Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
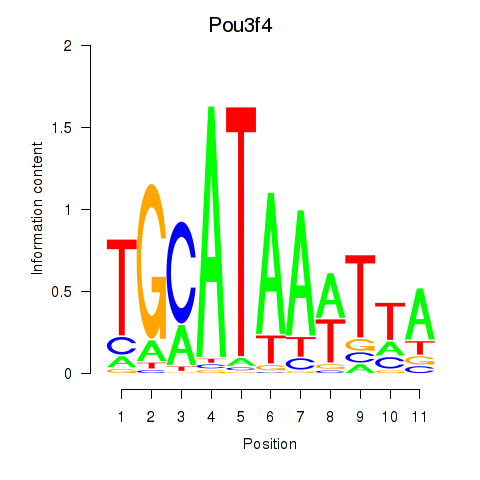
Results for Pou3f4
Z-value: 1.24
Transcription factors associated with Pou3f4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Pou3f4
|
ENSMUSG00000056854.5 | Pou3f4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Pou3f4 | mm39_v1_chrX_+_109857866_109857886 | 0.77 | 2.4e-15 | Click! |
Activity profile of Pou3f4 motif
Sorted Z-values of Pou3f4 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Pou3f4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr11_-_99134885 | 11.73 |
ENSMUST00000103132.10
ENSMUST00000038214.7 |
Krt222
|
keratin 222 |
| chr2_-_79959178 | 11.41 |
ENSMUST00000102654.11
ENSMUST00000102655.10 |
Pde1a
|
phosphodiesterase 1A, calmodulin-dependent |
| chr2_-_66271097 | 10.59 |
ENSMUST00000112371.9
ENSMUST00000138910.3 |
Scn1a
|
sodium channel, voltage-gated, type I, alpha |
| chr1_-_132318039 | 10.59 |
ENSMUST00000132435.2
|
Tmcc2
|
transmembrane and coiled-coil domains 2 |
| chr15_+_92059224 | 10.35 |
ENSMUST00000068378.6
|
Cntn1
|
contactin 1 |
| chr10_+_79650496 | 10.15 |
ENSMUST00000218857.2
ENSMUST00000220365.2 |
Palm
|
paralemmin |
| chr8_+_13034245 | 10.11 |
ENSMUST00000110873.10
ENSMUST00000173006.8 ENSMUST00000145067.8 |
Mcf2l
|
mcf.2 transforming sequence-like |
| chr11_+_92989229 | 9.56 |
ENSMUST00000107859.8
ENSMUST00000107861.8 ENSMUST00000042943.13 ENSMUST00000107858.9 |
Car10
|
carbonic anhydrase 10 |
| chr3_+_67799510 | 8.56 |
ENSMUST00000063263.5
ENSMUST00000182006.4 |
Iqcj
Iqschfp
|
IQ motif containing J Iqcj and Schip1 fusion protein |
| chr6_+_21215472 | 8.50 |
ENSMUST00000081542.6
|
Kcnd2
|
potassium voltage-gated channel, Shal-related family, member 2 |
| chr10_-_63926044 | 8.48 |
ENSMUST00000105439.2
|
Lrrtm3
|
leucine rich repeat transmembrane neuronal 3 |
| chr10_+_90412114 | 8.36 |
ENSMUST00000182427.8
ENSMUST00000182053.8 ENSMUST00000182113.8 |
Anks1b
|
ankyrin repeat and sterile alpha motif domain containing 1B |
| chr12_-_72283465 | 8.04 |
ENSMUST00000021497.16
ENSMUST00000137990.2 |
Rtn1
|
reticulon 1 |
| chr11_-_99313078 | 7.98 |
ENSMUST00000017741.4
|
Krt12
|
keratin 12 |
| chr1_-_38937190 | 7.92 |
ENSMUST00000193435.6
ENSMUST00000194361.6 |
Chst10
|
carbohydrate sulfotransferase 10 |
| chr13_+_93441307 | 7.90 |
ENSMUST00000080127.12
|
Homer1
|
homer scaffolding protein 1 |
| chr14_+_27598021 | 7.71 |
ENSMUST00000211684.2
ENSMUST00000210924.2 |
Erc2
|
ELKS/RAB6-interacting/CAST family member 2 |
| chr9_+_113641615 | 7.61 |
ENSMUST00000111838.10
ENSMUST00000166734.10 ENSMUST00000214522.2 ENSMUST00000163895.3 |
Clasp2
|
CLIP associating protein 2 |
| chr1_+_66360865 | 7.47 |
ENSMUST00000114013.8
|
Map2
|
microtubule-associated protein 2 |
| chr4_+_15881256 | 7.36 |
ENSMUST00000029876.2
|
Calb1
|
calbindin 1 |
| chr7_+_113365235 | 7.23 |
ENSMUST00000046687.16
|
Spon1
|
spondin 1, (f-spondin) extracellular matrix protein |
| chr8_+_94537910 | 7.16 |
ENSMUST00000138659.9
|
Gnao1
|
guanine nucleotide binding protein, alpha O |
| chr11_+_92990110 | 7.13 |
ENSMUST00000107863.4
|
Car10
|
carbonic anhydrase 10 |
| chr6_+_96090127 | 6.89 |
ENSMUST00000122120.8
|
Tafa1
|
TAFA chemokine like family member 1 |
| chr11_+_71640739 | 6.58 |
ENSMUST00000150531.2
|
Wscd1
|
WSC domain containing 1 |
| chr1_+_179788675 | 6.57 |
ENSMUST00000076687.12
ENSMUST00000097450.10 ENSMUST00000212756.2 |
Cdc42bpa
|
CDC42 binding protein kinase alpha |
| chr1_-_38937170 | 6.53 |
ENSMUST00000193441.6
|
Chst10
|
carbohydrate sulfotransferase 10 |
| chr18_+_37453427 | 6.51 |
ENSMUST00000078271.4
|
Pcdhb5
|
protocadherin beta 5 |
| chr16_+_94225942 | 6.45 |
ENSMUST00000141176.2
|
Ttc3
|
tetratricopeptide repeat domain 3 |
| chr3_+_102641822 | 6.36 |
ENSMUST00000029451.12
|
Tspan2
|
tetraspanin 2 |
| chr15_+_30457772 | 6.22 |
ENSMUST00000228282.2
|
Ctnnd2
|
catenin (cadherin associated protein), delta 2 |
| chr10_-_127025851 | 6.22 |
ENSMUST00000222006.2
ENSMUST00000019611.15 ENSMUST00000219245.2 |
Arhgef25
|
Rho guanine nucleotide exchange factor (GEF) 25 |
| chr1_-_38937061 | 6.18 |
ENSMUST00000027249.12
|
Chst10
|
carbohydrate sulfotransferase 10 |
| chr2_-_79959802 | 6.04 |
ENSMUST00000102653.8
|
Pde1a
|
phosphodiesterase 1A, calmodulin-dependent |
| chr13_-_91053440 | 5.78 |
ENSMUST00000109541.4
|
Atp6ap1l
|
ATPase, H+ transporting, lysosomal accessory protein 1-like |
| chr11_-_12362136 | 5.76 |
ENSMUST00000174874.8
|
Cobl
|
cordon-bleu WH2 repeat |
| chr3_-_116047148 | 5.63 |
ENSMUST00000090473.7
|
Gpr88
|
G-protein coupled receptor 88 |
| chr7_-_66915756 | 5.61 |
ENSMUST00000207715.2
|
Mef2a
|
myocyte enhancer factor 2A |
| chr4_-_41569500 | 5.55 |
ENSMUST00000108049.9
ENSMUST00000108052.10 ENSMUST00000108050.2 |
Fam219a
|
family with sequence similarity 219, member A |
| chr13_+_42834989 | 5.54 |
ENSMUST00000149235.8
|
Phactr1
|
phosphatase and actin regulator 1 |
| chr15_+_18819033 | 5.39 |
ENSMUST00000166873.9
|
Cdh10
|
cadherin 10 |
| chr8_-_72178340 | 5.25 |
ENSMUST00000153800.8
ENSMUST00000146100.8 |
Fcho1
|
FCH domain only 1 |
| chr5_-_52723607 | 5.22 |
ENSMUST00000199942.5
|
Lgi2
|
leucine-rich repeat LGI family, member 2 |
| chr8_+_63404228 | 5.16 |
ENSMUST00000118003.8
|
Spock3
|
sparc/osteonectin, cwcv and kazal-like domains proteoglycan 3 |
| chr18_+_37072232 | 5.13 |
ENSMUST00000115662.9
ENSMUST00000195590.2 |
Pcdha2
|
protocadherin alpha 2 |
| chr2_-_120075639 | 4.93 |
ENSMUST00000090071.5
|
Pla2g4e
|
phospholipase A2, group IVE |
| chr5_-_52723700 | 4.89 |
ENSMUST00000039750.7
|
Lgi2
|
leucine-rich repeat LGI family, member 2 |
| chr3_+_68479578 | 4.72 |
ENSMUST00000170788.9
|
Schip1
|
schwannomin interacting protein 1 |
| chr4_-_138053545 | 4.60 |
ENSMUST00000105817.4
|
Pink1
|
PTEN induced putative kinase 1 |
| chr6_-_102441628 | 4.59 |
ENSMUST00000032159.7
|
Cntn3
|
contactin 3 |
| chr15_-_100497863 | 4.57 |
ENSMUST00000073837.13
|
Pou6f1
|
POU domain, class 6, transcription factor 1 |
| chr5_+_57876401 | 4.57 |
ENSMUST00000094783.7
|
Pcdh7
|
protocadherin 7 |
| chr15_-_53765869 | 4.51 |
ENSMUST00000078673.14
|
Samd12
|
sterile alpha motif domain containing 12 |
| chr4_-_110144676 | 4.47 |
ENSMUST00000106598.8
ENSMUST00000102723.11 ENSMUST00000153906.2 |
Elavl4
|
ELAV like RNA binding protein 4 |
| chr11_+_87469211 | 4.42 |
ENSMUST00000107962.8
ENSMUST00000122067.8 |
Septin4
|
septin 4 |
| chr2_+_87610895 | 4.42 |
ENSMUST00000215394.2
|
Olfr152
|
olfactory receptor 152 |
| chr5_-_123804745 | 4.40 |
ENSMUST00000149410.2
|
Clip1
|
CAP-GLY domain containing linker protein 1 |
| chr8_+_63404395 | 4.39 |
ENSMUST00000119068.8
|
Spock3
|
sparc/osteonectin, cwcv and kazal-like domains proteoglycan 3 |
| chr1_-_25868592 | 4.33 |
ENSMUST00000135518.8
|
Adgrb3
|
adhesion G protein-coupled receptor B3 |
| chr3_-_19749489 | 4.29 |
ENSMUST00000061294.5
|
Crh
|
corticotropin releasing hormone |
| chr18_+_37637317 | 4.29 |
ENSMUST00000052179.8
|
Pcdhb20
|
protocadherin beta 20 |
| chr4_-_138053603 | 4.18 |
ENSMUST00000030536.13
|
Pink1
|
PTEN induced putative kinase 1 |
| chr7_+_100435548 | 4.12 |
ENSMUST00000216021.2
|
Fam168a
|
family with sequence similarity 168, member A |
| chr11_-_99280177 | 4.02 |
ENSMUST00000103131.5
|
Krt10
|
keratin 10 |
| chr9_-_58109564 | 3.95 |
ENSMUST00000163897.8
ENSMUST00000215950.2 |
Islr2
|
immunoglobulin superfamily containing leucine-rich repeat 2 |
| chr3_+_105881027 | 3.88 |
ENSMUST00000000573.9
|
Ovgp1
|
oviductal glycoprotein 1 |
| chr3_-_33136153 | 3.86 |
ENSMUST00000108225.10
|
Pex5l
|
peroxisomal biogenesis factor 5-like |
| chr18_+_37544717 | 3.74 |
ENSMUST00000051126.4
|
Pcdhb10
|
protocadherin beta 10 |
| chr11_+_29668563 | 3.73 |
ENSMUST00000060992.6
|
Rtn4
|
reticulon 4 |
| chr2_+_67004178 | 3.71 |
ENSMUST00000239009.2
ENSMUST00000238912.2 |
Xirp2
|
xin actin-binding repeat containing 2 |
| chr5_-_74226394 | 3.66 |
ENSMUST00000145016.3
|
Usp46
|
ubiquitin specific peptidase 46 |
| chr14_+_10123804 | 3.57 |
ENSMUST00000022262.6
ENSMUST00000224714.2 |
Fezf2
|
Fez family zinc finger 2 |
| chr13_+_93440572 | 3.56 |
ENSMUST00000109493.9
|
Homer1
|
homer scaffolding protein 1 |
| chr2_-_26184450 | 3.53 |
ENSMUST00000217256.2
ENSMUST00000227200.2 |
Ccdc187
|
coiled-coil domain containing 187 |
| chr12_+_103564479 | 3.47 |
ENSMUST00000190151.2
|
Ppp4r4
|
protein phosphatase 4, regulatory subunit 4 |
| chrX_-_42256694 | 3.38 |
ENSMUST00000115058.8
ENSMUST00000115059.8 |
Tenm1
|
teneurin transmembrane protein 1 |
| chr16_-_22258469 | 3.35 |
ENSMUST00000079601.13
|
Etv5
|
ets variant 5 |
| chr10_+_127226180 | 3.10 |
ENSMUST00000077046.12
ENSMUST00000105250.9 |
R3hdm2
|
R3H domain containing 2 |
| chr10_-_57408512 | 3.08 |
ENSMUST00000169122.8
|
Serinc1
|
serine incorporator 1 |
| chr5_+_114427227 | 2.93 |
ENSMUST00000169347.6
|
Myo1h
|
myosin 1H |
| chr1_+_179788037 | 2.92 |
ENSMUST00000097453.9
ENSMUST00000111117.8 |
Cdc42bpa
|
CDC42 binding protein kinase alpha |
| chr4_-_36056726 | 2.87 |
ENSMUST00000108124.4
|
Lingo2
|
leucine rich repeat and Ig domain containing 2 |
| chr15_+_100768806 | 2.86 |
ENSMUST00000201549.4
ENSMUST00000108908.6 |
Scn8a
|
sodium channel, voltage-gated, type VIII, alpha |
| chr15_+_100768776 | 2.86 |
ENSMUST00000108909.9
|
Scn8a
|
sodium channel, voltage-gated, type VIII, alpha |
| chr16_-_22258320 | 2.73 |
ENSMUST00000170393.2
|
Etv5
|
ets variant 5 |
| chrX_+_152506577 | 2.72 |
ENSMUST00000140575.8
ENSMUST00000208373.2 ENSMUST00000185492.7 ENSMUST00000149514.8 |
Nbdy
|
negative regulator of P-body association |
| chr10_-_75946790 | 2.60 |
ENSMUST00000120757.2
|
Slc5a4b
|
solute carrier family 5 (neutral amino acid transporters, system A), member 4b |
| chr13_+_83720484 | 2.45 |
ENSMUST00000196207.5
|
Mef2c
|
myocyte enhancer factor 2C |
| chrX_-_87159237 | 2.41 |
ENSMUST00000113966.8
ENSMUST00000113964.2 |
Il1rapl1
|
interleukin 1 receptor accessory protein-like 1 |
| chr19_-_57227742 | 2.40 |
ENSMUST00000111559.8
|
Ablim1
|
actin-binding LIM protein 1 |
| chr4_+_42114817 | 2.40 |
ENSMUST00000098123.4
|
Gm13304
|
predicted gene 13304 |
| chr15_+_100768551 | 2.37 |
ENSMUST00000082209.13
|
Scn8a
|
sodium channel, voltage-gated, type VIII, alpha |
| chr4_+_41903610 | 2.36 |
ENSMUST00000098128.4
|
Ccl21d
|
chemokine (C-C motif) ligand 21D |
| chr4_+_42255693 | 2.22 |
ENSMUST00000178864.3
|
Ccl21b
|
chemokine (C-C motif) ligand 21B (leucine) |
| chr3_+_68776884 | 2.14 |
ENSMUST00000054551.3
|
1110032F04Rik
|
RIKEN cDNA 1110032F04 gene |
| chr9_-_105398346 | 2.12 |
ENSMUST00000176770.8
ENSMUST00000085133.13 |
Atp2c1
|
ATPase, Ca++-sequestering |
| chr18_+_57601541 | 2.10 |
ENSMUST00000091892.4
ENSMUST00000209782.2 |
Ctxn3
|
cortexin 3 |
| chr7_-_12552764 | 2.04 |
ENSMUST00000108546.2
ENSMUST00000072222.8 |
Zfp329
|
zinc finger protein 329 |
| chr11_-_73215442 | 2.01 |
ENSMUST00000021119.9
|
Aspa
|
aspartoacylase |
| chr12_-_72964646 | 2.00 |
ENSMUST00000044000.12
|
4930447C04Rik
|
RIKEN cDNA 4930447C04 gene |
| chr7_-_85895409 | 1.97 |
ENSMUST00000165771.2
ENSMUST00000233075.2 ENSMUST00000233317.2 ENSMUST00000233312.2 ENSMUST00000233928.2 ENSMUST00000232799.2 ENSMUST00000233744.2 |
Vmn2r76
|
vomeronasal 2, receptor 76 |
| chr4_+_41569775 | 1.95 |
ENSMUST00000102963.10
|
Dnai1
|
dynein axonemal intermediate chain 1 |
| chr11_-_50732373 | 1.83 |
ENSMUST00000109133.2
ENSMUST00000109134.8 ENSMUST00000049625.2 |
Zfp879
|
zinc finger protein 879 |
| chr10_-_38998272 | 1.81 |
ENSMUST00000136546.8
|
Fam229b
|
family with sequence similarity 229, member B |
| chr7_+_107702486 | 1.79 |
ENSMUST00000104917.4
|
Olfr483
|
olfactory receptor 483 |
| chr2_+_67276338 | 1.79 |
ENSMUST00000239060.2
ENSMUST00000028410.4 ENSMUST00000112347.8 ENSMUST00000238878.2 |
Xirp2
|
xin actin-binding repeat containing 2 |
| chr15_+_100768507 | 1.72 |
ENSMUST00000201518.4
ENSMUST00000200933.4 |
Scn8a
|
sodium channel, voltage-gated, type VIII, alpha |
| chr7_-_103320398 | 1.70 |
ENSMUST00000062144.4
|
Olfr624
|
olfactory receptor 624 |
| chr2_+_81883566 | 1.62 |
ENSMUST00000047527.8
|
Zfp804a
|
zinc finger protein 804A |
| chr12_-_59108211 | 1.56 |
ENSMUST00000217863.2
ENSMUST00000021380.10 |
Trappc6b
|
trafficking protein particle complex 6B |
| chr3_-_63391300 | 1.51 |
ENSMUST00000192926.2
|
Strit1
|
small transmembrane regulator of ion transport 1 |
| chr17_+_18269686 | 1.49 |
ENSMUST00000176802.3
|
Vmn2r124
|
vomeronasal 2, receptor 124 |
| chr3_-_126918491 | 1.39 |
ENSMUST00000238781.2
|
Ank2
|
ankyrin 2, brain |
| chr6_-_145156517 | 1.29 |
ENSMUST00000111728.8
ENSMUST00000204105.2 ENSMUST00000060797.10 |
Casc1
|
cancer susceptibility candidate 1 |
| chr10_+_97482946 | 1.29 |
ENSMUST00000105285.4
|
Epyc
|
epiphycan |
| chr2_-_88990751 | 1.25 |
ENSMUST00000216833.2
ENSMUST00000216976.2 |
Olfr1224-ps1
|
olfactory receptor 1224, pseudogene 1 |
| chr6_-_113696809 | 1.23 |
ENSMUST00000203770.3
ENSMUST00000064993.8 |
Ghrl
|
ghrelin |
| chr15_-_81074921 | 1.23 |
ENSMUST00000131235.9
ENSMUST00000134469.9 ENSMUST00000239114.2 ENSMUST00000149582.8 |
Mrtfa
|
myocardin related transcription factor A |
| chr2_+_173579285 | 1.22 |
ENSMUST00000067530.6
|
Vapb
|
vesicle-associated membrane protein, associated protein B and C |
| chr11_+_93934940 | 1.22 |
ENSMUST00000132079.8
|
Spag9
|
sperm associated antigen 9 |
| chr1_+_106866678 | 1.20 |
ENSMUST00000112724.3
|
Serpinb12
|
serine (or cysteine) peptidase inhibitor, clade B (ovalbumin), member 12 |
| chr18_+_44237474 | 1.11 |
ENSMUST00000081271.7
|
Spink12
|
serine peptidase inhibitor, Kazal type 12 |
| chr7_-_5041427 | 1.09 |
ENSMUST00000062428.5
|
Zfp784
|
zinc finger protein 784 |
| chr8_+_107662352 | 1.09 |
ENSMUST00000212524.2
ENSMUST00000047425.5 |
Sntb2
|
syntrophin, basic 2 |
| chr11_-_74615496 | 1.08 |
ENSMUST00000021091.15
|
Pafah1b1
|
platelet-activating factor acetylhydrolase, isoform 1b, subunit 1 |
| chr6_-_101176147 | 1.08 |
ENSMUST00000239140.2
|
Pdzrn3
|
PDZ domain containing RING finger 3 |
| chr5_-_146157711 | 1.08 |
ENSMUST00000169407.9
ENSMUST00000161331.8 ENSMUST00000159074.3 ENSMUST00000067837.10 |
Rnf6
|
ring finger protein (C3H2C3 type) 6 |
| chr8_+_11808349 | 1.06 |
ENSMUST00000074856.13
ENSMUST00000098938.9 |
Arhgef7
|
Rho guanine nucleotide exchange factor (GEF7) |
| chr7_+_92390811 | 1.05 |
ENSMUST00000032879.15
|
Rab30
|
RAB30, member RAS oncogene family |
| chr4_+_136150835 | 1.01 |
ENSMUST00000088677.5
ENSMUST00000121571.8 ENSMUST00000117699.2 |
Htr1d
|
5-hydroxytryptamine (serotonin) receptor 1D |
| chr18_+_65831324 | 0.99 |
ENSMUST00000115097.8
ENSMUST00000117694.2 ENSMUST00000235962.2 |
Oacyl
|
O-acyltransferase like |
| chr9_-_58109627 | 0.99 |
ENSMUST00000216231.2
|
Islr2
|
immunoglobulin superfamily containing leucine-rich repeat 2 |
| chr5_-_108943211 | 0.95 |
ENSMUST00000004943.2
|
Tmed11
|
transmembrane p24 trafficking protein 11 |
| chr14_-_96756503 | 0.88 |
ENSMUST00000022666.9
|
Klhl1
|
kelch-like 1 |
| chr9_+_19533591 | 0.82 |
ENSMUST00000215372.2
|
Zfp317
|
zinc finger protein 317 |
| chr18_+_22478146 | 0.80 |
ENSMUST00000120223.8
ENSMUST00000097655.4 |
Asxl3
|
ASXL transcriptional regulator 3 |
| chr17_+_87270707 | 0.78 |
ENSMUST00000139344.2
|
Rhoq
|
ras homolog family member Q |
| chr7_+_86564302 | 0.77 |
ENSMUST00000233033.2
ENSMUST00000170835.4 |
Vmn2r78
|
vomeronasal 2, receptor 78 |
| chr13_-_23372145 | 0.74 |
ENSMUST00000228239.2
ENSMUST00000227950.2 ENSMUST00000226651.2 ENSMUST00000227679.2 ENSMUST00000228854.2 |
Vmn1r220
|
vomeronasal 1 receptor 220 |
| chr10_+_57521958 | 0.70 |
ENSMUST00000177473.8
|
Pkib
|
protein kinase inhibitor beta, cAMP dependent, testis specific |
| chr7_-_48105989 | 0.68 |
ENSMUST00000188918.2
|
Mrgprb1
|
MAS-related GPR, member B1 |
| chr9_-_58109526 | 0.66 |
ENSMUST00000216297.2
|
Islr2
|
immunoglobulin superfamily containing leucine-rich repeat 2 |
| chr17_+_87270504 | 0.65 |
ENSMUST00000024956.15
|
Rhoq
|
ras homolog family member Q |
| chr9_+_19533374 | 0.63 |
ENSMUST00000213725.2
ENSMUST00000208694.2 |
Zfp317
|
zinc finger protein 317 |
| chr2_+_110551927 | 0.60 |
ENSMUST00000111017.9
|
Muc15
|
mucin 15 |
| chr4_+_13784749 | 0.60 |
ENSMUST00000098256.4
|
Runx1t1
|
RUNX1 translocation partner 1 |
| chr17_-_59298338 | 0.59 |
ENSMUST00000025064.14
|
Pdzph1
|
PDZ and pleckstrin homology domains 1 |
| chr7_-_90125178 | 0.59 |
ENSMUST00000032843.9
|
Tmem126b
|
transmembrane protein 126B |
| chr4_+_155048571 | 0.57 |
ENSMUST00000030931.11
ENSMUST00000070953.11 |
Pank4
|
pantothenate kinase 4 |
| chr11_+_93935021 | 0.56 |
ENSMUST00000075695.13
ENSMUST00000092777.11 |
Spag9
|
sperm associated antigen 9 |
| chr2_+_30254239 | 0.53 |
ENSMUST00000077977.14
ENSMUST00000140075.9 ENSMUST00000142801.8 ENSMUST00000100214.10 |
Miga2
|
mitoguardin 2 |
| chrX_-_153362356 | 0.48 |
ENSMUST00000101137.3
|
Samt2
|
spermatogenesis associated multipass transmembrane protein 2 |
| chr17_-_22127073 | 0.48 |
ENSMUST00000106026.3
|
Zfp995
|
zinc finger protein 995 |
| chr7_-_85525474 | 0.46 |
ENSMUST00000077478.8
ENSMUST00000233729.2 ENSMUST00000233079.2 ENSMUST00000233315.2 ENSMUST00000233115.2 ENSMUST00000233748.2 ENSMUST00000232965.2 ENSMUST00000232897.2 |
Vmn2r73
|
vomeronasal 2, receptor 73 |
| chr10_+_129336682 | 0.46 |
ENSMUST00000056002.5
ENSMUST00000218901.2 |
Olfr790
|
olfactory receptor 790 |
| chr2_+_154149122 | 0.40 |
ENSMUST00000088921.6
|
Bpifb9b
|
BPI fold containing family B, member 9B |
| chrX_+_95498965 | 0.40 |
ENSMUST00000033553.14
|
Heph
|
hephaestin |
| chr13_+_104424359 | 0.39 |
ENSMUST00000065766.7
|
Adamts6
|
a disintegrin-like and metallopeptidase (reprolysin type) with thrombospondin type 1 motif, 6 |
| chr13_-_21257469 | 0.38 |
ENSMUST00000058168.2
|
Olfr1370
|
olfactory receptor 1370 |
| chr10_-_129509659 | 0.36 |
ENSMUST00000213294.2
ENSMUST00000216067.3 ENSMUST00000203424.2 |
Olfr801
|
olfactory receptor 801 |
| chr2_-_88534814 | 0.35 |
ENSMUST00000216928.2
ENSMUST00000216977.2 |
Olfr1196
|
olfactory receptor 1196 |
| chr10_+_57521930 | 0.35 |
ENSMUST00000177325.8
|
Pkib
|
protein kinase inhibitor beta, cAMP dependent, testis specific |
| chr7_+_104274001 | 0.33 |
ENSMUST00000215454.3
ENSMUST00000213297.2 |
Olfr657
|
olfactory receptor 657 |
| chr8_-_86091946 | 0.33 |
ENSMUST00000034133.14
|
Mylk3
|
myosin light chain kinase 3 |
| chr13_-_22686415 | 0.28 |
ENSMUST00000078642.2
|
Vmn1r202
|
vomeronasal 1 receptor 202 |
| chr2_-_85266670 | 0.27 |
ENSMUST00000214679.2
ENSMUST00000217218.3 |
Olfr994
|
olfactory receptor 994 |
| chr19_+_13211616 | 0.25 |
ENSMUST00000064102.2
|
Olfr1463
|
olfactory receptor 1463 |
| chr1_-_5987617 | 0.25 |
ENSMUST00000044180.5
|
Npbwr1
|
neuropeptides B/W receptor 1 |
| chr11_+_93935066 | 0.22 |
ENSMUST00000103168.10
|
Spag9
|
sperm associated antigen 9 |
| chr1_+_43571536 | 0.22 |
ENSMUST00000202540.2
|
Nck2
|
non-catalytic region of tyrosine kinase adaptor protein 2 |
| chr6_-_57991216 | 0.20 |
ENSMUST00000228951.2
|
Vmn1r26
|
vomeronasal 1 receptor 26 |
| chr2_-_88600832 | 0.19 |
ENSMUST00000217588.3
|
Olfr1200
|
olfactory receptor 1200 |
| chr11_+_93935156 | 0.19 |
ENSMUST00000024979.15
|
Spag9
|
sperm associated antigen 9 |
| chr9_+_39722280 | 0.13 |
ENSMUST00000213975.2
|
Olfr970
|
olfactory receptor 970 |
| chr17_+_36857967 | 0.12 |
ENSMUST00000041964.7
|
H2-M11
|
histocompatibility 2, M region locus 11 |
| chr11_+_102775991 | 0.10 |
ENSMUST00000100369.4
|
Fam187a
|
family with sequence similarity 187, member A |
| chr6_+_66708376 | 0.09 |
ENSMUST00000077482.2
|
Vmn1r37
|
vomeronasal 1 receptor 37 |
| chrX_+_20529137 | 0.08 |
ENSMUST00000001989.9
|
Uba1
|
ubiquitin-like modifier activating enzyme 1 |
| chr3_+_85946145 | 0.07 |
ENSMUST00000238331.2
|
Sh3d19
|
SH3 domain protein D19 |
| chr7_-_103217838 | 0.04 |
ENSMUST00000214806.2
|
Olfr616
|
olfactory receptor 616 |
| chr7_+_101530524 | 0.04 |
ENSMUST00000106978.8
|
Anapc15
|
anaphase promoting complex C subunit 15 |
| chr15_+_84076423 | 0.04 |
ENSMUST00000023071.8
|
Samm50
|
SAMM50 sorting and assembly machinery component |
| chr11_-_69686756 | 0.04 |
ENSMUST00000045971.9
|
Chrnb1
|
cholinergic receptor, nicotinic, beta polypeptide 1 (muscle) |
| chr6_-_134864731 | 0.04 |
ENSMUST00000203762.4
ENSMUST00000215088.2 ENSMUST00000066107.10 |
Gpr19
|
G protein-coupled receptor 19 |
| chr16_+_25620652 | 0.03 |
ENSMUST00000115304.8
ENSMUST00000115305.2 ENSMUST00000040231.13 ENSMUST00000115306.8 |
Trp63
|
transformation related protein 63 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.8 | 7.4 | GO:0035502 | metanephric part of ureteric bud development(GO:0035502) |
| 1.7 | 10.1 | GO:0060160 | negative regulation of dopamine receptor signaling pathway(GO:0060160) |
| 1.5 | 8.8 | GO:1902958 | neuron intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:0036482) positive regulation of macromitophagy(GO:1901526) positive regulation of mitochondrial electron transport, NADH to ubiquinone(GO:1902958) regulation of hydrogen peroxide-induced neuron intrinsic apoptotic signaling pathway(GO:1903383) negative regulation of hydrogen peroxide-induced neuron intrinsic apoptotic signaling pathway(GO:1903384) positive regulation of mitophagy in response to mitochondrial depolarization(GO:1904925) |
| 1.4 | 4.3 | GO:0008628 | hormone-mediated apoptotic signaling pathway(GO:0008628) |
| 1.4 | 10.0 | GO:0061743 | motor learning(GO:0061743) |
| 1.1 | 5.6 | GO:0055005 | ventricular cardiac myofibril assembly(GO:0055005) |
| 1.0 | 4.1 | GO:1905053 | regulation of base-excision repair(GO:1905051) positive regulation of base-excision repair(GO:1905053) |
| 0.9 | 4.4 | GO:0044861 | protein transport into plasma membrane raft(GO:0044861) |
| 0.9 | 3.5 | GO:0080163 | regulation of protein serine/threonine phosphatase activity(GO:0080163) |
| 0.8 | 5.8 | GO:0001757 | somite specification(GO:0001757) |
| 0.8 | 9.5 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.8 | 6.4 | GO:0014004 | microglia differentiation(GO:0014004) microglia development(GO:0014005) |
| 0.8 | 17.5 | GO:0046069 | cGMP catabolic process(GO:0046069) |
| 0.8 | 3.9 | GO:2000360 | negative regulation of binding of sperm to zona pellucida(GO:2000360) |
| 0.8 | 7.6 | GO:1903690 | negative regulation of wound healing, spreading of epidermal cells(GO:1903690) |
| 0.8 | 8.4 | GO:1900383 | regulation of synaptic plasticity by receptor localization to synapse(GO:1900383) |
| 0.8 | 10.6 | GO:0019227 | neuronal action potential propagation(GO:0019227) action potential propagation(GO:0098870) |
| 0.7 | 2.0 | GO:0010705 | meiotic DNA double-strand break processing involved in reciprocal meiotic recombination(GO:0010705) |
| 0.6 | 6.1 | GO:0060762 | regulation of branching involved in mammary gland duct morphogenesis(GO:0060762) |
| 0.6 | 11.5 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.5 | 8.0 | GO:0061303 | cornea development in camera-type eye(GO:0061303) |
| 0.5 | 2.1 | GO:0055071 | cellular manganese ion homeostasis(GO:0030026) Golgi calcium ion homeostasis(GO:0032468) manganese ion homeostasis(GO:0055071) |
| 0.5 | 3.6 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) cerebral cortex GABAergic interneuron migration(GO:0021853) interneuron migration(GO:1904936) |
| 0.5 | 3.9 | GO:0016560 | protein import into peroxisome matrix, docking(GO:0016560) |
| 0.4 | 3.1 | GO:1904222 | regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904217) positive regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904219) positive regulation of serine C-palmitoyltransferase activity(GO:1904222) |
| 0.4 | 8.5 | GO:1902004 | positive regulation of beta-amyloid formation(GO:1902004) |
| 0.4 | 3.7 | GO:0071787 | endoplasmic reticulum tubular network assembly(GO:0071787) |
| 0.4 | 20.6 | GO:0007616 | long-term memory(GO:0007616) |
| 0.3 | 10.6 | GO:0007097 | nuclear migration(GO:0007097) establishment of nucleus localization(GO:0040023) |
| 0.3 | 1.2 | GO:0044830 | modulation by host of viral RNA genome replication(GO:0044830) negative regulation by virus of viral protein levels in host cell(GO:0046725) |
| 0.3 | 2.0 | GO:0006083 | acetate metabolic process(GO:0006083) |
| 0.3 | 2.2 | GO:0090074 | negative regulation of protein homodimerization activity(GO:0090074) |
| 0.3 | 4.3 | GO:0008343 | adult feeding behavior(GO:0008343) |
| 0.2 | 18.3 | GO:0019228 | neuronal action potential(GO:0019228) |
| 0.2 | 4.0 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.2 | 2.4 | GO:0003138 | primary heart field specification(GO:0003138) sinoatrial valve development(GO:0003172) sinoatrial valve morphogenesis(GO:0003185) |
| 0.2 | 10.0 | GO:0035025 | positive regulation of Rho protein signal transduction(GO:0035025) |
| 0.2 | 2.6 | GO:1904659 | glucose transmembrane transport(GO:1904659) |
| 0.2 | 10.6 | GO:0042982 | amyloid precursor protein metabolic process(GO:0042982) |
| 0.2 | 4.7 | GO:0001553 | luteinization(GO:0001553) |
| 0.2 | 2.4 | GO:0097105 | presynaptic membrane assembly(GO:0097105) |
| 0.2 | 7.2 | GO:0007212 | dopamine receptor signaling pathway(GO:0007212) |
| 0.2 | 7.5 | GO:0007026 | negative regulation of microtubule depolymerization(GO:0007026) |
| 0.2 | 1.4 | GO:0036309 | protein localization to M-band(GO:0036309) protein localization to T-tubule(GO:0036371) |
| 0.2 | 1.1 | GO:0044314 | protein K27-linked ubiquitination(GO:0044314) |
| 0.2 | 4.9 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.1 | 3.5 | GO:0034453 | microtubule anchoring(GO:0034453) |
| 0.1 | 10.4 | GO:0010765 | positive regulation of sodium ion transport(GO:0010765) |
| 0.1 | 1.0 | GO:0014827 | intestine smooth muscle contraction(GO:0014827) |
| 0.1 | 6.2 | GO:0060997 | dendritic spine morphogenesis(GO:0060997) |
| 0.1 | 4.9 | GO:0009395 | phospholipid catabolic process(GO:0009395) |
| 0.1 | 5.6 | GO:0045773 | positive regulation of axon extension(GO:0045773) |
| 0.1 | 6.5 | GO:0070936 | protein K48-linked ubiquitination(GO:0070936) |
| 0.1 | 17.4 | GO:0007416 | synapse assembly(GO:0007416) |
| 0.1 | 1.1 | GO:2000394 | positive regulation of lamellipodium morphogenesis(GO:2000394) |
| 0.1 | 5.4 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.1 | 3.4 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.1 | 1.1 | GO:2000480 | negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
| 0.0 | 1.4 | GO:1903077 | negative regulation of protein localization to plasma membrane(GO:1903077) negative regulation of protein localization to cell periphery(GO:1904376) |
| 0.0 | 0.2 | GO:1903898 | positive regulation of translation in response to endoplasmic reticulum stress(GO:0036493) negative regulation of PERK-mediated unfolded protein response(GO:1903898) |
| 0.0 | 0.3 | GO:0060298 | regulation of vascular permeability involved in acute inflammatory response(GO:0002528) positive regulation of sarcomere organization(GO:0060298) |
| 0.0 | 4.4 | GO:0003281 | ventricular septum development(GO:0003281) |
| 0.0 | 4.5 | GO:0043488 | regulation of mRNA stability(GO:0043488) |
| 0.0 | 1.7 | GO:0070098 | chemokine-mediated signaling pathway(GO:0070098) |
| 0.0 | 0.0 | GO:0010482 | epidermal cell division(GO:0010481) regulation of epidermal cell division(GO:0010482) cloacal septation(GO:0060197) squamous basal epithelial stem cell differentiation involved in prostate gland acinus development(GO:0060529) |
| 0.0 | 0.6 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.0 | 2.4 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.0 | 3.3 | GO:0031532 | actin cytoskeleton reorganization(GO:0031532) |
| 0.0 | 0.9 | GO:0021680 | cerebellar Purkinje cell layer development(GO:0021680) |
| 0.0 | 2.3 | GO:0002244 | hematopoietic progenitor cell differentiation(GO:0002244) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.5 | 7.6 | GO:1903754 | cortical microtubule plus-end(GO:1903754) cytoplasmic microtubule plus-end(GO:1904511) |
| 0.8 | 7.5 | GO:0097442 | CA3 pyramidal cell dendrite(GO:0097442) |
| 0.8 | 20.4 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.8 | 10.1 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.8 | 3.9 | GO:0030312 | external encapsulating structure(GO:0030312) |
| 0.7 | 8.8 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.6 | 5.8 | GO:1990357 | terminal web(GO:1990357) |
| 0.5 | 3.9 | GO:0017071 | intracellular cyclic nucleotide activated cation channel complex(GO:0017071) |
| 0.4 | 4.4 | GO:0044352 | pinosome(GO:0044352) macropinosome(GO:0044354) |
| 0.3 | 4.5 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.3 | 8.5 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.2 | 4.3 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.2 | 12.8 | GO:0043034 | costamere(GO:0043034) |
| 0.2 | 8.0 | GO:0005790 | smooth endoplasmic reticulum(GO:0005790) |
| 0.2 | 23.7 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.2 | 2.0 | GO:0000801 | central element(GO:0000801) |
| 0.2 | 4.3 | GO:0043196 | varicosity(GO:0043196) |
| 0.1 | 7.7 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.1 | 3.7 | GO:0071782 | endoplasmic reticulum tubular network(GO:0071782) |
| 0.1 | 17.4 | GO:0042641 | actomyosin(GO:0042641) |
| 0.1 | 1.1 | GO:0000322 | storage vacuole(GO:0000322) |
| 0.1 | 7.2 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.1 | 1.1 | GO:0000235 | astral microtubule(GO:0000235) |
| 0.1 | 10.0 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.1 | 1.6 | GO:0030008 | TRAPP complex(GO:0030008) |
| 0.1 | 14.9 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.1 | 7.4 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.1 | 4.9 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.0 | 1.6 | GO:0016459 | myosin complex(GO:0016459) |
| 0.0 | 6.4 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 6.9 | GO:0030016 | myofibril(GO:0030016) |
| 0.0 | 1.0 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.0 | 6.0 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 21.2 | GO:0005887 | integral component of plasma membrane(GO:0005887) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 6.9 | 20.6 | GO:0016232 | HNK-1 sulfotransferase activity(GO:0016232) |
| 1.7 | 8.5 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 1.6 | 17.5 | GO:0004117 | calmodulin-dependent cyclic-nucleotide phosphodiesterase activity(GO:0004117) calcium- and calmodulin-regulated 3',5'-cyclic-GMP phosphodiesterase activity(GO:0048101) |
| 1.4 | 10.1 | GO:0031750 | D3 dopamine receptor binding(GO:0031750) |
| 1.1 | 4.3 | GO:0051431 | corticotropin-releasing hormone receptor 2 binding(GO:0051431) |
| 1.0 | 8.8 | GO:0055131 | C3HC4-type RING finger domain binding(GO:0055131) |
| 0.9 | 7.4 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.9 | 7.2 | GO:0031821 | G-protein coupled serotonin receptor binding(GO:0031821) |
| 0.9 | 11.5 | GO:0031802 | type 5 metabotropic glutamate receptor binding(GO:0031802) |
| 0.8 | 7.5 | GO:0005519 | cytoskeletal regulatory protein binding(GO:0005519) |
| 0.7 | 2.0 | GO:0019807 | aspartoacylase activity(GO:0019807) |
| 0.7 | 20.4 | GO:0031402 | sodium ion binding(GO:0031402) |
| 0.6 | 3.9 | GO:0005052 | peroxisome matrix targeting signal-1 binding(GO:0005052) |
| 0.5 | 2.1 | GO:0015410 | manganese-transporting ATPase activity(GO:0015410) |
| 0.4 | 7.6 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.4 | 9.5 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.4 | 3.9 | GO:0004568 | chitinase activity(GO:0004568) |
| 0.4 | 1.2 | GO:0033149 | FFAT motif binding(GO:0033149) |
| 0.3 | 10.1 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.3 | 5.3 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.2 | 3.1 | GO:0022889 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.2 | 0.7 | GO:0031768 | ghrelin receptor binding(GO:0031768) |
| 0.2 | 1.1 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.2 | 2.6 | GO:0005412 | glucose:sodium symporter activity(GO:0005412) |
| 0.1 | 4.9 | GO:0004623 | phospholipase A2 activity(GO:0004623) |
| 0.1 | 5.5 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.1 | 2.2 | GO:0048273 | MAP-kinase scaffold activity(GO:0005078) mitogen-activated protein kinase p38 binding(GO:0048273) |
| 0.1 | 5.8 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 4.5 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.1 | 1.9 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.1 | 5.6 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.1 | 2.4 | GO:0003680 | AT DNA binding(GO:0003680) |
| 0.1 | 0.6 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 0.1 | 6.6 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.1 | 5.5 | GO:0051393 | alpha-actinin binding(GO:0051393) |
| 0.1 | 1.4 | GO:0005522 | profilin binding(GO:0005522) |
| 0.1 | 5.6 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.1 | 1.0 | GO:0051378 | serotonin binding(GO:0051378) |
| 0.1 | 8.6 | GO:0004860 | protein kinase inhibitor activity(GO:0004860) |
| 0.0 | 7.3 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.0 | 1.7 | GO:0008009 | chemokine activity(GO:0008009) |
| 0.0 | 2.4 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 7.2 | GO:0003774 | motor activity(GO:0003774) |
| 0.0 | 3.5 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
| 0.0 | 0.9 | GO:0042923 | neuropeptide binding(GO:0042923) |
| 0.0 | 6.6 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 0.4 | GO:0016724 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.0 | 3.7 | GO:0004843 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) |
| 0.0 | 1.4 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.0 | 9.5 | GO:0000287 | magnesium ion binding(GO:0000287) |
| 0.0 | 0.3 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.0 | 19.8 | GO:0005198 | structural molecule activity(GO:0005198) |
| 0.0 | 1.3 | GO:0048487 | beta-tubulin binding(GO:0048487) |
| 0.0 | 1.1 | GO:0050681 | androgen receptor binding(GO:0050681) |
| 0.0 | 5.9 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.0 | 3.5 | GO:0008017 | microtubule binding(GO:0008017) |
| 0.0 | 0.1 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.0 | 1.3 | GO:0005550 | pheromone binding(GO:0005550) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 7.2 | PID S1P S1P4 PATHWAY | S1P4 pathway |
| 0.3 | 17.4 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.2 | 9.9 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.1 | 5.5 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.1 | 8.1 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.1 | 10.0 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.1 | 4.4 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.1 | 6.2 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.1 | 9.5 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.1 | 15.9 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.1 | 1.4 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.0 | 3.7 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 2.2 | PID ARF6 TRAFFICKING PATHWAY | Arf6 trafficking events |
| 0.0 | 3.6 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 17.5 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.2 | 10.4 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.2 | 9.8 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.2 | 8.1 | REACTOME ERK MAPK TARGETS | Genes involved in ERK/MAPK targets |
| 0.1 | 5.4 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.1 | 7.2 | REACTOME G PROTEIN ACTIVATION | Genes involved in G-protein activation |
| 0.1 | 8.5 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 1.0 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.1 | 2.4 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |
| 0.0 | 4.3 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.6 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.0 | 0.7 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.0 | 3.7 | REACTOME P75 NTR RECEPTOR MEDIATED SIGNALLING | Genes involved in p75 NTR receptor-mediated signalling |
| 0.0 | 1.2 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.2 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.0 | 2.0 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.0 | 2.3 | REACTOME G ALPHA Q SIGNALLING EVENTS | Genes involved in G alpha (q) signalling events |