Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
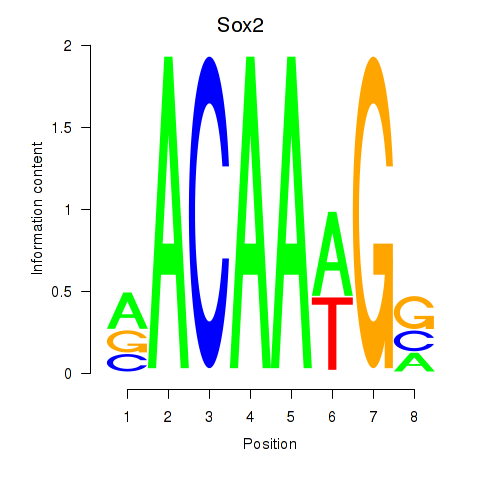
Results for Sox2
Z-value: 2.86
Transcription factors associated with Sox2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Sox2
|
ENSMUSG00000074637.8 | Sox2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Sox2 | mm39_v1_chr3_+_34704554_34704554 | 0.06 | 6.2e-01 | Click! |
Activity profile of Sox2 motif
Sorted Z-values of Sox2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Sox2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.5 | 16.5 | GO:0045660 | positive regulation of neutrophil differentiation(GO:0045660) |
| 4.8 | 14.4 | GO:0061107 | seminal vesicle development(GO:0061107) |
| 4.7 | 14.0 | GO:0071929 | alpha-tubulin acetylation(GO:0071929) |
| 4.5 | 13.6 | GO:1900039 | positive regulation of cellular response to hypoxia(GO:1900039) |
| 3.9 | 19.4 | GO:0034164 | negative regulation of toll-like receptor 9 signaling pathway(GO:0034164) |
| 3.6 | 10.9 | GO:0002581 | negative regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002581) |
| 3.5 | 10.4 | GO:0002305 | gamma-delta intraepithelial T cell differentiation(GO:0002304) CD8-positive, gamma-delta intraepithelial T cell differentiation(GO:0002305) |
| 3.0 | 12.0 | GO:1902309 | negative regulation of peptidyl-serine dephosphorylation(GO:1902309) |
| 2.9 | 11.8 | GO:1904209 | regulation of chemokine (C-C motif) ligand 2 secretion(GO:1904207) positive regulation of chemokine (C-C motif) ligand 2 secretion(GO:1904209) |
| 2.9 | 8.8 | GO:0034118 | erythrocyte aggregation(GO:0034117) regulation of erythrocyte aggregation(GO:0034118) |
| 2.8 | 11.3 | GO:1900535 | medium-chain fatty-acyl-CoA catabolic process(GO:0036114) long-chain fatty-acyl-CoA catabolic process(GO:0036116) palmitic acid metabolic process(GO:1900533) palmitic acid biosynthetic process(GO:1900535) |
| 2.8 | 8.4 | GO:0043973 | histone H3-K4 acetylation(GO:0043973) |
| 2.7 | 21.8 | GO:1900147 | Schwann cell migration(GO:0036135) regulation of Schwann cell migration(GO:1900147) |
| 2.7 | 8.1 | GO:0045074 | interleukin-10 biosynthetic process(GO:0042091) regulation of interleukin-10 biosynthetic process(GO:0045074) |
| 2.7 | 2.7 | GO:1902462 | regulation of mesenchymal stem cell proliferation(GO:1902460) positive regulation of mesenchymal stem cell proliferation(GO:1902462) |
| 2.7 | 8.0 | GO:1904020 | regulation of G-protein coupled receptor internalization(GO:1904020) |
| 2.7 | 8.0 | GO:0090673 | endothelial cell-matrix adhesion(GO:0090673) |
| 2.6 | 10.3 | GO:0032972 | regulation of muscle filament sliding speed(GO:0032972) |
| 2.5 | 17.8 | GO:0033088 | negative regulation of immature T cell proliferation in thymus(GO:0033088) |
| 2.5 | 15.2 | GO:1903182 | regulation of SUMO transferase activity(GO:1903182) positive regulation of SUMO transferase activity(GO:1903755) |
| 2.3 | 9.1 | GO:0033375 | protein localization to secretory granule(GO:0033366) protein localization to mast cell secretory granule(GO:0033367) protease localization to mast cell secretory granule(GO:0033368) maintenance of protein location in mast cell secretory granule(GO:0033370) T cell secretory granule organization(GO:0033371) maintenance of protease location in mast cell secretory granule(GO:0033373) protein localization to T cell secretory granule(GO:0033374) protease localization to T cell secretory granule(GO:0033375) maintenance of protein location in T cell secretory granule(GO:0033377) maintenance of protease location in T cell secretory granule(GO:0033379) granzyme B localization to T cell secretory granule(GO:0033380) maintenance of granzyme B location in T cell secretory granule(GO:0033382) |
| 2.3 | 20.6 | GO:0030035 | microspike assembly(GO:0030035) |
| 2.3 | 9.1 | GO:0050717 | positive regulation of interleukin-1 alpha secretion(GO:0050717) |
| 2.3 | 16.0 | GO:0090235 | regulation of metaphase plate congression(GO:0090235) |
| 2.2 | 6.7 | GO:0036145 | dendritic cell homeostasis(GO:0036145) |
| 2.1 | 8.6 | GO:0061152 | trachea submucosa development(GO:0061152) trachea gland development(GO:0061153) |
| 2.0 | 5.9 | GO:0060448 | dichotomous subdivision of terminal units involved in lung branching(GO:0060448) |
| 1.9 | 9.7 | GO:0043490 | malate-aspartate shuttle(GO:0043490) |
| 1.9 | 9.5 | GO:0003349 | epicardium-derived cardiac endothelial cell differentiation(GO:0003349) |
| 1.9 | 5.6 | GO:0019417 | sulfur oxidation(GO:0019417) |
| 1.9 | 7.4 | GO:1901003 | negative regulation of fermentation(GO:1901003) |
| 1.8 | 3.6 | GO:0032971 | regulation of muscle filament sliding(GO:0032971) |
| 1.8 | 14.5 | GO:0071802 | negative regulation of podosome assembly(GO:0071802) |
| 1.7 | 7.0 | GO:1904158 | axonemal central apparatus assembly(GO:1904158) |
| 1.7 | 8.6 | GO:0031022 | nuclear migration along microfilament(GO:0031022) |
| 1.6 | 6.6 | GO:1904565 | response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904565) cellular response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904566) |
| 1.6 | 4.9 | GO:0061762 | CAMKK-AMPK signaling cascade(GO:0061762) |
| 1.6 | 11.3 | GO:0002727 | natural killer cell cytokine production(GO:0002370) regulation of natural killer cell cytokine production(GO:0002727) |
| 1.5 | 6.1 | GO:0051562 | negative regulation of mitochondrial calcium ion concentration(GO:0051562) |
| 1.5 | 7.6 | GO:0038032 | termination of G-protein coupled receptor signaling pathway(GO:0038032) |
| 1.5 | 5.8 | GO:1902164 | mRNA localization resulting in posttranscriptional regulation of gene expression(GO:0010609) positive regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902164) |
| 1.4 | 7.1 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 1.4 | 5.6 | GO:2000312 | regulation of kainate selective glutamate receptor activity(GO:2000312) |
| 1.4 | 4.2 | GO:0060392 | negative regulation of SMAD protein import into nucleus(GO:0060392) |
| 1.4 | 8.4 | GO:0006438 | valyl-tRNA aminoacylation(GO:0006438) |
| 1.4 | 9.5 | GO:0043553 | negative regulation of phosphatidylinositol 3-kinase activity(GO:0043553) |
| 1.4 | 10.8 | GO:0002681 | somatic diversification of T cell receptor genes(GO:0002568) somatic recombination of T cell receptor gene segments(GO:0002681) T cell receptor V(D)J recombination(GO:0033153) |
| 1.3 | 8.0 | GO:1903964 | monounsaturated fatty acid metabolic process(GO:1903964) monounsaturated fatty acid biosynthetic process(GO:1903966) |
| 1.3 | 66.4 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 1.3 | 4.0 | GO:0034378 | chylomicron assembly(GO:0034378) |
| 1.3 | 3.9 | GO:0051466 | positive regulation of corticotropin-releasing hormone secretion(GO:0051466) |
| 1.3 | 7.7 | GO:0006172 | ADP biosynthetic process(GO:0006172) |
| 1.3 | 7.7 | GO:0035120 | post-embryonic appendage morphogenesis(GO:0035120) post-embryonic limb morphogenesis(GO:0035127) post-embryonic forelimb morphogenesis(GO:0035128) telomeric repeat-containing RNA transcription(GO:0097393) telomeric repeat-containing RNA transcription from RNA pol II promoter(GO:0097394) regulation of telomeric RNA transcription from RNA pol II promoter(GO:1901580) negative regulation of telomeric RNA transcription from RNA pol II promoter(GO:1901581) positive regulation of telomeric RNA transcription from RNA pol II promoter(GO:1901582) |
| 1.3 | 3.8 | GO:0061723 | glycophagy(GO:0061723) |
| 1.3 | 12.6 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 1.2 | 5.0 | GO:1900191 | biofilm formation(GO:0042710) single-species biofilm formation(GO:0044010) single-species biofilm formation in or on host organism(GO:0044407) membrane disruption in other organism(GO:0051673) regulation of single-species biofilm formation(GO:1900190) negative regulation of single-species biofilm formation(GO:1900191) regulation of single-species biofilm formation in or on host organism(GO:1900228) negative regulation of single-species biofilm formation in or on host organism(GO:1900229) |
| 1.2 | 3.7 | GO:0071707 | immunoglobulin heavy chain V-D-J recombination(GO:0071707) |
| 1.2 | 13.5 | GO:1903608 | protein localization to cytoplasmic stress granule(GO:1903608) |
| 1.2 | 23.7 | GO:1902004 | positive regulation of beta-amyloid formation(GO:1902004) |
| 1.2 | 17.7 | GO:1900028 | negative regulation of ruffle assembly(GO:1900028) |
| 1.2 | 5.9 | GO:1903898 | negative regulation of PERK-mediated unfolded protein response(GO:1903898) |
| 1.2 | 4.7 | GO:0002528 | regulation of vascular permeability involved in acute inflammatory response(GO:0002528) |
| 1.2 | 6.9 | GO:1903054 | negative regulation of extracellular matrix organization(GO:1903054) |
| 1.1 | 5.7 | GO:1902775 | mitochondrial large ribosomal subunit assembly(GO:1902775) |
| 1.1 | 3.4 | GO:2001160 | regulation of histone H3-K79 methylation(GO:2001160) |
| 1.1 | 4.5 | GO:1902268 | negative regulation of polyamine transmembrane transport(GO:1902268) |
| 1.1 | 21.0 | GO:0033089 | positive regulation of T cell differentiation in thymus(GO:0033089) positive regulation of thymocyte aggregation(GO:2000400) |
| 1.1 | 1.1 | GO:0040030 | regulation of molecular function, epigenetic(GO:0040030) |
| 1.1 | 8.8 | GO:0033184 | positive regulation of histone ubiquitination(GO:0033184) |
| 1.1 | 4.3 | GO:0060279 | positive regulation of ovulation(GO:0060279) |
| 1.1 | 4.3 | GO:0051586 | positive regulation of neurotransmitter uptake(GO:0051582) positive regulation of dopamine uptake involved in synaptic transmission(GO:0051586) positive regulation of catecholamine uptake involved in synaptic transmission(GO:0051944) |
| 1.1 | 12.7 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 1.0 | 4.2 | GO:0021586 | pons maturation(GO:0021586) |
| 1.0 | 20.8 | GO:0061179 | negative regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0061179) |
| 1.0 | 6.2 | GO:1903265 | positive regulation of tumor necrosis factor-mediated signaling pathway(GO:1903265) |
| 1.0 | 4.1 | GO:1905154 | negative regulation of clathrin-mediated endocytosis(GO:1900186) negative regulation of synaptic vesicle recycling(GO:1903422) negative regulation of membrane invagination(GO:1905154) calcium ion regulated lysosome exocytosis(GO:1990927) |
| 1.0 | 3.1 | GO:0048239 | negative regulation of DNA recombination at telomere(GO:0048239) regulation of DNA recombination at telomere(GO:0072695) |
| 1.0 | 12.3 | GO:0035879 | lactate transport(GO:0015727) lactate transmembrane transport(GO:0035873) plasma membrane lactate transport(GO:0035879) |
| 1.0 | 9.1 | GO:0038026 | reelin-mediated signaling pathway(GO:0038026) |
| 1.0 | 3.0 | GO:0072385 | minus-end-directed organelle transport along microtubule(GO:0072385) |
| 1.0 | 4.9 | GO:0061086 | negative regulation of histone H3-K27 methylation(GO:0061086) |
| 1.0 | 9.8 | GO:0030206 | chondroitin sulfate biosynthetic process(GO:0030206) |
| 1.0 | 7.6 | GO:0061366 | behavioral response to chemical pain(GO:0061366) behavioral response to formalin induced pain(GO:0061368) |
| 0.9 | 2.8 | GO:2000097 | negative regulation of lipoprotein oxidation(GO:0034443) regulation of smooth muscle cell-matrix adhesion(GO:2000097) |
| 0.9 | 18.0 | GO:1902083 | negative regulation of peptidyl-cysteine S-nitrosylation(GO:1902083) |
| 0.9 | 9.4 | GO:0098887 | neurotransmitter receptor transport, endosome to postsynaptic membrane(GO:0098887) |
| 0.9 | 8.4 | GO:0042699 | follicle-stimulating hormone signaling pathway(GO:0042699) |
| 0.9 | 16.8 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.9 | 5.6 | GO:0033030 | negative regulation of neutrophil apoptotic process(GO:0033030) |
| 0.9 | 6.5 | GO:0061083 | regulation of protein refolding(GO:0061083) negative regulation of protein refolding(GO:0061084) |
| 0.9 | 15.8 | GO:0090527 | actin filament reorganization(GO:0090527) |
| 0.9 | 12.0 | GO:0051152 | positive regulation of smooth muscle cell differentiation(GO:0051152) |
| 0.9 | 1.8 | GO:0036166 | phenotypic switching(GO:0036166) |
| 0.9 | 6.4 | GO:0045200 | establishment or maintenance of neuroblast polarity(GO:0045196) establishment of neuroblast polarity(GO:0045200) |
| 0.9 | 2.7 | GO:0061511 | centriole elongation(GO:0061511) |
| 0.9 | 12.6 | GO:0031274 | positive regulation of pseudopodium assembly(GO:0031274) |
| 0.9 | 22.1 | GO:0007202 | activation of phospholipase C activity(GO:0007202) |
| 0.9 | 21.7 | GO:0030213 | hyaluronan biosynthetic process(GO:0030213) |
| 0.9 | 16.4 | GO:0006337 | nucleosome disassembly(GO:0006337) |
| 0.9 | 7.7 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.8 | 13.5 | GO:0070933 | histone H4 deacetylation(GO:0070933) |
| 0.8 | 7.6 | GO:0032596 | protein transport into membrane raft(GO:0032596) |
| 0.8 | 2.5 | GO:0031283 | negative regulation of cGMP metabolic process(GO:0030824) negative regulation of cGMP biosynthetic process(GO:0030827) negative regulation of guanylate cyclase activity(GO:0031283) |
| 0.8 | 2.5 | GO:1901074 | regulation of engulfment of apoptotic cell(GO:1901074) |
| 0.8 | 3.3 | GO:0033575 | protein glycosylation at cell surface(GO:0033575) protein galactosylation at cell surface(GO:0033580) protein galactosylation(GO:0042125) |
| 0.8 | 2.4 | GO:0034670 | chemotaxis to arachidonic acid(GO:0034670) response to arachidonic acid(GO:1904550) |
| 0.8 | 7.2 | GO:0060753 | regulation of mast cell chemotaxis(GO:0060753) |
| 0.8 | 2.3 | GO:0014862 | regulation of the force of skeletal muscle contraction(GO:0014728) regulation of skeletal muscle contraction by chemo-mechanical energy conversion(GO:0014862) |
| 0.8 | 11.6 | GO:0034242 | negative regulation of syncytium formation by plasma membrane fusion(GO:0034242) |
| 0.8 | 2.3 | GO:0046985 | positive regulation of hemoglobin biosynthetic process(GO:0046985) |
| 0.8 | 3.8 | GO:0019086 | late viral transcription(GO:0019086) |
| 0.7 | 6.7 | GO:0060633 | negative regulation of transcription initiation from RNA polymerase II promoter(GO:0060633) |
| 0.7 | 3.6 | GO:0046013 | regulation of T cell homeostatic proliferation(GO:0046013) |
| 0.7 | 19.9 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.7 | 2.8 | GO:0061188 | regulation of chromatin silencing at rDNA(GO:0061187) negative regulation of chromatin silencing at rDNA(GO:0061188) |
| 0.7 | 2.8 | GO:0045399 | positive regulation of interleukin-3 production(GO:0032752) interleukin-3 biosynthetic process(GO:0042223) regulation of interleukin-3 biosynthetic process(GO:0045399) positive regulation of interleukin-3 biosynthetic process(GO:0045401) positive regulation of granulocyte macrophage colony-stimulating factor biosynthetic process(GO:0045425) cellular response to molecule of fungal origin(GO:0071226) |
| 0.7 | 3.5 | GO:2000048 | negative regulation of cell-cell adhesion mediated by cadherin(GO:2000048) |
| 0.7 | 2.1 | GO:1901642 | purine nucleoside transmembrane transport(GO:0015860) nucleoside transmembrane transport(GO:1901642) |
| 0.7 | 3.4 | GO:0002326 | B cell lineage commitment(GO:0002326) |
| 0.7 | 9.4 | GO:0051574 | positive regulation of histone H3-K9 methylation(GO:0051574) |
| 0.7 | 8.7 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.7 | 2.0 | GO:0071846 | actin filament debranching(GO:0071846) |
| 0.7 | 0.7 | GO:1904009 | response to monosodium glutamate(GO:1904008) cellular response to monosodium glutamate(GO:1904009) |
| 0.6 | 3.9 | GO:1900041 | negative regulation of interleukin-2 secretion(GO:1900041) |
| 0.6 | 5.2 | GO:1902951 | negative regulation of dendritic spine maintenance(GO:1902951) |
| 0.6 | 1.9 | GO:0060217 | hemangioblast cell differentiation(GO:0060217) |
| 0.6 | 1.9 | GO:0002572 | pro-T cell differentiation(GO:0002572) |
| 0.6 | 2.6 | GO:0002949 | tRNA threonylcarbamoyladenosine modification(GO:0002949) |
| 0.6 | 17.7 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.6 | 12.6 | GO:2000480 | negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
| 0.6 | 7.6 | GO:0030050 | vesicle transport along actin filament(GO:0030050) |
| 0.6 | 4.4 | GO:2000195 | negative regulation of female gonad development(GO:2000195) |
| 0.6 | 6.8 | GO:0006527 | arginine catabolic process(GO:0006527) |
| 0.6 | 1.9 | GO:1990428 | miRNA transport(GO:1990428) |
| 0.6 | 4.9 | GO:0006868 | glutamine transport(GO:0006868) |
| 0.6 | 6.0 | GO:0016198 | axon choice point recognition(GO:0016198) |
| 0.6 | 1.8 | GO:2001200 | positive regulation of dendritic cell differentiation(GO:2001200) |
| 0.6 | 1.2 | GO:0070934 | CRD-mediated mRNA stabilization(GO:0070934) |
| 0.6 | 7.7 | GO:1901898 | negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.6 | 2.4 | GO:0003290 | atrial septum secundum morphogenesis(GO:0003290) |
| 0.6 | 9.4 | GO:0072697 | protein localization to cell cortex(GO:0072697) |
| 0.6 | 13.5 | GO:0000920 | cell separation after cytokinesis(GO:0000920) |
| 0.6 | 2.3 | GO:1901491 | negative regulation of lymphangiogenesis(GO:1901491) |
| 0.6 | 2.3 | GO:0046898 | response to cycloheximide(GO:0046898) |
| 0.6 | 7.0 | GO:1904672 | regulation of somatic stem cell population maintenance(GO:1904672) |
| 0.6 | 4.7 | GO:0043314 | negative regulation of neutrophil degranulation(GO:0043314) negative regulation of blood vessel remodeling(GO:0060313) negative regulation of neutrophil activation(GO:1902564) |
| 0.6 | 4.0 | GO:2000427 | positive regulation of apoptotic cell clearance(GO:2000427) |
| 0.6 | 6.2 | GO:0001781 | neutrophil apoptotic process(GO:0001781) |
| 0.6 | 4.0 | GO:0032687 | negative regulation of interferon-alpha production(GO:0032687) |
| 0.6 | 1.7 | GO:0002631 | regulation of granuloma formation(GO:0002631) negative regulation of granuloma formation(GO:0002632) regulation of toll-like receptor 5 signaling pathway(GO:0034147) negative regulation of toll-like receptor 5 signaling pathway(GO:0034148) negative regulation of nucleotide-binding oligomerization domain containing 1 signaling pathway(GO:0070429) |
| 0.6 | 2.3 | GO:0046351 | disaccharide biosynthetic process(GO:0046351) |
| 0.6 | 4.5 | GO:2000271 | positive regulation of fibroblast apoptotic process(GO:2000271) |
| 0.6 | 2.2 | GO:1990091 | sodium-dependent self proteolysis(GO:1990091) |
| 0.6 | 1.7 | GO:0042668 | auditory receptor cell fate determination(GO:0042668) negative regulation of forebrain neuron differentiation(GO:2000978) |
| 0.6 | 4.4 | GO:1990086 | lens fiber cell apoptotic process(GO:1990086) |
| 0.6 | 1.7 | GO:0030887 | positive regulation of myeloid dendritic cell activation(GO:0030887) |
| 0.6 | 1.7 | GO:0003245 | cardiac muscle tissue growth involved in heart morphogenesis(GO:0003245) |
| 0.5 | 1.6 | GO:0060166 | olfactory pit development(GO:0060166) |
| 0.5 | 6.5 | GO:0060346 | bone trabecula formation(GO:0060346) |
| 0.5 | 2.2 | GO:1901187 | regulation of ephrin receptor signaling pathway(GO:1901187) |
| 0.5 | 4.3 | GO:0089711 | L-glutamate transmembrane transport(GO:0089711) |
| 0.5 | 2.7 | GO:0006235 | dTTP biosynthetic process(GO:0006235) pyrimidine deoxyribonucleoside triphosphate biosynthetic process(GO:0009212) dTTP metabolic process(GO:0046075) |
| 0.5 | 2.7 | GO:0055059 | asymmetric neuroblast division(GO:0055059) regulation of timing of neuron differentiation(GO:0060164) |
| 0.5 | 1.1 | GO:1904616 | regulation of actin filament binding(GO:1904529) regulation of actin binding(GO:1904616) |
| 0.5 | 4.8 | GO:1990253 | cellular response to leucine starvation(GO:1990253) |
| 0.5 | 3.2 | GO:0007412 | axon target recognition(GO:0007412) |
| 0.5 | 3.7 | GO:0045163 | clustering of voltage-gated potassium channels(GO:0045163) |
| 0.5 | 3.7 | GO:0051012 | microtubule sliding(GO:0051012) |
| 0.5 | 3.6 | GO:0042796 | snRNA transcription from RNA polymerase III promoter(GO:0042796) |
| 0.5 | 2.0 | GO:0055011 | atrial cardiac muscle cell differentiation(GO:0055011) atrial cardiac muscle cell development(GO:0055014) |
| 0.5 | 6.6 | GO:0043117 | positive regulation of vascular permeability(GO:0043117) |
| 0.5 | 2.5 | GO:0042631 | cellular response to water deprivation(GO:0042631) |
| 0.5 | 1.4 | GO:0060821 | inactivation of X chromosome by DNA methylation(GO:0060821) |
| 0.5 | 1.4 | GO:1904684 | negative regulation of metalloendopeptidase activity(GO:1904684) |
| 0.5 | 1.9 | GO:0021993 | initiation of neural tube closure(GO:0021993) |
| 0.5 | 1.9 | GO:0007023 | post-chaperonin tubulin folding pathway(GO:0007023) |
| 0.5 | 1.4 | GO:0002270 | plasmacytoid dendritic cell activation(GO:0002270) negative regulation of apoptotic cell clearance(GO:2000426) |
| 0.5 | 4.1 | GO:0046208 | spermine catabolic process(GO:0046208) |
| 0.5 | 9.2 | GO:0071786 | endoplasmic reticulum tubular network organization(GO:0071786) |
| 0.5 | 10.1 | GO:0043383 | negative T cell selection(GO:0043383) |
| 0.5 | 3.2 | GO:0051013 | microtubule severing(GO:0051013) |
| 0.5 | 4.5 | GO:0090286 | cytoskeletal anchoring at nuclear membrane(GO:0090286) |
| 0.5 | 1.8 | GO:1900739 | regulation of protein insertion into mitochondrial membrane involved in apoptotic signaling pathway(GO:1900739) positive regulation of protein insertion into mitochondrial membrane involved in apoptotic signaling pathway(GO:1900740) |
| 0.4 | 1.3 | GO:0046832 | negative regulation of nucleobase-containing compound transport(GO:0032240) negative regulation of RNA export from nucleus(GO:0046832) |
| 0.4 | 5.8 | GO:0097435 | fibril organization(GO:0097435) |
| 0.4 | 9.2 | GO:0006620 | posttranslational protein targeting to membrane(GO:0006620) |
| 0.4 | 5.3 | GO:0016191 | synaptic vesicle uncoating(GO:0016191) |
| 0.4 | 2.6 | GO:0061589 | calcium activated phosphatidylserine scrambling(GO:0061589) |
| 0.4 | 2.1 | GO:0060221 | retinal rod cell differentiation(GO:0060221) |
| 0.4 | 1.3 | GO:0007228 | positive regulation of hh target transcription factor activity(GO:0007228) |
| 0.4 | 1.7 | GO:0060709 | glycogen cell differentiation involved in embryonic placenta development(GO:0060709) |
| 0.4 | 7.1 | GO:0036159 | inner dynein arm assembly(GO:0036159) |
| 0.4 | 4.6 | GO:0035507 | regulation of myosin-light-chain-phosphatase activity(GO:0035507) |
| 0.4 | 2.9 | GO:0060468 | prevention of polyspermy(GO:0060468) |
| 0.4 | 6.2 | GO:0008343 | adult feeding behavior(GO:0008343) |
| 0.4 | 5.7 | GO:0072600 | establishment of protein localization to Golgi(GO:0072600) |
| 0.4 | 8.5 | GO:0046069 | cGMP catabolic process(GO:0046069) |
| 0.4 | 9.6 | GO:0031115 | negative regulation of microtubule polymerization(GO:0031115) |
| 0.4 | 4.4 | GO:0098789 | pre-mRNA cleavage required for polyadenylation(GO:0098789) |
| 0.4 | 2.0 | GO:0009301 | snRNA transcription(GO:0009301) |
| 0.4 | 4.7 | GO:0043615 | astrocyte cell migration(GO:0043615) |
| 0.4 | 5.1 | GO:1901339 | regulation of store-operated calcium channel activity(GO:1901339) |
| 0.4 | 7.3 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.4 | 3.5 | GO:0010792 | DNA double-strand break processing involved in repair via single-strand annealing(GO:0010792) |
| 0.4 | 8.0 | GO:0001675 | acrosome assembly(GO:0001675) |
| 0.4 | 1.1 | GO:0014878 | response to electrical stimulus involved in regulation of muscle adaptation(GO:0014878) |
| 0.4 | 1.1 | GO:0072708 | response to sorbitol(GO:0072708) response to dithiothreitol(GO:0072720) |
| 0.4 | 9.3 | GO:0072520 | seminiferous tubule development(GO:0072520) |
| 0.4 | 4.4 | GO:0010606 | positive regulation of cytoplasmic mRNA processing body assembly(GO:0010606) |
| 0.4 | 7.8 | GO:0007141 | male meiosis I(GO:0007141) |
| 0.4 | 5.9 | GO:0010739 | positive regulation of protein kinase A signaling(GO:0010739) |
| 0.4 | 2.6 | GO:0097324 | melanocyte migration(GO:0097324) |
| 0.4 | 2.9 | GO:0006561 | proline biosynthetic process(GO:0006561) |
| 0.4 | 2.2 | GO:0002913 | positive regulation of T cell anergy(GO:0002669) positive regulation of lymphocyte anergy(GO:0002913) |
| 0.4 | 1.4 | GO:2000297 | negative regulation of synapse maturation(GO:2000297) |
| 0.4 | 1.8 | GO:0031509 | telomeric heterochromatin assembly(GO:0031509) negative regulation of chromosome condensation(GO:1902340) |
| 0.4 | 7.4 | GO:0044406 | adhesion of symbiont to host(GO:0044406) |
| 0.4 | 10.5 | GO:0016242 | negative regulation of macroautophagy(GO:0016242) |
| 0.3 | 6.6 | GO:0035090 | maintenance of apical/basal cell polarity(GO:0035090) |
| 0.3 | 2.8 | GO:0001672 | regulation of chromatin assembly or disassembly(GO:0001672) |
| 0.3 | 3.1 | GO:1902952 | positive regulation of dendritic spine maintenance(GO:1902952) |
| 0.3 | 4.4 | GO:0035845 | photoreceptor cell outer segment organization(GO:0035845) |
| 0.3 | 3.4 | GO:0002098 | tRNA wobble uridine modification(GO:0002098) |
| 0.3 | 3.1 | GO:0019532 | oxalate transport(GO:0019532) |
| 0.3 | 3.4 | GO:1902514 | regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1902514) |
| 0.3 | 11.8 | GO:0034243 | regulation of transcription elongation from RNA polymerase II promoter(GO:0034243) |
| 0.3 | 1.7 | GO:0048687 | positive regulation of sprouting of injured axon(GO:0048687) positive regulation of axon extension involved in regeneration(GO:0048691) |
| 0.3 | 2.3 | GO:0010936 | negative regulation of macrophage cytokine production(GO:0010936) |
| 0.3 | 2.0 | GO:0097118 | neuroligin clustering involved in postsynaptic membrane assembly(GO:0097118) |
| 0.3 | 2.6 | GO:0090074 | negative regulation of protein homodimerization activity(GO:0090074) |
| 0.3 | 3.9 | GO:0051901 | positive regulation of mitochondrial depolarization(GO:0051901) |
| 0.3 | 2.3 | GO:0009313 | ganglioside catabolic process(GO:0006689) oligosaccharide catabolic process(GO:0009313) |
| 0.3 | 2.6 | GO:1900086 | positive regulation of peptidyl-tyrosine autophosphorylation(GO:1900086) |
| 0.3 | 2.6 | GO:0070417 | cellular response to cold(GO:0070417) |
| 0.3 | 2.8 | GO:0019372 | lipoxygenase pathway(GO:0019372) |
| 0.3 | 3.4 | GO:0006335 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.3 | 3.7 | GO:0090050 | positive regulation of cell migration involved in sprouting angiogenesis(GO:0090050) |
| 0.3 | 1.2 | GO:0001928 | regulation of exocyst assembly(GO:0001928) |
| 0.3 | 2.7 | GO:0061577 | calcium ion transmembrane transport via high voltage-gated calcium channel(GO:0061577) |
| 0.3 | 1.2 | GO:0070889 | platelet alpha granule organization(GO:0070889) |
| 0.3 | 0.6 | GO:2000017 | positive regulation of determination of dorsal identity(GO:2000017) |
| 0.3 | 4.3 | GO:0016576 | histone dephosphorylation(GO:0016576) |
| 0.3 | 1.7 | GO:0071698 | olfactory placode formation(GO:0030910) olfactory placode development(GO:0071698) olfactory placode morphogenesis(GO:0071699) |
| 0.3 | 11.8 | GO:0043551 | regulation of phosphatidylinositol 3-kinase activity(GO:0043551) |
| 0.3 | 2.3 | GO:0033484 | nitric oxide homeostasis(GO:0033484) |
| 0.3 | 2.2 | GO:0016560 | protein import into peroxisome matrix, docking(GO:0016560) |
| 0.3 | 2.2 | GO:0061470 | T follicular helper cell differentiation(GO:0061470) |
| 0.3 | 1.7 | GO:1901842 | negative regulation of high voltage-gated calcium channel activity(GO:1901842) |
| 0.3 | 11.0 | GO:0051290 | protein heterotetramerization(GO:0051290) |
| 0.3 | 9.0 | GO:0061157 | mRNA destabilization(GO:0061157) |
| 0.3 | 3.0 | GO:0000022 | mitotic spindle elongation(GO:0000022) mitotic spindle midzone assembly(GO:0051256) |
| 0.3 | 0.5 | GO:0071225 | cellular response to muramyl dipeptide(GO:0071225) |
| 0.3 | 4.6 | GO:0070816 | phosphorylation of RNA polymerase II C-terminal domain(GO:0070816) |
| 0.3 | 1.9 | GO:0035902 | response to immobilization stress(GO:0035902) |
| 0.3 | 3.0 | GO:0032784 | regulation of DNA-templated transcription, elongation(GO:0032784) |
| 0.3 | 9.9 | GO:0030261 | chromosome condensation(GO:0030261) |
| 0.3 | 1.6 | GO:1990166 | positive regulation of maintenance of mitotic sister chromatid cohesion(GO:0034184) protein localization to site of double-strand break(GO:1990166) |
| 0.3 | 7.2 | GO:0008210 | estrogen metabolic process(GO:0008210) |
| 0.3 | 2.4 | GO:0040031 | snRNA modification(GO:0040031) |
| 0.3 | 1.9 | GO:0007527 | adult somatic muscle development(GO:0007527) |
| 0.3 | 0.8 | GO:0040010 | positive regulation of growth rate(GO:0040010) |
| 0.3 | 3.4 | GO:0048681 | negative regulation of axon regeneration(GO:0048681) |
| 0.3 | 0.3 | GO:0006295 | nucleotide-excision repair, DNA incision, 3'-to lesion(GO:0006295) |
| 0.3 | 0.8 | GO:0014028 | notochord formation(GO:0014028) |
| 0.3 | 9.9 | GO:1902042 | negative regulation of extrinsic apoptotic signaling pathway via death domain receptors(GO:1902042) |
| 0.3 | 2.5 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.3 | 1.0 | GO:0031053 | primary miRNA processing(GO:0031053) |
| 0.3 | 5.3 | GO:2000785 | regulation of autophagosome assembly(GO:2000785) |
| 0.3 | 11.3 | GO:0031648 | protein destabilization(GO:0031648) |
| 0.2 | 0.5 | GO:0099566 | regulation of postsynaptic cytosolic calcium ion concentration(GO:0099566) |
| 0.2 | 1.5 | GO:0086103 | adrenergic receptor signaling pathway involved in heart process(GO:0086023) G-protein coupled receptor signaling pathway involved in heart process(GO:0086103) |
| 0.2 | 6.9 | GO:0007095 | mitotic G2 DNA damage checkpoint(GO:0007095) |
| 0.2 | 1.5 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 0.2 | 1.0 | GO:0021691 | cerebellar Purkinje cell layer maturation(GO:0021691) |
| 0.2 | 2.4 | GO:0045719 | negative regulation of glycogen biosynthetic process(GO:0045719) |
| 0.2 | 1.0 | GO:0021914 | negative regulation of smoothened signaling pathway involved in ventral spinal cord patterning(GO:0021914) |
| 0.2 | 2.2 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 0.2 | 0.7 | GO:0001880 | Mullerian duct regression(GO:0001880) |
| 0.2 | 1.7 | GO:1902669 | positive regulation of axon guidance(GO:1902669) |
| 0.2 | 1.0 | GO:2000301 | negative regulation of synaptic vesicle exocytosis(GO:2000301) |
| 0.2 | 0.5 | GO:0070885 | negative regulation of calcineurin-NFAT signaling cascade(GO:0070885) |
| 0.2 | 5.5 | GO:1903019 | negative regulation of glycoprotein metabolic process(GO:1903019) |
| 0.2 | 1.7 | GO:0060023 | soft palate development(GO:0060023) |
| 0.2 | 2.4 | GO:0031468 | nuclear envelope reassembly(GO:0031468) |
| 0.2 | 1.9 | GO:1903715 | regulation of aerobic respiration(GO:1903715) |
| 0.2 | 4.7 | GO:0035584 | calcium-mediated signaling using intracellular calcium source(GO:0035584) |
| 0.2 | 8.6 | GO:0044342 | type B pancreatic cell proliferation(GO:0044342) |
| 0.2 | 3.2 | GO:0045161 | neuronal ion channel clustering(GO:0045161) |
| 0.2 | 1.8 | GO:1904378 | maintenance of unfolded protein(GO:0036506) maintenance of unfolded protein involved in ERAD pathway(GO:1904378) |
| 0.2 | 1.4 | GO:0000722 | telomere maintenance via recombination(GO:0000722) |
| 0.2 | 9.8 | GO:0050852 | T cell receptor signaling pathway(GO:0050852) |
| 0.2 | 2.1 | GO:0006002 | fructose 6-phosphate metabolic process(GO:0006002) |
| 0.2 | 4.3 | GO:0045589 | regulation of regulatory T cell differentiation(GO:0045589) |
| 0.2 | 2.7 | GO:2000121 | regulation of removal of superoxide radicals(GO:2000121) |
| 0.2 | 1.1 | GO:0007341 | penetration of zona pellucida(GO:0007341) |
| 0.2 | 0.9 | GO:0019509 | L-methionine biosynthetic process from methylthioadenosine(GO:0019509) |
| 0.2 | 2.9 | GO:1990126 | retrograde transport, endosome to plasma membrane(GO:1990126) |
| 0.2 | 3.1 | GO:0034498 | early endosome to Golgi transport(GO:0034498) |
| 0.2 | 3.6 | GO:0043653 | mitochondrial fragmentation involved in apoptotic process(GO:0043653) |
| 0.2 | 0.4 | GO:0001922 | B-1 B cell homeostasis(GO:0001922) |
| 0.2 | 7.5 | GO:0002467 | germinal center formation(GO:0002467) |
| 0.2 | 6.4 | GO:0050901 | leukocyte tethering or rolling(GO:0050901) |
| 0.2 | 3.5 | GO:1902187 | negative regulation of viral release from host cell(GO:1902187) |
| 0.2 | 2.1 | GO:0046642 | negative regulation of alpha-beta T cell proliferation(GO:0046642) |
| 0.2 | 1.9 | GO:2001135 | regulation of endocytic recycling(GO:2001135) |
| 0.2 | 0.9 | GO:0097119 | postsynaptic density protein 95 clustering(GO:0097119) |
| 0.2 | 6.6 | GO:0031111 | negative regulation of microtubule polymerization or depolymerization(GO:0031111) |
| 0.2 | 2.3 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.2 | 3.8 | GO:0070570 | regulation of neuron projection regeneration(GO:0070570) |
| 0.2 | 5.2 | GO:0050860 | negative regulation of T cell receptor signaling pathway(GO:0050860) |
| 0.2 | 1.0 | GO:0030576 | Cajal body organization(GO:0030576) |
| 0.2 | 0.6 | GO:0003162 | atrioventricular node development(GO:0003162) |
| 0.2 | 0.8 | GO:0060988 | lipid tube assembly(GO:0060988) |
| 0.2 | 1.8 | GO:0070236 | negative regulation of activation-induced cell death of T cells(GO:0070236) |
| 0.2 | 2.6 | GO:0035092 | sperm chromatin condensation(GO:0035092) |
| 0.2 | 1.6 | GO:0036158 | outer dynein arm assembly(GO:0036158) |
| 0.2 | 2.0 | GO:0014842 | regulation of skeletal muscle satellite cell proliferation(GO:0014842) |
| 0.2 | 1.2 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 0.2 | 1.0 | GO:0090271 | positive regulation of fibroblast growth factor production(GO:0090271) |
| 0.2 | 2.5 | GO:0001574 | ganglioside biosynthetic process(GO:0001574) |
| 0.2 | 3.6 | GO:0001967 | suckling behavior(GO:0001967) |
| 0.2 | 1.1 | GO:0060296 | regulation of cilium movement involved in cell motility(GO:0060295) regulation of cilium beat frequency involved in ciliary motility(GO:0060296) regulation of cilium-dependent cell motility(GO:1902019) |
| 0.2 | 0.4 | GO:1903116 | positive regulation of actin filament-based movement(GO:1903116) |
| 0.2 | 0.4 | GO:0045625 | regulation of T-helper 1 cell differentiation(GO:0045625) |
| 0.2 | 3.0 | GO:1903861 | positive regulation of dendrite extension(GO:1903861) |
| 0.2 | 2.6 | GO:0006968 | cellular defense response(GO:0006968) |
| 0.2 | 2.4 | GO:1903546 | protein localization to photoreceptor outer segment(GO:1903546) |
| 0.2 | 15.9 | GO:0051225 | spindle assembly(GO:0051225) |
| 0.2 | 6.6 | GO:1901385 | regulation of voltage-gated calcium channel activity(GO:1901385) |
| 0.2 | 4.6 | GO:0042118 | endothelial cell activation(GO:0042118) |
| 0.2 | 1.3 | GO:0042090 | interleukin-12 biosynthetic process(GO:0042090) regulation of interleukin-12 biosynthetic process(GO:0045075) |
| 0.2 | 2.4 | GO:1901621 | negative regulation of smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:1901621) |
| 0.2 | 0.4 | GO:0003273 | cell migration involved in endocardial cushion formation(GO:0003273) |
| 0.2 | 0.7 | GO:0000105 | histidine biosynthetic process(GO:0000105) |
| 0.2 | 2.2 | GO:0035988 | chondrocyte proliferation(GO:0035988) |
| 0.2 | 1.4 | GO:0061143 | alveolar primary septum development(GO:0061143) |
| 0.2 | 2.2 | GO:0090161 | Golgi ribbon formation(GO:0090161) |
| 0.2 | 2.9 | GO:0033148 | positive regulation of intracellular estrogen receptor signaling pathway(GO:0033148) |
| 0.2 | 2.5 | GO:0030220 | platelet formation(GO:0030220) |
| 0.2 | 3.7 | GO:0097094 | craniofacial suture morphogenesis(GO:0097094) |
| 0.2 | 9.1 | GO:0034605 | cellular response to heat(GO:0034605) |
| 0.2 | 2.3 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.2 | 2.6 | GO:0051823 | regulation of synapse structural plasticity(GO:0051823) |
| 0.2 | 1.8 | GO:0046007 | negative regulation of activated T cell proliferation(GO:0046007) |
| 0.2 | 3.0 | GO:0071260 | cellular response to mechanical stimulus(GO:0071260) |
| 0.2 | 1.0 | GO:0046826 | negative regulation of protein export from nucleus(GO:0046826) |
| 0.2 | 4.0 | GO:1900271 | regulation of long-term synaptic potentiation(GO:1900271) |
| 0.2 | 1.0 | GO:2000680 | regulation of rubidium ion transport(GO:2000680) |
| 0.2 | 3.4 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 0.2 | 5.3 | GO:0042220 | response to cocaine(GO:0042220) |
| 0.2 | 0.5 | GO:0006432 | phenylalanyl-tRNA aminoacylation(GO:0006432) |
| 0.2 | 4.2 | GO:1901739 | regulation of myoblast fusion(GO:1901739) positive regulation of myoblast fusion(GO:1901741) |
| 0.2 | 3.4 | GO:0070229 | negative regulation of lymphocyte apoptotic process(GO:0070229) |
| 0.2 | 2.2 | GO:0022010 | central nervous system myelination(GO:0022010) axon ensheathment in central nervous system(GO:0032291) |
| 0.2 | 2.9 | GO:0000291 | nuclear-transcribed mRNA catabolic process, exonucleolytic(GO:0000291) |
| 0.2 | 0.9 | GO:0090168 | Golgi reassembly(GO:0090168) |
| 0.2 | 2.2 | GO:0010763 | positive regulation of fibroblast migration(GO:0010763) |
| 0.2 | 1.4 | GO:0018344 | protein geranylgeranylation(GO:0018344) |
| 0.2 | 0.8 | GO:0050916 | sensory perception of sweet taste(GO:0050916) |
| 0.2 | 4.0 | GO:0006346 | methylation-dependent chromatin silencing(GO:0006346) |
| 0.2 | 3.0 | GO:0034453 | microtubule anchoring(GO:0034453) |
| 0.1 | 0.6 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.1 | 3.4 | GO:2000401 | regulation of lymphocyte migration(GO:2000401) |
| 0.1 | 2.2 | GO:0035855 | megakaryocyte development(GO:0035855) |
| 0.1 | 2.8 | GO:0014049 | positive regulation of glutamate secretion(GO:0014049) |
| 0.1 | 0.6 | GO:0044313 | protein K6-linked deubiquitination(GO:0044313) |
| 0.1 | 1.0 | GO:0072718 | response to cisplatin(GO:0072718) |
| 0.1 | 1.1 | GO:1902101 | positive regulation of mitotic metaphase/anaphase transition(GO:0045842) positive regulation of mitotic sister chromatid separation(GO:1901970) positive regulation of metaphase/anaphase transition of cell cycle(GO:1902101) |
| 0.1 | 2.5 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.1 | 3.3 | GO:0035825 | reciprocal meiotic recombination(GO:0007131) reciprocal DNA recombination(GO:0035825) |
| 0.1 | 6.7 | GO:0045104 | intermediate filament cytoskeleton organization(GO:0045104) |
| 0.1 | 1.5 | GO:1900383 | regulation of synaptic plasticity by receptor localization to synapse(GO:1900383) |
| 0.1 | 6.5 | GO:0001541 | ovarian follicle development(GO:0001541) |
| 0.1 | 1.0 | GO:0061343 | cell adhesion involved in heart morphogenesis(GO:0061343) |
| 0.1 | 0.5 | GO:0010159 | specification of organ position(GO:0010159) |
| 0.1 | 3.9 | GO:0006349 | regulation of gene expression by genetic imprinting(GO:0006349) |
| 0.1 | 3.9 | GO:0000460 | maturation of 5.8S rRNA(GO:0000460) |
| 0.1 | 2.1 | GO:0033753 | ribosomal subunit export from nucleus(GO:0000054) ribosome localization(GO:0033750) establishment of ribosome localization(GO:0033753) rRNA-containing ribonucleoprotein complex export from nucleus(GO:0071428) |
| 0.1 | 1.3 | GO:0032261 | purine nucleotide salvage(GO:0032261) IMP salvage(GO:0032264) |
| 0.1 | 0.4 | GO:0060809 | CD8-positive, alpha-beta T cell differentiation involved in immune response(GO:0002302) mesodermal to mesenchymal transition involved in gastrulation(GO:0060809) |
| 0.1 | 2.4 | GO:0042659 | regulation of cell fate specification(GO:0042659) |
| 0.1 | 1.3 | GO:2000144 | positive regulation of DNA-templated transcription, initiation(GO:2000144) |
| 0.1 | 0.8 | GO:0033140 | negative regulation of peptidyl-serine phosphorylation of STAT protein(GO:0033140) |
| 0.1 | 0.1 | GO:0034139 | regulation of toll-like receptor 3 signaling pathway(GO:0034139) |
| 0.1 | 8.0 | GO:0006367 | transcription initiation from RNA polymerase II promoter(GO:0006367) |
| 0.1 | 0.6 | GO:0001783 | B cell apoptotic process(GO:0001783) |
| 0.1 | 1.1 | GO:0015074 | DNA integration(GO:0015074) |
| 0.1 | 3.0 | GO:0009303 | rRNA transcription(GO:0009303) |
| 0.1 | 0.3 | GO:0010957 | negative regulation of vitamin D biosynthetic process(GO:0010957) |
| 0.1 | 0.4 | GO:2000742 | anterior head development(GO:0097065) regulation of anterior head development(GO:2000742) positive regulation of anterior head development(GO:2000744) |
| 0.1 | 4.4 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.1 | 13.5 | GO:0051028 | mRNA transport(GO:0051028) |
| 0.1 | 2.3 | GO:0034497 | protein localization to pre-autophagosomal structure(GO:0034497) |
| 0.1 | 1.7 | GO:0071578 | zinc II ion transmembrane import(GO:0071578) |
| 0.1 | 58.5 | GO:0007283 | spermatogenesis(GO:0007283) |
| 0.1 | 1.6 | GO:0033008 | positive regulation of mast cell activation involved in immune response(GO:0033008) positive regulation of mast cell degranulation(GO:0043306) |
| 0.1 | 1.0 | GO:0090190 | positive regulation of mesonephros development(GO:0061213) regulation of mesonephros development(GO:0061217) regulation of branching involved in ureteric bud morphogenesis(GO:0090189) positive regulation of branching involved in ureteric bud morphogenesis(GO:0090190) |
| 0.1 | 1.3 | GO:0046477 | glycosylceramide catabolic process(GO:0046477) |
| 0.1 | 2.0 | GO:0007194 | negative regulation of adenylate cyclase activity(GO:0007194) negative regulation of cyclase activity(GO:0031280) |
| 0.1 | 1.8 | GO:1990090 | response to nerve growth factor(GO:1990089) cellular response to nerve growth factor stimulus(GO:1990090) |
| 0.1 | 1.7 | GO:0006353 | DNA-templated transcription, termination(GO:0006353) |
| 0.1 | 12.9 | GO:0006261 | DNA-dependent DNA replication(GO:0006261) |
| 0.1 | 2.4 | GO:0010824 | regulation of centrosome duplication(GO:0010824) |
| 0.1 | 0.8 | GO:1900246 | positive regulation of RIG-I signaling pathway(GO:1900246) |
| 0.1 | 2.6 | GO:0002026 | regulation of the force of heart contraction(GO:0002026) |
| 0.1 | 1.3 | GO:0070389 | chaperone cofactor-dependent protein refolding(GO:0070389) |
| 0.1 | 1.5 | GO:0043248 | proteasome assembly(GO:0043248) |
| 0.1 | 0.6 | GO:0018094 | protein polyglycylation(GO:0018094) |
| 0.1 | 4.5 | GO:0045071 | negative regulation of viral genome replication(GO:0045071) |
| 0.1 | 3.1 | GO:0000245 | spliceosomal complex assembly(GO:0000245) |
| 0.1 | 1.3 | GO:0050957 | equilibrioception(GO:0050957) |
| 0.1 | 0.4 | GO:0002430 | complement receptor mediated signaling pathway(GO:0002430) |
| 0.1 | 1.0 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.1 | 4.0 | GO:0001578 | microtubule bundle formation(GO:0001578) |
| 0.1 | 0.9 | GO:0070914 | UV-damage excision repair(GO:0070914) |
| 0.1 | 2.8 | GO:1904031 | positive regulation of cyclin-dependent protein kinase activity(GO:1904031) |
| 0.1 | 1.2 | GO:0072675 | osteoclast fusion(GO:0072675) |
| 0.1 | 0.7 | GO:0045585 | regulation of cytotoxic T cell differentiation(GO:0045583) positive regulation of cytotoxic T cell differentiation(GO:0045585) |
| 0.1 | 0.3 | GO:0001543 | ovarian follicle rupture(GO:0001543) |
| 0.1 | 0.7 | GO:0043247 | telomere maintenance in response to DNA damage(GO:0043247) |
| 0.1 | 1.5 | GO:0007019 | microtubule depolymerization(GO:0007019) |
| 0.1 | 1.3 | GO:0071539 | protein localization to centrosome(GO:0071539) |
| 0.1 | 0.2 | GO:0010593 | negative regulation of lamellipodium assembly(GO:0010593) |
| 0.1 | 1.0 | GO:0031290 | retinal ganglion cell axon guidance(GO:0031290) |
| 0.1 | 2.0 | GO:0009299 | mRNA transcription(GO:0009299) |
| 0.1 | 5.3 | GO:0051865 | protein autoubiquitination(GO:0051865) |
| 0.1 | 0.8 | GO:0032328 | alanine transport(GO:0032328) |
| 0.1 | 5.2 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.1 | 1.7 | GO:1903427 | negative regulation of reactive oxygen species biosynthetic process(GO:1903427) |
| 0.1 | 0.8 | GO:0048934 | peripheral nervous system neuron differentiation(GO:0048934) peripheral nervous system neuron development(GO:0048935) |
| 0.1 | 1.7 | GO:0043982 | histone H4-K5 acetylation(GO:0043981) histone H4-K8 acetylation(GO:0043982) |
| 0.1 | 0.7 | GO:1900113 | negative regulation of histone H3-K9 trimethylation(GO:1900113) |
| 0.1 | 5.0 | GO:0042100 | B cell proliferation(GO:0042100) |
| 0.1 | 1.6 | GO:0006054 | N-acetylneuraminate metabolic process(GO:0006054) |
| 0.1 | 1.2 | GO:0048671 | negative regulation of collateral sprouting(GO:0048671) |
| 0.1 | 1.3 | GO:1902043 | positive regulation of extrinsic apoptotic signaling pathway via death domain receptors(GO:1902043) |
| 0.1 | 1.5 | GO:0034389 | lipid particle organization(GO:0034389) |
| 0.1 | 0.3 | GO:2000395 | positive regulation of viral budding via host ESCRT complex(GO:1903774) regulation of ubiquitin-dependent endocytosis(GO:2000395) positive regulation of ubiquitin-dependent endocytosis(GO:2000397) |
| 0.1 | 2.1 | GO:0008333 | endosome to lysosome transport(GO:0008333) |
| 0.1 | 1.4 | GO:0032469 | endoplasmic reticulum calcium ion homeostasis(GO:0032469) |
| 0.1 | 0.2 | GO:0042494 | detection of bacterial lipoprotein(GO:0042494) |
| 0.1 | 12.5 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.1 | 0.8 | GO:0070050 | neuron cellular homeostasis(GO:0070050) |
| 0.1 | 0.4 | GO:1900119 | positive regulation of execution phase of apoptosis(GO:1900119) |
| 0.1 | 0.2 | GO:0072176 | nephric duct development(GO:0072176) nephric duct morphogenesis(GO:0072178) |
| 0.1 | 10.0 | GO:0060271 | cilium morphogenesis(GO:0060271) |
| 0.1 | 1.2 | GO:0090083 | regulation of inclusion body assembly(GO:0090083) |
| 0.1 | 0.2 | GO:0045048 | protein insertion into ER membrane(GO:0045048) |
| 0.1 | 3.5 | GO:0042147 | retrograde transport, endosome to Golgi(GO:0042147) |
| 0.1 | 0.3 | GO:0043486 | histone exchange(GO:0043486) |
| 0.1 | 0.5 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
| 0.1 | 1.0 | GO:0061462 | protein localization to lysosome(GO:0061462) |
| 0.1 | 1.5 | GO:0050832 | defense response to fungus(GO:0050832) |
| 0.1 | 1.0 | GO:0016180 | snRNA metabolic process(GO:0016073) snRNA processing(GO:0016180) |
| 0.1 | 2.5 | GO:0006986 | response to unfolded protein(GO:0006986) |
| 0.1 | 1.8 | GO:0009409 | response to cold(GO:0009409) |
| 0.1 | 0.8 | GO:0015816 | glycine transport(GO:0015816) |
| 0.1 | 0.8 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.1 | 1.6 | GO:0032620 | interleukin-17 production(GO:0032620) |
| 0.1 | 1.4 | GO:0035025 | positive regulation of Rho protein signal transduction(GO:0035025) |
| 0.1 | 0.7 | GO:0003338 | metanephros morphogenesis(GO:0003338) |
| 0.1 | 1.5 | GO:1903146 | regulation of mitophagy(GO:1903146) |
| 0.1 | 2.3 | GO:0033344 | cholesterol efflux(GO:0033344) |
| 0.1 | 1.6 | GO:0040015 | negative regulation of multicellular organism growth(GO:0040015) |
| 0.1 | 1.2 | GO:0016226 | iron-sulfur cluster assembly(GO:0016226) metallo-sulfur cluster assembly(GO:0031163) |
| 0.1 | 11.4 | GO:0007018 | microtubule-based movement(GO:0007018) |
| 0.1 | 1.9 | GO:0016601 | Rac protein signal transduction(GO:0016601) |
| 0.1 | 1.3 | GO:0046856 | phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.1 | 0.9 | GO:0021895 | cerebral cortex neuron differentiation(GO:0021895) |
| 0.1 | 0.6 | GO:2000114 | regulation of establishment of cell polarity(GO:2000114) |
| 0.1 | 0.9 | GO:0060670 | branching involved in labyrinthine layer morphogenesis(GO:0060670) |
| 0.1 | 2.1 | GO:0032012 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.1 | 0.5 | GO:2000467 | positive regulation of glycogen (starch) synthase activity(GO:2000467) |
| 0.1 | 0.6 | GO:0043044 | ATP-dependent chromatin remodeling(GO:0043044) |
| 0.1 | 0.5 | GO:1990573 | potassium ion import across plasma membrane(GO:1990573) |
| 0.0 | 0.7 | GO:0048535 | lymph node development(GO:0048535) |
| 0.0 | 1.4 | GO:0061001 | regulation of dendritic spine morphogenesis(GO:0061001) |
| 0.0 | 1.5 | GO:0070265 | necrotic cell death(GO:0070265) |
| 0.0 | 0.5 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 0.0 | 0.4 | GO:0061669 | spontaneous neurotransmitter secretion(GO:0061669) spontaneous synaptic transmission(GO:0098814) |
| 0.0 | 1.0 | GO:0043153 | entrainment of circadian clock by photoperiod(GO:0043153) |
| 0.0 | 0.9 | GO:0075522 | IRES-dependent viral translational initiation(GO:0075522) |
| 0.0 | 0.8 | GO:0031572 | G2 DNA damage checkpoint(GO:0031572) |
| 0.0 | 0.3 | GO:0036066 | protein O-linked fucosylation(GO:0036066) |
| 0.0 | 0.6 | GO:0061014 | positive regulation of mRNA catabolic process(GO:0061014) |
| 0.0 | 0.3 | GO:0060017 | parathyroid gland development(GO:0060017) |
| 0.0 | 0.5 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 0.0 | 2.0 | GO:0071219 | cellular response to molecule of bacterial origin(GO:0071219) cellular response to lipopolysaccharide(GO:0071222) |
| 0.0 | 1.3 | GO:0070059 | intrinsic apoptotic signaling pathway in response to endoplasmic reticulum stress(GO:0070059) |
| 0.0 | 0.1 | GO:0023016 | signal transduction by trans-phosphorylation(GO:0023016) |
| 0.0 | 0.2 | GO:1900037 | regulation of cellular response to hypoxia(GO:1900037) |
| 0.0 | 0.3 | GO:0010766 | negative regulation of sodium ion transport(GO:0010766) |
| 0.0 | 0.8 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.0 | 0.3 | GO:0006705 | mineralocorticoid biosynthetic process(GO:0006705) |
| 0.0 | 0.9 | GO:0010507 | negative regulation of autophagy(GO:0010507) |
| 0.0 | 0.4 | GO:0090200 | positive regulation of release of cytochrome c from mitochondria(GO:0090200) |
| 0.0 | 0.3 | GO:2000311 | regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000311) |
| 0.0 | 0.7 | GO:0032355 | response to estradiol(GO:0032355) |
| 0.0 | 0.4 | GO:0060997 | dendritic spine morphogenesis(GO:0060997) |
| 0.0 | 0.9 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.0 | 0.9 | GO:0001958 | endochondral ossification(GO:0001958) replacement ossification(GO:0036075) |
| 0.0 | 1.0 | GO:0001938 | positive regulation of endothelial cell proliferation(GO:0001938) |
| 0.0 | 0.4 | GO:0007614 | short-term memory(GO:0007614) |
| 0.0 | 3.8 | GO:0050871 | positive regulation of B cell activation(GO:0050871) |
| 0.0 | 11.2 | GO:0010498 | proteasomal protein catabolic process(GO:0010498) |
| 0.0 | 0.4 | GO:0035162 | embryonic hemopoiesis(GO:0035162) |
| 0.0 | 0.2 | GO:0060340 | positive regulation of type I interferon-mediated signaling pathway(GO:0060340) |
| 0.0 | 1.5 | GO:0002224 | toll-like receptor signaling pathway(GO:0002224) |
| 0.0 | 0.1 | GO:0010747 | positive regulation of plasma membrane long-chain fatty acid transport(GO:0010747) |
| 0.0 | 0.1 | GO:0060040 | retinal bipolar neuron differentiation(GO:0060040) |
| 0.0 | 1.2 | GO:1903749 | positive regulation of establishment of protein localization to mitochondrion(GO:1903749) |
| 0.0 | 0.1 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 0.0 | 0.1 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.0 | 1.4 | GO:0030168 | platelet activation(GO:0030168) |
| 0.0 | 0.4 | GO:0060384 | innervation(GO:0060384) |
| 0.0 | 0.7 | GO:0032007 | negative regulation of TOR signaling(GO:0032007) |
| 0.0 | 0.8 | GO:0043966 | histone H3 acetylation(GO:0043966) |
| 0.0 | 0.5 | GO:0085029 | extracellular matrix assembly(GO:0085029) |
| 0.0 | 0.2 | GO:1904659 | glucose transmembrane transport(GO:1904659) |
| 0.0 | 0.3 | GO:0006301 | postreplication repair(GO:0006301) |
| 0.0 | 0.4 | GO:0043666 | regulation of phosphoprotein phosphatase activity(GO:0043666) |
| 0.0 | 0.3 | GO:0038092 | nodal signaling pathway(GO:0038092) |
| 0.0 | 0.1 | GO:0035641 | locomotory exploration behavior(GO:0035641) |
| 0.0 | 0.1 | GO:0031119 | tRNA pseudouridine synthesis(GO:0031119) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.8 | 18.9 | GO:0044393 | microspike(GO:0044393) |
| 3.5 | 14.1 | GO:0008623 | CHRAC(GO:0008623) |
| 2.5 | 15.2 | GO:1990356 | sumoylated E2 ligase complex(GO:1990356) |
| 2.4 | 14.2 | GO:0042585 | germinal vesicle(GO:0042585) |
| 2.3 | 2.3 | GO:0035101 | FACT complex(GO:0035101) |
| 2.3 | 13.6 | GO:0044305 | calyx of Held(GO:0044305) |
| 2.1 | 12.6 | GO:0032777 | Piccolo NuA4 histone acetyltransferase complex(GO:0032777) |
| 2.1 | 6.3 | GO:0005673 | transcription factor TFIIE complex(GO:0005673) |
| 2.1 | 12.5 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 2.0 | 5.9 | GO:0060187 | cell pole(GO:0060187) |
| 1.5 | 13.5 | GO:0097165 | nuclear stress granule(GO:0097165) |
| 1.5 | 26.8 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 1.5 | 10.3 | GO:1990584 | cardiac Troponin complex(GO:1990584) |
| 1.4 | 17.8 | GO:0097427 | microtubule bundle(GO:0097427) |
| 1.3 | 7.6 | GO:0005971 | ribonucleoside-diphosphate reductase complex(GO:0005971) |
| 1.3 | 5.0 | GO:0097598 | sperm cytoplasmic droplet(GO:0097598) |
| 1.2 | 5.0 | GO:0000438 | core TFIIH complex portion of holo TFIIH complex(GO:0000438) |
| 1.2 | 6.1 | GO:0031251 | PAN complex(GO:0031251) |
| 1.1 | 12.6 | GO:0071598 | neuronal ribonucleoprotein granule(GO:0071598) |
| 1.1 | 3.3 | GO:0030905 | retromer, tubulation complex(GO:0030905) |
| 1.1 | 13.2 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 1.0 | 4.2 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 1.0 | 14.4 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 1.0 | 5.0 | GO:0044218 | other organism cell membrane(GO:0044218) other organism membrane(GO:0044279) |
| 1.0 | 4.8 | GO:1990131 | Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 1.0 | 11.4 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.9 | 9.1 | GO:0042629 | mast cell granule(GO:0042629) |
| 0.9 | 3.6 | GO:1902737 | dendritic filopodium(GO:1902737) |
| 0.9 | 8.0 | GO:0030485 | smooth muscle contractile fiber(GO:0030485) |
| 0.9 | 3.5 | GO:0016035 | zeta DNA polymerase complex(GO:0016035) |
| 0.8 | 2.5 | GO:0042272 | nuclear RNA export factor complex(GO:0042272) |
| 0.8 | 11.7 | GO:0001520 | outer dense fiber(GO:0001520) |
| 0.8 | 13.4 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.8 | 7.8 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 0.8 | 3.9 | GO:0097059 | CRLF-CLCF1 complex(GO:0097058) CNTFR-CLCF1 complex(GO:0097059) |
| 0.8 | 21.6 | GO:0030285 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.7 | 18.5 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.7 | 6.6 | GO:0097442 | CA3 pyramidal cell dendrite(GO:0097442) |
| 0.7 | 3.6 | GO:0035363 | histone locus body(GO:0035363) |
| 0.7 | 8.6 | GO:0034992 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.7 | 11.2 | GO:0042555 | MCM complex(GO:0042555) |
| 0.7 | 4.1 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 0.7 | 4.6 | GO:0070381 | endosome to plasma membrane transport vesicle(GO:0070381) |
| 0.6 | 1.9 | GO:0033193 | Lsd1/2 complex(GO:0033193) |
| 0.6 | 8.4 | GO:0036156 | inner dynein arm(GO:0036156) |
| 0.6 | 2.4 | GO:1990589 | ATF4-CREB1 transcription factor complex(GO:1990589) |
| 0.6 | 7.1 | GO:0044327 | dendritic spine head(GO:0044327) |
| 0.6 | 4.7 | GO:0000322 | storage vacuole(GO:0000322) |
| 0.6 | 8.0 | GO:0020005 | symbiont-containing vacuole membrane(GO:0020005) |
| 0.6 | 6.2 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.6 | 1.1 | GO:0031074 | nucleocytoplasmic shuttling complex(GO:0031074) |
| 0.6 | 1.7 | GO:0031933 | telomeric heterochromatin(GO:0031933) |
| 0.5 | 9.2 | GO:0001527 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.5 | 5.9 | GO:0031094 | platelet dense tubular network(GO:0031094) |
| 0.5 | 2.0 | GO:0071953 | elastic fiber(GO:0071953) |
| 0.5 | 6.6 | GO:0000137 | Golgi cis cisterna(GO:0000137) |
| 0.5 | 3.5 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.5 | 6.0 | GO:0098845 | postsynaptic endosome(GO:0098845) |
| 0.5 | 8.0 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.5 | 4.3 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.5 | 8.6 | GO:0090544 | BAF-type complex(GO:0090544) |
| 0.5 | 4.5 | GO:0002177 | manchette(GO:0002177) |
| 0.4 | 2.7 | GO:0005672 | transcription factor TFIIA complex(GO:0005672) |
| 0.4 | 13.4 | GO:0000786 | nucleosome(GO:0000786) |
| 0.4 | 3.9 | GO:0070545 | PeBoW complex(GO:0070545) |
| 0.4 | 1.3 | GO:0044294 | dendritic growth cone(GO:0044294) |
| 0.4 | 2.1 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 0.4 | 1.6 | GO:0072534 | perineuronal net(GO:0072534) |
| 0.4 | 1.6 | GO:0071149 | TEAD-2-YAP complex(GO:0071149) |
| 0.4 | 5.6 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.4 | 10.6 | GO:0097431 | mitotic spindle pole(GO:0097431) |
| 0.4 | 19.1 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.4 | 4.2 | GO:0000812 | Swr1 complex(GO:0000812) |
| 0.4 | 3.8 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
| 0.4 | 1.5 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.4 | 8.8 | GO:0031362 | anchored component of external side of plasma membrane(GO:0031362) |
| 0.4 | 5.1 | GO:0060091 | kinocilium(GO:0060091) |
| 0.4 | 3.6 | GO:0097451 | glial limiting end-foot(GO:0097451) |
| 0.4 | 6.0 | GO:0005639 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) |
| 0.3 | 3.1 | GO:0061689 | tricellular tight junction(GO:0061689) |
| 0.3 | 2.4 | GO:0097443 | sorting endosome(GO:0097443) |
| 0.3 | 1.0 | GO:0031904 | endosome lumen(GO:0031904) |
| 0.3 | 1.0 | GO:0070877 | microprocessor complex(GO:0070877) ribonuclease III complex(GO:1903095) |
| 0.3 | 4.3 | GO:0005642 | annulate lamellae(GO:0005642) |
| 0.3 | 4.0 | GO:1990907 | beta-catenin-TCF complex(GO:1990907) |
| 0.3 | 34.6 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.3 | 19.4 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.3 | 23.9 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.3 | 1.2 | GO:0005896 | interleukin-6 receptor complex(GO:0005896) |
| 0.3 | 18.2 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.3 | 2.4 | GO:0070775 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.3 | 14.5 | GO:0043596 | nuclear replication fork(GO:0043596) |
| 0.3 | 3.8 | GO:0016589 | NURF complex(GO:0016589) |
| 0.3 | 3.1 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.3 | 1.1 | GO:0043159 | acrosomal matrix(GO:0043159) |
| 0.3 | 8.8 | GO:0031527 | filopodium membrane(GO:0031527) |
| 0.3 | 2.0 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.3 | 1.7 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.3 | 2.2 | GO:0017071 | intracellular cyclic nucleotide activated cation channel complex(GO:0017071) |
| 0.3 | 9.7 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.3 | 4.1 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.3 | 7.5 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.3 | 2.3 | GO:0005786 | signal recognition particle, endoplasmic reticulum targeting(GO:0005786) |
| 0.3 | 2.6 | GO:0000408 | EKC/KEOPS complex(GO:0000408) |
| 0.3 | 4.9 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.3 | 2.8 | GO:0097542 | ciliary tip(GO:0097542) |
| 0.2 | 2.5 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.2 | 17.8 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.2 | 4.5 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.2 | 1.4 | GO:0005968 | Rab-protein geranylgeranyltransferase complex(GO:0005968) |
| 0.2 | 7.8 | GO:0005657 | replication fork(GO:0005657) |
| 0.2 | 4.1 | GO:0036056 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.2 | 4.8 | GO:0071004 | U2-type prespliceosome(GO:0071004) |
| 0.2 | 9.7 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.2 | 3.1 | GO:0000782 | telomere cap complex(GO:0000782) nuclear telomere cap complex(GO:0000783) |
| 0.2 | 1.3 | GO:0034709 | methylosome(GO:0034709) |
| 0.2 | 6.9 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 0.2 | 3.2 | GO:0097539 | ciliary transition fiber(GO:0097539) |
| 0.2 | 1.0 | GO:0044614 | nuclear pore cytoplasmic filaments(GO:0044614) |
| 0.2 | 2.3 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.2 | 4.4 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.2 | 0.6 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.2 | 1.8 | GO:0072379 | BAT3 complex(GO:0071818) ER membrane insertion complex(GO:0072379) |
| 0.2 | 1.2 | GO:0097452 | GAIT complex(GO:0097452) |
| 0.2 | 1.6 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.2 | 5.1 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.2 | 11.6 | GO:0005930 | axoneme(GO:0005930) |
| 0.2 | 4.4 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.2 | 1.8 | GO:0032391 | photoreceptor connecting cilium(GO:0032391) |
| 0.2 | 6.1 | GO:0005771 | multivesicular body(GO:0005771) |
| 0.2 | 2.3 | GO:0097136 | Bcl-2 family protein complex(GO:0097136) |
| 0.2 | 7.5 | GO:1990391 | DNA repair complex(GO:1990391) |
| 0.2 | 1.6 | GO:0005687 | U4 snRNP(GO:0005687) |
| 0.2 | 2.1 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 0.2 | 2.5 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.2 | 8.1 | GO:0005865 | striated muscle thin filament(GO:0005865) |
| 0.2 | 1.3 | GO:0005685 | U1 snRNP(GO:0005685) |
| 0.2 | 0.5 | GO:0009328 | phenylalanine-tRNA ligase complex(GO:0009328) |
| 0.2 | 1.3 | GO:0001652 | granular component(GO:0001652) |
| 0.2 | 4.5 | GO:0035686 | sperm fibrous sheath(GO:0035686) |
| 0.2 | 2.4 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.2 | 3.3 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.2 | 28.6 | GO:0005884 | actin filament(GO:0005884) |
| 0.2 | 1.4 | GO:0030130 | clathrin coat of trans-Golgi network vesicle(GO:0030130) |
| 0.2 | 3.5 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.1 | 5.1 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.1 | 0.6 | GO:1990696 | stereocilium membrane(GO:0060171) periciliary membrane compartment(GO:1990075) USH2 complex(GO:1990696) |
| 0.1 | 2.5 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.1 | 15.4 | GO:0005814 | centriole(GO:0005814) |
| 0.1 | 1.3 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.1 | 1.2 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.1 | 25.6 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.1 | 0.9 | GO:0031465 | Cul4B-RING E3 ubiquitin ligase complex(GO:0031465) |
| 0.1 | 4.0 | GO:0071782 | endoplasmic reticulum tubular network(GO:0071782) |
| 0.1 | 39.3 | GO:0005874 | microtubule(GO:0005874) |
| 0.1 | 0.4 | GO:0043512 | inhibin A complex(GO:0043512) |
| 0.1 | 6.8 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.1 | 10.8 | GO:0005875 | microtubule associated complex(GO:0005875) |
| 0.1 | 1.3 | GO:0000127 | transcription factor TFIIIC complex(GO:0000127) |
| 0.1 | 58.2 | GO:0005813 | centrosome(GO:0005813) |
| 0.1 | 1.1 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.1 | 10.6 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.1 | 1.5 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 0.1 | 0.2 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.1 | 0.7 | GO:0097226 | sperm mitochondrial sheath(GO:0097226) |
| 0.1 | 1.5 | GO:0043189 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.1 | 1.6 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 0.1 | 1.5 | GO:0000177 | cytoplasmic exosome (RNase complex)(GO:0000177) |
| 0.1 | 5.4 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.1 | 5.2 | GO:0035097 | histone methyltransferase complex(GO:0035097) |
| 0.1 | 6.8 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.1 | 6.8 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.1 | 1.2 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.1 | 1.8 | GO:0034704 | calcium channel complex(GO:0034704) |
| 0.1 | 2.3 | GO:0005844 | polysome(GO:0005844) |
| 0.1 | 0.2 | GO:0030289 | protein phosphatase 4 complex(GO:0030289) |
| 0.1 | 5.2 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.1 | 9.0 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) |
| 0.1 | 2.1 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.1 | 0.8 | GO:0070552 | BRISC complex(GO:0070552) |
| 0.1 | 8.1 | GO:0000151 | ubiquitin ligase complex(GO:0000151) |
| 0.1 | 9.9 | GO:0030139 | endocytic vesicle(GO:0030139) |
| 0.1 | 10.2 | GO:0055037 | recycling endosome(GO:0055037) |
| 0.1 | 4.3 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.1 | 2.7 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.1 | 8.8 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.1 | 3.1 | GO:0030904 | retromer complex(GO:0030904) |
| 0.1 | 0.9 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.1 | 5.9 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.1 | 1.0 | GO:0051233 | spindle midzone(GO:0051233) |
| 0.1 | 45.2 | GO:0048471 | perinuclear region of cytoplasm(GO:0048471) |
| 0.1 | 0.6 | GO:0071437 | invadopodium(GO:0071437) |
| 0.1 | 0.2 | GO:0005677 | chromatin silencing complex(GO:0005677) |
| 0.1 | 1.0 | GO:0046540 | U4/U6 x U5 tri-snRNP complex(GO:0046540) |
| 0.1 | 1.2 | GO:0002102 | podosome(GO:0002102) |
| 0.1 | 11.9 | GO:0015629 | actin cytoskeleton(GO:0015629) |
| 0.1 | 15.3 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.1 | 2.0 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.1 | 2.8 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.1 | 4.7 | GO:0032993 | protein-DNA complex(GO:0032993) |
| 0.1 | 0.3 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.1 | 0.7 | GO:0005790 | smooth endoplasmic reticulum(GO:0005790) |
| 0.1 | 2.5 | GO:0016592 | mediator complex(GO:0016592) |
| 0.1 | 11.1 | GO:0019867 | outer membrane(GO:0019867) organelle outer membrane(GO:0031968) |
| 0.1 | 1.4 | GO:0030175 | filopodium(GO:0030175) |
| 0.1 | 0.5 | GO:0005927 | muscle tendon junction(GO:0005927) |
| 0.1 | 2.0 | GO:0031985 | Golgi cisterna(GO:0031985) |
| 0.1 | 6.0 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.1 | 6.3 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.1 | 0.2 | GO:0097169 | AIM2 inflammasome complex(GO:0097169) |
| 0.1 | 1.8 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.0 | 1.1 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.0 | 2.4 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.0 | 4.8 | GO:0001669 | acrosomal vesicle(GO:0001669) |
| 0.0 | 2.9 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.0 | 39.3 | GO:0005730 | nucleolus(GO:0005730) |
| 0.0 | 0.8 | GO:0019897 | extrinsic component of plasma membrane(GO:0019897) |
| 0.0 | 3.5 | GO:0034702 | ion channel complex(GO:0034702) |
| 0.0 | 0.1 | GO:0048269 | methionine adenosyltransferase complex(GO:0048269) |
| 0.0 | 0.4 | GO:0032426 | stereocilium tip(GO:0032426) |
| 0.0 | 2.7 | GO:0097517 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.0 | 1.1 | GO:0005720 | nuclear heterochromatin(GO:0005720) |
| 0.0 | 0.6 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 2.3 | GO:0005681 | spliceosomal complex(GO:0005681) |
| 0.0 | 238.3 | GO:0005634 | nucleus(GO:0005634) |
| 0.0 | 3.8 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.0 | 0.4 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.0 | 1.7 | GO:0010008 | endosome membrane(GO:0010008) |
| 0.0 | 5.3 | GO:0005856 | cytoskeleton(GO:0005856) |
| 0.0 | 0.4 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.3 | 13.2 | GO:0038025 | reelin receptor activity(GO:0038025) |
| 2.7 | 26.6 | GO:0035374 | chondroitin sulfate binding(GO:0035374) |
| 2.5 | 15.2 | GO:0061656 | SUMO conjugating enzyme activity(GO:0061656) |
| 2.4 | 9.5 | GO:0036313 | phosphatidylinositol 3-kinase catalytic subunit binding(GO:0036313) |
| 2.3 | 7.0 | GO:0032190 | acrosin binding(GO:0032190) |
| 2.3 | 15.8 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 2.2 | 6.7 | GO:0050510 | N-acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase activity(GO:0050510) |
| 2.1 | 8.4 | GO:0031692 | alpha-1A adrenergic receptor binding(GO:0031691) alpha-1B adrenergic receptor binding(GO:0031692) follicle-stimulating hormone receptor binding(GO:0031762) |
| 1.9 | 3.9 | GO:0034046 | poly(G) binding(GO:0034046) |
| 1.9 | 13.3 | GO:1990254 | keratin filament binding(GO:1990254) |
| 1.9 | 9.4 | GO:0071987 | WD40-repeat domain binding(GO:0071987) |
| 1.6 | 6.5 | GO:0004105 | choline-phosphate cytidylyltransferase activity(GO:0004105) |
| 1.6 | 24.2 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 1.5 | 10.3 | GO:0030172 | troponin C binding(GO:0030172) troponin I binding(GO:0031013) |
| 1.4 | 5.8 | GO:0030023 | extracellular matrix constituent conferring elasticity(GO:0030023) |
| 1.4 | 8.4 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 1.4 | 8.4 | GO:0004832 | valine-tRNA ligase activity(GO:0004832) |
| 1.4 | 4.1 | GO:0052895 | norspermine:oxygen oxidoreductase activity(GO:0052894) N1-acetylspermine:oxygen oxidoreductase (N1-acetylspermidine-forming) activity(GO:0052895) |
| 1.4 | 23.3 | GO:0005522 | profilin binding(GO:0005522) |
| 1.3 | 8.0 | GO:0032896 | palmitoyl-CoA 9-desaturase activity(GO:0032896) |
| 1.3 | 5.3 | GO:0030156 | benzodiazepine receptor binding(GO:0030156) |
| 1.3 | 5.3 | GO:0072591 | citrate-L-glutamate ligase activity(GO:0072591) |
| 1.3 | 3.9 | GO:0097677 | STAT family protein binding(GO:0097677) |
| 1.3 | 7.6 | GO:0004748 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 1.2 | 8.6 | GO:0030284 | estrogen receptor activity(GO:0030284) |
| 1.2 | 3.6 | GO:0004911 | interleukin-2 receptor activity(GO:0004911) |
| 1.2 | 8.4 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 1.2 | 8.1 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 1.1 | 4.5 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
| 1.1 | 7.7 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 1.0 | 5.2 | GO:0042610 | CD8 receptor binding(GO:0042610) |
| 1.0 | 11.3 | GO:0036042 | long-chain fatty acyl-CoA binding(GO:0036042) |
| 1.0 | 22.1 | GO:0042288 | MHC class I protein binding(GO:0042288) |
| 0.9 | 5.5 | GO:0000099 | sulfur amino acid transmembrane transporter activity(GO:0000099) |
| 0.9 | 6.3 | GO:0047184 | 1-acylglycerophosphocholine O-acyltransferase activity(GO:0047184) |
| 0.9 | 32.7 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.9 | 5.2 | GO:0045322 | unmethylated CpG binding(GO:0045322) |
| 0.9 | 2.6 | GO:0061711 | N(6)-L-threonylcarbamoyladenine synthase(GO:0061711) |
| 0.8 | 5.1 | GO:0017077 | oxidative phosphorylation uncoupler activity(GO:0017077) |
| 0.8 | 8.3 | GO:0008889 | glycerophosphodiester phosphodiesterase activity(GO:0008889) |
| 0.8 | 3.3 | GO:1990460 | leptin receptor binding(GO:1990460) |
| 0.8 | 1.7 | GO:0032129 | histone deacetylase activity (H3-K9 specific)(GO:0032129) NAD-dependent histone deacetylase activity (H3-K9 specific)(GO:0046969) |
| 0.8 | 4.9 | GO:0015186 | L-glutamine transmembrane transporter activity(GO:0015186) |
| 0.8 | 2.5 | GO:0030226 | apolipoprotein receptor activity(GO:0030226) apolipoprotein A-I receptor activity(GO:0034188) phosphatidylserine-translocating ATPase activity(GO:0090556) |
| 0.8 | 8.2 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.8 | 3.2 | GO:0031694 | alpha-2A adrenergic receptor binding(GO:0031694) |
| 0.8 | 8.6 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.8 | 9.4 | GO:0032051 | clathrin light chain binding(GO:0032051) |
| 0.8 | 3.1 | GO:0010521 | telomerase inhibitor activity(GO:0010521) |
| 0.8 | 2.3 | GO:0019912 | cyclin-dependent protein kinase activating kinase activity(GO:0019912) |
| 0.8 | 2.3 | GO:0045142 | ATP-dependent DNA/RNA helicase activity(GO:0033680) triplex DNA binding(GO:0045142) |
| 0.8 | 2.3 | GO:0004461 | lactose synthase activity(GO:0004461) |
| 0.8 | 8.3 | GO:0048101 | calmodulin-dependent cyclic-nucleotide phosphodiesterase activity(GO:0004117) calcium- and calmodulin-regulated 3',5'-cyclic-GMP phosphodiesterase activity(GO:0048101) |
| 0.7 | 5.2 | GO:0004769 | steroid delta-isomerase activity(GO:0004769) |
| 0.7 | 5.9 | GO:0005166 | neurotrophin p75 receptor binding(GO:0005166) |
| 0.7 | 15.4 | GO:0015271 | outward rectifier potassium channel activity(GO:0015271) |
| 0.7 | 6.6 | GO:0005519 | cytoskeletal regulatory protein binding(GO:0005519) |
| 0.7 | 36.3 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.7 | 6.5 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.7 | 2.8 | GO:0004052 | arachidonate 12-lipoxygenase activity(GO:0004052) |
| 0.7 | 3.5 | GO:0034648 | histone demethylase activity (H3-dimethyl-K4 specific)(GO:0034648) |
| 0.7 | 20.7 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.7 | 43.7 | GO:0036002 | pre-mRNA binding(GO:0036002) |
| 0.7 | 6.0 | GO:0035381 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.7 | 8.5 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.6 | 5.7 | GO:0035033 | histone deacetylase regulator activity(GO:0035033) |
| 0.6 | 3.2 | GO:0034481 | chondroitin sulfotransferase activity(GO:0034481) |
| 0.6 | 5.6 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.6 | 6.8 | GO:0005168 | neurotrophin TRKA receptor binding(GO:0005168) |
| 0.6 | 4.3 | GO:0005087 | Ran guanyl-nucleotide exchange factor activity(GO:0005087) |
| 0.6 | 2.4 | GO:1990763 | arrestin family protein binding(GO:1990763) |
| 0.6 | 8.3 | GO:0005225 | volume-sensitive anion channel activity(GO:0005225) |
| 0.6 | 21.3 | GO:0008187 | poly-pyrimidine tract binding(GO:0008187) |
| 0.6 | 2.3 | GO:0070051 | fibrinogen binding(GO:0070051) |
| 0.6 | 2.3 | GO:0030519 | snoRNP binding(GO:0030519) |
| 0.6 | 3.4 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.6 | 3.4 | GO:0016300 | tRNA (uracil) methyltransferase activity(GO:0016300) |
| 0.6 | 4.5 | GO:0070699 | type II activin receptor binding(GO:0070699) |
| 0.6 | 8.4 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.6 | 1.1 | GO:0070883 | pre-miRNA binding(GO:0070883) |
| 0.6 | 7.7 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 0.5 | 4.9 | GO:0004716 | receptor signaling protein tyrosine kinase activity(GO:0004716) |
| 0.5 | 10.4 | GO:0016493 | C-C chemokine receptor activity(GO:0016493) |
| 0.5 | 3.2 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 0.5 | 25.9 | GO:0051959 | dynein light intermediate chain binding(GO:0051959) |
| 0.5 | 12.6 | GO:0004862 | cAMP-dependent protein kinase inhibitor activity(GO:0004862) |
| 0.5 | 6.3 | GO:0005229 | intracellular calcium activated chloride channel activity(GO:0005229) |
| 0.5 | 9.3 | GO:0043495 | protein anchor(GO:0043495) |
| 0.5 | 2.1 | GO:0005415 | nucleoside:sodium symporter activity(GO:0005415) |
| 0.5 | 18.0 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.5 | 14.5 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.5 | 2.5 | GO:0004735 | pyrroline-5-carboxylate reductase activity(GO:0004735) |
| 0.5 | 8.2 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.5 | 3.4 | GO:0047276 | N-acetyllactosaminide 3-alpha-galactosyltransferase activity(GO:0047276) |
| 0.5 | 1.9 | GO:0070052 | collagen V binding(GO:0070052) |
| 0.5 | 5.2 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.5 | 2.4 | GO:0005280 | hydrogen:amino acid symporter activity(GO:0005280) |
| 0.5 | 8.9 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.5 | 1.9 | GO:0003883 | CTP synthase activity(GO:0003883) |
| 0.5 | 17.7 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.5 | 11.1 | GO:0055103 | ligase regulator activity(GO:0055103) |
| 0.5 | 1.4 | GO:0000402 | open form four-way junction DNA binding(GO:0000401) crossed form four-way junction DNA binding(GO:0000402) |
| 0.5 | 6.4 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.5 | 2.3 | GO:0052795 | exo-alpha-sialidase activity(GO:0004308) alpha-sialidase activity(GO:0016997) exo-alpha-(2->3)-sialidase activity(GO:0052794) exo-alpha-(2->6)-sialidase activity(GO:0052795) exo-alpha-(2->8)-sialidase activity(GO:0052796) |
| 0.4 | 4.4 | GO:1990189 | peptide-serine-N-acetyltransferase activity(GO:1990189) |
| 0.4 | 7.1 | GO:0005172 | vascular endothelial growth factor receptor binding(GO:0005172) |
| 0.4 | 8.8 | GO:0030296 | protein tyrosine kinase activator activity(GO:0030296) |
| 0.4 | 2.6 | GO:1990269 | RNA polymerase II C-terminal domain phosphoserine binding(GO:1990269) |
| 0.4 | 2.2 | GO:0004305 | ethanolamine kinase activity(GO:0004305) |
| 0.4 | 33.2 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.4 | 11.3 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.4 | 35.7 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.4 | 1.3 | GO:0001034 | RNA polymerase III transcription factor activity, sequence-specific DNA binding(GO:0001034) |
| 0.4 | 1.3 | GO:0035851 | Krueppel-associated box domain binding(GO:0035851) |
| 0.4 | 3.0 | GO:0031821 | G-protein coupled serotonin receptor binding(GO:0031821) |
| 0.4 | 5.0 | GO:0052813 | phosphatidylinositol-4,5-bisphosphate 3-kinase activity(GO:0046934) phosphatidylinositol bisphosphate kinase activity(GO:0052813) |
| 0.4 | 10.7 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.4 | 2.1 | GO:0003872 | 6-phosphofructokinase activity(GO:0003872) |
| 0.4 | 2.5 | GO:0033192 | calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.4 | 8.8 | GO:0032041 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.4 | 22.0 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.4 | 1.2 | GO:0001160 | transcription termination site sequence-specific DNA binding(GO:0001147) transcription termination site DNA binding(GO:0001160) |
| 0.4 | 2.3 | GO:0030942 | endoplasmic reticulum signal peptide binding(GO:0030942) |
| 0.4 | 8.5 | GO:0032794 | GTPase activating protein binding(GO:0032794) |
| 0.4 | 0.8 | GO:0052724 | inositol hexakisphosphate kinase activity(GO:0000828) inositol hexakisphosphate 5-kinase activity(GO:0000832) inositol hexakisphosphate 1-kinase activity(GO:0052723) inositol hexakisphosphate 3-kinase activity(GO:0052724) |
| 0.4 | 20.0 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) |
| 0.4 | 2.2 | GO:0005052 | peroxisome matrix targeting signal-1 binding(GO:0005052) |
| 0.4 | 3.7 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.4 | 2.2 | GO:0051734 | ATP-dependent polydeoxyribonucleotide 5'-hydroxyl-kinase activity(GO:0046404) polydeoxyribonucleotide kinase activity(GO:0051733) ATP-dependent polynucleotide kinase activity(GO:0051734) |
| 0.4 | 2.2 | GO:0050656 | 3'-phosphoadenosine 5'-phosphosulfate binding(GO:0050656) |
| 0.4 | 4.0 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 0.4 | 11.9 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.4 | 15.0 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.4 | 3.6 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.4 | 3.9 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.3 | 2.1 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 0.3 | 4.9 | GO:0000900 | translation repressor activity, nucleic acid binding(GO:0000900) |
| 0.3 | 3.8 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.3 | 3.5 | GO:0000014 | single-stranded DNA endodeoxyribonuclease activity(GO:0000014) |
| 0.3 | 1.4 | GO:0004949 | cannabinoid receptor activity(GO:0004949) |
| 0.3 | 4.3 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.3 | 6.9 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 0.3 | 7.3 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.3 | 5.4 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.3 | 1.6 | GO:0030621 | U4 snRNA binding(GO:0030621) |
| 0.3 | 18.7 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.3 | 4.6 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.3 | 0.9 | GO:0033142 | progesterone receptor binding(GO:0033142) |
| 0.3 | 8.6 | GO:0030159 | receptor signaling complex scaffold activity(GO:0030159) |
| 0.3 | 4.2 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.3 | 2.7 | GO:1990459 | transferrin receptor binding(GO:1990459) |
| 0.3 | 9.2 | GO:0030552 | cAMP binding(GO:0030552) |
| 0.3 | 18.8 | GO:0004402 | histone acetyltransferase activity(GO:0004402) |
| 0.3 | 2.6 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.3 | 9.0 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.3 | 4.6 | GO:0001011 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) TFIIB-class transcription factor binding(GO:0001093) |
| 0.3 | 1.7 | GO:0005004 | GPI-linked ephrin receptor activity(GO:0005004) |
| 0.3 | 2.8 | GO:0030550 | acetylcholine receptor inhibitor activity(GO:0030550) |
| 0.3 | 6.6 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 0.3 | 1.3 | GO:0035614 | snRNA stem-loop binding(GO:0035614) |
| 0.3 | 7.8 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.3 | 2.1 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.3 | 5.3 | GO:0008432 | JUN kinase binding(GO:0008432) |
| 0.3 | 5.6 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.3 | 13.4 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.2 | 1.7 | GO:0004726 | non-membrane spanning protein tyrosine phosphatase activity(GO:0004726) |
| 0.2 | 5.7 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.2 | 24.6 | GO:0003725 | double-stranded RNA binding(GO:0003725) |
| 0.2 | 18.4 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.2 | 2.7 | GO:0031802 | type 5 metabotropic glutamate receptor binding(GO:0031802) |
| 0.2 | 14.1 | GO:0048487 | beta-tubulin binding(GO:0048487) |
| 0.2 | 14.2 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
| 0.2 | 1.9 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.2 | 2.8 | GO:0005127 | ciliary neurotrophic factor receptor binding(GO:0005127) |
| 0.2 | 0.9 | GO:0050436 | microfibril binding(GO:0050436) |
| 0.2 | 16.6 | GO:0002039 | p53 binding(GO:0002039) |
| 0.2 | 2.3 | GO:0051434 | BH3 domain binding(GO:0051434) |
| 0.2 | 9.7 | GO:1990782 | protein tyrosine kinase binding(GO:1990782) |
| 0.2 | 0.7 | GO:0016890 | site-specific endodeoxyribonuclease activity, specific for altered base(GO:0016890) |
| 0.2 | 0.7 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 0.2 | 2.5 | GO:1990405 | protein antigen binding(GO:1990405) |
| 0.2 | 11.1 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.2 | 1.7 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.2 | 0.8 | GO:0030144 | alpha-1,6-mannosylglycoprotein 6-beta-N-acetylglucosaminyltransferase activity(GO:0030144) |
| 0.2 | 1.2 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 0.2 | 0.6 | GO:0005026 | transforming growth factor beta receptor activity, type II(GO:0005026) |
| 0.2 | 1.2 | GO:0061629 | RNA polymerase II sequence-specific DNA binding transcription factor binding(GO:0061629) |
| 0.2 | 4.5 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.2 | 8.7 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.2 | 2.7 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.2 | 6.8 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.2 | 3.3 | GO:0016500 | protein-hormone receptor activity(GO:0016500) |
| 0.2 | 1.9 | GO:0016273 | arginine N-methyltransferase activity(GO:0016273) protein-arginine N-methyltransferase activity(GO:0016274) |
| 0.2 | 2.1 | GO:0004565 | beta-galactosidase activity(GO:0004565) |
| 0.2 | 43.6 | GO:0008017 | microtubule binding(GO:0008017) |
| 0.2 | 1.1 | GO:0032184 | SUMO polymer binding(GO:0032184) |
| 0.2 | 18.3 | GO:0005518 | collagen binding(GO:0005518) |
| 0.2 | 4.4 | GO:0031491 | nucleosome binding(GO:0031491) |
| 0.2 | 0.9 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.2 | 2.3 | GO:0017127 | cholesterol transporter activity(GO:0017127) |
| 0.2 | 1.4 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.2 | 1.0 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.2 | 19.9 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
| 0.2 | 0.5 | GO:0099530 | PLC activating G-protein coupled glutamate receptor activity(GO:0001639) G-protein coupled receptor activity involved in regulation of postsynaptic membrane potential(GO:0099530) |
| 0.2 | 0.6 | GO:0070736 | protein-glycine ligase activity, initiating(GO:0070736) |
| 0.2 | 2.2 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
| 0.2 | 8.3 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.2 | 5.5 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 0.2 | 0.5 | GO:0004350 | glutamate 5-kinase activity(GO:0004349) glutamate-5-semialdehyde dehydrogenase activity(GO:0004350) delta1-pyrroline-5-carboxylate synthetase activity(GO:0017084) amino acid kinase activity(GO:0019202) |
| 0.2 | 1.8 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.2 | 22.7 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.2 | 8.2 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.2 | 2.6 | GO:0044548 | S100 protein binding(GO:0044548) |
| 0.1 | 2.5 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.1 | 1.6 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.1 | 3.0 | GO:0071855 | neuropeptide receptor binding(GO:0071855) |
| 0.1 | 4.4 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.1 | 3.7 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.1 | 0.9 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.1 | 0.4 | GO:0004982 | N-formyl peptide receptor activity(GO:0004982) |
| 0.1 | 3.0 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.1 | 1.3 | GO:0047623 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.1 | 4.3 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.1 | 0.3 | GO:0000405 | bubble DNA binding(GO:0000405) |
| 0.1 | 1.9 | GO:0035198 | miRNA binding(GO:0035198) |
| 0.1 | 1.7 | GO:0042608 | T cell receptor binding(GO:0042608) |
| 0.1 | 1.5 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.1 | 0.6 | GO:1903763 | gap junction channel activity involved in cell communication by electrical coupling(GO:1903763) |
| 0.1 | 7.2 | GO:0070035 | ATP-dependent helicase activity(GO:0008026) purine NTP-dependent helicase activity(GO:0070035) |
| 0.1 | 5.5 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.1 | 9.3 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.1 | 0.7 | GO:0017151 | DEAD/H-box RNA helicase binding(GO:0017151) |
| 0.1 | 0.7 | GO:0004487 | methylenetetrahydrofolate dehydrogenase (NAD+) activity(GO:0004487) |
| 0.1 | 1.4 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.1 | 2.4 | GO:0022840 | leak channel activity(GO:0022840) narrow pore channel activity(GO:0022842) |
| 0.1 | 1.0 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
| 0.1 | 0.5 | GO:0003875 | ADP-ribosylarginine hydrolase activity(GO:0003875) |
| 0.1 | 4.1 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.1 | 1.6 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.1 | 3.5 | GO:0008080 | N-acetyltransferase activity(GO:0008080) |
| 0.1 | 5.2 | GO:0018024 | histone-lysine N-methyltransferase activity(GO:0018024) |
| 0.1 | 15.2 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.1 | 2.6 | GO:0070300 | phosphatidic acid binding(GO:0070300) |
| 0.1 | 3.0 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.1 | 21.7 | GO:0005516 | calmodulin binding(GO:0005516) |
| 0.1 | 1.1 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.1 | 5.7 | GO:0004386 | helicase activity(GO:0004386) |
| 0.1 | 1.9 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.1 | 1.0 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.1 | 3.8 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.1 | 1.5 | GO:0000983 | transcription factor activity, RNA polymerase II core promoter sequence-specific(GO:0000983) |
| 0.1 | 0.5 | GO:0070892 | lipoteichoic acid receptor activity(GO:0070892) |
| 0.1 | 5.5 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.1 | 5.3 | GO:0070888 | E-box binding(GO:0070888) |
| 0.1 | 0.9 | GO:1990829 | C-rich single-stranded DNA binding(GO:1990829) |
| 0.1 | 40.8 | GO:0003779 | actin binding(GO:0003779) |
| 0.1 | 3.0 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.1 | 1.8 | GO:0045028 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.1 | 2.8 | GO:0008536 | Ran GTPase binding(GO:0008536) |
| 0.1 | 1.6 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.1 | 2.5 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.1 | 7.4 | GO:0042393 | histone binding(GO:0042393) |
| 0.1 | 12.1 | GO:0008201 | heparin binding(GO:0008201) |
| 0.1 | 0.5 | GO:0004826 | phenylalanine-tRNA ligase activity(GO:0004826) |
| 0.1 | 0.2 | GO:0004802 | transketolase activity(GO:0004802) |
| 0.1 | 0.5 | GO:0070008 | serine-type exopeptidase activity(GO:0070008) |
| 0.1 | 4.0 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.1 | 1.3 | GO:0045499 | chemorepellent activity(GO:0045499) |
| 0.1 | 0.2 | GO:0047256 | beta-galactosyl-N-acetylglucosaminylgalactosylglucosyl-ceramide beta-1,3-acetylglucosaminyltransferase activity(GO:0008457) lactosylceramide 1,3-N-acetyl-beta-D-glucosaminyltransferase activity(GO:0047256) |
| 0.1 | 1.7 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.1 | 0.4 | GO:0016721 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 0.1 | 1.1 | GO:0031005 | filamin binding(GO:0031005) |
| 0.1 | 0.6 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.1 | 1.0 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.1 | 0.5 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.1 | 1.1 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
| 0.1 | 1.5 | GO:0004622 | lysophospholipase activity(GO:0004622) |
| 0.1 | 7.8 | GO:0042626 | ATPase activity, coupled to transmembrane movement of substances(GO:0042626) |
| 0.1 | 2.3 | GO:0070063 | RNA polymerase binding(GO:0070063) |
| 0.1 | 0.3 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.1 | 1.2 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.1 | 0.8 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.1 | 2.5 | GO:0004521 | endoribonuclease activity(GO:0004521) |
| 0.0 | 2.1 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.0 | 0.4 | GO:0005237 | inhibitory extracellular ligand-gated ion channel activity(GO:0005237) |
| 0.0 | 3.2 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.3 | GO:0050694 | galactose 3-O-sulfotransferase activity(GO:0050694) |
| 0.0 | 1.7 | GO:0033613 | activating transcription factor binding(GO:0033613) |
| 0.0 | 4.7 | GO:0047485 | protein N-terminus binding(GO:0047485) |
| 0.0 | 5.7 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.0 | 1.7 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.2 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.0 | 0.9 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.0 | 0.4 | GO:0031386 | protein tag(GO:0031386) |
| 0.0 | 4.4 | GO:0019887 | protein kinase regulator activity(GO:0019887) |
| 0.0 | 2.4 | GO:0008173 | RNA methyltransferase activity(GO:0008173) |
| 0.0 | 3.0 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.0 | 3.8 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 0.6 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.4 | GO:0008376 | acetylgalactosaminyltransferase activity(GO:0008376) |
| 0.0 | 11.8 | GO:0004674 | protein serine/threonine kinase activity(GO:0004674) |
| 0.0 | 1.4 | GO:0051183 | vitamin transporter activity(GO:0051183) |
| 0.0 | 0.7 | GO:0004993 | G-protein coupled serotonin receptor activity(GO:0004993) |
| 0.0 | 2.9 | GO:0051020 | GTPase binding(GO:0051020) |
| 0.0 | 0.9 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.0 | 0.3 | GO:0008417 | fucosyltransferase activity(GO:0008417) |
| 0.0 | 0.2 | GO:0030553 | cGMP binding(GO:0030553) |
| 0.0 | 0.2 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.0 | 0.3 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.0 | 3.9 | GO:0003713 | transcription coactivator activity(GO:0003713) |
| 0.0 | 0.1 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.0 | 0.7 | GO:0008009 | chemokine activity(GO:0008009) |
| 0.0 | 0.2 | GO:0005412 | glucose:sodium symporter activity(GO:0005412) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 17.9 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.8 | 3.3 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.8 | 17.7 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.7 | 2.0 | PID ER NONGENOMIC PATHWAY | Plasma membrane estrogen receptor signaling |
| 0.6 | 5.6 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 0.6 | 14.6 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.6 | 9.9 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
| 0.5 | 27.1 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.5 | 30.1 | PID EPHB FWD PATHWAY | EPHB forward signaling |
| 0.5 | 22.7 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.5 | 27.6 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.5 | 36.2 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.5 | 18.4 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.5 | 23.5 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.4 | 36.9 | PID TCR PATHWAY | TCR signaling in naïve CD4+ T cells |
| 0.4 | 20.5 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.4 | 10.1 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.4 | 6.0 | ST FAS SIGNALING PATHWAY | Fas Signaling Pathway |
| 0.4 | 7.2 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.4 | 17.2 | PID ATR PATHWAY | ATR signaling pathway |
| 0.3 | 1.0 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.3 | 20.7 | PID THROMBIN PAR1 PATHWAY | PAR1-mediated thrombin signaling events |
| 0.3 | 11.1 | ST TUMOR NECROSIS FACTOR PATHWAY | Tumor Necrosis Factor Pathway. |
| 0.3 | 11.4 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.3 | 28.4 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.2 | 4.0 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.2 | 3.0 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.2 | 1.3 | PID FOXO PATHWAY | FoxO family signaling |
| 0.2 | 2.4 | PID ERBB1 RECEPTOR PROXIMAL PATHWAY | EGF receptor (ErbB1) signaling pathway |
| 0.2 | 3.4 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.2 | 4.5 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.2 | 4.5 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.2 | 9.2 | PID HDAC CLASSII PATHWAY | Signaling events mediated by HDAC Class II |
| 0.2 | 8.6 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.2 | 0.7 | ST G ALPHA I PATHWAY | G alpha i Pathway |
| 0.2 | 15.7 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.2 | 15.1 | PID E2F PATHWAY | E2F transcription factor network |
| 0.2 | 5.2 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.2 | 1.7 | PID SHP2 PATHWAY | SHP2 signaling |
| 0.1 | 2.5 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.1 | 2.3 | PID MAPK TRK PATHWAY | Trk receptor signaling mediated by the MAPK pathway |
| 0.1 | 6.6 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.1 | 3.5 | PID MYC PATHWAY | C-MYC pathway |
| 0.1 | 5.5 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.1 | 7.3 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.1 | 3.1 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.1 | 1.4 | PID NFAT 3PATHWAY | Role of Calcineurin-dependent NFAT signaling in lymphocytes |
| 0.1 | 3.0 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.1 | 2.1 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.1 | 2.4 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.1 | 3.0 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.1 | 1.7 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.1 | 0.7 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.1 | 1.1 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.1 | 2.6 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.1 | 5.8 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.1 | 0.2 | SA B CELL RECEPTOR COMPLEXES | Antigen binding to B cell receptors activates protein tyrosine kinases, such as the Src family, which ultimate activate MAP kinases. |
| 0.1 | 0.8 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.1 | 3.2 | PID ERBB1 DOWNSTREAM PATHWAY | ErbB1 downstream signaling |
| 0.1 | 3.8 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.1 | 0.6 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.1 | 0.4 | PID TRAIL PATHWAY | TRAIL signaling pathway |
| 0.1 | 1.6 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 1.0 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.0 | 1.7 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 2.2 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
| 0.0 | 0.9 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 0.7 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.0 | 2.0 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.0 | 0.6 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.0 | 0.5 | ST ADRENERGIC | Adrenergic Pathway |
| 0.0 | 0.4 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.0 | 0.4 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.0 | 0.4 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.0 | 0.4 | PID CD8 TCR DOWNSTREAM PATHWAY | Downstream signaling in naïve CD8+ T cells |
| 0.0 | 0.7 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 0.2 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.0 | 0.2 | PID IFNG PATHWAY | IFN-gamma pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 55.6 | REACTOME GENERATION OF SECOND MESSENGER MOLECULES | Genes involved in Generation of second messenger molecules |
| 1.1 | 21.7 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 1.0 | 12.4 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.8 | 13.1 | REACTOME TANDEM PORE DOMAIN POTASSIUM CHANNELS | Genes involved in Tandem pore domain potassium channels |
| 0.8 | 12.7 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.8 | 25.5 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.7 | 18.6 | REACTOME CRMPS IN SEMA3A SIGNALING | Genes involved in CRMPs in Sema3A signaling |
| 0.7 | 4.3 | REACTOME PHOSPHORYLATION OF CD3 AND TCR ZETA CHAINS | Genes involved in Phosphorylation of CD3 and TCR zeta chains |
| 0.7 | 2.7 | REACTOME RNA POL I TRANSCRIPTION | Genes involved in RNA Polymerase I Transcription |
| 0.7 | 14.7 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |
| 0.6 | 33.2 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.6 | 21.2 | REACTOME PLATELET SENSITIZATION BY LDL | Genes involved in Platelet sensitization by LDL |
| 0.6 | 11.9 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.6 | 1.8 | REACTOME PD1 SIGNALING | Genes involved in PD-1 signaling |
| 0.6 | 9.1 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.5 | 2.2 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.5 | 9.2 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.5 | 0.5 | REACTOME ARMS MEDIATED ACTIVATION | Genes involved in ARMS-mediated activation |
| 0.5 | 11.1 | REACTOME RNA POL III CHAIN ELONGATION | Genes involved in RNA Polymerase III Chain Elongation |
| 0.5 | 15.1 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.5 | 44.3 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.4 | 3.3 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.4 | 21.7 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.4 | 3.1 | REACTOME CTNNB1 PHOSPHORYLATION CASCADE | Genes involved in Beta-catenin phosphorylation cascade |
| 0.4 | 2.3 | REACTOME SIGNALLING TO P38 VIA RIT AND RIN | Genes involved in Signalling to p38 via RIT and RIN |
| 0.4 | 17.0 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.4 | 24.4 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.4 | 7.7 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.3 | 5.2 | REACTOME ANDROGEN BIOSYNTHESIS | Genes involved in Androgen biosynthesis |
| 0.3 | 7.6 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.3 | 3.5 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 3 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 3 Promoter |
| 0.3 | 19.0 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.3 | 8.8 | REACTOME GPVI MEDIATED ACTIVATION CASCADE | Genes involved in GPVI-mediated activation cascade |
| 0.3 | 2.3 | REACTOME INFLAMMASOMES | Genes involved in Inflammasomes |
| 0.3 | 26.2 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.3 | 38.9 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.3 | 4.1 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.3 | 6.8 | REACTOME RNA POL I TRANSCRIPTION INITIATION | Genes involved in RNA Polymerase I Transcription Initiation |
| 0.3 | 21.5 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.3 | 5.0 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.3 | 6.7 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.3 | 0.3 | REACTOME BINDING AND ENTRY OF HIV VIRION | Genes involved in Binding and entry of HIV virion |
| 0.3 | 3.9 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.3 | 5.5 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.3 | 6.9 | REACTOME ACYL CHAIN REMODELLING OF PE | Genes involved in Acyl chain remodelling of PE |
| 0.3 | 3.2 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.3 | 5.2 | REACTOME IL RECEPTOR SHC SIGNALING | Genes involved in Interleukin receptor SHC signaling |
| 0.3 | 7.7 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.2 | 9.5 | REACTOME NITRIC OXIDE STIMULATES GUANYLATE CYCLASE | Genes involved in Nitric oxide stimulates guanylate cyclase |
| 0.2 | 2.2 | REACTOME MICRORNA MIRNA BIOGENESIS | Genes involved in MicroRNA (miRNA) Biogenesis |
| 0.2 | 13.0 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.2 | 2.4 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.2 | 2.6 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 0.2 | 7.3 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.2 | 33.7 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.2 | 5.2 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.2 | 4.3 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.2 | 1.2 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.2 | 3.6 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.2 | 1.3 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 2 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 2 Promoter |
| 0.2 | 1.8 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.2 | 2.1 | REACTOME COSTIMULATION BY THE CD28 FAMILY | Genes involved in Costimulation by the CD28 family |
| 0.2 | 5.1 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.2 | 14.3 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.2 | 9.2 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.2 | 7.3 | REACTOME P75 NTR RECEPTOR MEDIATED SIGNALLING | Genes involved in p75 NTR receptor-mediated signalling |
| 0.1 | 2.4 | REACTOME PKA MEDIATED PHOSPHORYLATION OF CREB | Genes involved in PKA-mediated phosphorylation of CREB |
| 0.1 | 3.3 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.1 | 8.3 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.1 | 13.0 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.1 | 1.4 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.1 | 1.7 | REACTOME P38MAPK EVENTS | Genes involved in p38MAPK events |
| 0.1 | 2.2 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.1 | 7.8 | REACTOME SIGNALING BY PDGF | Genes involved in Signaling by PDGF |
| 0.1 | 3.5 | REACTOME MEIOSIS | Genes involved in Meiosis |
| 0.1 | 2.3 | REACTOME ERK MAPK TARGETS | Genes involved in ERK/MAPK targets |
| 0.1 | 1.6 | REACTOME REGULATION OF IFNA SIGNALING | Genes involved in Regulation of IFNA signaling |
| 0.1 | 2.3 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.1 | 4.0 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.1 | 6.4 | REACTOME LATE PHASE OF HIV LIFE CYCLE | Genes involved in Late Phase of HIV Life Cycle |
| 0.1 | 2.2 | REACTOME NOD1 2 SIGNALING PATHWAY | Genes involved in NOD1/2 Signaling Pathway |
| 0.1 | 1.9 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.1 | 1.2 | REACTOME SIGNALING BY SCF KIT | Genes involved in Signaling by SCF-KIT |
| 0.1 | 4.0 | REACTOME REGULATION OF ORNITHINE DECARBOXYLASE ODC | Genes involved in Regulation of ornithine decarboxylase (ODC) |
| 0.1 | 0.5 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.1 | 2.5 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.1 | 1.3 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.1 | 0.3 | REACTOME FORMATION OF TRANSCRIPTION COUPLED NER TC NER REPAIR COMPLEX | Genes involved in Formation of transcription-coupled NER (TC-NER) repair complex |
| 0.0 | 3.5 | REACTOME DNA REPAIR | Genes involved in DNA Repair |
| 0.0 | 0.4 | REACTOME GLYCOPROTEIN HORMONES | Genes involved in Glycoprotein hormones |
| 0.0 | 1.6 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 1.1 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 1.2 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 1.5 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.0 | 0.7 | REACTOME SMAD2 SMAD3 SMAD4 HETEROTRIMER REGULATES TRANSCRIPTION | Genes involved in SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription |
| 0.0 | 0.6 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.0 | 1.9 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.0 | 0.4 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.0 | 1.9 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 0.2 | REACTOME TRANSMISSION ACROSS CHEMICAL SYNAPSES | Genes involved in Transmission across Chemical Synapses |
| 0.0 | 0.4 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.0 | 0.3 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.0 | 0.2 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |