Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
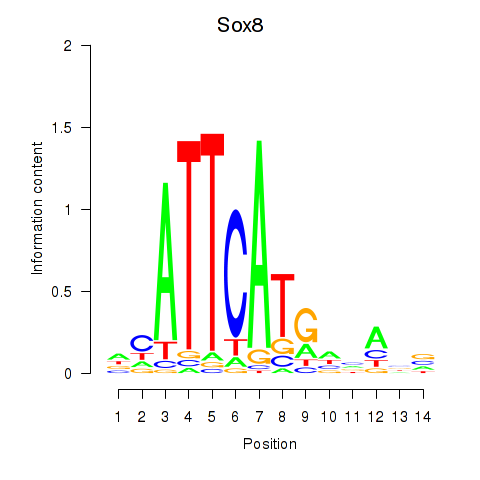
Results for Sox8
Z-value: 0.77
Transcription factors associated with Sox8
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Sox8
|
ENSMUSG00000024176.11 | Sox8 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Sox8 | mm39_v1_chr17_-_25789652_25789670 | 0.58 | 8.7e-08 | Click! |
Activity profile of Sox8 motif
Sorted Z-values of Sox8 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Sox8
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr13_-_51723473 | 8.46 |
ENSMUST00000239056.2
ENSMUST00000223543.3 |
Shc3
|
src homology 2 domain-containing transforming protein C3 |
| chr7_-_126399778 | 5.28 |
ENSMUST00000141355.4
|
Aldoa
|
aldolase A, fructose-bisphosphate |
| chr7_-_126399208 | 4.79 |
ENSMUST00000133514.8
ENSMUST00000151137.8 |
Aldoa
|
aldolase A, fructose-bisphosphate |
| chr4_+_11758147 | 4.51 |
ENSMUST00000029871.12
ENSMUST00000108303.2 |
Cdh17
|
cadherin 17 |
| chr7_-_126399574 | 4.20 |
ENSMUST00000106348.8
|
Aldoa
|
aldolase A, fructose-bisphosphate |
| chr9_+_36744016 | 3.97 |
ENSMUST00000214772.2
|
Fez1
|
fasciculation and elongation protein zeta 1 (zygin I) |
| chr13_+_93441307 | 3.90 |
ENSMUST00000080127.12
|
Homer1
|
homer scaffolding protein 1 |
| chr1_-_79838897 | 3.86 |
ENSMUST00000190724.2
|
Serpine2
|
serine (or cysteine) peptidase inhibitor, clade E, member 2 |
| chr3_+_65016750 | 3.81 |
ENSMUST00000049230.11
|
Kcnab1
|
potassium voltage-gated channel, shaker-related subfamily, beta member 1 |
| chr9_+_109881083 | 3.49 |
ENSMUST00000164930.8
ENSMUST00000199498.5 |
Map4
|
microtubule-associated protein 4 |
| chr1_+_109911467 | 3.28 |
ENSMUST00000172005.8
|
Cdh7
|
cadherin 7, type 2 |
| chr9_-_95697441 | 3.26 |
ENSMUST00000119760.2
|
Pls1
|
plastin 1 (I-isoform) |
| chr17_+_44445659 | 3.13 |
ENSMUST00000239215.2
|
Clic5
|
chloride intracellular channel 5 |
| chr5_-_103174794 | 2.93 |
ENSMUST00000128869.8
|
Mapk10
|
mitogen-activated protein kinase 10 |
| chr7_-_100306160 | 2.93 |
ENSMUST00000107046.8
ENSMUST00000107045.9 ENSMUST00000139708.9 |
Plekhb1
|
pleckstrin homology domain containing, family B (evectins) member 1 |
| chr10_-_33972503 | 2.75 |
ENSMUST00000069125.8
|
Calhm5
|
calcium homeostasis modulator family member 5 |
| chr9_+_65008735 | 2.72 |
ENSMUST00000213533.2
ENSMUST00000035499.5 ENSMUST00000077696.13 ENSMUST00000166273.2 |
Igdcc4
|
immunoglobulin superfamily, DCC subclass, member 4 |
| chrX_+_133305529 | 2.72 |
ENSMUST00000113224.9
ENSMUST00000113226.2 |
Drp2
|
dystrophin related protein 2 |
| chr2_-_73605387 | 2.69 |
ENSMUST00000166199.9
|
Chn1
|
chimerin 1 |
| chr12_-_115722081 | 2.68 |
ENSMUST00000103541.3
|
Ighv1-72
|
immunoglobulin heavy variable 1-72 |
| chr2_+_109522781 | 2.66 |
ENSMUST00000111050.10
|
Bdnf
|
brain derived neurotrophic factor |
| chr18_+_37085673 | 2.54 |
ENSMUST00000192512.6
ENSMUST00000192295.2 ENSMUST00000115661.5 |
Pcdha4
Gm42416
|
protocadherin alpha 4 predicted gene, 42416 |
| chr16_-_29360301 | 2.52 |
ENSMUST00000057018.15
ENSMUST00000182627.8 |
Atp13a4
|
ATPase type 13A4 |
| chr4_-_116263183 | 2.36 |
ENSMUST00000123072.8
ENSMUST00000144281.2 |
Mast2
|
microtubule associated serine/threonine kinase 2 |
| chr8_-_34237752 | 2.25 |
ENSMUST00000179364.3
|
Smim18
|
small integral membrane protein 18 |
| chr1_+_135710803 | 2.19 |
ENSMUST00000132795.8
|
Tnni1
|
troponin I, skeletal, slow 1 |
| chr19_+_4771089 | 2.09 |
ENSMUST00000238976.3
|
Sptbn2
|
spectrin beta, non-erythrocytic 2 |
| chrX_-_161426542 | 2.02 |
ENSMUST00000101102.2
|
Reps2
|
RALBP1 associated Eps domain containing protein 2 |
| chr18_+_37637317 | 2.02 |
ENSMUST00000052179.8
|
Pcdhb20
|
protocadherin beta 20 |
| chr4_+_127062924 | 1.94 |
ENSMUST00000046659.14
|
Dlgap3
|
DLG associated protein 3 |
| chrX_-_161426624 | 1.90 |
ENSMUST00000112334.8
|
Reps2
|
RALBP1 associated Eps domain containing protein 2 |
| chr4_-_58553553 | 1.79 |
ENSMUST00000107575.9
ENSMUST00000107574.8 ENSMUST00000147354.8 |
Lpar1
|
lysophosphatidic acid receptor 1 |
| chr12_-_115109539 | 1.79 |
ENSMUST00000192554.6
ENSMUST00000103522.3 |
Ighv1-52
|
immunoglobulin heavy variable 1-52 |
| chr11_-_46057224 | 1.70 |
ENSMUST00000020679.3
|
Nipal4
|
NIPA-like domain containing 4 |
| chr5_-_44259293 | 1.62 |
ENSMUST00000074113.13
|
Prom1
|
prominin 1 |
| chr10_-_128016135 | 1.62 |
ENSMUST00000238843.2
ENSMUST00000099139.9 |
Rbms2
|
RNA binding motif, single stranded interacting protein 2 |
| chr13_-_19579898 | 1.62 |
ENSMUST00000197565.3
ENSMUST00000221380.2 ENSMUST00000200323.3 ENSMUST00000199924.2 ENSMUST00000222869.2 |
Stard3nl
|
STARD3 N-terminal like |
| chr15_-_103123711 | 1.62 |
ENSMUST00000122182.2
ENSMUST00000108813.10 ENSMUST00000127191.2 |
Cbx5
|
chromobox 5 |
| chr12_-_115083839 | 1.55 |
ENSMUST00000103521.3
|
Ighv1-50
|
immunoglobulin heavy variable 1-50 |
| chr13_-_19579961 | 1.53 |
ENSMUST00000039694.13
|
Stard3nl
|
STARD3 N-terminal like |
| chr12_-_115172211 | 1.53 |
ENSMUST00000103526.3
|
Ighv1-55
|
immunoglobulin heavy variable 1-55 |
| chrX_+_157993303 | 1.48 |
ENSMUST00000112493.8
|
Rps6ka3
|
ribosomal protein S6 kinase polypeptide 3 |
| chr5_-_44259010 | 1.44 |
ENSMUST00000087441.11
|
Prom1
|
prominin 1 |
| chr13_+_93440572 | 1.43 |
ENSMUST00000109493.9
|
Homer1
|
homer scaffolding protein 1 |
| chr14_-_52628228 | 1.29 |
ENSMUST00000078171.2
|
Olfr1511
|
olfactory receptor 1511 |
| chr2_+_116709167 | 1.22 |
ENSMUST00000123598.8
ENSMUST00000155470.8 |
Tmco5
|
transmembrane and coiled-coil domains 5 |
| chr2_-_180844582 | 1.08 |
ENSMUST00000016511.6
|
Ptk6
|
PTK6 protein tyrosine kinase 6 |
| chr11_+_113550109 | 1.07 |
ENSMUST00000137878.2
|
Cog1
|
component of oligomeric golgi complex 1 |
| chr13_-_31134281 | 1.03 |
ENSMUST00000102946.8
|
Exoc2
|
exocyst complex component 2 |
| chr12_-_115425105 | 0.99 |
ENSMUST00000103532.3
|
Ighv1-62-3
|
immunoglobulin heavy variable 1-62-3 |
| chr14_+_21881794 | 0.97 |
ENSMUST00000152562.8
|
Vdac2
|
voltage-dependent anion channel 2 |
| chr1_+_136059101 | 0.97 |
ENSMUST00000075164.11
|
Kif21b
|
kinesin family member 21B |
| chr11_-_107685383 | 0.84 |
ENSMUST00000021066.4
|
Cacng4
|
calcium channel, voltage-dependent, gamma subunit 4 |
| chr5_+_53424471 | 0.84 |
ENSMUST00000147148.5
|
Smim20
|
small integral membrane protein 20 |
| chr17_-_26052335 | 0.76 |
ENSMUST00000044911.10
ENSMUST00000237186.2 |
Stub1
|
STIP1 homology and U-Box containing protein 1 |
| chr2_+_116709179 | 0.73 |
ENSMUST00000028834.3
|
Tmco5
|
transmembrane and coiled-coil domains 5 |
| chr17_+_26934617 | 0.72 |
ENSMUST00000062519.14
ENSMUST00000144221.2 ENSMUST00000142539.8 ENSMUST00000151681.2 |
Crebrf
|
CREB3 regulatory factor |
| chr17_-_40553176 | 0.71 |
ENSMUST00000026499.6
|
Crisp3
|
cysteine-rich secretory protein 3 |
| chr12_-_114843941 | 0.69 |
ENSMUST00000191862.6
ENSMUST00000103513.3 |
Ighv1-36
|
immunoglobulin heavy variable 1-36 |
| chr12_-_114451189 | 0.68 |
ENSMUST00000103493.3
|
Ighv1-4
|
immunoglobulin heavy variable 1-4 |
| chrX_-_23151771 | 0.67 |
ENSMUST00000115319.9
|
Klhl13
|
kelch-like 13 |
| chr19_-_33764859 | 0.59 |
ENSMUST00000148137.9
|
Lipo1
|
lipase, member O1 |
| chr18_+_65831324 | 0.51 |
ENSMUST00000115097.8
ENSMUST00000117694.2 ENSMUST00000235962.2 |
Oacyl
|
O-acyltransferase like |
| chr10_+_58207229 | 0.45 |
ENSMUST00000238939.2
|
Lims1
|
LIM and senescent cell antigen-like domains 1 |
| chr7_-_104646109 | 0.39 |
ENSMUST00000050599.5
|
Olfr672
|
olfactory receptor 672 |
| chr1_-_157072060 | 0.38 |
ENSMUST00000129880.8
|
Rasal2
|
RAS protein activator like 2 |
| chr7_+_106513407 | 0.38 |
ENSMUST00000098140.2
|
Olfr1532-ps1
|
olfactory receptor 1532, pseudogene 1 |
| chr19_-_40365318 | 0.28 |
ENSMUST00000239304.2
|
Sorbs1
|
sorbin and SH3 domain containing 1 |
| chr13_+_19526322 | 0.24 |
ENSMUST00000184430.2
|
Trgj4
|
T cell receptor gamma joining 4 |
| chr9_-_61883845 | 0.23 |
ENSMUST00000113990.2
|
Paqr5
|
progestin and adipoQ receptor family member V |
| chr9_+_108566513 | 0.22 |
ENSMUST00000192344.2
|
Prkar2a
|
protein kinase, cAMP dependent regulatory, type II alpha |
| chr7_-_86044743 | 0.21 |
ENSMUST00000053958.6
|
Olfr303
|
olfactory receptor 303 |
| chrX_-_71961890 | 0.20 |
ENSMUST00000152200.2
|
Cetn2
|
centrin 2 |
| chr11_+_114618209 | 0.17 |
ENSMUST00000069325.14
|
Dnai2
|
dynein axonemal intermediate chain 2 |
| chr4_+_154226819 | 0.17 |
ENSMUST00000030895.12
|
Wrap73
|
WD repeat containing, antisense to Trp73 |
| chr11_-_43638790 | 0.15 |
ENSMUST00000048578.3
ENSMUST00000109278.8 |
Ttc1
|
tetratricopeptide repeat domain 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 4.5 | GO:0002314 | germinal center B cell differentiation(GO:0002314) |
| 1.3 | 3.9 | GO:0061107 | seminal vesicle development(GO:0061107) |
| 0.9 | 2.7 | GO:0061193 | taste bud development(GO:0061193) |
| 0.8 | 3.3 | GO:1902896 | terminal web assembly(GO:1902896) |
| 0.8 | 14.3 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 0.6 | 3.1 | GO:0072139 | glomerular parietal epithelial cell differentiation(GO:0072139) |
| 0.5 | 3.5 | GO:0051012 | microtubule sliding(GO:0051012) |
| 0.4 | 1.8 | GO:1904565 | response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904565) cellular response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904566) |
| 0.4 | 2.9 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 0.3 | 5.3 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.2 | 4.0 | GO:1902902 | negative regulation of autophagosome assembly(GO:1902902) |
| 0.2 | 2.4 | GO:0042090 | interleukin-12 biosynthetic process(GO:0042090) regulation of interleukin-12 biosynthetic process(GO:0045075) |
| 0.2 | 0.8 | GO:0090034 | regulation of chaperone-mediated protein complex assembly(GO:0090034) positive regulation of chaperone-mediated protein complex assembly(GO:0090035) |
| 0.1 | 3.1 | GO:0002024 | diet induced thermogenesis(GO:0002024) |
| 0.1 | 1.0 | GO:0001927 | exocyst assembly(GO:0001927) |
| 0.1 | 3.8 | GO:1902260 | negative regulation of delayed rectifier potassium channel activity(GO:1902260) |
| 0.1 | 0.7 | GO:0038161 | prolactin signaling pathway(GO:0038161) |
| 0.1 | 2.2 | GO:0014883 | transition between fast and slow fiber(GO:0014883) |
| 0.1 | 1.7 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.1 | 2.1 | GO:0021692 | cerebellar Purkinje cell layer morphogenesis(GO:0021692) |
| 0.1 | 0.8 | GO:2000311 | regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000311) |
| 0.1 | 8.5 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.1 | 8.5 | GO:0035249 | synaptic transmission, glutamatergic(GO:0035249) |
| 0.1 | 2.7 | GO:0008045 | motor neuron axon guidance(GO:0008045) |
| 0.0 | 1.1 | GO:0060575 | intestinal epithelial cell differentiation(GO:0060575) |
| 0.0 | 0.5 | GO:2000346 | negative regulation of hepatocyte proliferation(GO:2000346) |
| 0.0 | 1.1 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 0.0 | 3.3 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.0 | 1.5 | GO:0002224 | toll-like receptor signaling pathway(GO:0002224) |
| 0.0 | 4.1 | GO:0050808 | synapse organization(GO:0050808) |
| 0.0 | 0.2 | GO:2000480 | negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 14.3 | GO:0035686 | sperm fibrous sheath(GO:0035686) |
| 0.4 | 3.1 | GO:0071914 | prominosome(GO:0071914) |
| 0.4 | 3.3 | GO:1990357 | terminal web(GO:1990357) |
| 0.3 | 3.9 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 0.3 | 2.1 | GO:0008091 | spectrin(GO:0008091) |
| 0.2 | 3.8 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.2 | 3.2 | GO:0044232 | organelle membrane contact site(GO:0044232) |
| 0.1 | 2.7 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.1 | 2.2 | GO:0005861 | troponin complex(GO:0005861) |
| 0.1 | 5.3 | GO:0043034 | costamere(GO:0043034) |
| 0.1 | 1.9 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.1 | 1.1 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.1 | 8.5 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.1 | 1.6 | GO:0010369 | chromocenter(GO:0010369) |
| 0.1 | 0.8 | GO:0031371 | ubiquitin conjugating enzyme complex(GO:0031371) |
| 0.1 | 0.2 | GO:0071942 | XPC complex(GO:0071942) |
| 0.0 | 0.3 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.0 | 1.0 | GO:0000145 | exocyst(GO:0000145) |
| 0.0 | 3.1 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.0 | 1.0 | GO:0046930 | pore complex(GO:0046930) |
| 0.0 | 3.5 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.0 | 1.8 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.0 | 0.4 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.0 | 0.8 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 2.9 | GO:0043204 | perikaryon(GO:0043204) |
| 0.0 | 0.2 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.0 | 1.0 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 1.3 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.0 | 4.5 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 2.7 | GO:0045211 | postsynaptic membrane(GO:0045211) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 14.3 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 1.1 | 4.5 | GO:0035673 | oligopeptide transmembrane transporter activity(GO:0035673) |
| 0.9 | 2.7 | GO:0005169 | neurotrophin TRKB receptor binding(GO:0005169) |
| 0.6 | 2.9 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.4 | 5.3 | GO:0031802 | type 5 metabotropic glutamate receptor binding(GO:0031802) |
| 0.3 | 1.8 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 0.1 | 3.8 | GO:0070402 | NADPH binding(GO:0070402) |
| 0.1 | 1.0 | GO:0015288 | porin activity(GO:0015288) |
| 0.1 | 8.5 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.1 | 2.5 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.1 | 1.7 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.1 | 0.8 | GO:0030911 | TPR domain binding(GO:0030911) |
| 0.1 | 4.0 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.1 | 8.5 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.1 | 2.7 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.0 | 1.0 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.0 | 3.2 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.0 | 3.1 | GO:0042805 | actinin binding(GO:0042805) |
| 0.0 | 1.9 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 1.5 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.0 | 3.1 | GO:0005254 | chloride channel activity(GO:0005254) |
| 0.0 | 3.9 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.2 | GO:0032795 | heterotrimeric G-protein binding(GO:0032795) |
| 0.0 | 1.6 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.0 | 1.1 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 5.2 | GO:0008017 | microtubule binding(GO:0008017) |
| 0.0 | 0.2 | GO:0008603 | cAMP-dependent protein kinase regulator activity(GO:0008603) |
| 0.0 | 0.8 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 10.5 | GO:0005509 | calcium ion binding(GO:0005509) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 14.3 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.1 | 11.1 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.1 | 2.9 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.1 | 1.5 | SA B CELL RECEPTOR COMPLEXES | Antigen binding to B cell receptors activates protein tyrosine kinases, such as the Src family, which ultimate activate MAP kinases. |
| 0.0 | 2.3 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.0 | 2.7 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.0 | 1.3 | PID INSULIN PATHWAY | Insulin Pathway |
| 0.0 | 1.8 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.0 | 0.8 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.0 | 3.9 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 8.5 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.3 | 14.3 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.2 | 2.9 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.1 | 3.3 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.1 | 3.8 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.0 | 1.5 | REACTOME ERK MAPK TARGETS | Genes involved in ERK/MAPK targets |
| 0.0 | 2.1 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 1.0 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.0 | 2.2 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 0.8 | REACTOME DOWNREGULATION OF TGF BETA RECEPTOR SIGNALING | Genes involved in Downregulation of TGF-beta receptor signaling |
| 0.0 | 0.8 | REACTOME TRAFFICKING OF AMPA RECEPTORS | Genes involved in Trafficking of AMPA receptors |
| 0.0 | 0.5 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 2.7 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 1.8 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |