamily a, polypeptide 31 |
| chr1_+_175459559 | 3.84 |
ENSMUST00000040250.15
|
Kmo
|
kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) |
| chr9_+_78164402 | 3.83 |
ENSMUST00000217203.2
|
Gm3776
|
predicted gene 3776 |
| chr5_-_130053120 | 3.75 |
ENSMUST00000161640.8
ENSMUST00000161884.2 ENSMUST00000161094.8 |
Asl
|
argininosuccinate lyase |
| chr3_-_146321341 | 3.75 |
ENSMUST00000200633.2
|
Dnase2b
|
deoxyribonuclease II beta |
| chr5_-_87686048 | 3.70 |
ENSMUST00000031199.11
|
Sult1b1
|
sulfotransferase family 1B, member 1 |
| chr10_-_75616788 | 3.68 |
ENSMUST00000173512.2
ENSMUST00000173537.2 |
Gm20441
Gstt3
|
predicted gene 20441 glutathione S-transferase, theta 3 |
| chr7_+_100966289 | 3.66 |
ENSMUST00000163799.9
ENSMUST00000164479.9 |
Stard10
|
START domain containing 10 |
| chr3_+_122688721 | 3.64 |
ENSMUST00000023820.6
|
Fabp2
|
fatty acid binding protein 2, intestinal |
| chr6_-_55152002 | 3.63 |
ENSMUST00000003569.6
|
Inmt
|
indolethylamine N-methyltransferase |
| chr13_+_4283729 | 3.62 |
ENSMUST00000081326.7
|
Akr1c19
|
aldo-keto reductase family 1, member C19 |
| chr6_+_138117519 | 3.60 |
ENSMUST00000120939.8
ENSMUST00000204628.3 ENSMUST00000140932.2 ENSMUST00000120302.8 |
Mgst1
|
microsomal glutathione S-transferase 1 |
| chr16_-_22847829 | 3.58 |
ENSMUST00000100046.9
|
Kng2
|
kininogen 2 |
| chr1_-_121260274 | 3.58 |
ENSMUST00000161068.2
|
Insig2
|
insulin induced gene 2 |
| chr11_-_110142565 | 3.57 |
ENSMUST00000044003.14
|
Abca6
|
ATP-binding cassette, sub-family A (ABC1), member 6 |
| chr17_-_47732798 | 3.56 |
ENSMUST00000073143.7
|
1700001C19Rik
|
RIKEN cDNA 1700001C19 gene |
| chr1_+_88093726 | 3.49 |
ENSMUST00000097659.5
|
Ugt1a5
|
UDP glucuronosyltransferase 1 family, polypeptide A5 |
| chr1_-_155688635 | 3.47 |
ENSMUST00000035325.15
|
Qsox1
|
quiescin Q6 sulfhydryl oxidase 1 |
| chrX_+_149330371 | 3.46 |
ENSMUST00000066337.13
ENSMUST00000112715.2 |
Alas2
|
aminolevulinic acid synthase 2, erythroid |
| chr15_+_10358643 | 3.45 |
ENSMUST00000022858.8
|
Agxt2
|
alanine-glyoxylate aminotransferase 2 |
| chr16_-_19385631 | 3.44 |
ENSMUST00000090062.2
|
Olfr169
|
olfactory receptor 169 |
| chr4_+_60003438 | 3.42 |
ENSMUST00000107517.8
ENSMUST00000107520.2 |
Mup6
|
major urinary protein 6 |
| chr5_-_139799953 | 3.40 |
ENSMUST00000044002.10
|
Tmem184a
|
transmembrane protein 184a |
| chr1_+_175459735 | 3.38 |
ENSMUST00000097458.4
|
Kmo
|
kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) |
| chr10_-_24712034 | 3.35 |
ENSMUST00000218044.2
ENSMUST00000020169.9 |
Enpp3
|
ectonucleotide pyrophosphatase/phosphodiesterase 3 |
| chr5_-_105491795 | 3.34 |
ENSMUST00000171587.2
|
Gbp11
|
guanylate binding protein 11 |
| chr11_-_32774562 | 3.30 |
ENSMUST00000109365.8
ENSMUST00000020508.4 |
Smim23
|
small integral membrane protein 23 |
| chr6_-_57957795 | 3.29 |
ENSMUST00000176572.3
|
Vmn1r25
|
vomeronasal 1 receptor 25 |
| chr1_-_162641495 | 3.28 |
ENSMUST00000144916.8
ENSMUST00000140274.2 |
Fmo4
|
flavin containing monooxygenase 4 |
| chr4_+_115420817 | 3.28 |
ENSMUST00000141033.8
ENSMUST00000030486.15 |
Cyp4a31
|
cytochrome P450, family 4, subfamily a, polypeptide 31 |
| chr5_-_87402659 | 3.27 |
ENSMUST00000075858.4
|
Ugt2b37
|
UDP glucuronosyltransferase 2 family, polypeptide B37 |
| chr15_+_77361241 | 3.26 |
ENSMUST00000060551.9
|
Apol10a
|
apolipoprotein L 10A |
| chr5_+_45650821 | 3.25 |
ENSMUST00000198534.2
|
Lap3
|
leucine aminopeptidase 3 |
| chr11_-_83469446 | 3.25 |
ENSMUST00000019266.6
|
Ccl9
|
chemokine (C-C motif) ligand 9 |
| chr7_+_25760922 | 3.21 |
ENSMUST00000005669.9
|
Cyp2b13
|
cytochrome P450, family 2, subfamily b, polypeptide 13 |
| chr3_-_113367891 | 3.20 |
ENSMUST00000142505.9
|
Amy1
|
amylase 1, salivary |
| chrX_+_153264998 | 3.19 |
ENSMUST00000026029.2
|
Samt4
|
spermatogenesis associated multipass transmembrane protein 4 |
| chr2_-_112089627 | 3.16 |
ENSMUST00000043970.2
|
Nutm1
|
NUT midline carcinoma, family member 1 |
| chr3_-_14843512 | 3.14 |
ENSMUST00000094365.11
|
Car1
|
carbonic anhydrase 1 |
| chr3_+_94840352 | 3.13 |
ENSMUST00000090839.12
|
Selenbp1
|
selenium binding protein 1 |
| chr1_+_134313676 | 3.11 |
ENSMUST00000162187.2
|
Mgat4f
|
MGAT4 family, member F |
| chr5_+_135703426 | 3.06 |
ENSMUST00000153515.8
|
Por
|
P450 (cytochrome) oxidoreductase |
| chr18_+_84869456 | 3.00 |
ENSMUST00000160180.9
|
Cyb5a
|
cytochrome b5 type A (microsomal) |
| chr14_+_66207163 | 2.98 |
ENSMUST00000153460.8
|
Clu
|
clusterin |
| chr3_+_62245765 | 2.97 |
ENSMUST00000079300.13
|
Arhgef26
|
Rho guanine nucleotide exchange factor (GEF) 26 |
| chr18_+_51250748 | 2.96 |
ENSMUST00000116639.4
|
Prr16
|
proline rich 16 |
| chr7_-_6489812 | 2.92 |
ENSMUST00000086318.5
ENSMUST00000209866.4 |
Olfr5
|
olfactory receptor 5 |
| chr11_-_11848107 | 2.88 |
ENSMUST00000178704.8
|
Ddc
|
dopa decarboxylase |
| chr16_+_16031182 | 2.80 |
ENSMUST00000039408.3
|
Pkp2
|
plakophilin 2 |
| chr1_+_180878797 | 2.76 |
ENSMUST00000036819.7
|
9130409I23Rik
|
RIKEN cDNA 9130409I23 gene |
| chr11_-_109986804 | 2.74 |
ENSMUST00000100287.9
|
Abca8a
|
ATP-binding cassette, sub-family A (ABC1), member 8a |
| chr2_-_62242562 | 2.73 |
ENSMUST00000047812.8
|
Dpp4
|
dipeptidylpeptidase 4 |
| chr9_+_18389653 | 2.71 |
ENSMUST00000213625.2
ENSMUST00000069218.5 |
Mbd3l1
|
methyl-CpG binding domain protein 3-like 1 |
| chr17_+_7592045 | 2.70 |
ENSMUST00000095726.11
ENSMUST00000128533.8 ENSMUST00000129709.8 ENSMUST00000147803.8 ENSMUST00000140192.8 ENSMUST00000138222.8 ENSMUST00000144861.2 |
Tcp10a
|
t-complex protein 10a |
| chr19_-_4489415 | 2.69 |
ENSMUST00000235680.2
ENSMUST00000117462.2 ENSMUST00000048197.10 |
Rhod
|
ras homolog family member D |
| chr1_+_177635997 | 2.68 |
ENSMUST00000194528.3
|
Catspere1
|
cation channel sperm associated auxiliary subunit epsilon 1 |
| chr11_-_89893707 | 2.67 |
ENSMUST00000020864.9
|
Pctp
|
phosphatidylcholine transfer protein |
| chr8_-_43981143 | 2.67 |
ENSMUST00000080135.5
|
Adam26b
|
a disintegrin and metallopeptidase domain 26B |
| chr19_-_4092218 | 2.65 |
ENSMUST00000237999.2
ENSMUST00000042700.12 |
Gstp2
|
glutathione S-transferase, pi 2 |
| chr1_-_37996838 | 2.62 |
ENSMUST00000027254.10
ENSMUST00000114894.2 |
Lyg1
|
lysozyme G-like 1 |
| chrX_-_67936659 | 2.58 |
ENSMUST00000074894.2
|
Gm6812
|
predicted gene 6812 |
| chr13_+_4278681 | 2.58 |
ENSMUST00000118663.9
|
Akr1c19
|
aldo-keto reductase family 1, member C19 |
| chr4_+_39450265 | 2.57 |
ENSMUST00000029955.5
|
1700009N14Rik
|
RIKEN cDNA 1700009N14 gene |
| chr2_-_111104451 | 2.57 |
ENSMUST00000214760.2
|
Olfr1277
|
olfactory receptor 1277 |
| chr1_+_88022776 | 2.56 |
ENSMUST00000150634.8
ENSMUST00000058237.14 |
Ugt1a7c
|
UDP glucuronosyltransferase 1 family, polypeptide A7C |
| chr8_+_56747613 | 2.56 |
ENSMUST00000034026.10
|
Hpgd
|
hydroxyprostaglandin dehydrogenase 15 (NAD) |
| chr19_-_12100560 | 2.56 |
ENSMUST00000213471.2
ENSMUST00000214676.2 |
Olfr76
|
olfactory receptor 76 |
| chr7_+_143420586 | 2.55 |
ENSMUST00000152910.3
ENSMUST00000207630.2 |
Acte1
|
actin, epsilon 1 |
| chr18_+_44334331 | 2.53 |
ENSMUST00000097586.3
|
Gm10542
|
predicted gene 10542 |
| chr2_+_89757653 | 2.52 |
ENSMUST00000213720.3
ENSMUST00000102609.3 |
Olfr1258
|
olfactory receptor 1258 |
| chr1_+_87998487 | 2.51 |
ENSMUST00000073772.5
|
Ugt1a9
|
UDP glucuronosyltransferase 1 family, polypeptide A9 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 11.4 | 34.3 | GO:0018879 | biphenyl metabolic process(GO:0018879) |
| 10.8 | 32.5 | GO:0009753 | response to jasmonic acid(GO:0009753) cellular response to jasmonic acid stimulus(GO:0071395) |
| 8.6 | 25.7 | GO:0009073 | tyrosine biosynthetic process(GO:0006571) aromatic amino acid family biosynthetic process(GO:0009073) aromatic amino acid family biosynthetic process, prephenate pathway(GO:0009095) |
| 7.8 | 23.3 | GO:0042450 | arginine biosynthetic process via ornithine(GO:0042450) |
| 6.7 | 20.2 | GO:0043387 | mycotoxin catabolic process(GO:0043387) aflatoxin catabolic process(GO:0046223) organic heteropentacyclic compound catabolic process(GO:1901377) |
| 5.3 | 15.9 | GO:2000863 | positive regulation of estrogen secretion(GO:2000863) |
| 3.9 | 129.1 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 3.7 | 11.1 | GO:0033306 | phytol metabolic process(GO:0033306) fatty alcohol metabolic process(GO:1903173) |
| 3.7 | 14.7 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 3.5 | 17.7 | GO:0051919 | positive regulation of fibrinolysis(GO:0051919) |
| 3.4 | 47.0 | GO:0052695 | cellular glucuronidation(GO:0052695) |
| 3.3 | 13.1 | GO:0006069 | ethanol oxidation(GO:0006069) |
| 3.0 | 9.0 | GO:0019626 | short-chain fatty acid catabolic process(GO:0019626) |
| 2.6 | 7.8 | GO:0042853 | glycine biosynthetic process, by transamination of glyoxylate(GO:0019265) L-alanine catabolic process(GO:0042853) |
| 2.6 | 10.4 | GO:0001970 | positive regulation of activation of membrane attack complex(GO:0001970) |
| 2.1 | 19.0 | GO:0006559 | L-phenylalanine catabolic process(GO:0006559) tyrosine catabolic process(GO:0006572) erythrose 4-phosphate/phosphoenolpyruvate family amino acid catabolic process(GO:1902222) |
| 1.9 | 21.3 | GO:0071394 | cellular response to testosterone stimulus(GO:0071394) |
| 1.9 | 25.0 | GO:0072378 | blood coagulation, fibrin clot formation(GO:0072378) |
| 1.9 | 7.5 | GO:0051344 | negative regulation of cyclic-nucleotide phosphodiesterase activity(GO:0051344) regulation of aminoacyl-tRNA ligase activity(GO:1903630) |
| 1.8 | 5.4 | GO:0090320 | chylomicron assembly(GO:0034378) regulation of chylomicron remnant clearance(GO:0090320) |
| 1.7 | 10.2 | GO:0008355 | olfactory learning(GO:0008355) |
| 1.6 | 4.7 | GO:0009812 | flavonoid metabolic process(GO:0009812) |
| 1.5 | 7.4 | GO:0009115 | xanthine catabolic process(GO:0009115) |
| 1.4 | 5.5 | GO:0010609 | mRNA localization resulting in posttranscriptional regulation of gene expression(GO:0010609) |
| 1.4 | 4.1 | GO:0002386 | immune response in mucosal-associated lymphoid tissue(GO:0002386) immunoglobulin transcytosis in epithelial cells(GO:0002414) |
| 1.3 | 4.0 | GO:0034769 | basement membrane disassembly(GO:0034769) |
| 1.3 | 3.9 | GO:0006553 | lysine metabolic process(GO:0006553) |
| 1.3 | 6.3 | GO:0019482 | beta-alanine metabolic process(GO:0019482) |
| 1.2 | 7.2 | GO:0019805 | quinolinate biosynthetic process(GO:0019805) |
| 1.1 | 4.5 | GO:0060697 | positive regulation of phospholipid catabolic process(GO:0060697) |
| 1.1 | 6.6 | GO:0008295 | spermidine biosynthetic process(GO:0008295) |
| 1.1 | 2.2 | GO:0090206 | negative regulation of cholesterol biosynthetic process(GO:0045541) negative regulation of cholesterol metabolic process(GO:0090206) |
| 1.1 | 4.3 | GO:0015855 | nucleobase transport(GO:0015851) pyrimidine nucleobase transport(GO:0015855) |
| 1.0 | 7.9 | GO:0000270 | peptidoglycan metabolic process(GO:0000270) peptidoglycan catabolic process(GO:0009253) |
| 0.9 | 1.8 | GO:0001969 | activation of membrane attack complex(GO:0001905) regulation of activation of membrane attack complex(GO:0001969) |
| 0.9 | 3.6 | GO:0071449 | cellular response to lipid hydroperoxide(GO:0071449) |
| 0.8 | 2.4 | GO:0002149 | hypochlorous acid metabolic process(GO:0002148) hypochlorous acid biosynthetic process(GO:0002149) |
| 0.8 | 3.1 | GO:0046210 | nitrate catabolic process(GO:0043602) nitric oxide catabolic process(GO:0046210) cellular organohalogen metabolic process(GO:0090345) cellular organofluorine metabolic process(GO:0090346) |
| 0.8 | 2.3 | GO:1902037 | negative regulation of hematopoietic stem cell differentiation(GO:1902037) |
| 0.7 | 11.2 | GO:0015747 | urate transport(GO:0015747) |
| 0.7 | 20.3 | GO:0035634 | response to stilbenoid(GO:0035634) |
| 0.7 | 5.1 | GO:0050747 | positive regulation of lipoprotein metabolic process(GO:0050747) |
| 0.7 | 2.1 | GO:0071211 | protein targeting to vacuole involved in autophagy(GO:0071211) |
| 0.7 | 1.4 | GO:0046226 | coumarin catabolic process(GO:0046226) |
| 0.7 | 6.9 | GO:0015712 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.7 | 10.7 | GO:0032000 | positive regulation of fatty acid beta-oxidation(GO:0032000) |
| 0.7 | 4.6 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.6 | 13.5 | GO:0042359 | vitamin D metabolic process(GO:0042359) |
| 0.6 | 6.7 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.6 | 6.6 | GO:0000255 | allantoin metabolic process(GO:0000255) |
| 0.6 | 4.0 | GO:0015842 | aminergic neurotransmitter loading into synaptic vesicle(GO:0015842) |
| 0.6 | 3.4 | GO:0018992 | germ-line sex determination(GO:0018992) |
| 0.6 | 31.2 | GO:0030195 | negative regulation of blood coagulation(GO:0030195) negative regulation of hemostasis(GO:1900047) |
| 0.6 | 1.7 | GO:0010482 | epidermal cell division(GO:0010481) regulation of epidermal cell division(GO:0010482) |
| 0.6 | 6.6 | GO:0034626 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.5 | 3.3 | GO:0042737 | drug catabolic process(GO:0042737) |
| 0.5 | 2.7 | GO:0036343 | psychomotor behavior(GO:0036343) |
| 0.5 | 8.1 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.5 | 4.8 | GO:0032286 | central nervous system myelin maintenance(GO:0032286) |
| 0.5 | 1.5 | GO:1990535 | neuron projection maintenance(GO:1990535) |
| 0.5 | 18.1 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.5 | 6.0 | GO:0070863 | positive regulation of protein exit from endoplasmic reticulum(GO:0070863) |
| 0.5 | 2.3 | GO:0006538 | glutamate catabolic process(GO:0006538) |
| 0.4 | 4.5 | GO:0061042 | vascular wound healing(GO:0061042) |
| 0.4 | 1.7 | GO:0010982 | regulation of high-density lipoprotein particle clearance(GO:0010982) positive regulation of lipoprotein particle clearance(GO:0010986) |
| 0.4 | 4.2 | GO:0045198 | establishment of epithelial cell apical/basal polarity(GO:0045198) |
| 0.4 | 2.5 | GO:0036112 | medium-chain fatty-acyl-CoA metabolic process(GO:0036112) |
| 0.4 | 0.4 | GO:0009155 | purine deoxyribonucleotide catabolic process(GO:0009155) |
| 0.4 | 1.1 | GO:0098501 | polynucleotide dephosphorylation(GO:0098501) |
| 0.4 | 1.9 | GO:0042412 | taurine biosynthetic process(GO:0042412) |
| 0.4 | 4.1 | GO:0070574 | cadmium ion transport(GO:0015691) cadmium ion transmembrane transport(GO:0070574) |
| 0.4 | 1.5 | GO:0048388 | endosomal lumen acidification(GO:0048388) |
| 0.4 | 2.2 | GO:0002528 | regulation of vascular permeability involved in acute inflammatory response(GO:0002528) |
| 0.4 | 3.7 | GO:0035360 | positive regulation of peroxisome proliferator activated receptor signaling pathway(GO:0035360) |
| 0.4 | 2.2 | GO:0014016 | neuroblast differentiation(GO:0014016) |
| 0.4 | 1.1 | GO:1904344 | regulation of gastric mucosal blood circulation(GO:1904344) positive regulation of gastric mucosal blood circulation(GO:1904346) gastric mucosal blood circulation(GO:1990768) |
| 0.3 | 1.7 | GO:0002225 | positive regulation of antimicrobial peptide production(GO:0002225) positive regulation of antibacterial peptide production(GO:0002803) |
| 0.3 | 1.0 | GO:0006550 | isoleucine metabolic process(GO:0006549) isoleucine catabolic process(GO:0006550) |
| 0.3 | 4.2 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.3 | 1.0 | GO:0032474 | otolith morphogenesis(GO:0032474) |
| 0.3 | 2.0 | GO:0002430 | complement receptor mediated signaling pathway(GO:0002430) |
| 0.3 | 1.3 | GO:0051005 | negative regulation of lipoprotein lipase activity(GO:0051005) |
| 0.3 | 1.0 | GO:0030221 | basophil differentiation(GO:0030221) |
| 0.3 | 2.2 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 0.3 | 1.0 | GO:0021524 | visceral motor neuron differentiation(GO:0021524) |
| 0.3 | 1.8 | GO:0009624 | response to nematode(GO:0009624) |
| 0.3 | 0.6 | GO:1903070 | negative regulation of ER-associated ubiquitin-dependent protein catabolic process(GO:1903070) |
| 0.3 | 0.9 | GO:0002355 | detection of tumor cell(GO:0002355) |
| 0.3 | 6.0 | GO:0045717 | negative regulation of fatty acid biosynthetic process(GO:0045717) |
| 0.3 | 2.1 | GO:1901475 | pyruvate transmembrane transport(GO:1901475) |
| 0.3 | 0.9 | GO:0014737 | positive regulation of transcription from RNA polymerase II promoter involved in unfolded protein response(GO:0006990) positive regulation of muscle atrophy(GO:0014737) |
| 0.3 | 1.7 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 0.3 | 2.0 | GO:0060468 | prevention of polyspermy(GO:0060468) |
| 0.3 | 0.6 | GO:0016561 | protein import into peroxisome matrix, translocation(GO:0016561) |
| 0.3 | 1.6 | GO:0051410 | detoxification of nitrogen compound(GO:0051410) cellular detoxification of nitrogen compound(GO:0070458) |
| 0.2 | 1.0 | GO:0090472 | negative regulation of low-density lipoprotein particle receptor catabolic process(GO:0032804) dibasic protein processing(GO:0090472) |
| 0.2 | 0.7 | GO:1904736 | negative regulation of electron carrier activity(GO:1904733) regulation of fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:1904735) negative regulation of fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:1904736) |
| 0.2 | 3.9 | GO:0016998 | cell wall macromolecule catabolic process(GO:0016998) |
| 0.2 | 1.7 | GO:0060309 | elastin catabolic process(GO:0060309) |
| 0.2 | 3.1 | GO:0007567 | parturition(GO:0007567) |
| 0.2 | 0.7 | GO:0042726 | flavin-containing compound metabolic process(GO:0042726) |
| 0.2 | 4.5 | GO:0051923 | sulfation(GO:0051923) |
| 0.2 | 9.9 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.2 | 10.8 | GO:0006953 | acute-phase response(GO:0006953) |
| 0.2 | 0.7 | GO:0051715 | cytolysis in other organism(GO:0051715) |
| 0.2 | 1.9 | GO:0051799 | negative regulation of hair follicle development(GO:0051799) |
| 0.2 | 0.7 | GO:0061763 | multivesicular body-lysosome fusion(GO:0061763) |
| 0.2 | 3.4 | GO:0019321 | pentose metabolic process(GO:0019321) |
| 0.2 | 0.7 | GO:0035720 | intraciliary anterograde transport(GO:0035720) |
| 0.2 | 0.9 | GO:0060785 | regulation of apoptosis involved in tissue homeostasis(GO:0060785) |
| 0.2 | 3.3 | GO:0009143 | nucleoside triphosphate catabolic process(GO:0009143) |
| 0.2 | 3.5 | GO:0042541 | hemoglobin biosynthetic process(GO:0042541) |
| 0.2 | 0.6 | GO:0090365 | regulation of mRNA modification(GO:0090365) negative regulation of mRNA modification(GO:0090367) |
| 0.2 | 250.3 | GO:0007608 | sensory perception of smell(GO:0007608) |
| 0.2 | 0.6 | GO:1900247 | cytoplasmic translational elongation(GO:0002182) regulation of cytoplasmic translational elongation(GO:1900247) negative regulation of cytoplasmic translational elongation(GO:1900248) |
| 0.2 | 0.4 | GO:0046122 | dGTP metabolic process(GO:0046070) purine deoxyribonucleoside metabolic process(GO:0046122) |
| 0.2 | 0.8 | GO:0016332 | establishment or maintenance of polarity of embryonic epithelium(GO:0016332) |
| 0.2 | 3.7 | GO:1900003 | regulation of serine-type endopeptidase activity(GO:1900003) negative regulation of serine-type endopeptidase activity(GO:1900004) regulation of serine-type peptidase activity(GO:1902571) negative regulation of serine-type peptidase activity(GO:1902572) |
| 0.2 | 1.2 | GO:0007468 | regulation of rhodopsin gene expression(GO:0007468) |
| 0.2 | 1.4 | GO:0051940 | regulation of dopamine uptake involved in synaptic transmission(GO:0051584) regulation of catecholamine uptake involved in synaptic transmission(GO:0051940) chaperone-mediated protein transport(GO:0072321) |
| 0.2 | 1.8 | GO:0006682 | galactosylceramide biosynthetic process(GO:0006682) galactolipid biosynthetic process(GO:0019375) |
| 0.2 | 2.0 | GO:0010886 | positive regulation of cholesterol storage(GO:0010886) |
| 0.2 | 1.0 | GO:0006545 | glycine biosynthetic process(GO:0006545) |
| 0.2 | 0.8 | GO:0061187 | histone H4-K20 demethylation(GO:0035574) regulation of chromatin silencing at rDNA(GO:0061187) negative regulation of chromatin silencing at rDNA(GO:0061188) |
| 0.2 | 1.4 | GO:0015919 | peroxisomal membrane transport(GO:0015919) protein import into peroxisome membrane(GO:0045046) |
| 0.2 | 0.6 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.2 | 0.8 | GO:0070173 | regulation of enamel mineralization(GO:0070173) |
| 0.2 | 0.6 | GO:0010845 | positive regulation of reciprocal meiotic recombination(GO:0010845) |
| 0.2 | 0.8 | GO:0015755 | fructose transport(GO:0015755) |
| 0.2 | 0.9 | GO:0043056 | forward locomotion(GO:0043056) |
| 0.2 | 0.5 | GO:0035606 | peptidyl-cysteine S-trans-nitrosylation(GO:0035606) |
| 0.2 | 1.1 | GO:0070895 | transposon integration(GO:0070893) regulation of transposon integration(GO:0070894) negative regulation of transposon integration(GO:0070895) |
| 0.2 | 2.0 | GO:0015879 | carnitine transport(GO:0015879) |
| 0.2 | 1.4 | GO:0007341 | penetration of zona pellucida(GO:0007341) |
| 0.2 | 0.7 | GO:0044339 | canonical Wnt signaling pathway involved in osteoblast differentiation(GO:0044339) |
| 0.2 | 0.5 | GO:0060605 | tube lumen cavitation(GO:0060605) salivary gland cavitation(GO:0060662) |
| 0.2 | 1.0 | GO:0090274 | reduction of food intake in response to dietary excess(GO:0002023) positive regulation of somatostatin secretion(GO:0090274) |
| 0.2 | 1.0 | GO:2000124 | regulation of endocannabinoid signaling pathway(GO:2000124) |
| 0.2 | 1.2 | GO:0007262 | STAT protein import into nucleus(GO:0007262) |
| 0.2 | 3.7 | GO:0006309 | apoptotic DNA fragmentation(GO:0006309) |
| 0.2 | 0.9 | GO:0042744 | hydrogen peroxide catabolic process(GO:0042744) |
| 0.2 | 13.6 | GO:0007566 | embryo implantation(GO:0007566) |
| 0.2 | 0.7 | GO:0006027 | glycosaminoglycan catabolic process(GO:0006027) |
| 0.2 | 2.0 | GO:2000189 | positive regulation of cholesterol homeostasis(GO:2000189) |
| 0.2 | 0.9 | GO:0060474 | positive regulation of sperm motility involved in capacitation(GO:0060474) |
| 0.2 | 0.5 | GO:0035425 | autocrine signaling(GO:0035425) paracrine signaling(GO:0038001) |
| 0.2 | 0.6 | GO:0060598 | dichotomous subdivision of terminal units involved in mammary gland duct morphogenesis(GO:0060598) |
| 0.2 | 4.8 | GO:0046688 | response to copper ion(GO:0046688) |
| 0.2 | 0.9 | GO:0006105 | succinate metabolic process(GO:0006105) |
| 0.2 | 1.7 | GO:0036444 | calcium ion transmembrane import into mitochondrion(GO:0036444) |
| 0.2 | 0.6 | GO:0022615 | protein to membrane docking(GO:0022615) |
| 0.1 | 1.0 | GO:0019364 | pyridine nucleotide catabolic process(GO:0019364) |
| 0.1 | 0.7 | GO:0046449 | creatinine metabolic process(GO:0046449) |
| 0.1 | 0.9 | GO:0061732 | mitochondrial acetyl-CoA biosynthetic process from pyruvate(GO:0061732) |
| 0.1 | 0.6 | GO:1904158 | axonemal central apparatus assembly(GO:1904158) |
| 0.1 | 5.3 | GO:0050892 | intestinal absorption(GO:0050892) |
| 0.1 | 2.2 | GO:0030259 | lipid glycosylation(GO:0030259) |
| 0.1 | 0.8 | GO:0051572 | negative regulation of histone H3-K4 methylation(GO:0051572) |
| 0.1 | 0.7 | GO:0090370 | negative regulation of cholesterol efflux(GO:0090370) |
| 0.1 | 2.2 | GO:0006098 | pentose-phosphate shunt(GO:0006098) |
| 0.1 | 0.4 | GO:0045645 | regulation of eosinophil differentiation(GO:0045643) positive regulation of eosinophil differentiation(GO:0045645) |
| 0.1 | 0.7 | GO:1901492 | positive regulation of lymphangiogenesis(GO:1901492) |
| 0.1 | 0.5 | GO:0045590 | negative regulation of regulatory T cell differentiation(GO:0045590) |
| 0.1 | 0.4 | GO:0003127 | detection of nodal flow(GO:0003127) detection of endogenous stimulus(GO:0009726) |
| 0.1 | 3.0 | GO:0045793 | positive regulation of cell size(GO:0045793) |
| 0.1 | 2.6 | GO:0000054 | ribosomal subunit export from nucleus(GO:0000054) ribosome localization(GO:0033750) establishment of ribosome localization(GO:0033753) |
| 0.1 | 1.4 | GO:0070244 | negative regulation of thymocyte apoptotic process(GO:0070244) interleukin-17 secretion(GO:0072615) |
| 0.1 | 1.7 | GO:0042761 | very long-chain fatty acid biosynthetic process(GO:0042761) |
| 0.1 | 0.9 | GO:0070309 | lens fiber cell morphogenesis(GO:0070309) |
| 0.1 | 0.9 | GO:0098532 | histone H3-K27 trimethylation(GO:0098532) |
| 0.1 | 0.9 | GO:0003383 | apical constriction(GO:0003383) |
| 0.1 | 0.5 | GO:1901162 | serotonin biosynthetic process(GO:0042427) primary amino compound biosynthetic process(GO:1901162) |
| 0.1 | 1.8 | GO:1903546 | protein localization to photoreceptor outer segment(GO:1903546) |
| 0.1 | 1.6 | GO:0070633 | transepithelial transport(GO:0070633) |
| 0.1 | 0.3 | GO:0042126 | nitrate metabolic process(GO:0042126) nitrate assimilation(GO:0042128) |
| 0.1 | 0.4 | GO:2000321 | positive regulation of T-helper 17 cell differentiation(GO:2000321) |
| 0.1 | 3.7 | GO:0006346 | methylation-dependent chromatin silencing(GO:0006346) |
| 0.1 | 0.9 | GO:0035093 | spermatogenesis, exchange of chromosomal proteins(GO:0035093) |
| 0.1 | 1.1 | GO:0042640 | anagen(GO:0042640) |
| 0.1 | 0.5 | GO:0070272 | proton-transporting ATP synthase complex assembly(GO:0043461) proton-transporting ATP synthase complex biogenesis(GO:0070272) |
| 0.1 | 7.8 | GO:0045454 | cell redox homeostasis(GO:0045454) |
| 0.1 | 1.3 | GO:0046007 | negative regulation of activated T cell proliferation(GO:0046007) |
