Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
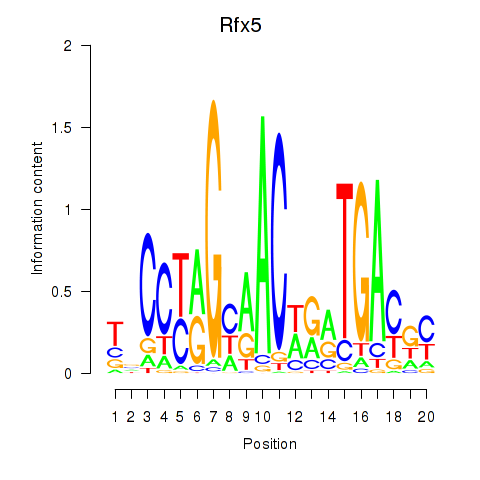
Results for Rfx5
Z-value: 1.75
Transcription factors associated with Rfx5
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Rfx5
|
ENSMUSG00000005774.13 | Rfx5 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Rfx5 | mm39_v1_chr3_+_94861386_94861537 | 0.49 | 1.1e-05 | Click! |
Activity profile of Rfx5 motif
Sorted Z-values of Rfx5 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Rfx5
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_+_34617784 | 27.22 |
ENSMUST00000114175.8
ENSMUST00000139063.8 ENSMUST00000078615.12 |
BC051142
|
cDNA sequence BC051142 |
| chr17_+_34364206 | 20.46 |
ENSMUST00000041982.9
ENSMUST00000171231.8 |
H2-DMb2
|
histocompatibility 2, class II, locus Mb2 |
| chr19_+_37364791 | 19.68 |
ENSMUST00000012587.4
|
Kif11
|
kinesin family member 11 |
| chr17_+_34457868 | 17.93 |
ENSMUST00000095342.11
ENSMUST00000167280.8 ENSMUST00000236838.2 |
H2-Ob
|
histocompatibility 2, O region beta locus |
| chr17_+_34482183 | 17.41 |
ENSMUST00000040828.7
ENSMUST00000237342.2 ENSMUST00000237866.2 |
H2-Ab1
|
histocompatibility 2, class II antigen A, beta 1 |
| chr18_+_60936910 | 17.38 |
ENSMUST00000097563.9
ENSMUST00000050487.16 ENSMUST00000167610.2 |
Cd74
|
CD74 antigen (invariant polypeptide of major histocompatibility complex, class II antigen-associated) |
| chr17_+_34416707 | 17.35 |
ENSMUST00000025196.9
|
Psmb8
|
proteasome (prosome, macropain) subunit, beta type 8 (large multifunctional peptidase 7) |
| chr7_+_126690525 | 15.40 |
ENSMUST00000056288.7
ENSMUST00000206102.2 |
AI467606
|
expressed sequence AI467606 |
| chr17_+_34524841 | 15.26 |
ENSMUST00000235530.2
|
H2-Eb1
|
histocompatibility 2, class II antigen E beta |
| chr17_+_34524884 | 14.89 |
ENSMUST00000074557.11
|
H2-Eb1
|
histocompatibility 2, class II antigen E beta |
| chr17_+_34372046 | 14.60 |
ENSMUST00000114232.4
|
H2-DMb1
|
histocompatibility 2, class II, locus Mb1 |
| chr17_+_34416689 | 14.03 |
ENSMUST00000173441.9
|
Psmb8
|
proteasome (prosome, macropain) subunit, beta type 8 (large multifunctional peptidase 7) |
| chr17_+_36172210 | 13.64 |
ENSMUST00000074259.15
ENSMUST00000174873.2 |
Nrm
|
nurim (nuclear envelope membrane protein) |
| chr17_+_35658131 | 12.39 |
ENSMUST00000071951.14
ENSMUST00000116598.10 ENSMUST00000078205.14 ENSMUST00000076256.8 |
H2-Q7
|
histocompatibility 2, Q region locus 7 |
| chr17_+_34544632 | 11.84 |
ENSMUST00000050325.10
|
H2-Eb2
|
histocompatibility 2, class II antigen E beta2 |
| chr17_+_34406523 | 11.83 |
ENSMUST00000170086.8
|
Tap1
|
transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) |
| chr17_+_35598583 | 11.57 |
ENSMUST00000081435.5
|
H2-Q4
|
histocompatibility 2, Q region locus 4 |
| chr17_+_34406762 | 11.57 |
ENSMUST00000041633.15
|
Tap1
|
transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) |
| chr17_-_37794434 | 11.14 |
ENSMUST00000016427.11
ENSMUST00000171139.3 |
H2-M2
|
histocompatibility 2, M region locus 2 |
| chr19_-_10460238 | 11.09 |
ENSMUST00000235392.2
ENSMUST00000237522.2 ENSMUST00000038842.5 |
Ppp1r32
|
protein phosphatase 1, regulatory subunit 32 |
| chr17_+_36172235 | 10.77 |
ENSMUST00000172931.2
|
Nrm
|
nurim (nuclear envelope membrane protein) |
| chr7_-_30364394 | 10.40 |
ENSMUST00000019697.9
|
Haus5
|
HAUS augmin-like complex, subunit 5 |
| chr10_-_128579879 | 9.86 |
ENSMUST00000026414.9
|
Dgka
|
diacylglycerol kinase, alpha |
| chr17_-_34219225 | 9.79 |
ENSMUST00000238098.2
ENSMUST00000087189.7 ENSMUST00000173075.3 ENSMUST00000172912.8 ENSMUST00000236740.2 ENSMUST00000025181.18 |
H2-K1
|
histocompatibility 2, K1, K region |
| chr17_+_34354556 | 9.36 |
ENSMUST00000236322.2
ENSMUST00000237402.2 |
H2-DMa
|
histocompatibility 2, class II, locus DMa |
| chr11_+_106167541 | 8.86 |
ENSMUST00000044462.4
|
Tcam1
|
testicular cell adhesion molecule 1 |
| chr17_-_34406193 | 8.69 |
ENSMUST00000173831.3
|
Psmb9
|
proteasome (prosome, macropain) subunit, beta type 9 (large multifunctional peptidase 2) |
| chr7_-_24145107 | 8.49 |
ENSMUST00000205776.2
|
Irgc1
|
immunity-related GTPase family, cinema 1 |
| chr17_+_37581103 | 8.21 |
ENSMUST00000038580.7
|
H2-M3
|
histocompatibility 2, M region locus 3 |
| chr14_+_40794817 | 7.62 |
ENSMUST00000189865.7
|
Dydc1
|
DPY30 domain containing 1 |
| chr9_-_21202545 | 7.51 |
ENSMUST00000215619.2
|
Cdkn2d
|
cyclin dependent kinase inhibitor 2D |
| chr4_-_126219465 | 7.50 |
ENSMUST00000102616.8
|
Tekt2
|
tektin 2 |
| chr6_-_124756645 | 7.27 |
ENSMUST00000147669.2
ENSMUST00000128697.8 ENSMUST00000032218.10 ENSMUST00000112475.9 |
Lrrc23
|
leucine rich repeat containing 23 |
| chr7_+_44926925 | 7.26 |
ENSMUST00000210861.2
|
Slc6a21
|
solute carrier family 6 member 21 |
| chr9_-_21202353 | 6.47 |
ENSMUST00000086374.8
|
Cdkn2d
|
cyclin dependent kinase inhibitor 2D |
| chr4_-_126219434 | 6.39 |
ENSMUST00000131113.8
|
Tekt2
|
tektin 2 |
| chr9_-_21202693 | 6.31 |
ENSMUST00000213407.2
|
Cdkn2d
|
cyclin dependent kinase inhibitor 2D |
| chr2_-_28453374 | 6.30 |
ENSMUST00000028161.6
|
Cel
|
carboxyl ester lipase |
| chr14_+_40794893 | 6.25 |
ENSMUST00000161837.2
|
Dydc1
|
DPY30 domain containing 1 |
| chr19_+_34194990 | 6.23 |
ENSMUST00000119603.2
|
Stambpl1
|
STAM binding protein like 1 |
| chr14_+_53607470 | 5.80 |
ENSMUST00000103652.5
|
Trav14n-3
|
T cell receptor alpha variable 14N-3 |
| chrX_+_44179759 | 5.78 |
ENSMUST00000040002.4
|
Prr32
|
proline rich 32 |
| chr17_-_25105277 | 5.48 |
ENSMUST00000234583.2
ENSMUST00000234968.2 ENSMUST00000044252.7 |
Nubp2
|
nucleotide binding protein 2 |
| chr16_-_31898088 | 5.39 |
ENSMUST00000023467.9
|
Pak2
|
p21 (RAC1) activated kinase 2 |
| chr5_+_139135966 | 5.07 |
ENSMUST00000026975.11
ENSMUST00000196441.2 |
Dnaaf5
|
dynein, axonemal assembly factor 5 |
| chr5_-_123663440 | 4.82 |
ENSMUST00000197682.6
|
Gm49027
|
predicted gene, 49027 |
| chr5_+_11392646 | 4.65 |
ENSMUST00000179482.2
|
Speer1
|
spermatogenesis associated glutamate (E)-rich protein 1 |
| chr5_+_11821653 | 4.65 |
ENSMUST00000178158.2
|
Gm8926
|
predicted gene 8926 |
| chr5_+_11733203 | 4.39 |
ENSMUST00000178989.2
|
Gm8922
|
predicted gene 8922 |
| chr9_-_51013361 | 4.37 |
ENSMUST00000170947.3
|
Hoatz
|
HOATZ cilia and flagella associated protein |
| chr5_+_11553780 | 4.31 |
ENSMUST00000179375.2
|
Gm8906
|
predicted gene 8906 |
| chrX_+_158480304 | 4.30 |
ENSMUST00000123433.8
|
Sh3kbp1
|
SH3-domain kinase binding protein 1 |
| chr4_-_115638209 | 4.24 |
ENSMUST00000106521.2
|
Tex38
|
testis expressed 38 |
| chr8_-_85794611 | 4.14 |
ENSMUST00000177563.2
|
Gm5741
|
predicted gene 5741 |
| chr5_+_11307124 | 4.11 |
ENSMUST00000177727.2
|
Gm8890
|
predicted gene 8890 |
| chr5_+_11178835 | 4.09 |
ENSMUST00000178863.2
|
Gm8879
|
predicted gene 8879 |
| chr5_+_11467593 | 4.08 |
ENSMUST00000179679.2
|
Gm8897
|
predicted gene 8897 |
| chr8_+_41205245 | 3.96 |
ENSMUST00000096663.5
|
Adam25
|
a disintegrin and metallopeptidase domain 25 (testase 2) |
| chr1_-_182929025 | 3.91 |
ENSMUST00000171366.7
|
Disp1
|
dispatched RND transporter family member 1 |
| chr4_-_116485118 | 3.91 |
ENSMUST00000030456.14
|
Nasp
|
nuclear autoantigenic sperm protein (histone-binding) |
| chr5_+_11234174 | 3.86 |
ENSMUST00000168407.3
|
Gm5861
|
predicted gene 5861 |
| chr8_+_106154266 | 3.84 |
ENSMUST00000067305.8
|
Lrrc36
|
leucine rich repeat containing 36 |
| chr8_-_105236506 | 3.81 |
ENSMUST00000064576.8
|
Terb1
|
telomere repeat binding bouquet formation protein 1 |
| chr10_-_123032821 | 3.40 |
ENSMUST00000219619.2
ENSMUST00000020334.9 |
Usp15
|
ubiquitin specific peptidase 15 |
| chr14_+_53886861 | 3.19 |
ENSMUST00000103660.4
|
Trav15-2-dv6-2
|
T cell receptor alpha variable 15-2-DV6-2 |
| chr8_+_82582953 | 3.17 |
ENSMUST00000109851.3
|
Inpp4b
|
inositol polyphosphate-4-phosphatase, type II |
| chr8_+_41246310 | 2.97 |
ENSMUST00000056331.8
|
Adam20
|
a disintegrin and metallopeptidase domain 20 |
| chr7_-_101570393 | 2.92 |
ENSMUST00000106965.8
ENSMUST00000106968.8 ENSMUST00000106967.8 |
Lrrc51
|
leucine rich repeat containing 51 |
| chr8_+_41276027 | 2.89 |
ENSMUST00000066814.7
|
Adam39
|
a disintegrin and metallopeptidase domain 39 |
| chr4_-_112291169 | 2.81 |
ENSMUST00000058605.3
|
Skint9
|
selection and upkeep of intraepithelial T cells 9 |
| chr13_+_25015653 | 2.67 |
ENSMUST00000038039.3
ENSMUST00000223804.2 |
Tdp2
|
tyrosyl-DNA phosphodiesterase 2 |
| chr1_-_53031814 | 2.31 |
ENSMUST00000190831.7
ENSMUST00000190726.2 |
1700019D03Rik
|
RIKEN cDNA 1700019D03 gene |
| chr7_+_100186399 | 2.19 |
ENSMUST00000120454.3
|
Coa4
|
cytochrome c oxidase assembly factor 4 |
| chr10_-_123032881 | 2.07 |
ENSMUST00000220377.2
|
Usp15
|
ubiquitin specific peptidase 15 |
| chr5_-_134485081 | 2.05 |
ENSMUST00000111244.5
|
Gtf2ird1
|
general transcription factor II I repeat domain-containing 1 |
| chr4_+_107825529 | 2.01 |
ENSMUST00000106713.5
ENSMUST00000238795.2 |
Slc1a7
|
solute carrier family 1 (glutamate transporter), member 7 |
| chr7_-_101570382 | 1.99 |
ENSMUST00000098236.9
|
Lrrc51
|
leucine rich repeat containing 51 |
| chr19_-_10460031 | 1.84 |
ENSMUST00000236733.2
|
Ppp1r32
|
protein phosphatase 1, regulatory subunit 32 |
| chr1_-_155022501 | 1.80 |
ENSMUST00000027744.10
|
Mr1
|
major histocompatibility complex, class I-related |
| chr14_+_55855484 | 1.70 |
ENSMUST00000002395.8
|
Rec8
|
REC8 meiotic recombination protein |
| chrX_+_111150171 | 1.64 |
ENSMUST00000164272.3
ENSMUST00000132037.2 |
4933403O08Rik
|
RIKEN cDNA 4933403O08 gene |
| chr7_-_45343785 | 1.56 |
ENSMUST00000040636.9
|
Sec1
|
secretory blood group 1 |
| chr7_-_33065747 | 1.54 |
ENSMUST00000179248.2
|
Scgb2b20
|
secretoglobin, family 2B, member 20 |
| chr8_+_94941192 | 1.43 |
ENSMUST00000079961.14
ENSMUST00000212824.2 |
Nup93
|
nucleoporin 93 |
| chr17_+_25105617 | 1.39 |
ENSMUST00000117890.8
ENSMUST00000168265.8 ENSMUST00000120943.8 ENSMUST00000068508.13 ENSMUST00000119829.8 |
Spsb3
|
splA/ryanodine receptor domain and SOCS box containing 3 |
| chr7_+_80764564 | 1.35 |
ENSMUST00000119083.2
|
Slc28a1
|
solute carrier family 28 (sodium-coupled nucleoside transporter), member 1 |
| chr10_-_7539009 | 1.24 |
ENSMUST00000163085.8
ENSMUST00000159917.8 |
Pcmt1
|
protein-L-isoaspartate (D-aspartate) O-methyltransferase 1 |
| chr2_+_121978156 | 1.17 |
ENSMUST00000102476.5
|
B2m
|
beta-2 microglobulin |
| chr5_-_124327776 | 1.05 |
ENSMUST00000159677.8
|
Pitpnm2
|
phosphatidylinositol transfer protein, membrane-associated 2 |
| chr5_-_36987917 | 1.00 |
ENSMUST00000031002.10
|
Man2b2
|
mannosidase 2, alpha B2 |
| chr7_+_100186259 | 0.95 |
ENSMUST00000054310.4
|
Coa4
|
cytochrome c oxidase assembly factor 4 |
| chr5_-_125371162 | 0.91 |
ENSMUST00000127148.2
|
Scarb1
|
scavenger receptor class B, member 1 |
| chr11_+_61395964 | 0.86 |
ENSMUST00000102657.10
|
B9d1
|
B9 protein domain 1 |
| chr1_+_134333506 | 0.70 |
ENSMUST00000027726.14
|
Cyb5r1
|
cytochrome b5 reductase 1 |
| chr1_-_74932266 | 0.65 |
ENSMUST00000006721.3
|
Cryba2
|
crystallin, beta A2 |
| chr15_+_36179676 | 0.50 |
ENSMUST00000171205.3
|
Spag1
|
sperm associated antigen 1 |
| chr11_+_72332167 | 0.27 |
ENSMUST00000045633.6
|
Mybbp1a
|
MYB binding protein (P160) 1a |
| chr5_-_140346132 | 0.17 |
ENSMUST00000196130.5
|
Snx8
|
sorting nexin 8 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.8 | 23.4 | GO:0046967 | cytosol to ER transport(GO:0046967) |
| 6.0 | 17.9 | GO:0002581 | negative regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002581) |
| 5.8 | 17.4 | GO:0002343 | peripheral B cell selection(GO:0002343) B cell affinity maturation(GO:0002344) |
| 4.8 | 92.0 | GO:0019886 | antigen processing and presentation of exogenous peptide antigen via MHC class II(GO:0019886) |
| 4.7 | 9.4 | GO:0002477 | antigen processing and presentation of exogenous peptide antigen via MHC class Ib(GO:0002477) antigen processing and presentation of exogenous protein antigen via MHC class Ib, TAP-dependent(GO:0002481) |
| 2.4 | 9.8 | GO:0002485 | antigen processing and presentation of endogenous peptide antigen via MHC class I via ER pathway(GO:0002484) antigen processing and presentation of endogenous peptide antigen via MHC class I via ER pathway, TAP-dependent(GO:0002485) |
| 1.5 | 19.7 | GO:0046602 | regulation of mitotic centrosome separation(GO:0046602) |
| 1.4 | 20.3 | GO:0048102 | autophagic cell death(GO:0048102) |
| 1.3 | 5.4 | GO:2001271 | negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 1.3 | 3.9 | GO:0035638 | patched ligand maturation(GO:0007225) signal maturation(GO:0035638) |
| 1.1 | 19.0 | GO:0036159 | inner dynein arm assembly(GO:0036159) |
| 1.1 | 5.5 | GO:0035616 | histone H2B conserved C-terminal lysine deubiquitination(GO:0035616) |
| 1.0 | 13.9 | GO:0006348 | chromatin silencing at telomere(GO:0006348) |
| 0.9 | 36.9 | GO:0002474 | antigen processing and presentation of peptide antigen via MHC class I(GO:0002474) |
| 0.7 | 40.1 | GO:0019882 | antigen processing and presentation(GO:0019882) |
| 0.7 | 9.9 | GO:0006654 | phosphatidic acid biosynthetic process(GO:0006654) |
| 0.5 | 7.2 | GO:0006707 | cholesterol catabolic process(GO:0006707) sterol catabolic process(GO:0016127) |
| 0.4 | 3.8 | GO:0044821 | meiotic telomere tethering at nuclear periphery(GO:0044821) meiotic attachment of telomere to nuclear envelope(GO:0070197) chromosome attachment to the nuclear envelope(GO:0097240) |
| 0.4 | 2.1 | GO:0014886 | transition between slow and fast fiber(GO:0014886) |
| 0.4 | 3.9 | GO:0006335 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.3 | 1.4 | GO:0015855 | nucleobase transport(GO:0015851) pyrimidine nucleobase transport(GO:0015855) |
| 0.2 | 5.5 | GO:0031163 | iron-sulfur cluster assembly(GO:0016226) metallo-sulfur cluster assembly(GO:0031163) |
| 0.2 | 1.4 | GO:0090521 | glomerular visceral epithelial cell migration(GO:0090521) |
| 0.2 | 2.0 | GO:0089711 | L-glutamate transmembrane transport(GO:0089711) |
| 0.2 | 3.1 | GO:0033617 | mitochondrial respiratory chain complex IV assembly(GO:0033617) |
| 0.2 | 7.3 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.1 | 3.2 | GO:0046855 | inositol phosphate dephosphorylation(GO:0046855) |
| 0.1 | 1.0 | GO:0006013 | mannose metabolic process(GO:0006013) |
| 0.1 | 10.4 | GO:0051225 | spindle assembly(GO:0051225) |
| 0.1 | 1.2 | GO:0046498 | S-adenosylhomocysteine metabolic process(GO:0046498) |
| 0.1 | 1.7 | GO:0007130 | synaptonemal complex assembly(GO:0007130) |
| 0.1 | 0.3 | GO:2000210 | positive regulation of anoikis(GO:2000210) |
| 0.0 | 0.2 | GO:0034498 | early endosome to Golgi transport(GO:0034498) |
| 0.0 | 2.7 | GO:0006302 | double-strand break repair(GO:0006302) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 10.6 | 127.3 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 6.7 | 40.1 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 3.0 | 44.9 | GO:0042611 | MHC protein complex(GO:0042611) |
| 2.9 | 23.4 | GO:0042825 | TAP complex(GO:0042825) |
| 2.5 | 20.3 | GO:0097129 | cyclin D2-CDK4 complex(GO:0097129) |
| 1.2 | 10.4 | GO:0070652 | HAUS complex(GO:0070652) |
| 1.1 | 24.4 | GO:0005652 | nuclear lamina(GO:0005652) |
| 0.7 | 13.9 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.4 | 3.8 | GO:0070187 | telosome(GO:0070187) |
| 0.3 | 5.5 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.3 | 19.7 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.3 | 1.7 | GO:0034991 | nuclear meiotic cohesin complex(GO:0034991) |
| 0.2 | 6.3 | GO:0042588 | zymogen granule(GO:0042588) |
| 0.1 | 0.3 | GO:0031074 | nucleocytoplasmic shuttling complex(GO:0031074) |
| 0.0 | 0.9 | GO:0036038 | MKS complex(GO:0036038) |
| 0.0 | 2.7 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 3.1 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.0 | 0.9 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.0 | 4.3 | GO:0030139 | endocytic vesicle(GO:0030139) |
| 0.0 | 2.3 | GO:0036126 | sperm flagellum(GO:0036126) |
| 0.0 | 1.4 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.0 | 1.2 | GO:0031526 | brush border membrane(GO:0031526) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.8 | 23.4 | GO:0015433 | peptide antigen-transporting ATPase activity(GO:0015433) tapasin binding(GO:0046980) |
| 3.1 | 79.7 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 2.3 | 33.8 | GO:0046977 | TAP binding(GO:0046977) |
| 2.1 | 6.3 | GO:0004771 | sterol esterase activity(GO:0004771) retinyl-palmitate esterase activity(GO:0050253) |
| 1.5 | 40.1 | GO:0004298 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 1.4 | 36.8 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 1.1 | 5.5 | GO:0061649 | ubiquitinated histone binding(GO:0061649) |
| 1.0 | 20.3 | GO:0004861 | cyclin-dependent protein serine/threonine kinase inhibitor activity(GO:0004861) |
| 0.8 | 3.2 | GO:0016316 | phosphatidylinositol-3,4-bisphosphate 4-phosphatase activity(GO:0016316) phosphatidylinositol-4,5-bisphosphate 4-phosphatase activity(GO:0034597) |
| 0.6 | 9.9 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.6 | 13.9 | GO:0042800 | histone methyltransferase activity (H3-K4 specific)(GO:0042800) |
| 0.5 | 1.6 | GO:0031127 | galactoside 2-alpha-L-fucosyltransferase activity(GO:0008107) alpha-(1,2)-fucosyltransferase activity(GO:0031127) |
| 0.5 | 1.4 | GO:0015389 | pyrimidine- and adenine-specific:sodium symporter activity(GO:0015389) |
| 0.4 | 10.5 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.3 | 3.9 | GO:0015197 | peptide transporter activity(GO:0015197) |
| 0.2 | 5.4 | GO:0030296 | protein tyrosine kinase activator activity(GO:0030296) |
| 0.2 | 1.2 | GO:0004719 | protein-L-isoaspartate (D-aspartate) O-methyltransferase activity(GO:0004719) |
| 0.2 | 0.9 | GO:0070506 | high-density lipoprotein particle receptor activity(GO:0070506) |
| 0.1 | 5.1 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.1 | 0.9 | GO:0008158 | hedgehog receptor activity(GO:0008158) |
| 0.1 | 5.5 | GO:0051539 | 4 iron, 4 sulfur cluster binding(GO:0051539) |
| 0.1 | 2.0 | GO:0005313 | L-glutamate transmembrane transporter activity(GO:0005313) |
| 0.1 | 12.8 | GO:0005178 | integrin binding(GO:0005178) |
| 0.1 | 2.3 | GO:0034237 | protein kinase A regulatory subunit binding(GO:0034237) |
| 0.1 | 3.9 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.1 | 7.3 | GO:0015171 | amino acid transmembrane transporter activity(GO:0015171) |
| 0.1 | 1.4 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.0 | 1.0 | GO:0004559 | alpha-mannosidase activity(GO:0004559) |
| 0.0 | 2.7 | GO:0030145 | manganese ion binding(GO:0030145) |
| 0.0 | 6.2 | GO:0008237 | metallopeptidase activity(GO:0008237) |
| 0.0 | 8.5 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.0 | 0.7 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 4.3 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.0 | 0.2 | GO:0004128 | cytochrome-b5 reductase activity, acting on NAD(P)H(GO:0004128) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 5.4 | PID MYC PATHWAY | C-MYC pathway |
| 0.1 | 23.4 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.1 | 9.9 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.1 | 2.7 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.0 | 4.3 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.0 | 1.2 | PID CD8 TCR DOWNSTREAM PATHWAY | Downstream signaling in naïve CD8+ T cells |
| 0.0 | 3.0 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 24.6 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.7 | 38.2 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |
| 0.4 | 11.8 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.3 | 20.3 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.3 | 5.4 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.2 | 9.9 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.2 | 17.4 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.2 | 3.2 | REACTOME SYNTHESIS OF PIPS AT THE EARLY ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the early endosome membrane |
| 0.1 | 4.3 | REACTOME EGFR DOWNREGULATION | Genes involved in EGFR downregulation |
| 0.1 | 2.0 | REACTOME GLUTAMATE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Glutamate Neurotransmitter Release Cycle |
| 0.0 | 0.9 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.0 | 1.4 | REACTOME TRANSPORT OF RIBONUCLEOPROTEINS INTO THE HOST NUCLEUS | Genes involved in Transport of Ribonucleoproteins into the Host Nucleus |
| 0.0 | 1.7 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.0 | 1.4 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |