Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
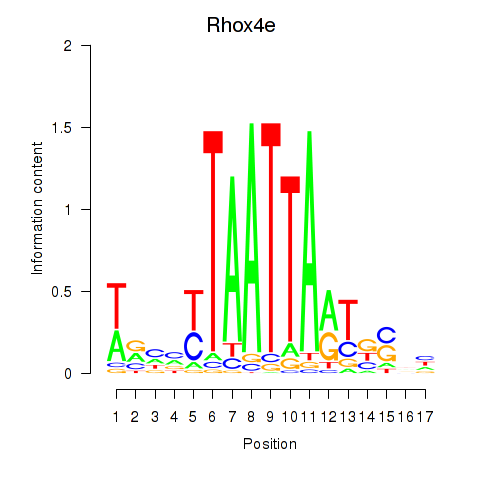
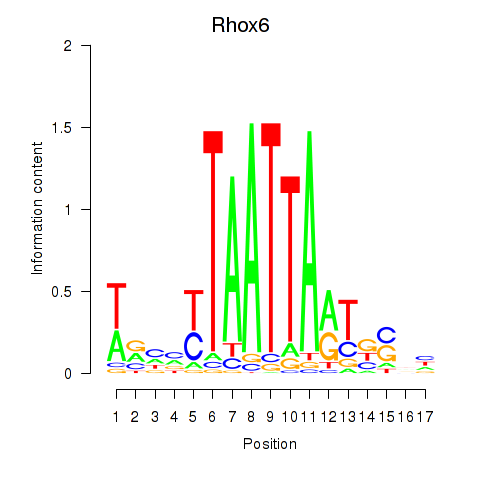
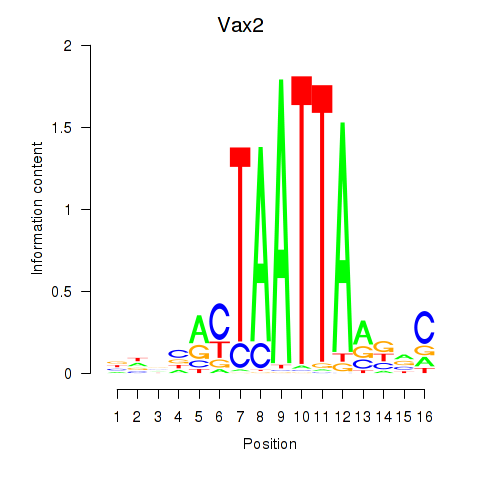
Results for Rhox4e_Rhox6_Vax2
Z-value: 1.21
Transcription factors associated with Rhox4e_Rhox6_Vax2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Rhox4e
|
ENSMUSG00000071770.6 | Rhox4e |
|
Rhox6
|
ENSMUSG00000006200.4 | Rhox6 |
|
Vax2
|
ENSMUSG00000034777.3 | Vax2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Rhox4e | mm39_v1_chrX_+_36739065_36739065 | -0.32 | 5.5e-03 | Click! |
| Rhox6 | mm39_v1_chrX_+_36915787_36915931 | 0.07 | 5.8e-01 | Click! |
| Vax2 | mm39_v1_chr6_+_83688213_83688270 | 0.03 | 7.8e-01 | Click! |
Activity profile of Rhox4e_Rhox6_Vax2 motif
Sorted Z-values of Rhox4e_Rhox6_Vax2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Rhox4e_Rhox6_Vax2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr7_+_51528788 | 16.85 |
ENSMUST00000107591.9
|
Gas2
|
growth arrest specific 2 |
| chr15_-_82678490 | 14.19 |
ENSMUST00000006094.6
|
Cyp2d26
|
cytochrome P450, family 2, subfamily d, polypeptide 26 |
| chr13_+_24023428 | 10.41 |
ENSMUST00000091698.12
ENSMUST00000110422.3 ENSMUST00000166467.9 |
Slc17a3
|
solute carrier family 17 (sodium phosphate), member 3 |
| chr5_+_90708962 | 10.28 |
ENSMUST00000094615.8
ENSMUST00000200765.2 |
Albfm1
|
albumin superfamily member 1 |
| chr19_-_39875192 | 9.68 |
ENSMUST00000168838.3
|
Cyp2c69
|
cytochrome P450, family 2, subfamily c, polypeptide 69 |
| chr19_-_39801188 | 9.55 |
ENSMUST00000162507.2
ENSMUST00000160476.9 ENSMUST00000239028.2 |
Cyp2c40
|
cytochrome P450, family 2, subfamily c, polypeptide 40 |
| chr19_-_39637489 | 9.21 |
ENSMUST00000067328.7
|
Cyp2c67
|
cytochrome P450, family 2, subfamily c, polypeptide 67 |
| chr16_+_22737128 | 8.80 |
ENSMUST00000170805.9
|
Fetub
|
fetuin beta |
| chr16_+_22737227 | 8.80 |
ENSMUST00000231880.2
|
Fetub
|
fetuin beta |
| chr16_+_22737050 | 8.72 |
ENSMUST00000231768.2
|
Fetub
|
fetuin beta |
| chrX_+_149330371 | 8.55 |
ENSMUST00000066337.13
ENSMUST00000112715.2 |
Alas2
|
aminolevulinic acid synthase 2, erythroid |
| chr2_+_163348728 | 7.98 |
ENSMUST00000143911.8
|
Hnf4a
|
hepatic nuclear factor 4, alpha |
| chr13_+_24023386 | 7.35 |
ENSMUST00000039721.14
|
Slc17a3
|
solute carrier family 17 (sodium phosphate), member 3 |
| chr2_+_172994841 | 6.71 |
ENSMUST00000029017.6
|
Pck1
|
phosphoenolpyruvate carboxykinase 1, cytosolic |
| chr10_+_127734384 | 6.57 |
ENSMUST00000047134.8
|
Sdr9c7
|
4short chain dehydrogenase/reductase family 9C, member 7 |
| chr3_-_67422821 | 6.38 |
ENSMUST00000054825.5
|
Rarres1
|
retinoic acid receptor responder (tazarotene induced) 1 |
| chr6_+_121815473 | 6.24 |
ENSMUST00000032228.9
|
Mug1
|
murinoglobulin 1 |
| chr10_-_53252210 | 6.18 |
ENSMUST00000095691.7
|
Cep85l
|
centrosomal protein 85-like |
| chr4_+_100336003 | 6.16 |
ENSMUST00000133493.9
ENSMUST00000092730.5 |
Ube2u
|
ubiquitin-conjugating enzyme E2U (putative) |
| chr4_+_43493344 | 5.99 |
ENSMUST00000030181.12
ENSMUST00000107922.3 |
Ccdc107
|
coiled-coil domain containing 107 |
| chr4_-_14621805 | 5.71 |
ENSMUST00000042221.14
|
Slc26a7
|
solute carrier family 26, member 7 |
| chr6_+_92793440 | 5.40 |
ENSMUST00000057977.4
|
A730049H05Rik
|
RIKEN cDNA A730049H05 gene |
| chr12_-_103660916 | 5.37 |
ENSMUST00000117053.8
|
Serpina1f
|
serine (or cysteine) peptidase inhibitor, clade A, member 1F |
| chr5_-_87240405 | 5.21 |
ENSMUST00000132667.2
ENSMUST00000145617.8 ENSMUST00000094649.11 |
Ugt2b36
|
UDP glucuronosyltransferase 2 family, polypeptide B36 |
| chr6_+_121983720 | 5.15 |
ENSMUST00000081777.8
|
Mug2
|
murinoglobulin 2 |
| chr4_-_14621669 | 5.03 |
ENSMUST00000143105.2
|
Slc26a7
|
solute carrier family 26, member 7 |
| chr11_+_58062467 | 4.94 |
ENSMUST00000020820.2
|
Mrpl22
|
mitochondrial ribosomal protein L22 |
| chrX_+_149372903 | 4.64 |
ENSMUST00000080884.11
|
Pfkfb1
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 |
| chr13_-_17869314 | 4.30 |
ENSMUST00000221598.2
ENSMUST00000068545.6 ENSMUST00000220514.2 |
Sugct
|
succinyl-CoA glutarate-CoA transferase |
| chr4_-_14621497 | 4.21 |
ENSMUST00000149633.2
|
Slc26a7
|
solute carrier family 26, member 7 |
| chr1_-_72323464 | 4.19 |
ENSMUST00000027381.13
|
Pecr
|
peroxisomal trans-2-enoyl-CoA reductase |
| chr4_+_15881256 | 4.18 |
ENSMUST00000029876.2
|
Calb1
|
calbindin 1 |
| chr1_-_72323407 | 4.05 |
ENSMUST00000097698.5
|
Pecr
|
peroxisomal trans-2-enoyl-CoA reductase |
| chr2_-_154916367 | 3.97 |
ENSMUST00000137242.2
ENSMUST00000054607.16 |
Ahcy
|
S-adenosylhomocysteine hydrolase |
| chr18_+_12874390 | 3.96 |
ENSMUST00000121018.8
ENSMUST00000119108.8 ENSMUST00000186263.2 ENSMUST00000191078.7 |
Cabyr
|
calcium-binding tyrosine-(Y)-phosphorylation regulated (fibrousheathin 2) |
| chr7_+_30193047 | 3.95 |
ENSMUST00000058280.13
ENSMUST00000133318.8 ENSMUST00000142575.8 ENSMUST00000131040.2 |
Prodh2
|
proline dehydrogenase (oxidase) 2 |
| chr11_-_100653754 | 3.93 |
ENSMUST00000107360.3
ENSMUST00000055083.4 |
Hcrt
|
hypocretin |
| chr15_-_74508197 | 3.88 |
ENSMUST00000023271.8
|
Mroh4
|
maestro heat-like repeat family member 4 |
| chr9_-_71070506 | 3.87 |
ENSMUST00000074465.9
|
Aqp9
|
aquaporin 9 |
| chr7_-_12829100 | 3.79 |
ENSMUST00000209822.3
ENSMUST00000235753.2 |
Vmn1r85
|
vomeronasal 1 receptor 85 |
| chr11_-_113600838 | 3.65 |
ENSMUST00000018871.8
|
Cpsf4l
|
cleavage and polyadenylation specific factor 4-like |
| chr8_+_46111703 | 3.58 |
ENSMUST00000134675.8
ENSMUST00000139869.8 ENSMUST00000126067.8 |
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr19_-_7943365 | 3.57 |
ENSMUST00000182102.8
ENSMUST00000075619.5 |
Slc22a27
|
solute carrier family 22, member 27 |
| chr18_+_12874368 | 3.44 |
ENSMUST00000235000.2
ENSMUST00000115857.9 |
Cabyr
|
calcium-binding tyrosine-(Y)-phosphorylation regulated (fibrousheathin 2) |
| chrX_+_102400061 | 3.42 |
ENSMUST00000116547.3
|
Chic1
|
cysteine-rich hydrophobic domain 1 |
| chr17_+_79919267 | 3.35 |
ENSMUST00000223924.2
|
Rmdn2
|
regulator of microtubule dynamics 2 |
| chr18_+_60345152 | 3.29 |
ENSMUST00000031549.6
|
Gm4951
|
predicted gene 4951 |
| chr14_-_45626237 | 3.09 |
ENSMUST00000227865.2
ENSMUST00000226856.2 ENSMUST00000226276.2 ENSMUST00000046191.9 |
Gnpnat1
|
glucosamine-phosphate N-acetyltransferase 1 |
| chr18_+_60515755 | 3.03 |
ENSMUST00000237185.2
|
Iigp1
|
interferon inducible GTPase 1 |
| chr6_+_37847721 | 2.99 |
ENSMUST00000031859.14
ENSMUST00000120428.8 |
Trim24
|
tripartite motif-containing 24 |
| chr18_-_10706701 | 2.98 |
ENSMUST00000002549.9
ENSMUST00000117726.9 ENSMUST00000117828.9 |
Abhd3
|
abhydrolase domain containing 3 |
| chr9_-_70842090 | 2.96 |
ENSMUST00000034731.10
|
Lipc
|
lipase, hepatic |
| chr5_-_87402659 | 2.95 |
ENSMUST00000075858.4
|
Ugt2b37
|
UDP glucuronosyltransferase 2 family, polypeptide B37 |
| chr2_-_27365633 | 2.93 |
ENSMUST00000138693.8
ENSMUST00000113941.9 ENSMUST00000077737.13 |
Brd3
|
bromodomain containing 3 |
| chr15_+_89218601 | 2.86 |
ENSMUST00000023282.9
|
Miox
|
myo-inositol oxygenase |
| chr8_+_46111778 | 2.86 |
ENSMUST00000143820.8
|
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr1_+_87983099 | 2.83 |
ENSMUST00000138182.8
ENSMUST00000113142.10 |
Ugt1a10
|
UDP glycosyltransferase 1 family, polypeptide A10 |
| chr17_-_59320257 | 2.83 |
ENSMUST00000174122.2
ENSMUST00000025065.12 |
Nudt12
|
nudix (nucleoside diphosphate linked moiety X)-type motif 12 |
| chr7_+_130375799 | 2.83 |
ENSMUST00000048453.7
ENSMUST00000208593.2 |
Btbd16
|
BTB (POZ) domain containing 16 |
| chr10_+_23770586 | 2.82 |
ENSMUST00000041416.8
|
Vnn1
|
vanin 1 |
| chr13_-_53627110 | 2.79 |
ENSMUST00000021922.10
|
Msx2
|
msh homeobox 2 |
| chr8_+_45960804 | 2.75 |
ENSMUST00000067065.14
ENSMUST00000124544.8 ENSMUST00000138049.9 ENSMUST00000132139.9 |
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr7_+_28869770 | 2.75 |
ENSMUST00000033886.8
ENSMUST00000209019.2 ENSMUST00000208330.2 |
Ggn
|
gametogenetin |
| chr8_+_45960931 | 2.65 |
ENSMUST00000171337.10
ENSMUST00000067107.15 |
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr7_-_126275529 | 2.60 |
ENSMUST00000106372.11
ENSMUST00000155419.3 ENSMUST00000106373.9 |
Sult1a1
|
sulfotransferase family 1A, phenol-preferring, member 1 |
| chr9_+_108216466 | 2.56 |
ENSMUST00000193987.2
|
Gpx1
|
glutathione peroxidase 1 |
| chr7_+_28869629 | 2.55 |
ENSMUST00000098609.4
|
Ggn
|
gametogenetin |
| chr9_-_85631361 | 2.55 |
ENSMUST00000039213.15
|
Ibtk
|
inhibitor of Bruton agammaglobulinemia tyrosine kinase |
| chr1_+_88234454 | 2.53 |
ENSMUST00000040210.14
|
Trpm8
|
transient receptor potential cation channel, subfamily M, member 8 |
| chr7_-_24423715 | 2.51 |
ENSMUST00000081657.6
|
Lypd11
|
Ly6/PLAUR domain containing 11 |
| chrM_+_10167 | 2.50 |
ENSMUST00000082414.1
|
mt-Nd4
|
mitochondrially encoded NADH dehydrogenase 4 |
| chr2_+_87725306 | 2.50 |
ENSMUST00000217436.2
|
Olfr1153
|
olfactory receptor 1153 |
| chr6_-_115569504 | 2.50 |
ENSMUST00000112957.2
|
Mkrn2os
|
makorin, ring finger protein 2, opposite strand |
| chr9_+_108216433 | 2.48 |
ENSMUST00000191997.2
|
Gpx1
|
glutathione peroxidase 1 |
| chr14_-_45626198 | 2.42 |
ENSMUST00000226590.2
|
Gnpnat1
|
glucosamine-phosphate N-acetyltransferase 1 |
| chr8_+_45960855 | 2.39 |
ENSMUST00000141039.8
|
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr17_-_37404764 | 2.38 |
ENSMUST00000087144.5
|
Olfr91
|
olfactory receptor 91 |
| chr8_+_96038143 | 2.35 |
ENSMUST00000161029.9
|
Tepp
|
testis, prostate and placenta expressed |
| chr9_+_21634779 | 2.34 |
ENSMUST00000034713.9
|
Ldlr
|
low density lipoprotein receptor |
| chr9_+_108216233 | 2.31 |
ENSMUST00000082429.8
|
Gpx1
|
glutathione peroxidase 1 |
| chr5_-_90788323 | 2.29 |
ENSMUST00000202784.4
ENSMUST00000031317.10 ENSMUST00000201370.2 |
Rassf6
|
Ras association (RalGDS/AF-6) domain family member 6 |
| chr10_-_128875765 | 2.19 |
ENSMUST00000204763.3
|
Olfr764-ps1
|
olfactory receptor 764, pseudogene 1 |
| chr6_-_41752111 | 2.19 |
ENSMUST00000214976.3
|
Olfr459
|
olfactory receptor 459 |
| chr1_+_87983189 | 2.09 |
ENSMUST00000173325.2
|
Ugt1a10
|
UDP glycosyltransferase 1 family, polypeptide A10 |
| chr19_+_3373285 | 2.09 |
ENSMUST00000025835.6
|
Cpt1a
|
carnitine palmitoyltransferase 1a, liver |
| chr7_+_114344920 | 2.03 |
ENSMUST00000136645.8
ENSMUST00000169913.8 |
Insc
|
INSC spindle orientation adaptor protein |
| chr2_-_111100733 | 2.01 |
ENSMUST00000099619.6
|
Olfr1277
|
olfactory receptor 1277 |
| chr15_+_59186876 | 1.98 |
ENSMUST00000022977.14
ENSMUST00000100640.5 |
Sqle
|
squalene epoxidase |
| chr8_+_96038224 | 1.95 |
ENSMUST00000098480.9
ENSMUST00000212056.2 |
Tepp
|
testis, prostate and placenta expressed |
| chr10_-_81243475 | 1.84 |
ENSMUST00000140916.8
|
Nfic
|
nuclear factor I/C |
| chr7_+_141765529 | 1.80 |
ENSMUST00000097943.2
|
Gm7579
|
predicted gene 7579 |
| chr16_-_45664664 | 1.78 |
ENSMUST00000036355.13
|
Phldb2
|
pleckstrin homology like domain, family B, member 2 |
| chr2_+_69727563 | 1.75 |
ENSMUST00000055758.16
ENSMUST00000112251.9 |
Ubr3
|
ubiquitin protein ligase E3 component n-recognin 3 |
| chrM_+_9870 | 1.75 |
ENSMUST00000084013.1
|
mt-Nd4l
|
mitochondrially encoded NADH dehydrogenase 4L |
| chr9_-_55419442 | 1.69 |
ENSMUST00000034866.9
|
Etfa
|
electron transferring flavoprotein, alpha polypeptide |
| chr2_-_168608949 | 1.67 |
ENSMUST00000075044.10
|
Sall4
|
spalt like transcription factor 4 |
| chr12_+_103524690 | 1.67 |
ENSMUST00000187155.7
|
Ppp4r4
|
protein phosphatase 4, regulatory subunit 4 |
| chr5_-_90788460 | 1.58 |
ENSMUST00000202704.4
|
Rassf6
|
Ras association (RalGDS/AF-6) domain family member 6 |
| chr16_-_45664591 | 1.57 |
ENSMUST00000076333.12
|
Phldb2
|
pleckstrin homology like domain, family B, member 2 |
| chr9_-_50472620 | 1.54 |
ENSMUST00000217236.2
|
Tex12
|
testis expressed 12 |
| chr2_+_69727599 | 1.53 |
ENSMUST00000131553.2
|
Ubr3
|
ubiquitin protein ligase E3 component n-recognin 3 |
| chr6_-_73446560 | 1.52 |
ENSMUST00000070163.6
|
4931417E11Rik
|
RIKEN cDNA 4931417E11 gene |
| chr5_-_3697806 | 1.51 |
ENSMUST00000119783.2
ENSMUST00000007559.15 |
Gatad1
|
GATA zinc finger domain containing 1 |
| chr7_+_30157704 | 1.50 |
ENSMUST00000126297.9
|
Nphs1
|
nephrosis 1, nephrin |
| chr15_-_66985760 | 1.49 |
ENSMUST00000092640.6
|
St3gal1
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 1 |
| chr9_-_50472605 | 1.44 |
ENSMUST00000034568.7
|
Tex12
|
testis expressed 12 |
| chr3_+_121746862 | 1.43 |
ENSMUST00000037958.14
ENSMUST00000196904.5 |
Arhgap29
|
Rho GTPase activating protein 29 |
| chr8_+_20478993 | 1.43 |
ENSMUST00000211848.2
|
Gm31371
|
predicted gene, 31371 |
| chr9_+_38755745 | 1.41 |
ENSMUST00000217350.2
|
Olfr924
|
olfactory receptor 924 |
| chr11_+_69729340 | 1.40 |
ENSMUST00000133967.8
ENSMUST00000094065.5 |
Tmem256
|
transmembrane protein 256 |
| chr12_+_116239006 | 1.38 |
ENSMUST00000090195.5
|
Gm11027
|
predicted gene 11027 |
| chr11_+_58468556 | 1.30 |
ENSMUST00000203418.4
|
Olfr325
|
olfactory receptor 325 |
| chr12_-_85871281 | 1.28 |
ENSMUST00000021676.12
ENSMUST00000142331.2 |
Erg28
|
ergosterol biosynthesis 28 |
| chr13_-_81859056 | 1.27 |
ENSMUST00000161920.2
ENSMUST00000048993.12 |
Polr3g
|
polymerase (RNA) III (DNA directed) polypeptide G |
| chr8_-_62355690 | 1.26 |
ENSMUST00000121785.9
ENSMUST00000034057.14 |
Palld
|
palladin, cytoskeletal associated protein |
| chr6_-_68968278 | 1.25 |
ENSMUST00000197966.2
|
Igkv4-81
|
immunoglobulin kappa variable 4-81 |
| chrX_-_159689993 | 1.24 |
ENSMUST00000069417.6
|
Gja6
|
gap junction protein, alpha 6 |
| chr15_+_102927366 | 1.23 |
ENSMUST00000165375.3
|
Hoxc4
|
homeobox C4 |
| chrX_-_137985960 | 1.20 |
ENSMUST00000033626.15
ENSMUST00000060824.4 |
Serpina7
|
serine (or cysteine) peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 7 |
| chr19_+_25384024 | 1.17 |
ENSMUST00000146647.3
|
Kank1
|
KN motif and ankyrin repeat domains 1 |
| chr5_-_66211842 | 1.13 |
ENSMUST00000200852.4
|
Rbm47
|
RNA binding motif protein 47 |
| chrX_-_8391193 | 1.12 |
ENSMUST00000115573.2
|
Gm14459
|
predicted gene 14459 |
| chr19_-_58781709 | 1.11 |
ENSMUST00000166692.2
|
1700019N19Rik
|
RIKEN cDNA 1700019N19 gene |
| chr9_+_80072361 | 1.06 |
ENSMUST00000184480.8
|
Myo6
|
myosin VI |
| chr2_-_89159994 | 1.05 |
ENSMUST00000213860.2
ENSMUST00000215679.2 |
Olfr1232
|
olfactory receptor 1232 |
| chr3_+_18002574 | 1.04 |
ENSMUST00000029080.5
|
Cypt12
|
cysteine-rich perinuclear theca 12 |
| chr2_-_164427367 | 1.04 |
ENSMUST00000109342.2
|
Wfdc6a
|
WAP four-disulfide core domain 6A |
| chr1_+_54289833 | 1.03 |
ENSMUST00000027128.11
|
Ccdc150
|
coiled-coil domain containing 150 |
| chr3_-_54962899 | 1.03 |
ENSMUST00000199144.5
|
Ccna1
|
cyclin A1 |
| chr12_-_79237722 | 1.03 |
ENSMUST00000085254.7
|
Rdh11
|
retinol dehydrogenase 11 |
| chr2_+_104961228 | 1.03 |
ENSMUST00000111098.8
ENSMUST00000111099.2 |
Wt1
|
Wilms tumor 1 homolog |
| chrX_+_110154017 | 1.02 |
ENSMUST00000210720.3
|
Cylc1
|
cylicin, basic protein of sperm head cytoskeleton 1 |
| chr7_+_45164042 | 1.01 |
ENSMUST00000210868.2
|
Tulp2
|
tubby-like protein 2 |
| chr9_-_19275301 | 1.00 |
ENSMUST00000214810.2
|
Olfr846
|
olfactory receptor 846 |
| chr8_-_41494890 | 0.99 |
ENSMUST00000051379.14
|
Mtus1
|
mitochondrial tumor suppressor 1 |
| chr9_+_20148415 | 0.96 |
ENSMUST00000086474.6
|
Olfr872
|
olfactory receptor 872 |
| chr12_+_78240999 | 0.96 |
ENSMUST00000211288.3
|
Gm6657
|
predicted gene 6657 |
| chr15_-_34356567 | 0.95 |
ENSMUST00000179647.2
|
9430069I07Rik
|
RIKEN cDNA 9430069I07 gene |
| chrM_+_7006 | 0.93 |
ENSMUST00000082405.1
|
mt-Co2
|
mitochondrially encoded cytochrome c oxidase II |
| chr10_-_126956991 | 0.92 |
ENSMUST00000080975.6
ENSMUST00000164259.9 |
Os9
|
amplified in osteosarcoma |
| chr11_-_99241924 | 0.92 |
ENSMUST00000017732.3
|
Krt27
|
keratin 27 |
| chr4_-_156340713 | 0.91 |
ENSMUST00000219393.2
|
Samd11
|
sterile alpha motif domain containing 11 |
| chr8_-_22550035 | 0.91 |
ENSMUST00000110738.3
|
Atp7b
|
ATPase, Cu++ transporting, beta polypeptide |
| chr11_-_99494134 | 0.91 |
ENSMUST00000072306.4
|
Gm11938
|
predicted gene 11938 |
| chr14_+_65504067 | 0.90 |
ENSMUST00000224629.2
|
Fbxo16
|
F-box protein 16 |
| chr1_+_173964312 | 0.90 |
ENSMUST00000053941.4
|
Olfr424
|
olfactory receptor 424 |
| chr19_-_44095840 | 0.89 |
ENSMUST00000119591.2
ENSMUST00000026217.11 |
Chuk
|
conserved helix-loop-helix ubiquitous kinase |
| chr11_-_99265721 | 0.89 |
ENSMUST00000006963.3
|
Krt28
|
keratin 28 |
| chr3_-_64049349 | 0.89 |
ENSMUST00000177151.9
|
Vmn2r2
|
vomeronasal 2, receptor 2 |
| chr1_+_85949827 | 0.89 |
ENSMUST00000131151.3
|
Spata3
|
spermatogenesis associated 3 |
| chr4_-_129121676 | 0.89 |
ENSMUST00000106051.8
|
C77080
|
expressed sequence C77080 |
| chr2_-_168609110 | 0.87 |
ENSMUST00000029061.12
ENSMUST00000103074.2 |
Sall4
|
spalt like transcription factor 4 |
| chr13_+_76115824 | 0.86 |
ENSMUST00000022081.3
|
Spata9
|
spermatogenesis associated 9 |
| chr4_-_133066594 | 0.86 |
ENSMUST00000043305.14
|
Wdtc1
|
WD and tetratricopeptide repeats 1 |
| chr8_+_46463633 | 0.85 |
ENSMUST00000110381.9
|
Lrp2bp
|
Lrp2 binding protein |
| chr8_-_122268693 | 0.85 |
ENSMUST00000034265.11
|
1700018B08Rik
|
RIKEN cDNA 1700018B08 gene |
| chr16_-_89368059 | 0.82 |
ENSMUST00000171542.2
|
Krtap11-1
|
keratin associated protein 11-1 |
| chr13_+_23191826 | 0.82 |
ENSMUST00000228758.2
ENSMUST00000228031.2 ENSMUST00000227573.2 |
Vmn1r213
|
vomeronasal 1 receptor 213 |
| chr19_+_60800012 | 0.81 |
ENSMUST00000128357.8
ENSMUST00000119633.8 ENSMUST00000025957.9 |
Dennd10
|
DENN domain containing 10 |
| chr18_+_38551960 | 0.81 |
ENSMUST00000236085.2
|
Ndfip1
|
Nedd4 family interacting protein 1 |
| chr18_+_38552011 | 0.81 |
ENSMUST00000025293.5
|
Ndfip1
|
Nedd4 family interacting protein 1 |
| chr17_-_57338468 | 0.80 |
ENSMUST00000007814.10
ENSMUST00000233480.2 |
Khsrp
|
KH-type splicing regulatory protein |
| chr13_+_28326465 | 0.79 |
ENSMUST00000017208.5
|
Prl5a1
|
prolactin family 5, subfamily a, member 1 |
| chr11_-_99907030 | 0.79 |
ENSMUST00000018399.3
|
Krt33a
|
keratin 33A |
| chr2_-_69542805 | 0.78 |
ENSMUST00000102706.4
ENSMUST00000073152.13 |
Fastkd1
|
FAST kinase domains 1 |
| chr3_-_75072319 | 0.77 |
ENSMUST00000124618.2
|
Zbbx
|
zinc finger, B-box domain containing |
| chr1_+_87111033 | 0.77 |
ENSMUST00000044533.9
|
Prss56
|
protease, serine 56 |
| chrX_+_9751861 | 0.77 |
ENSMUST00000067529.9
ENSMUST00000086165.4 |
Sytl5
|
synaptotagmin-like 5 |
| chr16_-_19341016 | 0.77 |
ENSMUST00000214315.2
|
Olfr167
|
olfactory receptor 167 |
| chr1_+_163757334 | 0.75 |
ENSMUST00000027876.11
ENSMUST00000170359.8 |
Scyl3
|
SCY1-like 3 (S. cerevisiae) |
| chr8_+_22329942 | 0.75 |
ENSMUST00000006745.4
|
Defb2
|
defensin beta 2 |
| chr7_-_103778992 | 0.74 |
ENSMUST00000053743.6
|
Ubqln5
|
ubiquilin 5 |
| chr12_+_55349422 | 0.74 |
ENSMUST00000021411.15
|
Prorp
|
protein only RNase P catalytic subunit |
| chr12_+_55350023 | 0.74 |
ENSMUST00000184766.8
ENSMUST00000183475.8 ENSMUST00000183654.2 |
Prorp
|
protein only RNase P catalytic subunit |
| chr12_+_80690985 | 0.73 |
ENSMUST00000219405.2
ENSMUST00000085245.7 |
Slc39a9
|
solute carrier family 39 (zinc transporter), member 9 |
| chr3_-_92441809 | 0.72 |
ENSMUST00000193521.2
|
2310046K23Rik
|
RIKEN cDNA 2310046K23 gene |
| chr4_-_131802606 | 0.72 |
ENSMUST00000146021.8
|
Epb41
|
erythrocyte membrane protein band 4.1 |
| chr11_+_90078485 | 0.72 |
ENSMUST00000124877.3
ENSMUST00000153734.2 ENSMUST00000134929.8 |
Gm45716
|
predicted gene 45716 |
| chr5_-_137015683 | 0.72 |
ENSMUST00000034953.14
ENSMUST00000085941.12 |
Znhit1
|
zinc finger, HIT domain containing 1 |
| chrX_-_73211444 | 0.71 |
ENSMUST00000114328.8
ENSMUST00000078060.4 |
Tex28
|
testis expressed 28 |
| chr2_+_24181208 | 0.71 |
ENSMUST00000058056.2
|
Il1f10
|
interleukin 1 family, member 10 |
| chr7_+_102287639 | 0.71 |
ENSMUST00000215995.2
ENSMUST00000214841.3 |
Olfr554
|
olfactory receptor 554 |
| chr10_-_129965752 | 0.70 |
ENSMUST00000215217.2
ENSMUST00000214192.2 |
Olfr824
|
olfactory receptor 824 |
| chr2_+_65676176 | 0.70 |
ENSMUST00000053910.10
|
Csrnp3
|
cysteine-serine-rich nuclear protein 3 |
| chr14_-_50558325 | 0.70 |
ENSMUST00000050928.3
|
Olfr734
|
olfactory receptor 734 |
| chr2_-_164480710 | 0.69 |
ENSMUST00000109336.2
|
Wfdc16
|
WAP four-disulfide core domain 16 |
| chr9_-_97252011 | 0.69 |
ENSMUST00000035026.5
|
Trim42
|
tripartite motif-containing 42 |
| chrX_-_49517375 | 0.69 |
ENSMUST00000215270.2
|
Olfr1324
|
olfactory receptor 1324 |
| chrX_-_137985979 | 0.68 |
ENSMUST00000152457.2
|
Serpina7
|
serine (or cysteine) peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 7 |
| chr8_+_10299288 | 0.68 |
ENSMUST00000214643.2
|
Myo16
|
myosin XVI |
| chr14_+_54669054 | 0.67 |
ENSMUST00000089688.6
ENSMUST00000225641.2 |
Mmp14
|
matrix metallopeptidase 14 (membrane-inserted) |
| chr13_+_22454692 | 0.67 |
ENSMUST00000228711.2
|
Vmn1r195
|
vomeronasal 1 receptor 195 |
| chr11_+_73489420 | 0.67 |
ENSMUST00000214228.2
|
Olfr384
|
olfactory receptor 384 |
| chr17_-_45970238 | 0.63 |
ENSMUST00000120717.8
|
Capn11
|
calpain 11 |
| chr13_-_23041731 | 0.63 |
ENSMUST00000228645.2
|
Vmn1r211
|
vomeronasal 1 receptor 211 |
| chr4_-_131802561 | 0.62 |
ENSMUST00000105970.8
ENSMUST00000105975.8 |
Epb41
|
erythrocyte membrane protein band 4.1 |
| chrX_-_134528431 | 0.62 |
ENSMUST00000113159.8
|
Pramex1
|
PRAME like, X-linked 1 |
| chr18_+_34675366 | 0.61 |
ENSMUST00000012426.3
|
Wnt8a
|
wingless-type MMTV integration site family, member 8A |
| chr11_-_99447672 | 0.60 |
ENSMUST00000092699.3
|
Krtap3-2
|
keratin associated protein 3-2 |
| chr4_-_150998857 | 0.59 |
ENSMUST00000105675.8
|
Park7
|
Parkinson disease (autosomal recessive, early onset) 7 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.7 | 8.2 | GO:0033306 | phytol metabolic process(GO:0033306) fatty alcohol metabolic process(GO:1903173) |
| 2.7 | 8.0 | GO:0042977 | regulation of activation of JAK2 kinase activity(GO:0010534) activation of JAK2 kinase activity(GO:0042977) negative regulation of activation of JAK2 kinase activity(GO:1902569) |
| 1.8 | 7.4 | GO:0010269 | response to symbiont(GO:0009608) response to symbiotic bacterium(GO:0009609) response to selenium ion(GO:0010269) |
| 1.7 | 6.7 | GO:0006114 | glycerol biosynthetic process(GO:0006114) positive regulation of transcription from RNA polymerase II promoter in response to acidic pH(GO:0061402) |
| 1.7 | 14.9 | GO:0019532 | oxalate transport(GO:0019532) |
| 1.4 | 21.3 | GO:0015747 | urate transport(GO:0015747) |
| 1.3 | 3.9 | GO:0010133 | proline catabolic process to glutamate(GO:0010133) |
| 1.3 | 5.2 | GO:0035502 | metanephric part of ureteric bud development(GO:0035502) |
| 1.1 | 5.5 | GO:0006041 | glucosamine metabolic process(GO:0006041) |
| 1.0 | 4.0 | GO:0002439 | chronic inflammatory response to antigenic stimulus(GO:0002439) |
| 1.0 | 3.9 | GO:0015851 | nucleobase transport(GO:0015851) pyrimidine nucleobase transport(GO:0015855) |
| 0.9 | 2.8 | GO:2001055 | positive regulation of mesenchymal cell apoptotic process(GO:2001055) |
| 0.9 | 28.4 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 0.8 | 3.3 | GO:0071596 | ubiquitin-dependent protein catabolic process via the N-end rule pathway(GO:0071596) |
| 0.6 | 1.9 | GO:0033189 | response to vitamin A(GO:0033189) |
| 0.6 | 3.0 | GO:0034371 | chylomicron remodeling(GO:0034371) |
| 0.6 | 2.3 | GO:0010899 | regulation of phosphatidylcholine catabolic process(GO:0010899) positive regulation of lipoprotein particle clearance(GO:0010986) receptor-mediated endocytosis of low-density lipoprotein particle involved in cholesterol transport(GO:0090118) positive regulation of lysosomal protein catabolic process(GO:1905167) |
| 0.5 | 3.0 | GO:0000711 | meiotic DNA repair synthesis(GO:0000711) |
| 0.5 | 3.9 | GO:0051970 | negative regulation of transmission of nerve impulse(GO:0051970) |
| 0.5 | 8.5 | GO:0042541 | hemoglobin biosynthetic process(GO:0042541) |
| 0.5 | 2.8 | GO:0006742 | NADP catabolic process(GO:0006742) |
| 0.5 | 26.3 | GO:0007339 | binding of sperm to zona pellucida(GO:0007339) |
| 0.5 | 4.6 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.4 | 1.3 | GO:0008204 | ergosterol biosynthetic process(GO:0006696) ergosterol metabolic process(GO:0008204) |
| 0.4 | 1.7 | GO:0080163 | regulation of protein serine/threonine phosphatase activity(GO:0080163) |
| 0.4 | 4.9 | GO:0052697 | flavonoid glucuronidation(GO:0052696) xenobiotic glucuronidation(GO:0052697) |
| 0.4 | 1.2 | GO:2000393 | negative regulation of lamellipodium morphogenesis(GO:2000393) |
| 0.4 | 3.0 | GO:0070562 | regulation of vitamin D receptor signaling pathway(GO:0070562) |
| 0.3 | 3.3 | GO:1903690 | negative regulation of wound healing, spreading of epidermal cells(GO:1903690) |
| 0.3 | 3.7 | GO:0098789 | pre-mRNA cleavage required for polyadenylation(GO:0098789) |
| 0.3 | 1.2 | GO:0016062 | adaptation of rhodopsin mediated signaling(GO:0016062) light adaption(GO:0036367) |
| 0.3 | 0.9 | GO:0015680 | intracellular copper ion transport(GO:0015680) |
| 0.3 | 11.7 | GO:0019369 | arachidonic acid metabolic process(GO:0019369) |
| 0.3 | 1.6 | GO:0048294 | negative regulation of isotype switching to IgE isotypes(GO:0048294) |
| 0.3 | 2.5 | GO:0050955 | thermoception(GO:0050955) |
| 0.2 | 2.9 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.2 | 1.0 | GO:1901317 | regulation of sperm motility(GO:1901317) |
| 0.2 | 13.8 | GO:0003301 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.2 | 7.4 | GO:0048240 | sperm capacitation(GO:0048240) |
| 0.2 | 1.2 | GO:0009786 | regulation of asymmetric cell division(GO:0009786) |
| 0.2 | 2.7 | GO:1902176 | negative regulation of oxidative stress-induced intrinsic apoptotic signaling pathway(GO:1902176) |
| 0.1 | 3.1 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.1 | 11.5 | GO:0007566 | embryo implantation(GO:0007566) |
| 0.1 | 0.4 | GO:0071211 | protein targeting to vacuole involved in autophagy(GO:0071211) lysosomal membrane organization(GO:0097212) positive regulation of protein folding(GO:1903334) |
| 0.1 | 0.9 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 0.1 | 2.1 | GO:0032000 | positive regulation of fatty acid beta-oxidation(GO:0032000) |
| 0.1 | 0.4 | GO:0002149 | hypochlorous acid metabolic process(GO:0002148) hypochlorous acid biosynthetic process(GO:0002149) |
| 0.1 | 0.5 | GO:0003275 | apoptotic process involved in outflow tract morphogenesis(GO:0003275) regulation of apoptotic process involved in outflow tract morphogenesis(GO:1902256) |
| 0.1 | 6.3 | GO:0035458 | cellular response to interferon-beta(GO:0035458) |
| 0.1 | 2.6 | GO:0051923 | sulfation(GO:0051923) |
| 0.1 | 2.8 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 0.1 | 0.5 | GO:0035036 | sperm-egg recognition(GO:0035036) |
| 0.1 | 0.5 | GO:0006566 | threonine metabolic process(GO:0006566) |
| 0.1 | 0.5 | GO:0046947 | hydroxylysine metabolic process(GO:0046946) hydroxylysine biosynthetic process(GO:0046947) |
| 0.1 | 0.9 | GO:0072641 | type I interferon secretion(GO:0072641) interferon-alpha secretion(GO:0072642) regulation of interferon-alpha secretion(GO:1902739) positive regulation of interferon-alpha secretion(GO:1902741) |
| 0.1 | 1.1 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.1 | 0.7 | GO:0070317 | negative regulation of G0 to G1 transition(GO:0070317) |
| 0.1 | 6.2 | GO:0016574 | histone ubiquitination(GO:0016574) |
| 0.1 | 0.6 | GO:0065001 | specification of axis polarity(GO:0065001) |
| 0.1 | 0.5 | GO:0043504 | mitochondrial DNA repair(GO:0043504) |
| 0.1 | 1.3 | GO:1904776 | regulation of protein localization to cell cortex(GO:1904776) positive regulation of protein localization to cell cortex(GO:1904778) |
| 0.1 | 15.4 | GO:0007050 | cell cycle arrest(GO:0007050) |
| 0.1 | 0.5 | GO:0035878 | nail development(GO:0035878) |
| 0.1 | 0.8 | GO:2000628 | regulation of miRNA metabolic process(GO:2000628) |
| 0.1 | 0.4 | GO:0035553 | oxidative single-stranded RNA demethylation(GO:0035553) |
| 0.1 | 3.0 | GO:0046470 | phosphatidylcholine metabolic process(GO:0046470) |
| 0.1 | 2.4 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.1 | 1.0 | GO:1903546 | protein localization to photoreceptor outer segment(GO:1903546) |
| 0.1 | 1.1 | GO:0071257 | cellular response to electrical stimulus(GO:0071257) |
| 0.1 | 2.7 | GO:0042773 | ATP synthesis coupled electron transport(GO:0042773) |
| 0.1 | 1.3 | GO:0071803 | positive regulation of podosome assembly(GO:0071803) |
| 0.1 | 0.4 | GO:1904749 | regulation of protein localization to nucleolus(GO:1904749) |
| 0.0 | 0.9 | GO:0045717 | negative regulation of fatty acid biosynthetic process(GO:0045717) |
| 0.0 | 0.7 | GO:0035988 | chondrocyte proliferation(GO:0035988) |
| 0.0 | 0.5 | GO:2000582 | regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000580) positive regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000582) |
| 0.0 | 0.3 | GO:0051013 | microtubule severing(GO:0051013) |
| 0.0 | 0.4 | GO:1903608 | protein localization to cytoplasmic stress granule(GO:1903608) |
| 0.0 | 0.2 | GO:0070173 | regulation of enamel mineralization(GO:0070173) |
| 0.0 | 10.7 | GO:0010466 | negative regulation of peptidase activity(GO:0010466) |
| 0.0 | 2.0 | GO:0060487 | lung epithelial cell differentiation(GO:0060487) |
| 0.0 | 1.5 | GO:0044062 | regulation of excretion(GO:0044062) |
| 0.0 | 2.0 | GO:0016126 | sterol biosynthetic process(GO:0016126) |
| 0.0 | 0.5 | GO:0048672 | positive regulation of collateral sprouting(GO:0048672) |
| 0.0 | 0.3 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.0 | 1.3 | GO:0006383 | transcription from RNA polymerase III promoter(GO:0006383) |
| 0.0 | 0.2 | GO:1902035 | positive regulation of hematopoietic stem cell proliferation(GO:1902035) |
| 0.0 | 1.0 | GO:0010758 | regulation of macrophage chemotaxis(GO:0010758) |
| 0.0 | 1.5 | GO:0018279 | protein N-linked glycosylation via asparagine(GO:0018279) |
| 0.0 | 1.8 | GO:0042475 | odontogenesis of dentin-containing tooth(GO:0042475) |
| 0.0 | 0.9 | GO:0031069 | hair follicle morphogenesis(GO:0031069) |
| 0.0 | 0.2 | GO:0009249 | tRNA 5'-leader removal(GO:0001682) protein lipoylation(GO:0009249) |
| 0.0 | 0.4 | GO:0003215 | cardiac right ventricle morphogenesis(GO:0003215) |
| 0.0 | 0.8 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.0 | 2.6 | GO:0051209 | release of sequestered calcium ion into cytosol(GO:0051209) negative regulation of sequestering of calcium ion(GO:0051283) |
| 0.0 | 0.6 | GO:0070306 | lens fiber cell differentiation(GO:0070306) |
| 0.0 | 1.7 | GO:0031497 | chromatin assembly(GO:0031497) |
| 0.0 | 0.4 | GO:0031365 | N-terminal protein amino acid modification(GO:0031365) |
| 0.0 | 0.2 | GO:0051457 | maintenance of protein location in nucleus(GO:0051457) |
| 0.0 | 0.4 | GO:2001240 | negative regulation of signal transduction in absence of ligand(GO:1901099) negative regulation of extrinsic apoptotic signaling pathway in absence of ligand(GO:2001240) |
| 0.0 | 0.4 | GO:1901998 | toxin transport(GO:1901998) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 4.6 | GO:0043540 | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase complex(GO:0043540) |
| 0.9 | 7.4 | GO:0097413 | Lewy body(GO:0097413) |
| 0.8 | 2.3 | GO:1990666 | PCSK9-LDLR complex(GO:1990666) |
| 0.8 | 6.2 | GO:0033503 | HULC complex(GO:0033503) |
| 0.4 | 1.7 | GO:0017133 | mitochondrial electron transfer flavoprotein complex(GO:0017133) electron transfer flavoprotein complex(GO:0045251) |
| 0.4 | 3.0 | GO:0005726 | perichromatin fibrils(GO:0005726) |
| 0.3 | 1.4 | GO:0097123 | cyclin A1-CDK2 complex(GO:0097123) |
| 0.3 | 7.4 | GO:0035686 | sperm fibrous sheath(GO:0035686) |
| 0.3 | 1.0 | GO:0043159 | acrosomal matrix(GO:0043159) |
| 0.2 | 3.0 | GO:0000801 | central element(GO:0000801) |
| 0.2 | 3.0 | GO:0020005 | symbiont-containing vacuole membrane(GO:0020005) |
| 0.2 | 0.4 | GO:0071920 | cleavage body(GO:0071920) |
| 0.2 | 3.3 | GO:0045180 | basal cortex(GO:0045180) |
| 0.2 | 3.7 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.1 | 0.9 | GO:0000836 | ER ubiquitin ligase complex(GO:0000835) Hrd1p ubiquitin ligase complex(GO:0000836) |
| 0.1 | 0.4 | GO:0033165 | interphotoreceptor matrix(GO:0033165) |
| 0.1 | 8.2 | GO:0031903 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.1 | 1.5 | GO:0036056 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.1 | 4.9 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.1 | 3.0 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.1 | 0.4 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.1 | 6.2 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.1 | 0.7 | GO:0044354 | pinosome(GO:0044352) macropinosome(GO:0044354) |
| 0.1 | 2.1 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.1 | 5.9 | GO:0070469 | respiratory chain(GO:0070469) |
| 0.1 | 0.9 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.1 | 0.4 | GO:0099524 | region of cytosol(GO:0099522) postsynaptic cytosol(GO:0099524) |
| 0.1 | 1.3 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.1 | 0.7 | GO:0000812 | Swr1 complex(GO:0000812) |
| 0.0 | 1.6 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.0 | 0.2 | GO:0030914 | STAGA complex(GO:0030914) |
| 0.0 | 14.2 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 20.8 | GO:0005789 | endoplasmic reticulum membrane(GO:0005789) |
| 0.0 | 0.4 | GO:0034709 | methylosome(GO:0034709) |
| 0.0 | 7.8 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.0 | 1.3 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.0 | 0.2 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.0 | 11.0 | GO:0014069 | postsynaptic density(GO:0014069) |
| 0.0 | 0.4 | GO:0005766 | primary lysosome(GO:0005766) azurophil granule(GO:0042582) |
| 0.0 | 4.2 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 1.1 | GO:0045334 | clathrin-coated endocytic vesicle(GO:0045334) |
| 0.0 | 1.7 | GO:0008287 | protein serine/threonine phosphatase complex(GO:0008287) phosphatase complex(GO:1903293) |
| 0.0 | 0.5 | GO:0002080 | acrosomal membrane(GO:0002080) |
| 0.0 | 13.4 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 6.9 | GO:0005743 | mitochondrial inner membrane(GO:0005743) |
| 0.0 | 1.0 | GO:0001917 | photoreceptor inner segment(GO:0001917) |
| 0.0 | 0.5 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 6.0 | GO:0048471 | perinuclear region of cytoplasm(GO:0048471) |
| 0.0 | 6.1 | GO:0000151 | ubiquitin ligase complex(GO:0000151) |
| 0.0 | 0.8 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.0 | 1.8 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.0 | 1.1 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.0 | 18.5 | GO:0005783 | endoplasmic reticulum(GO:0005783) |
| 0.0 | 1.2 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.0 | 0.1 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.0 | 36.8 | GO:0070062 | extracellular exosome(GO:0070062) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.8 | 8.5 | GO:0003870 | 5-aminolevulinate synthase activity(GO:0003870) N-succinyltransferase activity(GO:0016749) |
| 2.7 | 8.0 | GO:0070540 | stearic acid binding(GO:0070540) |
| 2.2 | 6.7 | GO:0004611 | phosphoenolpyruvate carboxykinase activity(GO:0004611) phosphoenolpyruvate carboxykinase (GTP) activity(GO:0004613) |
| 2.1 | 8.2 | GO:0019166 | trans-2-enoyl-CoA reductase (NADPH) activity(GO:0019166) |
| 1.8 | 17.8 | GO:0015562 | efflux transmembrane transporter activity(GO:0015562) |
| 1.2 | 26.3 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 1.0 | 5.9 | GO:0008970 | phosphatidylcholine 1-acylhydrolase activity(GO:0008970) |
| 0.9 | 14.9 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.8 | 3.9 | GO:0015205 | purine nucleobase transmembrane transporter activity(GO:0005345) pyrimidine nucleobase transmembrane transporter activity(GO:0005350) nucleobase transmembrane transporter activity(GO:0015205) glycerol channel activity(GO:0015254) |
| 0.8 | 28.4 | GO:0008392 | arachidonic acid epoxygenase activity(GO:0008392) |
| 0.7 | 14.2 | GO:0070330 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen(GO:0016712) aromatase activity(GO:0070330) |
| 0.7 | 2.1 | GO:1990698 | palmitoleoyltransferase activity(GO:1990698) |
| 0.6 | 4.0 | GO:0004013 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.5 | 4.3 | GO:0008410 | CoA-transferase activity(GO:0008410) |
| 0.5 | 4.2 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.5 | 2.6 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.5 | 4.6 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) |
| 0.5 | 2.8 | GO:0035529 | NADH pyrophosphatase activity(GO:0035529) |
| 0.4 | 3.0 | GO:0034056 | estrogen response element binding(GO:0034056) |
| 0.4 | 2.8 | GO:0034235 | GPI anchor binding(GO:0034235) |
| 0.3 | 1.0 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.3 | 2.6 | GO:0030292 | protein tyrosine kinase inhibitor activity(GO:0030292) |
| 0.3 | 13.1 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.3 | 2.3 | GO:0030229 | very-low-density lipoprotein particle receptor activity(GO:0030229) |
| 0.3 | 6.6 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
| 0.2 | 3.6 | GO:1901702 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.2 | 0.9 | GO:0032093 | SAM domain binding(GO:0032093) |
| 0.2 | 0.9 | GO:0004008 | copper-exporting ATPase activity(GO:0004008) copper-transporting ATPase activity(GO:0043682) |
| 0.2 | 7.4 | GO:0004602 | glutathione peroxidase activity(GO:0004602) |
| 0.2 | 2.9 | GO:0008199 | ferric iron binding(GO:0008199) |
| 0.2 | 1.5 | GO:0003836 | beta-galactoside (CMP) alpha-2,3-sialyltransferase activity(GO:0003836) |
| 0.1 | 0.9 | GO:0008384 | IkappaB kinase activity(GO:0008384) |
| 0.1 | 1.0 | GO:0044729 | double-stranded methylated DNA binding(GO:0010385) hemi-methylated DNA-binding(GO:0044729) |
| 0.1 | 0.6 | GO:0036470 | tyrosine 3-monooxygenase activator activity(GO:0036470) L-dopa decarboxylase activator activity(GO:0036478) |
| 0.1 | 0.5 | GO:0004793 | threonine aldolase activity(GO:0004793) L-allo-threonine aldolase activity(GO:0008732) |
| 0.1 | 4.0 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.1 | 19.8 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 3.9 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.1 | 2.9 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.1 | 0.5 | GO:0008475 | procollagen-lysine 5-dioxygenase activity(GO:0008475) |
| 0.1 | 0.4 | GO:0031686 | A1 adenosine receptor binding(GO:0031686) |
| 0.1 | 0.2 | GO:0008311 | double-stranded DNA 3'-5' exodeoxyribonuclease activity(GO:0008311) |
| 0.1 | 3.9 | GO:0016645 | oxidoreductase activity, acting on the CH-NH group of donors(GO:0016645) |
| 0.1 | 0.4 | GO:0033677 | DNA/RNA helicase activity(GO:0033677) |
| 0.1 | 0.4 | GO:0035515 | oxidative RNA demethylase activity(GO:0035515) |
| 0.1 | 0.2 | GO:0000171 | ribonuclease MRP activity(GO:0000171) |
| 0.1 | 3.7 | GO:0004521 | endoribonuclease activity(GO:0004521) |
| 0.1 | 0.5 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.1 | 0.8 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.1 | 0.4 | GO:0016679 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors(GO:0016679) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 0.1 | 5.5 | GO:0048029 | monosaccharide binding(GO:0048029) |
| 0.0 | 0.3 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 0.0 | 0.5 | GO:0086080 | protein binding involved in heterotypic cell-cell adhesion(GO:0086080) |
| 0.0 | 2.0 | GO:0016709 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen(GO:0016709) |
| 0.0 | 3.0 | GO:0019003 | GDP binding(GO:0019003) |
| 0.0 | 15.9 | GO:0008017 | microtubule binding(GO:0008017) |
| 0.0 | 1.3 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.0 | 1.6 | GO:0003899 | DNA-directed RNA polymerase activity(GO:0003899) |
| 0.0 | 0.5 | GO:0008297 | single-stranded DNA exodeoxyribonuclease activity(GO:0008297) |
| 0.0 | 2.1 | GO:0009055 | electron carrier activity(GO:0009055) |
| 0.0 | 0.7 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.0 | 9.4 | GO:0061630 | ubiquitin protein ligase activity(GO:0061630) |
| 0.0 | 4.9 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 1.2 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.0 | 0.2 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 0.0 | 0.5 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.0 | 0.6 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.0 | 0.4 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.0 | 1.5 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.0 | 0.4 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.0 | 0.5 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.0 | 0.4 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.0 | 0.4 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 1.6 | GO:0003774 | motor activity(GO:0003774) |
| 0.0 | 0.7 | GO:0031491 | nucleosome binding(GO:0031491) |
| 0.0 | 0.7 | GO:0016504 | peptidase activator activity(GO:0016504) |
| 0.0 | 1.2 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.0 | 14.1 | GO:0019904 | protein domain specific binding(GO:0019904) |
| 0.0 | 1.2 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 16.8 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.2 | 12.4 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 1.4 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.0 | 1.8 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 7.9 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.0 | 0.9 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.0 | 3.0 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 0.9 | PID HIV NEF PATHWAY | HIV-1 Nef: Negative effector of Fas and TNF-alpha |
| 0.0 | 1.3 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.0 | 2.7 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 0.7 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.0 | 1.1 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.0 | 1.0 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 0.7 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.0 | 0.6 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.0 | 0.2 | PID MYC PATHWAY | C-MYC pathway |
| 0.0 | 0.5 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 7.4 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.5 | 16.8 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.4 | 6.7 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.4 | 8.5 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.3 | 3.9 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.3 | 5.5 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.2 | 8.0 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.2 | 2.6 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.2 | 5.3 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.1 | 4.0 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.1 | 4.6 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 14.7 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.1 | 2.6 | REACTOME ACTIVATED AMPK STIMULATES FATTY ACID OXIDATION IN MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.1 | 0.9 | REACTOME NFKB ACTIVATION THROUGH FADD RIP1 PATHWAY MEDIATED BY CASPASE 8 AND10 | Genes involved in NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 |
| 0.1 | 1.4 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.1 | 2.0 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 1.5 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.1 | 3.0 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.1 | 1.1 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 0.0 | 2.2 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.0 | 1.5 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.0 | 0.9 | REACTOME DESTABILIZATION OF MRNA BY KSRP | Genes involved in Destabilization of mRNA by KSRP |
| 0.0 | 0.4 | REACTOME DESTABILIZATION OF MRNA BY AUF1 HNRNP D0 | Genes involved in Destabilization of mRNA by AUF1 (hnRNP D0) |
| 0.0 | 0.7 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.0 | 3.9 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 0.4 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 5.6 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.0 | 1.3 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 0.4 | REACTOME RNA POL I TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase I Transcription Termination |