| 0.1 | 1.7 | GO:0043518 | negative regulation of DNA damage response, signal transduction by p53 class mediator(GO:0043518) |
| 0.1 | 8.5 | GO:0019395 | fatty acid oxidation(GO:0019395) |
| 0.1 | 7.3 | GO:0006749 | glutathione metabolic process(GO:0006749) |
| 0.1 | 0.7 | GO:0046010 | positive regulation of circadian sleep/wake cycle, non-REM sleep(GO:0046010) |
| 0.1 | 0.2 | GO:0060734 | regulation of endoplasmic reticulum stress-induced eIF2 alpha phosphorylation(GO:0060734) |
| 0.1 | 2.8 | GO:0045475 | locomotor rhythm(GO:0045475) |
| 0.1 | 3.3 | GO:0045662 | negative regulation of myoblast differentiation(GO:0045662) |
| 0.1 | 2.1 | GO:0035404 | histone-serine phosphorylation(GO:0035404) |
| 0.1 | 0.3 | GO:0070212 | protein poly-ADP-ribosylation(GO:0070212) regulation of telomeric DNA binding(GO:1904742) |
| 0.1 | 28.5 | GO:0008202 | steroid metabolic process(GO:0008202) |
| 0.1 | 0.3 | GO:0021682 | nerve maturation(GO:0021682) |
| 0.1 | 0.8 | GO:0036371 | protein localization to M-band(GO:0036309) protein localization to T-tubule(GO:0036371) |
| 0.1 | 1.2 | GO:0006625 | protein targeting to peroxisome(GO:0006625) peroxisomal transport(GO:0043574) protein localization to peroxisome(GO:0072662) establishment of protein localization to peroxisome(GO:0072663) |
| 0.1 | 2.0 | GO:0030261 | chromosome condensation(GO:0030261) |
| 0.1 | 1.0 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.1 | 0.5 | GO:0090168 | Golgi reassembly(GO:0090168) |
| 0.1 | 0.2 | GO:0034147 | regulation of granuloma formation(GO:0002631) negative regulation of granuloma formation(GO:0002632) regulation of toll-like receptor 5 signaling pathway(GO:0034147) negative regulation of toll-like receptor 5 signaling pathway(GO:0034148) negative regulation of nucleotide-binding oligomerization domain containing 1 signaling pathway(GO:0070429) tolerance induction to lipopolysaccharide(GO:0072573) |
| 0.1 | 0.7 | GO:0032790 | ribosome disassembly(GO:0032790) |
| 0.1 | 0.9 | GO:0060510 | Type II pneumocyte differentiation(GO:0060510) |
| 0.1 | 1.2 | GO:0032825 | positive regulation of natural killer cell differentiation(GO:0032825) |
| 0.1 | 0.8 | GO:2000739 | mesenchymal stem cell differentiation(GO:0072497) regulation of mesenchymal stem cell differentiation(GO:2000739) positive regulation of mesenchymal stem cell differentiation(GO:2000741) |
| 0.1 | 0.9 | GO:0000963 | mitochondrial RNA processing(GO:0000963) |
| 0.1 | 1.5 | GO:0006054 | N-acetylneuraminate metabolic process(GO:0006054) |
| 0.1 | 1.6 | GO:0046085 | adenosine metabolic process(GO:0046085) |
| 0.1 | 0.3 | GO:0031204 | posttranslational protein targeting to membrane, translocation(GO:0031204) |
| 0.1 | 0.8 | GO:0044334 | canonical Wnt signaling pathway involved in positive regulation of epithelial to mesenchymal transition(GO:0044334) |
| 0.1 | 0.2 | GO:1902630 | regulation of membrane hyperpolarization(GO:1902630) |
| 0.1 | 0.2 | GO:1904580 | regulation of intracellular mRNA localization(GO:1904580) positive regulation of intracellular mRNA localization(GO:1904582) |
| 0.1 | 0.6 | GO:0017196 | N-terminal peptidyl-methionine acetylation(GO:0017196) |
| 0.1 | 0.3 | GO:0021691 | cerebellar Purkinje cell layer maturation(GO:0021691) |
| 0.1 | 0.3 | GO:0001692 | histamine metabolic process(GO:0001692) |
| 0.1 | 1.1 | GO:0006977 | DNA damage response, signal transduction by p53 class mediator resulting in cell cycle arrest(GO:0006977) |
| 0.1 | 0.4 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.1 | 1.3 | GO:0002523 | leukocyte migration involved in inflammatory response(GO:0002523) |
| 0.1 | 3.6 | GO:0097178 | ruffle assembly(GO:0097178) |
| 0.1 | 1.3 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 0.1 | 0.2 | GO:0019509 | L-methionine biosynthetic process from methylthioadenosine(GO:0019509) |
| 0.1 | 3.1 | GO:0006687 | glycosphingolipid metabolic process(GO:0006687) |
| 0.1 | 0.4 | GO:0045716 | positive regulation of low-density lipoprotein particle receptor biosynthetic process(GO:0045716) |
| 0.1 | 0.4 | GO:0097646 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.1 | 0.6 | GO:0098703 | calcium ion import across plasma membrane(GO:0098703) calcium ion import into cell(GO:1990035) |
| 0.1 | 3.1 | GO:0050873 | brown fat cell differentiation(GO:0050873) |
| 0.1 | 0.3 | GO:0032510 | endosome to lysosome transport via multivesicular body sorting pathway(GO:0032510) |
| 0.1 | 0.9 | GO:0030502 | negative regulation of bone mineralization(GO:0030502) |
| 0.1 | 0.7 | GO:0001696 | gastric acid secretion(GO:0001696) |
| 0.1 | 0.4 | GO:0006561 | proline biosynthetic process(GO:0006561) |
| 0.1 | 2.0 | GO:0042832 | defense response to protozoan(GO:0042832) |
| 0.0 | 1.5 | GO:0039694 | viral RNA genome replication(GO:0039694) RNA replication(GO:0039703) |
| 0.0 | 0.2 | GO:0051661 | maintenance of centrosome location(GO:0051661) |
| 0.0 | 0.5 | GO:0034983 | peptidyl-lysine deacetylation(GO:0034983) |
| 0.0 | 0.8 | GO:1904778 | regulation of protein localization to cell cortex(GO:1904776) positive regulation of protein localization to cell cortex(GO:1904778) |
| 0.0 | 10.7 | GO:0006631 | fatty acid metabolic process(GO:0006631) |
| 0.0 | 1.3 | GO:1904355 | positive regulation of telomere capping(GO:1904355) |
| 0.0 | 0.8 | GO:0033008 | positive regulation of mast cell activation involved in immune response(GO:0033008) positive regulation of mast cell degranulation(GO:0043306) |
| 0.0 | 0.6 | GO:0021554 | optic nerve development(GO:0021554) |
| 0.0 | 0.3 | GO:2000628 | regulation of miRNA metabolic process(GO:2000628) |
| 0.0 | 0.3 | GO:0019348 | dolichol metabolic process(GO:0019348) |
| 0.0 | 0.5 | GO:0060576 | intestinal epithelial cell development(GO:0060576) |
| 0.0 | 0.3 | GO:0009162 | deoxyribonucleoside monophosphate metabolic process(GO:0009162) |
| 0.0 | 0.8 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.0 | 2.0 | GO:0006730 | one-carbon metabolic process(GO:0006730) |
| 0.0 | 1.0 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.0 | 1.7 | GO:0007032 | endosome organization(GO:0007032) |
| 0.0 | 0.5 | GO:0001731 | formation of translation preinitiation complex(GO:0001731) |
| 0.0 | 0.8 | GO:2001241 | positive regulation of extrinsic apoptotic signaling pathway in absence of ligand(GO:2001241) |
| 0.0 | 2.2 | GO:0051496 | positive regulation of stress fiber assembly(GO:0051496) |
| 0.0 | 1.0 | GO:0010763 | positive regulation of fibroblast migration(GO:0010763) |
| 0.0 | 0.7 | GO:0031163 | iron-sulfur cluster assembly(GO:0016226) metallo-sulfur cluster assembly(GO:0031163) |
| 0.0 | 0.6 | GO:0043950 | positive regulation of cAMP-mediated signaling(GO:0043950) |
| 0.0 | 0.8 | GO:0031295 | T cell costimulation(GO:0031295) |
| 0.0 | 0.4 | GO:0016446 | somatic hypermutation of immunoglobulin genes(GO:0016446) |
| 0.0 | 1.7 | GO:2000134 | negative regulation of G1/S transition of mitotic cell cycle(GO:2000134) |
| 0.0 | 0.4 | GO:0033603 | positive regulation of dopamine secretion(GO:0033603) |
| 0.0 | 0.4 | GO:0000290 | deadenylation-dependent decapping of nuclear-transcribed mRNA(GO:0000290) |
| 0.0 | 0.6 | GO:0007614 | short-term memory(GO:0007614) |
| 0.0 | 1.5 | GO:0000266 | mitochondrial fission(GO:0000266) |
| 0.0 | 2.2 | GO:0007218 | neuropeptide signaling pathway(GO:0007218) |
| 0.0 | 0.5 | GO:0051044 | positive regulation of membrane protein ectodomain proteolysis(GO:0051044) |
| 0.0 | 0.1 | GO:0090214 | spongiotrophoblast layer developmental growth(GO:0090214) |
| 0.0 | 0.5 | GO:0008535 | respiratory chain complex IV assembly(GO:0008535) |
| 0.0 | 0.5 | GO:0016322 | neuron remodeling(GO:0016322) |
| 0.0 | 0.2 | GO:0034214 | protein hexamerization(GO:0034214) |
| 0.0 | 0.3 | GO:0016540 | protein autoprocessing(GO:0016540) |
| 0.0 | 0.1 | GO:0048698 | negative regulation of collateral sprouting in absence of injury(GO:0048698) |
| 0.0 | 0.3 | GO:0009072 | aromatic amino acid family metabolic process(GO:0009072) |
| 0.0 | 0.1 | GO:0042776 | mitochondrial ATP synthesis coupled proton transport(GO:0042776) |
| 0.0 | 0.3 | GO:0006465 | signal peptide processing(GO:0006465) |
| 0.0 | 0.2 | GO:0050957 | equilibrioception(GO:0050957) |
| 0.0 | 0.6 | GO:0031069 | hair follicle morphogenesis(GO:0031069) |
| 0.0 | 0.8 | GO:0051567 | histone H3-K9 methylation(GO:0051567) |
| 0.0 | 0.3 | GO:0071712 | ER-associated misfolded protein catabolic process(GO:0071712) |
| 0.0 | 0.7 | GO:0000132 | establishment of mitotic spindle orientation(GO:0000132) |
| 0.0 | 0.2 | GO:0006517 | protein deglycosylation(GO:0006517) |
| 0.0 | 0.7 | GO:0051281 | positive regulation of release of sequestered calcium ion into cytosol(GO:0051281) |
| 0.0 | 0.1 | GO:0045409 | negative regulation of interleukin-6 biosynthetic process(GO:0045409) |
| 0.0 | 0.4 | GO:0007130 | synaptonemal complex assembly(GO:0007130) |
| 0.0 | 0.3 | GO:0042572 | retinol metabolic process(GO:0042572) |
| 0.0 | 0.3 | GO:0045741 | positive regulation of epidermal growth factor-activated receptor activity(GO:0045741) |
| 0.0 | 0.3 | GO:0006298 | mismatch repair(GO:0006298) |
| 0.0 | 0.1 | GO:2000467 | positive regulation of glycogen (starch) synthase activity(GO:2000467) |
| 0.0 | 0.7 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.0 | 6.4 | GO:0051321 | meiotic cell cycle(GO:0051321) |
| 0.0 | 1.1 | GO:0042698 | ovulation cycle(GO:0042698) |
| 0.0 | 1.0 | GO:0007586 | digestion(GO:0007586) |
| 0.0 | 0.3 | GO:0070584 | mitochondrion morphogenesis(GO:0070584) |
| 0.0 | 0.6 | GO:0051497 | negative regulation of stress fiber assembly(GO:0051497) |
| 0.0 | 0.5 | GO:0031648 | protein destabilization(GO:0031648) |
| 0.0 | 0.6 | GO:0051180 | vitamin transport(GO:0051180) |
| 0.0 | 0.3 | GO:0051560 | mitochondrial calcium ion homeostasis(GO:0051560) |
| 0.0 | 0.1 | GO:0070131 | positive regulation of mitochondrial translation(GO:0070131) |
| 0.0 | 1.2 | GO:0050728 | negative regulation of inflammatory response(GO:0050728) |
| 0.0 | 0.5 | GO:0030513 | positive regulation of BMP signaling pathway(GO:0030513) |
| 0.0 | 0.2 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.0 | 0.1 | GO:0070127 | tRNA aminoacylation for mitochondrial protein translation(GO:0070127) |
| 0.0 | 0.1 | GO:0018094 | protein polyglycylation(GO:0018094) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 14.6 | GO:0005579 | membrane attack complex(GO:0005579) |
| 1.3 | 11.6 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 1.2 | 13.3 | GO:0045179 | apical cortex(GO:0045179) |
| 0.8 | 32.3 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.6 | 1.7 | GO:0097224 | sperm connecting piece(GO:0097224) |
| 0.5 | 2.1 | GO:0031088 | platelet dense granule membrane(GO:0031088) intrinsic component of autophagosome membrane(GO:0097636) integral component of autophagosome membrane(GO:0097637) |
| 0.5 | 2.7 | GO:0097057 | TRAF2-GSTP1 complex(GO:0097057) |
| 0.4 | 3.6 | GO:0032937 | SREBP-SCAP-Insig complex(GO:0032937) |
| 0.4 | 1.5 | GO:1990590 | ATF1-ATF4 transcription factor complex(GO:1990590) |
| 0.4 | 1.5 | GO:0043159 | acrosomal matrix(GO:0043159) |
| 0.4 | 2.9 | GO:0045293 | mRNA editing complex(GO:0045293) |
| 0.4 | 6.4 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.4 | 1.4 | GO:0070722 | Tle3-Aes complex(GO:0070722) |
| 0.4 | 1.4 | GO:0097543 | ciliary inversin compartment(GO:0097543) |
| 0.3 | 1.0 | GO:0032997 | Fc receptor complex(GO:0032997) Fc-epsilon receptor I complex(GO:0032998) |
| 0.3 | 1.2 | GO:0005745 | m-AAA complex(GO:0005745) |
| 0.3 | 1.2 | GO:0060171 | stereocilium membrane(GO:0060171) |
| 0.3 | 0.6 | GO:0030485 | smooth muscle contractile fiber(GO:0030485) |
| 0.3 | 2.2 | GO:0000836 | ER ubiquitin ligase complex(GO:0000835) Hrd1p ubiquitin ligase complex(GO:0000836) |
| 0.3 | 66.3 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.3 | 36.0 | GO:0000800 | lateral element(GO:0000800) |
| 0.2 | 0.7 | GO:0071007 | U2-type catalytic step 2 spliceosome(GO:0071007) |
| 0.2 | 0.7 | GO:1990730 | VCP-NSFL1C complex(GO:1990730) |
| 0.2 | 2.7 | GO:0036128 | CatSper complex(GO:0036128) |
| 0.2 | 1.5 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.2 | 3.7 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.2 | 2.5 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.2 | 1.2 | GO:0071547 | piP-body(GO:0071547) |
| 0.2 | 0.8 | GO:0070469 | respiratory chain(GO:0070469) |
| 0.2 | 0.6 | GO:0043512 | inhibin A complex(GO:0043512) |
| 0.2 | 17.5 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.2 | 2.4 | GO:0020005 | symbiont-containing vacuole membrane(GO:0020005) |
| 0.2 | 0.5 | GO:1902636 | kinociliary basal body(GO:1902636) |
| 0.1 | 4.7 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.1 | 2.4 | GO:0005766 | primary lysosome(GO:0005766) azurophil granule(GO:0042582) |
| 0.1 | 1.0 | GO:0070552 | BRISC complex(GO:0070552) |
| 0.1 | 1.1 | GO:0000275 | mitochondrial proton-transporting ATP synthase complex, catalytic core F(1)(GO:0000275) proton-transporting ATP synthase complex, catalytic core F(1)(GO:0045261) |
| 0.1 | 0.9 | GO:0045254 | pyruvate dehydrogenase complex(GO:0045254) |
| 0.1 | 2.0 | GO:0034362 | low-density lipoprotein particle(GO:0034362) |
| 0.1 | 0.8 | GO:0070695 | FHF complex(GO:0070695) |
| 0.1 | 1.0 | GO:0031362 | anchored component of external side of plasma membrane(GO:0031362) |
| 0.1 | 2.6 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.1 | 4.1 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.1 | 54.5 | GO:0005743 | mitochondrial inner membrane(GO:0005743) |
| 0.1 | 3.0 | GO:0000786 | nucleosome(GO:0000786) |
| 0.1 | 1.2 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.1 | 0.5 | GO:0070876 | SOSS complex(GO:0070876) |
| 0.1 | 2.7 | GO:0030057 | desmosome(GO:0030057) |
| 0.1 | 1.0 | GO:0036156 | inner dynein arm(GO:0036156) |
| 0.1 | 0.9 | GO:0043219 | lateral loop(GO:0043219) |
| 0.1 | 9.0 | GO:0005777 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.1 | 170.2 | GO:0005783 | endoplasmic reticulum(GO:0005783) |
| 0.1 | 2.8 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.1 | 0.8 | GO:0070369 | beta-catenin-TCF7L2 complex(GO:0070369) catenin-TCF7L2 complex(GO:0071664) |
| 0.1 | 0.2 | GO:0098855 | HCN channel complex(GO:0098855) |
| 0.1 | 1.0 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.1 | 1.0 | GO:0043196 | varicosity(GO:0043196) |
| 0.1 | 3.2 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.1 | 0.8 | GO:0016011 | dystroglycan complex(GO:0016011) |
| 0.1 | 1.9 | GO:0035686 | sperm fibrous sheath(GO:0035686) |
| 0.1 | 0.7 | GO:0044292 | dendrite terminus(GO:0044292) |
| 0.1 | 1.3 | GO:0005732 | small nucleolar ribonucleoprotein complex(GO:0005732) |
| 0.1 | 0.6 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.1 | 8.2 | GO:0001669 | acrosomal vesicle(GO:0001669) |
| 0.1 | 0.9 | GO:0042588 | zymogen granule(GO:0042588) |
| 0.0 | 1.1 | GO:0048770 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.0 | 1.0 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.0 | 9.6 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.0 | 6.7 | GO:0005764 | lytic vacuole(GO:0000323) lysosome(GO:0005764) |
| 0.0 | 3.1 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 1.6 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.0 | 0.3 | GO:0032389 | MutLalpha complex(GO:0032389) |
| 0.0 | 0.5 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.0 | 0.6 | GO:0090543 | Flemming body(GO:0090543) |
| 0.0 | 0.4 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.0 | 1.4 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 0.6 | GO:0031091 | platelet alpha granule(GO:0031091) |
| 0.0 | 0.2 | GO:0071546 | pi-body(GO:0071546) |
| 0.0 | 0.1 | GO:0032437 | cuticular plate(GO:0032437) |
| 0.0 | 1.0 | GO:0005763 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.0 | 146.4 | GO:0016021 | integral component of membrane(GO:0016021) |
| 0.0 | 2.2 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 0.5 | GO:0030140 | trans-Golgi network transport vesicle(GO:0030140) |
| 0.0 | 0.5 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.0 | 1.3 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.0 | 0.7 | GO:0005604 | basement membrane(GO:0005604) |
| 0.0 | 52.4 | GO:0070062 | extracellular exosome(GO:0070062) |
| 0.0 | 2.0 | GO:0000792 | heterochromatin(GO:0000792) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 10.8 | 32.5 | GO:0047006 | 17-alpha,20-alpha-dihydroxypregn-4-en-3-one dehydrogenase activity(GO:0047006) |
| 5.3 | 15.9 | GO:0080130 | L-phenylalanine:2-oxoglutarate aminotransferase activity(GO:0080130) |
| 4.9 | 14.8 | GO:0005009 | insulin-activated receptor activity(GO:0005009) |
| 4.9 | 19.6 | GO:0016743 | carboxyl- or carbamoyltransferase activity(GO:0016743) |
| 4.8 | 19.3 | GO:0071614 | linoleic acid epoxygenase activity(GO:0071614) |
| 4.4 | 13.1 | GO:0004024 | alcohol dehydrogenase activity, zinc-dependent(GO:0004024) |
| 4.3 | 25.7 | GO:0004505 | phenylalanine 4-monooxygenase activity(GO:0004505) |
| 3.9 | 130.0 | GO:0008392 | arachidonic acid epoxygenase activity(GO:0008392) |
| 3.8 | 156.7 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 3.4 | 13.5 | GO:1902271 | D3 vitamins binding(GO:1902271) |
| 3.0 | 18.0 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 2.8 | 11.1 | GO:0019166 | trans-2-enoyl-CoA reductase (NADPH) activity(GO:0019166) |
| 2.1 | 8.2 | GO:0047961 | glycine N-acyltransferase activity(GO:0047961) |
| 2.1 | 12.3 | GO:0032450 | maltose alpha-glucosidase activity(GO:0032450) |
| 2.0 | 16.4 | GO:0047035 | testosterone dehydrogenase (NAD+) activity(GO:0047035) |
| 2.0 | 40.8 | GO:0016712 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen(GO:0016712) |
| 2.0 | 5.9 | GO:0008119 | thiopurine S-methyltransferase activity(GO:0008119) |
| 1.9 | 7.4 | GO:0004031 | aldehyde oxidase activity(GO:0004031) oxidoreductase activity, acting on the aldehyde or oxo group of donors, oxygen as acceptor(GO:0016623) |
| 1.9 | 5.6 | GO:0008431 | vitamin E binding(GO:0008431) |
| 1.7 | 6.9 | GO:0050309 | glucose-6-phosphatase activity(GO:0004346) sugar-terminal-phosphatase activity(GO:0050309) |
| 1.6 | 9.3 | GO:0050656 | 3'-phosphoadenosine 5'-phosphosulfate binding(GO:0050656) |
| 1.5 | 12.3 | GO:0005186 | pheromone activity(GO:0005186) |
| 1.5 | 7.6 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) |
| 1.5 | 13.3 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 1.4 | 4.2 | GO:0016034 | maleylacetoacetate isomerase activity(GO:0016034) |
| 1.3 | 9.3 | GO:0004769 | steroid delta-isomerase activity(GO:0004769) |
| 1.3 | 7.8 | GO:0008453 | alanine-glyoxylate transaminase activity(GO:0008453) |
| 1.2 | 3.5 | GO:0016749 | 5-aminolevulinate synthase activity(GO:0003870) N-succinyltransferase activity(GO:0016749) |
| 1.1 | 3.4 | GO:0019150 | D-ribulokinase activity(GO:0019150) |
| 1.0 | 4.0 | GO:0036468 | aromatic-L-amino-acid decarboxylase activity(GO:0004058) L-dopa decarboxylase activity(GO:0036468) |
| 0.9 | 3.7 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 0.9 | 3.7 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.9 | 4.5 | GO:0097003 | adipokinetic hormone receptor activity(GO:0097003) |
| 0.9 | 2.7 | GO:0035731 | S-nitrosoglutathione binding(GO:0035730) dinitrosyl-iron complex binding(GO:0035731) |
| 0.9 | 5.3 | GO:0008745 | N-acetylmuramoyl-L-alanine amidase activity(GO:0008745) peptidoglycan receptor activity(GO:0016019) |
| 0.9 | 4.3 | GO:0015205 | purine nucleobase transmembrane transporter activity(GO:0005345) pyrimidine nucleobase transmembrane transporter activity(GO:0005350) nucleobase transmembrane transporter activity(GO:0015205) glycerol channel activity(GO:0015254) |
| 0.8 | 2.5 | GO:0016508 | long-chain-enoyl-CoA hydratase activity(GO:0016508) |
| 0.8 | 2.4 | GO:0019002 | GMP binding(GO:0019002) |
| 0.8 | 3.1 | GO:0016708 | nitric oxide dioxygenase activity(GO:0008941) oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of two atoms of oxygen into one donor(GO:0016708) iron-cytochrome-c reductase activity(GO:0047726) |
| 0.7 | 3.7 | GO:0004531 | deoxyribonuclease II activity(GO:0004531) |
| 0.7 | 2.2 | GO:0047936 | glucose 1-dehydrogenase [NAD(P)] activity(GO:0047936) |
| 0.7 | 2.2 | GO:0003990 | acetylcholinesterase activity(GO:0003990) |
| 0.7 | 3.6 | GO:0008172 | S-methyltransferase activity(GO:0008172) |
| 0.7 | 2.2 | GO:0005148 | prolactin receptor binding(GO:0005148) |
| 0.7 | 3.5 | GO:0004974 | leukotriene receptor activity(GO:0004974) |
| 0.7 | 7.0 | GO:0003854 | 3-beta-hydroxy-delta5-steroid dehydrogenase activity(GO:0003854) |
| 0.7 | 2.8 | GO:0042284 | sphingolipid delta-4 desaturase activity(GO:0042284) |
| 0.7 | 7.5 | GO:0016174 | NAD(P)H oxidase activity(GO:0016174) |
| 0.7 | 2.0 | GO:0004982 | N-formyl peptide receptor activity(GO:0004982) |
| 0.7 | 3.3 | GO:0004528 | phosphodiesterase I activity(GO:0004528) |
| 0.7 | 3.9 | GO:0016647 | oxidoreductase activity, acting on the CH-NH group of donors, oxygen as acceptor(GO:0016647) |
| 0.7 | 2.0 | GO:0017099 | very-long-chain-acyl-CoA dehydrogenase activity(GO:0017099) |
| 0.6 | 10.7 | GO:0070008 | serine-type exopeptidase activity(GO:0070008) |
| 0.6 | 4.4 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.6 | 10.6 | GO:0015143 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.6 | 6.8 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 0.6 | 4.8 | GO:0005534 | galactose binding(GO:0005534) |
| 0.6 | 6.6 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.6 | 5.4 | GO:0004768 | stearoyl-CoA 9-desaturase activity(GO:0004768) |
| 0.6 | 5.9 | GO:0004064 | arylesterase activity(GO:0004064) |
| 0.6 | 29.5 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.6 | 6.6 | GO:0009922 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.5 | 10.3 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
| 0.5 | 9.2 | GO:0008430 | selenium binding(GO:0008430) |
| 0.5 | 6.1 | GO:0048403 | brain-derived neurotrophic factor binding(GO:0048403) |
| 0.5 | 3.2 | GO:0004556 | alpha-amylase activity(GO:0004556) |
| 0.4 | 1.8 | GO:0047275 | glucosaminylgalactosylglucosylceramide beta-galactosyltransferase activity(GO:0047275) |
| 0.4 | 1.3 | GO:0031751 | D4 dopamine receptor binding(GO:0031751) |
| 0.4 | 5.1 | GO:0004322 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.4 | 1.1 | GO:0001729 | ceramide kinase activity(GO:0001729) |
| 0.4 | 1.9 | GO:0016774 | phosphotransferase activity, carboxyl group as acceptor(GO:0016774) |
| 0.4 | 6.5 | GO:0003796 | lysozyme activity(GO:0003796) |
| 0.4 | 2.2 | GO:0004784 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 0.4 | 1.1 | GO:0031768 | ghrelin receptor binding(GO:0031768) |
| 0.4 | 1.1 | GO:0004771 | sterol esterase activity(GO:0004771) |
| 0.3 | 1.7 | GO:0004165 | dodecenoyl-CoA delta-isomerase activity(GO:0004165) |
| 0.3 | 0.7 | GO:0016608 | growth hormone-releasing hormone activity(GO:0016608) |
| 0.3 | 3.9 | GO:0015280 | ligand-gated sodium channel activity(GO:0015280) |
| 0.3 | 0.9 | GO:0004774 | succinate-CoA ligase activity(GO:0004774) |
| 0.3 | 1.5 | GO:0008761 | UDP-N-acetylglucosamine 2-epimerase activity(GO:0008761) |
| 0.3 | 0.9 | GO:0030021 | extracellular matrix structural constituent conferring compression resistance(GO:0030021) structural constituent of tooth enamel(GO:0030345) |
| 0.3 | 2.1 | GO:0050833 | pyruvate transmembrane transporter activity(GO:0050833) |
| 0.3 | 3.5 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 0.3 | 1.1 | GO:0046403 | polynucleotide 3'-phosphatase activity(GO:0046403) |
| 0.3 | 0.9 | GO:0004739 | pyruvate dehydrogenase (acetyl-transferring) activity(GO:0004739) |
| 0.3 | 0.9 | GO:0004447 | iodide peroxidase activity(GO:0004447) |
| 0.3 | 1.7 | GO:0008970 | phosphatidylcholine 1-acylhydrolase activity(GO:0008970) |
| 0.3 | 3.3 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.3 | 1.1 | GO:0004441 | inositol-1,4-bisphosphate 1-phosphatase activity(GO:0004441) |
| 0.3 | 1.9 | GO:0004782 | sulfinoalanine decarboxylase activity(GO:0004782) |
| 0.3 | 2.3 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.2 | 1.0 | GO:0004793 | threonine aldolase activity(GO:0004793) L-allo-threonine aldolase activity(GO:0008732) |
| 0.2 | 1.4 | GO:0008142 | oxysterol binding(GO:0008142) |
| 0.2 | 1.2 | GO:0004060 | arylamine N-acetyltransferase activity(GO:0004060) |
| 0.2 | 265.0 | GO:0004984 | olfactory receptor activity(GO:0004984) |
| 0.2 | 0.9 | GO:0072541 | peroxynitrite reductase activity(GO:0072541) |
| 0.2 | 2.7 | GO:0008525 | phosphatidylcholine transporter activity(GO:0008525) |
| 0.2 | 1.0 | GO:0019767 | IgE receptor activity(GO:0019767) |
| 0.2 | 1.6 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.2 | 0.8 | GO:0001588 | dopamine neurotransmitter receptor activity, coupled via Gs(GO:0001588) |
| 0.2 | 1.0 | GO:0005124 | scavenger receptor binding(GO:0005124) |
| 0.2 | 0.8 | GO:0035575 | histone demethylase activity (H4-K20 specific)(GO:0035575) |
| 0.2 | 1.9 | GO:0004579 | oligosaccharyl transferase activity(GO:0004576) dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.2 | 1.3 | GO:0016812 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amides(GO:0016812) |
| 0.2 | 5.0 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.2 | 0.6 | GO:0033754 | indoleamine 2,3-dioxygenase activity(GO:0033754) |
| 0.2 | 1.3 | GO:0001851 | complement component C3b binding(GO:0001851) |
| 0.2 | 1.8 | GO:0008420 | CTD phosphatase activity(GO:0008420) |
| 0.2 | 1.8 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.2 | 0.7 | GO:0035800 | deubiquitinase activator activity(GO:0035800) |
| 0.2 | 20.4 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.2 | 2.8 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.2 | 1.7 | GO:0004415 | hyalurononglucosaminidase activity(GO:0004415) |
| 0.2 | 0.7 | GO:0031721 | hemoglobin alpha binding(GO:0031721) |
| 0.2 | 1.0 | GO:0015226 | amino-acid betaine transmembrane transporter activity(GO:0015199) carnitine transmembrane transporter activity(GO:0015226) |
| 0.2 | 1.0 | GO:0000832 | inositol hexakisphosphate kinase activity(GO:0000828) inositol hexakisphosphate 5-kinase activity(GO:0000832) inositol hexakisphosphate 1-kinase activity(GO:0052723) inositol hexakisphosphate 3-kinase activity(GO:0052724) |
| 0.2 | 26.3 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.2 | 1.1 | GO:0047276 | N-acetyllactosaminide 3-alpha-galactosyltransferase activity(GO:0047276) |
| 0.2 | 1.8 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.2 | 0.5 | GO:0004510 | tryptophan 5-monooxygenase activity(GO:0004510) |
| 0.2 | 0.3 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.2 | 0.6 | GO:0008559 | xenobiotic-transporting ATPase activity(GO:0008559) |
| 0.2 | 3.8 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.2 | 11.4 | GO:0016706 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, 2-oxoglutarate as one donor, and incorporation of one atom each of oxygen into both donors(GO:0016706) |
| 0.2 | 1.4 | GO:0003836 | beta-galactoside (CMP) alpha-2,3-sialyltransferase activity(GO:0003836) |
| 0.1 | 1.2 | GO:0047134 | protein-disulfide reductase activity(GO:0047134) |
| 0.1 | 0.9 | GO:0019958 | C-X-C chemokine binding(GO:0019958) |
| 0.1 | 1.0 | GO:0016453 | acetyl-CoA C-acetyltransferase activity(GO:0003985) C-acetyltransferase activity(GO:0016453) |
| 0.1 | 1.9 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.1 | 0.8 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
| 0.1 | 0.4 | GO:0004731 | purine-nucleoside phosphorylase activity(GO:0004731) |
| 0.1 | 5.6 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 0.1 | 4.0 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.1 | 0.4 | GO:0005137 | interleukin-5 receptor binding(GO:0005137) |
| 0.1 | 1.1 | GO:0003996 | acyl-CoA ligase activity(GO:0003996) |
| 0.1 | 2.6 | GO:0070403 | NAD+ binding(GO:0070403) |
| 0.1 | 0.4 | GO:0044715 | 8-oxo-dGDP phosphatase activity(GO:0044715) |
| 0.1 | 3.6 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.1 | 5.4 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.1 | 7.3 | GO:0004177 | aminopeptidase activity(GO:0004177) |
| 0.1 | 1.5 | GO:0015250 | water channel activity(GO:0015250) |
| 0.1 | 1.5 | GO:0003847 | 1-alkyl-2-acetylglycerophosphocholine esterase activity(GO:0003847) |
| 0.1 | 3.3 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.1 | 1.7 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.1 | 0.8 | GO:0070492 | oligosaccharide binding(GO:0070492) |
| 0.1 | 2.0 | GO:0055106 | ubiquitin-protein transferase regulator activity(GO:0055106) |
| 0.1 | 0.8 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.1 | 1.0 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.1 | 1.1 | GO:0004571 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) |
| 0.1 | 0.3 | GO:0003868 | 4-hydroxyphenylpyruvate dioxygenase activity(GO:0003868) |
| 0.1 | 0.5 | GO:0004449 | isocitrate dehydrogenase (NAD+) activity(GO:0004449) |
| 0.1 | 2.6 | GO:0008395 | steroid hydroxylase activity(GO:0008395) |
| 0.1 | 0.3 | GO:0018169 | ribosomal S6-glutamic acid ligase activity(GO:0018169) |
| 0.1 | 0.8 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.1 | 12.7 | GO:0052689 | carboxylic ester hydrolase activity(GO:0052689) |
| 0.1 | 0.9 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) ADP transmembrane transporter activity(GO:0015217) |
| 0.1 | 0.9 | GO:0008603 | cAMP-dependent protein kinase regulator activity(GO:0008603) |
| 0.1 | 0.8 | GO:0015929 | hexosaminidase activity(GO:0015929) |
| 0.1 | 0.4 | GO:0004735 | pyrroline-5-carboxylate reductase activity(GO:0004735) |
| 0.1 | 0.9 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.1 | 0.5 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 0.1 | 0.9 | GO:0031433 | telethonin binding(GO:0031433) |
| 0.1 | 0.3 | GO:0003976 | UDP-N-acetylglucosamine-lysosomal-enzyme N-acetylglucosaminephosphotransferase activity(GO:0003976) |
| 0.1 | 0.7 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.1 | 1.6 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.1 | 0.6 | GO:0070699 | type II activin receptor binding(GO:0070699) |
| 0.1 | 0.9 | GO:0000182 | rDNA binding(GO:0000182) |
| 0.1 | 0.3 | GO:0098809 | nitrite reductase activity(GO:0098809) |
| 0.1 | 1.5 | GO:0004866 | endopeptidase inhibitor activity(GO:0004866) |
| 0.1 | 1.0 | GO:0008301 | DNA binding, bending(GO:0008301) |
| 0.1 | 0.5 | GO:0033592 | RNA strand annealing activity(GO:0033592) |
| 0.1 | 1.1 | GO:0070513 | death domain binding(GO:0070513) |
| 0.1 | 1.0 | GO:0015299 | solute:proton antiporter activity(GO:0015299) |
| 0.1 | 0.6 | GO:0086080 | protein binding involved in heterotypic cell-cell adhesion(GO:0086080) |
| 0.1 | 0.8 | GO:1990829 | C-rich single-stranded DNA binding(GO:1990829) |
| 0.1 | 2.7 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.1 | 3.8 | GO:0019003 | GDP binding(GO:0019003) |
| 0.1 | 0.4 | GO:0034046 | poly(G) binding(GO:0034046) |
| 0.1 | 3.2 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.0 | 0.9 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
| 0.0 | 1.0 | GO:0001056 | RNA polymerase III activity(GO:0001056) |
| 0.0 | 0.9 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
| 0.0 | 4.6 | GO:0005506 | iron ion binding(GO:0005506) |
| 0.0 | 0.4 | GO:0010861 | thyroid hormone receptor activator activity(GO:0010861) |
| 0.0 | 4.3 | GO:0016811 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amides(GO:0016811) |
| 0.0 | 0.8 | GO:0008484 | sulfuric ester hydrolase activity(GO:0008484) |
| 0.0 | 0.5 | GO:0003956 | NAD(P)+-protein-arginine ADP-ribosyltransferase activity(GO:0003956) |
| 0.0 | 0.7 | GO:0001848 | complement binding(GO:0001848) |
| 0.0 | 0.7 | GO:0017070 | U6 snRNA binding(GO:0017070) |
| 0.0 | 0.1 | GO:0009383 | rRNA (cytosine-C5-)-methyltransferase activity(GO:0009383) |
| 0.0 | 0.4 | GO:0015288 | porin activity(GO:0015288) |
| 0.0 | 0.3 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.0 | 0.8 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.0 | 0.2 | GO:1900750 | glutathione binding(GO:0043295) oligopeptide binding(GO:1900750) |
| 0.0 | 1.5 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
| 0.0 | 0.3 | GO:0070883 | pre-miRNA binding(GO:0070883) |
| 0.0 | 0.3 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.0 | 0.6 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.0 | 0.2 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.0 | 0.5 | GO:0042288 | MHC class I protein binding(GO:0042288) |
| 0.0 | 0.2 | GO:0005222 | intracellular cAMP activated cation channel activity(GO:0005222) |
| 0.0 | 0.5 | GO:1990381 | ubiquitin-specific protease binding(GO:1990381) |
| 0.0 | 0.3 | GO:0015450 | P-P-bond-hydrolysis-driven protein transmembrane transporter activity(GO:0015450) |
| 0.0 | 0.3 | GO:0043237 | laminin-1 binding(GO:0043237) |
| 0.0 | 0.6 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
| 0.0 | 5.2 | GO:0042626 | ATPase activity, coupled to transmembrane movement of substances(GO:0042626) |
| 0.0 | 0.6 | GO:0031701 | angiotensin receptor binding(GO:0031701) type 1 angiotensin receptor binding(GO:0031702) |
| 0.0 | 0.2 | GO:0001601 | peptide YY receptor activity(GO:0001601) pancreatic polypeptide receptor activity(GO:0001602) |
| 0.0 | 0.9 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 0.1 | GO:0031177 | phosphopantetheine binding(GO:0031177) |
| 0.0 | 0.4 | GO:0098505 | G-rich strand telomeric DNA binding(GO:0098505) |
| 0.0 | 0.6 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 0.0 | 2.4 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.0 | 0.8 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) |
| 0.0 | 2.1 | GO:0002039 | p53 binding(GO:0002039) |
| 0.0 | 1.2 | GO:0051287 | NAD binding(GO:0051287) |
| 0.0 | 0.9 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.0 | 0.6 | GO:0004601 | peroxidase activity(GO:0004601) |
| 0.0 | 2.4 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.0 | 0.1 | GO:0016494 | C-X-C chemokine receptor activity(GO:0016494) |
| 0.0 | 2.0 | GO:0004843 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) |
| 0.0 | 0.8 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.4 | GO:0042166 | acetylcholine binding(GO:0042166) |
| 0.0 | 0.2 | GO:0042923 | neuropeptide binding(GO:0042923) |
| 0.0 | 0.6 | GO:0008188 | neuropeptide receptor activity(GO:0008188) |
| 0.0 | 0.1 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.0 | 0.2 | GO:0043139 | 5'-3' DNA helicase activity(GO:0043139) |
| 0.0 | 0.5 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.0 | 0.6 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 0.3 | GO:0016799 | hydrolase activity, hydrolyzing N-glycosyl compounds(GO:0016799) |
| 0.0 | 2.6 | GO:0017171 | serine-type peptidase activity(GO:0008236) serine hydrolase activity(GO:0017171) |
| 0.0 | 0.5 | GO:0016876 | aminoacyl-tRNA ligase activity(GO:0004812) ligase activity, forming carbon-oxygen bonds(GO:0016875) ligase activity, forming aminoacyl-tRNA and related compounds(GO:0016876) |
| 0.0 | 0.1 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.0 | 0.2 | GO:0015174 | basic amino acid transmembrane transporter activity(GO:0015174) |
| 0.0 | 0.1 | GO:0070735 | protein-glycine ligase activity(GO:0070735) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 21.8 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.6 | 1.2 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.5 | 0.5 | PID NFAT 3PATHWAY | Role of Calcineurin-dependent NFAT signaling in lymphocytes |
| 0.3 | 20.8 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 2.0 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.1 | 1.2 | ST INTERFERON GAMMA PATHWAY | Interferon gamma pathway. |
| 0.1 | 1.7 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.1 | 0.7 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.1 | 0.4 | ST TYPE I INTERFERON PATHWAY | Type I Interferon (alpha/beta IFN) Pathway |
| 0.1 | 1.8 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.1 | 1.1 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
| 0.0 | 10.2 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 1.5 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.0 | 2.2 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.0 | 0.9 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.0 | 3.7 | PID E2F PATHWAY | E2F transcription factor network |
| 0.0 | 0.3 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.0 | 1.7 | PID HIV NEF PATHWAY | HIV-1 Nef: Negative effector of Fas and TNF-alpha |
| 0.0 | 0.3 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 0.0 | 1.0 | PID CONE PATHWAY | Visual signal transduction: Cones |
| 0.0 | 0.7 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.0 | 2.2 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.0 | 0.6 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 1.7 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 0.6 | PID EPHB FWD PATHWAY | EPHB forward signaling |
| 0.0 | 0.8 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.0 | 0.3 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.0 | 0.8 | PID IL23 PATHWAY | IL23-mediated signaling events |
| 0.0 | 0.7 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 0.6 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.0 | 0.4 | PID AP1 PATHWAY | AP-1 transcription factor network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.1 | 12.3 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 2.8 | 8.5 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 1.2 | 28.2 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 1.2 | 13.1 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.8 | 11.6 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.7 | 22.5 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.7 | 14.2 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.7 | 9.3 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.6 | 7.0 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.6 | 7.8 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.5 | 5.9 | REACTOME PHASE II CONJUGATION | Genes involved in Phase II conjugation |
| 0.5 | 23.8 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.5 | 7.0 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 0.4 | 6.8 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.3 | 82.0 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.3 | 7.7 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.3 | 4.3 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.2 | 3.4 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.2 | 2.7 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.2 | 6.8 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.2 | 7.6 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.2 | 4.5 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.2 | 3.3 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.2 | 3.5 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.1 | 6.0 | REACTOME METAL ION SLC TRANSPORTERS | Genes involved in Metal ion SLC transporters |
| 0.1 | 5.0 | REACTOME STEROID HORMONES | Genes involved in Steroid hormones |
| 0.1 | 2.0 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.1 | 1.0 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.1 | 2.1 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.1 | 0.9 | REACTOME GLUCAGON SIGNALING IN METABOLIC REGULATION | Genes involved in Glucagon signaling in metabolic regulation |
| 0.1 | 7.1 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.1 | 0.6 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.1 | 1.7 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.1 | 1.1 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 0.1 | 0.7 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.1 | 1.0 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 0.1 | 1.4 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.1 | 1.2 | REACTOME FORMATION OF ATP BY CHEMIOSMOTIC COUPLING | Genes involved in Formation of ATP by chemiosmotic coupling |
| 0.1 | 0.1 | REACTOME FRS2 MEDIATED CASCADE | Genes involved in FRS2-mediated cascade |
| 0.1 | 0.7 | REACTOME IRAK1 RECRUITS IKK COMPLEX | Genes involved in IRAK1 recruits IKK complex |
| 0.1 | 0.8 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.1 | 0.9 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.0 | 1.0 | REACTOME RNA POL III CHAIN ELONGATION | Genes involved in RNA Polymerase III Chain Elongation |
| 0.0 | 0.9 | REACTOME TRAF3 DEPENDENT IRF ACTIVATION PATHWAY | Genes involved in TRAF3-dependent IRF activation pathway |
| 0.0 | 1.2 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.0 | 1.9 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 0.3 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.0 | 1.0 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 0.4 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.0 | 1.0 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.0 | 0.3 | REACTOME P53 DEPENDENT G1 DNA DAMAGE RESPONSE | Genes involved in p53-Dependent G1 DNA Damage Response |
| 0.0 | 0.4 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.0 | 0.4 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.0 | 0.4 | REACTOME TRANSLATION | Genes involved in Translation |
| 0.0 | 1.2 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.0 | 2.4 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.0 | 1.7 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.0 | 0.5 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 0.4 | REACTOME IL RECEPTOR SHC SIGNALING | Genes involved in Interleukin receptor SHC signaling |
| 0.0 | 0.6 | REACTOME TRANSPORT OF GLUCOSE AND OTHER SUGARS BILE SALTS AND ORGANIC ACIDS METAL IONS AND AMINE COMPOUNDS | Genes involved in Transport of glucose and other sugars, bile salts and organic acids, metal ions and amine compounds |