Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
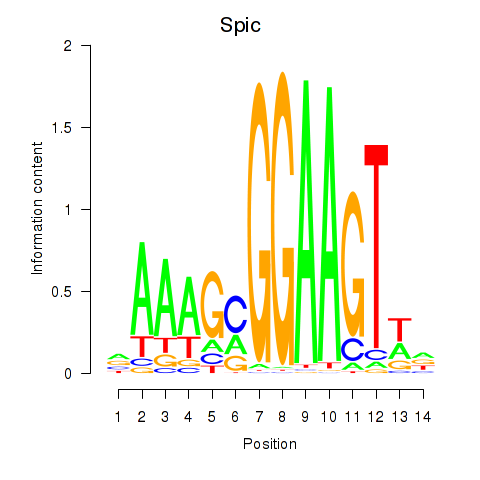
Results for Spic
Z-value: 3.42
Transcription factors associated with Spic
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Spic
|
ENSMUSG00000004359.17 | Spic |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Spic | mm39_v1_chr10_-_88518878_88518892 | 0.83 | 2.9e-19 | Click! |
Activity profile of Spic motif
Sorted Z-values of Spic motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Spic
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_-_136918671 | 28.13 |
ENSMUST00000032344.12
|
Arhgdib
|
Rho, GDP dissociation inhibitor (GDI) beta |
| chr6_-_136918844 | 27.99 |
ENSMUST00000204934.2
|
Arhgdib
|
Rho, GDP dissociation inhibitor (GDI) beta |
| chr7_+_43057611 | 25.61 |
ENSMUST00000005592.7
|
Siglecg
|
sialic acid binding Ig-like lectin G |
| chr7_-_126303351 | 25.35 |
ENSMUST00000106364.8
|
Coro1a
|
coronin, actin binding protein 1A |
| chrX_+_106132840 | 23.66 |
ENSMUST00000118666.8
ENSMUST00000053375.4 |
P2ry10
|
purinergic receptor P2Y, G-protein coupled 10 |
| chr11_-_79414542 | 23.63 |
ENSMUST00000179322.2
|
Evi2b
|
ecotropic viral integration site 2b |
| chr7_-_126303887 | 23.53 |
ENSMUST00000131415.8
|
Coro1a
|
coronin, actin binding protein 1A |
| chr7_+_28682253 | 22.20 |
ENSMUST00000085835.8
|
Map4k1
|
mitogen-activated protein kinase kinase kinase kinase 1 |
| chr4_+_130640436 | 22.13 |
ENSMUST00000151698.8
|
Laptm5
|
lysosomal-associated protein transmembrane 5 |
| chr11_-_115024807 | 21.89 |
ENSMUST00000106561.8
ENSMUST00000051264.14 ENSMUST00000106562.3 |
Cd300lf
|
CD300 molecule like family member F |
| chr6_-_136918885 | 21.86 |
ENSMUST00000111891.4
|
Arhgdib
|
Rho, GDP dissociation inhibitor (GDI) beta |
| chr7_-_44888220 | 21.58 |
ENSMUST00000210372.2
ENSMUST00000209779.2 ENSMUST00000098461.10 ENSMUST00000211373.2 |
Cd37
|
CD37 antigen |
| chr12_+_98234884 | 21.47 |
ENSMUST00000075072.6
|
Gpr65
|
G-protein coupled receptor 65 |
| chr6_+_48661503 | 21.34 |
ENSMUST00000067506.14
ENSMUST00000119575.8 ENSMUST00000121957.8 ENSMUST00000090070.6 |
Gimap4
|
GTPase, IMAP family member 4 |
| chr16_-_75706161 | 21.24 |
ENSMUST00000114239.9
|
Samsn1
|
SAM domain, SH3 domain and nuclear localization signals, 1 |
| chr1_-_144651157 | 20.37 |
ENSMUST00000027603.4
|
Rgs18
|
regulator of G-protein signaling 18 |
| chr7_+_28140352 | 20.01 |
ENSMUST00000078845.13
|
Gmfg
|
glia maturation factor, gamma |
| chr6_-_48685108 | 19.92 |
ENSMUST00000126422.3
ENSMUST00000119315.2 ENSMUST00000053661.7 |
Gimap6
|
GTPase, IMAP family member 6 |
| chr4_+_138503046 | 19.77 |
ENSMUST00000030528.9
|
Pla2g2d
|
phospholipase A2, group IID |
| chr1_+_171723231 | 19.68 |
ENSMUST00000097466.3
|
Gm10521
|
predicted gene 10521 |
| chr8_-_123159639 | 19.19 |
ENSMUST00000212600.2
|
Cyba
|
cytochrome b-245, alpha polypeptide |
| chr15_+_78128990 | 19.15 |
ENSMUST00000096357.12
|
Ncf4
|
neutrophil cytosolic factor 4 |
| chr3_+_87283687 | 19.08 |
ENSMUST00000163661.8
ENSMUST00000072480.9 |
Fcrl1
|
Fc receptor-like 1 |
| chr7_-_126303689 | 18.77 |
ENSMUST00000135087.8
|
Coro1a
|
coronin, actin binding protein 1A |
| chr7_+_142014546 | 18.67 |
ENSMUST00000105968.8
ENSMUST00000018963.11 ENSMUST00000105967.8 |
Lsp1
|
lymphocyte specific 1 |
| chr7_-_121732067 | 18.45 |
ENSMUST00000106469.8
ENSMUST00000063587.13 ENSMUST00000106468.7 ENSMUST00000098068.10 |
Palb2
|
partner and localizer of BRCA2 |
| chr10_+_79715448 | 18.36 |
ENSMUST00000006679.15
|
Prtn3
|
proteinase 3 |
| chr6_+_48661477 | 18.16 |
ENSMUST00000118802.8
|
Gimap4
|
GTPase, IMAP family member 4 |
| chr2_+_43638814 | 17.98 |
ENSMUST00000112824.8
ENSMUST00000055776.8 |
Arhgap15
|
Rho GTPase activating protein 15 |
| chr3_-_106697459 | 17.87 |
ENSMUST00000038845.10
|
Cd53
|
CD53 antigen |
| chr19_+_4204605 | 17.59 |
ENSMUST00000061086.9
|
Ptprcap
|
protein tyrosine phosphatase, receptor type, C polypeptide-associated protein |
| chr17_+_57586094 | 17.26 |
ENSMUST00000169220.9
ENSMUST00000005889.16 ENSMUST00000112870.5 |
Vav1
|
vav 1 oncogene |
| chr9_+_7558449 | 17.25 |
ENSMUST00000018765.4
|
Mmp8
|
matrix metallopeptidase 8 |
| chr7_+_28140450 | 17.21 |
ENSMUST00000135686.2
|
Gmfg
|
glia maturation factor, gamma |
| chr6_+_41039255 | 17.17 |
ENSMUST00000103266.3
|
Trbv5
|
T cell receptor beta, variable 5 |
| chr10_-_117118226 | 17.01 |
ENSMUST00000092163.9
|
Lyz2
|
lysozyme 2 |
| chr6_-_70237939 | 16.88 |
ENSMUST00000103386.3
|
Igkv6-23
|
immunoglobulin kappa variable 6-23 |
| chr9_-_20556031 | 16.85 |
ENSMUST00000148631.8
ENSMUST00000131128.2 ENSMUST00000151861.9 ENSMUST00000131343.8 ENSMUST00000086458.10 |
Fbxl12
|
F-box and leucine-rich repeat protein 12 |
| chr10_+_75778885 | 16.69 |
ENSMUST00000121151.2
|
Vpreb3
|
pre-B lymphocyte gene 3 |
| chr17_-_57385490 | 15.95 |
ENSMUST00000011623.9
|
Dennd1c
|
DENN/MADD domain containing 1C |
| chr13_+_30933209 | 15.88 |
ENSMUST00000021784.10
ENSMUST00000110307.3 ENSMUST00000222125.2 |
Irf4
|
interferon regulatory factor 4 |
| chr6_+_70348416 | 15.78 |
ENSMUST00000103391.4
|
Igkv6-17
|
immunoglobulin kappa variable 6-17 |
| chr8_+_95650315 | 15.65 |
ENSMUST00000153448.9
ENSMUST00000166802.9 ENSMUST00000074570.10 |
Adgrg5
|
adhesion G protein-coupled receptor G5 |
| chr7_-_19005721 | 15.63 |
ENSMUST00000032561.9
|
Vasp
|
vasodilator-stimulated phosphoprotein |
| chr5_-_134258435 | 15.50 |
ENSMUST00000016094.13
ENSMUST00000111275.8 ENSMUST00000144086.2 |
Ncf1
|
neutrophil cytosolic factor 1 |
| chr12_+_112727089 | 15.49 |
ENSMUST00000063888.5
|
Pld4
|
phospholipase D family, member 4 |
| chr6_-_70383976 | 15.42 |
ENSMUST00000103393.2
|
Igkv6-15
|
immunoglobulin kappa variable 6-15 |
| chr9_+_123822000 | 15.41 |
ENSMUST00000039171.9
|
Ccr3
|
chemokine (C-C motif) receptor 3 |
| chr9_+_88838953 | 15.35 |
ENSMUST00000098485.4
|
Bcl2a1a
|
B cell leukemia/lymphoma 2 related protein A1a |
| chr11_+_98798627 | 15.30 |
ENSMUST00000092706.13
|
Cdc6
|
cell division cycle 6 |
| chr6_+_129326927 | 15.25 |
ENSMUST00000065289.6
|
Clec12a
|
C-type lectin domain family 12, member a |
| chr4_+_130640611 | 14.93 |
ENSMUST00000156225.8
ENSMUST00000156742.8 |
Laptm5
|
lysosomal-associated protein transmembrane 5 |
| chr5_+_149202157 | 14.92 |
ENSMUST00000200806.4
|
Alox5ap
|
arachidonate 5-lipoxygenase activating protein |
| chr8_-_123159663 | 14.86 |
ENSMUST00000017604.10
|
Cyba
|
cytochrome b-245, alpha polypeptide |
| chr5_+_149201577 | 14.75 |
ENSMUST00000071130.5
|
Alox5ap
|
arachidonate 5-lipoxygenase activating protein |
| chr9_-_114610879 | 14.71 |
ENSMUST00000084867.9
ENSMUST00000216760.2 ENSMUST00000035009.16 |
Cmtm7
|
CKLF-like MARVEL transmembrane domain containing 7 |
| chr6_-_136918495 | 14.45 |
ENSMUST00000111892.8
|
Arhgdib
|
Rho, GDP dissociation inhibitor (GDI) beta |
| chr5_+_66018555 | 14.44 |
ENSMUST00000031106.8
|
Rhoh
|
ras homolog family member H |
| chr7_-_44888532 | 14.43 |
ENSMUST00000033063.15
|
Cd37
|
CD37 antigen |
| chr6_-_124715618 | 14.25 |
ENSMUST00000171549.9
|
Ptpn6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr15_+_78129040 | 14.07 |
ENSMUST00000133618.3
|
Ncf4
|
neutrophil cytosolic factor 4 |
| chr7_+_126184108 | 14.06 |
ENSMUST00000039522.8
|
Apobr
|
apolipoprotein B receptor |
| chr15_+_103362195 | 14.01 |
ENSMUST00000047405.9
|
Nckap1l
|
NCK associated protein 1 like |
| chrX_+_106299484 | 13.96 |
ENSMUST00000101294.9
ENSMUST00000118820.8 ENSMUST00000120971.8 |
Gpr174
|
G protein-coupled receptor 174 |
| chr3_-_105839980 | 13.96 |
ENSMUST00000098758.5
|
I830077J02Rik
|
RIKEN cDNA I830077J02 gene |
| chr2_-_129213050 | 13.93 |
ENSMUST00000028881.14
|
Il1b
|
interleukin 1 beta |
| chr13_+_102830029 | 13.78 |
ENSMUST00000022124.10
ENSMUST00000171267.2 ENSMUST00000167144.2 ENSMUST00000170878.2 |
Cd180
|
CD180 antigen |
| chr10_+_130158737 | 13.66 |
ENSMUST00000217702.2
ENSMUST00000042586.10 ENSMUST00000218605.2 |
Tespa1
|
thymocyte expressed, positive selection associated 1 |
| chr11_+_68322945 | 13.57 |
ENSMUST00000021283.8
|
Pik3r5
|
phosphoinositide-3-kinase regulatory subunit 5 |
| chr6_-_124715560 | 13.56 |
ENSMUST00000004377.15
|
Ptpn6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr9_+_89081262 | 13.24 |
ENSMUST00000068569.5
|
Bcl2a1b
|
B cell leukemia/lymphoma 2 related protein A1b |
| chr17_+_29579882 | 13.10 |
ENSMUST00000024810.8
|
Fgd2
|
FYVE, RhoGEF and PH domain containing 2 |
| chr6_-_124715542 | 13.09 |
ENSMUST00000174265.2
|
Ptpn6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr5_+_86219593 | 13.02 |
ENSMUST00000198435.5
ENSMUST00000031171.9 |
Stap1
|
signal transducing adaptor family member 1 |
| chr1_-_170755136 | 12.79 |
ENSMUST00000046322.14
ENSMUST00000159171.2 |
Fcrla
|
Fc receptor-like A |
| chr6_-_124710084 | 12.76 |
ENSMUST00000112484.10
|
Ptpn6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr7_-_44888465 | 12.75 |
ENSMUST00000210078.2
|
Cd37
|
CD37 antigen |
| chr13_+_24685508 | 12.71 |
ENSMUST00000238974.2
|
Ripor2
|
RHO family interacting cell polarization regulator 2 |
| chr19_+_6135013 | 12.60 |
ENSMUST00000025704.3
|
Cdca5
|
cell division cycle associated 5 |
| chr6_+_41139948 | 12.60 |
ENSMUST00000103275.4
|
Trbv17
|
T cell receptor beta, variable 17 |
| chr17_+_56056977 | 12.57 |
ENSMUST00000025004.7
|
Adgre4
|
adhesion G protein-coupled receptor E4 |
| chr1_+_171668173 | 12.57 |
ENSMUST00000136479.8
|
Cd84
|
CD84 antigen |
| chr2_+_152689881 | 12.47 |
ENSMUST00000164120.8
ENSMUST00000178997.8 ENSMUST00000109816.8 |
Tpx2
|
TPX2, microtubule-associated |
| chr19_+_24976864 | 12.43 |
ENSMUST00000025831.8
|
Dock8
|
dedicator of cytokinesis 8 |
| chr4_-_43454561 | 12.37 |
ENSMUST00000107926.8
ENSMUST00000107925.8 |
Cd72
|
CD72 antigen |
| chr11_-_70703365 | 12.20 |
ENSMUST00000074572.7
ENSMUST00000108534.9 |
Scimp
|
SLP adaptor and CSK interacting membrane protein |
| chr12_-_115611981 | 11.89 |
ENSMUST00000103540.3
ENSMUST00000199266.2 |
Ighv8-12
|
immunoglobulin heavy variable V8-12 |
| chr11_-_82911615 | 11.87 |
ENSMUST00000038141.15
ENSMUST00000092838.11 |
Slfn8
|
schlafen 8 |
| chr16_+_36755338 | 11.80 |
ENSMUST00000023531.15
|
Hcls1
|
hematopoietic cell specific Lyn substrate 1 |
| chr6_+_123206802 | 11.64 |
ENSMUST00000112554.9
ENSMUST00000024118.11 ENSMUST00000117130.8 |
Clec4n
|
C-type lectin domain family 4, member n |
| chrX_-_165992311 | 11.54 |
ENSMUST00000112172.4
|
Tmsb4x
|
thymosin, beta 4, X chromosome |
| chr7_-_126641593 | 11.39 |
ENSMUST00000032915.8
|
Kif22
|
kinesin family member 22 |
| chr19_+_11352862 | 11.20 |
ENSMUST00000188995.2
|
Ms4a4a
|
membrane-spanning 4-domains, subfamily A, member 4A |
| chr11_-_16958647 | 11.20 |
ENSMUST00000102881.10
|
Plek
|
pleckstrin |
| chr19_+_12438125 | 11.10 |
ENSMUST00000081035.9
|
Mpeg1
|
macrophage expressed gene 1 |
| chr6_-_131224305 | 11.04 |
ENSMUST00000032306.15
ENSMUST00000088867.7 |
Klra2
|
killer cell lectin-like receptor, subfamily A, member 2 |
| chr1_+_40123858 | 10.93 |
ENSMUST00000027243.13
|
Il1r2
|
interleukin 1 receptor, type II |
| chr17_+_8144822 | 10.92 |
ENSMUST00000036370.8
|
Tagap
|
T cell activation Rho GTPase activating protein |
| chr2_+_22664094 | 10.86 |
ENSMUST00000014290.15
|
Apbb1ip
|
amyloid beta (A4) precursor protein-binding, family B, member 1 interacting protein |
| chr14_-_56026266 | 10.73 |
ENSMUST00000168716.8
ENSMUST00000178399.3 ENSMUST00000022830.14 |
Ripk3
|
receptor-interacting serine-threonine kinase 3 |
| chr3_-_95799242 | 10.54 |
ENSMUST00000171519.2
ENSMUST00000036360.13 ENSMUST00000090476.10 |
BC028528
|
cDNA sequence BC028528 |
| chr7_+_127661835 | 10.46 |
ENSMUST00000106242.10
ENSMUST00000120355.8 ENSMUST00000106240.9 ENSMUST00000098015.10 |
Itgam
Gm49368
|
integrin alpha M predicted gene, 49368 |
| chr11_-_82911548 | 10.46 |
ENSMUST00000108152.9
|
Slfn8
|
schlafen 8 |
| chr2_+_152689913 | 10.38 |
ENSMUST00000028969.9
|
Tpx2
|
TPX2, microtubule-associated |
| chr19_-_40982576 | 10.34 |
ENSMUST00000117695.8
|
Blnk
|
B cell linker |
| chr1_+_152683627 | 10.33 |
ENSMUST00000027754.7
|
Ncf2
|
neutrophil cytosolic factor 2 |
| chr11_+_87684299 | 10.32 |
ENSMUST00000020779.11
|
Mpo
|
myeloperoxidase |
| chr7_-_104019046 | 10.30 |
ENSMUST00000106831.3
|
Trim30b
|
tripartite motif-containing 30B |
| chr6_-_124710030 | 10.28 |
ENSMUST00000173647.2
|
Ptpn6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr8_+_118225008 | 10.12 |
ENSMUST00000081232.9
|
Plcg2
|
phospholipase C, gamma 2 |
| chr13_-_110493665 | 9.91 |
ENSMUST00000058806.7
ENSMUST00000224534.2 |
Gapt
|
Grb2-binding adaptor, transmembrane |
| chr1_+_87548026 | 9.89 |
ENSMUST00000169754.8
ENSMUST00000042275.15 ENSMUST00000168783.8 |
Inpp5d
|
inositol polyphosphate-5-phosphatase D |
| chrX_-_56371953 | 9.86 |
ENSMUST00000176986.8
|
Arhgef6
|
Rac/Cdc42 guanine nucleotide exchange factor (GEF) 6 |
| chr11_-_82910912 | 9.79 |
ENSMUST00000130822.3
|
Slfn8
|
schlafen 8 |
| chr13_-_97274360 | 9.59 |
ENSMUST00000225410.2
|
Nsa2
|
NSA2 ribosome biogenesis homolog |
| chr16_+_91203123 | 9.54 |
ENSMUST00000023691.12
|
Il10rb
|
interleukin 10 receptor, beta |
| chr13_-_49462694 | 9.37 |
ENSMUST00000110087.9
|
Fgd3
|
FYVE, RhoGEF and PH domain containing 3 |
| chr7_+_127661807 | 9.37 |
ENSMUST00000064821.14
|
Itgam
|
integrin alpha M |
| chr3_+_95566082 | 9.33 |
ENSMUST00000037947.15
ENSMUST00000178686.2 |
Mcl1
|
myeloid cell leukemia sequence 1 |
| chr1_-_20890437 | 9.30 |
ENSMUST00000053266.11
|
Mcm3
|
minichromosome maintenance complex component 3 |
| chr4_-_133601990 | 9.30 |
ENSMUST00000168974.9
|
Rps6ka1
|
ribosomal protein S6 kinase polypeptide 1 |
| chr7_-_103964662 | 9.27 |
ENSMUST00000106837.8
ENSMUST00000106839.9 ENSMUST00000070943.7 |
Trim12a
|
tripartite motif-containing 12A |
| chr15_-_66684442 | 9.14 |
ENSMUST00000100572.10
|
Sla
|
src-like adaptor |
| chr19_+_11469386 | 9.08 |
ENSMUST00000079855.11
|
Gm8369
|
predicted gene 8369 |
| chr1_-_170755109 | 8.88 |
ENSMUST00000162136.2
ENSMUST00000162887.2 |
Fcrla
|
Fc receptor-like A |
| chr10_-_17898838 | 8.81 |
ENSMUST00000220433.2
|
Abracl
|
ABRA C-terminal like |
| chr7_-_4728081 | 8.63 |
ENSMUST00000086363.5
ENSMUST00000086364.11 |
Tmem150b
|
transmembrane protein 150B |
| chr6_-_70364222 | 8.62 |
ENSMUST00000103392.3
ENSMUST00000195945.2 |
Igkv8-16
|
immunoglobulin kappa variable 8-16 |
| chr1_-_144124871 | 8.60 |
ENSMUST00000189061.7
|
Rgs1
|
regulator of G-protein signaling 1 |
| chr7_-_3918484 | 8.58 |
ENSMUST00000038176.15
ENSMUST00000206077.2 ENSMUST00000090689.5 |
Lilra6
|
leukocyte immunoglobulin-like receptor, subfamily A (with TM domain), member 6 |
| chr8_+_85428059 | 8.57 |
ENSMUST00000238364.2
ENSMUST00000238562.2 ENSMUST00000037165.6 |
Lyl1
|
lymphoblastomic leukemia 1 |
| chrX_-_20928219 | 8.56 |
ENSMUST00000040628.6
ENSMUST00000115333.9 ENSMUST00000115334.8 |
Zfp182
|
zinc finger protein 182 |
| chr2_+_163916042 | 8.55 |
ENSMUST00000018353.14
|
Stk4
|
serine/threonine kinase 4 |
| chr3_+_87283767 | 8.54 |
ENSMUST00000194786.6
ENSMUST00000191666.2 |
Fcrl1
|
Fc receptor-like 1 |
| chr8_+_66838927 | 8.53 |
ENSMUST00000039540.12
ENSMUST00000110253.3 |
Marchf1
|
membrane associated ring-CH-type finger 1 |
| chr7_-_103937301 | 8.46 |
ENSMUST00000098179.9
|
Trim5
|
tripartite motif-containing 5 |
| chr5_-_138169509 | 8.40 |
ENSMUST00000153867.8
|
Mcm7
|
minichromosome maintenance complex component 7 |
| chr1_+_153628598 | 8.35 |
ENSMUST00000182538.3
|
Rnasel
|
ribonuclease L (2', 5'-oligoisoadenylate synthetase-dependent) |
| chr6_-_130314465 | 8.33 |
ENSMUST00000088017.5
ENSMUST00000111998.9 |
Klra3
|
killer cell lectin-like receptor, subfamily A, member 3 |
| chr11_-_79418500 | 8.31 |
ENSMUST00000154415.2
|
Evi2a
|
ecotropic viral integration site 2a |
| chr6_+_123206880 | 8.29 |
ENSMUST00000205129.2
|
Clec4n
|
C-type lectin domain family 4, member n |
| chr13_-_97274430 | 8.25 |
ENSMUST00000073456.9
|
Nsa2
|
NSA2 ribosome biogenesis homolog |
| chrX_+_8137372 | 8.21 |
ENSMUST00000127103.8
ENSMUST00000115591.8 |
Slc38a5
|
solute carrier family 38, member 5 |
| chr3_+_87283748 | 8.16 |
ENSMUST00000167200.7
|
Fcrl1
|
Fc receptor-like 1 |
| chr4_+_11758147 | 7.96 |
ENSMUST00000029871.12
ENSMUST00000108303.2 |
Cdh17
|
cadherin 17 |
| chr7_-_126641565 | 7.95 |
ENSMUST00000205806.2
|
Kif22
|
kinesin family member 22 |
| chr1_-_85239791 | 7.90 |
ENSMUST00000162421.2
|
C130026I21Rik
|
RIKEN cDNA C130026I21 gene |
| chr4_-_136620376 | 7.86 |
ENSMUST00000046332.6
|
C1qc
|
complement component 1, q subcomponent, C chain |
| chr7_-_104018989 | 7.84 |
ENSMUST00000164410.2
|
Trim30b
|
tripartite motif-containing 30B |
| chr6_-_56681657 | 7.74 |
ENSMUST00000176595.3
ENSMUST00000170382.5 ENSMUST00000203958.2 |
Lsm5
|
LSM5 homolog, U6 small nuclear RNA and mRNA degradation associated |
| chr15_-_63932176 | 7.72 |
ENSMUST00000226675.2
ENSMUST00000228226.2 ENSMUST00000227024.2 |
Cyrib
|
CYFIP related Rac1 interactor B |
| chr2_+_24235300 | 7.72 |
ENSMUST00000114485.9
ENSMUST00000114482.3 |
Il1rn
|
interleukin 1 receptor antagonist |
| chr7_-_43193842 | 7.72 |
ENSMUST00000039861.7
|
Cd33
|
CD33 antigen |
| chr11_-_106606076 | 7.71 |
ENSMUST00000080853.11
ENSMUST00000183610.8 ENSMUST00000103069.10 ENSMUST00000106796.9 |
Pecam1
|
platelet/endothelial cell adhesion molecule 1 |
| chr10_-_17898938 | 7.69 |
ENSMUST00000220110.2
|
Abracl
|
ABRA C-terminal like |
| chr13_-_67508655 | 7.66 |
ENSMUST00000081582.8
|
Zfp953
|
zinc finger protein 953 |
| chr9_+_89081407 | 7.60 |
ENSMUST00000138109.2
|
Gm29094
|
predicted gene 29094 |
| chr10_-_17898977 | 7.59 |
ENSMUST00000020002.9
|
Abracl
|
ABRA C-terminal like |
| chr14_-_47514308 | 7.58 |
ENSMUST00000111792.9
|
Wdhd1
|
WD repeat and HMG-box DNA binding protein 1 |
| chr3_-_100936859 | 7.38 |
ENSMUST00000147399.9
|
Cd101
|
CD101 antigen |
| chr6_+_29529275 | 7.34 |
ENSMUST00000163511.7
|
Irf5
|
interferon regulatory factor 5 |
| chr1_+_152683568 | 7.28 |
ENSMUST00000190323.7
|
Ncf2
|
neutrophil cytosolic factor 2 |
| chr4_-_86775602 | 7.26 |
ENSMUST00000102814.5
|
Rps6
|
ribosomal protein S6 |
| chr17_-_34174631 | 7.26 |
ENSMUST00000174609.9
ENSMUST00000008812.9 |
Rps18
|
ribosomal protein S18 |
| chr19_+_53517528 | 7.25 |
ENSMUST00000038287.7
|
Dusp5
|
dual specificity phosphatase 5 |
| chr4_-_136626073 | 7.24 |
ENSMUST00000046285.6
|
C1qa
|
complement component 1, q subcomponent, alpha polypeptide |
| chr14_-_47514248 | 7.23 |
ENSMUST00000187531.8
ENSMUST00000111790.2 |
Wdhd1
|
WD repeat and HMG-box DNA binding protein 1 |
| chrX_+_8137620 | 6.98 |
ENSMUST00000033512.11
|
Slc38a5
|
solute carrier family 38, member 5 |
| chr15_+_6599001 | 6.95 |
ENSMUST00000227175.2
|
Fyb
|
FYN binding protein |
| chr6_-_70412460 | 6.84 |
ENSMUST00000103394.2
|
Igkv6-14
|
immunoglobulin kappa variable 6-14 |
| chr15_-_78189917 | 6.84 |
ENSMUST00000096356.5
|
Csf2rb2
|
colony stimulating factor 2 receptor, beta 2, low-affinity (granulocyte-macrophage) |
| chr11_-_16952929 | 6.82 |
ENSMUST00000156101.2
|
Plek
|
pleckstrin |
| chr4_-_43454600 | 6.81 |
ENSMUST00000098105.4
ENSMUST00000098104.10 ENSMUST00000030179.11 |
Cd72
|
CD72 antigen |
| chr5_+_4242367 | 6.78 |
ENSMUST00000177258.2
|
Mterf1b
|
mitochondrial transcription termination factor 1b |
| chr10_+_115653152 | 6.76 |
ENSMUST00000080630.11
ENSMUST00000179196.3 ENSMUST00000035563.15 |
Tspan8
|
tetraspanin 8 |
| chr14_+_53791444 | 6.70 |
ENSMUST00000198297.2
|
Trav14-1
|
T cell receptor alpha variable 14-1 |
| chr6_-_124888643 | 6.65 |
ENSMUST00000032217.2
|
Lag3
|
lymphocyte-activation gene 3 |
| chr17_+_47905553 | 6.64 |
ENSMUST00000182846.3
|
Ccnd3
|
cyclin D3 |
| chr4_+_140428777 | 6.64 |
ENSMUST00000138808.8
ENSMUST00000038893.6 |
Rcc2
|
regulator of chromosome condensation 2 |
| chrX_-_100777806 | 6.52 |
ENSMUST00000056614.7
|
Cxcr3
|
chemokine (C-X-C motif) receptor 3 |
| chr5_-_138169253 | 6.48 |
ENSMUST00000139983.8
|
Mcm7
|
minichromosome maintenance complex component 7 |
| chr3_+_89986831 | 6.46 |
ENSMUST00000029549.16
|
Tpm3
|
tropomyosin 3, gamma |
| chr4_-_136613498 | 6.43 |
ENSMUST00000046384.9
|
C1qb
|
complement component 1, q subcomponent, beta polypeptide |
| chr18_-_35873257 | 6.30 |
ENSMUST00000235302.2
|
Sting1
|
stimulator of interferon response cGAMP interactor 1 |
| chr15_+_102368510 | 6.21 |
ENSMUST00000164688.2
|
Prr13
|
proline rich 13 |
| chr4_-_59549243 | 6.14 |
ENSMUST00000173699.8
ENSMUST00000173884.8 ENSMUST00000102883.11 ENSMUST00000174586.8 |
Ptbp3
|
polypyrimidine tract binding protein 3 |
| chr2_+_118943274 | 6.10 |
ENSMUST00000140939.8
ENSMUST00000028795.10 |
Rad51
|
RAD51 recombinase |
| chr7_+_126184141 | 6.10 |
ENSMUST00000137646.8
|
Apobr
|
apolipoprotein B receptor |
| chr13_-_58276353 | 5.98 |
ENSMUST00000007980.7
|
Hnrnpa0
|
heterogeneous nuclear ribonucleoprotein A0 |
| chr10_-_79938487 | 5.97 |
ENSMUST00000042771.8
|
Sbno2
|
strawberry notch 2 |
| chr8_+_93581946 | 5.92 |
ENSMUST00000046290.3
ENSMUST00000210099.2 |
Lpcat2
|
lysophosphatidylcholine acyltransferase 2 |
| chr9_+_106099797 | 5.81 |
ENSMUST00000062241.11
|
Tlr9
|
toll-like receptor 9 |
| chr14_+_80237691 | 5.76 |
ENSMUST00000228749.2
ENSMUST00000088735.4 |
Olfm4
|
olfactomedin 4 |
| chr10_-_89279751 | 5.74 |
ENSMUST00000219351.2
ENSMUST00000220128.2 ENSMUST00000099374.9 ENSMUST00000105298.3 |
Gas2l3
|
growth arrest-specific 2 like 3 |
| chr19_+_4282040 | 5.66 |
ENSMUST00000237518.2
|
Pold4
|
polymerase (DNA-directed), delta 4 |
| chr1_+_72750418 | 5.64 |
ENSMUST00000059980.11
|
Rpl37a
|
ribosomal protein L37a |
| chr1_-_149836974 | 5.63 |
ENSMUST00000190507.2
ENSMUST00000070200.15 |
Pla2g4a
|
phospholipase A2, group IVA (cytosolic, calcium-dependent) |
| chr13_-_67480588 | 5.60 |
ENSMUST00000109735.9
ENSMUST00000168892.9 |
Zfp595
|
zinc finger protein 595 |
| chr1_-_144124787 | 5.60 |
ENSMUST00000185714.2
|
Rgs1
|
regulator of G-protein signaling 1 |
| chr3_+_88528410 | 5.60 |
ENSMUST00000029694.14
ENSMUST00000176804.8 |
Arhgef2
|
rho/rac guanine nucleotide exchange factor (GEF) 2 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 12.4 | 37.2 | GO:0071846 | actin filament debranching(GO:0071846) |
| 11.4 | 34.1 | GO:0014805 | smooth muscle adaptation(GO:0014805) |
| 10.3 | 92.4 | GO:1901164 | negative regulation of trophoblast cell migration(GO:1901164) |
| 8.5 | 67.7 | GO:0061502 | uropod organization(GO:0032796) early endosome to recycling endosome transport(GO:0061502) |
| 8.0 | 63.9 | GO:0033277 | abortive mitotic cell cycle(GO:0033277) |
| 6.5 | 13.0 | GO:1902226 | regulation of macrophage colony-stimulating factor signaling pathway(GO:1902226) regulation of response to macrophage colony-stimulating factor(GO:1903969) regulation of cellular response to macrophage colony-stimulating factor stimulus(GO:1903972) |
| 6.0 | 18.0 | GO:0010925 | regulation of inositol-polyphosphate 5-phosphatase activity(GO:0010924) positive regulation of inositol-polyphosphate 5-phosphatase activity(GO:0010925) phospholipase C-inhibiting G-protein coupled receptor signaling pathway(GO:0030845) regulation of cell diameter(GO:0060305) |
| 5.5 | 33.1 | GO:0006742 | NADP catabolic process(GO:0006742) |
| 5.5 | 21.9 | GO:0035962 | TRIF-dependent toll-like receptor signaling pathway(GO:0035666) interleukin-4-mediated signaling pathway(GO:0035771) response to interleukin-13(GO:0035962) cellular response to interleukin-13(GO:0035963) negative regulation of apoptotic cell clearance(GO:2000426) |
| 5.5 | 16.4 | GO:0032685 | negative regulation of granulocyte macrophage colony-stimulating factor production(GO:0032685) |
| 5.3 | 15.9 | GO:0045404 | interleukin-10 biosynthetic process(GO:0042091) interleukin-13 biosynthetic process(GO:0042231) regulation of interleukin-10 biosynthetic process(GO:0045074) positive regulation of interleukin-4 biosynthetic process(GO:0045404) |
| 4.9 | 29.7 | GO:0002540 | leukotriene production involved in inflammatory response(GO:0002540) |
| 4.9 | 14.7 | GO:0002337 | B-1a B cell differentiation(GO:0002337) |
| 4.6 | 13.9 | GO:0060559 | positive regulation of vitamin metabolic process(GO:0046136) positive regulation of vitamin D biosynthetic process(GO:0060557) positive regulation of calcidiol 1-monooxygenase activity(GO:0060559) |
| 4.6 | 13.7 | GO:0010387 | COP9 signalosome assembly(GO:0010387) |
| 4.4 | 48.8 | GO:0030886 | negative regulation of myeloid dendritic cell activation(GO:0030886) |
| 4.4 | 26.1 | GO:0002322 | B cell proliferation involved in immune response(GO:0002322) |
| 4.3 | 8.7 | GO:0002649 | regulation of tolerance induction to self antigen(GO:0002649) |
| 4.2 | 12.7 | GO:1903904 | negative regulation of establishment of T cell polarity(GO:1903904) |
| 4.1 | 12.4 | GO:0090673 | endothelial cell-matrix adhesion(GO:0090673) |
| 3.7 | 18.7 | GO:2000660 | negative regulation of interleukin-1-mediated signaling pathway(GO:2000660) |
| 3.7 | 11.2 | GO:0006393 | termination of mitochondrial transcription(GO:0006393) |
| 3.7 | 3.7 | GO:0032701 | negative regulation of interleukin-18 production(GO:0032701) |
| 3.7 | 18.4 | GO:0097029 | mature conventional dendritic cell differentiation(GO:0097029) |
| 3.6 | 10.7 | GO:0002149 | hypochlorous acid metabolic process(GO:0002148) hypochlorous acid biosynthetic process(GO:0002149) |
| 3.5 | 10.6 | GO:0034240 | negative regulation of macrophage fusion(GO:0034240) |
| 3.5 | 14.0 | GO:0070358 | regulation of CD8-positive, alpha-beta T cell differentiation(GO:0043376) actin polymerization-dependent cell motility(GO:0070358) |
| 3.3 | 10.0 | GO:0098501 | polynucleotide dephosphorylation(GO:0098501) |
| 3.0 | 14.8 | GO:0070829 | heterochromatin maintenance(GO:0070829) |
| 2.9 | 5.8 | GO:0032661 | regulation of interleukin-18 production(GO:0032661) |
| 2.8 | 19.8 | GO:0014005 | microglia differentiation(GO:0014004) microglia development(GO:0014005) |
| 2.7 | 8.1 | GO:0070476 | rRNA (guanine-N7)-methylation(GO:0070476) |
| 2.7 | 10.7 | GO:2000449 | regulation of CD8-positive, alpha-beta T cell extravasation(GO:2000449) |
| 2.7 | 8.0 | GO:0002314 | germinal center B cell differentiation(GO:0002314) |
| 2.5 | 10.1 | GO:1904569 | regulation of selenocysteine incorporation(GO:1904569) |
| 2.5 | 12.6 | GO:0071922 | establishment of sister chromatid cohesion(GO:0034085) cohesin loading(GO:0071921) regulation of cohesin loading(GO:0071922) |
| 2.5 | 9.9 | GO:0045659 | negative regulation of neutrophil differentiation(GO:0045659) |
| 2.3 | 6.8 | GO:0033082 | regulation of extrathymic T cell differentiation(GO:0033082) |
| 2.2 | 6.7 | GO:0002434 | immune complex clearance(GO:0002434) |
| 2.2 | 6.6 | GO:0072356 | chromosome passenger complex localization to kinetochore(GO:0072356) |
| 2.1 | 8.5 | GO:0060800 | regulation of cell differentiation involved in embryonic placenta development(GO:0060800) |
| 2.1 | 4.1 | GO:0072717 | cellular response to actinomycin D(GO:0072717) |
| 2.0 | 24.4 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 2.0 | 9.9 | GO:0002309 | T cell proliferation involved in immune response(GO:0002309) |
| 1.9 | 9.3 | GO:1903378 | positive regulation of oxidative stress-induced neuron intrinsic apoptotic signaling pathway(GO:1903378) |
| 1.9 | 5.6 | GO:0034769 | basement membrane disassembly(GO:0034769) |
| 1.8 | 14.6 | GO:0010897 | negative regulation of triglyceride catabolic process(GO:0010897) |
| 1.8 | 10.9 | GO:0070895 | transposon integration(GO:0070893) regulation of transposon integration(GO:0070894) negative regulation of transposon integration(GO:0070895) |
| 1.7 | 5.1 | GO:0009224 | CMP salvage(GO:0006238) CMP biosynthetic process(GO:0009224) pyrimidine ribonucleotide salvage(GO:0010138) pyrimidine nucleotide salvage(GO:0032262) UMP salvage(GO:0044206) CMP metabolic process(GO:0046035) |
| 1.7 | 5.1 | GO:0001788 | antibody-dependent cellular cytotoxicity(GO:0001788) |
| 1.7 | 16.7 | GO:0051025 | negative regulation of immunoglobulin secretion(GO:0051025) |
| 1.6 | 19.8 | GO:0002361 | CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0002361) |
| 1.6 | 10.9 | GO:0002291 | T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:0002291) |
| 1.5 | 4.6 | GO:0014739 | positive regulation of muscle hyperplasia(GO:0014739) |
| 1.5 | 4.5 | GO:0060010 | Sertoli cell fate commitment(GO:0060010) |
| 1.5 | 6.0 | GO:0071348 | cellular response to interleukin-11(GO:0071348) |
| 1.4 | 4.3 | GO:1903722 | regulation of centriole elongation(GO:1903722) |
| 1.4 | 2.9 | GO:0034427 | nuclear-transcribed mRNA catabolic process, exonucleolytic, 3'-5'(GO:0034427) |
| 1.4 | 2.8 | GO:1902623 | negative regulation of neutrophil migration(GO:1902623) |
| 1.3 | 5.3 | GO:2000158 | positive regulation of ubiquitin-specific protease activity(GO:2000158) |
| 1.3 | 15.4 | GO:0048245 | eosinophil chemotaxis(GO:0048245) |
| 1.3 | 15.3 | GO:0002903 | negative regulation of B cell apoptotic process(GO:0002903) |
| 1.3 | 10.1 | GO:0032237 | activation of store-operated calcium channel activity(GO:0032237) |
| 1.2 | 4.9 | GO:0042539 | hypotonic salinity response(GO:0042539) cellular hypotonic salinity response(GO:0071477) |
| 1.2 | 45.1 | GO:0051482 | positive regulation of cytosolic calcium ion concentration involved in phospholipase C-activating G-protein coupled signaling pathway(GO:0051482) |
| 1.2 | 6.0 | GO:0035752 | lysosomal lumen pH elevation(GO:0035752) |
| 1.2 | 6.0 | GO:0038032 | termination of G-protein coupled receptor signaling pathway(GO:0038032) |
| 1.2 | 13.1 | GO:0001955 | blood vessel maturation(GO:0001955) |
| 1.2 | 5.9 | GO:0036151 | phosphatidylcholine acyl-chain remodeling(GO:0036151) |
| 1.2 | 2.4 | GO:0033634 | positive regulation of cell-cell adhesion mediated by integrin(GO:0033634) |
| 1.1 | 3.2 | GO:0071460 | cellular response to cell-matrix adhesion(GO:0071460) |
| 1.0 | 9.3 | GO:2000491 | positive regulation of hepatic stellate cell activation(GO:2000491) |
| 1.0 | 4.0 | GO:0045869 | negative regulation of single stranded viral RNA replication via double stranded DNA intermediate(GO:0045869) |
| 1.0 | 7.9 | GO:0045650 | negative regulation of macrophage differentiation(GO:0045650) |
| 0.9 | 6.6 | GO:0045085 | negative regulation of interleukin-2 biosynthetic process(GO:0045085) |
| 0.9 | 33.2 | GO:0045730 | respiratory burst(GO:0045730) |
| 0.9 | 17.0 | GO:0016998 | cell wall macromolecule catabolic process(GO:0016998) |
| 0.9 | 4.7 | GO:0042631 | cellular response to water deprivation(GO:0042631) |
| 0.9 | 2.7 | GO:2000554 | regulation of T-helper 1 cell cytokine production(GO:2000554) positive regulation of T-helper 1 cell cytokine production(GO:2000556) |
| 0.9 | 2.7 | GO:0035854 | basophil differentiation(GO:0030221) eosinophil fate commitment(GO:0035854) |
| 0.9 | 1.7 | GO:1904139 | microglial cell migration(GO:1904124) regulation of microglial cell migration(GO:1904139) |
| 0.9 | 4.3 | GO:0097309 | cap1 mRNA methylation(GO:0097309) |
| 0.9 | 3.4 | GO:0006269 | DNA replication, synthesis of RNA primer(GO:0006269) |
| 0.8 | 4.2 | GO:0017182 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 0.8 | 15.2 | GO:0015816 | glycine transport(GO:0015816) |
| 0.8 | 11.8 | GO:0045651 | positive regulation of granulocyte differentiation(GO:0030854) positive regulation of macrophage differentiation(GO:0045651) |
| 0.8 | 5.6 | GO:0031394 | positive regulation of prostaglandin biosynthetic process(GO:0031394) positive regulation of unsaturated fatty acid biosynthetic process(GO:2001280) |
| 0.8 | 2.4 | GO:0001180 | transcription initiation from RNA polymerase I promoter for nuclear large rRNA transcript(GO:0001180) |
| 0.8 | 2.4 | GO:0034165 | positive regulation of toll-like receptor 9 signaling pathway(GO:0034165) |
| 0.8 | 16.5 | GO:0030574 | collagen catabolic process(GO:0030574) |
| 0.8 | 12.4 | GO:0001771 | immunological synapse formation(GO:0001771) |
| 0.7 | 8.0 | GO:0031125 | rRNA 3'-end processing(GO:0031125) |
| 0.7 | 2.9 | GO:0090400 | stress-induced premature senescence(GO:0090400) |
| 0.7 | 24.6 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.7 | 5.6 | GO:0071802 | negative regulation of podosome assembly(GO:0071802) |
| 0.7 | 4.9 | GO:0015867 | ATP transport(GO:0015867) |
| 0.7 | 3.5 | GO:0043321 | regulation of natural killer cell degranulation(GO:0043321) negative regulation of CD4-positive, alpha-beta T cell proliferation(GO:2000562) |
| 0.7 | 18.5 | GO:0036342 | post-anal tail morphogenesis(GO:0036342) |
| 0.7 | 11.5 | GO:0051152 | positive regulation of smooth muscle cell differentiation(GO:0051152) |
| 0.7 | 18.9 | GO:0000460 | maturation of 5.8S rRNA(GO:0000460) |
| 0.6 | 19.3 | GO:0007080 | mitotic metaphase plate congression(GO:0007080) |
| 0.6 | 17.9 | GO:1901741 | positive regulation of myoblast fusion(GO:1901741) |
| 0.6 | 2.4 | GO:0061188 | regulation of chromatin silencing at rDNA(GO:0061187) negative regulation of chromatin silencing at rDNA(GO:0061188) |
| 0.6 | 9.1 | GO:0032494 | response to peptidoglycan(GO:0032494) |
| 0.6 | 5.3 | GO:0030321 | transepithelial chloride transport(GO:0030321) |
| 0.6 | 22.4 | GO:0050869 | negative regulation of B cell activation(GO:0050869) |
| 0.6 | 3.5 | GO:1990481 | mRNA pseudouridine synthesis(GO:1990481) |
| 0.6 | 24.9 | GO:0060236 | regulation of mitotic spindle organization(GO:0060236) |
| 0.6 | 19.9 | GO:0050832 | defense response to fungus(GO:0050832) |
| 0.6 | 4.5 | GO:0051533 | positive regulation of NFAT protein import into nucleus(GO:0051533) |
| 0.6 | 3.9 | GO:0031087 | deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
| 0.5 | 6.5 | GO:1900118 | negative regulation of execution phase of apoptosis(GO:1900118) |
| 0.5 | 4.1 | GO:1900264 | regulation of DNA-directed DNA polymerase activity(GO:1900262) positive regulation of DNA-directed DNA polymerase activity(GO:1900264) |
| 0.5 | 2.0 | GO:0002266 | follicular dendritic cell activation(GO:0002266) follicular dendritic cell differentiation(GO:0002268) |
| 0.5 | 7.2 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
| 0.5 | 1.9 | GO:0018103 | protein C-linked glycosylation(GO:0018103) peptidyl-tryptophan modification(GO:0018211) protein C-linked glycosylation via tryptophan(GO:0018317) protein C-linked glycosylation via 2'-alpha-mannosyl-L-tryptophan(GO:0018406) |
| 0.5 | 15.3 | GO:0000731 | DNA synthesis involved in DNA repair(GO:0000731) |
| 0.4 | 17.8 | GO:0045954 | positive regulation of natural killer cell mediated cytotoxicity(GO:0045954) |
| 0.4 | 3.4 | GO:0097428 | protein maturation by iron-sulfur cluster transfer(GO:0097428) |
| 0.4 | 3.0 | GO:1902410 | mitotic cytokinetic process(GO:1902410) |
| 0.4 | 14.7 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 0.4 | 3.7 | GO:0000056 | ribosomal small subunit export from nucleus(GO:0000056) |
| 0.4 | 2.4 | GO:1900157 | regulation of bone mineralization involved in bone maturation(GO:1900157) |
| 0.4 | 2.8 | GO:0051013 | microtubule severing(GO:0051013) |
| 0.4 | 2.7 | GO:2000234 | positive regulation of ribosome biogenesis(GO:0090070) positive regulation of rRNA processing(GO:2000234) |
| 0.4 | 3.5 | GO:0032625 | interleukin-21 production(GO:0032625) interleukin-21 secretion(GO:0072619) |
| 0.4 | 1.1 | GO:0030200 | heparan sulfate proteoglycan catabolic process(GO:0030200) |
| 0.4 | 6.3 | GO:0071578 | zinc II ion transmembrane import(GO:0071578) |
| 0.4 | 0.4 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 0.4 | 1.5 | GO:0021943 | formation of radial glial scaffolds(GO:0021943) |
| 0.4 | 8.4 | GO:2001275 | positive regulation of glucose import in response to insulin stimulus(GO:2001275) |
| 0.4 | 1.4 | GO:1904417 | regulation of xenophagy(GO:1904415) positive regulation of xenophagy(GO:1904417) |
| 0.3 | 2.8 | GO:2000346 | negative regulation of hepatocyte proliferation(GO:2000346) |
| 0.3 | 4.7 | GO:0007016 | cytoskeletal anchoring at plasma membrane(GO:0007016) |
| 0.3 | 1.7 | GO:0039534 | negative regulation of MDA-5 signaling pathway(GO:0039534) |
| 0.3 | 2.0 | GO:1901668 | regulation of superoxide dismutase activity(GO:1901668) |
| 0.3 | 0.3 | GO:0034136 | negative regulation of toll-like receptor 2 signaling pathway(GO:0034136) |
| 0.3 | 4.9 | GO:0060837 | blood vessel endothelial cell differentiation(GO:0060837) |
| 0.3 | 14.0 | GO:0043029 | T cell homeostasis(GO:0043029) |
| 0.3 | 6.0 | GO:0070935 | 3'-UTR-mediated mRNA stabilization(GO:0070935) |
| 0.3 | 4.6 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.3 | 13.2 | GO:0033119 | negative regulation of RNA splicing(GO:0033119) |
| 0.3 | 4.0 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.3 | 6.6 | GO:0030213 | hyaluronan biosynthetic process(GO:0030213) |
| 0.3 | 6.0 | GO:0035493 | SNARE complex assembly(GO:0035493) |
| 0.3 | 13.6 | GO:0043551 | regulation of phosphatidylinositol 3-kinase activity(GO:0043551) |
| 0.3 | 6.6 | GO:0070498 | interleukin-1-mediated signaling pathway(GO:0070498) |
| 0.3 | 3.1 | GO:2000601 | positive regulation of Arp2/3 complex-mediated actin nucleation(GO:2000601) |
| 0.3 | 2.7 | GO:0034244 | negative regulation of transcription elongation from RNA polymerase II promoter(GO:0034244) |
| 0.3 | 4.8 | GO:0090161 | Golgi ribbon formation(GO:0090161) |
| 0.3 | 2.4 | GO:0031145 | anaphase-promoting complex-dependent catabolic process(GO:0031145) |
| 0.3 | 1.3 | GO:0007256 | activation of JNKK activity(GO:0007256) |
| 0.2 | 3.0 | GO:1901096 | regulation of autophagosome maturation(GO:1901096) |
| 0.2 | 1.6 | GO:0046116 | queuosine biosynthetic process(GO:0008616) queuosine metabolic process(GO:0046116) |
| 0.2 | 5.2 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.2 | 8.3 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.2 | 1.2 | GO:0034379 | very-low-density lipoprotein particle assembly(GO:0034379) |
| 0.2 | 3.5 | GO:0001522 | pseudouridine synthesis(GO:0001522) |
| 0.2 | 4.4 | GO:0046597 | negative regulation of viral entry into host cell(GO:0046597) negative regulation of viral release from host cell(GO:1902187) |
| 0.2 | 4.8 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.2 | 4.4 | GO:0006516 | glycoprotein catabolic process(GO:0006516) |
| 0.2 | 11.2 | GO:0043124 | negative regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043124) |
| 0.2 | 1.1 | GO:0046784 | viral mRNA export from host cell nucleus(GO:0046784) |
| 0.2 | 1.4 | GO:0007406 | negative regulation of neuroblast proliferation(GO:0007406) |
| 0.2 | 0.5 | GO:0046038 | GMP catabolic process(GO:0046038) |
| 0.2 | 1.8 | GO:0046606 | negative regulation of centrosome duplication(GO:0010826) negative regulation of centrosome cycle(GO:0046606) |
| 0.2 | 36.1 | GO:1990830 | response to leukemia inhibitory factor(GO:1990823) cellular response to leukemia inhibitory factor(GO:1990830) |
| 0.2 | 5.2 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.2 | 23.8 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.1 | 13.1 | GO:0043507 | positive regulation of JUN kinase activity(GO:0043507) |
| 0.1 | 10.1 | GO:0045576 | mast cell activation(GO:0045576) |
| 0.1 | 1.4 | GO:1900246 | positive regulation of RIG-I signaling pathway(GO:1900246) |
| 0.1 | 3.3 | GO:0043981 | histone H4-K5 acetylation(GO:0043981) histone H4-K8 acetylation(GO:0043982) |
| 0.1 | 24.6 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.1 | 6.8 | GO:1900047 | negative regulation of blood coagulation(GO:0030195) negative regulation of hemostasis(GO:1900047) |
| 0.1 | 20.3 | GO:0008277 | regulation of G-protein coupled receptor protein signaling pathway(GO:0008277) |
| 0.1 | 2.7 | GO:0003283 | atrial septum development(GO:0003283) |
| 0.1 | 3.6 | GO:0030033 | microvillus assembly(GO:0030033) |
| 0.1 | 26.1 | GO:0002244 | hematopoietic progenitor cell differentiation(GO:0002244) |
| 0.1 | 3.9 | GO:0034508 | centromere complex assembly(GO:0034508) |
| 0.1 | 2.8 | GO:0032729 | positive regulation of interferon-gamma production(GO:0032729) |
| 0.1 | 11.1 | GO:0046847 | filopodium assembly(GO:0046847) |
| 0.1 | 2.7 | GO:1902751 | positive regulation of cell cycle G2/M phase transition(GO:1902751) |
| 0.1 | 1.4 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.1 | 0.6 | GO:0071698 | olfactory placode formation(GO:0030910) olfactory placode development(GO:0071698) olfactory placode morphogenesis(GO:0071699) |
| 0.1 | 0.6 | GO:0090521 | glomerular visceral epithelial cell migration(GO:0090521) |
| 0.1 | 1.4 | GO:0035330 | regulation of hippo signaling(GO:0035330) |
| 0.1 | 9.1 | GO:0038083 | peptidyl-tyrosine autophosphorylation(GO:0038083) |
| 0.1 | 18.9 | GO:0051607 | defense response to virus(GO:0051607) |
| 0.1 | 0.7 | GO:0010886 | positive regulation of cholesterol storage(GO:0010886) |
| 0.1 | 0.9 | GO:0031547 | brain-derived neurotrophic factor receptor signaling pathway(GO:0031547) |
| 0.1 | 11.0 | GO:0042113 | B cell activation(GO:0042113) |
| 0.1 | 1.3 | GO:0043649 | dicarboxylic acid catabolic process(GO:0043649) |
| 0.1 | 10.2 | GO:0007173 | epidermal growth factor receptor signaling pathway(GO:0007173) |
| 0.1 | 0.8 | GO:0071763 | nuclear membrane organization(GO:0071763) |
| 0.1 | 4.7 | GO:0002224 | toll-like receptor signaling pathway(GO:0002224) |
| 0.1 | 1.6 | GO:0050961 | detection of temperature stimulus involved in sensory perception(GO:0050961) detection of temperature stimulus involved in sensory perception of pain(GO:0050965) |
| 0.1 | 4.6 | GO:0050830 | defense response to Gram-positive bacterium(GO:0050830) |
| 0.1 | 1.0 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 0.1 | 4.2 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.1 | 3.3 | GO:0046039 | dopamine receptor signaling pathway(GO:0007212) GTP metabolic process(GO:0046039) |
| 0.1 | 7.7 | GO:0000956 | nuclear-transcribed mRNA catabolic process(GO:0000956) |
| 0.1 | 2.5 | GO:0031648 | protein destabilization(GO:0031648) |
| 0.1 | 0.4 | GO:0000469 | cleavage involved in rRNA processing(GO:0000469) |
| 0.1 | 1.6 | GO:0051084 | 'de novo' posttranslational protein folding(GO:0051084) |
| 0.1 | 2.9 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.1 | 2.0 | GO:1903861 | positive regulation of dendrite extension(GO:1903861) |
| 0.0 | 4.5 | GO:0032436 | positive regulation of proteasomal ubiquitin-dependent protein catabolic process(GO:0032436) |
| 0.0 | 9.9 | GO:0046777 | protein autophosphorylation(GO:0046777) |
| 0.0 | 7.1 | GO:0001843 | neural tube closure(GO:0001843) |
| 0.0 | 1.0 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.0 | 12.9 | GO:0006869 | lipid transport(GO:0006869) |
| 0.0 | 6.9 | GO:0019221 | cytokine-mediated signaling pathway(GO:0019221) |
| 0.0 | 3.7 | GO:0070374 | positive regulation of ERK1 and ERK2 cascade(GO:0070374) |
| 0.0 | 3.2 | GO:0032496 | response to lipopolysaccharide(GO:0032496) |
| 0.0 | 9.0 | GO:0006936 | muscle contraction(GO:0006936) |
| 0.0 | 0.3 | GO:0000470 | maturation of LSU-rRNA(GO:0000470) |
| 0.0 | 2.1 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 0.0 | 1.1 | GO:0045761 | regulation of adenylate cyclase activity(GO:0045761) |
| 0.0 | 0.1 | GO:2000298 | regulation of Rho-dependent protein serine/threonine kinase activity(GO:2000298) |
| 0.0 | 0.4 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.0 | 1.0 | GO:0046425 | regulation of JAK-STAT cascade(GO:0046425) regulation of STAT cascade(GO:1904892) |
| 0.0 | 1.8 | GO:0035023 | regulation of Rho protein signal transduction(GO:0035023) |
| 0.0 | 0.4 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 6.3 | 100.4 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 5.6 | 28.2 | GO:0005944 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) |
| 5.3 | 63.9 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 3.2 | 12.7 | GO:0060171 | stereocilium membrane(GO:0060171) |
| 3.1 | 15.3 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 2.6 | 67.7 | GO:0032426 | stereocilium tip(GO:0032426) |
| 1.9 | 13.2 | GO:0048237 | rough endoplasmic reticulum lumen(GO:0048237) |
| 1.8 | 22.8 | GO:0005818 | aster(GO:0005818) |
| 1.7 | 27.6 | GO:0042555 | MCM complex(GO:0042555) |
| 1.6 | 93.3 | GO:0001772 | immunological synapse(GO:0001772) |
| 1.4 | 12.4 | GO:0030485 | smooth muscle contractile fiber(GO:0030485) |
| 1.3 | 18.8 | GO:0031209 | SCAR complex(GO:0031209) |
| 1.1 | 15.9 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 1.1 | 10.7 | GO:0097342 | ripoptosome(GO:0097342) |
| 1.0 | 12.6 | GO:0008278 | cohesin complex(GO:0008278) |
| 1.0 | 5.2 | GO:0017177 | glucosidase II complex(GO:0017177) |
| 1.0 | 10.9 | GO:0042101 | T cell receptor complex(GO:0042101) |
| 1.0 | 5.8 | GO:0036019 | endolysosome(GO:0036019) |
| 1.0 | 2.9 | GO:0032156 | septin cytoskeleton(GO:0032156) |
| 1.0 | 2.9 | GO:0055087 | Ski complex(GO:0055087) |
| 0.9 | 13.7 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.9 | 20.2 | GO:0042627 | chylomicron(GO:0042627) |
| 0.9 | 3.5 | GO:0032021 | NELF complex(GO:0032021) |
| 0.9 | 7.7 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 0.8 | 8.5 | GO:1990462 | omegasome(GO:1990462) |
| 0.7 | 11.0 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 0.7 | 3.6 | GO:0070820 | tertiary granule(GO:0070820) |
| 0.7 | 9.3 | GO:0097136 | Bcl-2 family protein complex(GO:0097136) |
| 0.7 | 4.1 | GO:0005663 | DNA replication factor C complex(GO:0005663) |
| 0.7 | 10.7 | GO:0042582 | primary lysosome(GO:0005766) azurophil granule(GO:0042582) |
| 0.6 | 5.8 | GO:0042581 | specific granule(GO:0042581) |
| 0.5 | 8.8 | GO:0071012 | catalytic step 1 spliceosome(GO:0071012) |
| 0.5 | 21.3 | GO:0051233 | spindle midzone(GO:0051233) |
| 0.5 | 3.4 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.5 | 6.3 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.5 | 15.7 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.5 | 4.6 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
| 0.5 | 11.4 | GO:0046540 | U4/U6 x U5 tri-snRNP complex(GO:0046540) |
| 0.5 | 28.0 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.4 | 17.1 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.4 | 8.3 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.4 | 2.0 | GO:1990130 | Iml1 complex(GO:1990130) |
| 0.4 | 2.0 | GO:0033257 | Bcl3/NF-kappaB2 complex(GO:0033257) |
| 0.4 | 16.7 | GO:0008305 | integrin complex(GO:0008305) |
| 0.4 | 3.7 | GO:0005642 | annulate lamellae(GO:0005642) |
| 0.4 | 4.4 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.4 | 1.4 | GO:0097543 | ciliary inversin compartment(GO:0097543) |
| 0.3 | 4.3 | GO:0090543 | Flemming body(GO:0090543) |
| 0.3 | 3.1 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.3 | 10.1 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.3 | 20.0 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.2 | 28.2 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.2 | 6.9 | GO:0042588 | zymogen granule(GO:0042588) |
| 0.2 | 2.2 | GO:0000815 | ESCRT III complex(GO:0000815) |
| 0.2 | 3.7 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.2 | 21.4 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.2 | 12.0 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.2 | 2.3 | GO:0030897 | HOPS complex(GO:0030897) |
| 0.2 | 0.5 | GO:1902560 | GMP reductase complex(GO:1902560) |
| 0.2 | 2.4 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.2 | 1.0 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.2 | 4.6 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.2 | 2.5 | GO:0036513 | Derlin-1 retrotranslocation complex(GO:0036513) |
| 0.2 | 1.7 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.2 | 14.1 | GO:0001726 | ruffle(GO:0001726) |
| 0.1 | 4.3 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.1 | 12.1 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.1 | 0.4 | GO:0097232 | lamellar body membrane(GO:0097232) alveolar lamellar body membrane(GO:0097233) |
| 0.1 | 4.2 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.1 | 17.3 | GO:0030175 | filopodium(GO:0030175) |
| 0.1 | 6.1 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.1 | 0.6 | GO:0030121 | AP-1 adaptor complex(GO:0030121) |
| 0.1 | 4.6 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.1 | 1.1 | GO:0000347 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.1 | 1.3 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.1 | 12.0 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.1 | 4.6 | GO:0071339 | MLL1/2 complex(GO:0044665) MLL1 complex(GO:0071339) |
| 0.1 | 1.0 | GO:0008282 | ATP-sensitive potassium channel complex(GO:0008282) |
| 0.1 | 5.0 | GO:0031252 | cell leading edge(GO:0031252) |
| 0.1 | 21.3 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.1 | 19.0 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.1 | 4.9 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.1 | 59.7 | GO:0000323 | lytic vacuole(GO:0000323) lysosome(GO:0005764) |
| 0.1 | 49.6 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.1 | 6.4 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.1 | 11.0 | GO:0030863 | cortical cytoskeleton(GO:0030863) |
| 0.1 | 3.9 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.1 | 24.7 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.1 | 33.9 | GO:0009986 | cell surface(GO:0009986) |
| 0.1 | 0.6 | GO:0005850 | eukaryotic translation initiation factor 2 complex(GO:0005850) |
| 0.1 | 21.9 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.1 | 0.7 | GO:0090571 | RNA polymerase II transcription repressor complex(GO:0090571) |
| 0.1 | 0.8 | GO:0097539 | ciliary transition fiber(GO:0097539) |
| 0.1 | 2.8 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.1 | 1.5 | GO:0060170 | ciliary membrane(GO:0060170) |
| 0.1 | 5.6 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.1 | 87.0 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 2.3 | GO:0005776 | autophagosome(GO:0005776) |
| 0.0 | 61.5 | GO:0005887 | integral component of plasma membrane(GO:0005887) |
| 0.0 | 3.6 | GO:0030139 | endocytic vesicle(GO:0030139) |
| 0.0 | 1.9 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.0 | 0.4 | GO:0097450 | astrocyte end-foot(GO:0097450) glial limiting end-foot(GO:0097451) |
| 0.0 | 2.9 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.0 | 1.2 | GO:0005903 | brush border(GO:0005903) |
| 0.0 | 2.6 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 1.0 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.0 | 3.2 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 10.8 | GO:0000151 | ubiquitin ligase complex(GO:0000151) |
| 0.0 | 2.5 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.0 | 12.5 | GO:0005911 | cell-cell junction(GO:0005911) |
| 0.0 | 92.7 | GO:0005886 | plasma membrane(GO:0005886) |
| 0.0 | 3.5 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.0 | 0.3 | GO:0034709 | methylosome(GO:0034709) |
| 0.0 | 1.9 | GO:0000123 | histone acetyltransferase complex(GO:0000123) |
| 0.0 | 0.4 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 53.0 | GO:0005829 | cytosol(GO:0005829) |
| 0.0 | 0.4 | GO:0030686 | 90S preribosome(GO:0030686) |
| 0.0 | 2.1 | GO:0000794 | condensed nuclear chromosome(GO:0000794) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 10.3 | 92.4 | GO:0005094 | Rho GDP-dissociation inhibitor activity(GO:0005094) |
| 5.9 | 29.7 | GO:0004051 | arachidonate 5-lipoxygenase activity(GO:0004051) |
| 5.5 | 21.9 | GO:0005136 | interleukin-4 receptor binding(GO:0005136) |
| 5.1 | 66.3 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 4.9 | 63.9 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 4.3 | 13.0 | GO:0005157 | macrophage colony-stimulating factor receptor binding(GO:0005157) |
| 3.2 | 67.7 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 3.1 | 34.1 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 2.9 | 8.7 | GO:0005026 | transforming growth factor beta receptor activity, type II(GO:0005026) |
| 2.8 | 19.8 | GO:0001851 | complement component C3b binding(GO:0001851) |
| 2.7 | 10.9 | GO:0004910 | interleukin-1, Type II, blocking receptor activity(GO:0004910) |
| 2.7 | 37.2 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 2.5 | 22.8 | GO:0061676 | importin-alpha family protein binding(GO:0061676) |
| 2.5 | 20.2 | GO:0030229 | very-low-density lipoprotein particle receptor activity(GO:0030229) |
| 2.5 | 10.0 | GO:0046403 | polynucleotide 3'-phosphatase activity(GO:0046403) |
| 2.2 | 8.9 | GO:0004566 | beta-glucuronidase activity(GO:0004566) |
| 2.0 | 8.0 | GO:0035673 | oligopeptide transmembrane transporter activity(GO:0035673) |
| 2.0 | 5.9 | GO:0047192 | 1-alkylglycerophosphocholine O-acetyltransferase activity(GO:0047192) |
| 1.9 | 7.7 | GO:0005152 | interleukin-1 receptor antagonist activity(GO:0005152) |
| 1.7 | 11.8 | GO:0019770 | IgG receptor activity(GO:0019770) |
| 1.6 | 4.9 | GO:0046964 | 3'-phosphoadenosine 5'-phosphosulfate transmembrane transporter activity(GO:0046964) |
| 1.5 | 4.6 | GO:0035375 | zymogen binding(GO:0035375) |
| 1.5 | 30.7 | GO:0045028 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 1.5 | 10.7 | GO:0004704 | NF-kappaB-inducing kinase activity(GO:0004704) |
| 1.5 | 37.9 | GO:0051400 | BH domain binding(GO:0051400) |
| 1.4 | 9.8 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 1.4 | 22.2 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 1.4 | 9.5 | GO:0004920 | interleukin-10 receptor activity(GO:0004920) |
| 1.4 | 13.6 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 1.3 | 5.3 | GO:0035800 | deubiquitinase activator activity(GO:0035800) |
| 1.2 | 9.9 | GO:0016312 | inositol bisphosphate phosphatase activity(GO:0016312) PTB domain binding(GO:0051425) |
| 1.2 | 4.9 | GO:0035827 | rubidium ion transmembrane transporter activity(GO:0035827) |
| 1.2 | 14.6 | GO:0046934 | phosphatidylinositol-4,5-bisphosphate 3-kinase activity(GO:0046934) phosphatidylinositol bisphosphate kinase activity(GO:0052813) |
| 1.2 | 28.1 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 1.1 | 3.4 | GO:0003896 | DNA primase activity(GO:0003896) |
| 1.1 | 6.6 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 1.1 | 15.4 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 1.1 | 15.2 | GO:0015187 | glycine transmembrane transporter activity(GO:0015187) |
| 1.0 | 15.6 | GO:0005522 | profilin binding(GO:0005522) |
| 0.9 | 17.0 | GO:0003796 | lysozyme activity(GO:0003796) |
| 0.9 | 6.5 | GO:0019958 | C-X-C chemokine binding(GO:0019958) |
| 0.9 | 2.8 | GO:0001847 | opsonin receptor activity(GO:0001847) |
| 0.9 | 10.6 | GO:0031730 | CCR5 chemokine receptor binding(GO:0031730) |
| 0.8 | 10.1 | GO:0035368 | selenocysteine insertion sequence binding(GO:0035368) |
| 0.8 | 4.5 | GO:0035014 | phosphatidylinositol 3-kinase regulator activity(GO:0035014) |
| 0.7 | 10.3 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.7 | 5.1 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.7 | 4.3 | GO:0004483 | mRNA (nucleoside-2'-O-)-methyltransferase activity(GO:0004483) |
| 0.7 | 6.1 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.7 | 48.6 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.7 | 25.4 | GO:0004623 | phospholipase A2 activity(GO:0004623) |
| 0.6 | 8.3 | GO:0001069 | regulatory region RNA binding(GO:0001069) |
| 0.6 | 5.7 | GO:0016936 | galactoside binding(GO:0016936) |
| 0.6 | 3.7 | GO:0042834 | peptidoglycan binding(GO:0042834) |
| 0.6 | 4.5 | GO:0070290 | N-acylphosphatidylethanolamine-specific phospholipase D activity(GO:0070290) |
| 0.6 | 14.0 | GO:0045125 | bioactive lipid receptor activity(GO:0045125) |
| 0.6 | 2.2 | GO:0005459 | UDP-galactose transmembrane transporter activity(GO:0005459) |
| 0.5 | 5.3 | GO:0000295 | adenine nucleotide transmembrane transporter activity(GO:0000295) purine ribonucleotide transmembrane transporter activity(GO:0005346) ATP transmembrane transporter activity(GO:0005347) purine nucleotide transmembrane transporter activity(GO:0015216) ADP transmembrane transporter activity(GO:0015217) |
| 0.5 | 15.3 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.5 | 2.8 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 0.4 | 7.2 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.4 | 18.3 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.4 | 4.1 | GO:0043142 | single-stranded DNA-dependent ATPase activity(GO:0043142) |
| 0.4 | 18.0 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.4 | 3.3 | GO:0031821 | G-protein coupled serotonin receptor binding(GO:0031821) |
| 0.4 | 1.6 | GO:0043184 | vascular endothelial growth factor receptor 2 binding(GO:0043184) |
| 0.4 | 7.0 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.4 | 8.5 | GO:0008329 | signaling pattern recognition receptor activity(GO:0008329) pattern recognition receptor activity(GO:0038187) |
| 0.3 | 2.4 | GO:0001179 | RNA polymerase I transcription factor binding(GO:0001179) |
| 0.3 | 1.0 | GO:0008281 | sulfonylurea receptor activity(GO:0008281) |
| 0.3 | 20.2 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.3 | 11.2 | GO:0001968 | fibronectin binding(GO:0001968) |
| 0.3 | 4.7 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.3 | 1.2 | GO:0035402 | histone kinase activity (H3-T11 specific)(GO:0035402) |
| 0.3 | 3.7 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.3 | 1.7 | GO:0089720 | caspase binding(GO:0089720) |
| 0.3 | 5.6 | GO:0045309 | protein phosphorylated amino acid binding(GO:0045309) |
| 0.3 | 11.5 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.3 | 16.7 | GO:0070063 | RNA polymerase binding(GO:0070063) |
| 0.3 | 3.3 | GO:0043996 | histone acetyltransferase activity (H4-K5 specific)(GO:0043995) histone acetyltransferase activity (H4-K8 specific)(GO:0043996) histone acetyltransferase activity (H4-K16 specific)(GO:0046972) |
| 0.3 | 1.3 | GO:0030621 | U4 snRNA binding(GO:0030621) |
| 0.2 | 10.9 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.2 | 9.1 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.2 | 16.6 | GO:0016278 | lysine N-methyltransferase activity(GO:0016278) protein-lysine N-methyltransferase activity(GO:0016279) |
| 0.2 | 1.1 | GO:0016435 | rRNA (guanine) methyltransferase activity(GO:0016435) |
| 0.2 | 8.1 | GO:0008276 | protein methyltransferase activity(GO:0008276) |
| 0.2 | 2.4 | GO:0008381 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.2 | 31.4 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.2 | 4.3 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.2 | 2.9 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.2 | 79.8 | GO:0005525 | GTP binding(GO:0005525) |
| 0.2 | 15.7 | GO:0030295 | protein kinase activator activity(GO:0030295) |
| 0.2 | 61.2 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.2 | 7.4 | GO:0048365 | Rac GTPase binding(GO:0048365) |
| 0.2 | 0.5 | GO:0016657 | GMP reductase activity(GO:0003920) oxidoreductase activity, acting on NAD(P)H, nitrogenous group as acceptor(GO:0016657) |
| 0.2 | 9.3 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
| 0.2 | 12.1 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.2 | 33.2 | GO:0005085 | guanyl-nucleotide exchange factor activity(GO:0005085) |
| 0.2 | 6.8 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.2 | 2.7 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
| 0.2 | 2.2 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.1 | 6.0 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.1 | 2.7 | GO:0070742 | DNA binding, bending(GO:0008301) C2H2 zinc finger domain binding(GO:0070742) |
| 0.1 | 6.3 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.1 | 8.0 | GO:0008186 | RNA helicase activity(GO:0003724) ATP-dependent RNA helicase activity(GO:0004004) RNA-dependent ATPase activity(GO:0008186) |
| 0.1 | 13.8 | GO:0004896 | cytokine receptor activity(GO:0004896) |
| 0.1 | 0.4 | GO:0001160 | transcription termination site sequence-specific DNA binding(GO:0001147) transcription termination site DNA binding(GO:0001160) |
| 0.1 | 31.8 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.1 | 6.4 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) |
| 0.1 | 9.3 | GO:0004386 | helicase activity(GO:0004386) |
| 0.1 | 0.7 | GO:0001226 | RNA polymerase II transcription corepressor binding(GO:0001226) |
| 0.1 | 25.8 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.1 | 2.4 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 0.1 | 2.3 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.1 | 31.3 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.1 | 2.4 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.1 | 1.4 | GO:0017151 | DEAD/H-box RNA helicase binding(GO:0017151) |
| 0.1 | 1.0 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 0.1 | 15.8 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.1 | 3.2 | GO:0005035 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.1 | 11.2 | GO:0003690 | double-stranded DNA binding(GO:0003690) |
| 0.1 | 6.7 | GO:0051219 | phosphoprotein binding(GO:0051219) |
| 0.1 | 10.1 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.1 | 1.9 | GO:0000030 | mannosyltransferase activity(GO:0000030) |
| 0.1 | 1.6 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.1 | 17.3 | GO:0005125 | cytokine activity(GO:0005125) |
| 0.1 | 1.4 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.1 | 1.3 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
| 0.1 | 1.1 | GO:0008484 | sulfuric ester hydrolase activity(GO:0008484) |
| 0.1 | 9.0 | GO:0020037 | heme binding(GO:0020037) |
| 0.1 | 3.9 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.1 | 1.8 | GO:0051539 | 4 iron, 4 sulfur cluster binding(GO:0051539) |
| 0.1 | 1.1 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.1 | 0.7 | GO:0030169 | low-density lipoprotein particle binding(GO:0030169) |
| 0.0 | 0.2 | GO:0033142 | progesterone receptor binding(GO:0033142) |
| 0.0 | 8.9 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.0 | 25.0 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 0.0 | 1.0 | GO:0030331 | estrogen receptor binding(GO:0030331) |
| 0.0 | 1.2 | GO:0000062 | fatty-acyl-CoA binding(GO:0000062) |
| 0.0 | 0.3 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.0 | 22.9 | GO:0003682 | chromatin binding(GO:0003682) |
| 0.0 | 6.8 | GO:0000287 | magnesium ion binding(GO:0000287) |
| 0.0 | 18.9 | GO:0004930 | G-protein coupled receptor activity(GO:0004930) |
| 0.0 | 2.0 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.0 | 3.7 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 33.3 | GO:0001071 | nucleic acid binding transcription factor activity(GO:0001071) transcription factor activity, sequence-specific DNA binding(GO:0003700) |
| 0.0 | 7.7 | GO:0046982 | protein heterodimerization activity(GO:0046982) |
| 0.0 | 10.3 | GO:0003677 | DNA binding(GO:0003677) |
| 0.0 | 0.7 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 2.7 | GO:0030414 | peptidase inhibitor activity(GO:0030414) |
| 0.0 | 1.0 | GO:0016765 | transferase activity, transferring alkyl or aryl (other than methyl) groups(GO:0016765) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 26.2 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 1.3 | 94.5 | PID EPO PATHWAY | EPO signaling pathway |
| 1.2 | 36.9 | SA B CELL RECEPTOR COMPLEXES | Antigen binding to B cell receptors activates protein tyrosine kinases, such as the Src family, which ultimate activate MAP kinases. |
| 1.0 | 13.9 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 1.0 | 80.5 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.9 | 89.9 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.9 | 47.4 | PID ATR PATHWAY | ATR signaling pathway |
| 0.8 | 28.2 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.6 | 27.0 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.6 | 27.7 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.6 | 18.5 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.6 | 18.7 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.6 | 25.9 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.5 | 6.5 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 0.4 | 17.1 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.4 | 7.4 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.4 | 20.1 | PID THROMBIN PAR1 PATHWAY | PAR1-mediated thrombin signaling events |
| 0.4 | 20.2 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.3 | 10.7 | PID IL23 PATHWAY | IL23-mediated signaling events |
| 0.3 | 9.3 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.3 | 19.5 | PID PDGFRB PATHWAY | PDGFR-beta signaling pathway |
| 0.3 | 6.6 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.3 | 6.0 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.3 | 11.3 | PID TCR PATHWAY | TCR signaling in naïve CD4+ T cells |
| 0.3 | 15.9 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.3 | 7.3 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.3 | 2.8 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.3 | 18.5 | PID CXCR4 PATHWAY | CXCR4-mediated signaling events |
| 0.3 | 4.9 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.2 | 12.6 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.2 | 4.9 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.2 | 7.9 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.2 | 4.8 | SIG CD40PATHWAYMAP | Genes related to CD40 signaling |
| 0.2 | 34.8 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.1 | 4.3 | PID VEGFR1 2 PATHWAY | Signaling events mediated by VEGFR1 and VEGFR2 |
| 0.1 | 13.4 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.1 | 8.7 | PID E2F PATHWAY | E2F transcription factor network |
| 0.1 | 4.3 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.1 | 0.9 | ST INTERFERON GAMMA PATHWAY | Interferon gamma pathway. |
| 0.1 | 15.5 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.1 | 4.2 | PID P53 REGULATION PATHWAY | p53 pathway |
| 0.1 | 1.7 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 0.7 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.0 | 0.8 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.0 | 1.1 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.0 | 5.1 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.1 | 74.5 | REACTOME REGULATION OF IFNG SIGNALING | Genes involved in Regulation of IFNG signaling |
| 2.7 | 21.5 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 1.9 | 5.6 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 1.7 | 27.6 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 1.7 | 30.7 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 1.5 | 74.0 | REACTOME LATENT INFECTION OF HOMO SAPIENS WITH MYCOBACTERIUM TUBERCULOSIS | Genes involved in Latent infection of Homo sapiens with Mycobacterium tuberculosis |
| 1.4 | 25.4 | REACTOME ACYL CHAIN REMODELLING OF PS | Genes involved in Acyl chain remodelling of PS |
| 1.3 | 22.8 | REACTOME POL SWITCHING | Genes involved in Polymerase switching |
| 1.0 | 15.3 | REACTOME CDC6 ASSOCIATION WITH THE ORC ORIGIN COMPLEX | Genes involved in CDC6 association with the ORC:origin complex |
| 1.0 | 14.2 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION IN TLR7 8 OR 9 SIGNALING | Genes involved in TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling |
| 1.0 | 12.4 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.9 | 28.3 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.9 | 2.7 | REACTOME E2F ENABLED INHIBITION OF PRE REPLICATION COMPLEX FORMATION | Genes involved in E2F-enabled inhibition of pre-replication complex formation |
| 0.9 | 55.6 | REACTOME GPVI MEDIATED ACTIVATION CASCADE | Genes involved in GPVI-mediated activation cascade |
| 0.8 | 15.6 | REACTOME P130CAS LINKAGE TO MAPK SIGNALING FOR INTEGRINS | Genes involved in p130Cas linkage to MAPK signaling for integrins |
| 0.8 | 16.1 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 0.8 | 10.3 | REACTOME REGULATION OF SIGNALING BY CBL | Genes involved in Regulation of signaling by CBL |
| 0.7 | 8.9 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.7 | 36.3 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.7 | 18.5 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.6 | 143.2 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.6 | 12.7 | REACTOME RIP MEDIATED NFKB ACTIVATION VIA DAI | Genes involved in RIP-mediated NFkB activation via DAI |
| 0.6 | 4.5 | REACTOME TRAFFICKING AND PROCESSING OF ENDOSOMAL TLR | Genes involved in Trafficking and processing of endosomal TLR |
| 0.6 | 19.2 | REACTOME SEMA4D IN SEMAPHORIN SIGNALING | Genes involved in Sema4D in semaphorin signaling |
| 0.5 | 31.6 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.5 | 5.9 | REACTOME ACYL CHAIN REMODELLING OF PC | Genes involved in Acyl chain remodelling of PC |
| 0.5 | 11.7 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.5 | 8.7 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.5 | 15.1 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.5 | 8.3 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.4 | 6.1 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.4 | 26.5 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.4 | 21.9 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.4 | 11.7 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.3 | 5.2 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.3 | 6.7 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION | Genes involved in TRAF6 mediated IRF7 activation |
| 0.3 | 4.3 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.3 | 10.1 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.3 | 3.1 | REACTOME DEADENYLATION DEPENDENT MRNA DECAY | Genes involved in Deadenylation-dependent mRNA decay |
| 0.3 | 20.0 | REACTOME 3 UTR MEDIATED TRANSLATIONAL REGULATION | Genes involved in 3' -UTR-mediated translational regulation |
| 0.3 | 9.4 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.3 | 11.0 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.3 | 30.6 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.2 | 6.3 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.2 | 16.7 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.2 | 4.7 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.2 | 22.2 | REACTOME TOLL RECEPTOR CASCADES | Genes involved in Toll Receptor Cascades |
| 0.2 | 3.5 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.2 | 10.1 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.2 | 13.3 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.2 | 38.5 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.2 | 3.7 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.2 | 6.7 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.2 | 6.6 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.1 | 2.4 | REACTOME PHOSPHORYLATION OF THE APC C | Genes involved in Phosphorylation of the APC/C |
| 0.1 | 5.1 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.1 | 11.7 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.1 | 16.8 | REACTOME CLASS A1 RHODOPSIN LIKE RECEPTORS | Genes involved in Class A/1 (Rhodopsin-like receptors) |
| 0.1 | 1.1 | REACTOME FORMATION OF TRANSCRIPTION COUPLED NER TC NER REPAIR COMPLEX | Genes involved in Formation of transcription-coupled NER (TC-NER) repair complex |
| 0.1 | 3.4 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.1 | 1.3 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 7ALPHA HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 7alpha-hydroxycholesterol |
| 0.1 | 4.9 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.0 | 5.6 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.0 | 2.2 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.0 | 1.4 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.0 | 1.7 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 3.0 | REACTOME GLYCEROPHOSPHOLIPID BIOSYNTHESIS | Genes involved in Glycerophospholipid biosynthesis |
| 0.0 | 0.6 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 3 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 3 Promoter |
| 0.0 | 0.5 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.0 | 0.9 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |