Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
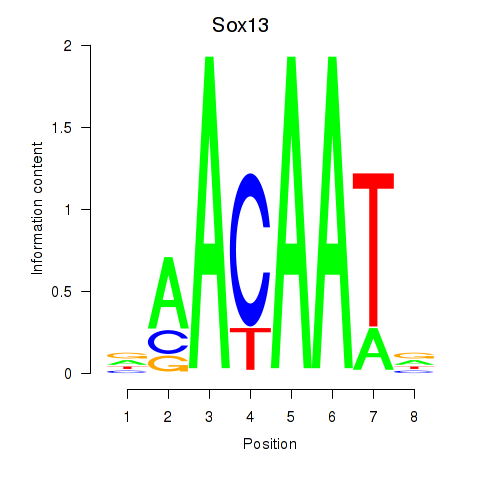
Results for Sox13
Z-value: 0.49
Transcription factors associated with Sox13
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Sox13
|
ENSMUSG00000070643.12 | Sox13 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Sox13 | mm39_v1_chr1_-_133352115_133352138 | 0.20 | 9.0e-02 | Click! |
Activity profile of Sox13 motif
Sorted Z-values of Sox13 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Sox13
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr11_-_99328969 | 6.52 |
ENSMUST00000017743.3
|
Krt20
|
keratin 20 |
| chr7_+_121818692 | 4.75 |
ENSMUST00000033152.5
|
Chp2
|
calcineurin-like EF hand protein 2 |
| chrX_+_156485570 | 2.30 |
ENSMUST00000112520.2
|
Smpx
|
small muscle protein, X-linked |
| chr8_-_62576140 | 2.05 |
ENSMUST00000034052.14
ENSMUST00000034054.9 |
Anxa10
|
annexin A10 |
| chr6_-_87312743 | 1.53 |
ENSMUST00000042025.12
ENSMUST00000205033.2 |
Antxr1
|
anthrax toxin receptor 1 |
| chr12_+_8062331 | 1.53 |
ENSMUST00000171239.2
|
Apob
|
apolipoprotein B |
| chr6_-_70435020 | 1.50 |
ENSMUST00000198184.2
|
Igkv6-13
|
immunoglobulin kappa variable 6-13 |
| chr6_-_87312681 | 1.30 |
ENSMUST00000204805.3
|
Antxr1
|
anthrax toxin receptor 1 |
| chr10_-_117074501 | 1.25 |
ENSMUST00000159193.8
ENSMUST00000020392.5 |
9530003J23Rik
|
RIKEN cDNA 9530003J23 gene |
| chr4_-_58499398 | 1.17 |
ENSMUST00000107570.2
|
Lpar1
|
lysophosphatidic acid receptor 1 |
| chrX_+_132751729 | 1.16 |
ENSMUST00000033602.9
|
Tnmd
|
tenomodulin |
| chr1_+_172327812 | 1.10 |
ENSMUST00000192460.2
|
Tagln2
|
transgelin 2 |
| chr6_-_68994064 | 1.07 |
ENSMUST00000103341.4
|
Igkv4-80
|
immunoglobulin kappa variable 4-80 |
| chr6_-_69377328 | 0.88 |
ENSMUST00000198345.2
|
Igkv4-62
|
immunoglobulin kappa variable 4-62 |
| chr19_+_41471067 | 0.83 |
ENSMUST00000067795.13
|
Lcor
|
ligand dependent nuclear receptor corepressor |
| chr3_-_141687987 | 0.67 |
ENSMUST00000029948.15
|
Bmpr1b
|
bone morphogenetic protein receptor, type 1B |
| chr2_-_104324035 | 0.67 |
ENSMUST00000111124.8
|
Hipk3
|
homeodomain interacting protein kinase 3 |
| chr6_+_48951872 | 0.60 |
ENSMUST00000184917.6
|
Doxl2
|
diamine oxidase-like protein 2 |
| chr8_+_3543131 | 0.57 |
ENSMUST00000061508.8
ENSMUST00000207318.2 |
Zfp358
|
zinc finger protein 358 |
| chr6_+_67701864 | 0.55 |
ENSMUST00000103304.3
|
Igkv1-133
|
immunoglobulin kappa variable 1-133 |
| chr12_-_104439589 | 0.51 |
ENSMUST00000021513.6
|
Gsc
|
goosecoid homeobox |
| chr3_-_75177378 | 0.44 |
ENSMUST00000039047.5
|
Serpini2
|
serine (or cysteine) peptidase inhibitor, clade I, member 2 |
| chr5_-_86666408 | 0.36 |
ENSMUST00000140095.2
ENSMUST00000134179.8 |
Tmprss11g
|
transmembrane protease, serine 11g |
| chr6_+_43150886 | 0.35 |
ENSMUST00000059512.5
|
Olfr13
|
olfactory receptor 13 |
| chr19_+_41471395 | 0.31 |
ENSMUST00000237208.2
ENSMUST00000238398.2 |
Lcor
|
ligand dependent nuclear receptor corepressor |
| chr9_+_40098375 | 0.30 |
ENSMUST00000062229.6
|
Olfr986
|
olfactory receptor 986 |
| chr6_+_131463363 | 0.25 |
ENSMUST00000159229.3
ENSMUST00000075020.10 |
Gm6619
5430401F13Rik
|
predicted gene 6619 RIKEN cDNA 5430401F13 gene |
| chr19_-_18978463 | 0.23 |
ENSMUST00000040153.15
ENSMUST00000112828.8 |
Rorb
|
RAR-related orphan receptor beta |
| chr9_+_38432876 | 0.20 |
ENSMUST00000216496.2
|
Olfr911-ps1
|
olfactory receptor 911, pseudogene 1 |
| chr6_-_93769426 | 0.18 |
ENSMUST00000204788.2
|
Magi1
|
membrane associated guanylate kinase, WW and PDZ domain containing 1 |
| chr17_-_24054717 | 0.13 |
ENSMUST00000059906.8
|
Prss33
|
protease, serine 33 |
| chr11_+_29668563 | 0.12 |
ENSMUST00000060992.6
|
Rtn4
|
reticulon 4 |
| chr10_+_57362512 | 0.11 |
ENSMUST00000220042.2
|
Hsf2
|
heat shock factor 2 |
| chr4_-_115638209 | 0.11 |
ENSMUST00000106521.2
|
Tex38
|
testis expressed 38 |
| chr10_+_57362457 | 0.09 |
ENSMUST00000079833.6
|
Hsf2
|
heat shock factor 2 |
| chr5_-_24782465 | 0.03 |
ENSMUST00000030795.10
|
Abcf2
|
ATP-binding cassette, sub-family F (GCN20), member 2 |
| chr2_-_35994072 | 0.01 |
ENSMUST00000112961.10
ENSMUST00000112966.10 |
Lhx6
|
LIM homeobox protein 6 |
| chr2_-_89448589 | 0.01 |
ENSMUST00000216129.2
|
Olfr1248
|
olfactory receptor 1248 |
| chr1_+_19173246 | 0.01 |
ENSMUST00000037294.8
|
Tfap2d
|
transcription factor AP-2, delta |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 2.8 | GO:1903054 | negative regulation of extracellular matrix organization(GO:1903054) |
| 0.3 | 1.2 | GO:1904565 | response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904565) cellular response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904566) |
| 0.2 | 6.5 | GO:0045109 | intermediate filament organization(GO:0045109) |
| 0.2 | 4.7 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade(GO:0070886) |
| 0.1 | 0.7 | GO:0048165 | ovarian cumulus expansion(GO:0001550) fused antrum stage(GO:0048165) |
| 0.1 | 1.5 | GO:0006642 | triglyceride mobilization(GO:0006642) |
| 0.1 | 1.3 | GO:0016998 | cell wall macromolecule catabolic process(GO:0016998) |
| 0.0 | 0.5 | GO:0014029 | neural crest formation(GO:0014029) |
| 0.0 | 1.2 | GO:0001886 | endothelial cell morphogenesis(GO:0001886) |
| 0.0 | 0.7 | GO:0043508 | negative regulation of JUN kinase activity(GO:0043508) |
| 0.0 | 0.2 | GO:0046549 | retinal cone cell development(GO:0046549) |
| 0.0 | 4.0 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.0 | 0.1 | GO:0071787 | endoplasmic reticulum tubular network assembly(GO:0071787) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.5 | GO:0034359 | mature chylomicron(GO:0034359) |
| 0.3 | 2.3 | GO:0005927 | muscle tendon junction(GO:0005927) |
| 0.2 | 2.8 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.0 | 6.5 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 0.3 | GO:0090543 | Flemming body(GO:0090543) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.2 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 0.1 | 1.5 | GO:0035473 | lipase binding(GO:0035473) |
| 0.1 | 1.3 | GO:0003796 | lysozyme activity(GO:0003796) |
| 0.1 | 0.7 | GO:0005025 | transforming growth factor beta receptor activity, type I(GO:0005025) |
| 0.0 | 0.2 | GO:0008502 | melatonin receptor activity(GO:0008502) |
| 0.0 | 2.8 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 2.1 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.0 | 0.2 | GO:0001162 | RNA polymerase II intronic transcription regulatory region sequence-specific DNA binding(GO:0001162) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.8 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.0 | 1.5 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 1.3 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.0 | 0.7 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 1.2 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.0 | 2.1 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.5 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 0.7 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |